Supplementary Figure 1
|
|
|
- Alexandra Allen
- 5 years ago
- Views:
Transcription
1 Supplementary Figure 1 Rarefaction Analysis and Sampling. a, Rarefaction plots show the expected levels of coverage for different subsamples of sequencing libraries for repertoires of clone size of at least 2 (darkest blue line), 10 (second darkest blue), 20 (red, the cut-off chosen for the overlap analysis), 30 (light blue), 40 (lighter blue) or 50 and greater (light grey) in the repertoire of D207. Separate plots are shown for each tissue. b, Sampling needed for clones of different sizes and coverage thresholds in D207. Bars indicate the number of additional libraries needed to sample 0.5,0.75,0.9 and 0.95 (fractions) of all clones at different clone size cut-offs. Colors of the bars indicate the tissue. PBL = peripheral blood; BM = bone marrow; SPL = spleen; MLN = mesenteric lymph node.
2 Supplementary Figure 2
3 Cosine Similarity Analysis. a, Box plots of distribution of cosine similarity between all sample pairs within a tissue. Similarity is assessed for C 20 clones, C 20 clones without the top 10 clones in each tissue, all clones, or all clones without the top 10 copy number clones in each tissue. Boxes represent the first and third quartiles bisected by the median. Whiskers represent the most extreme data excluding outliers, where outliers (dots) are data beyond the third or first quartile by a distance exceeding 1.5 times the inter-quartile interval. The C 20 clone panel for D207 corresponds to Fig. 1b in the main manuscript. b, The cosine similarity of clonal populations between tissue pairs in D181 and D207 with and without blood clones. Each section represents a tissue and each arrow within each section represents the level of overlap (cosine similarity) between that tissue and clones from other tissues. Longer arrows indicate higher value of cosine similarity indicating more overlap of clonal populations between the tissues. Top: all C 20 clones (same as Fig. 1c). Bottom: C 20 overlap without blood clones; clones that have sequences in the blood were computationally removed from all tissues.
4 Supplementary Figure 3 Analysis of Clonal Networks in D207 Using Different Clone Size Definitions. Clonal overlap was evaluated using cosine similarities between different tissue pairs, as described in the Online Methods. Each wedge represents a tissue and each arrow represents the cosine similarity between the tissues (arrow colors are indicated in the legend, on the left). Three different definitions of clone size are used for the analysis: a, the copy number of all sequences in a clone in a given tissue; b, the total number of unique sequence instances in a clone in a given tissue; and c, the definition that we used in the manuscript (same image as Fig. 1c) - the number of independent sequencing library samples in which the clone appears, in a tissue.
5 D181 D207 Fraction of Clones Clone Size (binned) Supplementary Figure 4 Clone Sizes Grouped by Clone Category. Sizes of large clones (in binned clone sizes of sequence instances, see Online Methods), grouped by tissue distribution category in D181 (left) and D207 (right).
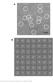
6 Supplementary Figure 5 Large Clone Tissue Distribution Types in D207 C 20 clones (including clones in the blood): a, Single tissue clones; b, Two-tissue clones; c, Regional (found in 3-5 tissues) and d, Global clones (found in 6-8 tissues). Tissues that contain only a single instance of a clone are not counted towards the distribution pattern of that clone. Each line is a clone. The position along the line represents a tissue. The size of the circles at each position represents the total number of instances the clones have in each tissue. The colored wedges represent the fraction of sequencing libraries that contain at least one instance of the clone in each tissue. A sample clone is shown for each tissue distribution pattern. The number to the left of each line is that clone s unique clone ID in The number to the right is the number of different clones with this tissue distribution pattern. p = peripheral blood; b = bone marrow; s = spleen; l = lung; m = mesenteric lymph node; j = jejunum; i= ileum; c = colon.
7 Supplementary Figure 6 Large Clone Tissue Distribution Types in D181. C 20 clones (including clones in the blood): a, Single tissue clones; b, Two-tissue clones; c, Regional (found in 3-5 tissues) and d, Global clones (found in 6-8 tissues). Tissues that contain only a single instance of a clone are not counted towards the distribution pattern of that clone. Each line is a clone. The position along the line represents a tissue. The size of the circles at each position represents the total number of instances the clones have in each tissue. The colored wedges represent the fraction of sequencing libraries that contain at least one instance of the clone in each tissue. A sample clone is shown for each tissue distribution pattern. The number to the left of each line is that clone s unique clone ID in The number to the right is the number of different clones with this tissue distribution pattern. p = peripheral blood; b = bone marrow; s = spleen; l = lung; m = mesenteric lymph node; j = jejunum; i= ileum; c = colon.
8 a 0.01% 0.005% 0.001% PB###BM##SPL##Lun##MLN#Jej###Ile##Col# PB###BM##SPL##Lun##MLN#Jej###Ile##Col# PB###BM##SPL##Lun##MLN#Jej###Ile##Col# b C20 Clone Number D181 C20 Clone Number D207 Single Double Regional Global 0 Bucket 85% 90% 0 Bucket 85% 90% c 85% 90% Supplementary Figure 7 Effects of Clone Size Thresholding on Clone Tissue Membership. a, Similar blood and GI tract networks are seen at different clone size cut-offs. Overlapping clones are plotted at different clone size cut-offs (greater than or equal to 0.01%, 0.005% and 0.001% of total copies of all of the sequencing libraries within each donor). Each horizontal line represents a clone. Networks of overlapping clones were visualized using MiXCR and VDJtools software (see Online Methods). Only clones from D207 that overlapped between two or more tissues are shown. PBL = peripheral blood; BM = bone marrow; SPL = spleen; Lun = lung; MLN = mesenteric lymph node; Jej = jejunum; Ile = ileum; Col = colon. b, Breadth of tissue membership using different CDR3 identity thresholds. The tissue distribution of C 20 clones in the two most deeply sequenced donors was studied using: 85% amino acid identity in CDR3 (the cut-off used in the main manuscript), 90% amino acid identity and no requirement for any CDR3 identity ( bucket ). All of the sequences were collapsed at these different CDR3 similarity thresholds into clones. Then C 20 clones were
9 analyzed for their tissue representation: single tissue, double (2 tissues), regional (3-5 tissues) and global (6-8 tissues). The C 20 clones were recalculated for each threshold. c, Comparable partitioning of clones from D207 into blood and GI tract networks at two different CDR3 amino acid similarity thresholds. Networks of C 20 clones with 85% CDR3 identity (left image is from Fig. 1c in the main manuscript) and 90% CDR3 amino acid identity (right panel) were analyzed using the cosine statistic (see Online Methods). Cosine similarity for two-tissue comparisons is plotted using the color scheme defined in the legend.
10 D207 D181 D145 PBL BM SPL Lun MLN Jej Ile Col PBL BM SPL Lun MLN Jej Ile Col PBL BM SPL Lun MLN Jej Ile Col D149 D168 D182 PBL BM SPL Lun MLN Jej Ile Col PBL BM SPL Lun MLN Jej Ile Col PBL BM SPL Lun MLN Jej Ile Col Supplementary Figure 8 Blood and GI Tract Networks are Present in All Donors. Clonal overlap was evaluated in all donors with clones that comprised 0.01% or more of total copies. Each horizontal line represents a clone. Networks of overlapping clones were visualized using MiXCR and VDJtools software (see Online Methods). Only clones that overlapped between two or more tissues are shown. The MLN sample in D168 was hypocellular. PBL = peripheral blood; BM = bone marrow; SPL = spleen; Lun = lung; MLN = mesenteric lymph node; Jej = jejunum; Ile = ileum; Col = colon.
