ACCEPTED. Extended-spectrum ß-lactamases of the CTX-M type now in. Switzerland
|
|
|
- Lambert Richardson
- 6 years ago
- Views:
Transcription
1 AAC Accepts, published online ahead of print on 0 April 00 Antimicrob. Agents Chemother. doi:./aac.0-0 Copyright 00, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved Revised AAC -0 Extended-spectrum ß-lactamases of the CTX-M type now in Switzerland Marie-Frédérique Lartigue, 1ψ Catherine Zinsius, 1 Aline Wenger, Jacques Bille, Laurent Poirel, 1 and Patrice Nordmann 1* Service de Bactériologie-Virologie, Hôpital de Bicêtre, Assistance Publique/ Hôpitaux de Paris, Faculté de Médecine Paris-Sud, Université Paris Sud, K.-Bicêtre, France, 1 and Institut de Microbiologie, Centre Hospitalier Universitaire Vaudois, CH Lausanne, Switzerland. Keywords: Extended spectrum ß-lactamases, CTX-M, Switzerland 1 1 Running title: CTX-Ms in Switzerland * Corresponding author. Mailing address: Service de Bactériologie-Virologie, Hôpital de Bicêtre, rue du Général Leclerc, K.-Bicêtre, France. Phone: Fax: nordmann.patrice@bct.aphp.fr 1
2 ψ Present address: Service de Bactériologie-Hygiène, CHRU Trousseau, UFR Médecine, Université François-Rabelais de Tours, 000 Tours Cedex, France.
3 1 Epidemiology of clavulanic-acid inhibited extended-spectrum ß-lactamases (ESBL) was investigated among infection-associated enterobacterial isolates at the University Hospital in Lausanne, Switzerland from January 00 to June 00. Out of non repetitive ESBL producers (prevalence rate of 0.%), produced CTX-M-like ESBLs. CTX-Ms were mostly from clonally non-related Escherichia coli isolates, from urinary infections and community-acquired patients. Pediatric patients (0 out of ) accounted for a large number of CTX-M producers. CTX- M-1 was the most frequent CTX-M-type enzyme. The plasmid-located bla CTX-M genes were associated to either ISEcp1 or ISCR1 insertion sequences. This study is the first published report of CTX-M-type ß-lactamases in Switzerland.
4 Plasmid-mediated extended-spectrum ß-lactamases (ESBLs) were first identified in a Klebsiella pneumoniae isolate in Germany in 1 (1). Then, ESBL-positive Enterobacteriaceae have been isolated worldwide mostly from hospitalized patients (1). These enzymes hydrolyze significantly expanded-spectrum cephalosporins such as cefotaxime, and ceftazidime, and the monobactam aztreonam, sparing carbapenems. Their activity is inhibited in vitro by clavulanic acid. Until the 000 s, most of the ESBLs were structurally related to the narrow-spectrum TEM- and SHV-type ß- lactamases, with single to several amino acid substitutions surrounding their active site (1). Beginning late s, novel types of ESBLs, the CTX-M enzymes, have emerged worldwide, mostly from Escherichia coli. The CTX-Ms are mostly from community- acquired isolates (, ). The over fifty CTX-M enzymes so far reported may be grouped into five main subgroups according to amino acid sequence identity (CTX-M- 1, -M-, -M-, -M-, and -M-) (). Most of the CTX-Ms hydrolyze cefotaxime better than ceftazidime. However, several CTX-Ms including CTX-M-1 (, 1, ), which is now the most widespread CTX-M enzyme worldwide (), hydrolyze ceftazidime better than cefotaxime (). Since those CTX-Ms are reported increasingly in France (, 1, 1, ), Italy (, 0), and recently Austria (), it was interesting to search for those enzymes in Switzerland, a country known to have a tight policy of antibiotic prescription and to have an overall low level of multidrug resistance in bacteria (). The aim of the present study was to estimate the prevalence and the type of the ESBLs produced by enterobacterial isolates among non-repetitive clinical isolates over an 1-month period from January 00 to June 00 at the University hospital of Lausanne, Switzerland.
