Detailed Contents of Practical Streptomyces Genetics ISBN
|
|
|
- Hester Byrd
- 6 years ago
- Views:
Transcription
1 Detailed Contents of Practical Streptomyces Genetics ISBN Chapter 1 General introduction to actinomycete biology Taxonomy of Streptomyces... 2 The genera of actinomycetes... 2 The genus Streptomyces... 2 Ecology of Streptomyces... 5 Streptomycetes as pathogens... 6 Some physiological features of primary metabolism in streptomycetes... 7 Carbon sources... 7 Nitrogen sources... 8 Amino acid catabolism... 9 Biosynthesis... 9 Some physiological novelties Antibiotic production by Streptomyces Streptomycetes as antibiotic producers Antibiosis in soil Physiology and regulation of antibiotic production 16 Developmental biology of Streptomyces The Streptomyces chromosome and its genetic elements 18 DNA base composition The chromosome Plasmids Transposable elements Phages Restriction and modification of Streptomyces DNA Genetic studies with streptomycetes and their near relatives Actinomycetes used for genetical studies Genetics and strain improvement for antibiotic and enzyme production Safety guidelines for recombinant DNA experiments with Streptomyces Chapter 2 Growth and preservation of Streptomyces Selective isolation of streptomycetes Making a Streptomyces spore suspension Plating out a Streptomyces spore suspension Streptomyces cultures on agar Growth of mycelium on cellophane overlays, nitrocellulose filters or agar Growth of Streptomyces mycelium in liquid Growth of Streptomyces for physiological studies Conditions for spore germination and reproducible exponential growth and antibiotic production by Streptomyces coelicolor Germination of Streptomyces spores Procedure Procedure Preparation of aerial mycelium and spores separately. 54 Aerial mycelium Spores Sporulation of Streptomyces in submerged cultures Preparation of protoplasts from Streptomyces lividans 66 and S. coelicolor A3(2) Preservation of Streptomyces strains and phages Preparation of lyophils Alternative I Alternative II Growing a culture from the lyophil Chapter 3 Microscopical methods Light microscopy Cellophane cultures Coverslip or slide cultures Impression preparations Staining for light microscopy DAPI Staining of Streptomyces mycelium.. 65 Staining for respiratory activity Immunofluorescent staining of Streptomyces mycelium Electron microscopy Electron microscopy of thin sections Sectioning of Streptomyces colonies Staining thin sections of Streptomyces colonies for glycogen Spatial visualisation of gene expression Chapter 4 Specialised biochemical techniques Preparation of cell-free extracts from Streptomyces Cell-free extracts from mycelium Cell-free extracts from spores Small-scale cell-free extract from spores Large-scale cell-free extract from spores Preparation of a coupled transcription-translation system from S. lividans Two-dimensional polyacrylamide gel electrophoresis and its application to Streptomyces proteins.. 83 In-vivo radioactive labelling of Streptomyces proteins 90 Two-step liquid cultivation and radioactive labelling method for Streptomyces Growth and radioactive labelling of Streptomyces on zirconium-silica beads Detection and assay of S. coelicolor (and S. lividans) antibiotics... 93
2 The S. coelicolor antibiotics Actinorhodin Undecylprodigiosin CDA Methylenomycin S. lividans as an antibiotic producer Secondary metabolite production in S. lividans transformants Chapter 5 Mutagenesis of Streptomyces by irradiation or chemicals General remarks about mutagenesis The optimal amount of mutagenesis Expression of mutations Spores or mycelium? Choice of mutagen Precautions Mutagenesis of Streptomyces spores by ultraviolet light (UV) Mutagenesis of Streptomyces spores by N-methyl-N nitro-n-nitrosoguanidine (NTG) Isolation of specific classes of mutant Resistant mutants Mutants that have to be isolated by screening Auxotrophs Other classes of mutants screened by replica plating Mutants recognised by visual examination. 106 Chapter 6 Transposon mutagenesis in Streptomyces General points about transposon mutagenesis Discovery of Streptomyces transposable elements 110 Streptomyces transposons suitable for mutagenesis of Streptomyces Use of heterologous transposons for mutagenising Streptomyces genes Transposon delivery vectors for Streptomyces Isolation of independent mutant strains Cloning transposon-tagged Streptomyces DNA in E. coli Tn4556 from S. fradiae A. Transposition of Tn4560 from puc1169 to a chromosomal location B. Protocol for inserting Tn4560 into S. coelicolor NF strains C. Tn4560 mutagenesis of SCP2* plasmids in S. lividans D. Tn4560 mutagenesis of pock-forming SCP2* in S. coelicolor IS493 from S. lividans E. Sectoring method for delivering IS493 derivatives using temperature-sensitive plasmids IS6100 from Mycobacterium fortuitum F. Transposon mutagenesis of S. lividans using psit Tn5493 derived from Tn G. Mutagenising S. lividans using pjoe H. Alternative method for isolating S. lividans with random insertions of Tn Chapter 7 In vivo genetic analysis by conjugation and protoplast fusion The modes of gene exchange in Streptomyces Mating Protoplast fusion Transduction Chromosomal recombination by transformation 128 Electroporation The practicalities of making crosses Quantitative analysis of crosses Recombination frequency Frequency of plasmid transfer Linkage mapping Establishing a genetic map by the "four-on-four" procedure The "classical" way of analysing the results of a "four-on-four" cross Analysing the results of a "four-on-four" cross by minimizing multiple crossovers Mapping a new marker by a single selection Plate-crosses Detection of conjugative plasmids by "pock" formation by transconjugants Protoplast fusion Fusion of protoplasts of S. coelicolor or S. lividans Genetic analysis by protoplast fusion Chapter 8 Preparation and analysis of genomic and plasmid DNA Isolation of genomic DNA Discussion of individual steps A. Kirby mix procedure for the isolation of genomic DNA B. Salting out procedure for the isolation of genomic DNA C. CTAB procedure for the isolation of genomic DNA Isolation of CCC plasmid DNA Discussion of individual steps A. Plasmid isolation by neutral lysis B. Standard CsCl-ethidium bromide gradient centrifugation C. CsCl density gradient for removing polysaccharides from DNA D. Plasmid isolation by alkaline lysis and potassium acetate precipitation E. Plasmid isolation by alkaline lysis and phenol
3 precipitation F. Plasmid purification using QIAGEN anion exchange column chromatography G. Plasmid isolation by the boiling method H. Alkaline denaturation of partially purified DNA samples I. Purification of samples using ethidium bromide J. Acid phenol (ph4) extraction K. Jurassic preps plasmid purification using guanidine thiocyanate Standard agarose gel electrophoresis Pulsed-field gel electrophoresis General considerations Standard procedure for preparing Streptomyces chromosomal DNA for PFGE Phenol wash procedure for preparation of actinomycete DNA for PFGE Digestion of DNA with restriction endonucleases in agarose blocks for PFGE Loading gels Running conditions for PFGE Removal of small DNA fragments Solutions Ammonium acetate EDTA ph Lithium chloride Sodium chloride Sodium hydroxide Potassium acetate and sodium acetate, 3M Tris-HCl buffer Chapter 9 General considerations about gene cloning in Streptomyces Features of Streptomyces genes Restriction-modification and other host factors Preparation of vector and target DNA How many clones are required for making a representative gene library? Choice and use of restriction endonucleases for making gene libraries Choice of cloning vector Vector host-range Size of the target DNA Integrating vectors Low copy number plasmid vectors High copy number plasmid vectors Plasmid versus phage vectors Positive selection vectors E. coli vectors containing Streptomyces selection markers Using bifunctional vectors that replicate in E. coli and Streptomyces Selective markers Antibiotic resistance Counterselectable markers Ligation conditions Transformation Finding the desired clone Sib-Selection Antibiotic biosynthetic genes Confirmation of clones Does the phenotype depend on the cloned DNA? 224 Is the cloned DNA rearranged? Is the promoter for a cloned gene present on the cloned DNA? What if the desired gene cannot be cloned? Assessing the quality of Streptomyces gene libraries. 226 Storing gene libraries Chapter 10 Introduction of DNA into Streptomyces Methods available Restriction barriers Use of single-stranded DNA Transformation and transfection in Streptomyces Polyethylene glycol (PEG) Transformation and transfection frequencies PEG-assisted transformation of Streptomyces protoplasts with plasmid DNA Standard procedure Rapid small-scale procedure Use of denatured DNA for protoplast transformation Spot-transformation Cosmid transformation Electroporation of mycelium Recognition, selection and screening of Streptomyces transformants Lethal zygosis reaction (pocks) Fertility Resistance markers Selection of antibiotic-resistant transformants by overlaying or flooding Soft agar overlays Flooding Detection of melanin-producing colonies Screening for plasmid DNA Colony hybridisation Using nitrocellulose filters Using Whatman 541 paper Preparation of colony replicas Hybridisation end labelling of oligonucleotide probes Using Whatman 540 paper Use of the Polymerase Chain Reaction (PCR) to identify transformants Conjugation from E. coli
