Thornhill et al. ME Supplementary Materials revision 08/2007 1
|
|
|
- Alexina Fleming
- 5 years ago
- Views:
Transcription
1 Thornhill et al. ME Supplementary Materials revision 08/ Supplementary Table 1 Summary of clones obtained from estimates of Taq polymerase/bacterial cloning error in Symbiodinium type A3 (culture 77). The dominant clone from this culture was subjected to a second round of PCR and cloning. Comparisons are relative to the parental sequence. Sample Name Same as Dominant Sequence? GenBank Accession # A3.Direct - EU A Yes EU A Yes EU A Yes EU A Yes EU A Yes EU A Yes EU A Yes EU A Yes EU A Yes EU A Yes EU A Yes EU A Yes EU A Yes EU A No EU A Yes EU A Yes EU A Yes EU A Yes EU A Yes EU A Yes EU A Yes EU A Yes EU A Yes EU A Yes EU A Yes EU A Yes EU A Yes EU A Yes EU A Yes EU074857
2 Thornhill et al. ME Supplementary Materials revision 08/ Supplementary Table 2 Summary of clones obtained from estimates of Taq polymerase/bacterial cloning error in Symbiodinium type C1 (culture 152). The dominant clone from this culture was subjected to a second round of PCR and cloning. Comparisons are relative to the parental sequence. Sample Name Same as Dominant Sequence? GenBank Accession # C1.Direct - EU C Yes EU C Yes EU C Yes EU C Yes EU C Yes EU C Yes EU C Yes EU C Yes EU C No EU C Yes EU C Yes EU C Yes EU C No EU C Yes EU C Yes EU C Yes EU C Yes EU C Yes EU C Yes EU C Yes EU C Yes EU C Yes EU C No EU C Yes EU C Yes EU C Yes EU C Yes EU C No EU C Yes EU C Yes EU074885
3 Thornhill et al. ME Supplementary Materials revision 08/ Supplementary Table 3 Summary of secondary structural disruptions and PCR-chimeras identified within the 5.8S-rDNA and ITS2 of bacterial clones originating from Symbiodinium type A3 (culture 77). A Y indicates that one or more disruptions were detected while a - denotes that no disruptions were identified. Sample Name Same as Dominant Sequence? GenBank Accession # 5.8S-rDNA Disruptions 1 ITS2 Disruptions 2 Suspected PCR Chimera A3.Direct - EU A Yes EU A Yes EU A Yes EU A No EU A No EU A Yes EU A No EU Y (124, 125) - - A Yes EU A Yes EU A No EU A Yes EU A Yes EU A Yes EU A No EU Y (110) - - A Yes EU A Yes EU A Yes EU A Yes EU A Yes EU A No EU Y (148) - A Yes EU A Yes EU Numbers corresponds to the nucleotide position listed in the alignment presented in Figure 3 of the manuscript. 2 Numbers corresponds to the nucleotide position listed in the Supplementary Figure 3 alignment.
4 Thornhill et al. ME Supplementary Materials revision 08/ Supplementary Table 4 Summary of secondary structural disruptions and PCR-chimeras identified within the 5.8S-rDNA and ITS2 of bacterial clones originating from Symbiodinium type B1 (culture 13). A Y indicates that one or more disruptions were detected while a - denotes that no disruptions were identified. Sample Name Same as Dominant Sequence? GenBank Accession # 5.8S-rDNA Disruptions 1 ITS2 Disruptions 2 Suspected PCR Chimera B1.Direct - EU B No EU B No EU B Yes EU B No EU B No EU Y (21) - - B Yes EU B No EU Y (30) - B No EU Y (142, 151) - B Yes EU B Yes EU B Yes EU B Yes EU B No EU Y (73) - - B Yes EU B No EU B Yes EU B Yes EU B Yes EU B No EU B No EU Y (7) - - B No EU B No EU Numbers corresponds to the nucleotide position listed in the alignment presented in Figure 3 of the manuscript. 2 Numbers corresponds to the nucleotide position listed in the alignment of Supplementary Figure 4.
5 Thornhill et al. ME Supplementary Materials revision 08/ Supplementary Table 5 Summary of secondary structural disruptions and PCR-chimeras identified within the 5.8S-rDNA and ITS2 of bacterial clones originating from Symbiodinium type C1 (culture 152). A Y indicates that one or more disruptions were detected while a - denotes that no disruptions were identified. Sample Name Same as Dominant Sequence? GenBank Accession # 5.8S-rDNA Disruptions 1 ITS2 Disruptions 2 Suspected PCR Chimera C1.Direct - EU C No EU Y (54, 60-68) - C Yes EU C No EU C No EU C Yes EU C No EU Y (95) - - C No EU C No EU C No EU C No EU Y (152) - C No EU C No EU Y (49, 123) - - C No EU C Yes EU C No EU C No EU C No EU Y (119) - - C No EU Y (54, 60-68) Y (C1/C1.1126) C Yes EU C No EU Y (26) - - C No EU C No EU Y (99) Numbers corresponds to the nucleotide position listed in the alignment presented in Figure 3 of the manuscript. 2 Numbers corresponds to the nucleotide position listed in the alignment of Supplementary Figure 5. 3 Names correspond to the parent sequences of a suspected PCR-chimera. See Supplementary Figure 1.
6 Thornhill et al. ME Supplementary Materials revision 08/ Supplementary Table 6 Summary of secondary structural disruptions and PCR-chimeras identified within the 5.8S-rDNA and ITS2 of bacterial clones originating from Symbiodinium type D1a (culture A001). A Y indicates that one or more disruptions were detected while a - denotes that no disruptions were identified. Sample Name Same as Dominant Sequence? GenBank Accession # 5.8S-rDNA Disruptions 1 ITS2 Disruptions 2 Suspected PCR Chimera D1a.Direct - EU D1a.2010 Yes EU D1a.2014 No EU Y (44-136) - - D1a.2017 No EU Y (48) - D1a.2019 Yes EU D1a.2023 No EU Y (48, 50, 93) - D1a.2025 Yes EU D1a.2027 No EU D1a.2028 No EU Y (44-136) Y (48) - D1a.2038 No EU D1a.2040 No EU Y (105) Y (48) - D1a.2042 No EU D1a.2050 No EU D1a.2052 No EU Y (150) - D1a.2056 Yes EU D1a.2062 No EU D1a.2064 No EU D1a.2066 No EU Y (121) - D1a.2068 No EU Numbers corresponds to the nucleotide position listed in the alignment presented in Figure 3 of the manuscript. 2 Numbers corresponds to the nucleotide position listed in the alignment presented in Figure 4 of the manuscript.
7 Thornhill et al. ME Supplementary Materials revision 08/ Supplementary Table 7 Summary of secondary structural disruptions and PCR-chimeras identified within the 5.8S-rDNA and ITS2 of bacterial clones originating from Symbiodinium type E2 (culture CCMP 421). A Y indicates that one or more disruptions were detected while a - denotes that no disruptions were identified. Same as Dominant Sequence? 5.8S-rDNA Disruptions 1 ITS2 Disruptions 2 Sample Name GenBank Accession # Suspected PCR Chimera 3 E2.Direct - EU E No EU Y (22) Y (49, 176, 195) Y (E2/E2.2094) E Yes EU E Yes EU E Yes EU E No EU Y (105) - - E No EU Y (157) - E No EU Y (51, 152, 176, 195) - E No EU Y (51, 152, 176, 195) - E No EU Y (49) - E No EU Y (49) - E No EU Y (49) Y (E2/E2.2094) E No EU E No EU Y (49) - E Yes EU E No EU E Yes EU E Yes EU E No EU E Yes EU E No EU Y (51, 152, 176, 190, 195) Y (E2/E2.2094) E No EU E No EU Y (119) - - E No EU Y (51, 152, 176, 195) - E No EU Y (51, 152, 176, 195) Y (E2/E2.2094) 1 Numbers corresponds to the nucleotide position listed in the alignment presented in Figure 3 of the manuscript. 2 Numbers corresponds to the nucleotide position listed in the alignment of Supplementary Figure 6. 3 Names correspond to the parent sequences of a suspected PCR-chimera. See Supplementary Figure 1.
