JUNIPERUS RECURVA VAR. UNCINATA, THE HOOKED BRANCHLET JUNIPER, A NEW VARIETY FROM NEPAL
|
|
|
- Jesse Day
- 5 years ago
- Views:
Transcription
1 Phytologia (December 2009) 91(3) 361 JUNIPERUS RECURVA VAR. UNCINATA, THE HOOKED BRANCHLET JUNIPER, A NEW VARIETY FROM NEPAL Robert P. Adams Biology Department, Baylor University, Waco, TX Robert_Adams@baylor.edu Ram P. Chaudhary Central Department of Botany, Tribhuvan University, Kirtipur, Kathmandu, Nepal R. Naresh Pandey Cincinnati Children Hospital, Cincinnati, OH Ram Lakhan Singh Biology Department, Baylor University, Waco, TX 76798, USA Robert_Adams@baylor.edu ABSTRACT Routine field work in Nepal led to the discovery of a new variety of J. recurva (J. recurva var. uncinata R. P. Adams), the hooked branchlet juniper, a shrub with short leaves and hooked or recurved branchlet tips, growing at 3900 m. Analyses of sequence data from nrdna and cpdna (petn-psbm) of Juniperus indica, J. i. var. caespitosa, J. i. var. rushforthiana, J. recurva and J. squamata confirmed the distinct nature of the new variety. Analyses of leaf terpenoids from the parents, and several intermediates between J. recurva and J. r. var. uncinata support hybridization. Phytologia 91(3): (December, 2009). KEY WORDS: Juniperus indica, J. i. var. caespitosa, J. i. var. rushforthiana, J. recurva, J. recurva var. uncinata R. P. Adams, var. nov., J. squamata, SNPs, nrdna (ITS), petn, psbm, sequences, terpenoids, taxonomy, hybridization. The junipers of the central Himalayans consist of J. communis L. var. saxatilis Pall., J. indica Bertol. var. caespitosa Farjon, J. i. var.
2 362 Phytologia (December 2009) 91(3) indica Bertol., J. i. var. rushforthiana R. P. Adams, J. recurva Buch.- Ham. ex D. Don, and J. squamata Buch.-Ham. ex D. Don (Adams, 2008). While collecting specimens of J. recurva in Nepal, we discovered a shrub that differs from typical J. recurva in having branchlet tips that are hook-shaped, having shorter leaves, and branching near the base. In addition, several plants were found that appear to be intermediate between typical J. recurva and the newly discovered 'hooked-tip' plants. The purpose of this paper is to report on analyses of J. recurva, the newly discovered hooked-tipped shrubs and compare these with closely related species (Juniperus indica, J. i. var. caespitosa, J. i. var. rushforthiana, J. squamata) using nrdna (ITS) and cp petn-psbm SNPs. In addition, the volatile leaf terpenoids from the J. recurva plants and putative hybrids were analyzed. MATERIALS AND METHODS Specimens (GenBank nrdna, petn-psbm) used in this study: J. indica var. indica, Adams , (GQ118641, GQ118648), L. Singh, Dumpa, Jomson, 2900 m, Nepal; J. indica var. caespitosa, Adams (GQ118642, GQ118649), R. Chaudhary, between Kyangjin Gompa and Langtang Glacier, 4000 m, Nepal; J. indica var. rushforthiana, Adams 8140, 8141, (GQ118643, GQ118650), K. Rushforth, Soe Tajitang, 3000 m and Lingshi, 3480 m, Bhutan, J. recurva var. recurva, Adams 7210, 7211, 7215, (GQ118644, GQ118651), between Sing Gompa and Cholan Pati lodge, m, Nepal; J. recurva var. uncinata, Adams (GQ118645, GQ118652) Lauri Binayak, 3900 m, Nepal, J. squamata var. squamata, Adams 6796, (GQ118646, GQ118653) Xian Botanical Garden, China, 7012, Kunming Botanical Garden, China, J. squamata f. wilsonii, Adams 5521 (GQ118647, GQ118654), Arnold Arboretum, acc a. Voucher specimens are deposited at BAYLU. One gram (fresh weight) of the foliage was placed in 20 g of activated silica gel and transported to the lab, thence stored at -20 o C until the DNA was extracted. DNA was extracted from juniper leaves by use of a Qiagen mini-plant kit as per manufacturer's instructions.
3 Phytologia (December 2009) 91(3) 363 Amplification and sequencing ITS (nrdna) and petn-psbm amplifications were performed in 30 µl reactions using 6 ng of genomic DNA, 1.5 units Epi-Centre Fail- Safe Taq polymerase, 15 µl 2x buffer E or K (final concentration: 50 mm KCl, 50 mm Tris-HCl (ph 8.3), 200 µm each dntp, plus Epi- Centre proprietary enhancers with mm MgCl 2 according to the buffer used) and 1.8 µm each primer. Gene Primers 2x buffer annealing program size bp nrdna ITSA-42F/ ITSB+57R K 50ºC (94-50x30) petn petn5f/psbm111r E 50ºC (94-50x30) Primers (5'-3'): ITS: ITSA = GGA AGG AGA AGT CGT AAC AAG G; ITSB = CTT TTC CTC CGC TTA TTG ATA TG. ITSA and ITSB primers from Blattner (1999). additional ITS primers (based on Juniperus sequences): ITSA-42F = GAT TGA ATG ATC CGG TGA AGT ITSB+57R = ATT TTC ATG CTG GGC TCT petn - psbm: petn5f = AAC GAA GCG AAA ATC AAT CA psbm111r = AAA GAG AGG GAT TCG TAT GGA petn and psbm primers were based on conserved sequences from Juniperus species. The following PCR conditions were used: MJ Research Programmable Thermal Cycler, 30 cycles, 94 o C (1 min.), 50 o C (2 min.), 72 o C (2 min.), with a final step of 72 o C (5 min.). The PCR reaction was subjected to purification by agarose gel electrophoresis (1.5% agarose, 70 v, 55 min.). In each case, the single band was excised and purified using a Qiagen QIAquick gel extraction kit. The gel purified DNA band with the appropriate primer was sent to McLab Inc. for sequencing. Sequences for both strands were edited and a consensus sequence was produced using Chromas, version 2.31 (Technelysium Pty Ltd.). Alignments were made using MAFFT ( manually corrected, and re-analyzed using NJ with 1000 bootstrap replications (
4 364 Phytologia (December 2009) 91(3) Isolation and analysis of Oils - see Adams et al. (2009). Data Analysis - Terpenoids (as per cent total oil) were coded and compared among the taxa by the Gower metric (Gower, 1971). Principal coordinate analysis was performed by factoring the associational matrix using the formulation of Gower (1966) and Veldman (1967). RESULTS AND DISCUSSION The hooked branchlet junipers were found at 3900 m on a south facing slope at timberline at Lauri Binayak (Fig. 1). A large population of J. recurva grew just west of Sing Gompa (Fig. 1) and other individual J. recurva plants were found between Sing Gompa and Cholan Pati lodge (Fig. 1). Plants morphologically intermediate between J. recurva and the hooked branchlets junipers were found around Cholan Pati between 3400 and 3600 m (Fig. 1). The small elevation change between Sing Gompa (3400 m) and Lauri Binayak (3800 m) appears sufficient to cause habitat differences that support typical J. recurva and the hookedtip juniper, as well as intermediate habitats around Cholan Pati. Figure 1. Site map of juniper collections between Sing Gompa and Lauri Binayak, Nepal.
5 Phytologia (December 2009) 91(3) 365 The hooked branchlet junipers near Lauri Binayak are recognized as a new variety of J. recurva: Juniperus recurva var. uncinata R. P. Adams, var. nov. TYPE: Nepal, 100 m s of Lauri Binayak, N ', E ', 3900 m, on south facing slope, 0.5 m x 0.5 m shrub, 1 Nov 1993, Adams 7212 (HOLOTYPE: BAYLU, TOPOTYPES: Adams 7213, 7214, BAYLU), Fig. 2). Junipero recurvae similis sed differt habitu fruticoso, apicibus ramulorum unciformibus vel recurvatis, et foliis brevibus (20 28 mm). Similar to Juniperus recurva, but differing in that the branchlet tips are hooked or recurved, the leaves are short (20-28 mm) and it grows as a shrub. The terminal branchlet tips of var. uncinata are striking as they form a hook at their tips (Fig. 2), in contrast to var. recurva that has lax tips that are not hooked (Fig. 3). The leaves of uncinata are mm long, compared to mm in var. recurva (Figs. 2, 3). In addition, var. uncinata is multi-stemmed at the base whereas var. recurva has a strong central axis, even when appearing as a shrub. It appears that the two varieties hybridize in the area around Cholan Pati. A putative hybrid (Adams 7211) is shown in Fig. 4. Note that the hooked branchlets are like those of var. uncinata, but the longer leaves are like var. recurva; this plant was a shrub, branched at the base, 1 m wide x 1 m tall. Due to the cutting of limbs for use as incense, many of the trees in this area appear as shrub-like. In addition to hybrids, individuals were found that appear to be back-crossed, or of a F 2 generation. Adams 7210 is from a shrub, 4 m wide x 0.7 m tall, with very small leaves, but the branchlet tips are scarcely hooked (Fig. 5), suggesting the presence of some genes from J. recurva var. recurva in an otherwise typical J. r. var. uncinata individual.
6 366 Phytologia (December 2009) 91(3) Fig. 2. Juniperus recurva var. uncinata, Holotype, Adams Fig. 3. Juniperus recurva var. recurva. Adams 7219.
7 Phytologia (December 2009) 91(3) 367 Fig. 4. Putative hybrid between Juniperus recurva var. recurva and J. r. var. uncinata, Adams Fig. 5. Putative backcross of J. recurva var. recurva to J. r. var. uncinata, Adams 7210.