4 Chapter 11 Plasmids and their use for gene cloning General properties of Streptomyces plasmids and their use for gene cloning Wild-type plasmids that have been used extensively to construct cloning vectors pij pjv psg SCP2* Higher copy number derivatives of SCP2*. 263 SCP2* vectors as delivery systems for gene disruptions SLP1 and psam Other integrating vectors List of special purpose vectors Bifunctional E. coli-streptomyces plasmids orit (RK2) vectors for conjugation between E. coli and Streptomyces cosmid vectors Expression vectors Vectors with promoterless reporter genes Positive selection vectors Integrating vectors Unstable and temperature-sensitive plasmids useful for gene replacement and transposon delivery Vectors without the tsr gene Vectors with resistance markers other than the common ones Vectors with blue/white selection (lacz ) in E. coli Chapter 12 Streptomyces phages The relevance of phages to Streptomyces genetics Occurrence, isolation and storage of Streptomyces phages Lytic and temperate phages Phages from soil Phages from lysogens Phages as industrial contaminants Streptomyces phage genetics In vivo physiological and genetic studies Deletion mutants and DNA packaging limitation 273 Phage DNA Uses of wild-type phages in the study of their hosts. 274 Transduction Localised mutagenesis using generalised transduction Restriction-modification systems Storage of Streptomyces phages Plaque assay of Streptomyces phages Single-plaque isolation of Streptomyces phages Preparation of high-titre Streptomyces phage stocks. 278 Isolation of new Streptomyces phages Isolation procedure I (direct method) Isolation procedure II (specific enrichment) Selection of potential transducing phages by pyrophosphate resistance Generalised transduction of S. venezuelae using SV1 phage Large-scale preparation of Streptomyces phage DNA 282 Small-scale preparation of Streptomyces phage DNA 285 Chapter 13 Cloning with phage vectors General features of C31 and its vector derivatives. 290 Shotgun cloning with C31 vectors Choice of C31 vectors for mutational cloning. 291 Choice of C31 vectors for screening by complementation of mutants or acquisition of new capabilities Ligation conditions Maximising and estimating insert frequency Construction and stability of lysogens Homogenotisation Application of C31 to gene fusions General features of the C31::xylE vectors KC862: a xyle-containing C31 derivative that gives yellow plaques only when carrying inserts with active promoters 296 A single copy number promoter-probe vector, KC Vectors for in situ fusions of xyle to chromo somally located transcription units Other phage-based cloning systems Prophage transformation with phage SAt-1 of S. azureus Phage-mediated transduction of plasmids Vectors based on other Streptomyces phages Use of integration functions of Streptomyces phages Transfection Transfer of Streptomyces phage DNA onto nitrocellulose filters for plaque hybridisation. 303 Preparation of C31 lysogens Procedure A Procedure B Low-tech method for detecting C31 derivatives containing resistance genes Use of glka counterselection to select deletions from, or loss of, C31 prophages
5 Chapter 14 Gene disruption and gene replacement Creating null mutants Method (a). Insertional inactivation via a single crossover Method (b). Insertional inactivation via double crossing over Method (c). Insertional inactivation using an inframe deletion Other factors affecting the choice of approach Polar effects The size of intervals used Mutation stability Method of delivery Non-replicating E. coli plasmids Temperature-sensitive replicons Phage vectors Unstable replicons Dealing with essential genes Introduction of point mutations and other subtle changes Counterselection of the delivery vector "Heterologous" disruptions and replacements Choice of resistance markers Selecting for single crossover intermediates during gene replacement A practical example of gene disruption Homogenotisation Problems arising from unintended homogenotisation events Cloning mutant alleles by homogenotisation Gene replacements involving whole gene clusters Mutational analysis of transcription units Chapter 15 Reporter systems Introduction The problems with lacz Choice of vector: plasmid, phage and transposon systems Antibiotic resistance genes as reporters: neo and cat. 342 The tyrosinase-encoding operon of S. glaucescens as a reporter system The whie (spore pigment) major operon as a reporter system The catechol 2,3-dioxygenase determinant, xyle, a readily quantified reporter gene for Streptomyces Detection of xyle expression in situ in colonies. 344 Detection of xyle expression in C31 plaques. 344 Assay of catechol 2,3-dioxygenase in cell-free extracts Use of the Vibrio harveyi luxab genes as a reporter system Reporter systems based on EGFP A reporter gene encoding a thermostable malate dehydrogenase Assay of thermostable malate dehydrogenase in cell-free extracts The ampc ( -lactamase) gene as a reporter The lac (secreted -galactosidase) gene of S. lividans as a reporter for transcription and secretion The redd gene of S. coelicolor as an easily scorable reporter of transcription in S. coelicolor and S. lividans Chapter 16 RNA methods General precautions when working with RNA Harvesting Streptomyces cultures for RNA isolation. 354 Isolation of RNA Isolation of RNA using modified Kirby mix, phenol/chloroform extraction and DNase I treatment Isolation of RNA using CsCl gradients Isolation of RNA using SDS and hot phenol Storage of RNA Assessing the quantity and quality of RNA preparations Spectrophotometry Agarose gel electrophoresis High resolution S1 nuclease mapping General strategies for making probes for high resolution S1 nuclease mapping Notes of caution in probe construction Specific activity of the probe How much probe to add to each hybridisation reaction High resolution S1 nuclease mapping of the 3 ends of transcripts Hybridisation solution The practicalities of high resolution S1 nuclease mapping Controls Generating sequencing ladders for high resolution S1 nuclease mapping Interpretation of results Low resolution S1 nuclease mapping Primer extension mapping Northern blotting In vitro transcription Streptomyces RNA polymerase purification Standard purification of Streptomyces RNA polymerase Purification of histidine-tagged RNA polymerase
6 Preparation of DNA-cellulose Chapter 17 Production and secretion of proteins by Streptomyces Transcription initiation Translation initiation Signal peptides Codon usage Regulated expression systems Culture conditions Levels of expression A selection of plasmids suitable for intracellular expression Plasmids suitable for secretion Optimising expression of Streptomyces genes in E. coli Changing the codon usage at the 5 end of a coding region E. coli vectors that have been used to overexpress streptomycete genes Chapter 18 Analysing Streptomyces DNA DNA sequencing Alternative and additional Maxam and Gilbert base-specific reactions for sequencing end-labelled DNA Sequence analysis Identifying protein coding regions FRAME analysis Codon preference Hidden Markov model Codon usage tables Accessing Streptomyces (actinomycete) sequences in the databases Design of oligonucleotides for use as probes and PCR primers PCR conditions Chapter 19 Media, buffers and suppliers Agar Media Minimal medium (MM) Complete medium (CM) Hickey-Tresner agar (HT agar) R2 Medium R2YE Medium R5 Medium Mannitol soya flour medium (MS) Supplemented minimal medium, solid (SMMS) 410 MMT Difco nutrient agar (DNA) Oxoid nutrient agar (ONA) Soft nutrient agar (SNA) L agar Liquid media Yeast extract-malt extract medium (YEME) Tryptone soya broth (TSB) Difco nutrient broth (DNB) L broth (LB) Supplemented liquid minimal medium (SMM). 413 Minimal liquid medium (NMMP) Labelling medium for Streptomyces X YT medium Growth factor supplements Buffers P (protoplast) Buffer T (transformation) buffer L (lysis) buffer TE Buffer SM Buffer SSC Addresses of suppliers Chapter 20 Genome maps and genetically marked strains S. coelicolor A3(2) Genetic/physical map Genetically marked strains Genome sequencing project S. lividans Genetic/physical map Genetically marked strains Genetic differences between S. coelicolor A3(2) and S. lividans S. griseus S. ambofaciens S. rimosus Plasmids SCP1 and SLP Chapter 21 Maps of DNA fragments Conventions used for the restriction maps List of restriction endonucleases, recognition sites and isoschizomers (Table) Lists of genes Alphabetical list of resistance and indicator genes, and other DNA fragments (Table) List of genes grouped according to function Resistance genes Counterselectable markers Indicator genes Other DNA fragments Resistance genes grouped according to antibiotic class Aminoglycosides Bialaphos, phosphinothricin Bleomycin, phleomycin Chloramphenicol