5 MATERIALS AND METHODS Bacterial isolates. ESBL-producing enterobacterial isolates resulted from the screening of, enterobacterial isolates obtained from infections samples that were sent to the Department of Microbiology of the Lausanne University Hospital, Switzerland, from January 00 to June 00. Isolates were first identified by using the Vitek system (biomérieux SA, Marcy-l Etoile, France). Electrocompetent E. coli TOP (Invitrogen, Cergy Pontoise, France) was used as a recipient strain in transformation experiments. E. coli NCTC 01, harboring 1-, -, -, and -kb plasmids, was used as a plasmid-containing reference strain (). Susceptibility testing and screening for ESBL-producing isolates. The antibiotic susceptibility of enterobacterial clinical isolates was determined by Vitek Advance Expert System (biomerieux) and by the disk diffusion method on Mueller- Hinton (MH) agar plates with ß-lactam and non-ß-lactam antibiotic-containing disks (Sanofi Diagnostics Pasteur, Marnes-La-Coquette, France), according to the Clinical and Laboratory Standards Institute guidelines (). The double-disk synergy test was performed with ceftriaxone, ceftazidime, aztreonam, cefpodoxime and amoxicillin- clavulanic acid disks using different disk spacings (0,, 0, and 0 mm) on MH agar plates, and the results were interpreted as described previously (). The Etest strips containing cefepime cefepime + clavulanic acid (AB Biodisk, Solna, Sweden) was also performed. Then, MICs were determined for selected ß-lactams by an agar dilution technique on MH agar with an inoculum of CFU per spot, as described previously (). MICs of several ß-lactams were determined alone or in combination with a fixed concentration of either clavulanic acid ( g/ml) or tazobactam ( g/ml). MIC values were interpreted according to the guidelines of the Clinical and Laboratory Standards Institute ().
6 PCR amplification for detection of ESBL genes, analysis of genetic their environment, and sequencing. Under standard PCR conditions (0), a series of primers was used for detection of several Ambler class A ß-lactamase genes (Table 1). Detection was performed for genes encoding TEM (PRETEM-1 and PRETEM-), SHV (OS- and OS-), and CTX-M (CTXM-A1 and CTXM-A) (, ). For each reaction, 0. µg of whole-cell DNA of the ESBL-possessing enterobacterial isolates or 0. µg of plasmid DNA from E. coli TOP electroporants was used. The genetic environment of the bla CTX-M genes was characterized by PCR (1) since different genetic elements are associated with the bla CTX-M genes such as ISEcp1- like insertion sequences and the ISCR1 element comprising orf1 which is embedded in a sul1-type integron (). Whole-cell DNA of the isolates was extracted as described previously (1). The regions located upstream of the bla CTX-M genes were amplified with primers annealing to ISEcp1, to orf1 of ISCR1 together with primers for the bla CTX-M genes (). The sequences located downstream to the bla CTX-M genes were studied by PCR experiments with the forward primer CTX-MA1 and the reverse primer CTX-M-preB or IS0Bint. When the orf1 gene was found upstream of bla CTX-M, a PCR experiment was performed with primer CTX-MA1 and reverse primer qaced-1b in order to search for a sul1-type integron structure as previously described with a duplication (even partial) of the ISCR1 element (1, ). For direct DNA sequencing, PCR products were purified using PCR purification columns (Qiagen, Courtaboeuf, France). The sizes of the sequenced bla SHV, bla TEM, bla CTX-M genes were -, 1-, -bp, respectively. Sequencing reactions were performed using specific primers and an automated sequencer (ABI ; Applied Biosystems, Foster City, Calif.). The nucleotide and deduced protein sequences were analyzed with software available over the Internet at the National Center of