11
12 Supplementary Figure 9 Clonal Lineage Networks in D207 with and without Blood Clones Each horizontal line represents a clone with at least 20 unique sequence instances in one tissue that overlaps with at least one other tissue, each column represents a tissue. All sub plots are identical, but are colored differently to highlight clones that reside in different tissues. In each instance, the clones that overlap in a specific tissue are colored as indicated by the tissue listed at the top of each panel. All other clones are denoted in black. a, Blood tissue clones highlighted; b, GI tract clones highlighted. Top row: all C 20 clones. Bottom row: blood C 20 clones computationally excluded (any clone with sequences found in the blood is removed from the analysis). Clones are ordered by their tissue membership.
13
14 Supplementary Figure 10 Clonal Lineage Networks in D181 with and without Blood Clones. Each horizontal line represents a clone with at least 20 unique sequence instances in one tissue that overlaps with at least one other tissue, each column represents a tissue. All sub plots are identical, but are colored differently to highlight clones that reside in different tissues. In each instance, the clones that overlap in a specific tissue are colored as indicated by the tissue listed at the top of each panel. All other clones are denoted in black. a, Blood tissue clones highlighted; b, GI tract clones highlighted. Top row: all C 20 clones. Bottom row: blood C 20 clones computationally excluded (any clone with sequences found in the blood is removed from the analysis). Clones are ordered by their tissue membership.
15 Supplementary Figure 11 Sequence Mixing within Clonal Lineages. a (inset), Numbers of clones used for the clumpiness analysis. Shown are the numbers of clones with nodes in each pair of tissues. This analysis is restricted to C 20 clones in D207; these numbers correspond to the clones analyzed in Fig. 3b. b, Sequence mixing within clonal lineages varies between different tissue types. The frequency distributions of clumpiness values between two tissues for all clones with nodes in each tissue pair (combinations are listed at the top of each plot). This analysis was performed on C 20 clones in D207.
16 Supplementary Table 1: PCR Primers. Shown are the nucleotide sequences of the primers used for PCR amplifications (see Methods for PCR details).
17 Supplementary Table 2: Donor Tissue Amplicon Libraries. Shown are the numbers of sequencing libraries for each donor tissue. LD = leader primer mixes; FR1 = framework region 1 primer mix.
An atlas of B-cell clonal distribution in the human body
 An atlas of B-cell clonal distribution in the human body 27 Nature America, Inc., part of Springer Nature. All rights reserved. Wenzhao Meng,8, Bochao Zhang 2,8, Gregory W Schwartz 2, Aaron M Rosenfeld
An atlas of B-cell clonal distribution in the human body 27 Nature America, Inc., part of Springer Nature. All rights reserved. Wenzhao Meng,8, Bochao Zhang 2,8, Gregory W Schwartz 2, Aaron M Rosenfeld
Nature Immunology: doi: /ni Supplementary Figure 1. Data-processing pipeline.
 Supplementary Figure 1 Data-processing pipeline. Steps for processing data from multiple sorted B cell populations derived from a single individual at a single time point are shown. Parameters used are
Supplementary Figure 1 Data-processing pipeline. Steps for processing data from multiple sorted B cell populations derived from a single individual at a single time point are shown. Parameters used are
SUPPLEMENTARY INFORMATION
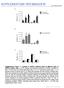 SUPPLEMENTARY INFORMATION doi:10.1038/nature12474 Supplementary Figure 1 Analysis of mtdna mutation loads in different types of mtdna mutator mice. a, PCR, cloning, and sequencing analysis of mtdna mutations
SUPPLEMENTARY INFORMATION doi:10.1038/nature12474 Supplementary Figure 1 Analysis of mtdna mutation loads in different types of mtdna mutator mice. a, PCR, cloning, and sequencing analysis of mtdna mutations
Mapping long-range promoter contacts in human cells with high-resolution capture Hi-C
 CORRECTION NOTICE Nat. Genet. 47, 598 606 (2015) Mapping long-range promoter contacts in human cells with high-resolution capture Hi-C Borbala Mifsud, Filipe Tavares-Cadete, Alice N Young, Robert Sugar,
CORRECTION NOTICE Nat. Genet. 47, 598 606 (2015) Mapping long-range promoter contacts in human cells with high-resolution capture Hi-C Borbala Mifsud, Filipe Tavares-Cadete, Alice N Young, Robert Sugar,
Nature Methods: doi: /nmeth Supplementary Figure 1. Pilot CrY2H-seq experiments to confirm strain and plasmid functionality.
 Supplementary Figure 1 Pilot CrY2H-seq experiments to confirm strain and plasmid functionality. (a) RT-PCR on HIS3 positive diploid cell lysate containing known interaction partners AT3G62420 (bzip53)
Supplementary Figure 1 Pilot CrY2H-seq experiments to confirm strain and plasmid functionality. (a) RT-PCR on HIS3 positive diploid cell lysate containing known interaction partners AT3G62420 (bzip53)
Revised: RG-RV2 by Fukuhara et al.
 Supplemental Figure 1 The generation of Spns2 conditional knockout mice. (A) Schematic representation of the wild type Spns2 locus (Spns2 + ), the targeted allele, the floxed allele (Spns2 f ) and the
Supplemental Figure 1 The generation of Spns2 conditional knockout mice. (A) Schematic representation of the wild type Spns2 locus (Spns2 + ), the targeted allele, the floxed allele (Spns2 f ) and the
Supplementary Figure and Table Legends
 1 Supplementary Figure and Table Legends Figure S1: Whole-animal metabolic analysis. 12 week old WT and Dvl1 / were singly housed in CLAMS cages (Comprehensive Laboratory Animals Monitoring System) for
1 Supplementary Figure and Table Legends Figure S1: Whole-animal metabolic analysis. 12 week old WT and Dvl1 / were singly housed in CLAMS cages (Comprehensive Laboratory Animals Monitoring System) for
Supporting Information
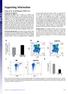 (xe number cells) LSK (% gated) Supporting Information Youm et al../pnas. SI Materials and Methods Quantification of sjtrecs. CD + T subsets were isolated from splenocytes using mouse CD + T cells positive
(xe number cells) LSK (% gated) Supporting Information Youm et al../pnas. SI Materials and Methods Quantification of sjtrecs. CD + T subsets were isolated from splenocytes using mouse CD + T cells positive
SUPPLEMENTARY INFORMATION
 Contents De novo assembly... 2 Assembly statistics for all 150 individuals... 2 HHV6b integration... 2 Comparison of assemblers... 4 Variant calling and genotyping... 4 Protein truncating variants (PTV)...
Contents De novo assembly... 2 Assembly statistics for all 150 individuals... 2 HHV6b integration... 2 Comparison of assemblers... 4 Variant calling and genotyping... 4 Protein truncating variants (PTV)...