7 Gyrase inhibitors, novobiocin, ciprofloxacin 450 Hygromycin Macrotetrolides, nonactin, tetranactin MLS (macrolide, lincosamide and streptogramin B) resistance Puromycin Spectinomycin, streptomycin Streptothricins Tetracyclines Thiostrepton and analogues Viomycin, capreomycin B Other resistances Antibiotics, antimetabolites, and suppliers (Table) Maps of DNA fragments Resistance genes Counterselectable markers Indicator genes Other DNA fragments Integrating plasmids derived from C31 and phage VWB (Table) Plasmid maps grouped according to the Streptomyces replicon (Table) E. coli plasmids (Table) Transposable elements (Table) Restriction maps Phage C31 and its derivatives Integrating plasmids derived from C31 and phage VWB Plasmid maps grouped according to the Streptomyces replicon pij101 derivatives pres1 derivatives pjv1 derivatives psg5 derivatives SCP2* derivatives psam2 and SLP1 derivatives E. coli plasmids Transposable elements IS117 derivatives IS493 derivatives Tn4556 derivatives Chapter 22 Maps of plasmids, transposons and phage genomes Conventions used for the restriction maps Lists of all restriction maps Phage C31 and its derivatives (Table) Tn5 derivative Index with 3000 entries
8 Practical Streptomyces Genetics Order form Please print this form, fill it in and mail it, with payment made out to the John Innes Centre Ltd, to: D.A. Hopwood (Streptomyces Manual) John Innes Centre Norwich NR4 7UH England in case of queries) Price per copy plus postage and packing* Postage and packing charges: UK and Europe: 7.00 per book Rest of the world: per book I should like to order copies of the book (publication date July 2000) Payment must be made in pounds sterling only, to the John Innes Centre Ltd by one of the following methods. I enclose a CHEQUE for drawn against a UK bank I enclose an INTERNATIONAL MONEY ORDER for payable through a UK designated Bank I enclose a BANKER's DRAFT for payable through a UK clearing bank Delivery address (this is your mailing label: please use typewriter or block capitals) Name: Address: (optional):
2054, Chap. 14, page 1
 2054, Chap. 14, page 1 I. Recombinant DNA technology (Chapter 14) A. recombinant DNA technology = collection of methods used to perform genetic engineering 1. genetic engineering = deliberate modification
2054, Chap. 14, page 1 I. Recombinant DNA technology (Chapter 14) A. recombinant DNA technology = collection of methods used to perform genetic engineering 1. genetic engineering = deliberate modification
Molecular Genetics Techniques. BIT 220 Chapter 20
 Molecular Genetics Techniques BIT 220 Chapter 20 What is Cloning? Recombinant DNA technologies 1. Producing Recombinant DNA molecule Incorporate gene of interest into plasmid (cloning vector) 2. Recombinant
Molecular Genetics Techniques BIT 220 Chapter 20 What is Cloning? Recombinant DNA technologies 1. Producing Recombinant DNA molecule Incorporate gene of interest into plasmid (cloning vector) 2. Recombinant
Lecture Four. Molecular Approaches I: Nucleic Acids
 Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
Design. Construction. Characterization
 Design Construction Characterization DNA mrna (messenger) A C C transcription translation C A C protein His A T G C T A C G Plasmids replicon copy number incompatibility selection marker origin of replication
Design Construction Characterization DNA mrna (messenger) A C C transcription translation C A C protein His A T G C T A C G Plasmids replicon copy number incompatibility selection marker origin of replication
Genetics Lecture Notes Lectures 13 16
 Genetics Lecture Notes 7.03 2005 Lectures 13 16 Lecture 13 Transposable elements Transposons are usually from 10 3 to 10 4 base pairs in length, depending on the transposon type. The key property of transposons
Genetics Lecture Notes 7.03 2005 Lectures 13 16 Lecture 13 Transposable elements Transposons are usually from 10 3 to 10 4 base pairs in length, depending on the transposon type. The key property of transposons
Regulation of enzyme synthesis
 Regulation of enzyme synthesis The lac operon is an example of an inducible operon - it is normally off, but when a molecule called an inducer is present, the operon turns on. The trp operon is an example
Regulation of enzyme synthesis The lac operon is an example of an inducible operon - it is normally off, but when a molecule called an inducer is present, the operon turns on. The trp operon is an example
Molecular Cell Biology - Problem Drill 11: Recombinant DNA
 Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
Reading Lecture 3: 24-25, 45, Lecture 4: 66-71, Lecture 3. Vectors. Definition Properties Types. Transformation
 Lecture 3 Reading Lecture 3: 24-25, 45, 55-66 Lecture 4: 66-71, 75-79 Vectors Definition Properties Types Transformation 56 VECTORS- Definition Vectors are carriers of a DNA fragment of interest Insert
Lecture 3 Reading Lecture 3: 24-25, 45, 55-66 Lecture 4: 66-71, 75-79 Vectors Definition Properties Types Transformation 56 VECTORS- Definition Vectors are carriers of a DNA fragment of interest Insert
Selected Techniques Part I
 1 Selected Techniques Part I Gel Electrophoresis Can be both qualitative and quantitative Qualitative About what size is the fragment? How many fragments are present? Is there in insert or not? Quantitative
1 Selected Techniques Part I Gel Electrophoresis Can be both qualitative and quantitative Qualitative About what size is the fragment? How many fragments are present? Is there in insert or not? Quantitative
T. A. Brown. Gene Cloning. & DNA Analysis. An Introduction. Seventh Edition
 T. A. Brown Gene Cloning & DNA Analysis An Introduction Seventh Edition GENE CLONING AND DNA ANALYSIS GENE CLONING AND DNA ANALYSIS An Introduction T.A. BROWN University of Manchester Manchester Seventh
T. A. Brown Gene Cloning & DNA Analysis An Introduction Seventh Edition GENE CLONING AND DNA ANALYSIS GENE CLONING AND DNA ANALYSIS An Introduction T.A. BROWN University of Manchester Manchester Seventh
Motivation From Protein to Gene
 MOLECULAR BIOLOGY 2003-4 Topic B Recombinant DNA -principles and tools Construct a library - what for, how Major techniques +principles Bioinformatics - in brief Chapter 7 (MCB) 1 Motivation From Protein
MOLECULAR BIOLOGY 2003-4 Topic B Recombinant DNA -principles and tools Construct a library - what for, how Major techniques +principles Bioinformatics - in brief Chapter 7 (MCB) 1 Motivation From Protein
Chapter 4. Recombinant DNA Technology
 Chapter 4 Recombinant DNA Technology 5. Plasmid Cloning Vectors Plasmid Plasmids Self replicating Double-stranded Mostly circular DNA ( 500 kb) Linear : Streptomyces, Borrelia burgdorferi Replicon
Chapter 4 Recombinant DNA Technology 5. Plasmid Cloning Vectors Plasmid Plasmids Self replicating Double-stranded Mostly circular DNA ( 500 kb) Linear : Streptomyces, Borrelia burgdorferi Replicon
2014 Pearson Education, Inc. CH 8: Recombinant DNA Technology
 CH 8: Recombinant DNA Technology Biotechnology the use of microorganisms to make practical products Recombinant DNA = DNA from 2 different sources What is Recombinant DNA Technology? modifying genomes
CH 8: Recombinant DNA Technology Biotechnology the use of microorganisms to make practical products Recombinant DNA = DNA from 2 different sources What is Recombinant DNA Technology? modifying genomes
CH 8: Recombinant DNA Technology
 CH 8: Recombinant DNA Technology Biotechnology the use of microorganisms to make practical products Recombinant DNA = DNA from 2 different sources What is Recombinant DNA Technology? modifying genomes
CH 8: Recombinant DNA Technology Biotechnology the use of microorganisms to make practical products Recombinant DNA = DNA from 2 different sources What is Recombinant DNA Technology? modifying genomes
