Supplemental Text: Detailed protocol for generating Cas9-mediated fluorescent protein knock-ins with a self-excising selection cassette (SEC)
|
|
|
- Shauna Powell
- 6 years ago
- Views:
Transcription
1 Supplemental Text: Detailed protocol for generating Cas9-mediated fluorescent protein knock-ins with a self-excising selection cassette (SEC) Before the Experiment Choose the Cas9 target site 1) Identify a bp region in which the Cas9 target site should be located. We generally use a 200 bp window centered on the start codon (for N-terminal tags) or stop codon (for C- terminal tags. 2) Submit this genomic sequence to the Zhang lab s CRISPR design tool at Make sure you have selected C. elegans as the genome for checking specificity. 3) The design tool returns a list of potential targeting sequences, ranked in order of predicted specificity. We always try to choose target sites with a specificity score >95, and in most cases find a site that scores 98 or 99 (100 indicates perfect specificity). If there are several candidate sites with high specificity, choose the site that is closest to the desired insertion site (start or stop codon). The best case scenario is to have the insertion site within the guide sequence and within 10 bp of the PAM (NGG motif), so that insertion of will disrupt the target site (Figure P1A). If this is not possible, then choose a Cas9 target sequence that is within the coding region of your gene. For the targeting sequence you choose, copy and paste the list of potential off-target sites into a Word document or Excel sheet and save this list for future reference. Add the target sequence to the Cas9 sgrna construct 1) The design tool returns target sites of the form 5 N 20 -NGG-3, where N is any base. You need to insert the N 20 sequence into the Cas9 sgrna construct (pdd162, Addgene #47549). We use NEB s Q5 Site-Directed Mutagenesis Kit to do this. Use forward primer 5 -N 20 GTTTTAGAGCTAGAAATAGCAAGT-3, where N 20 is your 20 bp targeting sequence from the design tool, and reverse primer 5 -CAAGACATCTCGCAATAGG-3. 2) IMPORTANT: Do not include the PAM (NGG motif) in your primers for the Cas9-sgRNA construct. The NGG motif must be present in the target DNA, but it is not part of the sgrna. 7 SI
2 3) We use sequencing primer 5 -GGTGTGAAATACCGCACAGA-3 to verify correct insertion of the targeting sequence. Design primers to add homology arms to an FP SEC vector Figure 1C shows our strategy for cloning homology arms into FP SEC vectors. Homology arms are generated by PCR and inserted in place of the ccdb negative selection markers, which are flanked by restriction sites. We chose these particular restriction sites so that no residual sequence is left behind after addition of the homology arms. ccdb negative selection makes this cloning strategy exceptionally robust and efficient: in a pilot experiment, we generated 8 repair templates in parallel in a single afternoon. 7/8 of these reactions yielded >80% correct clones; the remaining reaction also yielded correct clones, albeit at a lower frequency. You need to design four primers: two for each homology arm. These primers will amplify the homology arms and add sequence overlaps for Gibson assembly to the ends of each arm. If FP::SEC insertion will not disrupt the Cas9 target site, your primers will also need to introduce silent mutations to prevent Cas9 from cutting the repair template. You have a choice of two possible pairs of restriction enzymes to digest the FP SEC vector: AvrIISpeI or ClaISpeI. If AvrII and SpeI are used, the repair template will include a flexible linker between the 5 homology arm and FP (this is useful for generating C-terminal tags). If ClaI and SpeI are used, the 5 homology arm will be fused directly to the FP, with no added sequence (this is useful for N-terminal tags, or when no flexible linker is desired). Figure P1 shows sample primer designs for each situation. Detailed primer design instructions: 1) First, decide whether additional mutations are needed to prevent Cas9 from cutting the repair template. We make additional mutations whenever the insertion site is not within the 10 bp of the target sequence closest to the PAM. Additional mutations are made using synonymous codons so that the amino acid sequence is not altered (See Figure P1B for an example). If possible, the simplest and most effective approach is to mutate the PAM (NGG motif), since this motif is absolutely required for cleavage of a substrate by Cas9. If a PAM mutation is not feasible, introduce as many mutations as possible (at least 5-6) in the target sequence. 8 SI
3 A 1) Design homology arm primers Upstream vector sequence 5 -acgttgtaaaacgacggccagtcgccggca Genome Cas9 Target (On reverse strand) PRIMER 1 3xFlag PRIMER 3 5 -CGTGATTACAAGGATGACGATGACAAGAGA M E H N D F R V... ctaaacatgcttggcttttatgc-3 PAM ATGGAACACAACGACTTTCG attagctaaacatgcttggcttttatgcat...~500 bp...tccaggctgaccgaga ctatatggaacacaacgactttcgtg...~500 bp...cgcagcaaggactaggagctcctg gtccgactggctct--gata 3 -gtcgttcctgatcctcg TACCAGAGGTTCCCTCTCCTCCTGTTGTA-5 tattgtaccagtatcgacaaaggacacact-5 mneongreen PRIMER 2 PRIMER 4 Downstream vector sequence Cut site 2) Gibson assemble to digested vector 5 -acgttgtaaaacgacggccagtcgccggcactaaacatgcttggcttttatgc...5 Homology arm...caggctgaccgagactatatggtctccaagggagaggaggacaacat acgttgtaaaacgacggccagtcgccggca ATGGTCTCCAAGGGAGAGGAGGACAACAT...3 Upstream vector sequence ctagttacccgaagt...ccdb...cctcaggagcatcg mneongreen SpeI ClaI 5 -CGTGATTACAAGGATGACGATGACAAGAGAATGGAACACAACGACTTTCG...3 Homology arm...cagcaaggactaggagcataacatggtcatagctgtttcctgtgtga CGTGATTACAAGGATGACGATGACAAGAGA ataacatggtcatagctgtttcctgtgtga...3 3xFlag SpeI ctagttcgacctgca...ccdb...gttcctaggaaatcg ClaI Downstream vector sequence 3) Final repair template M V S K G E E D N M R D Y K D D D D K R M E H N D F R V Homology arm...caggctgaccgagactatatggtctccaagggagaggaggacaacat...cgtgattacaaggatgacgatgacaagagaatggaacacaacgactttcg...3 Homology arm... B 1) Design homology arm primers Upstream vector sequence 5 -acgttgtaaaacgacggccagtcgccggca Genome PRIMER 1 3xFlag PRIMER 3 5 -CGTGATTACAAGGATGACGATGACAAGAGA AGATGCGTCAAGGCAGA-3... D E D R D S V R N TAAcctctaaagtttttaattccgg GGATAAGATGCGTCAAGGCAGAAC...~500 bp...atgaagaccgcga CTCAGTTCGCAACTAAcctctaaagtttttaattccggat...~500 bp...tacctgaaatgcaagttaaagattttgacttt CTTCTaGCGCT--GAGTCAAGCGTTG 3 -ggactttacgttcaatttctaaaac PAM CCTCGTAGCCCTCGGAGTCCTCGTAGCTAC-5 ctttagctattgtaccagtatcgacaaagg-5 Cas9 Target Flexible linker mneongreen Cut site (On reverse strand) PRIMER 2 PRIMER 4 Downstream vector sequence 2) Gibson assemble to digested vector Silent Mutation Destroys PAM 5 -acgttgtaaaacgacggccagtcgccggcaagatgcgtcaaggcaga...5 Homology arm...gaagatcgcgactcagttcgcaacggagcatcgggagcctcaggagcatcgatg acgttgtaaaacgacggccagtcgccggca GGAGCATCGGGAGCCTCAGGAGCATCGATG...3 Upstream vector sequence ctagttacccgaagt...ccdb...gcaggtcgacctag Flexible linker mneongreen SpeI AvrII 5 -CGTGATTACAAGGATGACGATGACAAGAGATAAcctctaaagtttttaattccgg...3 Homology arm...cctgaaatgcaagttaaagattttggaaatcgataacatggtcatagctgtttcc CGTGATTACAAGGATGACGATGACAAGAGA gaaatcgataacatggtcatagctgtttcc...3 3xFlag ctagttcgacctgca...ccdb...acttcgggttcctag Downstream vector sequence SpeI AvrII 3) Final repair template... E D R D S V R N G A S G A S G A S M V S R D Y K D D D D K R *...5 Homology arm...gaagatcgcgactcagttcgcaacggagcatcgggagcctcaggagcatcgatggtctcc...cgtgattacaaggatgacgatgacaagagataacctctaaagtttttaattccgg...3 Homology arm... Flexible linker mneongreen 3xFlag Figure P1: Primer design examples. (A) Primers for an N-terminal tag. In this example, insertion disrupts the Cas9 target site, so no mutations are required. (B) Primers for a C-terminal tag, including the optional flexible linker that is built into the FP SEC vectors. In this example, a silent PAM mutation is introduced to block Cas9 cleavage. 9 SI