7 Biotechnology Information website ( Plasmid transfer and analysis. Plasmid DNAs of enterobacterial isolates were extracted using the Kieser technique (1). They were then electroporated into E. coli TOP, and recombinant strains were selected onto cefotaxime-containing (1 µg/ml) Trypticase Soy agar plates. Plasmid DNAs of these transformants were detected by electrophoresis on a 0.% agarose gel. Hybridization. DNA-DNA hybridizations were performed as described with a Southern transfer of an agarose gel containing plasmid DNA from E. coli TOP electroporants (). The probes consisted in a ca. 00-bp PCR fragment generated from isolate producing CTX-M-1, isolate producing CTX-M-1, and isolate producing CTX-M-. They were internal to the bla CTX-M genes (1). Labeling of the probe and signal detection were carried out using a nonradioactive labeling and detection kit according to the manufacturer instructions (Amersham Pharmacia Biotech). PFGE. Whole-cell DNAs embedded in 1% agarose plugs (Bio-Rad) were digested with XbaI restriction enzyme (Amersham Pharmacia Biotech) and separated in a 1% pulsed-field-certified agarose gel (Bio-Rad) by using a CHEF DRII system (Bio- Rad), as described previously (). Pulsed field gel electrophoresis (PFGE) was run at 1 C, with a V/cm current, a switch angle of, and for E. coli, a run time of 1 h followed by a run time of 1 h, with two linear switch ramps of and 1 s and 1 to s, and for K. pneumoniae, switch times of to s for 0 h. After migration, gels were stained in a 0. mg/ml ethidium bromide solution, and PFGE results were analyzed according to the criteria of Tenover et al. (). RESULTS
8 Epidemiology and PCR detection of ß-lactamase genes. A total of nonrepetitive ESBL-positive isolates were collected from patients. The prevalence of ESBL producers was 0.%. Any enterobacterial isolate flagged as a possible ESBL producer by Vitek Advance Expert System was tested by the double-disk test and by the Etest strip containing cefepime/cefepime + clavulanic acid. They were E. coli (n = 1), Klebsiella pneumoniae (n = 1), and Enterobacter cloacae (n = ) Proteus mirabilis (n =), and Klebsiella oxytoca (n=1). The sex ratio of patients were /, female/man. The isolates were from hospitalized patients (/), outpatients (/), and significantly from the pediatrics (1/). Out of the hospitalized patients, isolates were obtained within the h of their hospitalization. Therefore, community- acquired isolates were from 0 patients ( hospitalized patients + outpatients). Three patients had two different ESBL (+) isolates (E. coli and K. pneumoniae in two cases, and E. coli and P. mirabilis in one case) that were collected at different periods of time. Moreover, a same ESBL (+) isolate was identified from an urinary sample of a woman and from the eyes of her twins. The ESBL (+) isolates were mostly from urine (%), from pus (%) but also from respiratory tract or blood. Most of the patients were treated successfully with a carbapenem-containing antibiotic regimen (data not shown). PCR experiments with primers specific for the bla CTX-M, bla TEM, and bla SHV genes yielded bla CTX-M -positive results for out of the isolates (Table ). A bla CTX-M -gene was identified in 1 out of the 0 community-acquired isolates. The bla TEM and bla SHV genes encoding ESBLs were identified in all the 1 bla CTX-M -negative isolates. A single isolate (E. coli 1) expressed two different ESBLs. Molecular identification of bla CTX-M, bla TEM, and bla SHV genes and genetic environment of bla CTX-M genes. Seventy nine per cent of ESBLs were CTX-M
9 enzymes (/) (Table ). Most of them (%, /) belonged to the CTX-M-1 group: CTX-M-1 being predominant (/). Twenty per cent of the CTX-Ms were CTX-M--like (/) being either CTX-M-1 or CTX-M-. A single isolate produced CTX-M-. A single E. coli isolate produced both CTX-M-1 and CTX-M-1. Sixty per cent of the CTX-M-producers isolates produced also the narrow-spectrum ß-lactamase TEM-1. E. coli was the main ESBL-producer. It was identified in % of the CTX-M- positive isolates and CTX-M enzymes accounted for 1% of the ESBLs in that species. The prevalence of CTX-M among ESBL types was also high in K. pneumoniae (%). Only one out of the six E. cloacae isolates produced a CTX-M enzyme. The other ESBLs were mostly of the SHV-type in E. cloacae and K. pneumoniae (Table ). The genetic structure surrounding the bla CTX-M genes was then determined. ISEcp1 was identified upstream of out of the bla CTX-M (+) genes (1%). The right boundary of ISEcp1 was located between to 0 bp upstream of the start codon of bla CTX-M genes. ISEcp1 was identified at 0 bp upstream of bla CTX-M-1 gene and at bp upstream of bla CTX-M-1 genes. A -bp region with an identical sequence was found between ISEcp1 and the start codon of bla CTX-M- -like genes (bla CTX-M-, and bla CTX-M- 1). Downstream of four out of six bla CTX-M-1 genes (1, 1,, and 1), an IS0-like element was found as already described for this gene (). ISEcp1 was not identified upstream of the bla CTX-M- and bla CTX-M- genes, as previously reported (1). ISCR1 was found in isolates harboring bla CTX-M- and bla CTX-M- genes (Figure). Antibiotic susceptibility results. The isolates were resistant to amino-, ureido-, and carboxy-penicillins, and clavulanic acid and tazobactam addition partially restored the activity of those antibiotics (Table ). Isolates producing CTX-Ms were resistant to cefotaxime whereas this was not always the case for TEM- and SHV producers. The
