RIPTIDE HIGH THROUGHPUT RAPID LIBRARY PREP (HT-RLP)
|
|
|
- Joel Morrison
- 6 years ago
- Views:
Transcription
1 Application Note: RIPTIDE HIGH THROUGHPUT RAPID LIBRARY PREP (HT-RLP) Introduction: Innovations in DNA sequencing during the 21st century have revolutionized our ability to obtain nucleotide information at an ever decreasing cost. Illumina and BGI/Complete Genomics are predicting a human genome will cost as little as $100 within the next ten years, representing a 10-fold increase in sequence output per sequencer run. The technology continues to increase the number of reads per sequencer run and therefore the number of samples that can be multiplexed. However, labs are unable to keep pace with sequencing capacity, and sample preparation, specifically library preperation, is the bottleneck. The average list price of library prep today is $30-$50 (USD) per sample and typically does not include ancillary consumables such as that for DNA fragmentation and purification. The steps involved in the most widely used library prep protocols include: 1. Fragmentation, 2. End-repair, 3. A-tailing, 4. Adapter ligation, 5. Amplification and 6. Size selection. Typically, there are a number of clean-up steps involved and every step is performed on individual samples. An alternative approach using transposase mediated tagmentation combines Key Benefits: High throughput: Each kit produces up to 960 samples per day Simple workflow: 5 pipette steps per sample (no fragmentation, repair or ligation) High quality data: Tunable to diverse gc content Low cost: Contact igenomx for introductory pricing (info@igenomx.com) Table 1: Throughput. Technology Daily throughput (manual)* Daily throughput (automated) steps per sample Fragment/ligate Transposase igenomx <5 * One technician fragmentation and adapter addition in a single step, followed by transposon removal, PCR amplification and size selection. Figure 1: Workflow of Riptide high throughput rapid library prep (HT-RLP)
2 A number of groups have adapted the Nextera technology from Illumina for use in plasmid and bacterial library prep for high throughput sequencing. This approach requires expensive liquid handling equipment, precise quantification of template: enzyme ratios, and results in a loss of chromosome or molecule ends prior to sequencing. While these teams have managed to reduce the reagent cost through volume reduction, the savings have come at the cost of sequence coverage and uniformity, limiting the actual ability for increased sample throughput in multiplexed sequencer runs due to the oversequencing penalty required for completeness and variant identification. igenomx High Throughput Rapid Library Prep (HT-RLP) matches the throughput and cost potential offered by the latest high throughput DNA sequencers. Study Design: Experimental Design 96 technical replicates of E. Coli strain MG6550 were used as the template. 50ng of isolated gdna was used per sample in a 10uL reaction in each well of a 96-well microtiter plate (Table 2). Workflow igenomx mastermix, including primers, NTPs, buffers and polymerase are added to each well. DNA is denatured and igenomx random primers are hybridized to template DNA. Primer construct includes: 5 - Illumina P5 adapter- 8bp sample barcode N12-3. A 30 minute primer extension reaction is performed (no cycling). Extended products are terminated using a small ratio of biotinylatedddntps to native NTPs. ddntp termination controls fragment length, reduces the potential for chimera formation or intramolecular interactions, and minimizes bias during random priming. Aliquots from each well are then pooled without normalization. The product pool is subjected to a bead cleanup to remove small products and unincorporated biotin-ddntps. Products from the 96 sample library pool are then captured on streptavidin coated beads, washed and converted into a dual-adapter library through a second strand displacing primer extension reaction. The primer that is bound closest to the bead displaces upstream products. All displaced product and excess reactants are washed away. PCR is used to amplify material and incorporate full length Illumina adapters. Final products from the 96 sample pool are size selected and ready for flow cell loading and sequencing. The full protocol takes approximately 3 hours with less than 20 minutes of hands-on time for all 96 samples. Multiple 96 sample plates can be processed in serial or parallel for up to 960 samples. There is no chemical or physical fragmentation, no end repair, no A-tailing, and no ligation steps involved. Individual samples do not need normalization prior to pooling and there are no timedependent reactions. An optional index barcode can be used for pooling multiple plates of 96 samples on a single flow cell. 2 x 300 paired end sequencing was performed using the Illumina MiSeq and v3 chemistry. The igenomx BaseSpace QC application was used for analysis. Library construct is 5 - P5 adapter sample barcode insert P7 adapter 3 (Figure 2). Table 2: Design of representative data. Organism Genome size E Coli MG Mb (Haploid) %GC 50% DNA input/replicate 50ng # of replicates 96 Sequencer/chemistry Illumina MiSeq/2x300 Figure 2: Schematic of library construct.
3 Results: Greater than 66% of bases sequenced were at a quality of Q30 or greater. On average, each sample received 483,756 reads with 90% of samples having read counts within a 4-fold range. This was achieved without normalization of individual sample yields (Figure 3). The mean depth of coverage per sample was 19x with a range of 8x to 47x. Median depth was 16x across all 96 samples with a average standard deviation of coverage of 12x. Reference coverage averaged 99.48% for all 96 samples with a range of 98.65% to 99.74%. Summary statistics are found in Table 3. Coverage uniformity is high in that each sample had greater than 90% of bases covered within 1 standard deviation of the mean (Figure 4). The prep contains reagents for both low and mid/ high GC organisms as well as an option for samples with unknown GC content. Optimized GC coverage of multiple organisms can be found below. All samples sequenced to a mean depth of approximately 20x result in greater than 99% genome coverage. Data on sequence coverage (Table 4) and GC performance (Figure 5-11) can be found in the following appendix. Table 3: Mapping and coverage summary statistics. Metric Average across 96 samples Range (low to high for all 96) % mapped 99.18% 98.98%-99.29% Mean coverage 19x 8x-47x Figure 4: Coverage histogram. Random downsampling to 100,000 reads, all 96 samples are plotted Discussion: High throughput multiplexed sequencing is now matched with high throughput multiplex library prep. The features include low sample input, high throughput, automationready workflow with dramatically reduced cost. There is no DNA fragmentation, end repair, A-tailing, ligation and the associated clean-up steps with each of these enzymatic processes. The technology allows for 96 samples processed in three hours and over 1,000 samples per day without the need for expensive capital equipment. The assay is automation friendly. The cost, throughput and performance may justify the conversion of many PCR and Sanger sequencing tests over to high throughput NGS technology. Median coverage 16x 3x-44x Stan. Dev. coverage 12x 4x-21x % at 1x 99.48% 98.65%-99.74% To learn more, or to purchase Riptide, visit igenomx.com/products % at 5x 95.09% 74.93%-99.21% % at 10x 77.73% 27.38%-98.69%