4 2) Each homology arm should be bp long. The positions of the two primers most proximal to the FP^SEC^3xFlag module (i.e., the reverse primer for the 5 homology arm and the forward primer for the 3 homology arm) are fixed by the need to insert FP^SEC^3xFlag at a specific location. The positions of the distal primers are more flexible. We design the proximal primers first based on our desired insertion site, and then use Primer-BLAST to pick the best possible distal primers. 3) Decide with FP you want to insert, and consult Table P1 for the sequence that needs to be added to the end of each primer to allow Gibson assembly. 4) Ideally, the primer length should be less than 60 bp, because longer primers are much more expensive and fail more often. If you find you need a longer primer because your Cas9 target site is far away from the insertion site, it might be more cost effective to purchase a synthetic DNA fragment (we like IDT s gblocks) containing the homology arm instead of using PCR. 5) Before ordering primers, double check that the will be in frame with your gene of interest. Add homology arms to the repair template 1) Prepare the vector: o Grow bacteria carrying the FP SEC vector and miniprep the plasmid DNA. Note that, prior to replacing the ccdb elements with homology arms, FP SEC vectors must be grown in cells that are resistant to ccdb (ccdb Survival cells). o Digest an entire miniprep of FP SEC vector overnight at 37 C (consult Table P1 for which construct and enzymes to use). o Purify the digested vector using a PCR cleanup spin column to remove the enzymes. Process the entire digested vector as one sample; do not attempt to gel purify individual bands. o The digested, purified vector may be stored at 4 C for at least a few months and reused to construct multiple repair templates. 2) Prepare the homology arms: o Generate two PCR products (the homology arms) using genomic DNA as the template and the primers you designed above. o Mix the two PCR products and purify them together on a single PCR cleanup spin column. 10 SI
5 N-terminal mneongreen::3xflag Digest vector pdd268 with ClaI and SpeI 5' arm reverse primer: 5'- CATGTTGTCCTCCTCTCCCTTGGAGACCAT- (Cas9 target mutations)- (Homology arm sequence)- 3' C-terminal mneongreen::3xflag (with flexible linker) Digest vector pdd268 with AvrII and SpeI N-terminal GFP::3xFlag Digest vector pdd282 with ClaI and SpeI 5' arm reverse primer: 5'- TCCAGTGAACAATTCTTCTCCTTTACTCAT- (Cas9 target mutations)- (Homology arm sequence)- 3' C-terminal GFP::3xFlag (with flexible linker) Digest vector pdd282 with AvrII and SpeI N-terminal YPET::3xFlag Digest vector pdd283 with ClaI and SpeI 5' arm reverse primer: 5'- TCCTGTAAATAACTCTTCTCCTTTTGACAT- (Cas9 target mutations)- (Homology arm sequence)- 3' C-terminal YPET::3xFlag (with flexible linker) Digest vector pdd283 with AvrII and SpeI N-terminal TagRFP-T::3xFlag Digest vector pdd284 with ClaI and SpeI 5' arm forward primer: 5'- gtcacgacgttgtaaaacgacggccagtcg- (Homology arm sequence)- 3' 5' arm reverse primer: 5'- CTTGATGAGCTCCTCTCCCTTGGAGACCAT- (Cas9 target mutations)- (Homology arm sequence)- 3' C-terminal TagRFP-T::3xFlag (with flexible linker) Digest vector pdd284 with AvrII and SpeI 5' arm forward primer: 5'- gtcacgacgttgtaaaacgacggccagtcg- (Homology arm sequence)- 3' N-terminal mkate2::3xflag Digest vector pdd285 with ClaI and SpeI 5' arm forward primer: 5'- gtcacgacgttgtaaaacgacggccagtcg- (Homology arm sequence)- 3' 5' arm reverse primer: 5'- CATGTTTTCTTTAATGAGCTCGGAGACCAT- (Cas9 target mutations)- (Homology arm sequence)- 3' C-terminal mkate2::3xflag (with flexible linker) Digest vector pdd285 with AvrII and SpeI 5' arm forward primer: 5'- gtcacgacgttgtaaaacgacggccagtcg- (Homology arm sequence)- 3' Table P1: Primers for insertion of homology arms into FP::SEC vectors. 11 SI
6 3) Mix 1 µl of vector, 4 µl of homology arms and 5 µl of isothemal assembly enzyme mix (we use NEBuilder HiFi DNA assembly mix from NEB). Incubate 50 C or as directed by the enzyme manufacturer. Transform 2 µl of the reaction to suitable competent cells. 4) Isolate DNA from 3-6 clones and sequence with M13 Forward and Reverse primers to verify correct insertion of the homology arms. This cloning procedure is efficient enough that screening clones prior to sequencing is not necessary. Injections to Generate Knock-ins Day 0: Injection 1) Prepare an injection mix containing the following: 10 ng/µl homologous repair template 50 ng/µl Cas9-sgRNA construct with your targeting sequence Fluorescent co-injection markers (to label extrachromosomal arrays): o 10 ng/µl pgh8 (Prab-3::mCherry neuronal co-injection marker; Addgene #19359) o 5 ng/µl pcfj104 (Pmyo-3::mCherry body wall muscle co-injection marker; Addgene #19328) o 2.5 ng/µl pcfj90 (Pmyo-2::mCherry pharyngeal co-injection marker; Addgene #19327) Prepare plasmid DNA using Invitrogen s PureLink mini-prep kit, which gives high injection efficiencies. 2) Inject the mixture into the gonads of young adult worms of strain N2 (or substitute any strain you like). 3) Transfer the injected worms to new seeded plates (three animals per plate works well in our hands). Use regular NGM plates (no drug) at this stage. Also make a control plate with uninjected worms, so that when you do the drug selection it can serve as a negative control. 4) Put the plates at 25 C and let the worms lay eggs without selection for 2-3 days. Day 2 or 3: Add hygromycin Prepare and filter sterilize a 5 mg/ml hygromycin solution in water. For 6 cm plates poured with 10 ml agar plates, pipet 500 µl of drug onto the surface of each plate of worms, for a final concentration of ~250 µg/ml (if using different size plates, adjust the volume accordingly). Swirl gently so that the solution covers the entire surface of the plate, then let it dry. Put the worms back at 25ºC. Note: In our hands, it does not make any difference whether we add the drug on 12 SI
7 the second or third day after injection, but the drug must be added no later than the third day in order to kill untransformed F1 progeny before they reproduce and overcrowd the plates. Day 6 or 7: Pick initial knock-in worms 1) Examine the plates and identify those that contain Roller (Rol) animals that survived the hygromycin treatment. Knock-in plates should be obvious: there should be lots of animals, they should look totally healthy, and L3 and older worms should be Rol (the Rol phenotype is not expressed in L1 or L2 larvae). Do not waste your time picking from plates that have only a few, sick-looking worms. 2) Candidate knock-in animals are L4/adults that 1) survive hygromycin selection; 2) are Rol; and 3) lack the red fluorescent extrachromosomal array markers. Note that we occasionally see a plate with many wild-type worms that survived selection, but do not pick these they typically carry extrachromosomal arrays or rearrangements. Also note that, in our experience, rare non-fluorescent animals on plates with lots of mcherry() animals (i.e., lots of array animals) are usually false positives. 3) Single 5-10 candidate knock-in adults to new plates without hygromycin. If you do not see any candidate knock-ins at this stage, or if you have fewer lines than you d like, wait 3 days and then examine the plates again. We sometimes find knock-ins 9-10 days after injection (when the F3 are young adults) that were missed during the first round of screening. Day 9 or 10: Look for homozygous plates Look for plates where 100% of L4s and adults are Rollers. These are homozygous knock-in animals. They can be maintained indefinitely, outcrossed if desired, or mated to another genetic background. The strong Rol phenotype makes it very easy to follow the knock-in in crosses (but note that Rol males mate poorly). You can also take L1s from these plates and proceed directly to heat shock to remove the selectable markers. It is straightforward to generate lethal mutations with our strategy, because knock-in alleles can be isolated and maintained as heterozygotes. You will know that your knock-in is lethal if you see only heterozygous plates (i.e., plates with ~1/4 wild-type worms and 1/4 dead embryos). You should expect your initial knock-in to be lethal if you are making an N-terminal tag on an essential gene, because the initial knock-in is a transcriptional null mutation. 13 SI