10 CTX-M-1 producers were resistant also to ceftazidime except two isolates (E. coli, and P. mirabilis ). Susceptibilities to cefepime and cefpirome varied whereas they were constant for carbapenems. Disk diffusion susceptibility testing indicated that overall resistance rate of ESBL producers were,, 0,, 0% for amikacin, sulfonamides, ciprofloxacin, tetracyclines and chloramphenicol, respectively, with no significant difference between CTX-M and non-ctx-m producers (data not shown). PFGE. Analysis of XbaI-restricted DNA of CTX-M-producing E. coli and K. pneumoniae isolates showed that the bla CTX-M -positive isolates were mostly non- clonally related. Only three E. coli isolates (isolates, 1, and ) were clonally related or identical. Those isolates produced CTX-M-1 and two of them (isolates and 1) were identified from patients hospitalized in the same ward. Analysis of the PFGE profiles of CTX-M-1-producing K. pneumoniae isolates showed that three isolates (0, 1, and ) were undistinguishable. They were from urine of a mother of premature twins and from conjonctivitis of her twins. Plasmid analysis and hybridization. Analysis of plasmid DNAs of the CTX-M- producing isolates were extracted and analyzed by gel electrophoresis. Their sizes ranged from to kb with one to five plasmids per isolate. The plasmid DNAs hybridized with any of the three internal probes for bla CTX-M-1-like, bla CTX-M-, and bla CTX- M--like (data not shown). A single hybridization signal was evidenced in each isolate, except for E. coli isolate 1 exhibiting two hybridization signals. These results showed that all except one CTX-M producers contained a single bla CTX-M gene. Those ß- lactamase genes were located on 0- and -kb plasmids. Four out of the seven bla CTX- M-1 genes were located on a similar-in-size plasmid (ca. 0 kb) although the CTX-M producers were not clonally related. The bla CTX-M-1 genes were located mostly on plasmids varying in size. The bla CTX-M-1 and bla CTX-M-1 genes of E. coli isolate 1 were
11 located on two different plasmids of 0-kb and 1-kb, respectively. These results indicated that the spread of bla CTX-M genes resulted from both clonal and plasmid spread. DISCUSSION The results of this work provided insights into the molecular epidemiology of the spread of ESBLs in enterobacterial isolates responsible for infections at a University hospital in Switzerland. The prevalence rate of ESBL producers was 0.% which is a very similar value to that reported in the nearby-located country Austria in 00 (). However this rate is lower than that reported in the Netherlands (.% in Amsterdam, 00 [1]), in France (.% at Bicetre hospital, 00, P Nordmann, personal data), Italy (1.%, 00) (), and the United States (.%, ) (1). Variable prevalence rates of ESBL producers may be related in part to differences in antibiotic policy (more quinolone used than ß-lactams for treating community patients in Switzerland []) but also to differences in urbanization sizes (Paris area or Amsterdam versus Lausanne for example). The bla CTX-M genes were widespread among ESBL producers in (% of ESBL-producers) which was a slightly higher value than that reported at Bicêtre hospital (%) (personal data), in Amsterdam (%) (1), and in Austria (%) (). A recent study performed in several Swiss laboratories between 001 and 00 identified.% of CTX-Ms among ESBLs producers (). ESBLs of E. coli were mostly CTX-Ms at Lausanne (1%), at Bicêtre (%, personal data), in Austria (%, []), in Amsterdam (%, [1]). Moreover, / of the CTX-M producers were E. coli. The prevalence of CTX-Ms among ESBL(+) K. pneumoniae was also high (%) as in Paris (%), whereas this value was much more lower in Austria (0%) and in Italy (1.%) (0).