4 Appendix Table 4: Coverage for all samples. Sample/ Barcode % Duplication Mean Median STD Dev % 1x %5x %10x AT GC
5 Table 4: Coverage for all samples (continued). Sample/ Barcode % Duplication Mean Median STD Dev % 1x %5x %10x AT GC
6 Table 4: Coverage for all samples (continued). Sample/ Barcode % Duplication Mean Median STD Dev % 1x %5x %10x AT GC Figure 5: Clostridium difficile (29.1%GC). Figure 6: Staphylococcus aureus (32.7%GC). Figure 7: Helicobacter pylori (39%GC). Figure 8: Escherichia coli (50.8%GC). Figure 9: Klebsiella pneumoniae (57.6%GC). Figure 10: Mycobacterium tuberculosis (65.6%GC). igenomx.com
NextGen Sequencing Technologies Sequencing overview
 Outline Conventional NextGen High-throughput sequencing (Next-Gen sequencing) technologies. Illumina sequencing in detail. Quality control. Sequence coverage. Multiplexing. FASTQ files. Shendure and Ji
Outline Conventional NextGen High-throughput sequencing (Next-Gen sequencing) technologies. Illumina sequencing in detail. Quality control. Sequence coverage. Multiplexing. FASTQ files. Shendure and Ji
Lab methods: Exome / Genome. Ewart de Bruijn
 Lab methods: Exome / Genome 27 06 2013 Ewart de Bruijn Library prep is only a small part of the complete DNA analysis workflow DNA isolation library prep enrichment flowchip prep sequencing bioinformatics
Lab methods: Exome / Genome 27 06 2013 Ewart de Bruijn Library prep is only a small part of the complete DNA analysis workflow DNA isolation library prep enrichment flowchip prep sequencing bioinformatics
HyperCap, an automatable workflow on the Agilent Bravo B
 Automation Note February 2018 HyperCap, an automatable workflow on the Agilent Bravo B 1. OVERVIEW As the demand for next-generation sequencing (NGS) grows, laboratories must adapt to manage increased
Automation Note February 2018 HyperCap, an automatable workflow on the Agilent Bravo B 1. OVERVIEW As the demand for next-generation sequencing (NGS) grows, laboratories must adapt to manage increased
Integrated NGS Sample Preparation Solutions for Limiting Amounts of RNA and DNA. March 2, Steven R. Kain, Ph.D. ABRF 2013
 Integrated NGS Sample Preparation Solutions for Limiting Amounts of RNA and DNA March 2, 2013 Steven R. Kain, Ph.D. ABRF 2013 NuGEN s Core Technologies Selective Sequence Priming Nucleic Acid Amplification
Integrated NGS Sample Preparation Solutions for Limiting Amounts of RNA and DNA March 2, 2013 Steven R. Kain, Ph.D. ABRF 2013 NuGEN s Core Technologies Selective Sequence Priming Nucleic Acid Amplification
High-yield, Scalable Library Preparation with the NEBNext Ultra II FS DNA Library Prep Kit
 be INSPIRED drive DISCOVERY stay GENUINE TECHNICAL NOTE High-yield, Scalable Library Preparation with the NEBNext Ultra II FS DNA Library Prep Kit Improving performance, ease of use and reliability of
be INSPIRED drive DISCOVERY stay GENUINE TECHNICAL NOTE High-yield, Scalable Library Preparation with the NEBNext Ultra II FS DNA Library Prep Kit Improving performance, ease of use and reliability of
Functional Genomics Research Stream. Research Meetings: November 2 & 3, 2009 Next Generation Sequencing
 Functional Genomics Research Stream Research Meetings: November 2 & 3, 2009 Next Generation Sequencing Current Issues Research Meetings: Meet with me this Thursday or Friday. (bring laboratory notebook
Functional Genomics Research Stream Research Meetings: November 2 & 3, 2009 Next Generation Sequencing Current Issues Research Meetings: Meet with me this Thursday or Friday. (bring laboratory notebook
Deep Sequencing technologies
 Deep Sequencing technologies Gabriela Salinas 30 October 2017 Transcriptome and Genome Analysis Laboratory http://www.uni-bc.gwdg.de/index.php?id=709 Microarray and Deep-Sequencing Core Facility University
Deep Sequencing technologies Gabriela Salinas 30 October 2017 Transcriptome and Genome Analysis Laboratory http://www.uni-bc.gwdg.de/index.php?id=709 Microarray and Deep-Sequencing Core Facility University
Next-generation sequencing technologies
 Next-generation sequencing technologies NGS applications Illumina sequencing workflow Overview Sequencing by ligation Short-read NGS Sequencing by synthesis Illumina NGS Single-molecule approach Long-read
Next-generation sequencing technologies NGS applications Illumina sequencing workflow Overview Sequencing by ligation Short-read NGS Sequencing by synthesis Illumina NGS Single-molecule approach Long-read
Incorporating Molecular ID Technology. Accel-NGS 2S MID Indexing Kits
 Incorporating Molecular ID Technology Accel-NGS 2S MID Indexing Kits Molecular Identifiers (MIDs) MIDs are indices used to label unique library molecules MIDs can assess duplicate molecules in sequencing
Incorporating Molecular ID Technology Accel-NGS 2S MID Indexing Kits Molecular Identifiers (MIDs) MIDs are indices used to label unique library molecules MIDs can assess duplicate molecules in sequencing
Cat. Nos. NT09115, NT091120, NT , NT , and NTBC0950 DISCONTINUED
 Nextera DNA Sample Prep Kit (Roche Titanium-compatible) Cat. Nos. NT09115, NT091120, NT0911-50, NT0911-96, and NTBC0950 DISCONTINUED The Nextera DNA Sample Prep Kit is designed to prepare genomic DNA libraries
Nextera DNA Sample Prep Kit (Roche Titanium-compatible) Cat. Nos. NT09115, NT091120, NT0911-50, NT0911-96, and NTBC0950 DISCONTINUED The Nextera DNA Sample Prep Kit is designed to prepare genomic DNA libraries
SureSelect XT HS. Target Enrichment
 SureSelect XT HS Target Enrichment What Is It? SureSelect XT HS joins the SureSelect library preparation reagent family as Agilent s highest sensitivity hybrid capture-based library prep and target enrichment
SureSelect XT HS Target Enrichment What Is It? SureSelect XT HS joins the SureSelect library preparation reagent family as Agilent s highest sensitivity hybrid capture-based library prep and target enrichment
Nextera DNA Sample Prep Kit (Roche FLX-compatible)
 Cat. Nos. FL09115, FL091120, and FLBC0950 The is designed to prepare genomic DNA libraries compatible with the Roche 454 Genome Sequencer FLX System, with standard FLX chemistry. Nextera technology* employs
Cat. Nos. FL09115, FL091120, and FLBC0950 The is designed to prepare genomic DNA libraries compatible with the Roche 454 Genome Sequencer FLX System, with standard FLX chemistry. Nextera technology* employs
Effective Miniaturization of Illumina Nextera XT Library Prep for Multiplexed Whole Genome Sequencing and Microbiome Applications
 APPLICATION NOTE Effective Miniaturization of Illumina Nextera XT Library Prep for Multiplexed Whole Genome Sequencing and Microbiome Applications Jefferson Lai, John Lesnick, Noël Ruppert Labcyte Inc.
APPLICATION NOTE Effective Miniaturization of Illumina Nextera XT Library Prep for Multiplexed Whole Genome Sequencing and Microbiome Applications Jefferson Lai, John Lesnick, Noël Ruppert Labcyte Inc.