8 Thornhill et al. ME Supplementary Materials revision 08/ Supplementary Table 8 Summary of secondary structural/processing site disruptions and PCR-chimeras identified within the 5.8SrDNA and ITS2 of bacterial clones originating from Symbiodinium populations of Hawaiian Porites spp. corals (data from Apprill & Gates 2007). A Y indicates that one or more disruptions were detected while a - denotes that no disruptions were identified. Sample Name 5.8S Disruptions 1 ITS2 & Conserved Processing Site Disruptions 2 Suspected PCR Chimera 3 GenBank Accession # C15 AY KB1 DQ Y ( , 110, 114) Y (78, 98, 110, 119, , 123) - KB2 DQ Y ( , 110, 114, 123) Y (78, 98, 110, 121) - KB3 DQ Y ( , 114) Y (55, 78, 110, 121) - KB4 DQ Y ( , 114) Y (55, 78, 110, 121) - KB102C_5B DQ Y ( , 114) - Y (KB4/C15) KB102C_8B DQ Y ( , 114) Y (78, 110, 121) - KB119_16 DQ Y (94) Y ( ) - KB129_4 DQ KB135_42 DQ Y ( , 110, 114) Y (78, 98, 104, 110, 121) - KB135_63 DQ Y ( , 110, 114) Y (78) Y (KB1/C15) KB135_89 DQ Y ( , 114) Y (78, 110, 121) - KB162R_3 DQ Y ( , 110, 114) Y (78, 98, 110, 119, , 123) - KB162R_4 DQ Y ( , 110, 114) Y (78, 98, 110, 119, , 123, 182) - KB179A_18 DQ KB179B_21 DQ Y ( , 114, 128) Y (78, 110, 121) - KB179B_24 DQ Y (40, 154) - KB179B_7 DQ Y ( , 114) Y (55) Y (KB4/C15) KB184_1 DQ Y ( , 110, 114) Y (78, 98, 110, 119, , 123) - KB184_3 DQ Y ( , 110, 114) Y (18, 78, 98, 110, 119, , 123) - KB184L_1 DQ Y ( , 110, 114) Y (78, 98, 110, 119, , 123) - KB184RL_4 DQ Y ( , 110, 114, 123) Y (78, 98, 110, 121) - KB184RR_3 DQ Y ( , 110, 114, 123) Y (78, 98, 110, 121) - KB184RR_4 DQ Y ( , 110) Y (33, 78, 98, 110, 119, , 123) - 1 Numbers corresponds to the nucleotide position listed in the alignment presented in Figure 3 of the manuscript. 2 Numbers corresponds to the nucleotide position listed in the alignment presented in Figure 5 of the manuscript. 3 Names correspond to the parent sequences of a suspected PCR-chimera. See Supplementary Figure 7. 8
9 Thornhill et al. ME Supplementary Materials revision 08/ Supplementary Figure Legends Supplementary Figure 1: Putative PCR-chimeras detected in the bacterial cloned dataset of Symbiodinium type C1 (culture 152) and type E2 (culture CCMP 421). Suspected chimeric sequences include clones: A) C1_1178, B) E2_2074, C) E2_2122, D) E2_2130, and E) E2_2102. For each, an alignment of the parent sequences and putative chimera is shown. Alignments include only those nucleotide positions that differ; the number at the top of an alignment (read vertically downwards) designates site position on an alignment that including all clones from that culture. Dashes (-) indicate an insertion/deletion (indel) while an arrow ( ) denotes the break point between parent sequences in the putative chimera. Nucleotide positions in the chimera differing from either parent sequence are indicated in gray; these are likely a result of Taq polymerase error or failure to recover the actual parental sequence(s). Diagrams are presented below each alignment indicating the portions of the parent molecules contributing to each chimera. Supplementary Figure 2: Unrooted maximum parsimony phylogenies of ITS2 region sequence variants generated via PCR and bacterial cloning from two clonal Symbiodinium cultures: (A) culture 77 ( type A3; GenBank Accession Numbers EU EU074938), (B) culture 152 ( type C1; GenBank Accession Numbers EU EU074961). For each polytomy, numbers of point substitutions and/or indels are related to branch length. Indels were weighted as a single change regardless of length. Numbers of clones with identical sequences are indicated by circle size at each node; identical sequences recovered more than three times have the corresponding number of replicates (in parentheses) following clone name. Supplementary Figures 3 6: Secondary structural analyses and alignments of ITS2 sequences from clonal isolates of Symbiodinium types : A3 (culture 77, Supplemental Figure 3), B1 (culture 13, Supplemental Figure 4), C1 (culture 152, Supplemental Figure 5), and E2 (culture CCMP 421, Supplemental Figure 6). Boundaries of the helices are labeled and correspond to those above the nucleotide alignment. The sequence generated directly from the PCR product is used as the template against which the cloned sequences are compared (see Materials and Methods). A nucleotide (A, C, G, or U) in place of a dot (.) denotes a substitution at that position. Dashes (-) indicate an insertion/deletion (indel). Numbers listed under Mut. Pos. represent positions where mutational differences exist between the direct- and clone-derived sequences. For these, effects to secondary structural stability of ITS2 are inferred (i.e., Effect SS ): N = no influence, + = positive influence, - = negative influence, ~ = weakened interaction (G-U pairing). Arrows identify variable sequence positions on the secondary structure diagram. Characters depicted in gray represent a highly conserved rrna processing site. Supplementary Figure 7: Putative PCR-chimeras detected in the bacterial cloned dataset of Symbiodinium populations of Hawaiian Porites spp. corals (data from Apprill & Gates 2007). Suspected chimeric sequences include clones: A) KB102C_5B, B) KB135_63, C) KB179B_7. For each, an alignment of the parent sequences and putative chimera is shown. Alignments include only those nucleotide positions that differ; the number at the top of an alignment (read vertically downwards) designates site position on an alignment that including all clones from Apprill & Gates (2007). Dashes (-) indicate an insertion/deletion (indel) while an arrow ( ) denotes the break point between parent sequences in the putative chimera. Nucleotide positions 9
10 Thornhill et al. ME Supplementary Materials revision 08/ in the chimera differing from either parent sequence are indicated in gray; these are likely a result of Taq polymerase error or failure to recover the actual parental sequence(s). Diagrams are presented below each alignment indicating the portions of the parent molecules contributing to each chimera. 10
11 Thornhill et al. ME Supplementary Materials revision 08/ Supplementary Figure 1 11
12 Thornhill et al. ME Supplementary Materials revision 08/ Supplementary Figure 2 12
13 Thornhill et al. ME Supplementary Materials revision 08/ Supplementary Figure 3 13
14 Thornhill et al. ME Supplementary Materials revision 08/ Supplementary Figure 4 14
15 Thornhill et al. ME Supplementary Materials revision 08/ Supplementary Figure 5 15
16 Thornhill et al. ME Supplementary Materials revision 08/ Supplementary Figure 6 16
17 Thornhill et al. ME Supplementary Materials revision 08/ Supplementary Figure 7 17
Factors affecting PCR
 Lec. 11 Dr. Ahmed K. Ali Factors affecting PCR The sequences of the primers are critical to the success of the experiment, as are the precise temperatures used in the heating and cooling stages of the
Lec. 11 Dr. Ahmed K. Ali Factors affecting PCR The sequences of the primers are critical to the success of the experiment, as are the precise temperatures used in the heating and cooling stages of the
Bundle 6 Test Review
 Bundle 6 Test Review DNA vs. RNA DNA Replication Gene Mutations- Protein Synthesis 1. Label the different components and complete the complimentary base pairing. What is this molecule called? Deoxyribonucleic
Bundle 6 Test Review DNA vs. RNA DNA Replication Gene Mutations- Protein Synthesis 1. Label the different components and complete the complimentary base pairing. What is this molecule called? Deoxyribonucleic
All subjects gave written informed consent before participating in this study, which
 Supplementary Information Materials and Methods Sequence generation and analysis All subjects gave written informed consent before participating in this study, which was approved by the Washington University