8 368 Phytologia (December 2009) 91(3) To gather additional information on the relationships within the J. recurva complex and other related taxa, the nrdna (ITS region) was sequenced. This resulted in 1106 to 1141 bp of sequence data, with 19 mutational events that included 5 indels. A 29 bp deletion was found in all samples of J. recurva vars. recurva and uncinata. A 4 bp deletion was found in all var. uncinata and J. squamata samples. Eight of the mutational events were single occurrences and not deemed of taxonomic interest. This resulted in 11 events (simply noted as SNPs for discussion) that were used for analysis. An associational matrix was constructed using these 11 SNPs; the resulting minimum spanning network is shown in figure 6 (left). No variation was found among J. recurva var. recurva and only one SNP was found in J. r. var. uncinata Figure 6. Left: Minimum spanning network based on 11 nrdna SNPs. Right: Minimum spanning network based on 24 petn-psbm SNPs. The numbers next to the links are the number of SNPs. Dashed line ( ) is the second nearest link.
9 Phytologia (December 2009) 91(3) 369 (Fig. 6, left). No variation was found among the three J. squamata samples. However, the three varieties of J. indica showed considerable differences. Sequencing the cp petn-spacer-psbm region resulted in bp with 35 mutational events that included 9 indels. Five of the indels and 6 of the nucleotide mutations were single occurrence events (autapomorphies) and eliminated, leaving 24 informative mutational events that were utilized (herein called SNPs). An associational matrix was constructed using these 24 SNPs, and the resulting minimum spanning network is shown in figure 6 (right). Interestingly, again, no variation was found within J. recurva var. uncinata or among J. r. var. recurva (Fig. 6, right). However, the three J. squamata samples showed considerable variation. The three varieties of J. indica, as before, showed considerable differences, with J. recurva var. recurva being placed in the midst of the three J. indica varieties (Fig. 6, right). The intermediate plants (7210, 7211, 7218) are plotted differently in each data set. Plant 7210 has 3 nrdna mutations that separate it from all J. recurva plants (Fig. 6, left), but its cp petn-psbm sequences are identical to J. recurva var. uncinata (Fig. 6, right) suggesting that the male (pollen and cp transmitting) parent was J. r. var. uncinata. Plant 7211 is identical in its nrdna to two of the J. recurva var. uncinata plants (Fig. 6, left), whereas, its cp petn-psbm sequence is identical to J. recurva var. recurva plants (Fig. 6, right), suggesting that the male parent was J. r. var. recurva. If this is correct, then gene flow seems to be bi-directional. Plant 7218 is intermediate in its nrdna between J. r. var. recurva and var. uncinata (Fig. 6, left) and has 4 SNPs separating it from J. r. var. recurva (Fig. 6, right). A minimum spanning network based on combining the 11 nrdna and 24 petn-psbm SNPs shows that J. recurva var. recurva is well separated from J. r. var. uncinata (Fig. 7). The varieties of J. indica are nearly equidistant. The accessions of J. squamata show considerable differences. The intermediate individuals largely follow the cp petn-psbm differences in that each are associated with either J. r. var. recurva (7211, 7218) or J. r. var. uncinata (7210) (Fig. 7). Although J. r. var.
10 370 Phytologia (December 2009) 91(3) Figure 7. Minimum spanning network based on 11 nrdna and 24 cp petn-psbm SNPs. The dotted lines show the 2nd nearest link. Numbers next to the links are the number of SNP differences. uncinata appears to be as distinct from J. r. var. recurva as other species. However, due to the likely hybridization and intergradation, it seems unwise to recognize it at the specific level at this time.
11 Phytologia (December 2009) 91(3) 371 Figure 8. NJ tree based on nrdna and cp petn-psbm sequences. The numbers on the branches are bootstrap probabilities (1000 reps.). Another method to examine the sequence differences is by adding an outgroup (J. scopulorum, J. virginiana) to the data set and computing a NJ tree. Figure 8 shows a NJ tree based on combined nrdna and petn-psbm sequences. Notice, although the species group
12 372 Phytologia (December 2009) 91(3) into clades, these clades are generally not well supported. The intermediate plants grouped with J. r. var. recurva (7211) or J. r. var. uncinata (7210), reflecting the male chloroplast donor. Intermediate plant 7218 is not closely associated with any clade. There is weak support for J. recurva var. uncinata and J. squamata as a clade (Fig. 8, 47). Examination of the leaves of the taxa concerned, show (Fig. 9) that the leaves of J. recurva var. uncinata are very similar to those of J. recurva var. recurva and not like those of J. squamata. In addition, the gene-flow between var. recurva and var. uncinata suggest a very close relationship. It is also noteworthy that the leaves of J. indica and its varieties are scale-like and differ from the decurrent leaves of J. recurva var. recurva (Fig. 9) such that the placement of var. recurva within the J. indica clade is inconsistent with leaf morphology. Perhaps there are ancestral gene-patterns of J. indica that are still expressed in J. recurva var. recurva for the two gene regions sequenced. Incomplete lineage sorting and gene coalescence Degnan and Rosenberg (2009) define incomplete lineage sorting as "the failure of two or more lineages in a population to coalesce, leading to the possibility that at least one of the lineages first coalesces with a lineage from a less closely related population." Syring et al. (2007, Table 6) suggest that the number of years for monophyly to be more likely than paraphyly in Pinus may be from 1.7 to 24 Myr and it may take from 5.4 to 76 Myr for a species to attain a complete genomewide coalescence. The oldest known fossil of Juniperus is from Europe (dated at 35 m yr bp, Kvacek, 2002). The Himalayas began to form with the collision of the Indian and Eurasian plates about m yr bp (USGS, 2009). It may be that J. indica and J. recurva have had insufficient evolutionary time to fully complete lineage sorting (at least for the two gene regions sequenced in this study). Leaf terpenoids and analysis of hybridization A more detailed examination of J. r. var. recurva, J. r. var. uncinata and putative hybrids was undertaken by examining the volatile leaf terpenoids. Table 1 shows the compositions of the oils of J. recurva from the study area (Fig. 1). The oils of var. recurva are high in α- pinene, sabinene, δ-3-carene, limonene, β-phellandrene and elemol
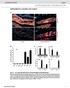
13 Phytologia (December 2009) 91(3) 373 Figure 9. Leaves and female cones of taxa in this study. with moderate amounts of myrcene, terpinolene, terpinen-4-ol, pregeijerene B, γ-eudesmol and abietadiene (Table 1).
14 374 Phytologia (December 2009) 91(3) The leaf oils of var. uncinata are very high in δ-3-carene ( %) and high in terpinolene, trans-muurola-4(14),5-diene, cubebol, δ- cadinene, with moderate amounts of α-pinene, sabinene, limonene, β- phellandrene, cis-muurola-3,5-diene, and trans-cadina-1(6),4-diene (Table 1). Several compounds were present in var. recurva, but absent in var. uncinata: α-thujene, cis-sabinene hydrate, trans-sabinene hydrate, hexenyl isovalerate, linalyl acetate, germacrene B, humulene epoxide II, and γ-eudesmol. Two compounds were found in all var. uncinata oils but were absent in var. recurva oils: cis-limonene oxide and β- oplopenone (Table 1). The intermediate plants (7210, 7211, 7218) often had intermediate values in their composition. Note α-thujene, α-pinene, δ-3- carene, terpinen-4-ol, cis-muurola-3,5-diene, trans-cadina-1(6),4-diene, trans-muurola-4(14),5-diene, δ-cadinene, elemol, γ-eudesmol, and abietadiene (Table 1). Two of the compounds exhibited a 'gigantism' in being markedly higher concentration in some of the intermediate plants: myrcene and β-phellandrene (Table 1). Thirty eight of the terpenoids that were larger than trace amounts, and displayed fidelity within the taxa (boldface in Table 1), were used to compute an association matrix. Factoring the matrix resulted in two eigenroots that were larger than the average diagonal value, and likely biologically significant. The first three eigenroots (47.3, 16.0, 10.3 % of the variance) were used to plot the individuals (Fig. 10). Juniperus recurva var. recurva individuals formed a looser cluster than those of var. uncinata. Two of the intermediate plants (7211, 7218) form the classical U triangle between parents and hybrids, exactly the ordination position that is expected for hybrids (see Adams, 1982, for a detailed examination of multivariate analysis of both synthetic and putative hybrids). The third intermediate plant (7210) is much closer to J. r. var. uncinata and behaves as a back-cross or F 2 generation plant in this ordination (Adams, 1982). It is interesting to note that none of the analyses of sequence data had the sensitivity to detect these hybrids, and/or back-crossed, individuals.
15 Phytologia (December 2009) 91(3) 375 Pleines, Jakob and Blattner (2008) state "As tree-based methods are mostly insufficient to depict relationships within species, network approaches are better suitable to infer gene or locus genealogies. Problematic for phylogeographic studies are alleles shared among multiple species, which could result from either hybridization of incomplete lineage sorting." The results from the present study show that analysis of hybridization and introgression using multivariate statistical methods of quantitative data is more informative than the treebased method utilized. Figure 10. PCO of J. recurva individuals based on 38 terpenoids. The dashed line is the minimum spanning network and the values next to the links are the distance values between OTUs. In summary, the recognition of the new variety, J. recurva var. uncinata, is supported by morphology, terpenoids and DNA sequence data. Additional research is needed to clarify the variation found in J. squamata.