12 Most of ESBL-producing E. coli isolates (%) have been isolated from urinary tract infections, as found in other studies (,, ). More than one-third of CTX-Mproducing isolates were from community-acquired infections, as reported from other European countries (,, ). Most of the CTX-Ms identified in Switzerland belonged to the CTX-M group 1 (CTX-M-1, ) as reported in France (Bicêtre hospital, [personal data], and Center and South of France [1]), in Austria (), and in Amsterdam (1). CTX-M-1 was the predominant CTX-M in Lausanne (%), as elsewhere in Western Europe (, 1, 0, ). Twenty per cent of the CTX-M enzymes were CTX-M--like in Lausanne, % in Austria, and only % in Bicêtre hospital. Moreover, CTX-M- and the genetically- related CTX-M-1 were the more prevalent CTX-M-types isolated from clinical samples in Spain until 00 (). PFGE analysis showed that the spread of bla CTX-M (+) E. coli and K. pneumoniae isolates was not related to spread of single clones whereas reported in several studies from other countries (UK, France, Italy, Spain) (, 0, 1, ). Concerning non ß-lactam antimicrobial susceptibility of CTX-M producers, high rates of resistance were observed especially for ciprofloxacin (0%), as described in other countries such as in Austria (% ), in Canada (% ), and in Italy (% 0). However, the resistance rate to ciprofloxacin of isolates from pediatric patients was lower (%) than that from isolates from adults (%) which is consistent with a low usage of fluoroquinolones in pediatrics. We identified here the spread of community-acquired CTX-M producers in Switzerland with CTX-M-1 being the most prevalent type as observed now in other European countries. One of the most interesting finding of this study would be the size of the reservoir of CTX-M producers in pediatrics. Finally, spread of CTX-M producers 1
13 in community-acquired E. coli in Switzerland in a similar manner as observed its neighbouring countries may indicate difficulty for controlling those emerging resistance determinants whatever the antibiotic policy is. 1
14 ACKNOWLEDGEMENTS This work was funded by a grant from the Ministère de l'education Nationale et de la Recherche (UPRES-EA), Université Paris XI, France, and by the European Community (th PCRD, LSHM-CT-00-0). L.P. is a researcher from the INSERM, France. We thank Dr Cristina BELLINI for help for the reviewing of the medical charts. 1
15 REFERENCES Al Naiemi, N., A. Bart, M. D. de Jong, C. M. Vandenbroucke-Grauls, P. J. Rietra, Y. J. Debets-Ossenkopp, P. C. Wever, L. Spanjaard, A. J. Bos, and B. Duim. 00. Widely distributed and predominant CTX-M extended-spectrum ß-lactamases in Amsterdam, The Netherlands. J. Clin. Microbiol. : Bonnet, R. 00. Growing group of extended-spectrum ß-lactamases: the CTX- M enzymes. Antimicrob. Agents Chemother. :1-1.. Bou, G., M. Cartelle, M. Tomas, D. Canle, F. Molina, R. Moure, J. M. Eiros, and A. Guerrero. 00. Identification and broad dissemination of the CTX-M-1 ß-lactamase in different Escherichia coli strains in the North- west area of Spain. J. Clin. Microbiol. 