Functional analysis using EBI Metagenomics
|
|
|
- Michael Blair
- 6 years ago
- Views:
Transcription
1 Functional analysis using EBI Metagenomics Contents Tutorial information... 2 Tutorial learning objectives... 2 An introduction to functional analysis using EMG... 3 What are protein signatures?... 3 Assigning functional information to metagenomic sequences... 4 Finding functional information about samples on the EMG website... 5 Exercises... 9 Part 1 Browsing analysed data via the EMG website... 9 Part 2 - Analysing EMG data using STAMP Alex Mitchell (mitchell@ebi.ac.uk) V1.4, March
2 Tutorial information Course description This tutorial provides an introduction to functional analysis of metagenomic data, using the EBI Metagenomics (EMG) resource. You will learn about the underlying analysis software and how it is used to infer information about the function of predicted coding sequences, and how to find and download data for use with 3 rd party analysis software. Course level Suitable for graduate-level scientists and above Pre-requisites Basic knowledge of biology and basic Unix skills Subject area Genes, Genomes and Metagenomes; Sequence Analysis Target audience Scientists interested in metagenome analysis Resources required Internet access (a current browser such as the latest version of Firefox, Chrome or Safari) and the STAMP analysis package. Approximate time needed 60 minutes Tutorial learning objectives After completing this course, you should: understand how EMG provides functional analysis of metagenomic data sets know where to find and how to interpret analysis results for samples on the EMG website know how to download the raw sample data and analysis results for use with 3 rd party visualisation and statistical analysis packages 2
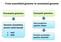
3 An introduction to functional analysis using EMG The EBI Metagenomics resource (EMG) provides functional analysis of predicted coding sequences (pcds) from metagenomic data sets using the InterPro database. InterPro is a sequence analysis resource that predicts protein family membership, along with the presence of important domains and sites. It does this by combining predictive models known as protein signatures from a number of different databases into a single searchable resource. InterPro curators manually integrate the different signatures, providing names and descriptive abstracts and, whenever possible, adding Gene Ontology (GO) terms. Links are also provided to pathway databases, such as KEGG, MetaCyc and Reactome, and to structural resources, such as SCOP, CATH and PDB. What are protein signatures? Protein signatures are obtained by modelling the conservation of amino acids at specific positions within a group of related proteins (i.e., a protein family), or within the domains/sites shared by a group of proteins. InterPro s different member databases use different computational methods to produce protein signatures, and they each have their own particular focus of interest: structural and/or functional domains, protein families, or protein features, such as active sites or binding sites (see Figure 1). 3
4 Figure 1. InterPro member databases grouped by the methods (indicated in white coloured text) used to construct their signatures. Their focus of interest is shown in blue text. Only a subset of the InterPro member databases are used by EMG: Gene3D, TIGRFAMs, Pfam, PRINTS and PROSITE patterns. These databases were selected since, together, they provide both high coverage and offer detailed functional analysis, and have underlying algorithms that can cope with the vast amounts of fragmentary sequence data found in metagenomic datasets. Assigning functional information to metagenomic sequences Whilst InterPro matches to metagenomic sequence sets are informative in their own right, EMG offers an additional type of analysis in the form of Gene Ontology (GO) terms. The Gene Ontology is made up of 3 structured controlled vocabularies that describe gene products in terms of their associated biological processes, cellular components and molecular functions in a species-independent manner. By using GO terms, scientists working on different species or using different databases can compare datasets, since they have a precisely defined name and meaning 4
5 for a particular concept. Terms in the Gene Ontology are ordered into hierarchies, with less specific terms towards the top and more specific terms towards the bottom (see Figure 2). Figure 2. An example of GO terms organised into a hierarchy, with terms becoming less specific as the hierarchy is ascended (e.g., alpha-tubulin binding is a type of cytoskeletal binding, which is a type of protein binding). Note that a GO term can have more than one parent term. The Gene Ontology also allows for different types of relationships between terms (such as has part of or regulates ). The EMG analysis pipeline only uses the straightforward is a relationships. More information about the GO project can be found at As part of the EMG analysis pipeline, GO terms for molecular function, biological process and cellular component are associated to pcds in a sample by via the InterPro2GO mapping service. This works as follows: InterPro entries are given GO terms by curators if the terms can be accurately applied to all of the proteins matching that entry. Sequences searched against InterPro are then associated with GO terms by virtue of the entries they match - a protein that matches one InterPro entry with the GO term kinase activity and another InterPro entry with the GO term zinc ion binding will be annotated with both GO terms. Finding functional information about samples on the EMG website Functional analysis of samples within projects on the EMG website ( can be accessed by clicking on the Functional Analysis tab found toward the top of any sample page (see Figure 3 below). 5
6 Figure 3. A Functional analysis tab can be found towards the top of each sample page. Clicking on this tab brings up a page displaying sequence features (the number of reads with pcds, the number of pcds with InterPro matches, etc), InterPro match information and GO term annotation for the sample, as shown in Figure 4 below. 6
7 Figure 4. Functional analysis of metagenomics data, as shown on the EMG website. A) InterPro match information for the predicted coding sequences in the sample is shown. The number of InterPro matches are displayed graphically, and as a table that has a text search facility. B) The GO terms predicted for the sample are displayed. Different graphical representations are available, and can be selected by clicking on the Switch view icons. The Gene Ontology terms displayed graphically on the web site have been slimmed with a special GO slim developed for metagenomic data sets. GO slims are cut-down versions of the Gene Ontology, containing a subset of the terms in the whole GO. They give a broad overview of the ontology content without the detail of the specific fine-grained terms. The full data sets used to generate both the InterPro and GO overview charts, along with a host of additional data (processed reads, pcds, reads encoding 16S rrnas, taxonomic analyses, etc) can be downloaded for further analysis by clicking the Download tab, found towards the top of the page (see Figure 5). 7
8 Figure 5. Each sample has a download tab, where the full set of sequences, analyses, summaries and raw data can be downloaded. 8
9 Exercises Part 1 Browsing analysed data via the EMG website Open the Metagenomics Portal homepage in a web browser ( From the Projects list (click the Projects tab, or View all projects, find and click on HOT station, Central North Pacific Gyre, ALOH. You should now have a Project overview page describing the project, related publications, and links to the samples that the project contains. Question 1: What publications are associated with this study? Scroll down to Associated Samples and open the HOT Station ALOHA, 25 m sample. You should now have a Sample Overview page, describing various meta-data associated with the sample, such as the geographic location from which it was isolated, its collection date, and so on. Question 2: What is the latitude, longitude and depth at which the sample was collected? Question 3: What geographic location does this correspond to? Question 4: What environmental ontology (ENVO) identifer and name has the sample material been annotated with? The page has a number of tabs towards the top (Overview, Quality control, Taxonomy analysis, Functional analysis). Click on the Download tab. Click the file labelled Predicted CDS (FASTA) link, and save this file to your desktop. Find the file, and either double click on it to open it, or examine it using less by typing the following commands in a terminal window: 9