16 376 Phytologia (December 2009) 91(3) ACKNOWLEDGEMENTS Thanks to Guy Nesom for the Latin description and to Andrea Schwarzbach, Randall Terry and Billie Turner for manuscript reviews. Thanks to Tonya Yanke for lab assistance. This research was supported in part with funds from Baylor University. LITERATURE CITED Adams, R. P Statistical character weighting and similarity stability. Brittonia 27: Adams, R. P A comparison of multivariate methods for the detection of hybridization. Taxon 31: Adams, R. P Systematics of smooth leaf margin Juniperus of the western hemisphere based on leaf essential oils and RAPD DNA fingerprinting. Biochem. Syst. Ecol. 28: Adams, R. P Junipers of the World: The genus Juniperus. 2nd Ed., Trafford Publ., Vancouver, B. C. Adams, R. P., B. R. Ruiz, M. Nogales and S. S. Fontinha Geographic variation and systematics of Juniperus phoenicea L. from Madeira and the Canary Islands: Analyses of volatile leaf oils. Phytologia 91(1): Blattner, F. R Direct amplification of the entire ITS region from poorly preserved plant material using recombinant PCR. BioTechniques 27: Degnan, J. H. and N. A. Rosenberg Gene tree discordance, phylogenetic inference and the multispecies coalescent. Review. Trends in Ecol. and Evol. (in press). Gower, J. C Some distance properties of latent root and vector methods used in multivariate analysis. Biometrika 53: Gower, J. C A general coefficient of similarity and some of its properties. Biometrics 27: Kvacek, Z A new juniper from the Palaeogene of central Europe. Feddes Repert. 111: 7-8. Pleines, T., S. S. Jakob and F. R. Blattner Application of noncoding DNA regions in infraspecific analyses. Plant Syst. Evol. Online Review, 1 Aug 2008,
17 Phytologia (December 2009) 91(3) 377 Syring, J., K. Farrell, R. Businsky, R. Cronn and A. Liston Widespread genealogical nonmonophyly in species of Pinus subgenus strobus. Syst. Biol. 56: USGS The Himalayas: Two Continents Collide. Veldman D. J., Fortran programming for the behavioral sciences. Holt, Rinehart and Winston Publ., NY.
18 378 Phytologia (December 2009) 91(3) Table 1. Comparison of volatile leaf oils of J. recurva var. recurva, J. r. var. uncinata and intermediates. var. recurva intermediate plants var. uncinata AI Compound tricyclene t t - - t α-thujene t α-pinene α-fenchene verbenene sabinene β-pinene t 0.1 t myrcene δ-3-carene α-terpinene t t 1020 p-cymene t t t limonene β-phellandrene γ-terpinene t t t 1065 cis-sabinene hydrate t t cis-linalool oxide(furanoid) t terpinolene ,7-epoxy myrcene t - - t trans-sabinene hydrate t t t linalool t t t
19 Phytologia (December 2009) 91(3) 379 Table 1. Continued isopentyl-isovalerate methyl-3-butenmethyl butanoate t t trans-p-menth-2-en-1-ol t t t t cis-limonene oxide t t cis-p-menth-2-en-1-ol t t t trans-limonene oxide t t 0.1 t t p-mentha-1,5-dien-8-ol t t t 0.2 t t t 1174 terpinen-4-ol t t 1179 p-cymen-8-ol 0.3 t t t - t p-cymen-9-ol t t - t α-terpineol verbenone t citronellol t trans-chrysanthenyl acetate hexenyl isovalerate carvacrol, methyl ether - - t t - t piperitone t t linalyl acetate methyl citronellate t pregeijerene B t t bornyl acetate t
20 380 Phytologia (December 2009) 91(3) Table 1. Continued trans-linalool oxide acetate (pyranoid) t t 1298 carvacrol t trans-carvyl acetate t α-cubebene α-terpinyl acetate α-copaene β-cubebene methyl eugenol t (E)-caryophyllene t t cis-muurola-3,5-diene α-humulene - t t t cis-cadina-1(6),4-diene t - t 1475 trans-cadina-1(6),4-diene t γ-muurolene t t t 0.1 t t t t t 1480 germacrene D t - t 1493 trans-muurola-4(14),5-diene epi-cubebol α-muurolene γ-cadinene cubebol δ-cadinene
21 Phytologia (December 2009) 91(3) 381 Table 1. Continued zonarene trans-cadina-1,4-diene α-cadinene t t elemol t cis-murol-5-en-5-α-ol t 0.1 t t 1559 germacrene B germacrene-d-4-ol caryophyllene oxide t - t 0.1 t t t trans-muurol-5-en-4-α-ol t β -oplopenone t humulene epoxide II t ,10-di-epi-cubenol t epi-cubenol γ-eudesmol t epi-α-cadinol epi-α-muurolol α-muurolol t t t 0.3 t t β -eudesmol t t α-eudesmol α -cadinol bulnesol t t - -
22 382 Phytologia (December 2009) 91(3) Table 1. Continued shyobunol t α-acetoxyelemol t t abietatriene t manool abietadiene t t 2282 sempervirol t epi-abietal t trans-totarol t trans-ferruginol t
THE TAXNOMIC AFFINITY OF A JUNIPER POPULATION FROM COLONIA PACHECO, MEXICO
 132 Phytologia (April 2011) 93(1) THE TAXNOMIC AFFINITY OF A JUNIPER POPULATION FROM COLONIA PACHECO, MEXICO Robert P. Adams Biology Department, Baylor University, Box 97388, Waco, TX 76798, USA, email
132 Phytologia (April 2011) 93(1) THE TAXNOMIC AFFINITY OF A JUNIPER POPULATION FROM COLONIA PACHECO, MEXICO Robert P. Adams Biology Department, Baylor University, Box 97388, Waco, TX 76798, USA, email
Robert P. Adams Biology Department, Baylor University Waco, TX, 76798, USA
 Phytologia (December 2010) 92(3) 413 DISCOVERY OF A NEW POPULATION OF JUNIPERUS GRACILIOR VAR. URBANIANA FROM THE DOMINICAN REPUBLIC: ANALYSES OF LEAF TERPENOIDS AND SNPS FROM nrdna AND trnc-trnd Robert
Phytologia (December 2010) 92(3) 413 DISCOVERY OF A NEW POPULATION OF JUNIPERUS GRACILIOR VAR. URBANIANA FROM THE DOMINICAN REPUBLIC: ANALYSES OF LEAF TERPENOIDS AND SNPS FROM nrdna AND trnc-trnd Robert
The volatile leaf oil of Juniperus microsperma and its taxonomy
 Phytologia (February 2013) 95(1) 87 The volatile leaf oil of Juniperus microsperma and its taxonomy Robert P. Adams Biology Department, Baylor University, Box 727, Gruver, TX 79040, USA email Robert_Adams@baylor.edu
Phytologia (February 2013) 95(1) 87 The volatile leaf oil of Juniperus microsperma and its taxonomy Robert P. Adams Biology Department, Baylor University, Box 727, Gruver, TX 79040, USA email Robert_Adams@baylor.edu
Hybridization between Juniperus grandis, J. occidentalis and J. osteosperma in northwest Nevada I: Terpenes, Leviathan Mine, Nevada
 58 Phytologia (February 2013) 95(1) Hybridization between Juniperus grandis, J. occidentalis and J. osteosperma in northwest Nevada I: Terpenes, Leviathan Mine, Nevada Robert P. Adams Biology Department,
58 Phytologia (February 2013) 95(1) Hybridization between Juniperus grandis, J. occidentalis and J. osteosperma in northwest Nevada I: Terpenes, Leviathan Mine, Nevada Robert P. Adams Biology Department,
Leaf essential oils of Juniperus in central and southern Iran
 288 Phytologia (November 2013) 95(4) Leaf essential oils of Juniperus in central and southern Iran Robert P. Adams Biology Department, Baylor University, Box 97388,Waco, TX 76798, USA, Robert_Adams@baylor.edu
288 Phytologia (November 2013) 95(4) Leaf essential oils of Juniperus in central and southern Iran Robert P. Adams Biology Department, Baylor University, Box 97388,Waco, TX 76798, USA, Robert_Adams@baylor.edu
DNA FROM HERBARIUM SPECIMENS: I. CORRELATION OF DNA SIZE WITH SPECIMEN AGE
 346 Phytologia (December 2010) 92(3) DNA FROM HERBARIUM SPECIMENS: I. CORRELATION OF DNA SIZE WITH SPECIMEN AGE Robert P. Adams Biology Department, Baylor University, Box 97388, Waco, TX 76798, USA, email
346 Phytologia (December 2010) 92(3) DNA FROM HERBARIUM SPECIMENS: I. CORRELATION OF DNA SIZE WITH SPECIMEN AGE Robert P. Adams Biology Department, Baylor University, Box 97388, Waco, TX 76798, USA, email
JUNIPERUS COMPACTA (CUPRESSACEAE) A NEW SPECIES FROM MEXICO
 Phytologia (December 2007) 89(3) 361 JUNIPERUS COMPACTA (CUPRESSACEAE) A NEW SPECIES FROM MEXICO Robert P. Adams Biology Department, Baylor University, Waco, TX 76798, USA Robert_Adams@baylor.edu Andrea
Phytologia (December 2007) 89(3) 361 JUNIPERUS COMPACTA (CUPRESSACEAE) A NEW SPECIES FROM MEXICO Robert P. Adams Biology Department, Baylor University, Waco, TX 76798, USA Robert_Adams@baylor.edu Andrea
DNA FROM HERBARIUM SPECIMENS: II. CORRELATION OF DNA DEGRADATION WITH HUMIDITY
 Phytologia (December 2011) 93(3) 351 DNA FROM HERBARIUM SPECIMENS: II. CORRELATION OF DNA DEGRADATION WITH HUMIDITY Robert P. Adams Biology Department, Baylor University, Box 97388, Waco, TX 76798, USA,
Phytologia (December 2011) 93(3) 351 DNA FROM HERBARIUM SPECIMENS: II. CORRELATION OF DNA DEGRADATION WITH HUMIDITY Robert P. Adams Biology Department, Baylor University, Box 97388, Waco, TX 76798, USA,
Inheritance of nrdna in artificial hybrids of Hesperocyparis arizonica x H. macrocarpa
 Phytologia (Oct 6, 2016) 98(4) 277 Inheritance of nrdna in artificial hybrids of Hesperocyparis arizonica x H. macrocarpa Robert P. Adams Biology Department, Baylor University, Box 97388, Waco, TX 76798,
Phytologia (Oct 6, 2016) 98(4) 277 Inheritance of nrdna in artificial hybrids of Hesperocyparis arizonica x H. macrocarpa Robert P. Adams Biology Department, Baylor University, Box 97388, Waco, TX 76798,
Geographic variation in J. phoenicea var. phoenicea from throughout its range: Analysis of nrdna and the petn-psbm cp region.