Supplemental Figure 1
 Supplemental Figure 1 A P1 NOD - SCID(X*X*)(0/0) NOD - SCID(X*Y)(A*0201/A*0201) B F1 (X*X)(A*0201/0) (X*X*)(0/0) F2 (X*X*)(0/0) (X*Y)(0/0) (X*X*)(A*0201/0) (X*X*)(A*0201/0) F3 (X*X*)(0/0) (X*X)(A*0201/0)
Supplemental Figure 1 A P1 NOD - SCID(X*X*)(0/0) NOD - SCID(X*Y)(A*0201/A*0201) B F1 (X*X)(A*0201/0) (X*X*)(0/0) F2 (X*X*)(0/0) (X*Y)(0/0) (X*X*)(A*0201/0) (X*X*)(A*0201/0) F3 (X*X*)(0/0) (X*X)(A*0201/0)
Supplementary figures
 Mucida et al. Supplementary material Supplementary figures Supplementary Figure 1. Oral administration of OVA suppresses Th2 differentiation, Germinal Center (GC) formation and immunoglobulin class switching
Mucida et al. Supplementary material Supplementary figures Supplementary Figure 1. Oral administration of OVA suppresses Th2 differentiation, Germinal Center (GC) formation and immunoglobulin class switching
SUPPLEMENTARY MATERIAL AND METHODS
 SUPPLEMENTARY MATERIAL AND METHODS Amplification of HEV ORF1, ORF2 and ORF3 genome regions Total RNA was extracted from 200 µl EDTA plasma using Cobas AmpliPrep total nucleic acid isolation kit (Roche,
SUPPLEMENTARY MATERIAL AND METHODS Amplification of HEV ORF1, ORF2 and ORF3 genome regions Total RNA was extracted from 200 µl EDTA plasma using Cobas AmpliPrep total nucleic acid isolation kit (Roche,
Supplementary Information for Single-cell sequencing of the small-rna transcriptome
 Supplementary Information for Single-cell sequencing of the small-rna transcriptome Omid R. Faridani 1,6,*, Ilgar Abdullayev 1,2,6, Michael Hagemann-Jensen 1,3, John P. Schell 4, Fredrik Lanner 4,5 and
Supplementary Information for Single-cell sequencing of the small-rna transcriptome Omid R. Faridani 1,6,*, Ilgar Abdullayev 1,2,6, Michael Hagemann-Jensen 1,3, John P. Schell 4, Fredrik Lanner 4,5 and
Applications of the Ion AmpliSeq Immune Repertoire Assay Plus TCRβ
 Applications of the Ion AmpliSeq Immune Repertoire Assay Plus TCRβ Timothy Looney, PhD Staff Scientist, Clinical Next-Generation Sequencing Division Thermo Fisher Scientific The world leader in serving
Applications of the Ion AmpliSeq Immune Repertoire Assay Plus TCRβ Timothy Looney, PhD Staff Scientist, Clinical Next-Generation Sequencing Division Thermo Fisher Scientific The world leader in serving
(Hadlock, Journal of Virology, 2000), (Keck, Journal of Virology, 2008) CBH-5 Domain B 412, 416, 417, 418, 420, 421, 422, 423, 483, 484, 485, 488,
 Supplementary Table. mabs analyzed mab Antigenic Critical Binding Residues by Citations Domains Alanine Scanning 2 CBH-4B Domain A NA (Hadlock, Journal of Virology, 2), (Keck, Journal of Virology, 24)
Supplementary Table. mabs analyzed mab Antigenic Critical Binding Residues by Citations Domains Alanine Scanning 2 CBH-4B Domain A NA (Hadlock, Journal of Virology, 2), (Keck, Journal of Virology, 24)
Parts of a standard FastQC report
 FastQC FastQC, written by Simon Andrews of Babraham Bioinformatics, is a very popular tool used to provide an overview of basic quality control metrics for raw next generation sequencing data. There are
FastQC FastQC, written by Simon Andrews of Babraham Bioinformatics, is a very popular tool used to provide an overview of basic quality control metrics for raw next generation sequencing data. There are
Applications of AmpliSeq-based Ion Torrent TCRB Immune Repertoire Sequencing
 Applications of AmpliSeq-based Ion Torrent TCRB Immune Repertoire Sequencing Timothy Looney, PhD Staff Scientist, Clinical Next-Generation Sequencing Division Thermo Fisher Scientific The world leader
Applications of AmpliSeq-based Ion Torrent TCRB Immune Repertoire Sequencing Timothy Looney, PhD Staff Scientist, Clinical Next-Generation Sequencing Division Thermo Fisher Scientific The world leader
with different microfluidic device parameters. Distribution of number of transcripts (y-axis) detected across nuclei, ranked in decreasing
 Supplementary Figure 1 DroNc-seq benchmarking experiments. (a) Overview of DroNc-seq. (b) CAD schema of DroNc-seq microfluidic device. (c) Comparison of library complexity from experiments with different
Supplementary Figure 1 DroNc-seq benchmarking experiments. (a) Overview of DroNc-seq. (b) CAD schema of DroNc-seq microfluidic device. (c) Comparison of library complexity from experiments with different
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for Integrative Biology. This journal is The Royal Society of Chemistry 2015 Electronic Supplementary Information Table S1. Definition of quantitative cellular features
Electronic Supplementary Material (ESI) for Integrative Biology. This journal is The Royal Society of Chemistry 2015 Electronic Supplementary Information Table S1. Definition of quantitative cellular features
Supplementary Figure 1
 Supplementary Figure 1 Concept of barcoding to suppress error in sequencing. Each template DNA molecule is barcoded with a random and unique sequence (marked as red, turquoise and green). All PCR generated
Supplementary Figure 1 Concept of barcoding to suppress error in sequencing. Each template DNA molecule is barcoded with a random and unique sequence (marked as red, turquoise and green). All PCR generated
Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions. Supplementary Material
 Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions Joshua N. Burton 1, Andrew Adey 1, Rupali P. Patwardhan 1, Ruolan Qiu 1, Jacob O. Kitzman 1, Jay Shendure 1 1 Department
Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions Joshua N. Burton 1, Andrew Adey 1, Rupali P. Patwardhan 1, Ruolan Qiu 1, Jacob O. Kitzman 1, Jay Shendure 1 1 Department
Supplementary information: Optimization of design and production strategies for novel adeno-associated viral display peptide libraries
 Supplementary information: Optimization of design and production strategies for novel adeno-associated viral display peptide libraries Jakob Körbelin, Agnes Hunger, Malik Alawi, Timo Sieber, Mascha Binder,
Supplementary information: Optimization of design and production strategies for novel adeno-associated viral display peptide libraries Jakob Körbelin, Agnes Hunger, Malik Alawi, Timo Sieber, Mascha Binder,
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Optimization of Patch-seq protocol.