B. Incorrect! Ligation is also a necessary step for cloning.
 Genetics - Problem Drill 15: The Techniques in Molecular Genetics No. 1 of 10 1. Which of the following is not part of the normal process of cloning recombinant DNA in bacteria? (A) Restriction endonuclease
Genetics - Problem Drill 15: The Techniques in Molecular Genetics No. 1 of 10 1. Which of the following is not part of the normal process of cloning recombinant DNA in bacteria? (A) Restriction endonuclease
Learning Objectives :
 Learning Objectives : Understand the basic differences between genomic and cdna libraries Understand how genomic libraries are constructed Understand the purpose for having overlapping DNA fragments in
Learning Objectives : Understand the basic differences between genomic and cdna libraries Understand how genomic libraries are constructed Understand the purpose for having overlapping DNA fragments in
Chapter 15 Recombinant DNA and Genetic Engineering. Restriction Enzymes Function as Nature s Pinking Shears
 Chapter 15 Recombinant DNA and Genetic Engineering In this chapter you will learn How restriction enzyme work and why they are essential to DNA technology. About various procedures such as cloning and
Chapter 15 Recombinant DNA and Genetic Engineering In this chapter you will learn How restriction enzyme work and why they are essential to DNA technology. About various procedures such as cloning and
Recombinant DNA Technology
 Recombinant DNA Technology Common General Cloning Strategy Target DNA from donor organism extracted, cut with restriction endonuclease and ligated into a cloning vector cut with compatible restriction
Recombinant DNA Technology Common General Cloning Strategy Target DNA from donor organism extracted, cut with restriction endonuclease and ligated into a cloning vector cut with compatible restriction
VOLUME 2. Molecular Clonin g A LABORATORY MANUA L THIRD EDITIO N. Joseph Sambrook. David W. Russell
 VOLUME 2 Molecular Clonin g A LABORATORY MANUA L THIRD EDITIO N Joseph Sambrook David W. Russell Chapter 8 In Vitro Amplification of DNA by the Polymerase 8. 1 Chain Reaction 1 The Basic Polymerase Chain
VOLUME 2 Molecular Clonin g A LABORATORY MANUA L THIRD EDITIO N Joseph Sambrook David W. Russell Chapter 8 In Vitro Amplification of DNA by the Polymerase 8. 1 Chain Reaction 1 The Basic Polymerase Chain
GENETICS EXAM 3 FALL a) is a technique that allows you to separate nucleic acids (DNA or RNA) by size.
 Student Name: All questions are worth 5 pts. each. GENETICS EXAM 3 FALL 2004 1. a) is a technique that allows you to separate nucleic acids (DNA or RNA) by size. b) Name one of the materials (of the two
Student Name: All questions are worth 5 pts. each. GENETICS EXAM 3 FALL 2004 1. a) is a technique that allows you to separate nucleic acids (DNA or RNA) by size. b) Name one of the materials (of the two
7. (10pts.) The diagram below shows the immunity region of phage l. The genes are:
 7. (10pts.) The diagram below shows the immunity region of phage l. The genes are: ci codes for ci repressor. The ci857 allele is temperature sensitive. cro codes for Cro protein. N codes for the transcription
7. (10pts.) The diagram below shows the immunity region of phage l. The genes are: ci codes for ci repressor. The ci857 allele is temperature sensitive. cro codes for Cro protein. N codes for the transcription
4 Mutant Hunts - To Select or to Screen (Perhaps Even by Brute Force)
 Genetic Techniques for Biological Research Corinne A. Michels Copyright q 2002 John Wiley & Sons, Ltd ISBNs: 0-471-89921-6 (Hardback); 0-470-84662-3 (Electronic) 4 Mutant Hunts - To Select or to Screen
Genetic Techniques for Biological Research Corinne A. Michels Copyright q 2002 John Wiley & Sons, Ltd ISBNs: 0-471-89921-6 (Hardback); 0-470-84662-3 (Electronic) 4 Mutant Hunts - To Select or to Screen
DNA miniprep by Alkaline Lysis (activity)
 DNA miniprep by Alkaline Lysis (activity) Contents 1 Alkaline Lysis 2 Exercise 1: Plasmid DNA Mini-Prep by Alkaline Lysis 3 Identification of Plasmid DNA 4 Exercise 2: Restriction Digestion Identification
DNA miniprep by Alkaline Lysis (activity) Contents 1 Alkaline Lysis 2 Exercise 1: Plasmid DNA Mini-Prep by Alkaline Lysis 3 Identification of Plasmid DNA 4 Exercise 2: Restriction Digestion Identification
Molecular Techniques Third-year Biology
 PLANNING Genetics Lab practices Molecular Techniques. Genetics Lab practices protocol. 2015-16 PCR-DIRECTED MUTAGENESIS, MOLECULAR CLONING AND RESTRICTION ANALYSIS Sessions 1 & 2 (2x3 hours): PCR-directed
PLANNING Genetics Lab practices Molecular Techniques. Genetics Lab practices protocol. 2015-16 PCR-DIRECTED MUTAGENESIS, MOLECULAR CLONING AND RESTRICTION ANALYSIS Sessions 1 & 2 (2x3 hours): PCR-directed
Please sign below if you wish to have your grades posted by the last five digits of your SSN
 BIO 226R EXAM III (Sample) PRINT YOUR NAME Please sign below if you wish to have your grades posted by the last five digits of your Signature BIO 226R Exam III has 8 pages, and 26 questions. There are
BIO 226R EXAM III (Sample) PRINT YOUR NAME Please sign below if you wish to have your grades posted by the last five digits of your Signature BIO 226R Exam III has 8 pages, and 26 questions. There are
Recombinant DNA Technology. The Role of Recombinant DNA Technology in Biotechnology. yeast. Biotechnology. Recombinant DNA technology.
 PowerPoint Lecture Presentations prepared by Mindy Miller-Kittrell, North Carolina State University C H A P T E R 8 Recombinant DNA Technology The Role of Recombinant DNA Technology in Biotechnology Biotechnology?
PowerPoint Lecture Presentations prepared by Mindy Miller-Kittrell, North Carolina State University C H A P T E R 8 Recombinant DNA Technology The Role of Recombinant DNA Technology in Biotechnology Biotechnology?
Molecular Biology: Gene cloning
 Molecular Biology: Gene cloning Author: Prof Marinda Oosthuizen Licensed under a Creative Commons Attribution license. CLONING VECTORS The central component of a gene cloning experiment is the vector or
Molecular Biology: Gene cloning Author: Prof Marinda Oosthuizen Licensed under a Creative Commons Attribution license. CLONING VECTORS The central component of a gene cloning experiment is the vector or
Mcbio 316: Exam 3. (10) 2. Compare and contrast operon vs gene fusions.
 Mcbio 316: Exam 3 Name (15) 1. Transposons provide useful tools for genetic analysis. List 5 different uses of transposon insertions. ANSWER: Many answers are possible, however, if multiple items on the
Mcbio 316: Exam 3 Name (15) 1. Transposons provide useful tools for genetic analysis. List 5 different uses of transposon insertions. ANSWER: Many answers are possible, however, if multiple items on the
Unit 8: Genomics Guided Reading Questions (150 pts total)
 Name: AP Biology Biology, Campbell and Reece, 7th Edition Adapted from chapter reading guides originally created by Lynn Miriello Chapter 18 The Genetics of Viruses and Bacteria Unit 8: Genomics Guided
Name: AP Biology Biology, Campbell and Reece, 7th Edition Adapted from chapter reading guides originally created by Lynn Miriello Chapter 18 The Genetics of Viruses and Bacteria Unit 8: Genomics Guided
Chapter 20 Recombinant DNA Technology. Copyright 2009 Pearson Education, Inc.
 Chapter 20 Recombinant DNA Technology Copyright 2009 Pearson Education, Inc. 20.1 Recombinant DNA Technology Began with Two Key Tools: Restriction Enzymes and DNA Cloning Vectors Recombinant DNA refers
Chapter 20 Recombinant DNA Technology Copyright 2009 Pearson Education, Inc. 20.1 Recombinant DNA Technology Began with Two Key Tools: Restriction Enzymes and DNA Cloning Vectors Recombinant DNA refers
By two mechanisms: Mutation Genetic Recombination
 Genetics (see text pages 257-259, 267-298) Remember what it is we want to address: How is it that prokaryotes gain new genetic ability? The cells are haploid and reproduce by fission...so how does an genetic
Genetics (see text pages 257-259, 267-298) Remember what it is we want to address: How is it that prokaryotes gain new genetic ability? The cells are haploid and reproduce by fission...so how does an genetic
Lac Operon contains three structural genes and is controlled by the lac repressor: (1) LacY protein transports lactose into the cell.