8 Note: It is impossible to tell whether two strains that originated from the same injection plate derive from independent insertion events or a single insertion event. Therefore, although we single 5-10 worms from each plate in the previous step, we keep only one line from each plate. Selectable marker removal Figure P2 shows the overall scheme for selectable marker removal. In most cases, the initial knock-in is homozygous viable and marker removal is extremely simple (Figure P2A). If the initial knock-in is lethal, marker removal is slightly more complicated because heterozygous knock-in animals segregate wild-type animals at each generation, which makes it impossible to identify animals that have excised the marker based on wild-type phenotype alone (Figure P2). In this situation, there are two choices. If your knock-in strain is visibly fluorescent, you can simply heat shock heterozygotes and identify animals that have excised the marker based on a wild-type phenotype plus visible fluorescence (Figure P2B). If fluorescence in your knock-in strain is too dim to see by eye, you need to mate in a GFP-marked balancer chromosome first (Figure P2C). Mate males carrying an appropriate GFP-marked balancer to Rol knock-in hermaphrodites. Pick GFP-positive, Rol animals from the F1 progeny. These animals should now no longer segregate wild-type progeny in the absence of heat shock (Figure P2C). Use these balanced knock-in worms for subsequent steps. Day 0: Heat shock 1) Pick 6-8 L1/L2 larvae to each of three new plates. It is possible to perform marker excision using older animals, but using young larvae results in the highest efficiency because the germ cells have not yet begun to divide. 2) Heat shock the plates at 34 C for 4 hours (or at 32 C for 4-5 hours) in an air incubator to activate expression of hs::cre. After heat shock, return the plates to 20 C or 25 C. Day 5-7: Pick knock-in animals that have lost the marker Pick wild-type worms to new plates. The animals you will pick will be the F1 progeny of the L1/L2 larvae that you heat shocked in the previous step. Be careful not to pick these animals too early, since the Rol phenotype conferred by sqt-1(d) does not appear until L3. To be safe, we only pick L4 and adult animals at this step. 14 SI
9 A Rol hs::cre Rol WT B Rol hs::cre Dead Rol (mng) WT WT (mng) WT (mng) C Rol X GFP Rol, GFP hs::cre Dead Rol, GFP Balancer Phenotype WT, GFP WT Figure P2: Genetic schemes for marker self-excision. (A) For homozygous viable knock-ins, the situation is simple: after heat shock, any wild-type worms will have lost both copies of SEC. (B) If the knock-in is homozygous lethal, the strain produces 1/4 wild-type progeny at each generation. This makes it impossible to unambiguously identify worms that have lost SEC based on wild-type phenotype alone, although knock-ins may be identifiable if they show visible fluorescence. (C) A simple, 1-step cross to introduce a GFP balancer chromosome results in a strain that does not segregate any wild-type progeny. Heat shock-induced marker excision in this background generates wild-type animals that can be easily and unambiguously identified. 15 SI
Supplementary Information for: Streamlined genome engineering with a self-excising drug selection cassette
 Supplementary Information for: Streamlined genome engineering with a self-excising drug selection cassette Daniel J. Dickinson, Ariel M. Pani, Jennifer K. Heppert, Christopher D. Higgins and Bob Goldstein
Supplementary Information for: Streamlined genome engineering with a self-excising drug selection cassette Daniel J. Dickinson, Ariel M. Pani, Jennifer K. Heppert, Christopher D. Higgins and Bob Goldstein
Protocol for cloning SEC-based repair templates using Gibson assembly and ccdb negative selection
 Protocol for cloning SEC-based repair templates using Gibson assembly and ccdb negative selection Written by Dan Dickinson (daniel.dickinson@austin.utexas.edu) and last updated January 2018. A version
Protocol for cloning SEC-based repair templates using Gibson assembly and ccdb negative selection Written by Dan Dickinson (daniel.dickinson@austin.utexas.edu) and last updated January 2018. A version
Construct Design and Cloning Guide for Cas9-triggered homologous recombination
 Construct Design and Cloning Guide for Cas9-triggered homologous recombination Written by Dan Dickinson (ddickins@live.unc.edu) and last updated December 2013. Reference: Dickinson DJ, Ward JD, Reiner
Construct Design and Cloning Guide for Cas9-triggered homologous recombination Written by Dan Dickinson (ddickins@live.unc.edu) and last updated December 2013. Reference: Dickinson DJ, Ward JD, Reiner
Protocols for cloning SEC-based repair templates using SapTrap assembly
 Protocols for cloning SEC-based repair templates using SapTrap assembly Written by Dan Dickinson (ddickins@live.unc.edu) and last updated July 2016. Overview SapTrap (Schwartz and Jorgensen, 2016) is a
Protocols for cloning SEC-based repair templates using SapTrap assembly Written by Dan Dickinson (ddickins@live.unc.edu) and last updated July 2016. Overview SapTrap (Schwartz and Jorgensen, 2016) is a
File S1. Detailed protocol for CRISPR/Cas9 mediated genome editing with dual marker cassettes
 File S1 Detailed protocol for CRISPR/Cas9 mediated genome editing with dual marker cassettes A) Preparing sgrna vectors We ve modified the strategy to create new sgrna vectors starting with the klp 12
File S1 Detailed protocol for CRISPR/Cas9 mediated genome editing with dual marker cassettes A) Preparing sgrna vectors We ve modified the strategy to create new sgrna vectors starting with the klp 12
Protocols for cloning SEC-based repair templates using SapTrap assembly
 Protocols for cloning SEC-based repair templates using SapTrap assembly Written by Dan Dickinson (daniel.dickinson@austin.utexas.edu) and last updated January 2018. Overview SapTrap (Schwartz and Jorgensen,
Protocols for cloning SEC-based repair templates using SapTrap assembly Written by Dan Dickinson (daniel.dickinson@austin.utexas.edu) and last updated January 2018. Overview SapTrap (Schwartz and Jorgensen,
Mos1 insertion. MosTIC protocol-11/2006. repair template - Mos1 transposase expression - Mos1 excision - DSB formation. homolog arm.
 MosTIC (Mos1 excision induced Transgene Instructed gene Conversion) Valérie Robert (vrobert@biologie.ens.fr) and Jean-Louis Bessereau (jlbesse@biologie.ens.fr) (November 2006) Introduction: MosTIC (Robert
MosTIC (Mos1 excision induced Transgene Instructed gene Conversion) Valérie Robert (vrobert@biologie.ens.fr) and Jean-Louis Bessereau (jlbesse@biologie.ens.fr) (November 2006) Introduction: MosTIC (Robert
Cold Fusion Cloning Kit. Cat. #s MC100A-1, MC101A-1. User Manual
 Fusion Cloning technology Cold Fusion Cloning Kit Store the master mixture and positive controls at -20 C Store the competent cells at -80 C. (ver. 120909) A limited-use label license covers this product.
Fusion Cloning technology Cold Fusion Cloning Kit Store the master mixture and positive controls at -20 C Store the competent cells at -80 C. (ver. 120909) A limited-use label license covers this product.
Creating pentr vectors by BP reaction
 Creating pentr vectors by BP reaction Tanya Lepikhova and Rafael Martinez 15072011 Overview: 1. Design primers to add the attb sites to gene of interest 2. Perform PCR with a high fidelity DNA polymerase
Creating pentr vectors by BP reaction Tanya Lepikhova and Rafael Martinez 15072011 Overview: 1. Design primers to add the attb sites to gene of interest 2. Perform PCR with a high fidelity DNA polymerase
GenBuilder TM Plus Cloning Kit User Manual
 GenBuilder TM Plus Cloning Kit User Manual Cat. No. L00744 Version 11242017 Ⅰ. Introduction... 2 I.1 Product Information... 2 I.2 Kit Contents and Storage... 2 I.3 GenBuilder Cloning Kit Workflow... 2
GenBuilder TM Plus Cloning Kit User Manual Cat. No. L00744 Version 11242017 Ⅰ. Introduction... 2 I.1 Product Information... 2 I.2 Kit Contents and Storage... 2 I.3 GenBuilder Cloning Kit Workflow... 2
GenBuilder TM Plus Cloning Kit User Manual
 GenBuilder TM Plus Cloning Kit User Manual Cat.no L00744 Version 11242017 Ⅰ. Introduction... 2 I.1 Product Information... 2 I.2 Kit Contents and Storage... 2 I.3 GenBuilder Cloning Kit Workflow... 2 Ⅱ.
GenBuilder TM Plus Cloning Kit User Manual Cat.no L00744 Version 11242017 Ⅰ. Introduction... 2 I.1 Product Information... 2 I.2 Kit Contents and Storage... 2 I.3 GenBuilder Cloning Kit Workflow... 2 Ⅱ.