 Phytologia (Oct 1, 2014) 96(4) 247 Geographic variation in J. phoenicea var. phoenicea from throughout its range: Analysis of nrdna and the petn-psbm cp region. Robert P. Adams Biology Department, Baylor
Phytologia (Oct 1, 2014) 96(4) 247 Geographic variation in J. phoenicea var. phoenicea from throughout its range: Analysis of nrdna and the petn-psbm cp region. Robert P. Adams Biology Department, Baylor
LOW VARIABILITY OF DNA FINGERPRINTS OF TEXAS SNOWBELLS: CONSERVATION IMPLICATIONS
 198 LOW VARIABILITY OF DNA FINGERPRINTS OF TEXAS SNOWBELLS: CONSERVATION IMPLICATIONS Robert P. Adams Biology Department, Baylor University, Waco, TX 76706 Robert_Adams@baylor.edu and Jackie Poole Wildlife
198 LOW VARIABILITY OF DNA FINGERPRINTS OF TEXAS SNOWBELLS: CONSERVATION IMPLICATIONS Robert P. Adams Biology Department, Baylor University, Waco, TX 76706 Robert_Adams@baylor.edu and Jackie Poole Wildlife
VARIATION IN nrdna AND cpdna OF JUNIPERUS CALIFORNICA, J. GRANDIS, J. OCCIDENTALIS AND J. OSTEOSPERMA (CUPRESSACEAE)
 266 VARIATION IN nrdna AND cpdna OF JUNIPERUS CALIFORNICA, J. GRANDIS, J. OCCIDENTALIS AND J. OSTEOSPERMA (CUPRESSACEAE) Robert P. Adams Biology Department, Baylor University, Box 97388,Waco, TX 76798,
266 VARIATION IN nrdna AND cpdna OF JUNIPERUS CALIFORNICA, J. GRANDIS, J. OCCIDENTALIS AND J. OSTEOSPERMA (CUPRESSACEAE) Robert P. Adams Biology Department, Baylor University, Box 97388,Waco, TX 76798,
GEOGRAPHIC VARIATION IN THE LEAF ESSENTIAL OILS OF HESPEROCYPARIS ARIZONICA AND H. GLABRA
 366 Phytologia (December 2010) 92(3) GEOGRAPHIC VARIATION IN THE LEAF ESSENTIAL OILS OF HESPEROCYPARIS ARIZONICA AND H. GLABRA Robert P. Adams Biology Department, Baylor University, Box 97388, Waco, TX
366 Phytologia (December 2010) 92(3) GEOGRAPHIC VARIATION IN THE LEAF ESSENTIAL OILS OF HESPEROCYPARIS ARIZONICA AND H. GLABRA Robert P. Adams Biology Department, Baylor University, Box 97388, Waco, TX
Geographical variation in the leaf volatile oils of Grindelia ciliata and G. adenodonta (Asteraceae)
 112 Phytologia (Apr 4, 2016) 98(2) Geographical variation in the leaf volatile oils of Grindelia ciliata and G. adenodonta (Asteraceae) Robert P. Adams Biology Department, Baylor University, Waco, TX 76798,
112 Phytologia (Apr 4, 2016) 98(2) Geographical variation in the leaf volatile oils of Grindelia ciliata and G. adenodonta (Asteraceae) Robert P. Adams Biology Department, Baylor University, Waco, TX 76798,
Inheritance of nrdna in artificial hybrids of Cryptomeria japonica cv. Haara and C. japonica cv. Kumotooshi
 Phytologia (Jan 5, 2016) 98(1) 37 Inheritance of nrdna in artificial hybrids of Cryptomeria japonica cv. Haara and C. japonica cv. Kumotooshi Robert P. Adams Biology Department, Baylor University, Box
Phytologia (Jan 5, 2016) 98(1) 37 Inheritance of nrdna in artificial hybrids of Cryptomeria japonica cv. Haara and C. japonica cv. Kumotooshi Robert P. Adams Biology Department, Baylor University, Box
DNA BARCODING A JUNIPER: THE CASE OF THE SOUTH TEXAS DUVAL COUNTY JUNIPER AND SERRATE JUNIPERS OF NORTH AMERICA
 146 DNA BARCODING A JUNIPER: THE CASE OF THE SOUTH TEXAS DUVAL COUNTY JUNIPER AND SERRATE JUNIPERS OF NORTH AMERICA Robert P. Adams Biology Department, Baylor University, Box 97388,Waco, TX 76798, USA,
146 DNA BARCODING A JUNIPER: THE CASE OF THE SOUTH TEXAS DUVAL COUNTY JUNIPER AND SERRATE JUNIPERS OF NORTH AMERICA Robert P. Adams Biology Department, Baylor University, Box 97388,Waco, TX 76798, USA,
Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH).
 Bisulfite Treatment of DNA Dilute DNA sample to 2µg DNA in 50µl ddh 2 O. Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH). Incubate in a 37ºC water bath for 30 minutes. To 55µl samples
Bisulfite Treatment of DNA Dilute DNA sample to 2µg DNA in 50µl ddh 2 O. Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH). Incubate in a 37ºC water bath for 30 minutes. To 55µl samples
Y-chromosomal haplogroup typing Using SBE reaction
 Schematic of multiplex PCR followed by SBE reaction Multiplex PCR Exo SAP purification SBE reaction 5 A 3 ddatp ddgtp 3 T 5 A G 3 T 5 3 5 G C 5 3 3 C 5 ddttp ddctp 5 T 3 T C 3 A 5 3 A 5 5 C 3 3 G 5 3 G
Schematic of multiplex PCR followed by SBE reaction Multiplex PCR Exo SAP purification SBE reaction 5 A 3 ddatp ddgtp 3 T 5 A G 3 T 5 3 5 G C 5 3 3 C 5 ddttp ddctp 5 T 3 T C 3 A 5 3 A 5 5 C 3 3 G 5 3 G
Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC
 Supplementary Appendixes Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC ACG TAG CTC CGG CTG GA-3 for vimentin, /5AmMC6/TCC CTC GCG CGT GGC TTC CGC
Supplementary Appendixes Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC ACG TAG CTC CGG CTG GA-3 for vimentin, /5AmMC6/TCC CTC GCG CGT GGC TTC CGC
Arabidopsis actin depolymerizing factor AtADF4 mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB
 Arabidopsis actin depolymerizing factor mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB Files in this Data Supplement: Supplemental Table S1 Supplemental Table
Arabidopsis actin depolymerizing factor mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB Files in this Data Supplement: Supplemental Table S1 Supplemental Table
Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR
 Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR 1 The problem We wish to clone a yet unknown gene from a known
Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR 1 The problem We wish to clone a yet unknown gene from a known
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
Supplemental Data Supplemental Figure 1.