 Regulation of gene expression a. Expression of most genes can be turned off and on, usually by controlling the initiation of transcription. b. Lactose degradation in E. coli (Negative Control) Lac Operon
Regulation of gene expression a. Expression of most genes can be turned off and on, usually by controlling the initiation of transcription. b. Lactose degradation in E. coli (Negative Control) Lac Operon
GENETIC ENGINEERING worksheet
 Section A: Genetic Engineering Overview 1. What is genetic engineering? 2. Put the steps of genetic engineering in order. Recombinant product is isolated, purified and analyzed before marketing. The DNA
Section A: Genetic Engineering Overview 1. What is genetic engineering? 2. Put the steps of genetic engineering in order. Recombinant product is isolated, purified and analyzed before marketing. The DNA
Chapter 20 Biotechnology
 Chapter 20 Biotechnology Manipulation of DNA In 2007, the first entire human genome had been sequenced. The ability to sequence an organisms genomes were made possible by advances in biotechnology, (the
Chapter 20 Biotechnology Manipulation of DNA In 2007, the first entire human genome had been sequenced. The ability to sequence an organisms genomes were made possible by advances in biotechnology, (the
BIOTECHNOLOGY : PRINCIPLES AND PROCESSES
 CHAPTER 11 BIOTECHNOLOGY : PRINCIPLES AND PROCESSES POINTS TO REMEMBER Bacteriophage : A virus that infects bacteria. Bioreactor : A large vessel in which raw materials are biologically converted into
CHAPTER 11 BIOTECHNOLOGY : PRINCIPLES AND PROCESSES POINTS TO REMEMBER Bacteriophage : A virus that infects bacteria. Bioreactor : A large vessel in which raw materials are biologically converted into
462 BCH. Biotechnology & Genetic engineering. (Practical)
 462 BCH Biotechnology & Genetic engineering (Practical) Nora Aljebrin Office: Building 5, 3 rd floor, 5T304 E.mail: naljebrin@ksu.edu.sa All lectures and lab sheets are available on my website: Fac.ksu.edu.sa\naljebrin
462 BCH Biotechnology & Genetic engineering (Practical) Nora Aljebrin Office: Building 5, 3 rd floor, 5T304 E.mail: naljebrin@ksu.edu.sa All lectures and lab sheets are available on my website: Fac.ksu.edu.sa\naljebrin
Genetic Adaptation II. Microbial Physiology Module 3
 Genetic Adaptation II Microbial Physiology Module 3 Topics Topic 4: Topic 5: Transposable Elements Exchange of Genetic Material Between Organisms Topic 5a: Protection Against Foreign DNA Aims and Objectives
Genetic Adaptation II Microbial Physiology Module 3 Topics Topic 4: Topic 5: Transposable Elements Exchange of Genetic Material Between Organisms Topic 5a: Protection Against Foreign DNA Aims and Objectives
XactEdit Cas9 Nuclease with NLS User Manual
 XactEdit Cas9 Nuclease with NLS User Manual An RNA-guided recombinant endonuclease for efficient targeted DNA cleavage Catalog Numbers CE1000-50K, CE1000-50, CE1000-250, CE1001-250, CE1001-1000 Table of
XactEdit Cas9 Nuclease with NLS User Manual An RNA-guided recombinant endonuclease for efficient targeted DNA cleavage Catalog Numbers CE1000-50K, CE1000-50, CE1000-250, CE1001-250, CE1001-1000 Table of
Lectures of Dr.Mohammad Alfaham. The Bacterial Genetics
 Lectures of Dr.Mohammad Alfaham The Bacterial Genetics is the total collection of genes carried by a bacterium both on its chromosome and on its extrachromosomal genetic elements (plasmids) A Gene A gene
Lectures of Dr.Mohammad Alfaham The Bacterial Genetics is the total collection of genes carried by a bacterium both on its chromosome and on its extrachromosomal genetic elements (plasmids) A Gene A gene
CONTENTS. LIST OF FIGURES... i. LIST OF TABLES... iii. SUMMARY OF THE WORK DONE... vi
 CONTENTS SECTION ~ LIST OF FIGURES... i LIST OF TABLES... iii LIST OF ABBREVIATIONS USED... ~... iv SUMMARY OF THE WORK DONE... vi 1. INTROD UCTION... :.................................... 1 1.1. AN OVERVIEW
CONTENTS SECTION ~ LIST OF FIGURES... i LIST OF TABLES... iii LIST OF ABBREVIATIONS USED... ~... iv SUMMARY OF THE WORK DONE... vi 1. INTROD UCTION... :.................................... 1 1.1. AN OVERVIEW
2 nd year Medical Students - JU Bacterial genetics. Dr. Hamed Al Zoubi Associate Professor of Medical Microbiology. MBBS / J.U.S.
 2 nd year Medical Students - JU Bacterial genetics Dr. Hamed Al Zoubi Associate Professor of Medical Microbiology. MBBS / J.U.S.T MSc, PhD/ UK Bacterial genetics ILOs: bacterial genome and replication
2 nd year Medical Students - JU Bacterial genetics Dr. Hamed Al Zoubi Associate Professor of Medical Microbiology. MBBS / J.U.S.T MSc, PhD/ UK Bacterial genetics ILOs: bacterial genome and replication
Biotechnology. Chapter 20. Biology Eighth Edition Neil Campbell and Jane Reece. PowerPoint Lecture Presentations for
 Chapter 20 Biotechnology PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from Joan Sharp Copyright
Chapter 20 Biotechnology PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from Joan Sharp Copyright
CELL BIOLOGY - CLUTCH CH TECHNIQUES IN CELL BIOLOGY.
 !! www.clutchprep.com CONCEPT: LIGHT MICROSCOPE A light microscope uses light waves and magnification to view specimens Can be used to visualize transparent, and some cellular components - Can visualize
!! www.clutchprep.com CONCEPT: LIGHT MICROSCOPE A light microscope uses light waves and magnification to view specimens Can be used to visualize transparent, and some cellular components - Can visualize
ml recombinant E. coli cultures (at a density of A 600 units per ml)
 for purification of plasmid DNA, from bacterial cultures Cat. No. 1 754 777 (50 purifications) Cat. No. 1 754 785 (50 purifications) Principle Alkaline lysis releases plasmid DNA from bacteria and RNase
for purification of plasmid DNA, from bacterial cultures Cat. No. 1 754 777 (50 purifications) Cat. No. 1 754 785 (50 purifications) Principle Alkaline lysis releases plasmid DNA from bacteria and RNase
Competency 1: The student will demonstrate knowledge of the safety regulations that must be implemented at the biotechnology laboratory by:
 Common Course Number: BSC-4420-L Course Title: Biotechnology Laboratory Catalog Course Description: This course provides students with practical, hands-on laboratory experiences to supplement the BSC-4330
Common Course Number: BSC-4420-L Course Title: Biotechnology Laboratory Catalog Course Description: This course provides students with practical, hands-on laboratory experiences to supplement the BSC-4330
Micro 411/Genome Sci411 Exam Three
 Name Question 1. 20pts Diagram the initiation of recombination by RecBCD starting with double stranded DNA. Make sure to label all enzymes, the location of the Chi site and where RecA is required. A. 5
Name Question 1. 20pts Diagram the initiation of recombination by RecBCD starting with double stranded DNA. Make sure to label all enzymes, the location of the Chi site and where RecA is required. A. 5
Bootcamp: Molecular Biology Techniques and Interpretation
 Bootcamp: Molecular Biology Techniques and Interpretation Bi8 Winter 2016 Today s outline Detecting and quantifying nucleic acids and proteins: Basic nucleic acid properties Hybridization PCR and Designing
Bootcamp: Molecular Biology Techniques and Interpretation Bi8 Winter 2016 Today s outline Detecting and quantifying nucleic acids and proteins: Basic nucleic acid properties Hybridization PCR and Designing
STANDARD CLONING PROCEDURES. Shotgun cloning (using a plasmid vector and E coli as a host).