0: Brigante, G., F. Luzzaro, M. Perilli, G. Lombardi, A. Coli, G. M. Rossolini, G. Amicosante, and A. Toniolo. 00. Evolution of CTX-M- type ß-lactamases in isolates of Escherichia coli infecting hospital and community patients. Int. J. Antimicrob. Agents. :1-1.. Bruderer, T., M. Jutzi, H. Adler, and R. Frei. 00. The CTX-M is the most predominant group of extended-spectrum ß-lactamases (ESBL) in Switzerland. Swiss Society for Microbiology. th Annual Assembly, Lausanne, March -, P.. Canton, R., and T. M. Coque. 00. The CTX-M ß-lactamase pandemic. Curr. Opin. Microbiol. :-.. Clinical and Laboratory Standards Institute. 00. Performance standards for antimicrobial susceptibility testing; fifteenth informational supplement. 1
16 M0-S1. Clinical and Laboratory Standards Institute, Wayne, PA, USA.. Eckert, C., V. Gautier, M. Saladin-Allard, N. Hidri, C. Verdet, Z. Ould- Hocine, G. Barnaud, F. Delisle, A. Rossier, T. Lambert, A. Philippon, and G. Arlet. 00. Dissemination of CTX-M-type ß-lactamases among clinical isolates of Enterobacteriaceae in Paris, France. Antimicrob. Agents Chemother. :1-1.. Eisner, A., E. J. Fagan, G. Feierl, H. H. Kessler, E. Marth, D. M. Livermore, and N. Woodford. 00 Emergence of Enterobacteriaceae isolates producing CTX-M extended-spectrum ß-lactamase in Austria. Antimicrob. Agents Chemother. 0:-.. Filippini, M., G. Masiero, and K. Moschetti. 00. Socioeconomic determinants of regional differences in outpatient antibiotic consumption: evidence from Switzerland. Health Policy. :-.. Jarlier, V., M. H. Nicolas, G. Fournier, and A. Philippon. 1. Extended broad-spectrum ß-lactamases conferring transferable resistance to newer ß- lactam agents in Enterobacteriaceae: hospital prevalence and susceptibility patterns. Rev. Infect. Dis. :-. 1. Karim, A., L. Poirel, S. Nagarajan, and P. Nordmann Plasmidmediated extended-spectrum ß-lactamase (CTX-M- like) from India and gene association with insertion sequence ISEcp1. FEMS Microbiol. Lett. 01: Kieser, T. 1. Factors affecting the isolation of CCC DNA from Streptomyces lividans and Escherichia coli. Plasmid 1: Knothe, H., P. Shah, V. Krcmery, M. Antal, and S. Mitsuhashi. 1. Transferable resistance to cefotaxime, cefoxitin, cefamandole and cefuroxime in 1
17 clinical isolates of Klebsiella pneumoniae and Serratia marcescens. Infection : Lartigue, M. F., N. Fortineau, and P. Nordmann. 00. Spread of novel expanded-spectrum ß-lactamases in Enterobacteriaceae in a university hospital in the Paris area, France. Clin. Microbiol. Infect. : Lartigue, M. F., L. Poirel, and P. Nordmann. 00. Diversity of genetic environment of bla CTX-M genes. FEMS Microbiol. Lett. : Lavigne, J. P., H. Marchandin, J. Delmas, N. Bouziges, E. Lecaillon, L. Cavalie, H. Jean-Pierre, R. Bonnet, and A. Sotto. 00. qnra in CTX-M- producing Escherichia coli from France. Antimicrob. Agents Chemother. 0:-. 1. Medeiros, A. A. 1. Evolution and dissemination of ß-lactamases accelerated by generations of ß-lactam antibiotics. Clin. Infect. Dis. (Suppl):S Moland, E. S., N. D. Hanson, J. A. Black, A. Hossain, W. Song, and K. S. Thomson. 