Nextera DNA Sample Prep Kit
 Nextera DNA Sample Prep Kit (Roche FLX-compatible) Cat. Nos. FL09115, FL091120, and FLBC0950 * Covered by patents issued and pending. Connect with Epicentre on our blog (epicentral.blogspot.com), Facebook
Nextera DNA Sample Prep Kit (Roche FLX-compatible) Cat. Nos. FL09115, FL091120, and FLBC0950 * Covered by patents issued and pending. Connect with Epicentre on our blog (epicentral.blogspot.com), Facebook
Biotool DNA library prep kit V2 for Illumina
 Biotool DNA library prep kit V2 for Illumina Description Biotool DNA library prep kit V2 for Illumina is developed specially for the Illumina high-throughput sequencing platform, and generates sequencing-ready
Biotool DNA library prep kit V2 for Illumina Description Biotool DNA library prep kit V2 for Illumina is developed specially for the Illumina high-throughput sequencing platform, and generates sequencing-ready
ACCEL-NGS 2S DNA LIBRARY KITS
 ACCEL-NGS 2S DNA LIBRARY KITS Accel-NGS 2S DNA Library Kits produce high quality libraries with an all-inclusive, easy-to-use format. The kits contain all reagents necessary to build high complexity libraries
ACCEL-NGS 2S DNA LIBRARY KITS Accel-NGS 2S DNA Library Kits produce high quality libraries with an all-inclusive, easy-to-use format. The kits contain all reagents necessary to build high complexity libraries
Increased transcription detection with the NEBNext Single Cell/Low Input RNA Library Prep Kit
 be INSPIRED drive DISCOVERY stay GENUINE TECHNICAL NOTE Increased transcription detection with the NEBNext Single Cell/Low Input RNA Library Prep Kit Highly sensitive, robust generation of high quality
be INSPIRED drive DISCOVERY stay GENUINE TECHNICAL NOTE Increased transcription detection with the NEBNext Single Cell/Low Input RNA Library Prep Kit Highly sensitive, robust generation of high quality
Illumina s Suite of Targeted Resequencing Solutions
 Illumina s Suite of Targeted Resequencing Solutions Colin Baron Sr. Product Manager Sequencing Applications 2011 Illumina, Inc. All rights reserved. Illumina, illuminadx, Solexa, Making Sense Out of Life,
Illumina s Suite of Targeted Resequencing Solutions Colin Baron Sr. Product Manager Sequencing Applications 2011 Illumina, Inc. All rights reserved. Illumina, illuminadx, Solexa, Making Sense Out of Life,
Increase Sequencing Efficiency with the SeqCap EZ Prime Exome
 Sequencing Solutions Technical Note June 2018 How To Increase Sequencing Efficiency with the SeqCap EZ Prime Exome Applications Whole Exome Sequencing Products SeqCap EZ Prime Exome SeqCap EZ HyperCap
Sequencing Solutions Technical Note June 2018 How To Increase Sequencing Efficiency with the SeqCap EZ Prime Exome Applications Whole Exome Sequencing Products SeqCap EZ Prime Exome SeqCap EZ HyperCap
Introduction Bioo Scientific
 Next Generation Sequencing Catalog 2014-2015 Introduction Bioo Scientific Bioo Scientific is a global life science company headquartered in Austin, TX, committed to providing innovative products and superior
Next Generation Sequencing Catalog 2014-2015 Introduction Bioo Scientific Bioo Scientific is a global life science company headquartered in Austin, TX, committed to providing innovative products and superior
Fully Automated Library Quantification for Illumina Sequencing on the NGS STAR
 Fully Automated Library Quantification for Illumina Sequencing on the NGS STAR Introduction Hamilton Robotics, an industry leader in liquid handling and laboratory automation equipment, has partnered with
Fully Automated Library Quantification for Illumina Sequencing on the NGS STAR Introduction Hamilton Robotics, an industry leader in liquid handling and laboratory automation equipment, has partnered with
NEXT GENERATION SEQUENCING. Innovative solutions for library
 NEXT GENERATION SEQUENCING Innovative solutions for library pr NEXT GENERATION SOLUTIONS Dramatic improvements to commercial next generation sequencing (NGS) platforms have resulted in spectacular reductions
NEXT GENERATION SEQUENCING Innovative solutions for library pr NEXT GENERATION SOLUTIONS Dramatic improvements to commercial next generation sequencing (NGS) platforms have resulted in spectacular reductions
SURESELECTXT LOW INPUT TARGET ENRICHMENT
 SURESELECTXT LOW INPUT TARGET ENRICHMENT Low Input FFPE Optimized Streamlined Workflow SureSelect XT Low Input What is it? SureSelect XT Low Input is a low-input, FFPE-optimized library preparation kit.
SURESELECTXT LOW INPUT TARGET ENRICHMENT Low Input FFPE Optimized Streamlined Workflow SureSelect XT Low Input What is it? SureSelect XT Low Input is a low-input, FFPE-optimized library preparation kit.
Complete protocol in 110 minutes Enzymatic fragmentation without sonication One-step fragmentation/tagging to save time
 Molecular Cloning Laboratories Manual Version 1.2 Product name: MCNext UT DNA Sample Prep Kit Cat #: MCUDS-4, MCUDS-24, MCUDS-96 Description: This protocol explains how to prepare up to 96 pooled indexed
Molecular Cloning Laboratories Manual Version 1.2 Product name: MCNext UT DNA Sample Prep Kit Cat #: MCUDS-4, MCUDS-24, MCUDS-96 Description: This protocol explains how to prepare up to 96 pooled indexed
Next Generation Sequencing. Jeroen Van Houdt - Leuven 13/10/2017
 Next Generation Sequencing Jeroen Van Houdt - Leuven 13/10/2017 Landmarks in DNA sequencing 1953 Discovery of DNA double helix structure 1977 A Maxam and W Gilbert "DNA seq by chemical degradation" F Sanger"DNA
Next Generation Sequencing Jeroen Van Houdt - Leuven 13/10/2017 Landmarks in DNA sequencing 1953 Discovery of DNA double helix structure 1977 A Maxam and W Gilbert "DNA seq by chemical degradation" F Sanger"DNA
Illumina Sequencing Overview
 Illumina Sequencing Overview Part # 15045845_Rev.C 2013 Illumina, Inc. All rights reserved. Illumina, IlluminaDx, BaseSpace, BeadArray, BeadXpress, cbot, CSPro, DASL, DesignStudio, Eco, GAIIx, Genetic
Illumina Sequencing Overview Part # 15045845_Rev.C 2013 Illumina, Inc. All rights reserved. Illumina, IlluminaDx, BaseSpace, BeadArray, BeadXpress, cbot, CSPro, DASL, DesignStudio, Eco, GAIIx, Genetic
MHC Region. MHC expression: Class I: All nucleated cells and platelets Class II: Antigen presenting cells
 DNA based HLA typing methods By: Yadollah Shakiba, MD, PhD MHC Region MHC expression: Class I: All nucleated cells and platelets Class II: Antigen presenting cells Nomenclature of HLA Alleles Assigned
DNA based HLA typing methods By: Yadollah Shakiba, MD, PhD MHC Region MHC expression: Class I: All nucleated cells and platelets Class II: Antigen presenting cells Nomenclature of HLA Alleles Assigned
Get to Know Your DNA. Every Single Fragment.