10 cd ~/Desktop less HOT_Station_ALOHA,_25m_depth_pCDS.faa Have a look at one or two of the many sequences it contains. We will look at the analysis results for this entire batch of sequences, displayed on the EMG website, in a moment. First, we will attempt analyse just one of the predicted coding sequences using InterPro (the analysis results on the EMG website summarise these kind of results for tens or hundreds of thousands of sequences). In a new tab or window, open your web browser and navigate to Copy and paste the following sequence into the text box on the InterPro home page where it says Analyse your sequence : >SRR _1_115_+ ENNQEIKIIRNYINEFNLTGFIVGIPLDEKGQMTNQAI Press Search and wait for your results. Your sequence will be run through the InterProScan analysis software, which attempts to match it against all of the different signatures in the InterProScan database. contain? Question 5: What feature does InterProScan predict your sequence to Clicking on the InterPro entry name or IPR accession number will take you to the InterPro entry page for your result, where more information can be found. Question 6: What GO terms is the feature associated with? Return to the sample page for HOT station, Central North Pacific Gyre, ALOH 25 m. Now we are going to look at the functional analysis results for all of the pcds in the sample. First, we will find the number of sequences that made it through to the functional analysis section of the pipeline. 10
11 Click on the Quality control tab. This page displays a series of charts, showing how many sequences passed each quality control step, how many reads were left after clustering, and so on. Question 7: After all of the quality filtering steps are complete, how many reads were submitted for analysis by the pipeline? Next, we will look at the results of the functional predictions for the pcds. These can be found under the Functional analysis tab. Click on the Functional analysis tab and examine the InterPro match section. The top part of this page shows a sequence feature summary, showing the number of reads with predicted coding sequences (pcds), the number of pcds with InterPro matches, etc. Question 8: How many of the reads that passed the quality control stage had predicted coding sequences (pcds)? Question 9: How many pcds have InterProScan hits? Scroll down the page to the InterPro match summary section. pcds? Question 10: How many different InterPro entries were matched by the Question 11: Why is this figure different to the number of pcds that have InterProScan hits? Next we will examine the GO terms predicted by InterPro for the pcds in the sample. Scroll down to the GO term annotation section of the page and examine the 3 bar charts, which correspond to the 3 different components of the Gene Ontology. Question 12: What are the top 3 biological process terms predicted for the pcds from the sample? 11
12 Selecting the pie chart representation of GO terms makes it easier to visualise the data to find the answer. Now we will look at the taxonomic analysis for this sample. Click on the Taxonomy Analysis tab and examine the phylum composition graph and table. Question 13: How many different OTUs were found in this sample? Question 14: What are the top 3 phyla in the sample? Select the Krona chart view of the data using the icon. This brings up an interactive chart that can be used to analyse data at different taxonomic ranks. Question 15: What proportion of cyanobacteria in this sample are made up of Synechococcaceae? Now we will compare these analyses with those for a sample taken at 500 m from the same geographical location. In a new tab or window, find and open the HOT station, Central North Pacific Gyre, ALOH project page again. Select the sample HOT Station ALOHA, 500m and examine the information under the Functional analysis tabs. Question 16: How many pcds were in this sample? Question 17: How many of the pcdss have an InterPro match? sample? Question 18: How many different InterPro entries are matched by this sample? Question 19: Are these figures broadly comparable to the ones for the 25 m 12
13 Scroll to the bottom of the page and examine the GO term annotation for the day 500 m sample. Question 20: Are there visible differences between the GO terms for this sample and the 25 m sample? Could there be any biological explanation for this? Selecting the bar chart representation of GO terms makes it easier to compare different data sets. Return to the HOT station project page and open the taxonomy analysis results for all 4 of the samples, each in a new window. Question 21: How does the taxonomic composition change at different ocean sampling depths? Are any trends in the data consistent with your answer to question 18? Part 2 - Analysing EMG data using STAMP Whilst EMG does not currently support direct comparison of multiple samples, it is possible to download the underlying data for use with other visualisation and/or statistical analysis tools, such as STAMP (Statistical Analysis of Metagenomic Profiles: The Downloads tab for a given sample lists the files with the necessary data, although a slight amount of data wrangling is usually required to reformat files, etc. We are going perform a detailed comparison of the GO slimmed biological process results for the samples at 25 m and 500 m using STAMP. To do this, open the HOT station, Central North Pacific Gyre, ALOH project page again. Select the sample HOT Station ALOHA, 25 m study and download the GO slim annotation (CSV) file from the download tab. Repeat this for the HOT Station ALOHA, 500 m study. You should now have the following 2 files in your downloads folder: HOT_Station_ALOHA,_25m_depth_GO_slim.csv HOT_Station_ALOHA,_500m_depth,_MG_GO_slim.csv (don t worry about the file name inconsistency that is something we need to fix in the database!) 13
14 Drag or copy these files to your desktop, and then launch a terminal window, so that you can manipulate the files on the command line. Navigate to the Desktop directory by typing cd ~/Desktop/ Next, type ls to make sure you can see the files, and take a look at their contents using the less command. First we are going to wrangle data from the 25 m sample, using a mixture of grep to pull out the lines containing the phrase biological_process and awk to split up and reorder the fields. awk is a text-processing language that is useful for extracting or transforming documents and reformatting text. You can find out more about it here: Enter the following command (all on one line): grep 'biological_process' HOT_Station_ALOHA,_25m_depth_GO_slim.csv awk -F\" '{print $6 "\t" $4 "\t" $8 }' > biological_process_25m.txt This should produce a new file called biological_process_25m.txt. Use less to examine this file. Question 22: What are the differences between this file and the HOT_Station_ALOHA,_25m_depth_GO_slim.csv file? Next, we are going to add the biological process data from the 500 m sample to this file. Once again, will use grep to pull out the relevant lines in the source file, awk to process the lines and output the correct fields, and paste to merge the data with the contents of the file we produced above. Enter the following command (all on one line): grep 'biological_process' HOT_Station_ALOHA,_500m_depth,_MG_GO_slim.csv awk -F\" '{print $8 }' paste biological_process_25m.txt - > biological_process_25mvs500m.txt This should create another new file, this time called biological_process_25mvs500m.txt. Use less to examine this file. 14
15 Question 23: What is the difference between this file and the biological_process_25m.txt file? Finally, we need to add a simple header to the file, so that STAMP can process it properly. To do this, we could open the file and add the header manually, but here we will do it on the command line using the printf command to write the header, cat to print the contents of the file, and directing the output to a new file with the > operator. Enter the following command (all on one line): printf 'Classification\tSubclassifcation\t25m\t500m\n' cat - biological_process_25mvs500m.txt > biological_process_25mvs500m.spf This command should have produced a third file called biological_process_25mvs500m.spf that STAMP can read. Launch STAMP by typing STAMP into the terminal window. Load the biological_process_25mvs500m.spf file into STAMP (you will need to set Profile level to Subclassification and select Two samples from the tabs on the left hand side of the page to view the file once it is loaded). Try experimenting with different plot types and statistical tests. Question 24: Are there any observable differences in biological process GO terms between the 2 samples? For your convenience, three files containing the HOT Station GO term results for all 3 branches of the Gene Ontology (molecular function, biological process and cellular component) at each sample depth have been pre-generated and formatted for use with STAMP. These are located in the STAMP_data folder on the desktop (filenames: HOT_Station_ALOHA_bp.spf, HOT_Station_ALOHA_mf.spf and HOT_Station_ALOHA_cc.spf). Try loading these files into STAMP, as you did with the previous file, and exploring the results. You can select different samples to compare (25, 75, 125 and 500 m) using the sample 1 and sample 2 drop down menus. 15