Supplementary Information Materials and Methods Sequence generation and analysis All subjects gave written informed consent before participating in this study, which was approved by the Washington University
Supplementary Figure 1
 Supplementary Figure 1 Rarefaction Analysis and Sampling. a, Rarefaction plots show the expected levels of coverage for different subsamples of sequencing libraries for repertoires of clone size of at
Supplementary Figure 1 Rarefaction Analysis and Sampling. a, Rarefaction plots show the expected levels of coverage for different subsamples of sequencing libraries for repertoires of clone size of at
Some types of Mutagenesis
 Mutagenesis What Is a Mutation? Genetic information is encoded by the sequence of the nucleotide bases in DNA of the gene. The four nucleotides are: adenine (A), thymine (T), guanine (G), and cytosine
Mutagenesis What Is a Mutation? Genetic information is encoded by the sequence of the nucleotide bases in DNA of the gene. The four nucleotides are: adenine (A), thymine (T), guanine (G), and cytosine
SUPPLEMENTARY MATERIAL AND METHODS
 SUPPLEMENTARY MATERIAL AND METHODS Amplification of HEV ORF1, ORF2 and ORF3 genome regions Total RNA was extracted from 200 µl EDTA plasma using Cobas AmpliPrep total nucleic acid isolation kit (Roche,
SUPPLEMENTARY MATERIAL AND METHODS Amplification of HEV ORF1, ORF2 and ORF3 genome regions Total RNA was extracted from 200 µl EDTA plasma using Cobas AmpliPrep total nucleic acid isolation kit (Roche,
Bellerophon; a program to detect chimeric sequences in multiple sequence
 Revised ms: BIOINF-03-0817 Bellerophon; a program to detect chimeric sequences in multiple sequence alignments. Thomas Huber 1 *, Geoffrey Faulkner 1 and Philip Hugenholtz 2 1 ComBinE group, Advanced Computational
Revised ms: BIOINF-03-0817 Bellerophon; a program to detect chimeric sequences in multiple sequence alignments. Thomas Huber 1 *, Geoffrey Faulkner 1 and Philip Hugenholtz 2 1 ComBinE group, Advanced Computational
Supplementary information:
 Supplementary information: Distinct amino acids in the C-linker domain of the plant K + subcellular localization and activity at the plasma membrane channel KAT2 determine its Nieves-Cordones et al. Part
Supplementary information: Distinct amino acids in the C-linker domain of the plant K + subcellular localization and activity at the plasma membrane channel KAT2 determine its Nieves-Cordones et al. Part
Biological Sciences 50 Practice Exam 2
 NAME: Fall 2005 TF: Biological Sciences 50 Practice Exam 2 A. Be sure to write your name on the top of each of page of the examination. B. Write each answer only on the same page as the pertinent question.
NAME: Fall 2005 TF: Biological Sciences 50 Practice Exam 2 A. Be sure to write your name on the top of each of page of the examination. B. Write each answer only on the same page as the pertinent question.
Nature Biotechnology: doi: /nbt.4199
 Supplementary Figure 1 Base editing activities of engineered A3A-BE3 variants with mutations designed to disrupt non-specific interactions with substrate ssdna. Graphs illustrating the frequencies of C
Supplementary Figure 1 Base editing activities of engineered A3A-BE3 variants with mutations designed to disrupt non-specific interactions with substrate ssdna. Graphs illustrating the frequencies of C
(Hadlock, Journal of Virology, 2000), (Keck, Journal of Virology, 2008) CBH-5 Domain B 412, 416, 417, 418, 420, 421, 422, 423, 483, 484, 485, 488,
 Supplementary Table. mabs analyzed mab Antigenic Critical Binding Residues by Citations Domains Alanine Scanning 2 CBH-4B Domain A NA (Hadlock, Journal of Virology, 2), (Keck, Journal of Virology, 24)
Supplementary Table. mabs analyzed mab Antigenic Critical Binding Residues by Citations Domains Alanine Scanning 2 CBH-4B Domain A NA (Hadlock, Journal of Virology, 2), (Keck, Journal of Virology, 24)
Solutions to Quiz II
 MIT Department of Biology 7.014 Introductory Biology, Spring 2005 Solutions to 7.014 Quiz II Class Average = 79 Median = 82 Grade Range % A 90-100 27 B 75-89 37 C 59 74 25 D 41 58 7 F 0 40 2 Question 1
MIT Department of Biology 7.014 Introductory Biology, Spring 2005 Solutions to 7.014 Quiz II Class Average = 79 Median = 82 Grade Range % A 90-100 27 B 75-89 37 C 59 74 25 D 41 58 7 F 0 40 2 Question 1
on November 21, 2018 by guest
 AEM Accepts, published online ahead of print on 30 November 2012 Appl. Environ. Microbiol. doi:10.1128/aem.03171-12 Copyright 2012, American Society for Microbiology. All Rights Reserved. 1 2 3 4 5 6 7
AEM Accepts, published online ahead of print on 30 November 2012 Appl. Environ. Microbiol. doi:10.1128/aem.03171-12 Copyright 2012, American Society for Microbiology. All Rights Reserved. 1 2 3 4 5 6 7
03-511/711 Computational Genomics and Molecular Biology, Fall
 03-511/711 Computational Genomics and Molecular Biology, Fall 2011 1 Study questions These study problems are intended to help you to review for the final exam. This is not an exhaustive list of the topics
03-511/711 Computational Genomics and Molecular Biology, Fall 2011 1 Study questions These study problems are intended to help you to review for the final exam. This is not an exhaustive list of the topics
Supplementary Figure 1. Design of the control microarray. a, Genomic DNA from the
 Supplementary Information Supplementary Figures Supplementary Figure 1. Design of the control microarray. a, Genomic DNA from the strain M8 of S. ruber and a fosmid containing the S. ruber M8 virus M8CR4
Supplementary Information Supplementary Figures Supplementary Figure 1. Design of the control microarray. a, Genomic DNA from the strain M8 of S. ruber and a fosmid containing the S. ruber M8 virus M8CR4
* NOTE: If Lesson 2 has been performed, the DNA model should be uncoiled (and taken off its stand or tubing) to perform the steps on Pages
 OBJECTIVES Students will be able to: 1. Understand the DNA replication process. 2. Identify reasons for replication of DNA. 3. Understand the need for an exact copy of DNA. 4. Understand the function of
OBJECTIVES Students will be able to: 1. Understand the DNA replication process. 2. Identify reasons for replication of DNA. 3. Understand the need for an exact copy of DNA. 4. Understand the function of
Simple Deletion: a vector- and marker-free method to generate and isolate site-directed
 Electronic supplementary materials Simple Deletion: a vector- and marker-free method to generate and isolate site-directed deletion mutants Yasuhiro Inoue 1, Seiji Tsuge 2 1 National Agriculture and Food
Electronic supplementary materials Simple Deletion: a vector- and marker-free method to generate and isolate site-directed deletion mutants Yasuhiro Inoue 1, Seiji Tsuge 2 1 National Agriculture and Food
Pre-AP Biology DNA and Biotechnology Study Guide #1
 Last Name: First Name: Per. Pre-AP Biology DNA and Biotechnology Study Guide #1 Structure of DNA: Number of strands. Parallel or antiparallel?. Rosalind Franklin s x-ray crystallography image indicated
Last Name: First Name: Per. Pre-AP Biology DNA and Biotechnology Study Guide #1 Structure of DNA: Number of strands. Parallel or antiparallel?. Rosalind Franklin s x-ray crystallography image indicated
Bio Rad PCR Song Lyrics
 Bio Rad PCR Song Lyrics There was a time when to amplify DNA, You had to grow tons and tons of tiny cells. (Oooh) Then along came a guy named Dr. Kary Mullis, Said you can amplify in vitro just as well.
Bio Rad PCR Song Lyrics There was a time when to amplify DNA, You had to grow tons and tons of tiny cells. (Oooh) Then along came a guy named Dr. Kary Mullis, Said you can amplify in vitro just as well.