Conversion of plasmids into Gateway compatible cloning
 Conversion of plasmids into Gateway compatible cloning Rafael Martinez 14072011 Overview: 1. Select the right Gateway cassette (A, B or C). 2. Design primers to amplify the right Gateway cassette from
Conversion of plasmids into Gateway compatible cloning Rafael Martinez 14072011 Overview: 1. Select the right Gateway cassette (A, B or C). 2. Design primers to amplify the right Gateway cassette from
Analysis of gene function
 Genome 371, 22 February 2010, Lecture 12 Analysis of gene function Gene knockouts PHASE TWO: INTERPRETATION I THINK I FOUND A CORNER PIECE. 3 BILLION PIECES Analysis of a disease gene Gene knockout or
Genome 371, 22 February 2010, Lecture 12 Analysis of gene function Gene knockouts PHASE TWO: INTERPRETATION I THINK I FOUND A CORNER PIECE. 3 BILLION PIECES Analysis of a disease gene Gene knockout or
Targeted Protein Degradation
 The PITCh dtag vectors for rapid protein degradation of endogenous targets The dissection of complex biological systems requires target-specific control of protein function or abundance. Genetic perturbations
The PITCh dtag vectors for rapid protein degradation of endogenous targets The dissection of complex biological systems requires target-specific control of protein function or abundance. Genetic perturbations
Ben- Shahar Lab CRISPR/RMCE Protocol May 2015
 Ben- Shahar Lab CRISPR/RMCE Protocol May 2015 1. Before you begin a. Useful CRISPR overview for beginners: https://www.addgene.org/crispr/guide/ b. This protocol combines techniques described in these
Ben- Shahar Lab CRISPR/RMCE Protocol May 2015 1. Before you begin a. Useful CRISPR overview for beginners: https://www.addgene.org/crispr/guide/ b. This protocol combines techniques described in these
Nature Biotechnology: doi: /nbt.4166
 Supplementary Figure 1 Validation of correct targeting at targeted locus. (a) by immunofluorescence staining of 2C-HR-CRISPR microinjected embryos cultured to the blastocyst stage. Embryos were stained
Supplementary Figure 1 Validation of correct targeting at targeted locus. (a) by immunofluorescence staining of 2C-HR-CRISPR microinjected embryos cultured to the blastocyst stage. Embryos were stained
Supplementary Methods pcfd5 cloning protocol
 Supplementary Methods cloning protocol vermilion trna grna trna grna U6:3 Terminator AmpR attb is a vector for expressing one or multiple trna-flanked Cas9 grnas under the control of the strong, ubiquitous
Supplementary Methods cloning protocol vermilion trna grna trna grna U6:3 Terminator AmpR attb is a vector for expressing one or multiple trna-flanked Cas9 grnas under the control of the strong, ubiquitous
Data Sheet Quick PCR Cloning Kit
 Data Sheet Quick PCR Cloning Kit 6044 Cornerstone Ct. West, Ste. E DESCRIPTION: The Quick PCR Cloning Kit is a simple and highly efficient method to insert any gene or DNA fragment into a vector, without
Data Sheet Quick PCR Cloning Kit 6044 Cornerstone Ct. West, Ste. E DESCRIPTION: The Quick PCR Cloning Kit is a simple and highly efficient method to insert any gene or DNA fragment into a vector, without
1a. What is the ratio of feathered to unfeathered shanks in the offspring of the above cross?
 Problem Set 5 answers 1. Whether or not the shanks of chickens contains feathers is due to two independently assorting genes. Individuals have unfeathered shanks when they are homozygous for recessive
Problem Set 5 answers 1. Whether or not the shanks of chickens contains feathers is due to two independently assorting genes. Individuals have unfeathered shanks when they are homozygous for recessive
Cat # Box1 Box2. DH5a Competent E. coli cells CCK-20 (20 rxns) 40 µl 40 µl 50 µl x 20 tubes. Choo-Choo Cloning TM Enzyme Mix
 Molecular Cloning Laboratories User Manual Version 3.3 Product name: Choo-Choo Cloning Kits Cat #: CCK-10, CCK-20, CCK-096, CCK-384 Description: Choo-Choo Cloning is a highly efficient directional PCR
Molecular Cloning Laboratories User Manual Version 3.3 Product name: Choo-Choo Cloning Kits Cat #: CCK-10, CCK-20, CCK-096, CCK-384 Description: Choo-Choo Cloning is a highly efficient directional PCR
Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX
 Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX INTRODUCTION The CRISPR/Cas genome editing system consists of a single guide RNA
Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX INTRODUCTION The CRISPR/Cas genome editing system consists of a single guide RNA
Polymerase Chain Reaction
 Polymerase Chain Reaction Amplify your insert or verify its presence 3H Taq platinum PCR mix primers Ultrapure Water PCR tubes PCR machine A. Insert amplification For insert amplification, use the Taq
Polymerase Chain Reaction Amplify your insert or verify its presence 3H Taq platinum PCR mix primers Ultrapure Water PCR tubes PCR machine A. Insert amplification For insert amplification, use the Taq
GenBuilder TM Cloning Kit User Manual
 GenBuilder TM Cloning Kit User Manual Cat.no L00701 Version 11242017 Ⅰ. Introduction... 2 I.1 Product Information... 2 I.2 Kit Contents and Storage... 2 I.3 GenBuilder Cloning Kit Workflow... 2 Ⅱ. DNA
GenBuilder TM Cloning Kit User Manual Cat.no L00701 Version 11242017 Ⅰ. Introduction... 2 I.1 Product Information... 2 I.2 Kit Contents and Storage... 2 I.3 GenBuilder Cloning Kit Workflow... 2 Ⅱ. DNA
peco TM -T7-nGST, Eco cloning Kit User Manual (Patent pending)
 peco TM -T7-nGST, Eco cloning Kit User Manual (Patent pending) Cloning PCR products for E Coli expression of N-term GST-tagged protein Cat# Contents Amounts Application IC-1004 peco-t7-ngst vector built-in
peco TM -T7-nGST, Eco cloning Kit User Manual (Patent pending) Cloning PCR products for E Coli expression of N-term GST-tagged protein Cat# Contents Amounts Application IC-1004 peco-t7-ngst vector built-in
Protocols for cell lines using CRISPR/CAS
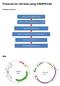 Protocols for cell lines using CRISPR/CAS Procedure overview Map Preparation of CRISPR/CAS plasmids Expression vectors for guide RNA (U6-gRNA) and Cas9 gene (CMV-p-Cas9) are ampicillin-resist ant and stable
Protocols for cell lines using CRISPR/CAS Procedure overview Map Preparation of CRISPR/CAS plasmids Expression vectors for guide RNA (U6-gRNA) and Cas9 gene (CMV-p-Cas9) are ampicillin-resist ant and stable
Amplifying the ALU intron for Hardy- Weinberg Analysis Part 1
 Bio 212 Lab Name: Amplifying the ALU intron for Hardy- Weinberg Analysis Part 1 OBJECTIVES: Review the following terms and concepts presented in Biology 211: enzymes, DNA structure and replication, role
Bio 212 Lab Name: Amplifying the ALU intron for Hardy- Weinberg Analysis Part 1 OBJECTIVES: Review the following terms and concepts presented in Biology 211: enzymes, DNA structure and replication, role
Guide-it sgrna In Vitro Transcription and Screening Systems User Manual
 Clontech Laboratories, Inc. Guide-it sgrna In Vitro Transcription and Screening Systems User Manual Cat. Nos. 631438, 631439 & 631440 (042114) Clontech Laboratories, Inc. A Takara Bio Company 1290 Terra
Clontech Laboratories, Inc. Guide-it sgrna In Vitro Transcription and Screening Systems User Manual Cat. Nos. 631438, 631439 & 631440 (042114) Clontech Laboratories, Inc. A Takara Bio Company 1290 Terra
CELLTECHGEN For Research Only. Construction of all-in-one vector for Lenti-virus system (Example: Lenti-EF1 -Cas9-EGFP-U6 sgrna vector)
 Construction of all-in-one vector for Lenti-virus system (Example: Lenti-EF1 -Cas9-EGFP-U6 sgrna vector) Catalog number: CTG-CAS9-18 Introduction The vector Lenti-EF1 -Cas9-EGFP-U6 sgrna is designed for
Construction of all-in-one vector for Lenti-virus system (Example: Lenti-EF1 -Cas9-EGFP-U6 sgrna vector) Catalog number: CTG-CAS9-18 Introduction The vector Lenti-EF1 -Cas9-EGFP-U6 sgrna is designed for
SUPPLEMENTARY INFORMATION
 Gene replacements and insertions in rice by intron targeting using CRISPR Cas9 Table of Contents Supplementary Figure 1. sgrna-induced targeted mutations in the OsEPSPS gene in rice protoplasts. Supplementary