16 Question 25: What trends can you identify in the molecular function, biological process and cellular component GO terms across the different sampling depths? 16
I AM NOT A METAGENOMIC EXPERT. I am merely the MESSENGER. Blaise T.F. Alako, PhD EBI Ambassador
 I AM NOT A METAGENOMIC EXPERT I am merely the MESSENGER Blaise T.F. Alako, PhD EBI Ambassador blaise@ebi.ac.uk Hubert Denise Alex Mitchell Peter Sterk Sarah Hunter http://www.ebi.ac.uk/metagenomics Blaise
I AM NOT A METAGENOMIC EXPERT I am merely the MESSENGER Blaise T.F. Alako, PhD EBI Ambassador blaise@ebi.ac.uk Hubert Denise Alex Mitchell Peter Sterk Sarah Hunter http://www.ebi.ac.uk/metagenomics Blaise
BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers
 BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers Web resources: NCBI database: http://www.ncbi.nlm.nih.gov/ Ensembl database: http://useast.ensembl.org/index.html UCSC
BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers Web resources: NCBI database: http://www.ncbi.nlm.nih.gov/ Ensembl database: http://useast.ensembl.org/index.html UCSC
Week 1 BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers
 Week 1 BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers Web resources: NCBI database: http://www.ncbi.nlm.nih.gov/ Ensembl database: http://useast.ensembl.org/index.html
Week 1 BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers Web resources: NCBI database: http://www.ncbi.nlm.nih.gov/ Ensembl database: http://useast.ensembl.org/index.html
Gene List Enrichment Analysis
 Outline Gene List Enrichment Analysis George Bell, Ph.D. BaRC Hot Topics March 16, 2010 Why do enrichment analysis? Main types Selecting or ranking genes Annotation sources Statistics Remaining issues
Outline Gene List Enrichment Analysis George Bell, Ph.D. BaRC Hot Topics March 16, 2010 Why do enrichment analysis? Main types Selecting or ranking genes Annotation sources Statistics Remaining issues
How to view Results with Scaffold. Proteomics Shared Resource
 How to view Results with Scaffold Proteomics Shared Resource Starting out Download Scaffold from http://www.proteomes oftware.com/proteom e_software_prod_sca ffold_download.html Follow installation instructions
How to view Results with Scaffold Proteomics Shared Resource Starting out Download Scaffold from http://www.proteomes oftware.com/proteom e_software_prod_sca ffold_download.html Follow installation instructions
Browser Exercises - I. Alignments and Comparative genomics
 Browser Exercises - I Alignments and Comparative genomics 1. Navigating to the Genome Browser (GBrowse) Note: For this exercise use http://www.tritrypdb.org a. Navigate to the Genome Browser (GBrowse)
Browser Exercises - I Alignments and Comparative genomics 1. Navigating to the Genome Browser (GBrowse) Note: For this exercise use http://www.tritrypdb.org a. Navigate to the Genome Browser (GBrowse)
Microarray Data Analysis in GeneSpring GX 11. Month ##, 200X
 Microarray Data Analysis in GeneSpring GX 11 Month ##, 200X Agenda Genome Browser GO GSEA Pathway Analysis Network building Find significant pathways Extract relations via NLP Data Visualization Options
Microarray Data Analysis in GeneSpring GX 11 Month ##, 200X Agenda Genome Browser GO GSEA Pathway Analysis Network building Find significant pathways Extract relations via NLP Data Visualization Options
From Variants to Pathways: Agilent GeneSpring GX s Variant Analysis Workflow
 From Variants to Pathways: Agilent GeneSpring GX s Variant Analysis Workflow Technical Overview Import VCF Introduction Next-generation sequencing (NGS) studies have created unanticipated challenges with
From Variants to Pathways: Agilent GeneSpring GX s Variant Analysis Workflow Technical Overview Import VCF Introduction Next-generation sequencing (NGS) studies have created unanticipated challenges with
KnetMiner USER TUTORIAL
 KnetMiner USER TUTORIAL Keywan Hassani-Pak ROTHAMSTED RESEARCH 10 NOVEMBER 2017 About KnetMiner KnetMiner, with a silent "K" and standing for Knowledge Network Miner, is a suite of open-source software
KnetMiner USER TUTORIAL Keywan Hassani-Pak ROTHAMSTED RESEARCH 10 NOVEMBER 2017 About KnetMiner KnetMiner, with a silent "K" and standing for Knowledge Network Miner, is a suite of open-source software
Knowledge-Guided Analysis with KnowEnG Lab
 Han Sinha Song Weinshilboum Knowledge-Guided Analysis with KnowEnG Lab KnowEnG Center Powerpoint by Charles Blatti Knowledge-Guided Analysis KnowEnG Center 2017 1 Exercise In this exercise we will be doing
Han Sinha Song Weinshilboum Knowledge-Guided Analysis with KnowEnG Lab KnowEnG Center Powerpoint by Charles Blatti Knowledge-Guided Analysis KnowEnG Center 2017 1 Exercise In this exercise we will be doing
Tutorial. Whole Metagenome Functional Analysis (beta) Sample to Insight. November 21, 2017
 Whole Metagenome Functional Analysis (beta) November 21, 2017 Sample to Insight QIAGEN Aarhus Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 www.qiagenbioinformatics.com AdvancedGenomicsSupport@qiagen.com
Whole Metagenome Functional Analysis (beta) November 21, 2017 Sample to Insight QIAGEN Aarhus Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 www.qiagenbioinformatics.com AdvancedGenomicsSupport@qiagen.com
A tutorial introduction into the MIPS PlantsDB barley&wheat database instances
 transplant 2 nd user training workshop Poznan, Poland, June, 27 th, 2013 A tutorial introduction into the MIPS PlantsDB barley&wheat database instances TUTORIAL ANSWERS Please direct any questions related
transplant 2 nd user training workshop Poznan, Poland, June, 27 th, 2013 A tutorial introduction into the MIPS PlantsDB barley&wheat database instances TUTORIAL ANSWERS Please direct any questions related
NCBI web resources I: databases and Entrez
 NCBI web resources I: databases and Entrez Yanbin Yin Most materials are downloaded from ftp://ftp.ncbi.nih.gov/pub/education/ 1 Homework assignment 1 Two parts: Extract the gene IDs reported in table
NCBI web resources I: databases and Entrez Yanbin Yin Most materials are downloaded from ftp://ftp.ncbi.nih.gov/pub/education/ 1 Homework assignment 1 Two parts: Extract the gene IDs reported in table
A WEB-BASED TOOL FOR GENOMIC FUNCTIONAL ANNOTATION, STATISTICAL ANALYSIS AND DATA MINING
 A WEB-BASED TOOL FOR GENOMIC FUNCTIONAL ANNOTATION, STATISTICAL ANALYSIS AND DATA MINING D. Martucci a, F. Pinciroli a,b, M. Masseroli a a Dipartimento di Bioingegneria, Politecnico di Milano, Milano,
A WEB-BASED TOOL FOR GENOMIC FUNCTIONAL ANNOTATION, STATISTICAL ANALYSIS AND DATA MINING D. Martucci a, F. Pinciroli a,b, M. Masseroli a a Dipartimento di Bioingegneria, Politecnico di Milano, Milano,
Bioinformatics & Protein Structural Analysis. Bioinformatics & Protein Structural Analysis. Learning Objective. Proteomics
 The molecular structures of proteins are complex and can be defined at various levels. These structures can also be predicted from their amino-acid sequences. Protein structure prediction is one of the
The molecular structures of proteins are complex and can be defined at various levels. These structures can also be predicted from their amino-acid sequences. Protein structure prediction is one of the
Introduction to DNA-Sequencing
 informatics.sydney.edu.au sih.info@sydney.edu.au The Sydney Informatics Hub provides support, training, and advice on research data, analyses and computing. Talk to us about your computing infrastructure,
informatics.sydney.edu.au sih.info@sydney.edu.au The Sydney Informatics Hub provides support, training, and advice on research data, analyses and computing. Talk to us about your computing infrastructure,
1 BASIC CHARTING. 1.1 Introduction
 1 BASIC CHARTING 1.1 Introduction This section covers the basic principles of how to create and modify a chart in Excel. With Excel 2016, the charting process is user-friendly and offers many ways to amplify
1 BASIC CHARTING 1.1 Introduction This section covers the basic principles of how to create and modify a chart in Excel. With Excel 2016, the charting process is user-friendly and offers many ways to amplify
 Genome Annotation Genome annotation What is the function of each part of the genome? Where are the genes? What is the mrna sequence (transcription, splicing) What is the protein sequence? What does
Genome Annotation Genome annotation What is the function of each part of the genome? Where are the genes? What is the mrna sequence (transcription, splicing) What is the protein sequence? What does
Getting Started with Workforce Detail Reporting
 The Cal Answers Workforce Detail Dashboards enable you to access detailed information about the Positions and Employees appointed into Jobs in your organization. The table below provides descriptions and