 Supplementary Figure 1 Optimization of Patch-seq protocol. Pilot experiments were carried out to test the effect of various protocol modifications on cdna yield, including addition of RNase inhibitor to
Supplementary Figure 1 Optimization of Patch-seq protocol. Pilot experiments were carried out to test the effect of various protocol modifications on cdna yield, including addition of RNase inhibitor to
Nature Biotechnology: doi: /nbt.4103
 Supplementary Figure 1 Cell types, spatial context and gene expression patterns within the juvenile zebrafish brain. a. Number of cells per cluster from each of the six biological replicates of whole brains
Supplementary Figure 1 Cell types, spatial context and gene expression patterns within the juvenile zebrafish brain. a. Number of cells per cluster from each of the six biological replicates of whole brains
SUPPLEMENTARY INFORMATION
 DOI: 1.138/ncb3342 EV O/E 13 4 B Relative expression 1..8.6.4.2 shctl Pcdh2_1 C Pcdh2_2 Number of shrns 12 8 4 293T mterc +/+ G3 mterc -/- D % of GFP + cells 1 8 6 4 2 Per2 p=.2 p>
DOI: 1.138/ncb3342 EV O/E 13 4 B Relative expression 1..8.6.4.2 shctl Pcdh2_1 C Pcdh2_2 Number of shrns 12 8 4 293T mterc +/+ G3 mterc -/- D % of GFP + cells 1 8 6 4 2 Per2 p=.2 p>
Who pairs with whom? High-throughput sequencing of the human paired heavy and light chain repertoire
 Who pairs with whom? High-throughput sequencing of the human paired heavy and light chain repertoire Technical Journal Club September 15 th Christina Müller Background - antibody repertoire is the sum
Who pairs with whom? High-throughput sequencing of the human paired heavy and light chain repertoire Technical Journal Club September 15 th Christina Müller Background - antibody repertoire is the sum
Supplementary Figures
 Supplementary Figures Supplementary Figure 1: Output of pipeline a, Each dot corresponds to a tumor, with vertical position indicating the total number of fusions identified. Tumor types are ordered by
Supplementary Figures Supplementary Figure 1: Output of pipeline a, Each dot corresponds to a tumor, with vertical position indicating the total number of fusions identified. Tumor types are ordered by
MassArray Analytical Tools
 MassArray Analytical Tools Reid F. Thompson and John M. Greally April 30, 08 Contents Introduction Changes for MassArray in current BioC release 3 3 Optimal Amplicon Design 4 4 Conversion Controls 0 5
MassArray Analytical Tools Reid F. Thompson and John M. Greally April 30, 08 Contents Introduction Changes for MassArray in current BioC release 3 3 Optimal Amplicon Design 4 4 Conversion Controls 0 5
Thornhill et al. ME Supplementary Materials revision 08/2007 1
 Thornhill et al. ME 07-415 Supplementary Materials revision 08/2007 1 Supplementary Table 1 Summary of clones obtained from estimates of Taq polymerase/bacterial cloning error in Symbiodinium type A3 (culture
Thornhill et al. ME 07-415 Supplementary Materials revision 08/2007 1 Supplementary Table 1 Summary of clones obtained from estimates of Taq polymerase/bacterial cloning error in Symbiodinium type A3 (culture
RNA-Seq Analysis. August Strand Genomics, Inc All rights reserved.
 RNA-Seq Analysis August 2014 Strand Genomics, Inc. 2014. All rights reserved. Contents Introduction... 3 Sample import... 3 Quantification... 4 Novel exon... 5 Differential expression... 12 Differential
RNA-Seq Analysis August 2014 Strand Genomics, Inc. 2014. All rights reserved. Contents Introduction... 3 Sample import... 3 Quantification... 4 Novel exon... 5 Differential expression... 12 Differential
Supplementary Material for
 www.sciencemag.org/cgi/content/full/science.aaa6090/dc1 Supplementary Material for Spatially resolved, highly multiplexed RNA profiling in single cells Kok Hao Chen, Alistair N. Boettiger, Jeffrey R. Moffitt,
www.sciencemag.org/cgi/content/full/science.aaa6090/dc1 Supplementary Material for Spatially resolved, highly multiplexed RNA profiling in single cells Kok Hao Chen, Alistair N. Boettiger, Jeffrey R. Moffitt,
points in a line over time.
 Chart types Published: 2018-07-07 Dashboard charts in the ExtraHop system offer multiple ways to visualize metric data, which can help you answer questions about your network behavior. You select a chart
Chart types Published: 2018-07-07 Dashboard charts in the ExtraHop system offer multiple ways to visualize metric data, which can help you answer questions about your network behavior. You select a chart
Tutorial. Primer Design. Sample to Insight. November 27, 2015
 Primer Design November 27, 2015 Sample to Insight CLC bio, a QIAGEN Company Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 www.clcbio.com support-clcbio@qiagen.com Primer Design
Primer Design November 27, 2015 Sample to Insight CLC bio, a QIAGEN Company Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 www.clcbio.com support-clcbio@qiagen.com Primer Design
SUPPLEMENTARY INFORMATION. Supplementary Figures 1-8
 SUPPLEMENTARY INFORMATION Supplementary Figures 1-8 Supplementary Figure 1. TFAM residues contacting the DNA minor groove (A) TFAM contacts on nonspecific DNA. Leu58, Ile81, Asn163, Pro178, and Leu182
SUPPLEMENTARY INFORMATION Supplementary Figures 1-8 Supplementary Figure 1. TFAM residues contacting the DNA minor groove (A) TFAM contacts on nonspecific DNA. Leu58, Ile81, Asn163, Pro178, and Leu182
Supporting Information
 Supporting Information Table S1. Overview of samples used for sequencing, and the number of sequences obtained from each sample. Visit 1 is day 0, Visit 2 is day 7, Visit 3 is day 28, and Visit 4 is day
Supporting Information Table S1. Overview of samples used for sequencing, and the number of sequences obtained from each sample. Visit 1 is day 0, Visit 2 is day 7, Visit 3 is day 28, and Visit 4 is day
Supplementary Figure 1 An overview of pirna biogenesis during fetal mouse reprogramming. (a) (b)
 Supplementary Figure 1 An overview of pirna biogenesis during fetal mouse reprogramming. (a) A schematic overview of the production and amplification of a single pirna from a transposon transcript. The
Supplementary Figure 1 An overview of pirna biogenesis during fetal mouse reprogramming. (a) A schematic overview of the production and amplification of a single pirna from a transposon transcript. The
Application Note. NGS Analysis of B-Cell Receptors & Antibodies by AptaAnalyzer -BCR
 Reduce to the Best Application Note NGS Analysis of B-Cell Receptors & Antibodies by AptaAnalyzer -BCR The software AptaAnalyzer harnesses next generation sequencing (NGS) data to monitor the immune response
Reduce to the Best Application Note NGS Analysis of B-Cell Receptors & Antibodies by AptaAnalyzer -BCR The software AptaAnalyzer harnesses next generation sequencing (NGS) data to monitor the immune response
Nature Biotechnology: doi: /nbt Supplementary Figure 1. MBQC base beta diversity, major protocol variables, and taxonomic profiles.