Gene replacements and insertions in rice by intron targeting using CRISPR Cas9 Table of Contents Supplementary Figure 1. sgrna-induced targeted mutations in the OsEPSPS gene in rice protoplasts. Supplementary
Using CRISPR for genetic alteration
 Using CRISPR for genetic alteration Joffrey Mianné. j.mianne@har.mrc.ac.uk Mary Lyon Centre, MRC Harwell. CRISPR/Cas origins Origin of the CRISPR/Cas system: Clustered-Regularly Interspaced Short Palindromic
Using CRISPR for genetic alteration Joffrey Mianné. j.mianne@har.mrc.ac.uk Mary Lyon Centre, MRC Harwell. CRISPR/Cas origins Origin of the CRISPR/Cas system: Clustered-Regularly Interspaced Short Palindromic
Protocol CRISPR Genome Editing In Cell Lines
 Protocol CRISPR Genome Editing In Cell Lines Protocol 2: HDR donor plasmid applications (gene knockout, gene mutagenesis, gene tagging, Safe Harbor ORF knock-in) Notes: 1. sgrna validation: GeneCopoeia
Protocol CRISPR Genome Editing In Cell Lines Protocol 2: HDR donor plasmid applications (gene knockout, gene mutagenesis, gene tagging, Safe Harbor ORF knock-in) Notes: 1. sgrna validation: GeneCopoeia
Supplemental Protocols Gagnon et al., 2014
 Last updated April 15th 2014 There may be a more up-to-date version of this protocol on our website http://labs.mcb.harvard.edu/schier/index.html Protocol Page Flowchart for Cas9-mediated mutagenesis 1
Last updated April 15th 2014 There may be a more up-to-date version of this protocol on our website http://labs.mcb.harvard.edu/schier/index.html Protocol Page Flowchart for Cas9-mediated mutagenesis 1
Protocol for Genome Editing via the RNA-guided Cas9 Nuclease in. Zebrafish Embryos 1
 Protocol for Genome Editing via the RNA-guided Cas9 Nuclease in Zebrafish Embryos 1 1. In vitro synthesis of capped Cas9 mrna The full length of humanized Cas9 cdnas with double NLS were cloned into pxt7
Protocol for Genome Editing via the RNA-guided Cas9 Nuclease in Zebrafish Embryos 1 1. In vitro synthesis of capped Cas9 mrna The full length of humanized Cas9 cdnas with double NLS were cloned into pxt7
Amplicons, Heteroduplexes and Enzymes - Proper Processing Elevates Detection of CRISPR Gene Editing Events
 Amplicons, Heteroduplexes and Enzymes - Proper Processing Elevates Detection of CRISPR Gene Editing Events Steve Siembieda, MS MBA VP Commercialization ABRF Conference February 2015 What Is CRISPR? Clustered
Amplicons, Heteroduplexes and Enzymes - Proper Processing Elevates Detection of CRISPR Gene Editing Events Steve Siembieda, MS MBA VP Commercialization ABRF Conference February 2015 What Is CRISPR? Clustered
Use of In-Fusion Cloning for Simple and Efficient Assembly of Gene Constructs No restriction enzymes or ligation reactions necessary
 No restriction enzymes or ligation reactions necessary Background The creation of genetic circuits and artificial biological systems typically involves the use of modular genetic components biological
No restriction enzymes or ligation reactions necessary Background The creation of genetic circuits and artificial biological systems typically involves the use of modular genetic components biological
Designing and creating your gene knockout Background The rada gene was identified as a gene, that when mutated, caused cells to become hypersensitive
 Designing and creating your gene knockout Background The rada gene was identified as a gene, that when mutated, caused cells to become hypersensitive to ionizing radiation. However, why these mutants are
Designing and creating your gene knockout Background The rada gene was identified as a gene, that when mutated, caused cells to become hypersensitive to ionizing radiation. However, why these mutants are
HE Swift Cloning Kit
 HE Swift Cloning Kit For high-efficient cloning of PCR products either blunt or sticky-end Kit Contents Contents VTT-BB05 phe Vector (35 ng/µl) 20 µl T4 DNA Ligase (3 U/µl) 20 µl 2 Reaction Buffer 100
HE Swift Cloning Kit For high-efficient cloning of PCR products either blunt or sticky-end Kit Contents Contents VTT-BB05 phe Vector (35 ng/µl) 20 µl T4 DNA Ligase (3 U/µl) 20 µl 2 Reaction Buffer 100
Guide-it sgrna In Vitro Transcription and Screening Systems User Manual
 Guide-it sgrna In Vitro Transcription and Screening Systems User Manual Cat. Nos. 632638, 632639, 632635, 632636, 632637 (040618) 1290 Terra Bella Avenue, Mountain View, CA 94043, USA U.S. Technical Support:
Guide-it sgrna In Vitro Transcription and Screening Systems User Manual Cat. Nos. 632638, 632639, 632635, 632636, 632637 (040618) 1290 Terra Bella Avenue, Mountain View, CA 94043, USA U.S. Technical Support:
Rapid selection of transgenic C. elegans using antibiotic resistance
 nature methods Rapid selection of transgenic C. elegans using antibiotic resistance Jennifer I Semple, Rosa Garcia-Verdugo & Ben Lehner Supplementary figures and text: Supplementary Figure 1 Supplementary
nature methods Rapid selection of transgenic C. elegans using antibiotic resistance Jennifer I Semple, Rosa Garcia-Verdugo & Ben Lehner Supplementary figures and text: Supplementary Figure 1 Supplementary
Supplementary Materials for
 www.sciencemag.org/cgi/content/full/science.aaa5945/dc1 Supplementary Materials for The mutagenic chain reaction: A method for converting heterozygous to homozygous mutations Valentino M. Gantz* and Ethan
www.sciencemag.org/cgi/content/full/science.aaa5945/dc1 Supplementary Materials for The mutagenic chain reaction: A method for converting heterozygous to homozygous mutations Valentino M. Gantz* and Ethan
Figure 1. Map of cloning vector pgem T-Easy (bacterial plasmid DNA)
 Texas A&M University-Corpus Christi CHEM4402 Biochemistry II Laboratory Laboratory 6: Ligation & Bacterial Transformation (Bring your text and laptop to class if you wish to work on your assignment during
Texas A&M University-Corpus Christi CHEM4402 Biochemistry II Laboratory Laboratory 6: Ligation & Bacterial Transformation (Bring your text and laptop to class if you wish to work on your assignment during
Schematic representation of the endogenous PALB2 locus and gene-disruption constructs
 Supplementary Figures Supplementary Figure 1. Generation of PALB2 -/- and BRCA2 -/- /PALB2 -/- DT40 cells. (A) Schematic representation of the endogenous PALB2 locus and gene-disruption constructs carrying
Supplementary Figures Supplementary Figure 1. Generation of PALB2 -/- and BRCA2 -/- /PALB2 -/- DT40 cells. (A) Schematic representation of the endogenous PALB2 locus and gene-disruption constructs carrying
Guide-it Indel Identification Kit User Manual
 Clontech Laboratories, Inc. Guide-it Indel Identification Kit User Manual Cat. No. 631444 (120114) Clontech Laboratories, Inc. A Takara Bio Company 1290 Terra Bella Avenue, Mountain View, CA 94043, USA
Clontech Laboratories, Inc. Guide-it Indel Identification Kit User Manual Cat. No. 631444 (120114) Clontech Laboratories, Inc. A Takara Bio Company 1290 Terra Bella Avenue, Mountain View, CA 94043, USA
CELLTECHGEN For Research Only. Construction of sgrna expression vector for Lenti-virus system (Example: Lenti-U6 sgrna-ef1 -Puro vector)
 Construction of sgrna expression vector for Lenti-virus system (Example: Lenti-U6 sgrna-ef1 -Puro vector) Catalog number: CTG-CAS9-11 Introduction The vector Lenti-U6 sgrna-ef1 -Puro is designed for expression
Construction of sgrna expression vector for Lenti-virus system (Example: Lenti-U6 sgrna-ef1 -Puro vector) Catalog number: CTG-CAS9-11 Introduction The vector Lenti-U6 sgrna-ef1 -Puro is designed for expression
ksierzputowska.com Research Title: Using novel TALEN technology to engineer precise mutations in the genome of D. melanogaster
 Research Title: Using novel TALEN technology to engineer precise mutations in the genome of D. melanogaster Research plan: Specific aims: 1. To successfully engineer transgenic Drosophila expressing TALENs
Research Title: Using novel TALEN technology to engineer precise mutations in the genome of D. melanogaster Research plan: Specific aims: 1. To successfully engineer transgenic Drosophila expressing TALENs
(A) and (B) Experiments 2 and 3 (also see Table 1). Simultaneous small deletion (24bp) and restriction enzyme site (RE) insertion.