The Cal Answers Workforce Detail Dashboards enable you to access detailed information about the Positions and Employees appointed into Jobs in your organization. The table below provides descriptions and
Kyoto Encyclopedia of Genes and Genomes (KEGG)
 NPTEL Biotechnology -Systems Biology Kyoto Encyclopedia of Genes and Genomes (KEGG) Dr. M. Vijayalakshmi School of Chemical and Biotechnology SASTRA University Joint Initiative of IITs and IISc Funded
NPTEL Biotechnology -Systems Biology Kyoto Encyclopedia of Genes and Genomes (KEGG) Dr. M. Vijayalakshmi School of Chemical and Biotechnology SASTRA University Joint Initiative of IITs and IISc Funded
David Crossman, Ph.D. UAB Heflin Center for Genomic Science. Immersion Course
 David Crossman, Ph.D. UAB Heflin Center for Genomic Science Immersion Course What to do with your list of genes Apply a Systems Biology approach to data mine and analyze your data Tools and databases available
David Crossman, Ph.D. UAB Heflin Center for Genomic Science Immersion Course What to do with your list of genes Apply a Systems Biology approach to data mine and analyze your data Tools and databases available
The Gene Ontology Annotation (GOA) project application of GO in SWISS-PROT, TrEMBL and InterPro
 Comparative and Functional Genomics Comp Funct Genom 2003; 4: 71 74. Published online in Wiley InterScience (www.interscience.wiley.com). DOI: 10.1002/cfg.235 Conference Review The Gene Ontology Annotation
Comparative and Functional Genomics Comp Funct Genom 2003; 4: 71 74. Published online in Wiley InterScience (www.interscience.wiley.com). DOI: 10.1002/cfg.235 Conference Review The Gene Ontology Annotation
Analysis of Microarray Data
 Analysis of Microarray Data Lecture 3: Visualization and Functional Analysis George Bell, Ph.D. Senior Bioinformatics Scientist Bioinformatics and Research Computing Whitehead Institute Outline Review
Analysis of Microarray Data Lecture 3: Visualization and Functional Analysis George Bell, Ph.D. Senior Bioinformatics Scientist Bioinformatics and Research Computing Whitehead Institute Outline Review
Introduction to Cognos Analytics and Report Navigation Training. IBM Cognos Analytics 11
 Introduction to Cognos Analytics and Report Navigation Training IBM Cognos Analytics 11 Applicable for former IBM Cognos 10 report users who access CBMS Cognos to run and view reports March 2018 This training
Introduction to Cognos Analytics and Report Navigation Training IBM Cognos Analytics 11 Applicable for former IBM Cognos 10 report users who access CBMS Cognos to run and view reports March 2018 This training
Analysis of Microarray Data
 Analysis of Microarray Data Lecture 3: Visualization and Functional Analysis George Bell, Ph.D. Bioinformatics Scientist Bioinformatics and Research Computing Whitehead Institute Outline Review Visualizing
Analysis of Microarray Data Lecture 3: Visualization and Functional Analysis George Bell, Ph.D. Bioinformatics Scientist Bioinformatics and Research Computing Whitehead Institute Outline Review Visualizing
static MM_Index snap(mm_index corect, MM_Index ligct, int imatch0, int *moleatoms, i
 BIOLUMINATE static MM_Index snap(mm_index corect, MM_Index ligct, int imatch0, int *moleatoms, int *refcoreatoms){int ncoreat = :vector mappings; PhpCoreMapping mapping; for COMMON(glidelig).
BIOLUMINATE static MM_Index snap(mm_index corect, MM_Index ligct, int imatch0, int *moleatoms, int *refcoreatoms){int ncoreat = :vector mappings; PhpCoreMapping mapping; for COMMON(glidelig).
Pivot Table Tutorial Using Ontario s Public Sector Salary Disclosure Data
 Pivot Table Tutorial Using Ontario s Public Sector Salary Disclosure Data Now that have become more familiar with downloading data in Excel format (xlsx) or a text or csv format (txt, csv), it s time to
Pivot Table Tutorial Using Ontario s Public Sector Salary Disclosure Data Now that have become more familiar with downloading data in Excel format (xlsx) or a text or csv format (txt, csv), it s time to
EMPLOYEE JOB AID GETTING STARTED IN WORKDAY KEY ICONS ICON FUNCTION DESCRIPTION
 EMPLOYEE JOB AID GETTING STARTED IN WORKDAY This job aid covers the basics to get started using Workday and the features that will help you use the system. Workday s proven processes simplify the way we
EMPLOYEE JOB AID GETTING STARTED IN WORKDAY This job aid covers the basics to get started using Workday and the features that will help you use the system. Workday s proven processes simplify the way we
How to view Results with. Proteomics Shared Resource
 How to view Results with Scaffold 3.0 Proteomics Shared Resource An overview This document is intended to walk you through Scaffold version 3.0. This is an introductory guide that goes over the basics
How to view Results with Scaffold 3.0 Proteomics Shared Resource An overview This document is intended to walk you through Scaffold version 3.0. This is an introductory guide that goes over the basics
Section 1 : About the Resource Database
 Section 1 : About the Resource Database The Resource Database brings together community-identified resources in a shared, searchable database which is integrated with the Client Registry. Since this single
Section 1 : About the Resource Database The Resource Database brings together community-identified resources in a shared, searchable database which is integrated with the Client Registry. Since this single
COMPUTER RESOURCES II:
 COMPUTER RESOURCES II: Using the computer to analyze data, using the internet, and accessing online databases Bio 210, Fall 2006 Linda S. Huang, Ph.D. University of Massachusetts Boston In the first computer
COMPUTER RESOURCES II: Using the computer to analyze data, using the internet, and accessing online databases Bio 210, Fall 2006 Linda S. Huang, Ph.D. University of Massachusetts Boston In the first computer
This practical aims to walk you through the process of text searching DNA and protein databases for sequence entries.
 PRACTICAL 1: BLAST and Sequence Alignment The EBI and NCBI websites, two of the most widely used life science web portals are introduced along with some of the principal databases: the NCBI Protein database,
PRACTICAL 1: BLAST and Sequence Alignment The EBI and NCBI websites, two of the most widely used life science web portals are introduced along with some of the principal databases: the NCBI Protein database,
SeattleSNPs Interactive Tutorial: Database Inteface Entrez, dbsnp, HapMap, Perlegen
 SeattleSNPs Interactive Tutorial: Database Inteface Entrez, dbsnp, HapMap, Perlegen The tutorial is designed to take you through the steps necessary to access SNP data from the primary database resources:
SeattleSNPs Interactive Tutorial: Database Inteface Entrez, dbsnp, HapMap, Perlegen The tutorial is designed to take you through the steps necessary to access SNP data from the primary database resources:
Genome Annotation - 2. Qi Sun Bioinformatics Facility Cornell University
 Genome Annotation - 2 Qi Sun Bioinformatics Facility Cornell University Output from Maker GFF file: Annotated gene, transcripts, and CDS FASTA file: Predicted transcript sequences Predicted protein sequences
Genome Annotation - 2 Qi Sun Bioinformatics Facility Cornell University Output from Maker GFF file: Annotated gene, transcripts, and CDS FASTA file: Predicted transcript sequences Predicted protein sequences
Protein Bioinformatics Part I: Access to information
 Protein Bioinformatics Part I: Access to information 260.655 April 6, 2006 Jonathan Pevsner, Ph.D. pevsner@kennedykrieger.org Outline [1] Proteins at NCBI RefSeq accession numbers Cn3D to visualize structures
Protein Bioinformatics Part I: Access to information 260.655 April 6, 2006 Jonathan Pevsner, Ph.D. pevsner@kennedykrieger.org Outline [1] Proteins at NCBI RefSeq accession numbers Cn3D to visualize structures
FACULTY OF BIOCHEMISTRY AND MOLECULAR MEDICINE
 FACULTY OF BIOCHEMISTRY AND MOLECULAR MEDICINE BIOMOLECULES COURSE: COMPUTER PRACTICAL 1 Author of the exercise: Prof. Lloyd Ruddock Edited by Dr. Leila Tajedin 2017-2018 Assistant: Leila Tajedin (leila.tajedin@oulu.fi)
FACULTY OF BIOCHEMISTRY AND MOLECULAR MEDICINE BIOMOLECULES COURSE: COMPUTER PRACTICAL 1 Author of the exercise: Prof. Lloyd Ruddock Edited by Dr. Leila Tajedin 2017-2018 Assistant: Leila Tajedin (leila.tajedin@oulu.fi)
BGGN 213: Foundations of Bioinformatics (Fall 2017)
 BGGN 213: Foundations of Bioinformatics (Fall 2017) Course Instructor: Dr. Barry J. Grant ( bjgrant@ucsd.edu ) Course Website: https://bioboot.github.io/bggn213_f17/ DRAFT: 2017-08-10 (15:02:30 PDT on
BGGN 213: Foundations of Bioinformatics (Fall 2017) Course Instructor: Dr. Barry J. Grant ( bjgrant@ucsd.edu ) Course Website: https://bioboot.github.io/bggn213_f17/ DRAFT: 2017-08-10 (15:02:30 PDT on
Ganatum: a graphical single-cell RNA-seq analysis pipeline
 Ganatum: a graphical single-cell RNA-seq analysis pipeline User Manual February 28, 2017 University of Hawaii 2017 Contents 1. Introduction... 1 2. Upload... 1 3. Batch-effect removal... 4 4. Outlier removal...