The Structure of DNA
 The Structure of DNA Questions to Ponder 1) How is the genetic info copied? 2) How does DNA store the genetic information? 3) How is the genetic info passed from generation to generation? The Structure
The Structure of DNA Questions to Ponder 1) How is the genetic info copied? 2) How does DNA store the genetic information? 3) How is the genetic info passed from generation to generation? The Structure
Information on barcode decoding
 Information on barcode decoding The Bioneer version 1.0 haploid deletion library (Bioneer catalog number M-1030H) was supplied as glycerol frozen stocks in thirty-one 96-well plates. According to the information
Information on barcode decoding The Bioneer version 1.0 haploid deletion library (Bioneer catalog number M-1030H) was supplied as glycerol frozen stocks in thirty-one 96-well plates. According to the information
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/4/2/eaao0665/dc1 Supplementary Materials for Variant ribosomal RNA alleles are conserved and exhibit tissue-specific expression Matthew M. Parks, Chad M. Kurylo,
advances.sciencemag.org/cgi/content/full/4/2/eaao0665/dc1 Supplementary Materials for Variant ribosomal RNA alleles are conserved and exhibit tissue-specific expression Matthew M. Parks, Chad M. Kurylo,
Mutations during meiosis and germ line division lead to genetic variation between individuals
 Mutations during meiosis and germ line division lead to genetic variation between individuals Types of mutations: point mutations indels (insertion/deletion) copy number variation structural rearrangements
Mutations during meiosis and germ line division lead to genetic variation between individuals Types of mutations: point mutations indels (insertion/deletion) copy number variation structural rearrangements
1. DNA replication. (a) Why is DNA replication an essential process?
 ame Section 7.014 Problem Set 3 Please print out this problem set and record your answers on the printed copy. Answers to this problem set are to be turned in to the box outside 68120 by 5:00pm on Friday
ame Section 7.014 Problem Set 3 Please print out this problem set and record your answers on the printed copy. Answers to this problem set are to be turned in to the box outside 68120 by 5:00pm on Friday
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Fredricks DN, Fiedler TL, Marrazzo JM. Molecular identification
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Fredricks DN, Fiedler TL, Marrazzo JM. Molecular identification
PCR in the Classroom. UC Davis - PCR Workshop Friday, September 26, 2003
 PCR in the Classroom UC Davis - PCR Workshop Friday, September 26, 2003 A little history In 1983, Kary B. Mullis conceived the procedure. He went on to Cetus Corp in Emeryville, CA where it was developed
PCR in the Classroom UC Davis - PCR Workshop Friday, September 26, 2003 A little history In 1983, Kary B. Mullis conceived the procedure. He went on to Cetus Corp in Emeryville, CA where it was developed
IN E. COLI WHAT IS THE FUNCTION OF DNA POLYMERASE III
 10 January, 2018 IN E. COLI WHAT IS THE FUNCTION OF DNA POLYMERASE III Document Filetype: PDF 312 KB 0 IN E. COLI WHAT IS THE FUNCTION OF DNA POLYMERASE III The actual replication enzyme in E. Both will
10 January, 2018 IN E. COLI WHAT IS THE FUNCTION OF DNA POLYMERASE III Document Filetype: PDF 312 KB 0 IN E. COLI WHAT IS THE FUNCTION OF DNA POLYMERASE III The actual replication enzyme in E. Both will
BIOLOGY - CLUTCH CH.20 - BIOTECHNOLOGY.
 !! www.clutchprep.com CONCEPT: DNA CLONING DNA cloning is a technique that inserts a foreign gene into a living host to replicate the gene and produce gene products. Transformation the process by which
!! www.clutchprep.com CONCEPT: DNA CLONING DNA cloning is a technique that inserts a foreign gene into a living host to replicate the gene and produce gene products. Transformation the process by which
Nature Immunology: doi: /ni Supplementary Figure 1. Data-processing pipeline.
 Supplementary Figure 1 Data-processing pipeline. Steps for processing data from multiple sorted B cell populations derived from a single individual at a single time point are shown. Parameters used are
Supplementary Figure 1 Data-processing pipeline. Steps for processing data from multiple sorted B cell populations derived from a single individual at a single time point are shown. Parameters used are
Analysis in Forensic Science
 Chapter 16 Gene Cloning & DNA Analysis in Forensic Science 1. DNA analysis in identification of crime suspects 2. Studying kinship by DNA profiling 3. Sex identification by DNA analysis Forensic science
Chapter 16 Gene Cloning & DNA Analysis in Forensic Science 1. DNA analysis in identification of crime suspects 2. Studying kinship by DNA profiling 3. Sex identification by DNA analysis Forensic science
Technical University of Denmark. Written examination, 29 May 2012 Course name: Life Science. Course number: Aids allowed: Written material
 1 Technical University of Denmark Written examination, 29 May 2012 Course name: Life Science Course number: 27008 Aids allowed: Written material Exam duration: 4 hours Weighting: The exam set consists
1 Technical University of Denmark Written examination, 29 May 2012 Course name: Life Science Course number: 27008 Aids allowed: Written material Exam duration: 4 hours Weighting: The exam set consists
SUPPLEMENTARY INFORMATION
 Contents De novo assembly... 2 Assembly statistics for all 150 individuals... 2 HHV6b integration... 2 Comparison of assemblers... 4 Variant calling and genotyping... 4 Protein truncating variants (PTV)...
Contents De novo assembly... 2 Assembly statistics for all 150 individuals... 2 HHV6b integration... 2 Comparison of assemblers... 4 Variant calling and genotyping... 4 Protein truncating variants (PTV)...
My Question Paper. Maximum Mark. Mark Awarded. Question. Total Mark. Page 1 of 27 WJEC/CBAC 2016 pdfcrowd.com
 My Question Paper Question Maximum Mark Mark Awarded 1 10 2 4 3 10 4 10 5 10 6 12 7 5 8 6 9 10 Total Mark Page 1 of 27 WJEC/CBAC 2016 1. Page 2 of 27 WJEC/CBAC 2016 2. Page 3 of 27 WJEC/CBAC 2016 3. The
My Question Paper Question Maximum Mark Mark Awarded 1 10 2 4 3 10 4 10 5 10 6 12 7 5 8 6 9 10 Total Mark Page 1 of 27 WJEC/CBAC 2016 1. Page 2 of 27 WJEC/CBAC 2016 2. Page 3 of 27 WJEC/CBAC 2016 3. The
7.1 Techniques for Producing and Analyzing DNA. SBI4U Ms. Ho-Lau
 7.1 Techniques for Producing and Analyzing DNA SBI4U Ms. Ho-Lau What is Biotechnology? From Merriam-Webster: the manipulation of living organisms or their components to produce useful usually commercial
7.1 Techniques for Producing and Analyzing DNA SBI4U Ms. Ho-Lau What is Biotechnology? From Merriam-Webster: the manipulation of living organisms or their components to produce useful usually commercial
NB536: Bioinformatics
 NB536: Bioinformatics Instructor Prof. Jong Kyoung Kim Department of New Biology Office: E4-613 E-mail: jkkim@dgist.ac.kr Homepage: https://scg.dgist.ac.kr Course website https://scg.dgist.ac.kr/index.php/courses
NB536: Bioinformatics Instructor Prof. Jong Kyoung Kim Department of New Biology Office: E4-613 E-mail: jkkim@dgist.ac.kr Homepage: https://scg.dgist.ac.kr Course website https://scg.dgist.ac.kr/index.php/courses
Worksheet Structure of DNA and Replication
 Eastern Intermediate High School Honors Biology Name: Period: Date: Worksheet Structure of DN and Replication Directions: Label the diagram below with the following choices: Nucleotide Deoxyribose Phosphate
Eastern Intermediate High School Honors Biology Name: Period: Date: Worksheet Structure of DN and Replication Directions: Label the diagram below with the following choices: Nucleotide Deoxyribose Phosphate
Chapter 4 DNA Structure & Gene Expression
 Biology 12 Name: Cell Biology Per: Date: Chapter 4 DNA Structure & Gene Expression Complete using BC Biology 12, pages 108-153 4.1 DNA Structure pages 112-114 1. DNA stands for and is the genetic material
Biology 12 Name: Cell Biology Per: Date: Chapter 4 DNA Structure & Gene Expression Complete using BC Biology 12, pages 108-153 4.1 DNA Structure pages 112-114 1. DNA stands for and is the genetic material
Student Exercise on Polymerase Chain Reaction (PCR) Prepared by the Office of Biotechnology, Iowa State University. Part I. Part III.