 Supplementary Figure 1 MBQC base beta diversity, major protocol variables, and taxonomic profiles. A) Multidimensional scaling of MBQC sample Bray-Curtis dissimilarities (see Fig. 1). Labels indicate centroids
Supplementary Figure 1 MBQC base beta diversity, major protocol variables, and taxonomic profiles. A) Multidimensional scaling of MBQC sample Bray-Curtis dissimilarities (see Fig. 1). Labels indicate centroids
Supplemental Data. Guo et al. (2015). Plant Cell /tpc
 Supplemental Figure 1. The Mutant exb1-d Displayed Pleiotropic Phenotypes and Produced Branches in the Axils of Cotyledons. (A) Branches were developed in exb1-d but not in wild-type plants. (B) and (C)
Supplemental Figure 1. The Mutant exb1-d Displayed Pleiotropic Phenotypes and Produced Branches in the Axils of Cotyledons. (A) Branches were developed in exb1-d but not in wild-type plants. (B) and (C)
Nature Methods: doi: /nmeth Supplementary Figure 1
 Supplementary Figure 1 Workflow for multimodal analysis using sc-gem on a programmable microfluidic device (Fluidigm). 1) Cells are captured and lysed, 2) RNA from lysed single cell is reverse-transcribed
Supplementary Figure 1 Workflow for multimodal analysis using sc-gem on a programmable microfluidic device (Fluidigm). 1) Cells are captured and lysed, 2) RNA from lysed single cell is reverse-transcribed
User Bulletin. Veriti 96-Well Thermal Cycler AmpFlSTR Kit Validation. Overview
 User Bulletin Veriti 96-Well Thermal Cycler AmpFlSTR Kit Validation June 2009 SUBJECT: AmpFlSTR PCR Amplification Kit validation on the Veriti 96-Well Thermal Cycler with 0.2 ml sample block format In
User Bulletin Veriti 96-Well Thermal Cycler AmpFlSTR Kit Validation June 2009 SUBJECT: AmpFlSTR PCR Amplification Kit validation on the Veriti 96-Well Thermal Cycler with 0.2 ml sample block format In
Supplementary Figure 1: (a) 3D illustration of the mobile phone microscopy attachment, containing a 3D movable sample holder (red piece for z
 Supplementary Figure 1: (a) 3D illustration of the mobile phone microscopy attachment, containing a 3D movable sample holder (red piece for z movement and blue piece for x-y movement). Illumination sources
Supplementary Figure 1: (a) 3D illustration of the mobile phone microscopy attachment, containing a 3D movable sample holder (red piece for z movement and blue piece for x-y movement). Illumination sources
Wu et al., Determination of genetic identity in therapeutic chimeric states. We used two approaches for identifying potentially suitable deletion loci
 SUPPLEMENTARY METHODS AND DATA General strategy for identifying deletion loci We used two approaches for identifying potentially suitable deletion loci for PDP-FISH analysis. In the first approach, we
SUPPLEMENTARY METHODS AND DATA General strategy for identifying deletion loci We used two approaches for identifying potentially suitable deletion loci for PDP-FISH analysis. In the first approach, we
Supporting Information
 Supporting Information Tomura et al. 1.173/pnas.8227815 WT CFSE Con A stimulation Before After 2 3 1 CFSE Kaede- Tg No photoconversion Photoconversion Kaede Red Fig. S1. Dilution of photoconverted Kaede
Supporting Information Tomura et al. 1.173/pnas.8227815 WT CFSE Con A stimulation Before After 2 3 1 CFSE Kaede- Tg No photoconversion Photoconversion Kaede Red Fig. S1. Dilution of photoconverted Kaede
Supplementary Figure 1. Generation of B2M -/- ESCs. Nature Biotechnology: doi: /nbt.3860
 Supplementary Figure 1 Generation of B2M -/- ESCs. (a) Maps of the B2M alleles in cells with the indicated B2M genotypes. Probes and restriction enzymes used in Southern blots are indicated (H, Hind III;
Supplementary Figure 1 Generation of B2M -/- ESCs. (a) Maps of the B2M alleles in cells with the indicated B2M genotypes. Probes and restriction enzymes used in Southern blots are indicated (H, Hind III;
initial single-cell analysis, with a pragmatic focus on surface markers with the highest potential for
 Supplementary Figure 1: Summary of the exclusionary approach to surface marker selection for initial single-cell analysis, with a pragmatic focus on surface markers with the highest potential for protein
Supplementary Figure 1: Summary of the exclusionary approach to surface marker selection for initial single-cell analysis, with a pragmatic focus on surface markers with the highest potential for protein
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Number and length distributions of the inferred fosmids.
 Supplementary Figure 1 Number and length distributions of the inferred fosmids. Fosmid were inferred by mapping each pool s sequence reads to hg19. We retained only those reads that mapped to within a
Supplementary Figure 1 Number and length distributions of the inferred fosmids. Fosmid were inferred by mapping each pool s sequence reads to hg19. We retained only those reads that mapped to within a
Map-Based Cloning of Qualitative Plant Genes
 Map-Based Cloning of Qualitative Plant Genes Map-based cloning using the genetic relationship between a gene and a marker as the basis for beginning a search for a gene Chromosome walking moving toward
Map-Based Cloning of Qualitative Plant Genes Map-based cloning using the genetic relationship between a gene and a marker as the basis for beginning a search for a gene Chromosome walking moving toward
Nature Biotechnology: doi: /nbt.4086
 Ag (-) anti-cd3 p815 p815-hcd2 Ag (-) anti-cd3 p815 p815-hcd2 Ag (-) anti-cd3 p815 p815-hcd2 Ag (-) anti-cd3 p815 p815-hcd2 Ag (-) anti-cd3 p815 p815-hcd2 Ag (-) anti-cd3 p815 p815-hcd2 Ag (-) anti-cd3
Ag (-) anti-cd3 p815 p815-hcd2 Ag (-) anti-cd3 p815 p815-hcd2 Ag (-) anti-cd3 p815 p815-hcd2 Ag (-) anti-cd3 p815 p815-hcd2 Ag (-) anti-cd3 p815 p815-hcd2 Ag (-) anti-cd3 p815 p815-hcd2 Ag (-) anti-cd3
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10772 Supplementary Figures: Supplementary Figure 1. Location of CNS1 within of the Foxp3 locus highlighting CNS1 and indicating transcription factor binding motifs downstream of TCR,
doi:10.1038/nature10772 Supplementary Figures: Supplementary Figure 1. Location of CNS1 within of the Foxp3 locus highlighting CNS1 and indicating transcription factor binding motifs downstream of TCR,
Genome-wide genetic screening with chemically-mutagenized haploid embryonic stem cells
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 Supplementary Information Genome-wide genetic screening with chemically-mutagenized haploid embryonic stem cells Josep V. Forment 1,2, Mareike Herzog
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 Supplementary Information Genome-wide genetic screening with chemically-mutagenized haploid embryonic stem cells Josep V. Forment 1,2, Mareike Herzog
Multiple layers of B cell memory with different effector functions. Ismail Dogan, Barbara Bertocci, Valérie Vilmont, Frédéric Delbos,
 Multiple layers of B cell memory with different effector functions Ismail Dogan, Barbara Bertocci, Valérie Vilmont, Frédéric Delbos, Jérome Mégret, Sébastien Storck, Claude-Agnès Reynaud & Jean-Claude
Multiple layers of B cell memory with different effector functions Ismail Dogan, Barbara Bertocci, Valérie Vilmont, Frédéric Delbos, Jérome Mégret, Sébastien Storck, Claude-Agnès Reynaud & Jean-Claude
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature11809 Supplementary Figure 1. Antibiotic treatment reduces intestinal bacterial load and allows access of non-pathogenic bacteria to the MLN, inducing intestinal
SUPPLEMENTARY INFORMATION doi:10.1038/nature11809 Supplementary Figure 1. Antibiotic treatment reduces intestinal bacterial load and allows access of non-pathogenic bacteria to the MLN, inducing intestinal
Metagenomic species profiling using universal phylogenetic marker genes
 Metagenomic species profiling using universal phylogenetic marker genes Shinichi Sunagawa, Daniel R. Mende, Georg Zeller, Fernando Izquierdo-Carrasco, Simon A. Berger, Jens Roat Kultima, Luis Pedro Coelho,
Metagenomic species profiling using universal phylogenetic marker genes Shinichi Sunagawa, Daniel R. Mende, Georg Zeller, Fernando Izquierdo-Carrasco, Simon A. Berger, Jens Roat Kultima, Luis Pedro Coelho,
SUPPLEMENTARY FIG. S2. Expression of single HLA loci in shns- and shb 2 m-transduced MKs. Expression of HLA class I antigens (HLA-ABC) as well as
 Supplementary Data Supplementary Methods Flow cytometric analysis of HLA class I single locus expression Expression of single HLA loci (HLA-A and HLA-B) by shns- and shb 2 m-transduced megakaryocytes (MKs)
Supplementary Data Supplementary Methods Flow cytometric analysis of HLA class I single locus expression Expression of single HLA loci (HLA-A and HLA-B) by shns- and shb 2 m-transduced megakaryocytes (MKs)
SUPPLEMENTARY INFORMATION
 SUPPLEMENRY INFORMION doi:.38/nature In vivo nucleosome mapping D4+ Lymphocytes radient-based and I-bead cell sorting D8+ Lymphocytes ranulocytes Lyse the cells Isolate and sequence mononucleosome cores
SUPPLEMENRY INFORMION doi:.38/nature In vivo nucleosome mapping D4+ Lymphocytes radient-based and I-bead cell sorting D8+ Lymphocytes ranulocytes Lyse the cells Isolate and sequence mononucleosome cores
Nature Methods: doi: /nmeth Supplementary Figure 1. Construction of a sensitive TetR mediated auxotrophic off-switch.