 HaloPlex HS NGS Target Enrichment System Get to Know Your DNA. Every Single Fragment. High sensitivity detection of rare variants using molecular barcodes How Does Molecular Barcoding Work? HaloPlex HS
HaloPlex HS NGS Target Enrichment System Get to Know Your DNA. Every Single Fragment. High sensitivity detection of rare variants using molecular barcodes How Does Molecular Barcoding Work? HaloPlex HS
Targeted Sequencing Using Droplet-Based Microfluidics. Keith Brown Director, Sales
 Targeted Sequencing Using Droplet-Based Microfluidics Keith Brown Director, Sales brownk@raindancetech.com Who we are: is a Provider of Microdroplet-based Solutions The Company s RainStorm TM Technology
Targeted Sequencing Using Droplet-Based Microfluidics Keith Brown Director, Sales brownk@raindancetech.com Who we are: is a Provider of Microdroplet-based Solutions The Company s RainStorm TM Technology
High Throughput Sequencing Technologies. UCD Genome Center Bioinformatics Core Monday 15 June 2015
 High Throughput Sequencing Technologies UCD Genome Center Bioinformatics Core Monday 15 June 2015 Sequencing Explosion www.genome.gov/sequencingcosts http://t.co/ka5cvghdqo Sequencing Explosion 2011 PacBio
High Throughput Sequencing Technologies UCD Genome Center Bioinformatics Core Monday 15 June 2015 Sequencing Explosion www.genome.gov/sequencingcosts http://t.co/ka5cvghdqo Sequencing Explosion 2011 PacBio
dbcamplicons pipeline Amplicons
 dbcamplicons pipeline Amplicons Matthew L. Settles Genome Center Bioinformatics Core University of California, Davis settles@ucdavis.edu; bioinformatics.core@ucdavis.edu Microbial community analysis Goal:
dbcamplicons pipeline Amplicons Matthew L. Settles Genome Center Bioinformatics Core University of California, Davis settles@ucdavis.edu; bioinformatics.core@ucdavis.edu Microbial community analysis Goal:
Nature Methods Optimal enzymes for amplifying sequencing libraries
 Nature Methods Optimal enzymes for amplifying sequencing libraries Michael A Quail, Thomas D Otto, Yong Gu, Simon R Harris, Thomas F Skelly, Jacqueline A McQuillan, Harold P Swerdlow & Samuel O Oyola Supplementary
Nature Methods Optimal enzymes for amplifying sequencing libraries Michael A Quail, Thomas D Otto, Yong Gu, Simon R Harris, Thomas F Skelly, Jacqueline A McQuillan, Harold P Swerdlow & Samuel O Oyola Supplementary
Improved Chemistry for NGS Library Cleanup and Size Selection Speakers: Charles Cowles, PhD & Curtis Knox
 Improved Chemistry for NGS Library Cleanup and Size Selection Speakers: Charles Cowles, PhD & Curtis Knox Promega Corporation Agenda What is size-selective purification and how is it used? Why is there
Improved Chemistry for NGS Library Cleanup and Size Selection Speakers: Charles Cowles, PhD & Curtis Knox Promega Corporation Agenda What is size-selective purification and how is it used? Why is there
dbcamplicons pipeline Amplicons
 dbcamplicons pipeline Amplicons Matthew L. Settles Genome Center Bioinformatics Core University of California, Davis settles@ucdavis.edu; bioinformatics.core@ucdavis.edu Microbial community analysis Goal:
dbcamplicons pipeline Amplicons Matthew L. Settles Genome Center Bioinformatics Core University of California, Davis settles@ucdavis.edu; bioinformatics.core@ucdavis.edu Microbial community analysis Goal:
Comparison of ExoSAP-IT and ExoSAP-IT Express reagents to alternative PCR cleanup methods
 WHITE PAPER ExoSAP-IT PCR cleanup reagents Comparison of ExoSAP-IT and ExoSAP-IT Express reagents to alternative PCR cleanup methods Abstract Here we present superior workflow advantages of enzymatic PCR
WHITE PAPER ExoSAP-IT PCR cleanup reagents Comparison of ExoSAP-IT and ExoSAP-IT Express reagents to alternative PCR cleanup methods Abstract Here we present superior workflow advantages of enzymatic PCR
NEBNext Direct Custom Ready Panels
 LIBRARY PREPARATION NEBNext Direct Custom Ready Panels Instruction Manual NEB #E6631S/L/X 8/24/96 reactions Version 1.0 4/18 be INSPIRED drive DISCOVERY stay GENUINE i This product is intended for research
LIBRARY PREPARATION NEBNext Direct Custom Ready Panels Instruction Manual NEB #E6631S/L/X 8/24/96 reactions Version 1.0 4/18 be INSPIRED drive DISCOVERY stay GENUINE i This product is intended for research
solid S Y S T E M s e q u e n c i n g See the Difference Discover the Quality Genome
 solid S Y S T E M s e q u e n c i n g See the Difference Discover the Quality Genome See the Difference With a commitment to your peace of mind, Life Technologies provides a portfolio of robust and scalable
solid S Y S T E M s e q u e n c i n g See the Difference Discover the Quality Genome See the Difference With a commitment to your peace of mind, Life Technologies provides a portfolio of robust and scalable
HALOPLEX PCR TARGET ENRICHMENT & LIBRARY PREPARATION PROTOCOL. Version 1.0.3, April 2011 For research use only
 HALOPLEX PCR TARGET ENRICHMENT & LIBRARY PREPARATION PROTOCOL Version 1.0.3, April 2011 For research use only HALOPLEX PCR TARGET ENRICHMENT & LIBRARY PREPARATION PROTOCOL Halo Genomics AB, 2011 No part
HALOPLEX PCR TARGET ENRICHMENT & LIBRARY PREPARATION PROTOCOL Version 1.0.3, April 2011 For research use only HALOPLEX PCR TARGET ENRICHMENT & LIBRARY PREPARATION PROTOCOL Halo Genomics AB, 2011 No part
Unbiased Quantitative RNA & DNA Specialty Sequencing Solutions
 Unbiased Quantitative RNA & DNA Specialty Sequencing Solutions The Heliscope single molecule sequencer is the first genetic analyzer to harness the power of single molecule sequencing. True single molecule
Unbiased Quantitative RNA & DNA Specialty Sequencing Solutions The Heliscope single molecule sequencer is the first genetic analyzer to harness the power of single molecule sequencing. True single molecule
High Throughput Sequencing Technologies. J Fass UCD Genome Center Bioinformatics Core Monday June 16, 2014
 High Throughput Sequencing Technologies J Fass UCD Genome Center Bioinformatics Core Monday June 16, 2014 Sequencing Explosion www.genome.gov/sequencingcosts http://t.co/ka5cvghdqo Sequencing Explosion
High Throughput Sequencing Technologies J Fass UCD Genome Center Bioinformatics Core Monday June 16, 2014 Sequencing Explosion www.genome.gov/sequencingcosts http://t.co/ka5cvghdqo Sequencing Explosion
SANGER SEQUENCING WHITE PAPER
 AAAT T C SANGER SEQUENCING WHITE PAPER FACTORS AFFECTING SANGER SEQUENCING ROBUSTNESS AND DATA QUALITY PATRICK WARNER, SHEA ANDERSON, ARCHANA DESHPANDE, DARYL M. GOHL, KENNETH B. BECKMAN UNIVERSITY OF
AAAT T C SANGER SEQUENCING WHITE PAPER FACTORS AFFECTING SANGER SEQUENCING ROBUSTNESS AND DATA QUALITY PATRICK WARNER, SHEA ANDERSON, ARCHANA DESHPANDE, DARYL M. GOHL, KENNETH B. BECKMAN UNIVERSITY OF
Whole-genome sequencing (WGS) of microbes employing nextgeneration sequencing (NGS) technologies enables pathogen
 Application Note Microbial whole-genome sequencing A novel, single-tube enzymatic fragmentation and library construction method enables fast turnaround times and improved data quality for microbial whole-genome
Application Note Microbial whole-genome sequencing A novel, single-tube enzymatic fragmentation and library construction method enables fast turnaround times and improved data quality for microbial whole-genome
Novel methods for RNA and DNA- Seq analysis using SMART Technology. Andrew Farmer, D. Phil. Vice President, R&D Clontech Laboratories, Inc.