Ganatum: a graphical single-cell RNA-seq analysis pipeline User Manual February 28, 2017 University of Hawaii 2017 Contents 1. Introduction... 1 2. Upload... 1 3. Batch-effect removal... 4 4. Outlier removal...
Function Prediction of Proteins from their Sequences with BAR 3.0
 Open Access Annals of Proteomics and Bioinformatics Short Communication Function Prediction of Proteins from their Sequences with BAR 3.0 Giuseppe Profiti 1,2, Pier Luigi Martelli 2 and Rita Casadio 2
Open Access Annals of Proteomics and Bioinformatics Short Communication Function Prediction of Proteins from their Sequences with BAR 3.0 Giuseppe Profiti 1,2, Pier Luigi Martelli 2 and Rita Casadio 2
Functional Annotation: Preliminary Results
 Functional Annotation: Preliminary Results Vani Rajan Gena Tang Neha Varghese Kevin Lee Gabriel Mitchell Tripp Jones Robert Petit Shaupu Qin Outline Motivation Naming scheme Preliminary Program Results
Functional Annotation: Preliminary Results Vani Rajan Gena Tang Neha Varghese Kevin Lee Gabriel Mitchell Tripp Jones Robert Petit Shaupu Qin Outline Motivation Naming scheme Preliminary Program Results
GenePainter 2 - Documentation
 GenePainter 2 - Documentation Authors Stefanie Mühlhausen, Björn Hammesfahr, Florian Odronitz, Marcel Hellkamp, Stephan Waack, Martin Kollmar How to cite Stefanie Mühlhausen, Marcel Hellkamp & Martin Kollmar
GenePainter 2 - Documentation Authors Stefanie Mühlhausen, Björn Hammesfahr, Florian Odronitz, Marcel Hellkamp, Stephan Waack, Martin Kollmar How to cite Stefanie Mühlhausen, Marcel Hellkamp & Martin Kollmar
Workday: Frequently Asked Questions (FAQ)
 Workday: Frequently Asked Questions (FAQ) Human Resources This document is intended for Scholastic employees, at all levels, to be able to quickly answer questions they may have as they begin to use the
Workday: Frequently Asked Questions (FAQ) Human Resources This document is intended for Scholastic employees, at all levels, to be able to quickly answer questions they may have as they begin to use the
A tutorial introduction into the MIPS PlantsDB barley&wheat databases. Manuel Spannagl&Kai Bader transplant user training Poznan June 2013
 A tutorial introduction into the MIPS PlantsDB barley&wheat databases Manuel Spannagl&Kai Bader transplant user training Poznan June 2013 MIPS PlantsDB tutorial - some exercises Please go to: http://mips.helmholtz-muenchen.de/plant/genomes.jsp
A tutorial introduction into the MIPS PlantsDB barley&wheat databases Manuel Spannagl&Kai Bader transplant user training Poznan June 2013 MIPS PlantsDB tutorial - some exercises Please go to: http://mips.helmholtz-muenchen.de/plant/genomes.jsp
Introduction to Bioinformatics CPSC 265. What is bioinformatics? Textbooks
 Introduction to Bioinformatics CPSC 265 Thanks to Jonathan Pevsner, Ph.D. Textbooks Johnathan Pevsner, who I stole most of these slides from (thanks!) has written a textbook, Bioinformatics and Functional
Introduction to Bioinformatics CPSC 265 Thanks to Jonathan Pevsner, Ph.D. Textbooks Johnathan Pevsner, who I stole most of these slides from (thanks!) has written a textbook, Bioinformatics and Functional
Massachusetts Institute of Technology Computational Evolutionary Biology, Fall, 2005 Laboratory 3: Detecting selection
 Massachusetts Institute of Technology 6.877 Computational Evolutionary Biology, Fall, 2005 Laboratory 3: Detecting selection Handed out: November 28 Due: December 14 Introduction In this laboratory we
Massachusetts Institute of Technology 6.877 Computational Evolutionary Biology, Fall, 2005 Laboratory 3: Detecting selection Handed out: November 28 Due: December 14 Introduction In this laboratory we
Sequence Based Function Annotation. Qi Sun Bioinformatics Facility Biotechnology Resource Center Cornell University
 Sequence Based Function Annotation Qi Sun Bioinformatics Facility Biotechnology Resource Center Cornell University Usage scenarios for sequence based function annotation Function prediction of newly cloned
Sequence Based Function Annotation Qi Sun Bioinformatics Facility Biotechnology Resource Center Cornell University Usage scenarios for sequence based function annotation Function prediction of newly cloned
quick start guide A quick start guide inflow support GET STARTED WITH INFLOW
 GET STARTED WITH INFLOW quick start guide Welcome to the inflow Community! This quick start guide includes all the important stuff to get you tracking your inventory before you know it! Just follow along
GET STARTED WITH INFLOW quick start guide Welcome to the inflow Community! This quick start guide includes all the important stuff to get you tracking your inventory before you know it! Just follow along
THE HOME DEPOT. Vendor SSR Training Guide
 THE HOME DEPOT Vendor SSR Training Guide REVISION HISTORY: Application and Program Version Changes Modified By 1.0 Document Created: January 2014 IPR Solutions 2.0 Added information regarding Prebuilt
THE HOME DEPOT Vendor SSR Training Guide REVISION HISTORY: Application and Program Version Changes Modified By 1.0 Document Created: January 2014 IPR Solutions 2.0 Added information regarding Prebuilt
Top Tips to Streamline Data Migration and Validation
 Top Tips to Streamline Data Migration and Validation Top Tips for Streamlining Data Migration and Validation Data migration and validation is a critical step when implementing new HR systems. Most systems
Top Tips to Streamline Data Migration and Validation Top Tips for Streamlining Data Migration and Validation Data migration and validation is a critical step when implementing new HR systems. Most systems
From assembled genome to annotated genome
 From assembled genome to annotated genome Procaryotic genomes Eucaryotic genomes Genome annotation servers (web based) 1. RAST 2. NCBI Gene prediction pipeline: Maker Function annotation pipeline: Blast2GO
From assembled genome to annotated genome Procaryotic genomes Eucaryotic genomes Genome annotation servers (web based) 1. RAST 2. NCBI Gene prediction pipeline: Maker Function annotation pipeline: Blast2GO
Finding and Exporting Data. Search
 Finding and Exporting Data Not sure what tool to use to find and export data? VectorBase Search is used to find data for simple queries for one or few genes. In contrast, BioMart is used to retrieve data
Finding and Exporting Data Not sure what tool to use to find and export data? VectorBase Search is used to find data for simple queries for one or few genes. In contrast, BioMart is used to retrieve data
Using the Genome Browser: A Practical Guide. Travis Saari
 Using the Genome Browser: A Practical Guide Travis Saari What is it for? Problem: Bioinformatics programs produce an overwhelming amount of data Difficult to understand anything from the raw data Data