 Supplementary Figure 1 Construction of a sensitive TetR mediated auxotrophic off-switch. A Production of the Tet repressor in yeast when conjugated to either the LexA4 or LexA8 promoter DNA binding sequences.
Supplementary Figure 1 Construction of a sensitive TetR mediated auxotrophic off-switch. A Production of the Tet repressor in yeast when conjugated to either the LexA4 or LexA8 promoter DNA binding sequences.
Supplementary Figure 1. Characterization of the POP2 transcriptional and post-transcriptional regulatory elements. (A) POP2 nucleotide sequence
 1 5 6 7 8 9 10 11 1 1 1 Supplementary Figure 1. Characterization of the POP transcriptional and post-transcriptional regulatory elements. (A) POP nucleotide sequence depicting the consensus sequence for
1 5 6 7 8 9 10 11 1 1 1 Supplementary Figure 1. Characterization of the POP transcriptional and post-transcriptional regulatory elements. (A) POP nucleotide sequence depicting the consensus sequence for
Table S1. Oligonucleotide primer sequences
 Table S1. Oligonucleotide primer sequences Primer Name DNA Sequence (5 -> 3 ) Description CD19c CD19d Cre7 Sfpi1lox1 Sfpi1lox2 Sfpi1lox3 5 -AACCAGTCAACACCCTTCC-3 5 -CCAGACTAGATACAGACCAG-3 5 -TCAGCTACACCAGAGACGG-3
Table S1. Oligonucleotide primer sequences Primer Name DNA Sequence (5 -> 3 ) Description CD19c CD19d Cre7 Sfpi1lox1 Sfpi1lox2 Sfpi1lox3 5 -AACCAGTCAACACCCTTCC-3 5 -CCAGACTAGATACAGACCAG-3 5 -TCAGCTACACCAGAGACGG-3
irweb: Data Analysis Guide
 We Find Health in Your Diversity. irepertoire Inc. irweb: Data Analysis Guide Immune Repertoire NGS Data Analysis for irepertoire Reagent Systems For Research Use Only. Not To Used For Clinical Diagnostics.
We Find Health in Your Diversity. irepertoire Inc. irweb: Data Analysis Guide Immune Repertoire NGS Data Analysis for irepertoire Reagent Systems For Research Use Only. Not To Used For Clinical Diagnostics.
Nature Methods: doi: /nmeth Supplementary Figure 1. DMS-MaPseq data are highly reproducible at elevated DMS concentrations.
 Supplementary Figure 1 DMS-MaPseq data are highly reproducible at elevated DMS concentrations. a, Correlation of Gini index for 202 yeast mrna regions with 15x coverage at 2.5% or 5% v/v DMS concentrations
Supplementary Figure 1 DMS-MaPseq data are highly reproducible at elevated DMS concentrations. a, Correlation of Gini index for 202 yeast mrna regions with 15x coverage at 2.5% or 5% v/v DMS concentrations
Nature Methods: doi: /nmeth.4396
 Supplementary Figure 1 Comparison of technical replicate consistency between and across the standard ATAC-seq method, DNase-seq, and Omni-ATAC. (a) Heatmap-based representation of ATAC-seq quality control
Supplementary Figure 1 Comparison of technical replicate consistency between and across the standard ATAC-seq method, DNase-seq, and Omni-ATAC. (a) Heatmap-based representation of ATAC-seq quality control
Determining presence/absence threshold for your dataset
 Determining presence/absence threshold for your dataset In PanCGHweb there are two ways to determine the presence/absence calling threshold. One is based on Receiver Operating Curves (ROC) generated for
Determining presence/absence threshold for your dataset In PanCGHweb there are two ways to determine the presence/absence calling threshold. One is based on Receiver Operating Curves (ROC) generated for
TECH NOTE SMARTer T-cell receptor profiling in single cells
 TECH NOTE SMARTer T-cell receptor profiling in single cells Flexible workflow: Illumina-ready libraries from FACS or manually sorted single cells Ease of use: Optimized indexing allows for pooling 96 cells
TECH NOTE SMARTer T-cell receptor profiling in single cells Flexible workflow: Illumina-ready libraries from FACS or manually sorted single cells Ease of use: Optimized indexing allows for pooling 96 cells
Figure 1. FasterDB SEARCH PAGE corresponding to human WNK1 gene. In the search page, gene searching, in the mouse or human genome, can be done: 1- By
 1 2 3 Figure 1. FasterD SERCH PGE corresponding to human WNK1 gene. In the search page, gene searching, in the mouse or human genome, can be done: 1- y keywords (ENSEML ID, HUGO gene name, synonyms or
1 2 3 Figure 1. FasterD SERCH PGE corresponding to human WNK1 gene. In the search page, gene searching, in the mouse or human genome, can be done: 1- y keywords (ENSEML ID, HUGO gene name, synonyms or
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Fig. 1 shrna mediated knockdown of ZRSR2 in K562 and 293T cells. (a) ZRSR2 transcript levels in stably transduced K562 cells were determined using qrt-pcr. GAPDH was
Supplementary Figure 1 Supplementary Fig. 1 shrna mediated knockdown of ZRSR2 in K562 and 293T cells. (a) ZRSR2 transcript levels in stably transduced K562 cells were determined using qrt-pcr. GAPDH was
Identify sampling methods and recognize biased samples
 9-1 Samples and Surveys Identify sampling methods and recognize biased samples Vocabulary population (p. 462) sample (p. 462) biased sample (p. 463) random sample (p. 462) systematic sample (p. 462) stratified
9-1 Samples and Surveys Identify sampling methods and recognize biased samples Vocabulary population (p. 462) sample (p. 462) biased sample (p. 463) random sample (p. 462) systematic sample (p. 462) stratified
Supplementary Figure 1
 Supplementary Figure 1 Ex2 promotor region Cre IRES cherry pa Ex4 Ex5 Ex1 untranslated Ex3 Ex5 untranslated EYFP pa Rosa26 STOP loxp loxp Cre recombinase EYFP pa Rosa26 loxp 1 kb Interleukin-9 fate reporter
Supplementary Figure 1 Ex2 promotor region Cre IRES cherry pa Ex4 Ex5 Ex1 untranslated Ex3 Ex5 untranslated EYFP pa Rosa26 STOP loxp loxp Cre recombinase EYFP pa Rosa26 loxp 1 kb Interleukin-9 fate reporter
Towards detection of minimal residual disease in multiple myeloma through circulating tumour DNA sequence analysis
 Towards detection of minimal residual disease in multiple myeloma through circulating tumour DNA sequence analysis Trevor Pugh, PhD, FACMG Princess Margaret Cancer Centre, University Health Network Dept.
Towards detection of minimal residual disease in multiple myeloma through circulating tumour DNA sequence analysis Trevor Pugh, PhD, FACMG Princess Margaret Cancer Centre, University Health Network Dept.