 Supplemental Data Supplemental Figure 1. Silique arrangement in the wild-type, jhs, and complemented lines. Wild-type (WT) (A), the jhs1 mutant (B,C), and the jhs1 mutant complemented with JHS1 (Com) (D)
Supplemental Data Supplemental Figure 1. Silique arrangement in the wild-type, jhs, and complemented lines. Wild-type (WT) (A), the jhs1 mutant (B,C), and the jhs1 mutant complemented with JHS1 (Com) (D)
Cat. # Product Size DS130 DynaExpress TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1
 Product Name: Kit Component TA PCR Cloning Kit (ptakn-2) Cat. # Product Size DS130 TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1 2 Ligation Buffer
Product Name: Kit Component TA PCR Cloning Kit (ptakn-2) Cat. # Product Size DS130 TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1 2 Ligation Buffer
Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis
 1 2 3 4 5 6 7 8 9 10 11 12 Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis Information Research). Exons
1 2 3 4 5 6 7 8 9 10 11 12 Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis Information Research). Exons
Gene synthesis by circular assembly amplification
 Gene synthesis by circular assembly amplification Duhee Bang & George M Church Supplementary figures and text: Supplementary Figure 1. Dpo4 gene (1.05kb) construction by various methods. Supplementary
Gene synthesis by circular assembly amplification Duhee Bang & George M Church Supplementary figures and text: Supplementary Figure 1. Dpo4 gene (1.05kb) construction by various methods. Supplementary
Multiple evidences of past evolution are hidden in nrdna of Juniperus arizonica and J. coahuilensis populations in the trans-pecos, Texas region
 38 Phytologia (Jan 19, 2017) 99(1) Multiple evidences of past evolution are hidden in nrdna of Juniperus arizonica and J. coahuilensis populations in the trans-pecos, Texas region Robert P. Adams Biology
38 Phytologia (Jan 19, 2017) 99(1) Multiple evidences of past evolution are hidden in nrdna of Juniperus arizonica and J. coahuilensis populations in the trans-pecos, Texas region Robert P. Adams Biology
Sequencing of DNA lesions facilitated by site-specific excision via base. excision repair DNA glycosylases yielding ligatable gaps
 Supporting information Sequencing of DNA lesions facilitated by site-specific excision via base excision repair DNA glycosylases yielding ligatable gaps Jan Riedl, Aaron M. Fleming, and Cynthia J. Burrows*
Supporting information Sequencing of DNA lesions facilitated by site-specific excision via base excision repair DNA glycosylases yielding ligatable gaps Jan Riedl, Aaron M. Fleming, and Cynthia J. Burrows*
Supporting Information. Copyright Wiley-VCH Verlag GmbH & Co. KGaA, Weinheim, 2006
 Supporting Information Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Supporting Information for Expanding the Genetic
Supporting Information Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Supporting Information for Expanding the Genetic
Disease and selection in the human genome 3
 Disease and selection in the human genome 3 Ka/Ks revisited Please sit in row K or forward RBFD: human populations, adaptation and immunity Neandertal Museum, Mettman Germany Sequence genome Measure expression
Disease and selection in the human genome 3 Ka/Ks revisited Please sit in row K or forward RBFD: human populations, adaptation and immunity Neandertal Museum, Mettman Germany Sequence genome Measure expression
strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular
![strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular](/thumbs/72/66361212.jpg) Additional file 2 Identification of AOX1 in P. pastoris GS115 with a Mut s phenotype Results and Discussion The HBsAg producing strain was originally identified as a Mut s (methanol utilization slow) strain
Additional file 2 Identification of AOX1 in P. pastoris GS115 with a Mut s phenotype Results and Discussion The HBsAg producing strain was originally identified as a Mut s (methanol utilization slow) strain
Supplementary Figure 1A A404 Cells +/- Retinoic Acid
 Supplementary Figure 1A A44 Cells +/- Retinoic Acid 1 1 H3 Lys4 di-methylation SM-actin VEC cfos (-) RA (+) RA 14 1 1 8 6 4 H3 Lys79 di-methylation SM-actin VEC cfos (-) RA (+) RA Supplementary Figure
Supplementary Figure 1A A44 Cells +/- Retinoic Acid 1 1 H3 Lys4 di-methylation SM-actin VEC cfos (-) RA (+) RA 14 1 1 8 6 4 H3 Lys79 di-methylation SM-actin VEC cfos (-) RA (+) RA Supplementary Figure
RPA-AB RPA-C Supplemental Figure S1: SDS-PAGE stained with Coomassie Blue after protein purification.
 RPA-AB RPA-C (a) (b) (c) (d) (e) (f) Supplemental Figure S: SDS-PAGE stained with Coomassie Blue after protein purification. (a) RPA; (b) RPA-AB; (c) RPA-CDE; (d) RPA-CDE core; (e) RPA-DE; and (f) RPA-C
RPA-AB RPA-C (a) (b) (c) (d) (e) (f) Supplemental Figure S: SDS-PAGE stained with Coomassie Blue after protein purification. (a) RPA; (b) RPA-AB; (c) RPA-CDE; (d) RPA-CDE core; (e) RPA-DE; and (f) RPA-C
MacBlunt PCR Cloning Kit Manual
 MacBlunt PCR Cloning Kit Manual Shipping and Storage MacBlunt PCR Cloning Kits are shipped on dry ice. Each kit contains a box with cloning reagents and an attached bag with Eco-Blue Competent Cells (optional).
MacBlunt PCR Cloning Kit Manual Shipping and Storage MacBlunt PCR Cloning Kits are shipped on dry ice. Each kit contains a box with cloning reagents and an attached bag with Eco-Blue Competent Cells (optional).
Supplemental Data. mir156-regulated SPL Transcription. Factors Define an Endogenous Flowering. Pathway in Arabidopsis thaliana
 Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
Supporting Online Information
 Supporting Online Information Isolation of Human Genomic DNA Sequences with Expanded Nucleobase Selectivity Preeti Rathi, Sara Maurer, Grzegorz Kubik and Daniel Summerer* Department of Chemistry and Chemical
Supporting Online Information Isolation of Human Genomic DNA Sequences with Expanded Nucleobase Selectivity Preeti Rathi, Sara Maurer, Grzegorz Kubik and Daniel Summerer* Department of Chemistry and Chemical
Legends for supplementary figures 1-3
 High throughput resistance profiling of Plasmodium falciparum infections based on custom dual indexing and Illumina next generation sequencing-technology Sidsel Nag 1,2 *, Marlene D. Dalgaard 3, Poul-Erik
High throughput resistance profiling of Plasmodium falciparum infections based on custom dual indexing and Illumina next generation sequencing-technology Sidsel Nag 1,2 *, Marlene D. Dalgaard 3, Poul-Erik
Engineering D66N mutant using quick change site directed mutagenesis. Harkewal Singh 09/01/2010
 Engineering D66N mutant using quick change site directed mutagenesis Harkewal Singh 09/01/2010 1 1- What is quick change site directed mutagenesis? 2- An overview of the kit contents. 3- A brief information
Engineering D66N mutant using quick change site directed mutagenesis Harkewal Singh 09/01/2010 1 1- What is quick change site directed mutagenesis? 2- An overview of the kit contents. 3- A brief information
ORFs and genes. Please sit in row K or forward
 ORFs and genes Please sit in row K or forward https://www.flickr.com/photos/teseum/3231682806/in/photostream/ Question: why do some strains of Vibrio cause cholera and others don t? Methods Mechanisms
ORFs and genes Please sit in row K or forward https://www.flickr.com/photos/teseum/3231682806/in/photostream/ Question: why do some strains of Vibrio cause cholera and others don t? Methods Mechanisms
SUPPLEMENTARY MATERIALS AND METHODS. E. coli strains, plasmids, and growth conditions. Escherichia coli strain P90C (1)
 SUPPLEMENTARY MATERIALS AND METHODS E. coli strains, plasmids, and growth conditions. Escherichia coli strain P90C (1) dinb::kan (lab stock) derivative was used as wild-type. MG1655 alka tag dinb (2) is
SUPPLEMENTARY MATERIALS AND METHODS E. coli strains, plasmids, and growth conditions. Escherichia coli strain P90C (1) dinb::kan (lab stock) derivative was used as wild-type. MG1655 alka tag dinb (2) is
Hes6. PPARα. PPARγ HNF4 CD36
 SUPPLEMENTARY INFORMATION Supplementary Table Positions and Sequences of ChIP primers -63 AGGTCACTGCCA -79 AGGTCTGCTGTG Hes6-0067 GGGCAaAGTTCA ACOT -395 GGGGCAgAGTTCA PPARα -309 GGCTCAaAGTTCAaGTTCA CPTa
SUPPLEMENTARY INFORMATION Supplementary Table Positions and Sequences of ChIP primers -63 AGGTCACTGCCA -79 AGGTCTGCTGTG Hes6-0067 GGGCAaAGTTCA ACOT -395 GGGGCAgAGTTCA PPARα -309 GGCTCAaAGTTCAaGTTCA CPTa
Supplemental Data. Bennett et al. (2010). Plant Cell /tpc
 BRN1 ---------MSSSNGGVPPGFRFHPTDEELLHYYLKKKISYEKFEMEVIKEVDLNKIEPWDLQDRCKIGSTPQNEWYFFSHKDRKYPTGS 81 BRN2 --------MGSSSNGGVPPGFRFHPTDEELLHYYLKKKISYQKFEMEVIREVDLNKLEPWDLQERCKIGSTPQNEWYFFSHKDRKYPTGS 82 SMB
BRN1 ---------MSSSNGGVPPGFRFHPTDEELLHYYLKKKISYEKFEMEVIKEVDLNKIEPWDLQDRCKIGSTPQNEWYFFSHKDRKYPTGS 81 BRN2 --------MGSSSNGGVPPGFRFHPTDEELLHYYLKKKISYQKFEMEVIREVDLNKLEPWDLQERCKIGSTPQNEWYFFSHKDRKYPTGS 82 SMB
PCR-based Markers and Cut Flower Longevity in Carnation
 PCRbased Markers and Cut Flower Longevity in Carnation Laura De Benedetti, Luca Braglia, Simona Bruna, Gianluca Burchi *, Antonio Mercuri and Tito Schiva Istituto Sperimentale per la Floricoltura, Corso
PCRbased Markers and Cut Flower Longevity in Carnation Laura De Benedetti, Luca Braglia, Simona Bruna, Gianluca Burchi *, Antonio Mercuri and Tito Schiva Istituto Sperimentale per la Floricoltura, Corso