 STANDARD CLONING PROCEDURES Shotgun cloning (using a plasmid vector and E coli as a host). 1) Digest donor DNA and plasmid DNA with the same restriction endonuclease 2) Mix the fragments together and treat
STANDARD CLONING PROCEDURES Shotgun cloning (using a plasmid vector and E coli as a host). 1) Digest donor DNA and plasmid DNA with the same restriction endonuclease 2) Mix the fragments together and treat
The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity
 Promega Notes Magazine Number 62, 1997, p. 02 The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity By Christine Andrews and Scott Lesley Promega
Promega Notes Magazine Number 62, 1997, p. 02 The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity By Christine Andrews and Scott Lesley Promega
XXII DNA cloning and sequencing. Outline
 XXII DNA cloning and sequencing 1) Deriving DNA for cloning Outline 2) Vectors; forming recombinant DNA; cloning DNA; and screening for clones containing recombinant DNA [replica plating and autoradiography;
XXII DNA cloning and sequencing 1) Deriving DNA for cloning Outline 2) Vectors; forming recombinant DNA; cloning DNA; and screening for clones containing recombinant DNA [replica plating and autoradiography;
Mutagenesis for Studying Gene Function Spring, 2007 Guangyi Wang, Ph.D. POST103B
 Mutagenesis for Studying Gene Function Spring, 2007 Guangyi Wang, Ph.D. POST103B guangyi@hawaii.edu http://www.soest.hawaii.edu/marinefungi/ocn403webpage.htm Overview of Last Lecture DNA microarray hybridization
Mutagenesis for Studying Gene Function Spring, 2007 Guangyi Wang, Ph.D. POST103B guangyi@hawaii.edu http://www.soest.hawaii.edu/marinefungi/ocn403webpage.htm Overview of Last Lecture DNA microarray hybridization
AmpliScribe T7-Flash Transcription Kit
 AmpliScribe T7-Flash Transcription Kit Cat. Nos. ASF3257 and ASF3507 Available exclusively thru Lucigen. lucigen.com/epibio www.lucigen.com MA191E AmpliScribe T7-Flash Transcription Kit 12/2016 1 1. Introduction
AmpliScribe T7-Flash Transcription Kit Cat. Nos. ASF3257 and ASF3507 Available exclusively thru Lucigen. lucigen.com/epibio www.lucigen.com MA191E AmpliScribe T7-Flash Transcription Kit 12/2016 1 1. Introduction
DNA recombination without ligase: TOPO TA Cloning. topoisomerase
 DNA recombination without ligase: TOPO TA Cloning topoisomerase Cloning strategies, cloning in bacteria other than E.coli Mitesh Shrestha Cloning strategies Cloning in bacteria other than E.coli Convenient
DNA recombination without ligase: TOPO TA Cloning topoisomerase Cloning strategies, cloning in bacteria other than E.coli Mitesh Shrestha Cloning strategies Cloning in bacteria other than E.coli Convenient
Contents... vii. List of Figures... xii. List of Tables... xiv. Abbreviatons... xv. Summary... xvii. 1. Introduction In vitro evolution...
 vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
FROM EXPERIMENTS IN BACTERIAL GENETICS AND GENE TECHNIQUE
 Uppsala 2001-04-01 REPORT FROM EXPERIMENTS IN BACTERIAL GENETICS AND GENE TECHNIQUE Laboratory assistants: Maria Jönsson Amera Gibreel Students: Contents ASSIGNMENT:... 3 INTRODUCTION:... 3 MATERIAL AND
Uppsala 2001-04-01 REPORT FROM EXPERIMENTS IN BACTERIAL GENETICS AND GENE TECHNIQUE Laboratory assistants: Maria Jönsson Amera Gibreel Students: Contents ASSIGNMENT:... 3 INTRODUCTION:... 3 MATERIAL AND
The Biotechnology Toolbox
 Chapter 15 The Biotechnology Toolbox Cutting and Pasting DNA Cutting DNA Restriction endonuclease or restriction enzymes Cellular protection mechanism for infected foreign DNA Recognition and cutting specific
Chapter 15 The Biotechnology Toolbox Cutting and Pasting DNA Cutting DNA Restriction endonuclease or restriction enzymes Cellular protection mechanism for infected foreign DNA Recognition and cutting specific
Biotechnolog y and DNA Technology
 PowerPoint Lecture Presentations prepared by Bradley W. Christian, McLennan Community College C H A P T E R 9 Biotechnolog y and DNA Technology Introduction to Biotechnology Biotechnology: the use of microorganisms,
PowerPoint Lecture Presentations prepared by Bradley W. Christian, McLennan Community College C H A P T E R 9 Biotechnolog y and DNA Technology Introduction to Biotechnology Biotechnology: the use of microorganisms,
2054, Chap. 13, page 1
 2054, Chap. 13, page 1 I. Microbial Recombination and Plasmids (Chapter 13) A. recombination = process of combining genetic material from 2 organisms to produce a genotype different from either parent
2054, Chap. 13, page 1 I. Microbial Recombination and Plasmids (Chapter 13) A. recombination = process of combining genetic material from 2 organisms to produce a genotype different from either parent
Amgen Laboratory Series. Tabs C and E
 Amgen Laboratory Series Tabs C and E Chapter 2A Goals Describe the characteristics of plasmids Explain how plasmids are used in cloning a gene Describe the function of restriction enzymes Explain how to
Amgen Laboratory Series Tabs C and E Chapter 2A Goals Describe the characteristics of plasmids Explain how plasmids are used in cloning a gene Describe the function of restriction enzymes Explain how to
BS1940 Course Topics Fall 2001 Drs. Hatfull and Arndt
 BS1940 Course Topics Fall 2001 Drs. Hatfull and Arndt Introduction to molecular biology Combining genetics, biochemistry, structural chemistry Information flow in biological systems: The Central Dogma
BS1940 Course Topics Fall 2001 Drs. Hatfull and Arndt Introduction to molecular biology Combining genetics, biochemistry, structural chemistry Information flow in biological systems: The Central Dogma
GENETICS - CLUTCH CH.5 GENETICS OF BACTERIA AND VIRUSES.
 !! www.clutchprep.com CONCEPT: WORKING WITH MICROORGANISMS Bacteria are easy to with in a laboratory setting They are fast dividing, take up little space, and are easily grown in a lab - Plating is when
!! www.clutchprep.com CONCEPT: WORKING WITH MICROORGANISMS Bacteria are easy to with in a laboratory setting They are fast dividing, take up little space, and are easily grown in a lab - Plating is when
Enhancers mutations that make the original mutant phenotype more extreme. Suppressors mutations that make the original mutant phenotype less extreme
 Interactomics and Proteomics 1. Interactomics The field of interactomics is concerned with interactions between genes or proteins. They can be genetic interactions, in which two genes are involved in the
Interactomics and Proteomics 1. Interactomics The field of interactomics is concerned with interactions between genes or proteins. They can be genetic interactions, in which two genes are involved in the
TRANSMISSIBLE GENETIC ELEMENTS: PLASMIDS, TRANSPOSONS & INTEGRONS
 TRANSMISSIBLE GENETIC ELEMENTS: PLASMIDS, TRANSPOSONS & INTEGRONS MOBILE GENETIC ELEMENTS 1 PLASMIDS -MOST OFTEN CIRCULAR MOLECULES OF DOUBLE-STRANDED DNA -VARY WIDELY IN SIZE -SELF-TRANSMISSIBLE ELEMENTS
TRANSMISSIBLE GENETIC ELEMENTS: PLASMIDS, TRANSPOSONS & INTEGRONS MOBILE GENETIC ELEMENTS 1 PLASMIDS -MOST OFTEN CIRCULAR MOLECULES OF DOUBLE-STRANDED DNA -VARY WIDELY IN SIZE -SELF-TRANSMISSIBLE ELEMENTS
DNA Cloning with Cloning Vectors
 Cloning Vectors A M I R A A. T. A L - H O S A R Y L E C T U R E R O F I N F E C T I O U S D I S E A S E S F A C U L T Y O F V E T. M E D I C I N E A S S I U T U N I V E R S I T Y - E G Y P T DNA Cloning
Cloning Vectors A M I R A A. T. A L - H O S A R Y L E C T U R E R O F I N F E C T I O U S D I S E A S E S F A C U L T Y O F V E T. M E D I C I N E A S S I U T U N I V E R S I T Y - E G Y P T DNA Cloning
CHAPTER 20 DNA TECHNOLOGY AND GENOMICS. Section A: DNA Cloning
 Section A: DNA Cloning 1. DNA technology makes it possible to clone genes for basic research and commercial applications: an overview 2. Restriction enzymes are used to make recombinant DNA 3. Genes can
Section A: DNA Cloning 1. DNA technology makes it possible to clone genes for basic research and commercial applications: an overview 2. Restriction enzymes are used to make recombinant DNA 3. Genes can
Supplementary Information
 Supplementary Information Supplementary Figures Figure S1. Study of mgtl translation in vitro. (A) Detection of 5 LR RNA using wild-type and anti-sd (91-95) substituted templates in a transcription-translation
Supplementary Information Supplementary Figures Figure S1. Study of mgtl translation in vitro. (A) Detection of 5 LR RNA using wild-type and anti-sd (91-95) substituted templates in a transcription-translation