Using the Genome Browser: A Practical Guide Travis Saari What is it for? Problem: Bioinformatics programs produce an overwhelming amount of data Difficult to understand anything from the raw data Data
VTK Finance Tab. Troop Leader Training
 VTK Finance Tab Troop Leader Training 1 Welcome, Troop Leaders and Troop Treasurers to the Volunteer Tool Kit Finance Tab Training. The VTK Finance Tab is a NEW on-line form for Troops to submit their
VTK Finance Tab Troop Leader Training 1 Welcome, Troop Leaders and Troop Treasurers to the Volunteer Tool Kit Finance Tab Training. The VTK Finance Tab is a NEW on-line form for Troops to submit their
Annotation Walkthrough Workshop BIO 173/273 Genomics and Bioinformatics Spring 2013 Developed by Justin R. DiAngelo at Hofstra University
 Annotation Walkthrough Workshop NAME: BIO 173/273 Genomics and Bioinformatics Spring 2013 Developed by Justin R. DiAngelo at Hofstra University A Simple Annotation Exercise Adapted from: Alexis Nagengast,
Annotation Walkthrough Workshop NAME: BIO 173/273 Genomics and Bioinformatics Spring 2013 Developed by Justin R. DiAngelo at Hofstra University A Simple Annotation Exercise Adapted from: Alexis Nagengast,
Discover the Microbes Within: The Wolbachia Project. Bioinformatics Lab
 Bioinformatics Lab ACTIVITY AT A GLANCE "Understanding nature's mute but elegant language of living cells is the quest of modern molecular biology. From an alphabet of only four letters representing the
Bioinformatics Lab ACTIVITY AT A GLANCE "Understanding nature's mute but elegant language of living cells is the quest of modern molecular biology. From an alphabet of only four letters representing the
Tivoli Workload Scheduler
 Tivoli Workload Scheduler Dynamic Workload Console Version 9 Release 2 Quick Reference for Typical Scenarios Table of Contents Introduction... 4 Intended audience... 4 Scope of this publication... 4 User
Tivoli Workload Scheduler Dynamic Workload Console Version 9 Release 2 Quick Reference for Typical Scenarios Table of Contents Introduction... 4 Intended audience... 4 Scope of this publication... 4 User
quick start guide Introducing Discus Guides to key features Welcome to Discus How your account works Creating a DISC profile Inviting a candidate
 www.discusonline.com Introducing Discus Welcome to Discus How your account works Guides to key features Creating a DISC profile Inviting a candidate Reading a DISC report Profiling a job Exploring working
www.discusonline.com Introducing Discus Welcome to Discus How your account works Guides to key features Creating a DISC profile Inviting a candidate Reading a DISC report Profiling a job Exploring working
Analyzing an individual sequence in the Sequence Editor
 BioNumerics Tutorial: Analyzing an individual sequence in the Sequence Editor 1 Aim The Sequence editor window is a convenient tool implemented in BioNumerics to edit and analyze nucleotide and amino acid
BioNumerics Tutorial: Analyzing an individual sequence in the Sequence Editor 1 Aim The Sequence editor window is a convenient tool implemented in BioNumerics to edit and analyze nucleotide and amino acid
Manager Self Service User Reference Guide for Hiring Managers
 Manager Self Service User Reference Guide for Hiring Managers Contents 1. Sign In... 2 2. Set General Preferences... 4 3. Create a Requisition... 8 4. Approve Requisition... 13 5. Manage Candidates...
Manager Self Service User Reference Guide for Hiring Managers Contents 1. Sign In... 2 2. Set General Preferences... 4 3. Create a Requisition... 8 4. Approve Requisition... 13 5. Manage Candidates...
What s New in Discovery Studio 2.5.5
 What s New in Discovery Studio 2.5.5 Discovery Studio takes modeling and simulations to the next level. It brings together the power of validated science on a customizable platform for drug discovery research.
What s New in Discovery Studio 2.5.5 Discovery Studio takes modeling and simulations to the next level. It brings together the power of validated science on a customizable platform for drug discovery research.
Bioinformatics for Proteomics. Ann Loraine
 Bioinformatics for Proteomics Ann Loraine aloraine@uab.edu What is bioinformatics? The science of collecting, processing, organizing, storing, analyzing, and mining biological information, especially data
Bioinformatics for Proteomics Ann Loraine aloraine@uab.edu What is bioinformatics? The science of collecting, processing, organizing, storing, analyzing, and mining biological information, especially data
ONLINE BIOINFORMATICS RESOURCES
 Dedan Githae Email: d.githae@cgiar.org BecA-ILRI Hub; Nairobi, Kenya 16 May, 2014 ONLINE BIOINFORMATICS RESOURCES Introduction to Molecular Biology and Bioinformatics (IMBB) 2014 The larger picture.. Lower
Dedan Githae Email: d.githae@cgiar.org BecA-ILRI Hub; Nairobi, Kenya 16 May, 2014 ONLINE BIOINFORMATICS RESOURCES Introduction to Molecular Biology and Bioinformatics (IMBB) 2014 The larger picture.. Lower
OnCorps Reports 2.0, Standard Reports. Site Supervisor Tutorial
 OnCorps Reports 2.0, Standard Reports Site Supervisor Tutorial i Table of Contents Table of Contents Welcome to OnCorps Reports... 1 Getting Started... 2 System Requirements... 3 Logging Into and Logging
OnCorps Reports 2.0, Standard Reports Site Supervisor Tutorial i Table of Contents Table of Contents Welcome to OnCorps Reports... 1 Getting Started... 2 System Requirements... 3 Logging Into and Logging
Institutional Research & Effectiveness Power BI Quick Start Guide
 Accessing Stetson s Institutional Research Power BI Reports 1. Request Power BI License If you are a new user, you must request a Power BI license. Contact Institutional Research to request a license.
Accessing Stetson s Institutional Research Power BI Reports 1. Request Power BI License If you are a new user, you must request a Power BI license. Contact Institutional Research to request a license.
Tutorial for Stop codon reassignment in the wild
 Tutorial for Stop codon reassignment in the wild Learning Objectives This tutorial has two learning objectives: 1. Finding evidence of stop codon reassignment on DNA fragments. 2. Detecting and confirming
Tutorial for Stop codon reassignment in the wild Learning Objectives This tutorial has two learning objectives: 1. Finding evidence of stop codon reassignment on DNA fragments. 2. Detecting and confirming
my.scouting Tools Training-Home Trend Chart Training Summary Report
 my.scouting Tools Training-Home my.scouting Tools is best experienced using the latest version of Google Chrome or Mozilla Firefox. Also works with the latest version of Safari, and Internet Explorer (v11).
my.scouting Tools Training-Home my.scouting Tools is best experienced using the latest version of Google Chrome or Mozilla Firefox. Also works with the latest version of Safari, and Internet Explorer (v11).