00. Prevalence of newer ß-lactamases in gram-negative clinical isolates collected in the United States from 001 to 00. J. Clin. Microbiol. : Mugnaioli, C., F. Luzzaro, F. De Luca, G. Brigante, M. Perilli, G. Amicosante, S. Stefani, A. Toniolo, and G. M. Rossolini. 00. CTX-Mtype extended-spectrum ß-lactamases in Italy: molecular epidemiology of an emerging countrywide problem. Antimicrob. Agents Chemother. 0: Oteo, J., C. Navarro, E. Cercenado, A. Delgado-Iribarren, I. Wilhelmi, B. Orden, C. Garcia, S. Miguelanez, M. Perez-Vazquez, S. Garcia- Cobos, B. Aracil, V. Bautista, and J. Campos. 00. Spread of Escherichia coli strains with high-level cefotaxime and ceftazidime resistance 1
18 between the community, long-term facilities, and hospital institutions. J. Clin. Microbiol. :-.. Pitout, J. D., P. Nordmann, K. B. Laupland, and L. Poirel. 00. Emergence of Enterobacteriaceae producing extended-spectrum ß-lactamases (ESBLs) in the community. J. Antimicrob. Chemother. :-.. Poirel, L., M. Gniadkowski, and P. Nordmann. 00. Biochemical analysis of the ceftazidime-hydrolysing extended-spectrum ß-lactamase CTX-M-1 and of its structurally related ß-lactamase CTX-M-. J. Antimicrob. Chemother. 0:1-.. Poirel, L., O. Menuteau, N. Agoli, C. Cattoen, and P. Nordmann. 00. Outbreak of extended spectrum ß-lactamase VEB-1-producing isolates of Acinetobacter baumannii in a French hospital. J. Clin. Microbiol. 1:-.. Poirel, L., T. Naas, I. Le Thomas, A. Karim, E. Bingen, and P. Nordmann CTX-M-type extended-spectrum ß-lactamase that hydrolyzes ceftazidime through a single amino acid substitution in the omega loop. Antimicrob. Agents Chemother. :-1.. Saladin, M., V. T. Cao, T. Lambert, J. L. Donay, J. L. Herrmann, Z. Ould- Hocine, C. Verdet, F. Delisle, A. Philippon, and G. Arlet. 00. Diversity of CTX-M ß-lactamases and their promoter regions from Enterobacteriaceae isolated in three Parisian hospitals. FEMS Microbiol. Lett. 0:-1.. Sambrook, J. E., F. Fritsch, and T. Maniatis. 1. Molecular cloning: a laboratory manual. nd ed. Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press.. Tenover, F. C., R. D. Arbeit, R. V. Goering, P. A. Mickelsen, B. E. Murray, D. H. Persing, and B. Swaminathan. 1. Interpreting chromosomal DNA 1
19 restriction patterns produced by pulsed-field gel electrophoresis: criteria for bacterial strain typing. J. Clin. Microbiol. :-.. Woodford, N., M. E. Ward, M. E. Kaufmann, J. Turton, E. J. Fagan, D. James, A. P. Johnson, R. Pike, M. Warner, T. Cheasty, A. Pearson, S. Harry, J. B. Leach, A. Loughrey, J. A. Lowes, R. E. Warren, and D. M. Livermore. 00. Community and hospital spread of Escherichia coli producing CTX-M extended-spectrum ß-lactamases in the UK. J. Antimicrob. Chemother. :-. 1