 Novel methods for RNA and DNA- Seq analysis using SMART Technology Andrew Farmer, D. Phil. Vice President, R&D Clontech Laboratories, Inc. Agenda Enabling Single Cell RNA-Seq using SMART Technology SMART
Novel methods for RNA and DNA- Seq analysis using SMART Technology Andrew Farmer, D. Phil. Vice President, R&D Clontech Laboratories, Inc. Agenda Enabling Single Cell RNA-Seq using SMART Technology SMART
NGS Library Construction Kit User Guide
 NGS Library Construction Kit User Guide Catalog Number BX2000-08M REV. 1.0 11.21.16 Part number 40023 LEGAL NOTICES Technical Services Limited Use Label License Limited Warranty Trademark Information Regulatory
NGS Library Construction Kit User Guide Catalog Number BX2000-08M REV. 1.0 11.21.16 Part number 40023 LEGAL NOTICES Technical Services Limited Use Label License Limited Warranty Trademark Information Regulatory
Overview of Next Generation Sequencing technologies. Céline Keime
 Overview of Next Generation Sequencing technologies Céline Keime keime@igbmc.fr Next Generation Sequencing < Second generation sequencing < General principle < Sequencing by synthesis - Illumina < Sequencing
Overview of Next Generation Sequencing technologies Céline Keime keime@igbmc.fr Next Generation Sequencing < Second generation sequencing < General principle < Sequencing by synthesis - Illumina < Sequencing
APPLICATION NOTE
 APPLICATION NOTE www.swiftbiosci.com Approaching Single-Cell Sequencing by Understanding NGS Library Complexity and Bias Abstract Demands are growing on genomics to deliver higher quality sequencing data
APPLICATION NOTE www.swiftbiosci.com Approaching Single-Cell Sequencing by Understanding NGS Library Complexity and Bias Abstract Demands are growing on genomics to deliver higher quality sequencing data
EPIGENTEK. EpiNext DNA Library Preparation Kit (Illumina) Base Catalog # P-1051 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE
 EpiNext DNA Library Preparation Kit (Illumina) Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE Uses: The EpiNext DNA Library Preparation Kit (Illumina) is suitable for preparing a DNA library
EpiNext DNA Library Preparation Kit (Illumina) Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE Uses: The EpiNext DNA Library Preparation Kit (Illumina) is suitable for preparing a DNA library
DNA concentration and purity were initially measured by NanoDrop 2000 and verified on Qubit 2.0 Fluorometer.
 DNA Preparation and QC Extraction DNA was extracted from whole blood or flash frozen post-mortem tissue using a DNA mini kit (QIAmp #51104 and QIAmp#51404, respectively) following the manufacturer s recommendations.
DNA Preparation and QC Extraction DNA was extracted from whole blood or flash frozen post-mortem tissue using a DNA mini kit (QIAmp #51104 and QIAmp#51404, respectively) following the manufacturer s recommendations.
CM581A2: NEXT GENERATION SEQUENCING PLATFORMS AND LIBRARY GENERATION
 CM581A2: NEXT GENERATION SEQUENCING PLATFORMS AND LIBRARY GENERATION Fall 2015 Instructors: Coordinator: Carol Wilusz, Associate Professor MIP, CMB Instructor: Dan Sloan, Assistant Professor, Biology,
CM581A2: NEXT GENERATION SEQUENCING PLATFORMS AND LIBRARY GENERATION Fall 2015 Instructors: Coordinator: Carol Wilusz, Associate Professor MIP, CMB Instructor: Dan Sloan, Assistant Professor, Biology,
High Throughput Sequencing Technologies. J Fass UCD Genome Center Bioinformatics Core Monday September 15, 2014
 High Throughput Sequencing Technologies J Fass UCD Genome Center Bioinformatics Core Monday September 15, 2014 Sequencing Explosion www.genome.gov/sequencingcosts http://t.co/ka5cvghdqo Sequencing Explosion
High Throughput Sequencing Technologies J Fass UCD Genome Center Bioinformatics Core Monday September 15, 2014 Sequencing Explosion www.genome.gov/sequencingcosts http://t.co/ka5cvghdqo Sequencing Explosion
HaloPlex HS. Get to Know Your DNA. Every Single Fragment. Kevin Poon, Ph.D.
 HaloPlex HS Get to Know Your DNA. Every Single Fragment. Kevin Poon, Ph.D. Sr. Global Product Manager Diagnostics & Genomics Group Agilent Technologies For Research Use Only. Not for Use in Diagnostic
HaloPlex HS Get to Know Your DNA. Every Single Fragment. Kevin Poon, Ph.D. Sr. Global Product Manager Diagnostics & Genomics Group Agilent Technologies For Research Use Only. Not for Use in Diagnostic
NEXTFLEX ChIP-Seq Kit (For Illumina Platforms) Catalog #NOVA (Kit contains 8 reactions) Bioo Scientific Corp V15.
 NEXTFLEX ChIP-Seq Kit (For Illumina Platforms) Catalog #NOVA-5143-01 (Kit contains 8 reactions) Bioo Scientific Corp. 2015-2018 V15.07 This product is for research use only. Not for use in diagnostic procedures.
NEXTFLEX ChIP-Seq Kit (For Illumina Platforms) Catalog #NOVA-5143-01 (Kit contains 8 reactions) Bioo Scientific Corp. 2015-2018 V15.07 This product is for research use only. Not for use in diagnostic procedures.
Absolute Human Telomere Length Quantification qpcr Assay Kit (AHTLQ) Catalog # reactions
 Absolute Human Telomere Length Quantification qpcr Assay Kit (AHTLQ) Catalog #8918 100 reactions Product Description Telomeres are repetitive nucleotide elements at the ends of chromosomes that protect
Absolute Human Telomere Length Quantification qpcr Assay Kit (AHTLQ) Catalog #8918 100 reactions Product Description Telomeres are repetitive nucleotide elements at the ends of chromosomes that protect
Sample Quality Control in Agilent NGS Solutions
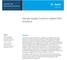 pplication Note Next Generation Sequencing Sample Quality Control in gilent NGS Solutions uthors Madhurima iswas and Shweta Sharma gilent Technologies, Inc. La Jolla, C US Weiwei Liu, Tracy Liu, and David
pplication Note Next Generation Sequencing Sample Quality Control in gilent NGS Solutions uthors Madhurima iswas and Shweta Sharma gilent Technologies, Inc. La Jolla, C US Weiwei Liu, Tracy Liu, and David
TECH NOTE Ligation-Free ChIP-Seq Library Preparation
 TECH NOTE Ligation-Free ChIP-Seq Library Preparation The DNA SMART ChIP-Seq Kit Ligation-free template switching technology: Minimize sample handling in a single-tube workflow >> Simplified protocol with
TECH NOTE Ligation-Free ChIP-Seq Library Preparation The DNA SMART ChIP-Seq Kit Ligation-free template switching technology: Minimize sample handling in a single-tube workflow >> Simplified protocol with
Performance Characteristics drmid Dx for Illumina NGS systems
 Performance Characteristics drmid Dx for Illumina NGS systems MANUFACTURER Multiplicom N.V. Galileïlaan 18 2845 Niel BELGIUM Revision date: August, 2017 Page 1 of 7 TABLE OF CONTENTS 1. TEST PRINCIPLE...
Performance Characteristics drmid Dx for Illumina NGS systems MANUFACTURER Multiplicom N.V. Galileïlaan 18 2845 Niel BELGIUM Revision date: August, 2017 Page 1 of 7 TABLE OF CONTENTS 1. TEST PRINCIPLE...