 Figure S1 Schematic representation of deletion/insertion experiments on gene Black lines represent genomic DNA. Red lines represent inserted sequence and blue lines are homology arms on template DNA. Scissors
Figure S1 Schematic representation of deletion/insertion experiments on gene Black lines represent genomic DNA. Red lines represent inserted sequence and blue lines are homology arms on template DNA. Scissors
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature09937 a Name Position Primersets 1a 1b 2 3 4 b2 Phenotype Genotype b Primerset 1a D T C R I E 10000 8000 6000 5000 4000 3000 2500 2000 1500 1000 800 Donor (D)
SUPPLEMENTARY INFORMATION doi:10.1038/nature09937 a Name Position Primersets 1a 1b 2 3 4 b2 Phenotype Genotype b Primerset 1a D T C R I E 10000 8000 6000 5000 4000 3000 2500 2000 1500 1000 800 Donor (D)
Genome Sequence Assembly
 Genome Sequence Assembly Learning Goals: Introduce the field of bioinformatics Familiarize the student with performing sequence alignments Understand the assembly process in genome sequencing Introduction:
Genome Sequence Assembly Learning Goals: Introduce the field of bioinformatics Familiarize the student with performing sequence alignments Understand the assembly process in genome sequencing Introduction:
4 Mutant Hunts - To Select or to Screen (Perhaps Even by Brute Force)
 Genetic Techniques for Biological Research Corinne A. Michels Copyright q 2002 John Wiley & Sons, Ltd ISBNs: 0-471-89921-6 (Hardback); 0-470-84662-3 (Electronic) 4 Mutant Hunts - To Select or to Screen
Genetic Techniques for Biological Research Corinne A. Michels Copyright q 2002 John Wiley & Sons, Ltd ISBNs: 0-471-89921-6 (Hardback); 0-470-84662-3 (Electronic) 4 Mutant Hunts - To Select or to Screen
Genetics - Problem Drill 19: Dissection of Gene Function: Mutational Analysis of Model Organisms
 Genetics - Problem Drill 19: Dissection of Gene Function: Mutational Analysis of Model Organisms No. 1 of 10 1. The mouse gene knockout is based on. (A) Homologous recombination (B) Site-specific recombination
Genetics - Problem Drill 19: Dissection of Gene Function: Mutational Analysis of Model Organisms No. 1 of 10 1. The mouse gene knockout is based on. (A) Homologous recombination (B) Site-specific recombination
Supplementary Information
 Supplementary Information Vidigal and Ventura a wt locus 5 region 3 region CCTCTGCCACTGCGAGGGCGTCCAATGGTGCTTG(...)AACAGGTGGAATATCCCTACTCTA predicted deletion clone 1 clone 2 clone 3 CCTCTGCCACTGCGAGGGCGTC-AGGTGGAATATCCCTACTCTA
Supplementary Information Vidigal and Ventura a wt locus 5 region 3 region CCTCTGCCACTGCGAGGGCGTCCAATGGTGCTTG(...)AACAGGTGGAATATCCCTACTCTA predicted deletion clone 1 clone 2 clone 3 CCTCTGCCACTGCGAGGGCGTC-AGGTGGAATATCCCTACTCTA
Lab Book igem Stockholm Lysostaphin. Week 8
 Lysostaphin Week 8 Summarized below are the experiments conducted this week in chronological order. Click on the experiment name to view it. To go back to this summary, click Summary in the footer. Summary
Lysostaphin Week 8 Summarized below are the experiments conducted this week in chronological order. Click on the experiment name to view it. To go back to this summary, click Summary in the footer. Summary
Generation of Non-typeable Haemophilus influenzae Directed Gene Deletion Mutants Jeroen D. Langereis *
 Generation of Non-typeable Haemophilus influenzae Directed Gene Deletion Mutants Jeroen D. Langereis * Laboratory of Pediatric Infectious Diseases, Department of Pediatrics and Laboratory of Medical Immunology,
Generation of Non-typeable Haemophilus influenzae Directed Gene Deletion Mutants Jeroen D. Langereis * Laboratory of Pediatric Infectious Diseases, Department of Pediatrics and Laboratory of Medical Immunology,
Hetero-Stagger PCR Cloning Kit
 Product Name: Code No: Size: DynaExpress Hetero-Stagger PCR Cloning Kit DS150 20 reactions Kit Components: Box 1 (-20 ) phst-1 Vector, linearized Annealing Buffer Ligase Mixture phst Forward Sequence Primer
Product Name: Code No: Size: DynaExpress Hetero-Stagger PCR Cloning Kit DS150 20 reactions Kit Components: Box 1 (-20 ) phst-1 Vector, linearized Annealing Buffer Ligase Mixture phst Forward Sequence Primer
Biology 163 Laboratory in Genetics, Final Exam, Dec. 10, 2005
 1 Biology 163 Laboratory in Genetics, Final Exam, Dec. 10, 2005 Honor Pledge: I have neither given nor received any unauthorized help on this exam: Name Printed: Signature: 1. (2 pts) If you see the following
1 Biology 163 Laboratory in Genetics, Final Exam, Dec. 10, 2005 Honor Pledge: I have neither given nor received any unauthorized help on this exam: Name Printed: Signature: 1. (2 pts) If you see the following
California Institute of Technology. Directed evolution. Dr. F.H. Arnold s lab
 Directed evolution. Dr. F.H. Arnold s lab May 4, 1999 Mutagenic PCR -[Mn] The amount of Mn used in the reaction should be titrated to produce the desired mutagenic rate. Libraries that have close to 30%
Directed evolution. Dr. F.H. Arnold s lab May 4, 1999 Mutagenic PCR -[Mn] The amount of Mn used in the reaction should be titrated to produce the desired mutagenic rate. Libraries that have close to 30%
Gateway Cloning Protocol (Clough Lab Edition) This document is a modification of the Gateway cloning protocol developed by Manju in Chris Taylor's lab
 Gateway Cloning Protocol (Clough Lab Edition) This document is a modification of the Gateway cloning protocol developed by Manju in Chris Taylor's lab With the Gateway cloning system, a PCR fragment is
Gateway Cloning Protocol (Clough Lab Edition) This document is a modification of the Gateway cloning protocol developed by Manju in Chris Taylor's lab With the Gateway cloning system, a PCR fragment is
A protocol for the assembly of 2 or 3 sgrnas. Table of contents
 A protocol for the assembly of 2 or 3 sgrnas Table of contents Simplified protocol... 2 Table 1 Nomenclature of PCR products/primers... 3 Table 2 Structure features of primers... 3 Sequence of DT1T2-PCR
A protocol for the assembly of 2 or 3 sgrnas Table of contents Simplified protocol... 2 Table 1 Nomenclature of PCR products/primers... 3 Table 2 Structure features of primers... 3 Sequence of DT1T2-PCR
5 µl 10X NEB µl ddh 2 O 5 µl StuI (10 units/µl) 2.5 µl XhoI (20 units/µl) *********************************** Incubate at 37 C overnight.
 Flowchart_Cassette Mutagenesis (Steps 0. A-L) 0. Original plasmid pt7-αhl-rl3-d8 In our strategy, for double-mutant E111C-K147C, we will make first K147C by a couple of oligo-pairs shorter than 40 bp (so
Flowchart_Cassette Mutagenesis (Steps 0. A-L) 0. Original plasmid pt7-αhl-rl3-d8 In our strategy, for double-mutant E111C-K147C, we will make first K147C by a couple of oligo-pairs shorter than 40 bp (so
Ready_to_use Fast Seamless Cloning Kit. User Manual
 For general laboratory use. Not for use in diagnostic procedures. FOR IN VITRO USE ONLY. Ready_to_use Fast Seamless Cloning Kit User Manual 1 / 6 Tel: 021-58975266 Fax: 021-50800270 Email:tech@dogene.com
For general laboratory use. Not for use in diagnostic procedures. FOR IN VITRO USE ONLY. Ready_to_use Fast Seamless Cloning Kit User Manual 1 / 6 Tel: 021-58975266 Fax: 021-50800270 Email:tech@dogene.com
NZYGene Synthesis kit
 Kit components Component Concentration Amount NZYGene Synthesis kit Catalogue number: MB33901, 10 reactions GS DNA Polymerase 1U/ μl 30 μl Reaction Buffer for GS DNA Polymerase 10 150 μl dntp mix 2 mm
Kit components Component Concentration Amount NZYGene Synthesis kit Catalogue number: MB33901, 10 reactions GS DNA Polymerase 1U/ μl 30 μl Reaction Buffer for GS DNA Polymerase 10 150 μl dntp mix 2 mm
1a. What is the ratio of feathered to unfeathered shanks in the offspring of the above cross?