 Student Exercise on Polymerase Chain Reaction (PCR) Prepared by the Office of Biotechnology, Iowa State University Part I. Segment of interest (02) Original-2 5' 3' 1. The purpose of PCR is to make copies
Student Exercise on Polymerase Chain Reaction (PCR) Prepared by the Office of Biotechnology, Iowa State University Part I. Segment of interest (02) Original-2 5' 3' 1. The purpose of PCR is to make copies
Chapter 8: Recombinant DNA. Ways this technology touches us. Overview. Genetic Engineering
 Chapter 8 Recombinant DNA and Genetic Engineering Genetic manipulation Ways this technology touches us Criminal justice The Justice Project, started by law students to advocate for DNA testing of Death
Chapter 8 Recombinant DNA and Genetic Engineering Genetic manipulation Ways this technology touches us Criminal justice The Justice Project, started by law students to advocate for DNA testing of Death
September 19, synthesized DNA. Label all of the DNA strands with 5 and 3 labels, and clearly show which strand(s) contain methyl groups.
 KEY DNA Replication and Mutation September 19, 2011 1. Below is a short DNA sequence located on the E. coli chromosome. In class we talked about how during the process of DNA replication, an enzyme adds
KEY DNA Replication and Mutation September 19, 2011 1. Below is a short DNA sequence located on the E. coli chromosome. In class we talked about how during the process of DNA replication, an enzyme adds
Module 6 Microbial Genetics. Chapter 8
 Module 6 Microbial Genetics Chapter 8 Structure and function of the genetic material Genetics science of o Study of what genes are, how they determine the characteristics of an organism, how they carry
Module 6 Microbial Genetics Chapter 8 Structure and function of the genetic material Genetics science of o Study of what genes are, how they determine the characteristics of an organism, how they carry
4. Analysing genes II Isolate mutants*
 .. 4. Analysing s II Isolate mutants* Using the mutant to isolate the classify mutants by complementation analysis wild type study phenotype of mutants mutant 1 - use mutant to isolate sequence put individual
.. 4. Analysing s II Isolate mutants* Using the mutant to isolate the classify mutants by complementation analysis wild type study phenotype of mutants mutant 1 - use mutant to isolate sequence put individual
Laura Sims PhD UC Berkeley Forest Pathology and Mycology Lab
 Laura Sims PhD UC Berkeley Forest Pathology and Mycology Lab Outline What can molecular biology tell us about a pathogen? Tools and techniques used for diagnostics ELISA PCR Sequencing Sequence alignment
Laura Sims PhD UC Berkeley Forest Pathology and Mycology Lab Outline What can molecular biology tell us about a pathogen? Tools and techniques used for diagnostics ELISA PCR Sequencing Sequence alignment
B. Transgenic plants with strong phenotype (%)
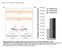 A. TCTAGTTGTTGTTGTTATGGTCTAGTTGTTGTTGTTATGGTCTAATTT AAATATGGTCTAAAGAAGAAGAATATGGTCTAAAGAAGAAGAATATGG 2XP35S STTM165 5 GGGGGATGAAGctaCCTGGTCCGA3 3 CCCCCUACUUC---GGACCAGGCU5 mir165 HindIII mir165 96 nt GTTGTTGTTGTTATGGTCTAGTTGTTGTTGTTATGGTCTAATTT
A. TCTAGTTGTTGTTGTTATGGTCTAGTTGTTGTTGTTATGGTCTAATTT AAATATGGTCTAAAGAAGAAGAATATGGTCTAAAGAAGAAGAATATGG 2XP35S STTM165 5 GGGGGATGAAGctaCCTGGTCCGA3 3 CCCCCUACUUC---GGACCAGGCU5 mir165 HindIII mir165 96 nt GTTGTTGTTGTTATGGTCTAGTTGTTGTTGTTATGGTCTAATTT
KEY CONCEPT DNA was identified as the genetic material through a series of experiments. Found live S with R bacteria and injected
 Section 1: Identifying DNA as the Genetic Material KEY CONCEPT DNA was identified as the genetic material through a series of experiments. VOCABULARY bacteriophage MAIN IDEA: Griffith finds a transforming
Section 1: Identifying DNA as the Genetic Material KEY CONCEPT DNA was identified as the genetic material through a series of experiments. VOCABULARY bacteriophage MAIN IDEA: Griffith finds a transforming
 http://fire.biol.wwu.edu/trent/trent/direct_detection_of_genotype.html 1 Like most other model organism Arabidopsis thaliana has a sequenced genome? What do we mean by sequenced genome? What sort of info
http://fire.biol.wwu.edu/trent/trent/direct_detection_of_genotype.html 1 Like most other model organism Arabidopsis thaliana has a sequenced genome? What do we mean by sequenced genome? What sort of info
11/17/14. Why would scientist want to make a mouse glow?
 11/17/14 Why would scientist want to make a mouse glow? 11/20 Your test today has ten words please use this time wisely. Chapter 8 Vocabulary Review Bacteriophage Viruses that infect bacteria, makes the
11/17/14 Why would scientist want to make a mouse glow? 11/20 Your test today has ten words please use this time wisely. Chapter 8 Vocabulary Review Bacteriophage Viruses that infect bacteria, makes the
Donor DNA Utilization during Gene Targeting with Zinc- finger Nucleases
 Donor DNA Utilization during Gene Targeting with Zinc- finger Nucleases Kelly J. Beumer, Jonathan K. Trautman, Kusumika Mukherjee and Dana Carroll Department of Biochemistry University of Utah School of
Donor DNA Utilization during Gene Targeting with Zinc- finger Nucleases Kelly J. Beumer, Jonathan K. Trautman, Kusumika Mukherjee and Dana Carroll Department of Biochemistry University of Utah School of
03-511/711 Computational Genomics and Molecular Biology, Fall
 03-511/711 Computational Genomics and Molecular Biology, Fall 2010 1 Study questions These study problems are intended to help you to review for the final exam. This is not an exhaustive list of the topics
03-511/711 Computational Genomics and Molecular Biology, Fall 2010 1 Study questions These study problems are intended to help you to review for the final exam. This is not an exhaustive list of the topics
Overview: The DNA Toolbox
 Overview: The DNA Toolbox Sequencing of the genomes of more than 7,000 species was under way in 2010 DNA sequencing has depended on advances in technology, starting with making recombinant DNA In recombinant
Overview: The DNA Toolbox Sequencing of the genomes of more than 7,000 species was under way in 2010 DNA sequencing has depended on advances in technology, starting with making recombinant DNA In recombinant
Combinatorial Evolution of Enzymes and Synthetic pathways Using
 Supplementary data Combinatorial Evolution of Enzymes and Synthetic pathways Using One-Step PCR Peng Jin,, Zhen Kang,,, *, Junli Zhang,, Linpei Zhang,, Guocheng Du,, and Jian Chen, The Key Laboratory of
Supplementary data Combinatorial Evolution of Enzymes and Synthetic pathways Using One-Step PCR Peng Jin,, Zhen Kang,,, *, Junli Zhang,, Linpei Zhang,, Guocheng Du,, and Jian Chen, The Key Laboratory of
Unit 1. DNA and the Genome
 Unit 1 DNA and the Genome National 5 Knowledge Learners should have a clear understanding of the following areas of content from their previous learning: *Cell division (mitosis) and chromosomes *Base
Unit 1 DNA and the Genome National 5 Knowledge Learners should have a clear understanding of the following areas of content from their previous learning: *Cell division (mitosis) and chromosomes *Base
3. The following sequence is destined to be translated into a protein: However, a mutation occurs that results in the molecule being altered to:
 1. Please identify the molecule below: 5 -ACTCGATTACGATACGA-3ʼ a) DNA b) mrna c) trna d) rrna e) It cannot be determined 2. If a complimentary strand of RNA were made to the molecule in question 1, what
1. Please identify the molecule below: 5 -ACTCGATTACGATACGA-3ʼ a) DNA b) mrna c) trna d) rrna e) It cannot be determined 2. If a complimentary strand of RNA were made to the molecule in question 1, what
Biology 100. ALE #10. From Gene to Protein and Biotechnology Practice Problems DNA
 Biology 100 Instructor: K. Marr Name Lab Section Group No. Quarter ALE #10. From Gene to Protein and Biotechnology Practice Problems Answer the following questions neatly and fully in the spaces provided.