PCR analysis was performed to show the presence and the integrity of the var1csa and var-
 Supplementary information: Methods: Table S1: Primer Name Nucleotide sequence (5-3 ) DBL3-F tcc ccg cgg agt gaa aca tca tgt gac tg DBL3-R gac tag ttt ctt tca ata aat cac tcg c DBL5-F cgc cct agg tgc ttc
Supplementary information: Methods: Table S1: Primer Name Nucleotide sequence (5-3 ) DBL3-F tcc ccg cgg agt gaa aca tca tgt gac tg DBL3-R gac tag ttt ctt tca ata aat cac tcg c DBL5-F cgc cct agg tgc ttc
Supporting information for Biochemistry, 1995, 34(34), , DOI: /bi00034a013
 Supporting information for Biochemistry, 1995, 34(34), 10807 10815, DOI: 10.1021/bi00034a013 LESNIK 10807-1081 Terms & Conditions Electronic Supporting Information files are available without a subscription
Supporting information for Biochemistry, 1995, 34(34), 10807 10815, DOI: 10.1021/bi00034a013 LESNIK 10807-1081 Terms & Conditions Electronic Supporting Information files are available without a subscription
Supplementary. Table 1: Oligonucleotides and Plasmids. complementary to positions from 77 of the SRα '- GCT CTA GAG AAC TTG AAG TAC AGA CTG C
 Supplementary Table 1: Oligonucleotides and Plasmids 913954 5'- GCT CTA GAG AAC TTG AAG TAC AGA CTG C 913955 5'- CCC AAG CTT ACA GTG TGG CCA TTC TGC TG 223396 5'- CGA CGC GTA CAG TGT GGC CAT TCT GCT G
Supplementary Table 1: Oligonucleotides and Plasmids 913954 5'- GCT CTA GAG AAC TTG AAG TAC AGA CTG C 913955 5'- CCC AAG CTT ACA GTG TGG CCA TTC TGC TG 223396 5'- CGA CGC GTA CAG TGT GGC CAT TCT GCT G
JUNIPERUS MARITIMA, THE SEASIDE JUNIPER, A NEW SPECIES FROM PUGET SOUND, NORTH AMERICA
 Phytologia (December 2007) 89(3) 263 JUNIPERUS MARITIMA, THE SEASIDE JUNIPER, A NEW SPECIES FROM PUGET SOUND, NORTH AMERICA Robert P. Adams Biology Department, Baylor University, Box 727, Gruver, TX 79040,
Phytologia (December 2007) 89(3) 263 JUNIPERUS MARITIMA, THE SEASIDE JUNIPER, A NEW SPECIES FROM PUGET SOUND, NORTH AMERICA Robert P. Adams Biology Department, Baylor University, Box 727, Gruver, TX 79040,
NESTED Sequence-based Typing (SBT) protocol for epidemiological typing of Legionella pneumophila directly from clinical samples
 NESTED Sequence-based Typing (SBT) protocol for epidemiological typing of Legionella pneumophila directly from clinical samples VERSION 2.0 SUMMARY This procedure describes the use of nested Sequence-Based
NESTED Sequence-based Typing (SBT) protocol for epidemiological typing of Legionella pneumophila directly from clinical samples VERSION 2.0 SUMMARY This procedure describes the use of nested Sequence-Based
II 0.95 DM2 (RPP1) DM3 (At3g61540) b
 Table S2. F 2 Segregation Ratios at 16 C, Related to Figure 2 Cross n c Phenotype Model e 2 Locus A Locus B Normal F 1 -like Enhanced d Uk-1/Uk-3 149 64 36 49 DM2 (RPP1) DM1 (SSI4) a Bla-1/Hh-0 F 3 111
Table S2. F 2 Segregation Ratios at 16 C, Related to Figure 2 Cross n c Phenotype Model e 2 Locus A Locus B Normal F 1 -like Enhanced d Uk-1/Uk-3 149 64 36 49 DM2 (RPP1) DM1 (SSI4) a Bla-1/Hh-0 F 3 111
PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells
 Supplementary Information for: PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells Ju Hye Jang 1, Hyun Kim 2, Mi Jung Jang 2, Ju Hyun Cho 1,2,* 1 Research Institute
Supplementary Information for: PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells Ju Hye Jang 1, Hyun Kim 2, Mi Jung Jang 2, Ju Hyun Cho 1,2,* 1 Research Institute
Project 07/111 Final Report October 31, Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines
 Project 07/111 Final Report October 31, 2007. Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines Project Leader: Dr Douglas C. Hodgins (519-824-4120 Ex 54758, fax 519-824-5930)
Project 07/111 Final Report October 31, 2007. Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines Project Leader: Dr Douglas C. Hodgins (519-824-4120 Ex 54758, fax 519-824-5930)
Materials Protein synthesis kit. This kit consists of 24 amino acids, 24 transfer RNAs, four messenger RNAs and one ribosome (see below).
 Protein Synthesis Instructions The purpose of today s lab is to: Understand how a cell manufactures proteins from amino acids, using information stored in the genetic code. Assemble models of four very
Protein Synthesis Instructions The purpose of today s lab is to: Understand how a cell manufactures proteins from amino acids, using information stored in the genetic code. Assemble models of four very
Supplementary Information. Construction of Lasso Peptide Fusion Proteins
 Supplementary Information Construction of Lasso Peptide Fusion Proteins Chuhan Zong 1, Mikhail O. Maksimov 2, A. James Link 2,3 * Departments of 1 Chemistry, 2 Chemical and Biological Engineering, and
Supplementary Information Construction of Lasso Peptide Fusion Proteins Chuhan Zong 1, Mikhail O. Maksimov 2, A. James Link 2,3 * Departments of 1 Chemistry, 2 Chemical and Biological Engineering, and
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/10/494/eaan6284/dc1 Supplementary Materials for Activation of master virulence regulator PhoP in acidic ph requires the Salmonella-specific protein UgtL Jeongjoon
www.sciencesignaling.org/cgi/content/full/10/494/eaan6284/dc1 Supplementary Materials for Activation of master virulence regulator PhoP in acidic ph requires the Salmonella-specific protein UgtL Jeongjoon
I. Things to keep in mind before you start this protocol
 Inverse PCR and Sequencing Protocol on 5 Fly Preps For recovery of sequences flanking piggybac elements This protocol is an adaptation of "Inverse PCR and Cycle Sequencing Protocols" by E. Jay Rehm Berkeley
Inverse PCR and Sequencing Protocol on 5 Fly Preps For recovery of sequences flanking piggybac elements This protocol is an adaptation of "Inverse PCR and Cycle Sequencing Protocols" by E. Jay Rehm Berkeley
Supplemental Information. Human Senataxin Resolves RNA/DNA Hybrids. Formed at Transcriptional Pause Sites. to Promote Xrn2-Dependent Termination
 Supplemental Information Molecular Cell, Volume 42 Human Senataxin Resolves RNA/DNA Hybrids Formed at Transcriptional Pause Sites to Promote Xrn2-Dependent Termination Konstantina Skourti-Stathaki, Nicholas
Supplemental Information Molecular Cell, Volume 42 Human Senataxin Resolves RNA/DNA Hybrids Formed at Transcriptional Pause Sites to Promote Xrn2-Dependent Termination Konstantina Skourti-Stathaki, Nicholas
evaluated with UAS CLB eliciting UAS CIT -N Libraries increase in the
 Supplementary Figures Supplementary Figure 1: Promoter scaffold library assemblies. Many ensembless of libraries were evaluated in this work. As a legend, the box outline color in top half of the figure
Supplementary Figures Supplementary Figure 1: Promoter scaffold library assemblies. Many ensembless of libraries were evaluated in this work. As a legend, the box outline color in top half of the figure
SUPPORTING INFORMATION
 SUPPORTING INFORMATION Investigation of the Biosynthesis of the Lasso Peptide Chaxapeptin Using an E. coli-based Production System Helena Martin-Gómez, Uwe Linne, Fernando Albericio, Judit Tulla-Puche,*
SUPPORTING INFORMATION Investigation of the Biosynthesis of the Lasso Peptide Chaxapeptin Using an E. coli-based Production System Helena Martin-Gómez, Uwe Linne, Fernando Albericio, Judit Tulla-Puche,*
A netlike rolling circle nucleic acid amplification technique
 Electronic Supplementary Material (ESI) for Analyst. This journal is The Royal Society of Chemistry 2014 Supplementary Information A netlike rolling circle nucleic acid amplification technique Xiaoli Zhu,
Electronic Supplementary Material (ESI) for Analyst. This journal is The Royal Society of Chemistry 2014 Supplementary Information A netlike rolling circle nucleic acid amplification technique Xiaoli Zhu,
VOLATILE CONSTITUENTS OF THE OILS FROM Povedadaphne quadriporata (LAURACEAE) FROM ALBERTO M. BRENES BIOLOGICAL PRESERVE, COSTA RICA
 Quim. Nova, Vol. 31, No. 3, 605-609, 2008 VOLATILE CONSTITUENTS OF THE OILS FROM Povedadaphne quadriporata (LAURACEAE) FROM ALBERTO M. BRENES BIOLOGICAL PRESERVE, COSTA RICA José F. Cicció* and Carlos
Quim. Nova, Vol. 31, No. 3, 605-609, 2008 VOLATILE CONSTITUENTS OF THE OILS FROM Povedadaphne quadriporata (LAURACEAE) FROM ALBERTO M. BRENES BIOLOGICAL PRESERVE, COSTA RICA José F. Cicció* and Carlos
SAY IT WITH DNA: Protein Synthesis Activity by Larry Flammer
 TEACHER S GUIDE SAY IT WITH DNA: Protein Synthesis Activity by Larry Flammer SYNOPSIS This activity uses the metaphor of decoding a secret message for the Protein Synthesis process. Students teach themselves
TEACHER S GUIDE SAY IT WITH DNA: Protein Synthesis Activity by Larry Flammer SYNOPSIS This activity uses the metaphor of decoding a secret message for the Protein Synthesis process. Students teach themselves
11th Meeting of the Science Working Group. Lima, Peru, October 2012 SWG-11-JM-11
 11th Meeting of the Science Working Group Lima, Peru, 15-19 October 2012 Russian population genetics studies of jack mackerel in the South Pacific P.K.Afanasiev M.A.Rabchun A.I.Glubokov Introduction. In
11th Meeting of the Science Working Group Lima, Peru, 15-19 October 2012 Russian population genetics studies of jack mackerel in the South Pacific P.K.Afanasiev M.A.Rabchun A.I.Glubokov Introduction. In
Supporting Information