Chapter 5. Objectives: Exploration of gene Recombinant DNA technology Genome sequencing Manipulation of Eukaryotic genes
 Chapter 5 Objectives: Exploration of gene Recombinant DNA technology Genome sequencing Manipulation of Eukaryotic genes Restriction enzymes - cleave DNA et specific sequence - found in prokaryotes, cleave
Chapter 5 Objectives: Exploration of gene Recombinant DNA technology Genome sequencing Manipulation of Eukaryotic genes Restriction enzymes - cleave DNA et specific sequence - found in prokaryotes, cleave
Combining Techniques to Answer Molecular Questions
 Combining Techniques to Answer Molecular Questions UNIT FM02 How to cite this article: Curr. Protoc. Essential Lab. Tech. 9:FM02.1-FM02.5. doi: 10.1002/9780470089941.etfm02s9 INTRODUCTION This manual is
Combining Techniques to Answer Molecular Questions UNIT FM02 How to cite this article: Curr. Protoc. Essential Lab. Tech. 9:FM02.1-FM02.5. doi: 10.1002/9780470089941.etfm02s9 INTRODUCTION This manual is
280 Index. Bacteriophage typing, discrimination, 18 reaction difference rule, 19, 23, 24 reproducibility, 18, 19 selection of bacteriophage, 17
 Index Antibiotic, 1-4 factors affecting activity, 3 minimum inhibitory concentration (MIC), 1, 3 solubility, 2, 4 Antibiotic resistance profiling, 1-4 antibiogram, 2 breakpoint method, 2 Antibodies, enzyme
Index Antibiotic, 1-4 factors affecting activity, 3 minimum inhibitory concentration (MIC), 1, 3 solubility, 2, 4 Antibiotic resistance profiling, 1-4 antibiogram, 2 breakpoint method, 2 Antibodies, enzyme
RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by. 5 AGACACAAACACCAUUGUCACACUCCACAGC; Rand-2 OMe,
 Materials and methods Oligonucleotides and DNA constructs RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by Dharmacon Inc. (Lafayette, CO). The sequences were: 122-2 OMe, 5
Materials and methods Oligonucleotides and DNA constructs RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by Dharmacon Inc. (Lafayette, CO). The sequences were: 122-2 OMe, 5
CHAPTER 9 DNA Technologies
 CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
The Mosaic Nature of Genomes
 The Mosaic Nature of Genomes n DNA sequence is not static Mutations of single bases Large deletions Large insertions of sequence n Transferred from other species n New functions useful in particular situations
The Mosaic Nature of Genomes n DNA sequence is not static Mutations of single bases Large deletions Large insertions of sequence n Transferred from other species n New functions useful in particular situations
BIOTECHNOLOGY. Sticky & blunt ends. Restriction endonucleases. Gene cloning an overview. DNA isolation & restriction
 BIOTECHNOLOGY RECOMBINANT DNA TECHNOLOGY Recombinant DNA technology involves sticking together bits of DNA from different sources. Made possible because DNA & the genetic code are universal. 2004 Biology
BIOTECHNOLOGY RECOMBINANT DNA TECHNOLOGY Recombinant DNA technology involves sticking together bits of DNA from different sources. Made possible because DNA & the genetic code are universal. 2004 Biology
MOLECULAR GENETICS: TRANSFORMATION AND CLONING adapted by Dr. D. L. Vogelien
 Introduction MOLECULAR GENETICS: TRANSFORMATION AND CLONING adapted by Dr. D. L. Vogelien The field of molecular genetics has resulted in a number of practical applications that have been of tremendous
Introduction MOLECULAR GENETICS: TRANSFORMATION AND CLONING adapted by Dr. D. L. Vogelien The field of molecular genetics has resulted in a number of practical applications that have been of tremendous
Chapter 10 (Part II) Gene Isolation and Manipulation
 Biology 234 J. G. Doheny Chapter 10 (Part II) Gene Isolation and Manipulation Practice Questions: Answer the following questions with one or two sentences. 1. What does PCR stand for? 2. What does the
Biology 234 J. G. Doheny Chapter 10 (Part II) Gene Isolation and Manipulation Practice Questions: Answer the following questions with one or two sentences. 1. What does PCR stand for? 2. What does the
Understanding the Cellular Mechanism of the Excess Microsporocytes I (EMSI) Gene. Andrew ElBardissi, The Pennsylvania State University
 Understanding the Cellular Mechanism of the Excess Microsporocytes I (EMSI) Gene Andrew ElBardissi, The Pennsylvania State University Abstract: Hong Ma, The Pennsylvania State University The Excess Microsporocytes
Understanding the Cellular Mechanism of the Excess Microsporocytes I (EMSI) Gene Andrew ElBardissi, The Pennsylvania State University Abstract: Hong Ma, The Pennsylvania State University The Excess Microsporocytes
PLNT2530 (2018) Unit 6b Sequence Libraries
 PLNT2530 (2018) Unit 6b Sequence Libraries Molecular Biotechnology (Ch 4) Analysis of Genes and Genomes (Ch 5) Unless otherwise cited or referenced, all content of this presenataion is licensed under the
PLNT2530 (2018) Unit 6b Sequence Libraries Molecular Biotechnology (Ch 4) Analysis of Genes and Genomes (Ch 5) Unless otherwise cited or referenced, all content of this presenataion is licensed under the
Methods for Working with DNA and RNA
 Methods for Working with DNA and RNA 1. Gel electrophoresis A. Materials: agarose (large DNAs) vs. acrylamide (high resolution, DNA sequencing) B. Separated by its sieving property and charge: both are
Methods for Working with DNA and RNA 1. Gel electrophoresis A. Materials: agarose (large DNAs) vs. acrylamide (high resolution, DNA sequencing) B. Separated by its sieving property and charge: both are
number Done by Corrected by Doctor Hamed Al Zoubi
 number 3 Done by Neda a Baniata Corrected by Waseem Abu Obeida Doctor Hamed Al Zoubi Note: it is important to refer to slides. Bacterial genetics *The main concepts we will talk about in this lecture:
number 3 Done by Neda a Baniata Corrected by Waseem Abu Obeida Doctor Hamed Al Zoubi Note: it is important to refer to slides. Bacterial genetics *The main concepts we will talk about in this lecture:
BIO 202 Midterm Exam Winter 2007
 BIO 202 Midterm Exam Winter 2007 Mario Chevrette Lectures 10-14 : Question 1 (1 point) Which of the following statements is incorrect. a) In contrast to prokaryotic DNA, eukaryotic DNA contains many repetitive
BIO 202 Midterm Exam Winter 2007 Mario Chevrette Lectures 10-14 : Question 1 (1 point) Which of the following statements is incorrect. a) In contrast to prokaryotic DNA, eukaryotic DNA contains many repetitive
Confirming the Phenotypes of E. coli Strains
 Confirming the Phenotypes of E. coli Strains INTRODUCTION Before undertaking any experiments, we need to confirm that the phenotypes of the E. coli strains we intend to use in the planned experiments correspond
Confirming the Phenotypes of E. coli Strains INTRODUCTION Before undertaking any experiments, we need to confirm that the phenotypes of the E. coli strains we intend to use in the planned experiments correspond
Development of Positive Control for Hepatitis B Virus
 Human Journals Research Article December 2015 Vol.:2, Issue:2 All rights are reserved by Saurabh Bandhavkar et al. Development of Positive Control for Hepatitis B Virus Keywords: Hepatitis B virus, pbluescript,
Human Journals Research Article December 2015 Vol.:2, Issue:2 All rights are reserved by Saurabh Bandhavkar et al. Development of Positive Control for Hepatitis B Virus Keywords: Hepatitis B virus, pbluescript,
Experimental genetics - I
 Experimental genetics - I Examples of diseases with genetic-links Hemophilia (complete loss or altered form of factor VIII): bleeding disorder Duchenne muscular dystrophy (altered form of dystrophin) muscle
Experimental genetics - I Examples of diseases with genetic-links Hemophilia (complete loss or altered form of factor VIII): bleeding disorder Duchenne muscular dystrophy (altered form of dystrophin) muscle
Site directed mutagenesis, Insertional and Deletion Mutagenesis. Mitesh Shrestha
 Site directed mutagenesis, Insertional and Deletion Mutagenesis Mitesh Shrestha Mutagenesis Mutagenesis (the creation or formation of a mutation) can be used as a powerful genetic tool. By inducing mutations
Site directed mutagenesis, Insertional and Deletion Mutagenesis Mitesh Shrestha Mutagenesis Mutagenesis (the creation or formation of a mutation) can be used as a powerful genetic tool. By inducing mutations
Chapter 20 DNA Technology & Genomics. If we can, should we?