Supplementary Figure 1
 Nucleotide Content E. coli End. Neb. Tr. End. Neb. Tr. Supplementary Figure 1 Fragmentation Site Profiles CRW1 End. Son. Tr. End. Son. Tr. Human PA1 Position Fragmentation site profiles. Nucleotide content
Nucleotide Content E. coli End. Neb. Tr. End. Neb. Tr. Supplementary Figure 1 Fragmentation Site Profiles CRW1 End. Son. Tr. End. Son. Tr. Human PA1 Position Fragmentation site profiles. Nucleotide content
Factors affecting PCR
 Lec. 11 Dr. Ahmed K. Ali Factors affecting PCR The sequences of the primers are critical to the success of the experiment, as are the precise temperatures used in the heating and cooling stages of the
Lec. 11 Dr. Ahmed K. Ali Factors affecting PCR The sequences of the primers are critical to the success of the experiment, as are the precise temperatures used in the heating and cooling stages of the
Supplementary Information for:
 Supplementary Information for: A streamlined and high-throughput targeting approach for human germline and cancer genomes using Oligonucleotide-Selective Sequencing Samuel Myllykangas 1, Jason D. Buenrostro
Supplementary Information for: A streamlined and high-throughput targeting approach for human germline and cancer genomes using Oligonucleotide-Selective Sequencing Samuel Myllykangas 1, Jason D. Buenrostro
BIOO LIFE SCIENCE PRODUCTS. NEXTflex TM 16S V4 Amplicon-Seq Kit 4 (Illumina Compatible) BIOO Scientific Corp V13.01
 BIOO LIFE SCIENCE PRODUCTS NEXTflex TM 16S V4 Amplicon-Seq Kit 4 (Illumina Compatible) Catalog #: 4201-01 (16 reactions) BIOO Scientific Corp. 2013 V13.01 TABLE OF CONTENTS GENERAL INFORMATION... 1 Product
BIOO LIFE SCIENCE PRODUCTS NEXTflex TM 16S V4 Amplicon-Seq Kit 4 (Illumina Compatible) Catalog #: 4201-01 (16 reactions) BIOO Scientific Corp. 2013 V13.01 TABLE OF CONTENTS GENERAL INFORMATION... 1 Product
KAPA hgdna QUANTIFICATION AND QC KIT:
 Poster Note As presented at AGBT 2015, Marco Island, FL KAPA hgdna QUANTIFICATION AND QC KIT: The KAPA Human Genomic DNA Quantification and QC Kit Enables Prediction of Sequencing Performance through User-Defined
Poster Note As presented at AGBT 2015, Marco Island, FL KAPA hgdna QUANTIFICATION AND QC KIT: The KAPA Human Genomic DNA Quantification and QC Kit Enables Prediction of Sequencing Performance through User-Defined
Samantha Court Genetic Technologist (Birmingham)
 National Genetic Technologist/Healthcare Science Practitioner Training Day Cardiff - 22 nd November 2017 Samantha Court Genetic Technologist (Birmingham) ccf DNA is released from all cells undergoing apoptosis
National Genetic Technologist/Healthcare Science Practitioner Training Day Cardiff - 22 nd November 2017 Samantha Court Genetic Technologist (Birmingham) ccf DNA is released from all cells undergoing apoptosis
High Throughput Sequencing Technologies. J Fass UCD Genome Center Bioinformatics Core Tuesday December 16, 2014
 High Throughput Sequencing Technologies J Fass UCD Genome Center Bioinformatics Core Tuesday December 16, 2014 Sequencing Explosion www.genome.gov/sequencingcosts http://t.co/ka5cvghdqo Sequencing Explosion
High Throughput Sequencing Technologies J Fass UCD Genome Center Bioinformatics Core Tuesday December 16, 2014 Sequencing Explosion www.genome.gov/sequencingcosts http://t.co/ka5cvghdqo Sequencing Explosion
JetSeq DNA Library Preparation Kit. Product Manual
 JetSeq DNA Library Preparation Kit Product Manual JetSeq DNA Library Preparation Kit JetSeq DNA Library Preparation Kit TABLE OF CONTENTS 1 Kit contents 04 2 Description 05 3 Storage 06 4 Safety information
JetSeq DNA Library Preparation Kit Product Manual JetSeq DNA Library Preparation Kit JetSeq DNA Library Preparation Kit TABLE OF CONTENTS 1 Kit contents 04 2 Description 05 3 Storage 06 4 Safety information
SWIFT 2S TURBO DNA LIBRARY KITS
 DT SHEET swiftbiosci.com SWIT 2S TURBO DN LIBRRY KITS The Easiest NGS Workflow for Routine Sequencing Highlights Simple, fast, and reliable Minimal steps and hands-on time with consistent fragmentation
DT SHEET swiftbiosci.com SWIT 2S TURBO DN LIBRRY KITS The Easiest NGS Workflow for Routine Sequencing Highlights Simple, fast, and reliable Minimal steps and hands-on time with consistent fragmentation
MiSeq. system applications
 MiSeq system applications Choose your application. Load, and go. Focused power. Speed and simplicity for targeted and small-genome sequencing. Optimized sample preparation kits, push-button sequencing,
MiSeq system applications Choose your application. Load, and go. Focused power. Speed and simplicity for targeted and small-genome sequencing. Optimized sample preparation kits, push-button sequencing,
10 RXN 50 RXN 500 RXN
 SeqPlex Enhanced DNA Amplification Kit for use with high throughput sequencing technologies Catalog Number SEQXE Storage Temperature 20 C TECHNICAL BULLETIN Product Description The SeqPlex DNA Amplification
SeqPlex Enhanced DNA Amplification Kit for use with high throughput sequencing technologies Catalog Number SEQXE Storage Temperature 20 C TECHNICAL BULLETIN Product Description The SeqPlex DNA Amplification
Updated August well High-multiplexing Tagmentation and Amplification Robyn Tanny August Company Kit Catalog Number.
 96-well High-multiplexing Tagmentation and Amplification Robyn Tanny August 2015 This protocol uses the following purchased reagents: Company Kit Catalog Number Illumina Nextera DNA Sample Preparation
96-well High-multiplexing Tagmentation and Amplification Robyn Tanny August 2015 This protocol uses the following purchased reagents: Company Kit Catalog Number Illumina Nextera DNA Sample Preparation
Introduction to the MiSeq
 Introduction to the MiSeq 2011 Illumina, Inc. All rights reserved. Illumina, illuminadx, BeadArray, BeadXpress, cbot, CSPro, DASL, Eco, Genetic Energy, GAIIx, Genome Analyzer, GenomeStudio, GoldenGate,
Introduction to the MiSeq 2011 Illumina, Inc. All rights reserved. Illumina, illuminadx, BeadArray, BeadXpress, cbot, CSPro, DASL, Eco, Genetic Energy, GAIIx, Genome Analyzer, GenomeStudio, GoldenGate,
NGS Sample QC with the Agilent 2200 TapeStation. Rainer Nitsche Application Engineer Agilent Technologies, Inc.
 NGS Sample QC with the Agilent 2200 TapeStation Rainer Nitsche Application Engineer Agilent Technologies, Inc. The Agilent 2100 Bioanalyzer First commercially available Lab-on-a-Chip product Introduced
NGS Sample QC with the Agilent 2200 TapeStation Rainer Nitsche Application Engineer Agilent Technologies, Inc. The Agilent 2100 Bioanalyzer First commercially available Lab-on-a-Chip product Introduced
Lecture 7. Next-generation sequencing technologies
 Lecture 7 Next-generation sequencing technologies Next-generation sequencing technologies General principles of short-read NGS Construct a library of fragments Generate clonal template populations Massively
Lecture 7 Next-generation sequencing technologies Next-generation sequencing technologies General principles of short-read NGS Construct a library of fragments Generate clonal template populations Massively
Multiplexed Strand-specific RNA-Seq Library Preparation for Illumina Sequencing Platforms
 Multiplexed Strand-specific RNA-Seq Library Preparation for Illumina Sequencing Platforms Important Things to know before you start: This protocol generates strand-specific reads, but may lead to slightly
Multiplexed Strand-specific RNA-Seq Library Preparation for Illumina Sequencing Platforms Important Things to know before you start: This protocol generates strand-specific reads, but may lead to slightly
Supplemental File 1: Modified Nextera XT DNA Sample Preparation Guide (Illumina, USA, Part # rev. C, October 2012).