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Shah NS, Auld SC, Brust JCM, et al. Transmission of extensively
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Shah NS, Auld SC, Brust JCM, et al. Transmission of extensively
DC enriched CD103. CD11b. CD11c. Spleen. DC enriched. CD11c. DC enriched. CD11c MLN
 CD CD11 + CD + CD11 d g CD CD11 + CD + CD11 CD + CD11 + Events (% of max) Events (% of max) CD + CD11 Events (% of max) Spleen CLN MLN Irf fl/fl e Irf fl/fl h Irf fl/fl DC enriched CD CD11 Spleen DC enriched
CD CD11 + CD + CD11 d g CD CD11 + CD + CD11 CD + CD11 + Events (% of max) Events (% of max) CD + CD11 Events (% of max) Spleen CLN MLN Irf fl/fl e Irf fl/fl h Irf fl/fl DC enriched CD CD11 Spleen DC enriched
Interferon- -producing immature myeloid cells confer protection. against severe invasive group A Streptococcus infections. Supplementary Information
 Supplementary Information Interferon- -producing immature myeloid cells confer protection against severe invasive group A Streptococcus infections Takayuki Matsumura, Manabu Ato, Tadayoshi Ikebe, Makoto
Supplementary Information Interferon- -producing immature myeloid cells confer protection against severe invasive group A Streptococcus infections Takayuki Matsumura, Manabu Ato, Tadayoshi Ikebe, Makoto
Nature Genetics: doi: /ng Supplementary Figure 1. Summary of genetic association data and their traits and gene mappings.
 Supplementary Figure 1 Summary of genetic association data and their traits and gene mappings. Distribution of the (a) number of publications or sources and (b) reported associations for each unique MeSH
Supplementary Figure 1 Summary of genetic association data and their traits and gene mappings. Distribution of the (a) number of publications or sources and (b) reported associations for each unique MeSH
Result Tables The Result Table, which indicates chromosomal positions and annotated gene names, promoter regions and CpG islands, is the best way for
 Result Tables The Result Table, which indicates chromosomal positions and annotated gene names, promoter regions and CpG islands, is the best way for you to discover methylation changes at specific genomic
Result Tables The Result Table, which indicates chromosomal positions and annotated gene names, promoter regions and CpG islands, is the best way for you to discover methylation changes at specific genomic
Erhard et al. (2013). Plant Cell /tpc
 Supplemental Figure 1. c1-hbr allele structure. Diagram of the c1-hbr allele found in stocks segregating 1:1 for rpd1-1 and rpd1-2 homozygous mutants showing the presence of a 363 base pair (bp) Heartbreaker
Supplemental Figure 1. c1-hbr allele structure. Diagram of the c1-hbr allele found in stocks segregating 1:1 for rpd1-1 and rpd1-2 homozygous mutants showing the presence of a 363 base pair (bp) Heartbreaker
Analyzing an individual sequence in the Sequence Editor
 BioNumerics Tutorial: Analyzing an individual sequence in the Sequence Editor 1 Aim The Sequence editor window is a convenient tool implemented in BioNumerics to edit and analyze nucleotide and amino acid
BioNumerics Tutorial: Analyzing an individual sequence in the Sequence Editor 1 Aim The Sequence editor window is a convenient tool implemented in BioNumerics to edit and analyze nucleotide and amino acid
Supplementary Figure 1. Details of the component modelling and steps in generating and validating a PTGS. The steps (boxes) and the flow of
 Supplementary Figure 1. Details of the component modelling and steps in generating and validating a PTGS. The steps (boxes) and the flow of information (arrows) through the modelling process, including
Supplementary Figure 1. Details of the component modelling and steps in generating and validating a PTGS. The steps (boxes) and the flow of information (arrows) through the modelling process, including
Systematic evaluation of spliced alignment programs for RNA- seq data
 Systematic evaluation of spliced alignment programs for RNA- seq data Pär G. Engström, Tamara Steijger, Botond Sipos, Gregory R. Grant, André Kahles, RGASP Consortium, Gunnar Rätsch, Nick Goldman, Tim
Systematic evaluation of spliced alignment programs for RNA- seq data Pär G. Engström, Tamara Steijger, Botond Sipos, Gregory R. Grant, André Kahles, RGASP Consortium, Gunnar Rätsch, Nick Goldman, Tim
How to view Results with Scaffold. Proteomics Shared Resource
 How to view Results with Scaffold Proteomics Shared Resource Starting out Download Scaffold from http://www.proteomes oftware.com/proteom e_software_prod_sca ffold_download.html Follow installation instructions
How to view Results with Scaffold Proteomics Shared Resource Starting out Download Scaffold from http://www.proteomes oftware.com/proteom e_software_prod_sca ffold_download.html Follow installation instructions
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Characterization of Hi-C/CHi-C dynamics and enhancer identification. (a) Scatterplot of Hi-C read counts supporting contacts between domain boundaries. Contacts enclosing domains
Supplementary Figure 1 Characterization of Hi-C/CHi-C dynamics and enhancer identification. (a) Scatterplot of Hi-C read counts supporting contacts between domain boundaries. Contacts enclosing domains
Excel 2011 Charts - Introduction Excel 2011 Series The University of Akron. Table of Contents COURSE OVERVIEW... 2
 Table of Contents COURSE OVERVIEW... 2 DISCUSSION... 2 OBJECTIVES... 2 COURSE TOPICS... 2 LESSON 1: CREATE A CHART QUICK AND EASY... 3 DISCUSSION... 3 CREATE THE CHART... 4 Task A Create the Chart... 4
Table of Contents COURSE OVERVIEW... 2 DISCUSSION... 2 OBJECTIVES... 2 COURSE TOPICS... 2 LESSON 1: CREATE A CHART QUICK AND EASY... 3 DISCUSSION... 3 CREATE THE CHART... 4 Task A Create the Chart... 4
Nature Methods: doi: /nmeth Supplementary Figure 1. Real-time deformability cytometry: contour detection and theoretical modeling.