 1. Whether or not the shanks of chickens contains feathers is due to two independently assorting genes. Individuals have unfeathered shanks when they are homozygous for recessive genes at two loci; the
1. Whether or not the shanks of chickens contains feathers is due to two independently assorting genes. Individuals have unfeathered shanks when they are homozygous for recessive genes at two loci; the
Simple protocol for gene editing using GenCrisprTM Cas9 nuclease
 Simple protocol for gene editing using GenCrisprTM Cas9 nuclease Contents Protocol Step 1: Choose the target DNA sequence Step 2: Design sgrna Step 3: Preparation for sgrna 3.1 In vitro transcription of
Simple protocol for gene editing using GenCrisprTM Cas9 nuclease Contents Protocol Step 1: Choose the target DNA sequence Step 2: Design sgrna Step 3: Preparation for sgrna 3.1 In vitro transcription of
7.012 Problem Set 5. Question 1
 Name Section 7.012 Problem Set 5 Question 1 While studying the problem of infertility, you attempt to isolate a hypothetical rabbit gene that accounts for the prolific reproduction of rabbits. After much
Name Section 7.012 Problem Set 5 Question 1 While studying the problem of infertility, you attempt to isolate a hypothetical rabbit gene that accounts for the prolific reproduction of rabbits. After much
Rapid amplification of cdna ends (RACE)
 Rapid amplification of cdna ends (RACE) Rapid amplification of cdna ends (RACE) is a technique used in molecular biology to obtain the full length sequence of an RNA transcript found within a cell. RACE
Rapid amplification of cdna ends (RACE) Rapid amplification of cdna ends (RACE) is a technique used in molecular biology to obtain the full length sequence of an RNA transcript found within a cell. RACE
Farnham Lab Protocol for CRISPR/Cas 9- mediated enhancer deletion using a puromycin selection vector Created by Yu (Phoebe) Guo
 Farnham Lab Protocol for CRISPR/Cas 9- mediated enhancer deletion using a puromycin selection vector Created by Yu (Phoebe) Guo 20151024 Part A. Design grna oligos 1. Design grnas Go to Optimized CRISPR
Farnham Lab Protocol for CRISPR/Cas 9- mediated enhancer deletion using a puromycin selection vector Created by Yu (Phoebe) Guo 20151024 Part A. Design grna oligos 1. Design grnas Go to Optimized CRISPR
Easi CRISPR for conditional and insertional alleles
 Easi CRISPR for conditional and insertional alleles C.B Gurumurthy, University Of Nebraska Medical Center Omaha, NE cgurumurthy@unmc.edu Types of Genome edits Gene disruption/inactivation Types of Genome
Easi CRISPR for conditional and insertional alleles C.B Gurumurthy, University Of Nebraska Medical Center Omaha, NE cgurumurthy@unmc.edu Types of Genome edits Gene disruption/inactivation Types of Genome
pgm-t Cloning Kit Cat. # : GVT202 Size : 20 Reactions Store at -20
 pgm-t Cloning Kit Cat. # : GVT202 Size : 20 Reactions Store at -20 1 Kit Contents Contents pgm-t Cloning Kit pgm-t Vector (50 ng/μl) 20 μl T4 DNA Ligase (3 U/μl) 20 μl 10X T4 DNA Ligation Buffer 30 μl
pgm-t Cloning Kit Cat. # : GVT202 Size : 20 Reactions Store at -20 1 Kit Contents Contents pgm-t Cloning Kit pgm-t Vector (50 ng/μl) 20 μl T4 DNA Ligase (3 U/μl) 20 μl 10X T4 DNA Ligation Buffer 30 μl
Perform HiFi PCR Purify PCR-mixture using PCR-clean up kit and determine concentration
 IGEM EXPERIMENT 2 CONSTRUCTION OF PSB#X#-T7CCDB Strategy Cloning Flow Chart (using restriction/ligation): 1. In silico design of cloning Design a cloning strategy (in case of insert/backbone): o Using
IGEM EXPERIMENT 2 CONSTRUCTION OF PSB#X#-T7CCDB Strategy Cloning Flow Chart (using restriction/ligation): 1. In silico design of cloning Design a cloning strategy (in case of insert/backbone): o Using
Supporting Information
 Supporting Information Development of a 2,4-Dinitrotoluene-Responsive Synthetic Riboswitch in E. coli cells Molly E. Davidson, Svetlana V. Harbaugh, Yaroslav G. Chushak, Morley O. Stone, Nancy Kelley-
Supporting Information Development of a 2,4-Dinitrotoluene-Responsive Synthetic Riboswitch in E. coli cells Molly E. Davidson, Svetlana V. Harbaugh, Yaroslav G. Chushak, Morley O. Stone, Nancy Kelley-
SUPPLEMENT MATERIALS FOR CURTIN,
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 SUPPLEMENT MATERIALS FOR CURTIN, et al. Validating genome-wide association candidates: Selecting, testing, and characterizing
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 SUPPLEMENT MATERIALS FOR CURTIN, et al. Validating genome-wide association candidates: Selecting, testing, and characterizing
EZ-10 SPIN COLUMN HANDBOOK
 EZ-0 SPIN COLUMN HANDBOOK EZ-0 Spin Column Plasmid DNA Mini Kit EZ-0 Spin Column PCR Products Purification Kit EZ-0 Spin Column DNA Gel Extraction Kit Version 20.0 Rev 3/23/205 ISO900 Certified 20 Konrad
EZ-0 SPIN COLUMN HANDBOOK EZ-0 Spin Column Plasmid DNA Mini Kit EZ-0 Spin Column PCR Products Purification Kit EZ-0 Spin Column DNA Gel Extraction Kit Version 20.0 Rev 3/23/205 ISO900 Certified 20 Konrad
CRISPR Genome Editing Embryo Microinjection Service Catalog
 CRISPR Genome Editing Embryo Microinjection Service Catalog CRISPR Genome Editing Services (Drosophila) Embryo Microinjection Services (Drosophila & Mosquito) ellgenetics.com +8862-2651-1809 +8862-2782-9911
CRISPR Genome Editing Embryo Microinjection Service Catalog CRISPR Genome Editing Services (Drosophila) Embryo Microinjection Services (Drosophila & Mosquito) ellgenetics.com +8862-2651-1809 +8862-2782-9911
Recombineering Manual
 Recombineering Manual Anthony Popkie The Phiel Laboratory The Research Institute at Nationwide Childrenʼs Hospital 1 BAC Transformation BACs may be transformed into either DY380, EL250 or EL350 cells.
Recombineering Manual Anthony Popkie The Phiel Laboratory The Research Institute at Nationwide Childrenʼs Hospital 1 BAC Transformation BACs may be transformed into either DY380, EL250 or EL350 cells.
Supplementary Information
 Supplementary Information Super-resolution imaging of fluorescently labeled, endogenous RNA Polymerase II in living cells with CRISPR/Cas9-mediated gene editing Won-Ki Cho 1, Namrata Jayanth 1, Susan Mullen
Supplementary Information Super-resolution imaging of fluorescently labeled, endogenous RNA Polymerase II in living cells with CRISPR/Cas9-mediated gene editing Won-Ki Cho 1, Namrata Jayanth 1, Susan Mullen
XactEdit Cas9 Nuclease with NLS User Manual
 XactEdit Cas9 Nuclease with NLS User Manual An RNA-guided recombinant endonuclease for efficient targeted DNA cleavage Catalog Numbers CE1000-50K, CE1000-50, CE1000-250, CE1001-250, CE1001-1000 Table of
XactEdit Cas9 Nuclease with NLS User Manual An RNA-guided recombinant endonuclease for efficient targeted DNA cleavage Catalog Numbers CE1000-50K, CE1000-50, CE1000-250, CE1001-250, CE1001-1000 Table of
Supplementary Figure 1. Diagram for CATCHA construct. Nature Biotechnology doi: /nbt.3444
 Supplementary Figure 1 Diagram for CATCHA construct. Supplementary Figure 2 Representative view of ebony (left) and non-ebony (right) F2 flies from experiments described in Fig. 1c. F0 #1 F0 #2 F0 #3 F0
Supplementary Figure 1 Diagram for CATCHA construct. Supplementary Figure 2 Representative view of ebony (left) and non-ebony (right) F2 flies from experiments described in Fig. 1c. F0 #1 F0 #2 F0 #3 F0
Recitation CHAPTER 9 DNA Technologies
 Recitation CHAPTER 9 DNA Technologies DNA Cloning: General Scheme A cloning vector and eukaryotic chromosomes are separately cleaved with the same restriction endonuclease. (A single chromosome is shown
Recitation CHAPTER 9 DNA Technologies DNA Cloning: General Scheme A cloning vector and eukaryotic chromosomes are separately cleaved with the same restriction endonuclease. (A single chromosome is shown
BRED: Bacteriophage Recombineering with Electroporated DNA
 BRED: Bacteriophage Recombineering with Electroporated DNA Introduction We have developed a system for generating mutations in lytically replicating mycobacteriophages that we have termed BRED: Bacteriophage
BRED: Bacteriophage Recombineering with Electroporated DNA Introduction We have developed a system for generating mutations in lytically replicating mycobacteriophages that we have termed BRED: Bacteriophage
Generation of gene knockout vectors for Dictyostelium discoideum
 Generation of gene knockout vectors for Dictyostelium discoideum Instruction manual Last date of revision April 2012 2015 Version PR29-0001 PR29-0003 www.stargate.com This manual can be downloaded under