 Supplemental File 1: Modified Nextera XT DNA Sample Preparation Guide (Illumina, USA, Part # 15031942 rev. C, October 2012). Required Kit content: Box1 ATM Amplicon Tagment Mix TD Tagment DNA Buffer NPM
Supplemental File 1: Modified Nextera XT DNA Sample Preparation Guide (Illumina, USA, Part # 15031942 rev. C, October 2012). Required Kit content: Box1 ATM Amplicon Tagment Mix TD Tagment DNA Buffer NPM
Chapter 7. DNA Microarrays
 Bioinformatics III Structural Bioinformatics and Genome Analysis Chapter 7. DNA Microarrays 7.9 Next Generation Sequencing 454 Sequencing Solexa Illumina Solid TM System Sequencing Process of determining
Bioinformatics III Structural Bioinformatics and Genome Analysis Chapter 7. DNA Microarrays 7.9 Next Generation Sequencing 454 Sequencing Solexa Illumina Solid TM System Sequencing Process of determining
Executive Summary. clinical supply services
 clinical supply services case study Development and NDA-level validation of quantitative polymerase chain reaction (qpcr) procedure for detection and quantification of residual E.coli genomic DNA Executive
clinical supply services case study Development and NDA-level validation of quantitative polymerase chain reaction (qpcr) procedure for detection and quantification of residual E.coli genomic DNA Executive
BIOO LIFE SCIENCE PRODUCTS
 BIOO LIFE SCIENCE PRODUCTS FOR REFERENCE PURPOSES This manual is for Reference Purposes Only. DO NOT use this protocol to run your assays. Periodically, optimizations and revisions are made to the kit
BIOO LIFE SCIENCE PRODUCTS FOR REFERENCE PURPOSES This manual is for Reference Purposes Only. DO NOT use this protocol to run your assays. Periodically, optimizations and revisions are made to the kit
High Throughput Illumina Nextera XT DNA Library Construction on the. Biomek FX P Dual Arm Multi-96 and Span-8 Workstation.
 High Throughput Illumina Nextera XT DNA Library Construction on the Biomek FX P Dual Arm Multi-96 and Span-8 Workstation Abstract Zach Smith, M.S. Beckman Coulter, Inc.; Jeremy Day, Ph.D. La Jolla Institute
High Throughput Illumina Nextera XT DNA Library Construction on the Biomek FX P Dual Arm Multi-96 and Span-8 Workstation Abstract Zach Smith, M.S. Beckman Coulter, Inc.; Jeremy Day, Ph.D. La Jolla Institute
DNA METHYLATION NH 2 H 3. Solutions using bisulfite conversion and immunocapture Ideal for NGS, Sanger sequencing, Pyrosequencing, and qpcr
 NH 2 DNA METHYLATION H 3 C Solutions using bisulfite conversion and immunocapture Ideal for NGS, Sanger sequencing, Pyrosequencing, and qpcr NH 2 N N H PAGE 3 Understanding DNA Methylation DNA methylation
NH 2 DNA METHYLATION H 3 C Solutions using bisulfite conversion and immunocapture Ideal for NGS, Sanger sequencing, Pyrosequencing, and qpcr NH 2 N N H PAGE 3 Understanding DNA Methylation DNA methylation
TECH NOTE Pushing the Limit: A Complete Solution for Generating Stranded RNA Seq Libraries from Picogram Inputs of Total Mammalian RNA
 TECH NOTE Pushing the Limit: A Complete Solution for Generating Stranded RNA Seq Libraries from Picogram Inputs of Total Mammalian RNA Stranded, Illumina ready library construction in
TECH NOTE Pushing the Limit: A Complete Solution for Generating Stranded RNA Seq Libraries from Picogram Inputs of Total Mammalian RNA Stranded, Illumina ready library construction in
QIAGEN s NGS Solutions for Biomarkers NGS & Bioinformatics team QIAGEN (Suzhou) Translational Medicine Co.,Ltd
 QIAGEN s NGS Solutions for Biomarkers NGS & Bioinformatics team QIAGEN (Suzhou) Translational Medicine Co.,Ltd 1 Our current NGS & Bioinformatics Platform 2 Our NGS workflow and applications 3 QIAGEN s
QIAGEN s NGS Solutions for Biomarkers NGS & Bioinformatics team QIAGEN (Suzhou) Translational Medicine Co.,Ltd 1 Our current NGS & Bioinformatics Platform 2 Our NGS workflow and applications 3 QIAGEN s
APPLICATION NOTE. Abstract. Introduction
 From minuscule amounts to magnificent results: reliable ChIP-seq data from 1, cells with the True MicroChIP and the MicroPlex Library Preparation kits Abstract Diagenode has developed groundbreaking solutions
From minuscule amounts to magnificent results: reliable ChIP-seq data from 1, cells with the True MicroChIP and the MicroPlex Library Preparation kits Abstract Diagenode has developed groundbreaking solutions
Absolute Mouse Telomere Length Quantification qpcr Assay Kit (AMTLQ) Catalog #M reactions
 Absolute Mouse Telomere Length Quantification qpcr Assay Kit (AMTLQ) Catalog #M8918 100 reactions Product Description Telomeres are repetitive nucleotide elements at the ends of chromosomes that protect
Absolute Mouse Telomere Length Quantification qpcr Assay Kit (AMTLQ) Catalog #M8918 100 reactions Product Description Telomeres are repetitive nucleotide elements at the ends of chromosomes that protect
Welcome to the NGS webinar series
 Welcome to the NGS webinar series Webinar 1 NGS: Introduction to technology, and applications NGS Technology Webinar 2 Targeted NGS for Cancer Research NGS in cancer Webinar 3 NGS: Data analysis for genetic
Welcome to the NGS webinar series Webinar 1 NGS: Introduction to technology, and applications NGS Technology Webinar 2 Targeted NGS for Cancer Research NGS in cancer Webinar 3 NGS: Data analysis for genetic
GreenMasterMix (2X) b i o s c i e n c e. G E N A X X O N b i o s c i e n c e. High ROX (500nM)
 G:\products\productflyer\pcr\polymerasen\hotstart\manu_m3052_green_en.docx GreenMasterMix (2) High RO (500nM) qpcr master mix with fluorescence dye and passive reference dye Contact & Technical support
G:\products\productflyer\pcr\polymerasen\hotstart\manu_m3052_green_en.docx GreenMasterMix (2) High RO (500nM) qpcr master mix with fluorescence dye and passive reference dye Contact & Technical support
Relative Mouse Telomere Length Quantification qpcr Assay Kit (RMTLQ) Catalog #M reactions
 Relative Mouse Telomere Length Quantification qpcr Assay Kit (RMTLQ) Catalog #M8908 100 reactions Product Description Telomeres are repetitive nucleotide elements at the ends of chromosomes that protect
Relative Mouse Telomere Length Quantification qpcr Assay Kit (RMTLQ) Catalog #M8908 100 reactions Product Description Telomeres are repetitive nucleotide elements at the ends of chromosomes that protect
ab High Sensitivity DNA Library Preparation Kit (For Illumina )
 ab185905 High Sensitivity DNA Library Preparation Kit (For Illumina ) Instructions for Use For the preparation of a DNA library using sub-nanogram amounts of DNA input for next generation sequencing applications