 Supporting Information Table S1. Oligonucleotide sequences used in this work Oligo DNA A B C D CpG-A CpG-B CpG-C CpG-D Sequence 5 ACA TTC CTA AGT CTG AAA CAT TAC AGC TTG CTA CAC GAG AAG AGC CGC CAT AGT
Supporting Information Table S1. Oligonucleotide sequences used in this work Oligo DNA A B C D CpG-A CpG-B CpG-C CpG-D Sequence 5 ACA TTC CTA AGT CTG AAA CAT TAC AGC TTG CTA CAC GAG AAG AGC CGC CAT AGT
Lecture 11: Gene Prediction
 Lecture 11: Gene Prediction Study Chapter 6.11-6.14 1 Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and end positions of genes in a genome Where are
Lecture 11: Gene Prediction Study Chapter 6.11-6.14 1 Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and end positions of genes in a genome Where are
2
 1 2 3 4 5 6 7 Supplemental Table 1. Magnaporthe oryzae strains generated in this study. Strain background Genotype Strain name Description Guy-11 H1:RFP H1:RFP Strain expressing Histone H1- encoding gene
1 2 3 4 5 6 7 Supplemental Table 1. Magnaporthe oryzae strains generated in this study. Strain background Genotype Strain name Description Guy-11 H1:RFP H1:RFP Strain expressing Histone H1- encoding gene
Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of
 Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of the cell line) were immunostained for HA, acetylated
Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of the cell line) were immunostained for HA, acetylated
Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system
 Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system Dr. Tim Welsink Molecular Biology Transient Gene Expression OUTLINE Short
Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system Dr. Tim Welsink Molecular Biology Transient Gene Expression OUTLINE Short
Volatile components of Japanese cedar cultivars as repellents related to resistance to Cryptomeria bark borer
 J Wood Sci (2002) 48:51-55 9 The Japan Wood Research Society 2002 Mitsuyoshi Yatagai 9 Hiroshi Makihara 9 Kihachiro Oba Volatile components of Japanese cedar cultivars as repellents related to resistance
J Wood Sci (2002) 48:51-55 9 The Japan Wood Research Society 2002 Mitsuyoshi Yatagai 9 Hiroshi Makihara 9 Kihachiro Oba Volatile components of Japanese cedar cultivars as repellents related to resistance
Multiplexing Genome-scale Engineering
 Multiplexing Genome-scale Engineering Harris Wang, Ph.D. Department of Systems Biology Department of Pathology & Cell Biology http://wanglab.c2b2.columbia.edu Rise of Genomics An Expanding Toolbox Esvelt
Multiplexing Genome-scale Engineering Harris Wang, Ph.D. Department of Systems Biology Department of Pathology & Cell Biology http://wanglab.c2b2.columbia.edu Rise of Genomics An Expanding Toolbox Esvelt
BioInformatics and Computational Molecular Biology. Course Website
 BioInformatics and Computational Molecular Biology Course Website http://bioinformatics.uchc.edu What is Bioinformatics Bioinformatics upgrades the information content of biological measurements. Discovery
BioInformatics and Computational Molecular Biology Course Website http://bioinformatics.uchc.edu What is Bioinformatics Bioinformatics upgrades the information content of biological measurements. Discovery
OPTIMIZING PROCEDURES TO OBTAIN RELIABLE DNA FINGERPRINTING DATA FOR USE IN SYSTEMATIC, ECOLOGICAL AND EVOLUTIONARY STUDIES.
 Phytologia (April 2008) 90(1) 83 OPTIMIZING PROCEDURES TO OBTAIN RELIABLE DNA FINGERPRINTING DATA FOR USE IN SYSTEMATIC, ECOLOGICAL AND EVOLUTIONARY STUDIES. Robert P. Adams Baylor University, Biology
Phytologia (April 2008) 90(1) 83 OPTIMIZING PROCEDURES TO OBTAIN RELIABLE DNA FINGERPRINTING DATA FOR USE IN SYSTEMATIC, ECOLOGICAL AND EVOLUTIONARY STUDIES. Robert P. Adams Baylor University, Biology
Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were
 1 Supplemental methods 2 3 4 5 6 7 8 9 1 11 12 13 14 15 16 17 18 19 21 22 23 Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were monitored by quantitative reverse-transcription
1 Supplemental methods 2 3 4 5 6 7 8 9 1 11 12 13 14 15 16 17 18 19 21 22 23 Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were monitored by quantitative reverse-transcription
Appendix 1a. Microsatellite analysis of P1-hyg, P2-neo and their. Amplified cdna (base pairs) P1-hyg P2-neo All progeny A A
 Supplementary information Appendix 1a. Microsatellite analysis of P1-hyg, P2-neo and their double drug resistant progeny (ABI 377). Primer set a,b Amplified cdna (base pairs) P1-hyg P2-neo All progeny
Supplementary information Appendix 1a. Microsatellite analysis of P1-hyg, P2-neo and their double drug resistant progeny (ABI 377). Primer set a,b Amplified cdna (base pairs) P1-hyg P2-neo All progeny
Genomics and Gene Recognition Genes and Blue Genes
 Genomics and Gene Recognition Genes and Blue Genes November 1, 2004 Prokaryotic Gene Structure prokaryotes are simplest free-living organisms studying prokaryotes can give us a sense what is the minimum
Genomics and Gene Recognition Genes and Blue Genes November 1, 2004 Prokaryotic Gene Structure prokaryotes are simplest free-living organisms studying prokaryotes can give us a sense what is the minimum
Genomic Sequencing. Genomic Sequencing. Maj Gen (R) Suhaib Ahmed, HI (M)
 Maj Gen (R) Suhaib Ahmed, HI (M) The process of determining the sequence of an unknown DNA is called sequencing. There are many approaches for DNA sequencing. In the last couple of decades automated Sanger
Maj Gen (R) Suhaib Ahmed, HI (M) The process of determining the sequence of an unknown DNA is called sequencing. There are many approaches for DNA sequencing. In the last couple of decades automated Sanger
Genetic Diversity of Gold-Spotted Pond Frog, Rana chosenica in Korea
 Genetic Diversity of Gold-Spotted Pond Frog, Rana chosenica in Korea Mi-Sook Min, Sun Kyung Park, Kelly Lasater, Hae Jun Baek, Dae Sik Park2, and Hang Lee 4 Seoul National University College of Veterinary
Genetic Diversity of Gold-Spotted Pond Frog, Rana chosenica in Korea Mi-Sook Min, Sun Kyung Park, Kelly Lasater, Hae Jun Baek, Dae Sik Park2, and Hang Lee 4 Seoul National University College of Veterinary
Supplemental material
 Supplemental material Diversity of O-antigen repeat-unit structures can account for the substantial sequence variation of Wzx translocases Yaoqin Hong and Peter R. Reeves School of Molecular Bioscience,
Supplemental material Diversity of O-antigen repeat-unit structures can account for the substantial sequence variation of Wzx translocases Yaoqin Hong and Peter R. Reeves School of Molecular Bioscience,
An engineered tryptophan zipper-type peptide as a molecular recognition scaffold
 SUPPLEMENTARY MATERIAL An engineered tryptophan zipper-type peptide as a molecular recognition scaffold Zihao Cheng and Robert E. Campbell* Supplementary Methods Library construction for FRET-based screening
SUPPLEMENTARY MATERIAL An engineered tryptophan zipper-type peptide as a molecular recognition scaffold Zihao Cheng and Robert E. Campbell* Supplementary Methods Library construction for FRET-based screening
Overexpression Normal expression Overexpression Normal expression. 26 (21.1%) N (%) P-value a N (%)
 SUPPLEMENTARY TABLES Table S1. Alteration of ZNF322A protein expression levels in relation to clinicopathological parameters in 123 Asian and 74 Caucasian lung cancer patients. Asian patients Caucasian
SUPPLEMENTARY TABLES Table S1. Alteration of ZNF322A protein expression levels in relation to clinicopathological parameters in 123 Asian and 74 Caucasian lung cancer patients. Asian patients Caucasian
Primer Design Workshop. École d'été en géné-que des champignons 2012 Dr. Will Hintz University of Victoria
 Primer Design Workshop École d'été en géné-que des champignons 2012 Dr. Will Hintz University of Victoria Scenario You have discovered the presence of a novel endophy5c organism living inside the cells
Primer Design Workshop École d'été en géné-que des champignons 2012 Dr. Will Hintz University of Victoria Scenario You have discovered the presence of a novel endophy5c organism living inside the cells
Supplemental Table 1. Primers used for PCR.
 Supplemental Table 1. Primers used for PCR. Gene Type Primer Sequence Genotyping and semi-quantitative RT-PCR F 5 -TTG CCC GAT CAC CAT CTG TA-3 rwa1-1 R 5 -TGT AGC GAT CAA GGC CTG ATC TAA-3 LB 5 -TAG CAT
Supplemental Table 1. Primers used for PCR. Gene Type Primer Sequence Genotyping and semi-quantitative RT-PCR F 5 -TTG CCC GAT CAC CAT CTG TA-3 rwa1-1 R 5 -TGT AGC GAT CAA GGC CTG ATC TAA-3 LB 5 -TAG CAT
Molecular Techniques Third-year Biology
 PLANNING Genetics Lab practices Molecular Techniques. Genetics Lab practices protocol. 2015-16 PCR-DIRECTED MUTAGENESIS, MOLECULAR CLONING AND RESTRICTION ANALYSIS Sessions 1 & 2 (2x3 hours): PCR-directed
PLANNING Genetics Lab practices Molecular Techniques. Genetics Lab practices protocol. 2015-16 PCR-DIRECTED MUTAGENESIS, MOLECULAR CLONING AND RESTRICTION ANALYSIS Sessions 1 & 2 (2x3 hours): PCR-directed
Supplementary Information
 Supplementary Information A general solution for opening double-stranded DNA for isothermal amplification Gangyi Chen, Juan Dong, Yi Yuan, Na Li, Xin Huang, Xin Cui* and Zhuo Tang* Supplementary Materials
Supplementary Information A general solution for opening double-stranded DNA for isothermal amplification Gangyi Chen, Juan Dong, Yi Yuan, Na Li, Xin Huang, Xin Cui* and Zhuo Tang* Supplementary Materials
Occurrence of viruses in apple and pear orchards in Latvia.