Biology 100 Instructor: K. Marr Name Lab Section Group No. Quarter ALE #10. From Gene to Protein and Biotechnology Practice Problems Answer the following questions neatly and fully in the spaces provided.
DNA REPLICATION REVIEW
 Biology Ms. Ye DNA REPLICATION REVIEW 1. Number the steps of DNA replication the correct order (1, 2, 3): Name Date Block Daughter strands are formed using complementary base pairing DNA unwinds The DNA
Biology Ms. Ye DNA REPLICATION REVIEW 1. Number the steps of DNA replication the correct order (1, 2, 3): Name Date Block Daughter strands are formed using complementary base pairing DNA unwinds The DNA
SUPPLEMENTAL MATERIALS
 SUPPLEMENL MERILS Eh-seq: RISPR epitope tagging hip-seq of DN-binding proteins Daniel Savic, E. hristopher Partridge, Kimberly M. Newberry, Sophia. Smith, Sarah K. Meadows, rian S. Roberts, Mark Mackiewicz,
SUPPLEMENL MERILS Eh-seq: RISPR epitope tagging hip-seq of DN-binding proteins Daniel Savic, E. hristopher Partridge, Kimberly M. Newberry, Sophia. Smith, Sarah K. Meadows, rian S. Roberts, Mark Mackiewicz,
DNA Structure and Replication, and Virus Structure and Replication Test Review
 DNA Structure and Replication, and Virus Structure and Replication Test Review What does DNA stand for? Deoxyribonucleic Acid DNA is what type of macromolecule? DNA is a nucleic acid The building blocks
DNA Structure and Replication, and Virus Structure and Replication Test Review What does DNA stand for? Deoxyribonucleic Acid DNA is what type of macromolecule? DNA is a nucleic acid The building blocks
Nucleic Acid Structure:
 Genetic Information In Microbes: The genetic material of bacteria and plasmids is DNA. Bacterial viruses (bacteriophages or phages) have DNA or RNA as genetic material. The two essential functions of genetic
Genetic Information In Microbes: The genetic material of bacteria and plasmids is DNA. Bacterial viruses (bacteriophages or phages) have DNA or RNA as genetic material. The two essential functions of genetic
SELECTED TECHNIQUES AND APPLICATIONS IN MOLECULAR GENETICS
 SELECTED TECHNIQUES APPLICATIONS IN MOLECULAR GENETICS Restriction Enzymes 15.1.1 The Discovery of Restriction Endonucleases p. 420 2 2, 3, 4, 6, 7, 8 Assigned Reading in Snustad 6th ed. 14.1.1 The Discovery
SELECTED TECHNIQUES APPLICATIONS IN MOLECULAR GENETICS Restriction Enzymes 15.1.1 The Discovery of Restriction Endonucleases p. 420 2 2, 3, 4, 6, 7, 8 Assigned Reading in Snustad 6th ed. 14.1.1 The Discovery
Biol 321 Spring 2013 Quiz 4 25 pts NAME
 Biol 321 Spring 2013 Quiz 4 25 pts NAME 1. (3 pts.) a. What is the name of this compound? BE EXPLICIT deoxyribose 5 b. Number the carbons on this structure: 4 1 3 2 2. (4 pts.) Circle True or False. If
Biol 321 Spring 2013 Quiz 4 25 pts NAME 1. (3 pts.) a. What is the name of this compound? BE EXPLICIT deoxyribose 5 b. Number the carbons on this structure: 4 1 3 2 2. (4 pts.) Circle True or False. If
Marker types. Potato Association of America Frederiction August 9, Allen Van Deynze
 Marker types Potato Association of America Frederiction August 9, 2009 Allen Van Deynze Use of DNA Markers in Breeding Germplasm Analysis Fingerprinting of germplasm Arrangement of diversity (clustering,
Marker types Potato Association of America Frederiction August 9, 2009 Allen Van Deynze Use of DNA Markers in Breeding Germplasm Analysis Fingerprinting of germplasm Arrangement of diversity (clustering,
Application Microfluidic Biochemical Analysis Devices (I) Date: 2013/06/07. Dr. Yi-Chung Tung. Polymerase Chain Reaction (PCR)
 Application Microfluidic Biochemical Analysis Devices (I) Date: 2013/06/07 Dr. Yi-Chung Tung The polymerase chain reaction (PCR) is a biochemical technology in molecular biology to amplify a single or
Application Microfluidic Biochemical Analysis Devices (I) Date: 2013/06/07 Dr. Yi-Chung Tung The polymerase chain reaction (PCR) is a biochemical technology in molecular biology to amplify a single or
Problem Set 8. Answer Key
 MCB 102 University of California, Berkeley August 11, 2009 Isabelle Philipp Online Document Problem Set 8 Answer Key 1. The Genetic Code (a) Are all amino acids encoded by the same number of codons? no
MCB 102 University of California, Berkeley August 11, 2009 Isabelle Philipp Online Document Problem Set 8 Answer Key 1. The Genetic Code (a) Are all amino acids encoded by the same number of codons? no
Enzyme that uses RNA as a template to synthesize a complementary DNA
 Biology 105: Introduction to Genetics PRACTICE FINAL EXAM 2006 Part I: Definitions Homology: Comparison of two or more protein or DNA sequence to ascertain similarities in sequences. If two genes have
Biology 105: Introduction to Genetics PRACTICE FINAL EXAM 2006 Part I: Definitions Homology: Comparison of two or more protein or DNA sequence to ascertain similarities in sequences. If two genes have
Sections 12.3, 13.1, 13.2
 Sections 12.3, 13.1, 13.2 Background: Watson & Crick recognized that base pairing in the double helix allows DNA to be copied, or replicated Each strand in the double helix has all the information to remake
Sections 12.3, 13.1, 13.2 Background: Watson & Crick recognized that base pairing in the double helix allows DNA to be copied, or replicated Each strand in the double helix has all the information to remake
Recombinant DNA recombinant DNA DNA cloning gene cloning
 DNA Technology Recombinant DNA In recombinant DNA, DNA from two different sources, often two species, are combined into the same DNA molecule. DNA cloning permits production of multiple copies of a specific
DNA Technology Recombinant DNA In recombinant DNA, DNA from two different sources, often two species, are combined into the same DNA molecule. DNA cloning permits production of multiple copies of a specific
Map-Based Cloning of Qualitative Plant Genes
 Map-Based Cloning of Qualitative Plant Genes Map-based cloning using the genetic relationship between a gene and a marker as the basis for beginning a search for a gene Chromosome walking moving toward
Map-Based Cloning of Qualitative Plant Genes Map-based cloning using the genetic relationship between a gene and a marker as the basis for beginning a search for a gene Chromosome walking moving toward
Flow of Genetic Information
 Flow of Genetic Information DNA Replication Links to the Next Generation Standards Scientific and Engineering Practices: Asking Questions (for science) and Defining Problems (for engineering) Developing
Flow of Genetic Information DNA Replication Links to the Next Generation Standards Scientific and Engineering Practices: Asking Questions (for science) and Defining Problems (for engineering) Developing
i-stop codon positions in the mcherry gene
 Supplementary Figure 1 i-stop codon positions in the mcherry gene The grnas (green) that can potentially generate stop codons from Trp (63 th and 98 th aa, upper panel) and Gln (47 th and 114 th aa, bottom
Supplementary Figure 1 i-stop codon positions in the mcherry gene The grnas (green) that can potentially generate stop codons from Trp (63 th and 98 th aa, upper panel) and Gln (47 th and 114 th aa, bottom