Exploring a fatal outbreak of Escherichia coli using PATRIC
 Exploring a fatal outbreak of Escherichia coli using PATRIC On May 19, 2011, the Robert Koch Institute, Germany's national-level public health authority, was informed about a cluster of three cases of
Exploring a fatal outbreak of Escherichia coli using PATRIC On May 19, 2011, the Robert Koch Institute, Germany's national-level public health authority, was informed about a cluster of three cases of
15/01/2014 Panasonic Parts Ordering User Guide
 15/01/2014 Panasonic Parts Ordering User Guide Panasonic UK, a branch of Panasonic Marketing Europe GmbH 1 15/01/14 Table of Contents Welcome to the Spare Parts Ordering System Bulk Orders 3 Parts Deliveries
15/01/2014 Panasonic Parts Ordering User Guide Panasonic UK, a branch of Panasonic Marketing Europe GmbH 1 15/01/14 Table of Contents Welcome to the Spare Parts Ordering System Bulk Orders 3 Parts Deliveries
Chapter 2: Access to Information
 Chapter 2: Access to Information Outline Introduction to biological databases Centralized databases store DNA sequences Contents of DNA, RNA, and protein databases Central bioinformatics resources: NCBI
Chapter 2: Access to Information Outline Introduction to biological databases Centralized databases store DNA sequences Contents of DNA, RNA, and protein databases Central bioinformatics resources: NCBI
TUTORIAL. Revised in Apr 2015
 TUTORIAL Revised in Apr 2015 Contents I. Overview II. Fly prioritizer Function prioritization III. Fly prioritizer Gene prioritization Gene Set Analysis IV. Human prioritizer Human disease prioritization
TUTORIAL Revised in Apr 2015 Contents I. Overview II. Fly prioritizer Function prioritization III. Fly prioritizer Gene prioritization Gene Set Analysis IV. Human prioritizer Human disease prioritization
Functional profiling of metagenomic short reads: How complex are complex microbial communities?
 Functional profiling of metagenomic short reads: How complex are complex microbial communities? Rohita Sinha Senior Scientist (Bioinformatics), Viracor-Eurofins, Lee s summit, MO Understanding reality,
Functional profiling of metagenomic short reads: How complex are complex microbial communities? Rohita Sinha Senior Scientist (Bioinformatics), Viracor-Eurofins, Lee s summit, MO Understanding reality,
Tutorial. In Silico Cloning. Sample to Insight. March 31, 2016
 In Silico Cloning March 31, 2016 Sample to Insight CLC bio, a QIAGEN Company Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 www.clcbio.com support-clcbio@qiagen.com In Silico Cloning
In Silico Cloning March 31, 2016 Sample to Insight CLC bio, a QIAGEN Company Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 www.clcbio.com support-clcbio@qiagen.com In Silico Cloning
Tutorial. Bisulfite Sequencing. Sample to Insight. September 15, 2016
 Bisulfite Sequencing September 15, 2016 Sample to Insight CLC bio, a QIAGEN Company Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 www.clcbio.com support-clcbio@qiagen.com Bisulfite
Bisulfite Sequencing September 15, 2016 Sample to Insight CLC bio, a QIAGEN Company Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 www.clcbio.com support-clcbio@qiagen.com Bisulfite
Importing Waypoints into Google Earth: Mac
 : Mac Grades 9-12 Overview: Locations stored in a GPS are called waypoints. Waypoints are coordinates that can be transferred from a GPS to a computer. In this activity the waypoint file will be saved
: Mac Grades 9-12 Overview: Locations stored in a GPS are called waypoints. Waypoints are coordinates that can be transferred from a GPS to a computer. In this activity the waypoint file will be saved
Introduction to Mainchain 3 Site Navigation... 5 Favorites... 5 Quick Track... 5 Dashboard & Widgets... 6 Searches
 User Guide Table of Contents Introduction to Mainchain 3 Site Navigation... 5 Favorites... 5 Quick Track... 5 Dashboard & Widgets... 6 Searches... 9 Track and Trace... 9 Domestic Consignment Tracking...
User Guide Table of Contents Introduction to Mainchain 3 Site Navigation... 5 Favorites... 5 Quick Track... 5 Dashboard & Widgets... 6 Searches... 9 Track and Trace... 9 Domestic Consignment Tracking...
Using Waypoints with ProMark3
 Using Waypoints with ProMark3 13 June 2007 Note: These instructions describe the latest versions of ProMark3 receiver firmware and the latest version of MobileMapper Office. You may download the latest
Using Waypoints with ProMark3 13 June 2007 Note: These instructions describe the latest versions of ProMark3 receiver firmware and the latest version of MobileMapper Office. You may download the latest
Halogen Goal / Objective Setting 1AP and 2AP Senior Development Officers Employee Guide
 Halogen Goal / Objective Setting 1AP and 2AP Senior Development Officers Employee Guide Contents Goal / Objective Setting Process... 2 Completing Online Objective Setting Form... 4 Reviewing and Keeping
Halogen Goal / Objective Setting 1AP and 2AP Senior Development Officers Employee Guide Contents Goal / Objective Setting Process... 2 Completing Online Objective Setting Form... 4 Reviewing and Keeping
PRACTICE USER GUIDE. For Practice Performance Registry. Rachel Bryan Updated
 PRACTICE USER GUIDE For Practice Performance Registry Rachel Bryan Rachel.bryan@dartnet.info Updated 01-21-2014 0 P a g e Practice User Guide Welcome, Practice User. This Guide will help you login and
PRACTICE USER GUIDE For Practice Performance Registry Rachel Bryan Rachel.bryan@dartnet.info Updated 01-21-2014 0 P a g e Practice User Guide Welcome, Practice User. This Guide will help you login and
Genomic Annotation Lab Exercise By Jacob Jipp and Marian Kaehler Luther College, Department of Biology Genomics Education Partnership 2010
 Genomic Annotation Lab Exercise By Jacob Jipp and Marian Kaehler Luther College, Department of Biology Genomics Education Partnership 2010 Genomics is a new and expanding field with an increasing impact
Genomic Annotation Lab Exercise By Jacob Jipp and Marian Kaehler Luther College, Department of Biology Genomics Education Partnership 2010 Genomics is a new and expanding field with an increasing impact
Genetics and Bioinformatics
 Genetics and Bioinformatics Kristel Van Steen, PhD 2 Montefiore Institute - Systems and Modeling GIGA - Bioinformatics ULg kristel.vansteen@ulg.ac.be Lecture 1: Setting the pace 1 Bioinformatics what s
Genetics and Bioinformatics Kristel Van Steen, PhD 2 Montefiore Institute - Systems and Modeling GIGA - Bioinformatics ULg kristel.vansteen@ulg.ac.be Lecture 1: Setting the pace 1 Bioinformatics what s
Briefly, this exercise can be summarised by the follow flowchart:
 Workshop exercise Data integration and analysis In this exercise, we would like to work out which GWAS (genome-wide association study) SNP associated with schizophrenia is most likely to be functional.
Workshop exercise Data integration and analysis In this exercise, we would like to work out which GWAS (genome-wide association study) SNP associated with schizophrenia is most likely to be functional.
Supervisor/Manager. 1. Go to Performance Management website: cornerstone.wm.edu. 2. Log in using your W&M network credentials
 1. Go to Performance Management website: cornerstone.wm.edu 2. Log in using your W&M network credentials 3. On the Welcome Page, click on the My Tasks box. Your name here November 2016: v1 Job Aid: Supervisor/Manager
1. Go to Performance Management website: cornerstone.wm.edu 2. Log in using your W&M network credentials 3. On the Welcome Page, click on the My Tasks box. Your name here November 2016: v1 Job Aid: Supervisor/Manager
2. The dropdown box has a number of databases that are searchable. Select the gene option and search for dihydrofolate reductase.
 Bioinformatics Introduction Worksheet The first part of this exercise is aimed at walking you through some of the key tools used by scientists to explore the relationship between genes and proteins throughout
Bioinformatics Introduction Worksheet The first part of this exercise is aimed at walking you through some of the key tools used by scientists to explore the relationship between genes and proteins throughout
SPORTS FACILITY AUDITOR (SFA) - FOOTBALL AUDITOR GUIDE
 SPORTS FACILITY AUDITOR (SFA) - FOOTBALL AUDITOR GUIDE 1 CONTENTS Section Page 1. Access 3 2. Main Menu 4 3. Community Facility Audits - Checklists 5-7 4. Community Facility Audits Site Name and Location
SPORTS FACILITY AUDITOR (SFA) - FOOTBALL AUDITOR GUIDE 1 CONTENTS Section Page 1. Access 3 2. Main Menu 4 3. Community Facility Audits - Checklists 5-7 4. Community Facility Audits Site Name and Location
Introduction to EMBL-EBI.