 INTERNATIONAL SCIENTIFIC CONFERENCE Sustainable Fruit Growing: From Plant to Product Occurrence of viruses in apple and pear orchards in Latvia. Neda P pola, Anna K le, Inga Moro ko Bi evska. The economically
INTERNATIONAL SCIENTIFIC CONFERENCE Sustainable Fruit Growing: From Plant to Product Occurrence of viruses in apple and pear orchards in Latvia. Neda P pola, Anna K le, Inga Moro ko Bi evska. The economically
Supplementary Figures
 Supplementary Figures Supplementary Fig. 1 Characterization of GSCs. a. Immunostaining of primary GSC spheres from GSC lines. Nestin (neural progenitor marker, red), TLX (green). Merged images of nestin,
Supplementary Figures Supplementary Fig. 1 Characterization of GSCs. a. Immunostaining of primary GSC spheres from GSC lines. Nestin (neural progenitor marker, red), TLX (green). Merged images of nestin,
Supporting Information
 Supporting Information Transfection of DNA Cages into Mammalian Cells Email: a.turberfield@physics.ox.ac.uk Table of Contents Supporting Figure 1 DNA tetrahedra used in transfection experiments 2 Supporting
Supporting Information Transfection of DNA Cages into Mammalian Cells Email: a.turberfield@physics.ox.ac.uk Table of Contents Supporting Figure 1 DNA tetrahedra used in transfection experiments 2 Supporting
ΔPDD1 x ΔPDD1. ΔPDD1 x wild type. 70 kd Pdd1. Pdd3
 Supplemental Fig. S1 ΔPDD1 x wild type ΔPDD1 x ΔPDD1 70 kd Pdd1 50 kd 37 kd Pdd3 Supplemental Fig. S1. ΔPDD1 strains express no detectable Pdd1 protein. Western blot analysis of whole-protein extracts
Supplemental Fig. S1 ΔPDD1 x wild type ΔPDD1 x ΔPDD1 70 kd Pdd1 50 kd 37 kd Pdd3 Supplemental Fig. S1. ΔPDD1 strains express no detectable Pdd1 protein. Western blot analysis of whole-protein extracts
Wet Lab Tutorial: Genelet Circuits
 Wet Lab Tutorial: Genelet Circuits DNA 17 This tutorial will introduce the in vitro transcriptional circuits toolkit. The tutorial will focus on the design, synthesis, and experimental testing of a synthetic
Wet Lab Tutorial: Genelet Circuits DNA 17 This tutorial will introduce the in vitro transcriptional circuits toolkit. The tutorial will focus on the design, synthesis, and experimental testing of a synthetic
Dierks Supplementary Fig. S1
 Dierks Supplementary Fig. S1 ITK SYK PH TH K42R wt K42R (kinase deficient) R29C E42K Y323F R29C E42K Y323F (reduced phospholipid binding) (enhanced phospholipid binding) (reduced Cbl binding) E42K Y323F
Dierks Supplementary Fig. S1 ITK SYK PH TH K42R wt K42R (kinase deficient) R29C E42K Y323F R29C E42K Y323F (reduced phospholipid binding) (enhanced phospholipid binding) (reduced Cbl binding) E42K Y323F
MCB421 FALL2005 EXAM#3 ANSWERS Page 1 of 12. ANSWER: Both transposon types form small duplications of adjacent host DNA sequences.
 Page 1 of 12 (10pts) 1. There are two mechanisms for transposition used by bacterial transposable elements: replicative (Tn3) and non-replicative (Tn5 and Tn10). Compare and contrast the two mechanisms
Page 1 of 12 (10pts) 1. There are two mechanisms for transposition used by bacterial transposable elements: replicative (Tn3) and non-replicative (Tn5 and Tn10). Compare and contrast the two mechanisms
Search for and Analysis of Single Nucleotide Polymorphisms (SNPs) in Rice (Oryza sativa, Oryza rufipogon) and Establishment of SNP Markers
 DNA Research 9, 163 171 (2002) Search for and Analysis of Single Nucleotide Polymorphisms (SNPs) in Rice (Oryza sativa, Oryza rufipogon) and Establishment of SNP Markers Shinobu Nasu, Junko Suzuki, Rieko
DNA Research 9, 163 171 (2002) Search for and Analysis of Single Nucleotide Polymorphisms (SNPs) in Rice (Oryza sativa, Oryza rufipogon) and Establishment of SNP Markers Shinobu Nasu, Junko Suzuki, Rieko
SUPPLEMENTARY INFORMATION
 1. RNA/DNA sequences used in this study 2. Height and stiffness measurements on hybridized molecules 3. Stiffness maps at varying concentrations of target DNA 4. Stiffness measurements on RNA/DNA hybrids.
1. RNA/DNA sequences used in this study 2. Height and stiffness measurements on hybridized molecules 3. Stiffness maps at varying concentrations of target DNA 4. Stiffness measurements on RNA/DNA hybrids.
NAME:... MODEL ANSWER... STUDENT NUMBER:... Maximum marks: 50. Internal Examiner: Hugh Murrell, Computer Science, UKZN
 COMP710, Bioinformatics with Julia, Test One, Thursday the 20 th of April, 2017, 09h30-11h30 1 NAME:...... MODEL ANSWER... STUDENT NUMBER:...... Maximum marks: 50 Internal Examiner: Hugh Murrell, Computer
COMP710, Bioinformatics with Julia, Test One, Thursday the 20 th of April, 2017, 09h30-11h30 1 NAME:...... MODEL ANSWER... STUDENT NUMBER:...... Maximum marks: 50 Internal Examiner: Hugh Murrell, Computer
BIOLOGY - CLUTCH CH.20 - BIOTECHNOLOGY.
 !! www.clutchprep.com CONCEPT: DNA CLONING DNA cloning is a technique that inserts a foreign gene into a living host to replicate the gene and produce gene products. Transformation the process by which
!! www.clutchprep.com CONCEPT: DNA CLONING DNA cloning is a technique that inserts a foreign gene into a living host to replicate the gene and produce gene products. Transformation the process by which
SUPPLEMENTAL MATERIAL GENOTYPING WITH MULTIPLEXING TARGETED RESEQUENCING
 SUPPLEMENTAL MATERIAL GENOTYPING WITH MULTIPLEXING TARGETED RESEQUENCING All of the patients and control subjects were sequenced and genotyped in the same way. Shotgun libraries of approximately 250 bp
SUPPLEMENTAL MATERIAL GENOTYPING WITH MULTIPLEXING TARGETED RESEQUENCING All of the patients and control subjects were sequenced and genotyped in the same way. Shotgun libraries of approximately 250 bp
dsxf - wt wt dsxf bp 2493bp Nature Biotechnology: doi: /nbt.4245 Supplementary Figure 1
 5 3 2493bp 2426bp attp GFP 3xP3 attp CDS dsxf - wt wt dsxf - Supplementary Figure 1 Molecular confirmation of the correct integration of the HDR-mediated event to generate dsxf PCRs were performed to verify
5 3 2493bp 2426bp attp GFP 3xP3 attp CDS dsxf - wt wt dsxf - Supplementary Figure 1 Molecular confirmation of the correct integration of the HDR-mediated event to generate dsxf PCRs were performed to verify
Protein Structure Analysis
 BINF 731 Protein Structure Analysis http://binf.gmu.edu/vaisman/binf731/ Iosif Vaisman COMPUTATIONAL BIOLOGY COMPUTATIONAL STRUCTURAL BIOLOGY COMPUTATIONAL MOLECULAR BIOLOGY BIOINFORMATICS STRUCTURAL BIOINFORMATICS
BINF 731 Protein Structure Analysis http://binf.gmu.edu/vaisman/binf731/ Iosif Vaisman COMPUTATIONAL BIOLOGY COMPUTATIONAL STRUCTURAL BIOLOGY COMPUTATIONAL MOLECULAR BIOLOGY BIOINFORMATICS STRUCTURAL BIOINFORMATICS
Electronic Supplementary Information. Transcription Monitoring Using Fused RNA with a Dye-Binding Light-Up Aptamer as a Tag: A Blue Fluorescent RNA
 Electronic Supplementary Information for Transcription Monitoring Using Fused RNA with a Dye-Binding Light-Up Aptamer as a Tag: A Blue Fluorescent RNA Shinsuke Sando,* Atsushi Narita, Masayoshi Hayami,
Electronic Supplementary Information for Transcription Monitoring Using Fused RNA with a Dye-Binding Light-Up Aptamer as a Tag: A Blue Fluorescent RNA Shinsuke Sando,* Atsushi Narita, Masayoshi Hayami,
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature07182 SUPPLEMENTAL FIGURES AND TABLES Fig. S1. myf5-expressing cells give rise to brown fat depots and skeletal muscle (a) Perirenal BAT from control (cre negative) and myf5-cre:r26r3-yfp
doi: 10.1038/nature07182 SUPPLEMENTAL FIGURES AND TABLES Fig. S1. myf5-expressing cells give rise to brown fat depots and skeletal muscle (a) Perirenal BAT from control (cre negative) and myf5-cre:r26r3-yfp
Engineering Escherichia coli for production of functionalized terpenoids using plant P450s
 Supplementary Information for Engineering Escherichia coli for production of functionalized terpenoids using plant P450s Michelle C. Y. Chang, Rachel A. Eachus, William Trieu, Dae-Kyun Ro, and Jay D. Keasling*
Supplementary Information for Engineering Escherichia coli for production of functionalized terpenoids using plant P450s Michelle C. Y. Chang, Rachel A. Eachus, William Trieu, Dae-Kyun Ro, and Jay D. Keasling*