ab185905 High Sensitivity DNA Library Preparation Kit (For Illumina ) Instructions for Use For the preparation of a DNA library using sub-nanogram amounts of DNA input for next generation sequencing applications
ab High Sensitivity DNA Library Preparation Kit (For Illumina )
 ab185905 High Sensitivity DNA Library Preparation Kit (For Illumina ) Instructions for Use For the preparation of a DNA library using sub-nanogram amounts of DNA input for next generation sequencing applications
ab185905 High Sensitivity DNA Library Preparation Kit (For Illumina ) Instructions for Use For the preparation of a DNA library using sub-nanogram amounts of DNA input for next generation sequencing applications
MASSEY GENOME SERVICE SEPTEMBER 2014 NEWSLETTER
 MASSEY GENOME SERVICE SEPTEMBER 2014 NEWSLETTER Welcome to the September 2014 issue of the Massey Genome Service (MGS) Newsletter. ILLUMINA MiSEQ NEXT GENERATION SEQUENCING A reminder, all enquiries regarding
MASSEY GENOME SERVICE SEPTEMBER 2014 NEWSLETTER Welcome to the September 2014 issue of the Massey Genome Service (MGS) Newsletter. ILLUMINA MiSEQ NEXT GENERATION SEQUENCING A reminder, all enquiries regarding
Genetics and Genomics in Medicine Chapter 3. Questions & Answers
 Genetics and Genomics in Medicine Chapter 3 Multiple Choice Questions Questions & Answers Question 3.1 Which of the following statements, if any, is false? a) Amplifying DNA means making many identical
Genetics and Genomics in Medicine Chapter 3 Multiple Choice Questions Questions & Answers Question 3.1 Which of the following statements, if any, is false? a) Amplifying DNA means making many identical
PCR Add-on Kit for Illumina Instruction Manual
 PCR Add-on Kit for Illumina Instruction Manual Catalog Numbers: 001 (SENSE mrna-seq Library Prep Kit V2 for Illumina) 009 (SENSE Total RNA-Seq Library Prep Kit for Illumina) 015 (QuantSeq 3 mrna-seq Library
PCR Add-on Kit for Illumina Instruction Manual Catalog Numbers: 001 (SENSE mrna-seq Library Prep Kit V2 for Illumina) 009 (SENSE Total RNA-Seq Library Prep Kit for Illumina) 015 (QuantSeq 3 mrna-seq Library
Data Sheet Quick PCR Cloning Kit
 Data Sheet Quick PCR Cloning Kit 6044 Cornerstone Ct. West, Ste. E DESCRIPTION: The Quick PCR Cloning Kit is a simple and highly efficient method to insert any gene or DNA fragment into a vector, without
Data Sheet Quick PCR Cloning Kit 6044 Cornerstone Ct. West, Ste. E DESCRIPTION: The Quick PCR Cloning Kit is a simple and highly efficient method to insert any gene or DNA fragment into a vector, without
NB536: Bioinformatics
 NB536: Bioinformatics Instructor Prof. Jong Kyoung Kim Department of New Biology Office: E4-613 E-mail: jkkim@dgist.ac.kr Homepage: https://scg.dgist.ac.kr Course website https://scg.dgist.ac.kr/index.php/courses
NB536: Bioinformatics Instructor Prof. Jong Kyoung Kim Department of New Biology Office: E4-613 E-mail: jkkim@dgist.ac.kr Homepage: https://scg.dgist.ac.kr Course website https://scg.dgist.ac.kr/index.php/courses
How is genome sequencing done?
 Click here to view Roche 454 Sequencing Genome Sequence FLX available at www.ssllc.com>> How is genome sequencing done? Using 454 Sequencing on the Genome Sequencer FLX System, DNA from a genome is converted
Click here to view Roche 454 Sequencing Genome Sequence FLX available at www.ssllc.com>> How is genome sequencing done? Using 454 Sequencing on the Genome Sequencer FLX System, DNA from a genome is converted
Twist Human Core Exome EF Multiplex Complete Kit, 16 Samples PN
 Twist Human Core Exome Enrichment Kit Twist Human Core Exome EF Multiplex Complete Kit, 16 Samples PN 100252 Reagents for preparing 16 exome-enriched libraries ready for sequencing from human genomic DNA
Twist Human Core Exome Enrichment Kit Twist Human Core Exome EF Multiplex Complete Kit, 16 Samples PN 100252 Reagents for preparing 16 exome-enriched libraries ready for sequencing from human genomic DNA
Genome Reagent Kits v2 Quick Reference Cards
 Chromium Genome Reagent Kits v2 Quick Reference Cards FOR USE WITH Chromium Genome Library Kit & Gel Bead Kit v2, 16 rxns PN-120258 Chromium Genome Chip Kit v2, 48 rxns PN-120257 Chromium i7 Multiplex
Chromium Genome Reagent Kits v2 Quick Reference Cards FOR USE WITH Chromium Genome Library Kit & Gel Bead Kit v2, 16 rxns PN-120258 Chromium Genome Chip Kit v2, 48 rxns PN-120257 Chromium i7 Multiplex
P HENIX. PHENIX PCR Enzyme Guide Tools For Life Science Discovery RESEARCH PRODUCTS
 PHENIX PCR Enzyme Guide PHENIX offers a broad line of premium quality PCR Enzymes. This PCR Enzyme Guide will help simplify your polymerase selection process. Each DNA Polymerase has different characteristics
PHENIX PCR Enzyme Guide PHENIX offers a broad line of premium quality PCR Enzymes. This PCR Enzyme Guide will help simplify your polymerase selection process. Each DNA Polymerase has different characteristics
BIOO LIFE SCIENCE PRODUCTS
 BIOO LIFE SCIENCE PRODUCTS FOR REFERENCE PURPOSES This manual is for Reference Purposes Only. DO NOT use this protocol to run your assays. Periodically, optimizations and revisions are made to the kit
BIOO LIFE SCIENCE PRODUCTS FOR REFERENCE PURPOSES This manual is for Reference Purposes Only. DO NOT use this protocol to run your assays. Periodically, optimizations and revisions are made to the kit
BIOO LIFE SCIENCE PRODUCTS
 BIOO LIFE SCIENCE PRODUCTS FOR REFERENCE PURPOSES This manual is for Reference Purposes Only. DO NOT use this protocol to run your assays. Periodically, optimizations and revisions are made to the kit
BIOO LIFE SCIENCE PRODUCTS FOR REFERENCE PURPOSES This manual is for Reference Purposes Only. DO NOT use this protocol to run your assays. Periodically, optimizations and revisions are made to the kit
Targeted Sequencing in the NBS Laboratory
 Targeted Sequencing in the NBS Laboratory Christopher Greene, PhD Newborn Screening and Molecular Biology Branch Division of Laboratory Sciences Gene Sequencing in Public Health Newborn Screening February
Targeted Sequencing in the NBS Laboratory Christopher Greene, PhD Newborn Screening and Molecular Biology Branch Division of Laboratory Sciences Gene Sequencing in Public Health Newborn Screening February
Exploring new frontiers with next-generation sequencing
 Spotlight: NGS Exploring new frontiers with next-generation sequencing Pushing the limits of discovery Next-generation sequencing (NGS) is being utilized for numerous new and exciting applications, such
Spotlight: NGS Exploring new frontiers with next-generation sequencing Pushing the limits of discovery Next-generation sequencing (NGS) is being utilized for numerous new and exciting applications, such