 Introduction to EMBL-EBI www.ebi.ac.uk What is EMBL-EBI? Part of EMBL Austria, Belgium, Croatia, Denmark, Finland, France, Germany, Greece, Iceland, Ireland, Israel, Italy, Luxembourg, the Netherlands,
Introduction to EMBL-EBI www.ebi.ac.uk What is EMBL-EBI? Part of EMBL Austria, Belgium, Croatia, Denmark, Finland, France, Germany, Greece, Iceland, Ireland, Israel, Italy, Luxembourg, the Netherlands,
UC LEARNING CENTER Manager Guide
 UC LEARNING CENTER Manager Guide Table of Contents Introduction 3 UCLC Overview 3 Browser Capability 3 Obtaining a Logon ID 4 Logging into the UC Learning Center 4 Help Contact 4 Navigation 5 Home Page
UC LEARNING CENTER Manager Guide Table of Contents Introduction 3 UCLC Overview 3 Browser Capability 3 Obtaining a Logon ID 4 Logging into the UC Learning Center 4 Help Contact 4 Navigation 5 Home Page
TUTORIAL: PCR ANALYSIS AND PRIMER DESIGN
 C HAPTER 8 TUTORIAL: PCR ANALYSIS AND PRIMER DESIGN Introduction This chapter introduces you to tools for designing and analyzing PCR primers and procedures. At the end of this tutorial session, you will
C HAPTER 8 TUTORIAL: PCR ANALYSIS AND PRIMER DESIGN Introduction This chapter introduces you to tools for designing and analyzing PCR primers and procedures. At the end of this tutorial session, you will
PIE Performance Process Dashboard & Grid
 PIE Performance Process Dashboard & Grid All Managers who have direct reports will have access to the Dashboards and Analytics functions in Reports under the main menu. Contents TASK : Running PIE Performance
PIE Performance Process Dashboard & Grid All Managers who have direct reports will have access to the Dashboards and Analytics functions in Reports under the main menu. Contents TASK : Running PIE Performance
Invoices 3.3 User Guide
 ! Invoices 3.3 User Guide We Make Software - Ecleti.com Invoices 2007-2018 Ecleti - Roberto Panetta all rights reserved Every effort has been made to ensure that the information in this manual is accurate.
! Invoices 3.3 User Guide We Make Software - Ecleti.com Invoices 2007-2018 Ecleti - Roberto Panetta all rights reserved Every effort has been made to ensure that the information in this manual is accurate.
GenMAPP Gene Database for Escherichia coli K12 Ec-K12-Std_External_ gdb ReadMe
 Page 1 of 6 GenMAPP Gene Database for Escherichia coli K12 Ec-K12-Std_External_20090529.gdb Last revised: 6/4/09 This document contains the following: 1. Overview of GenMAPP application and accessory programs
Page 1 of 6 GenMAPP Gene Database for Escherichia coli K12 Ec-K12-Std_External_20090529.gdb Last revised: 6/4/09 This document contains the following: 1. Overview of GenMAPP application and accessory programs
Motif Discovery in Drosophila
 Motif Discovery in Drosophila Wilson Leung Prerequisites Annotation of Transcription Start Sites in Drosophila Resources Web Site FlyBase The MEME Suite FlyFactorSurvey Web Address http://flybase.org http://meme-suite.org/
Motif Discovery in Drosophila Wilson Leung Prerequisites Annotation of Transcription Start Sites in Drosophila Resources Web Site FlyBase The MEME Suite FlyFactorSurvey Web Address http://flybase.org http://meme-suite.org/
systemsdock Operation Manual
 systemsdock Operation Manual Version 2.0 2016 April systemsdock is being developed by Okinawa Institute of Science and Technology http://www.oist.jp/ Integrated Open Systems Unit http://openbiology.unit.oist.jp/_new/
systemsdock Operation Manual Version 2.0 2016 April systemsdock is being developed by Okinawa Institute of Science and Technology http://www.oist.jp/ Integrated Open Systems Unit http://openbiology.unit.oist.jp/_new/
Internal UVM. How to register and set up your ilab account
 Internal UVM How to register and set up your ilab account This manual should serve as a guide to help you get your account set up. Occasionally, ilab makes changes to the site so the screen shots might
Internal UVM How to register and set up your ilab account This manual should serve as a guide to help you get your account set up. Occasionally, ilab makes changes to the site so the screen shots might
Why Use BLAST? David Form - August 15,
 Wolbachia Workshop 2017 Bioinformatics BLAST Basic Local Alignment Search Tool Finding Model Organisms for Study of Disease Can yeast be used as a model organism to study cystic fibrosis? BLAST Why Use
Wolbachia Workshop 2017 Bioinformatics BLAST Basic Local Alignment Search Tool Finding Model Organisms for Study of Disease Can yeast be used as a model organism to study cystic fibrosis? BLAST Why Use
First Data Personal Financial Manager (PFM) FAQ s
 Q: What browsers are supported with PFM? First Data Personal Financial Manager (PFM) FAQ s A: IE9 and 10, and current versions of Firefox, Chrome, and Safari are supported. Older versions may work but
Q: What browsers are supported with PFM? First Data Personal Financial Manager (PFM) FAQ s A: IE9 and 10, and current versions of Firefox, Chrome, and Safari are supported. Older versions may work but
Gantt chart for bpm'online. User manual
 Gantt chart for bpm'online User manual Contents Core functions 3 Installation 4 Setting up display of the Gantt chart 9 Set up the linear representation 9 Set up the hierarchical representation 10 Gantt
Gantt chart for bpm'online User manual Contents Core functions 3 Installation 4 Setting up display of the Gantt chart 9 Set up the linear representation 9 Set up the hierarchical representation 10 Gantt
Ensembl workshop. Thomas Randall, PhD bioinformatics.unc.edu. handouts, papers, datasets
 Ensembl workshop Thomas Randall, PhD tarandal@email.unc.edu bioinformatics.unc.edu www.unc.edu/~tarandal/ensembl handouts, papers, datasets Ensembl is a joint project between EMBL - EBI and the Sanger
Ensembl workshop Thomas Randall, PhD tarandal@email.unc.edu bioinformatics.unc.edu www.unc.edu/~tarandal/ensembl handouts, papers, datasets Ensembl is a joint project between EMBL - EBI and the Sanger
Collect, analyze and synthesize. Annotation. Annotation for D. virilis. Evidence Based Annotation. GEP goals: Evidence for Gene Models 08/22/2017
 Annotation Annotation for D. virilis Chris Shaffer July 2012 l Big Picture of annotation and then one practical example l This technique may not be the best with other projects (e.g. corn, bacteria) l
Annotation Annotation for D. virilis Chris Shaffer July 2012 l Big Picture of annotation and then one practical example l This technique may not be the best with other projects (e.g. corn, bacteria) l
Halogen Goal / Objective Setting 1AP and 2AP Program Managers Employee Guide
 Halogen Goal / Objective Setting 1AP and 2AP Program Managers Employee Guide Contents Goal / Objective Setting Process... 2 Completing Online Objective Setting Form... 4 Reviewing and Keeping Track of
Halogen Goal / Objective Setting 1AP and 2AP Program Managers Employee Guide Contents Goal / Objective Setting Process... 2 Completing Online Objective Setting Form... 4 Reviewing and Keeping Track of
Usermanual Active Tracing 3.0. Full Visibility. Complete chain of consignment tracking and paperless proof of delivery
 Usermanual Active Tracing 3.0 Full Visibility Complete chain of consignment tracking and paperless proof of delivery Benutzerhandbuch, November 2009 Seite 2 Table of contents Introduction... 3 1. Access...
Usermanual Active Tracing 3.0 Full Visibility Complete chain of consignment tracking and paperless proof of delivery Benutzerhandbuch, November 2009 Seite 2 Table of contents Introduction... 3 1. Access...