Next Gen Sequencing. Expansion of sequencing technology. Contents
 Next Gen Sequencing Contents 1 Expansion of sequencing technology 2 The Next Generation of Sequencing: High-Throughput Technologies 3 High Throughput Sequencing Applied to Genome Sequencing (TEDed CC BY-NC-ND
Next Gen Sequencing Contents 1 Expansion of sequencing technology 2 The Next Generation of Sequencing: High-Throughput Technologies 3 High Throughput Sequencing Applied to Genome Sequencing (TEDed CC BY-NC-ND
Reservoir of Bacterial Exotoxin Genes in the Environment
 Reservoir of Bacterial Exotoxin Genes in the Environment Veronica Casas, Joseph Magbanua, Gerico Sobrepeña, and Stanley Maloy * Center for Microbial Sciences San Diego State University 5500 Campanile Drive
Reservoir of Bacterial Exotoxin Genes in the Environment Veronica Casas, Joseph Magbanua, Gerico Sobrepeña, and Stanley Maloy * Center for Microbial Sciences San Diego State University 5500 Campanile Drive
8.1. DNA was identified as the genetic material through a series of experiments. Injected mice with R bacteria. Injected mice with S bacteria
 SECTION 8.1 IDENTIFYING DNA AS THE GENETIC MATERIAL Study Guide KEY CONCEPT DNA was identified as the genetic material through a series of experiments. VOCABULARY bacteriophage Griffith finds a transforming
SECTION 8.1 IDENTIFYING DNA AS THE GENETIC MATERIAL Study Guide KEY CONCEPT DNA was identified as the genetic material through a series of experiments. VOCABULARY bacteriophage Griffith finds a transforming
Identification of the Photoreceptor Transcriptional Co-Repressor SAMD11 as Novel Cause of. Autosomal Recessive Retinitis Pigmentosa
 Identification of the Photoreceptor Transcriptional Co-Repressor SAMD11 as Novel Cause of Autosomal Recessive Retinitis Pigmentosa Corton M 1,2 *, Avila-Fernández A 1,2, Campello L 3, Sánchez M 1,2, Benavides
Identification of the Photoreceptor Transcriptional Co-Repressor SAMD11 as Novel Cause of Autosomal Recessive Retinitis Pigmentosa Corton M 1,2 *, Avila-Fernández A 1,2, Campello L 3, Sánchez M 1,2, Benavides
DNA STRUCTURE. Nucleotides: Nitrogenous Bases (Carry the Genetic Code) Expectation Sheet: DNA & Cell Cycle. I can statements: Basic Information:
 Expectation Sheet: DNA & Cell Cycle NAME: Test is 11/8/17 I can statements: I can discuss how DNA is found in all organisms and that the structure is common to all living things. I can diagram and label
Expectation Sheet: DNA & Cell Cycle NAME: Test is 11/8/17 I can statements: I can discuss how DNA is found in all organisms and that the structure is common to all living things. I can diagram and label
Inhibition of HSV-1 Replication by Gene Editing Strategy. Running Title: Targeting HSV-1 Infection by CRISPR/Cas9
 SUPPLEMENTARY MATERIALS 2/4/16 Inhibition of HSV-1 Replication by Gene Editing Strategy Running Title: Targeting HSV-1 Infection by CRISPR/Cas9 Pamela C. Roehm 1,2,3*, Masoud Shekarabi 1, Hassen Wollebo
SUPPLEMENTARY MATERIALS 2/4/16 Inhibition of HSV-1 Replication by Gene Editing Strategy Running Title: Targeting HSV-1 Infection by CRISPR/Cas9 Pamela C. Roehm 1,2,3*, Masoud Shekarabi 1, Hassen Wollebo
Procomcure Biotech. PCR Reagents
 Procomcure Biotech PCR Reagents valid for 2018 VitaTaq DNA Polymerase is a standard Taq DNA polymerase suitable for all common PCR applications like colonypcr, cloning applications, high-throughput PCR
Procomcure Biotech PCR Reagents valid for 2018 VitaTaq DNA Polymerase is a standard Taq DNA polymerase suitable for all common PCR applications like colonypcr, cloning applications, high-throughput PCR
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Number and length distributions of the inferred fosmids.
 Supplementary Figure 1 Number and length distributions of the inferred fosmids. Fosmid were inferred by mapping each pool s sequence reads to hg19. We retained only those reads that mapped to within a
Supplementary Figure 1 Number and length distributions of the inferred fosmids. Fosmid were inferred by mapping each pool s sequence reads to hg19. We retained only those reads that mapped to within a
DNA & DNA Replication
 DNA & DNA Replication DNA Structure How did Watson and Crick contribute to our understanding of genetics? Watson and Crick developed the double helix model for DNA DNA Structure What is a double helix?
DNA & DNA Replication DNA Structure How did Watson and Crick contribute to our understanding of genetics? Watson and Crick developed the double helix model for DNA DNA Structure What is a double helix?
Supplementary Figures
 Supplementary Figures A B Supplementary Figure 1. Examples of discrepancies in predicted and validated breakpoint coordinates. A) Most frequently, predicted breakpoints were shifted relative to those derived
Supplementary Figures A B Supplementary Figure 1. Examples of discrepancies in predicted and validated breakpoint coordinates. A) Most frequently, predicted breakpoints were shifted relative to those derived
Supplemental Figure 1
 Supplemental Figure 1 Supplemental figure 1. Generation of gene targeted mice expressing an anti-id BCR. (A,B) Generation of the VDJ aid H KI mouse. (A) Targeting Construct. Top: Targeting construct for
Supplemental Figure 1 Supplemental figure 1. Generation of gene targeted mice expressing an anti-id BCR. (A,B) Generation of the VDJ aid H KI mouse. (A) Targeting Construct. Top: Targeting construct for
Recitation CHAPTER 9 DNA Technologies
 Recitation CHAPTER 9 DNA Technologies DNA Cloning: General Scheme A cloning vector and eukaryotic chromosomes are separately cleaved with the same restriction endonuclease. (A single chromosome is shown
Recitation CHAPTER 9 DNA Technologies DNA Cloning: General Scheme A cloning vector and eukaryotic chromosomes are separately cleaved with the same restriction endonuclease. (A single chromosome is shown
Marker Antibody Supplier. CD7 CD7 PE-CY 7 anti-human CD7 ebioscience, San Diego, USA
 Supplementary Table 1: Flurochrome labelled antibody used Marker Antibody Supplier CD3 CD4 CD8 CD25 CD26 CD127 CCR4 CCR7 Ki67 Viability stain Alexa Fluor 700 anti-human CD3 Fluorescein isothiocyanate antihuman
Supplementary Table 1: Flurochrome labelled antibody used Marker Antibody Supplier CD3 CD4 CD8 CD25 CD26 CD127 CCR4 CCR7 Ki67 Viability stain Alexa Fluor 700 anti-human CD3 Fluorescein isothiocyanate antihuman
Bioinformatics Support of Genome Sequencing Projects. Seminar in biology
 Bioinformatics Support of Genome Sequencing Projects Seminar in biology Introduction The Big Picture Biology reminder Enzyme for DNA manipulation DNA cloning DNA mapping Sequencing genomes Alignment of
Bioinformatics Support of Genome Sequencing Projects Seminar in biology Introduction The Big Picture Biology reminder Enzyme for DNA manipulation DNA cloning DNA mapping Sequencing genomes Alignment of
Nature Biotechnology: doi: /nbt Supplementary Figure 1. MBQC base beta diversity, major protocol variables, and taxonomic profiles.