 Chapter 20 DNA Technology & Genomics If we can, should we? Biotechnology Genetic manipulation of organisms or their components to make useful products Humans have been doing this for 1,000s of years plant
Chapter 20 DNA Technology & Genomics If we can, should we? Biotechnology Genetic manipulation of organisms or their components to make useful products Humans have been doing this for 1,000s of years plant
7.1 Techniques for Producing and Analyzing DNA. SBI4U Ms. Ho-Lau
 7.1 Techniques for Producing and Analyzing DNA SBI4U Ms. Ho-Lau What is Biotechnology? From Merriam-Webster: the manipulation of living organisms or their components to produce useful usually commercial
7.1 Techniques for Producing and Analyzing DNA SBI4U Ms. Ho-Lau What is Biotechnology? From Merriam-Webster: the manipulation of living organisms or their components to produce useful usually commercial
NCERT. 2. An enzyme catalysing the removal of nucleotides from the ends of DNA is: a. endonuclease b. exonuclease c. DNA ligase d.
 BIOTECHNOLOGY PRINCIPLES AND PROCESSES 75 CHAPTER 11 BIOTECHNOLOGY: PRINCIPLES AND PROCESSES 1. Rising of dough is due to: MULTIPLE-CHOICE QUESTIONS a. Multiplication of yeast b. Production of CO 2 c.
BIOTECHNOLOGY PRINCIPLES AND PROCESSES 75 CHAPTER 11 BIOTECHNOLOGY: PRINCIPLES AND PROCESSES 1. Rising of dough is due to: MULTIPLE-CHOICE QUESTIONS a. Multiplication of yeast b. Production of CO 2 c.
Vectors for Gene Cloning: Plasmids and Bacteriophages
 Vectors for Gene Cloning: Plasmids and Bacteriophages DNA molecule must be able to replicate within the host cell to be able to act as a vector for gene cloning, so that numerous copies of the recombinant
Vectors for Gene Cloning: Plasmids and Bacteriophages DNA molecule must be able to replicate within the host cell to be able to act as a vector for gene cloning, so that numerous copies of the recombinant
Purification and Characterization of a DNA Plasmid Part A CHEM 4581: Biochemistry Laboratory I Version: January 18, 2008
 Purification and Characterization of a DNA Plasmid Part A CHEM 4581: Biochemistry Laboratory I Version: January 18, 2008 INTRODUCTION DNA Plasmids. A plasmid is a small double-stranded, circular DNA molecule
Purification and Characterization of a DNA Plasmid Part A CHEM 4581: Biochemistry Laboratory I Version: January 18, 2008 INTRODUCTION DNA Plasmids. A plasmid is a small double-stranded, circular DNA molecule
B. Sc. III SEMESTER V BTT- 501: MOLECULAR BIOLOGY (CORE)
 B. Sc. III SEMESTER V BTT- 501: MOLECULAR BIOLOGY (CORE) Unit I: Genome Structure:Watson and Crick model of DNA; Genome organization with specific reference to prokaryotic and eukaryotic genomes; Genome
B. Sc. III SEMESTER V BTT- 501: MOLECULAR BIOLOGY (CORE) Unit I: Genome Structure:Watson and Crick model of DNA; Genome organization with specific reference to prokaryotic and eukaryotic genomes; Genome
SCREENING AND PRESERVATION OF DNA LIBRARIES
 MODULE 4 LECTURE 5 SCREENING AND PRESERVATION OF DNA LIBRARIES 4-5.1. Introduction Library screening is the process of identification of the clones carrying the gene of interest. Screening relies on a
MODULE 4 LECTURE 5 SCREENING AND PRESERVATION OF DNA LIBRARIES 4-5.1. Introduction Library screening is the process of identification of the clones carrying the gene of interest. Screening relies on a
Gene Cloning & DNA Analysis
 CSS451 CSS/HRT 451 Gene Cloning & DNA Analysis Chapter 4-5 T-DNA LB auxin cytokin opine Oncogenic genes RB vir genes ori opine catabolism Guo-qing Song Part 1 Basic principles Gene Cloning & DNA Analysis
CSS451 CSS/HRT 451 Gene Cloning & DNA Analysis Chapter 4-5 T-DNA LB auxin cytokin opine Oncogenic genes RB vir genes ori opine catabolism Guo-qing Song Part 1 Basic principles Gene Cloning & DNA Analysis
Biotechnology. Biotechnology is difficult to define but in general it s the use of biological systems to solve problems.
 MITE 2 S Biology Biotechnology Summer 2004 Austin Che Biotechnology is difficult to define but in general it s the use of biological systems to solve problems. Recombinant DNA consists of DNA assembled
MITE 2 S Biology Biotechnology Summer 2004 Austin Che Biotechnology is difficult to define but in general it s the use of biological systems to solve problems. Recombinant DNA consists of DNA assembled
Unit 2: Metabolism and Survival Sub-Topic (2.7) Genetic Control of Metabolism (2.8) Ethical considerations in the use of microorganisms
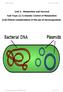 Unit 2: Metabolism and Survival Sub-Topic (2.7) Genetic Control of Metabolism (2.8) Ethical considerations in the use of microorganisms Duncanrig Secondary JHM&MHC 2015 Page 1 of 18 On completion of this
Unit 2: Metabolism and Survival Sub-Topic (2.7) Genetic Control of Metabolism (2.8) Ethical considerations in the use of microorganisms Duncanrig Secondary JHM&MHC 2015 Page 1 of 18 On completion of this
MMG 301, Lec. 25 Mutations and Bacteriophage
 MMG 301, Lec. 25 Mutations and Bacteriophage Questions for today: 1. What are mutations and how do they form? 2. How are mutant bacteria used in research? 3. What are the general properties of bacteriophage
MMG 301, Lec. 25 Mutations and Bacteriophage Questions for today: 1. What are mutations and how do they form? 2. How are mutant bacteria used in research? 3. What are the general properties of bacteriophage
7.03, 2005, Lecture 20 EUKARYOTIC GENES AND GENOMES I
 7.03, 2005, Lecture 20 EUKARYOTIC GENES AND GENOMES I For the last several lectures we have been looking at how one can manipulate prokaryotic genomes and how prokaryotic genes are regulated. In the next
7.03, 2005, Lecture 20 EUKARYOTIC GENES AND GENOMES I For the last several lectures we have been looking at how one can manipulate prokaryotic genomes and how prokaryotic genes are regulated. In the next
Purification of DNA from living cells
 Purification of DNA from living cells Total cell DNA & Plasmid DNA Grow and harvest bacterial culture Prepare cell extract Purify DNA from a cell extract Concentrate DNA samples Measure DNA concentration
Purification of DNA from living cells Total cell DNA & Plasmid DNA Grow and harvest bacterial culture Prepare cell extract Purify DNA from a cell extract Concentrate DNA samples Measure DNA concentration
Packaging of P22 DNA requires a pac site while packaging of lambda DNA requires a cos site. Briefly describe:
 1). (12 Points) Packaging of P22 DNA requires a pac site while packaging of lambda DNA requires a cos site. Briefly describe: 1. The mechanisms used by P22 and lambda to package DNA. P22 uses a headfull
1). (12 Points) Packaging of P22 DNA requires a pac site while packaging of lambda DNA requires a cos site. Briefly describe: 1. The mechanisms used by P22 and lambda to package DNA. P22 uses a headfull
Directe d Mutagenesis
 Directe d Mutagenesis A Practical Approac h M. J. McPHERSON 1. Mutagenesis facilitated by the removal or introduction of unique restriction sites 1 P. Carte r 1. Introduction to site-directed mutagenesis
Directe d Mutagenesis A Practical Approac h M. J. McPHERSON 1. Mutagenesis facilitated by the removal or introduction of unique restriction sites 1 P. Carte r 1. Introduction to site-directed mutagenesis
chapter eight: microbial genetics
 chapter eight: microbial genetics Revised 9/15/2016 the hereditary material Griffith 1927 & Avery, et al. 1944 the transforming principle coined by Griffith, identified by Avery the hereditary material
chapter eight: microbial genetics Revised 9/15/2016 the hereditary material Griffith 1927 & Avery, et al. 1944 the transforming principle coined by Griffith, identified by Avery the hereditary material