 Supplementary Figure 1 Real-time deformability cytometry: contour detection and theoretical modeling. (a) Image of cell deformed in constriction; contour (red) according to image analysis algorithm. Scale
Supplementary Figure 1 Real-time deformability cytometry: contour detection and theoretical modeling. (a) Image of cell deformed in constriction; contour (red) according to image analysis algorithm. Scale
Impact of gdna Integrity on the Outcome of DNA Methylation Studies
 Impact of gdna Integrity on the Outcome of DNA Methylation Studies Application Note Nucleic Acid Analysis Authors Emily Putnam, Keith Booher, and Xueguang Sun Zymo Research Corporation, Irvine, CA, USA
Impact of gdna Integrity on the Outcome of DNA Methylation Studies Application Note Nucleic Acid Analysis Authors Emily Putnam, Keith Booher, and Xueguang Sun Zymo Research Corporation, Irvine, CA, USA
Simultaneous profiling of transcriptome and DNA methylome from a single cell
 Additional file 1: Supplementary materials Simultaneous profiling of transcriptome and DNA methylome from a single cell Youjin Hu 1, 2, Kevin Huang 1, 3, Qin An 1, Guizhen Du 1, Ganlu Hu 2, Jinfeng Xue
Additional file 1: Supplementary materials Simultaneous profiling of transcriptome and DNA methylome from a single cell Youjin Hu 1, 2, Kevin Huang 1, 3, Qin An 1, Guizhen Du 1, Ganlu Hu 2, Jinfeng Xue
Supplementary Information Supplementary Figures
 Supplementary Information Supplementary Figures Figure S. The number of reads mapped to the and models for 76 human plasmablasts (AW-AW dataset) using bowtie reconstructed from (A) (B) (C) IMGT_mapped
Supplementary Information Supplementary Figures Figure S. The number of reads mapped to the and models for 76 human plasmablasts (AW-AW dataset) using bowtie reconstructed from (A) (B) (C) IMGT_mapped
i-stop codon positions in the mcherry gene
 Supplementary Figure 1 i-stop codon positions in the mcherry gene The grnas (green) that can potentially generate stop codons from Trp (63 th and 98 th aa, upper panel) and Gln (47 th and 114 th aa, bottom
Supplementary Figure 1 i-stop codon positions in the mcherry gene The grnas (green) that can potentially generate stop codons from Trp (63 th and 98 th aa, upper panel) and Gln (47 th and 114 th aa, bottom
Precise quantification of Ion Torrent libraries on the QuantStudio 3D Digital PCR System
 APPLICATION NOTE QuantStudio 3D Digital PCR System Precise quantification of Ion Torrent libraries on the QuantStudio 3D Digital PCR System Introduction The template preparation step in the next-generation
APPLICATION NOTE QuantStudio 3D Digital PCR System Precise quantification of Ion Torrent libraries on the QuantStudio 3D Digital PCR System Introduction The template preparation step in the next-generation
Accelerating skin wound healing by M-CSF through generating SSEA-1 and -3 stem cells. in the injured sites
 Accelerating skin wound healing by M-CSF through generating SSEA-1 and -3 stem cells in the injured sites Yunyuan Li, Reza Baradar Jalili, Aziz Ghahary Department of Surgery, University of British Columbia,
Accelerating skin wound healing by M-CSF through generating SSEA-1 and -3 stem cells in the injured sites Yunyuan Li, Reza Baradar Jalili, Aziz Ghahary Department of Surgery, University of British Columbia,
dbcamplicons pipeline Amplicons
 dbcamplicons pipeline Amplicons Matthew L. Settles Genome Center Bioinformatics Core University of California, Davis settles@ucdavis.edu; bioinformatics.core@ucdavis.edu Microbial community analysis Goal:
dbcamplicons pipeline Amplicons Matthew L. Settles Genome Center Bioinformatics Core University of California, Davis settles@ucdavis.edu; bioinformatics.core@ucdavis.edu Microbial community analysis Goal:
Supplementary Figure 1: Analysis of monocyte subsets and lineage relationships. (a) Gating strategy for definition of MDP and cmop populations in BM
 Supplementary Figure 1: Analysis of monocyte subsets and lineage relationships. (a) Gating strategy for definition of MDP and cmop populations in BM of Cx3cr1 GFP/+ mice related to Fig. 1a. MDP was defined
Supplementary Figure 1: Analysis of monocyte subsets and lineage relationships. (a) Gating strategy for definition of MDP and cmop populations in BM of Cx3cr1 GFP/+ mice related to Fig. 1a. MDP was defined
dbcamplicons pipeline Amplicons
 dbcamplicons pipeline Amplicons Matthew L. Settles Genome Center Bioinformatics Core University of California, Davis settles@ucdavis.edu; bioinformatics.core@ucdavis.edu Microbial community analysis Goal:
dbcamplicons pipeline Amplicons Matthew L. Settles Genome Center Bioinformatics Core University of California, Davis settles@ucdavis.edu; bioinformatics.core@ucdavis.edu Microbial community analysis Goal:
Supplementary information - High-throughput sequencing enhanced phage. display enables the identification of patient-specific epitope motifs in serum
 Supplementary information - High-throughput sequencing enhanced phage display enables the identification of patient-specific epitope motifs in serum Anders Christiansen 1, Jens V. Kringelum 2, Christian
Supplementary information - High-throughput sequencing enhanced phage display enables the identification of patient-specific epitope motifs in serum Anders Christiansen 1, Jens V. Kringelum 2, Christian
Nature Structural & Molecular Biology: doi: /nsmb.2996
 Supplementary Figure 1 Negative-stain EM of native cytoplasmic dynein. (a) A representative micrograph of negatively stained dynein, with the flexible dimeric molecules circled in white. (b) Reference-free
Supplementary Figure 1 Negative-stain EM of native cytoplasmic dynein. (a) A representative micrograph of negatively stained dynein, with the flexible dimeric molecules circled in white. (b) Reference-free
Distribution of human ILCs during chronic lung disease.
 Supplementary Figure 1 Distribution of human ILCs during chronic lung disease. Quantification of flow cytometric analysis of lung tissue from patients with COPD or IPF identifying (a) frequencies of total
Supplementary Figure 1 Distribution of human ILCs during chronic lung disease. Quantification of flow cytometric analysis of lung tissue from patients with COPD or IPF identifying (a) frequencies of total
RNA Sample Quality Assessment Test for the WT-Ovation FFPE System
 RNA Sample Quality Assessment Test for the WT-Ovation FFPE System TECHNICAL REPORT Introduction Gene expression analysis of clinical samples is challenging due to sample availability, amount, and integrity.
RNA Sample Quality Assessment Test for the WT-Ovation FFPE System TECHNICAL REPORT Introduction Gene expression analysis of clinical samples is challenging due to sample availability, amount, and integrity.
(A-B) P2ry14 expression was assessed by (A) genotyping (upper arrow: WT; lower
 Supplementary Figures S1. (A-B) P2ry14 expression was assessed by (A) genotyping (upper arrow: ; lower arrow: KO) and (B) q-pcr analysis with Lin- cells, The white vertical line in panel A indicates that
Supplementary Figures S1. (A-B) P2ry14 expression was assessed by (A) genotyping (upper arrow: ; lower arrow: KO) and (B) q-pcr analysis with Lin- cells, The white vertical line in panel A indicates that
Long-term dynamics of CA1 hippocampal place codes
 Long-term dynamics of CA1 hippocampal place codes Yaniv Ziv, Laurie D. Burns, Eric D. Cocker, Elizabeth O. Hamel, Kunal K. Ghosh, Lacey J. Kitch, Abbas El Gamal, and Mark J. Schnitzer Supplementary Fig.
Long-term dynamics of CA1 hippocampal place codes Yaniv Ziv, Laurie D. Burns, Eric D. Cocker, Elizabeth O. Hamel, Kunal K. Ghosh, Lacey J. Kitch, Abbas El Gamal, and Mark J. Schnitzer Supplementary Fig.
Supplementary Materials for
 www.sciencetranslationalmedicine.org/cgi/content/full/1/12/12ra23/dc1 Supplementary Materials for Measurement and Clinical Monitoring of Human Lymphocyte Clonality by Massively Parallel VDJ Pyrosequencing
www.sciencetranslationalmedicine.org/cgi/content/full/1/12/12ra23/dc1 Supplementary Materials for Measurement and Clinical Monitoring of Human Lymphocyte Clonality by Massively Parallel VDJ Pyrosequencing
Supplementary Figure 1 Strategy for parallel detection of DHSs and adjacent nucleosomes
 Supplementary Figure 1 Strategy for parallel detection of DHSs and adjacent nucleosomes DNase I cleavage DNase I DNase I digestion Sucrose gradient enrichment Small Large F1 F2...... F9 F1 F1 F2 F3 F4
Supplementary Figure 1 Strategy for parallel detection of DHSs and adjacent nucleosomes DNase I cleavage DNase I DNase I digestion Sucrose gradient enrichment Small Large F1 F2...... F9 F1 F1 F2 F3 F4