 Supplementary Figure 1 MBQC base beta diversity, major protocol variables, and taxonomic profiles. A) Multidimensional scaling of MBQC sample Bray-Curtis dissimilarities (see Fig. 1). Labels indicate centroids
Supplementary Figure 1 MBQC base beta diversity, major protocol variables, and taxonomic profiles. A) Multidimensional scaling of MBQC sample Bray-Curtis dissimilarities (see Fig. 1). Labels indicate centroids
High-resolution Identification and Separation of Living Cell Types by Multiple microrna-responsive Synthetic mrnas
 Supplementary information High-resolution Identification and Separation of Living Cell Types by Multiple microrna-responsive Synthetic mrnas Kei Endo *, Karin Hayashi, Hirohide Saito * Contents: 1. Supplementary
Supplementary information High-resolution Identification and Separation of Living Cell Types by Multiple microrna-responsive Synthetic mrnas Kei Endo *, Karin Hayashi, Hirohide Saito * Contents: 1. Supplementary
Nature Biotechnology: doi: /nbt.4103
 Supplementary Figure 1 Cell types, spatial context and gene expression patterns within the juvenile zebrafish brain. a. Number of cells per cluster from each of the six biological replicates of whole brains
Supplementary Figure 1 Cell types, spatial context and gene expression patterns within the juvenile zebrafish brain. a. Number of cells per cluster from each of the six biological replicates of whole brains
Supplemental Figure 1: Nisoldipine Concentration-Response Relationship on ipsc- CMs. Supplemental Figure 2: Exome Sequencing Prioritization Strategy
 9 10 11 1 1 1 1 1 1 1 19 0 1 9 0 1 Supplementary Appendix Table of Contents Figures: Supplemental Figure 1: Nisoldipine Concentration-Response Relationship on ipsc- CMs Supplemental Figure : Exome Sequencing
9 10 11 1 1 1 1 1 1 1 19 0 1 9 0 1 Supplementary Appendix Table of Contents Figures: Supplemental Figure 1: Nisoldipine Concentration-Response Relationship on ipsc- CMs Supplemental Figure : Exome Sequencing
Ch 12.DNA and RNA.Biology.Landis
 Identity Section 12 1 DNA (pages 287 294) This section tells about the experiments that helped scientists discover the relationship between genes and DNA. It also describes the chemical structure of the
Identity Section 12 1 DNA (pages 287 294) This section tells about the experiments that helped scientists discover the relationship between genes and DNA. It also describes the chemical structure of the
DNA and Biotechnology Form of DNA Form of DNA Form of DNA Form of DNA Replication of DNA Replication of DNA
 21 DNA and Biotechnology DNA and Biotechnology OUTLINE: Replication of DNA Gene Expression Mutations Regulating Gene Activity Genetic Engineering Genomics DNA (deoxyribonucleic acid) Double-stranded molecule
21 DNA and Biotechnology DNA and Biotechnology OUTLINE: Replication of DNA Gene Expression Mutations Regulating Gene Activity Genetic Engineering Genomics DNA (deoxyribonucleic acid) Double-stranded molecule
Name: Family: Date: Monday/Tuesday, March 9,
 Name: Family: Date: Monday/Tuesday, March 9,10 2015 Select the best answer for each question: Part 1: Multiple Choice (2 points each) 1. Protein Synthesis involves which two processes? a. DNA Replication
Name: Family: Date: Monday/Tuesday, March 9,10 2015 Select the best answer for each question: Part 1: Multiple Choice (2 points each) 1. Protein Synthesis involves which two processes? a. DNA Replication
Griffith and Transformation (pages ) 1. What hypothesis did Griffith form from the results of his experiments?
 Section 12 1 DNA (pages 287 294) This section tells about the experiments that helped scientists discover the relationship between genes and DNA. It also describes the chemical structure of the DNA molecule.
Section 12 1 DNA (pages 287 294) This section tells about the experiments that helped scientists discover the relationship between genes and DNA. It also describes the chemical structure of the DNA molecule.
BIO 304 Fall 2000 Exam II Name: ID #: 1. Fill in the blank with the best answer from the provided word bank. (2 pts each)
 1. Fill in the blank with the best answer from the provided word bank. (2 pts each) incomplete dominance conditional mutation penetrance expressivity pleiotropy Southern blotting hybridization epistasis
1. Fill in the blank with the best answer from the provided word bank. (2 pts each) incomplete dominance conditional mutation penetrance expressivity pleiotropy Southern blotting hybridization epistasis
1. The diagram below shows an error in the transcription of a DNA template to messenger RNA (mrna).
 1. The diagram below shows an error in the transcription of a DNA template to messenger RNA (mrna). Which statement best describes the error shown in the diagram? (A) The mrna strand contains the uracil
1. The diagram below shows an error in the transcription of a DNA template to messenger RNA (mrna). Which statement best describes the error shown in the diagram? (A) The mrna strand contains the uracil
Chapter 9. Topics - Genetics - Flow of Genetics - Regulation - Mutation - Recombination
 Chapter 9 Topics - Genetics - Flow of Genetics - Regulation - Mutation - Recombination 1 Genetics Genome Chromosome Gene Protein Genotype Phenotype 2 Terms and concepts gene Fundamental unit of heredity
Chapter 9 Topics - Genetics - Flow of Genetics - Regulation - Mutation - Recombination 1 Genetics Genome Chromosome Gene Protein Genotype Phenotype 2 Terms and concepts gene Fundamental unit of heredity
Nature Structural and Molecular Biology: doi: /nsmb.2847
 Supplementary Figure 1 Specificity coefficients and generality of the TRM code. (A) The ranking of TRM specificity is shown based on the percent enrichment of the dominant base at position +2 multiplied
Supplementary Figure 1 Specificity coefficients and generality of the TRM code. (A) The ranking of TRM specificity is shown based on the percent enrichment of the dominant base at position +2 multiplied
A. I think it is DNA or RNA (circle your answer) because: B. I think it is DNA or RNA (circle your answer) because:
 Name: Test Date: Block: Biology I: Unit 7 Molecular Genetics and Biotechnology Review for Unit Test Directions: You should use this as a guide to help you study for your test. You should also read through
Name: Test Date: Block: Biology I: Unit 7 Molecular Genetics and Biotechnology Review for Unit Test Directions: You should use this as a guide to help you study for your test. You should also read through
DNA DNA Profiling 18. Discuss the stages involved in DNA profiling 19. Define the process of DNA profiling 20. Give two uses of DNA profiling
 Name: 2.5 Genetics Objectives At the end of this sub section students should be able to: 2.5.1 Heredity and Variation 1. Discuss the diversity of organisms 2. Define the term species 3. Distinguish between
Name: 2.5 Genetics Objectives At the end of this sub section students should be able to: 2.5.1 Heredity and Variation 1. Discuss the diversity of organisms 2. Define the term species 3. Distinguish between
Human Genetic Variation. Ricardo Lebrón Dpto. Genética UGR
 Human Genetic Variation Ricardo Lebrón rlebron@ugr.es Dpto. Genética UGR What is Genetic Variation? Origins of Genetic Variation Genetic Variation is the difference in DNA sequences between individuals.
Human Genetic Variation Ricardo Lebrón rlebron@ugr.es Dpto. Genética UGR What is Genetic Variation? Origins of Genetic Variation Genetic Variation is the difference in DNA sequences between individuals.
INSECT-TRANSMITTED PROCARYOTES
 Fourteenth IOCV Conference, 2000 Insect-Transmitted Procaryotes INSECT-TRANSMITTED PROCARYOTES Improved Sensitivity in the Detection and Differentiation of Citrus Huanglongbing Bacteria from South Africa
Fourteenth IOCV Conference, 2000 Insect-Transmitted Procaryotes INSECT-TRANSMITTED PROCARYOTES Improved Sensitivity in the Detection and Differentiation of Citrus Huanglongbing Bacteria from South Africa
Genome Assembly Using de Bruijn Graphs. Biostatistics 666
 Genome Assembly Using de Bruijn Graphs Biostatistics 666 Previously: Reference Based Analyses Individual short reads are aligned to reference Genotypes generated by examining reads overlapping each position
Genome Assembly Using de Bruijn Graphs Biostatistics 666 Previously: Reference Based Analyses Individual short reads are aligned to reference Genotypes generated by examining reads overlapping each position