Generation of gene knockout vectors for Dictyostelium discoideum Instruction manual Last date of revision April 2012 2015 Version PR29-0001 PR29-0003 www.stargate.com This manual can be downloaded under
Lab Book igem Stockholm Lysostaphin. Week 9
 Lysostaphin Week 9 Summarized below are the experiments conducted this week in chronological order. Click on the experiment name to view it. To go back to this summary, click Summary in the footer. Summary
Lysostaphin Week 9 Summarized below are the experiments conducted this week in chronological order. Click on the experiment name to view it. To go back to this summary, click Summary in the footer. Summary
CLONING INVERTED REPEATS IN HIGH THROUGHPUT
 CLONING INVERTED REPEATS IN HIGH THROUGHPUT This protocol is calculated for cloning one 96 well plate 1. design primers: as standard we include a EcoRI restriction site on the 5 primer and XbaI on the
CLONING INVERTED REPEATS IN HIGH THROUGHPUT This protocol is calculated for cloning one 96 well plate 1. design primers: as standard we include a EcoRI restriction site on the 5 primer and XbaI on the
Ligation Independent Cloning (LIC) Procedure
 Ligation Independent Cloning (LIC) Procedure Ligation Independent Cloning (LIC) LIC cloning allows insertion of DNA fragments without using restriction enzymes into specific vectors containing engineered
Ligation Independent Cloning (LIC) Procedure Ligation Independent Cloning (LIC) LIC cloning allows insertion of DNA fragments without using restriction enzymes into specific vectors containing engineered
CRISPR/Cas9 Genome Editing: Transfection Methods
 CRISPR/ Genome Editing: Transfection Methods For over 20 years Mirus Bio has developed and manufactured high performance transfection products and technologies. That expertise is now being applied to the
CRISPR/ Genome Editing: Transfection Methods For over 20 years Mirus Bio has developed and manufactured high performance transfection products and technologies. That expertise is now being applied to the
Drug selection in worms
 Drug selection in worms The puromycin selection was developed at the Lehner lab: "Rapid selection of transgenic C. elegans using antibiotic resistance." Semple JI, Garcia- Verdugo R, Lehner B. Nature Methods
Drug selection in worms The puromycin selection was developed at the Lehner lab: "Rapid selection of transgenic C. elegans using antibiotic resistance." Semple JI, Garcia- Verdugo R, Lehner B. Nature Methods
3 Designing Primers for Site-Directed Mutagenesis
 3 Designing Primers for Site-Directed Mutagenesis 3.1 Learning Objectives During the next two labs you will learn the basics of site-directed mutagenesis: you will design primers for the mutants you designed
3 Designing Primers for Site-Directed Mutagenesis 3.1 Learning Objectives During the next two labs you will learn the basics of site-directed mutagenesis: you will design primers for the mutants you designed
MUTAGENESIS OF THE PROMOTER OF THE OIL PALM HOMOGENTISATE GERANYLGERANYL TRANSFERASE GENE (HGGT) BY PCR-DRIVEN OVERLAP EXTENSION
 ICBAA2017-30 MUTAGENESIS OF THE PROMOTER OF THE OIL PALM HOMOGENTISATE GERANYLGERANYL TRANSFERASE GENE (HGGT) BY PCR-DRIVEN OVERLAP EXTENSION Mohd Shahrul Nizwanshah Karim and Siti Nor Akmar Abdullah Laboratory
ICBAA2017-30 MUTAGENESIS OF THE PROMOTER OF THE OIL PALM HOMOGENTISATE GERANYLGERANYL TRANSFERASE GENE (HGGT) BY PCR-DRIVEN OVERLAP EXTENSION Mohd Shahrul Nizwanshah Karim and Siti Nor Akmar Abdullah Laboratory
Session 7 Glycerol Stocks & Sequencing Clones
 Session 7 Glycerol Stocks & Sequencing Clones Learning Objective: In this lab you will prepare several of your clones for DNA sequencing and make glycerol stock cultures as a stable and uniform starting
Session 7 Glycerol Stocks & Sequencing Clones Learning Objective: In this lab you will prepare several of your clones for DNA sequencing and make glycerol stock cultures as a stable and uniform starting
Introducing new DNA into the genome requires cloning the donor sequence, delivery of the cloned DNA into the cell, and integration into the genome.
 Key Terms Chapter 32: Genetic Engineering Cloning describes propagation of a DNA sequence by incorporating it into a hybrid construct that can be replicated in a host cell. A cloning vector is a plasmid
Key Terms Chapter 32: Genetic Engineering Cloning describes propagation of a DNA sequence by incorporating it into a hybrid construct that can be replicated in a host cell. A cloning vector is a plasmid
pdsipher and pdsipher -GFP shrna Vector User s Guide
 pdsipher and pdsipher -GFP shrna Vector User s Guide NOTE: PLEASE READ THE ENTIRE PROTOCOL CAREFULLY BEFORE USE Page 1. Introduction... 1 2. Vector Overview... 1 3. Vector Maps 2 4. Materials Provided...
pdsipher and pdsipher -GFP shrna Vector User s Guide NOTE: PLEASE READ THE ENTIRE PROTOCOL CAREFULLY BEFORE USE Page 1. Introduction... 1 2. Vector Overview... 1 3. Vector Maps 2 4. Materials Provided...
A) (5 points) As the starting step isolate genomic DNA from
 GS Final Exam Spring 00 NAME. bub ts is a recessive temperature sensitive mutation in yeast. At º C bub ts cells grow normally, but at º C they die. Use the information below to clone the wild-type BUB
GS Final Exam Spring 00 NAME. bub ts is a recessive temperature sensitive mutation in yeast. At º C bub ts cells grow normally, but at º C they die. Use the information below to clone the wild-type BUB
Supporting Information
 Supporting Information Kilian et al. 10.1073/pnas.1105861108 SI Materials and Methods Determination of the Electric Field Strength Required for Successful Electroporation. The transformation construct
Supporting Information Kilian et al. 10.1073/pnas.1105861108 SI Materials and Methods Determination of the Electric Field Strength Required for Successful Electroporation. The transformation construct
BSCI410-Liu/Spring 06 Exam #1 Feb. 23, 06
 Your Name: Your UID# 1. (20 points) Match following mutations with corresponding mutagens (X-RAY, Ds transposon excision, UV, EMS, Proflavin) a) Thymidine dimmers b) Breakage of DNA backbone c) Frameshift
Your Name: Your UID# 1. (20 points) Match following mutations with corresponding mutagens (X-RAY, Ds transposon excision, UV, EMS, Proflavin) a) Thymidine dimmers b) Breakage of DNA backbone c) Frameshift
pgm-t Cloning Kit For direct cloning of PCR products generated by Taq DNA polymerases For research use only Cat. # : GVT202 Size : 20 Reactions
 pgm-t Cloning Kit For direct cloning of PCR products generated by Taq DNA polymerases Cat. # : GVT202 Size : 20 Reactions Store at -20 For research use only 1 pgm-t Cloning Kit Cat. No.: GVT202 Kit Contents
pgm-t Cloning Kit For direct cloning of PCR products generated by Taq DNA polymerases Cat. # : GVT202 Size : 20 Reactions Store at -20 For research use only 1 pgm-t Cloning Kit Cat. No.: GVT202 Kit Contents
Nature Methods: doi: /nmeth Supplementary Figure 1. Neither driver nor effector alone displays expression of GFP
 Supplementary Figure 1 Neither driver nor effector alone displays expression of GFP Comparison of GFP fluorescence in a driver- only strain (a) or an effector-only strain (b) to their driver + effector
Supplementary Figure 1 Neither driver nor effector alone displays expression of GFP Comparison of GFP fluorescence in a driver- only strain (a) or an effector-only strain (b) to their driver + effector
Supplementary Information
 Single day construction of multi-gene circuits with 3G assembly Andrew D. Halleran 1, Anandh Swaminathan 2, and Richard M. Murray 1, 2 1. Bioengineering, California Institute of Technology, Pasadena, CA.
Single day construction of multi-gene circuits with 3G assembly Andrew D. Halleran 1, Anandh Swaminathan 2, and Richard M. Murray 1, 2 1. Bioengineering, California Institute of Technology, Pasadena, CA.
Experimental genetics - 2 Partha Roy
 Partha Roy Experimental genetics - 2 Making genetically altered animal 1) Gene knock-out k from: a) the entire animal b) selected cell-type/ tissue c) selected cell-type/tissue at certain time 2) Transgenic
Partha Roy Experimental genetics - 2 Making genetically altered animal 1) Gene knock-out k from: a) the entire animal b) selected cell-type/ tissue c) selected cell-type/tissue at certain time 2) Transgenic