20 TABLE 1. Sequences of primers used for detection of bla CTX-M genes and their genetic environment. Primer name Primer sequence Location CTX-MA1 '-scsatgtgcagyaccagtaa-' bla CTX-M gene CTX-MA '-ccg cra tat grt tgg tgg tg-' bla CTX-M gene, reverse primer CTXMA '-gcc gct caa tgt taa cgg-' bla CTX-M gene (CTX-M- group) bla CTX-M gene (CTX-M- group), CTXMB '-gaa acc gtg ggt tac gat-' reverse primer CTXMA '-ctg atg taa cac gga ttg ac-' bla CTX-M gene (CTX-M- group) bla CTX-M gene (CTX-M- group), CTXMC rev '-agc gcc cca tta ttg aga g-' reverse primer TOHOb rev '-tta cag ccc ttc ggc gat-' bla CTX-M gene (CTX-M- group), reverse primer CTXMpréB '-cac ttt gcc gtc gtc taa ggc g-' bla CTX-M gene (CTX-M-1 group), reverse primer ISEcpPROM+ '-tgc tct gtg gat aac ttg c -' ISEcpPROM- '-gca gtc taa att ctt cgt g-' ISEcp1 upstream of the promoter ISEcp1 downstream of the promoter IS0Bint '-gct ttt tga ctt tcc act cgc-' IS0B transposase, reverse primer Orf1-D '-ctc acg ccc tgg caa ggt tt-' Orf1 Orf1-D '-ctt ttg ccc tag ctg cgg t-' Orf1, reverse primer Orf1-'ext '-cag ctg gta gag cag cgt c-' -end of Orf 1, reverse primer OS '-tta tct ccc tgt tag cca cc-' bla SHV gene OS '-gat ttg ctg att tcg ccg g-' bla SHV gene, reverse primer PRETEM-1 '-gta tcc gct cat gag aca ata-' bla TEM gene '-tct aaa gta tat atg agt aaa ctt ggt PRETEM- ctg-' bla TEM gene, reverse primer 0
21 TABLE. Characterization of aquired ß-lactamases in ESBL (+) isolates ß-Lactamases a Isolate CTX-M type TEM type SHV type E. cloacae 1 CTX-M- E. cloacae TEM-1 SHV- E. cloacae SHV- E. cloacae SHV E. cloacae TEM-1 SHV- E. cloacae TEM-1 SHV- E. coli CTX-M-1 TEM-1 E. coli CTX-M-1 TEM-1 E. coli CTX-M-1 E. coli CTX-M-1 TEM-1 E. coli CTX-M-1 E. coli 1 CTX-M-1 E. coli 1 CTX-M-1 E. coli 1 CTX-M-1, CTX-M-1 TEM-1 E. coli 1 CTX-M-1 E. coli 1 CTX-M-1 TEM-1 E. coli 1 CTX-M-1 TEM-1 E. coli 1 CTX-M-1 TEM-1 E. coli 1 TEM- E. coli 0 CTX-M-1 TEM-1 E. coli 1 CTX-M-1 TEM-1 1
22 E. coli CTX-M-1 E. coli CTX-M-1 TEM-1 E. coli TEM-1 SHV- E. coli CTX-M- E. coli CTX-M-1 E. coli CTX-M-1 TEM-1 E. coli CTX-M-1 TEM-1 E. coli CTX-M-1 TEM-1 E. coli 0 CTX-M-1 E. coli 1 TEM- E. coli CTX-M-1 E. coli CTX-M-1 E. coli CTX-M-1 E. coli CTX-M-1 TEM-1 E. coli CTX-M-1 E. coli CTX-M-1 K. oxytoca CTX-M-1 TEM-1 K. pneumoniae CTX-M-1 TEM-1 K. pneumoniae 0 CTX-M- K. pneumoniae 1 TEM-1 SHV- K. pneumoniae TEM-1/TEM- SHV- K. pneumoniae SHV- K. pneumoniae CTX-M-1 TEM-1 K. pneumoniae SHV-a K. pneumoniae CTX-M-1
23 K. pneumoniae CTX-M-1 TEM-1 K. pneumoniae CTX-M-1 1 K. pneumoniae CTX-M-1 K. pneumoniae 0 CTX-M-1 TEM-1 K. pneumoniae 1 CTX-M-1 TEM-1 K. pneumoniae CTX-M-1 TEM-1 K. pneumoniae CTX-M-1 K. pneumoniae CTX-M-1 TEM-1 K. pneumoniae CTX-M-1 TEM-1 P. mirabilis CTX-M-1 TEM-1 P. mirabilis CTX-M-1 TEM-1 a The ESBL names are boldened.
24 TABLE. MICs of ß-lactams for ESBL (+) isolates. ß-Lactam(s) MIC (µg/ml : range) for isolates producing : CTX-M-1 CTX-M-1 CTX-M- CTX-M- CTX-M-1 TEM-ES SHV-ES Amoxicillin > > > > > > > Amoxicillin and clavulanic acid a 1 - -> Ticarcillin > > > > > > > Ticarcillin and clavulanic acid Piperacillin > > - > Piperacillin and tazobactam b Cephalotin > > > > > > Cefoxitin Ceftazidime - 1 -> Cefotaxime >
25 Cefepime Aztreonam Imipenem Ertapenem a Clavulanic acid was at a fixed concentration of µg/ml. b Tazobactam was at a fixed concentration of µg/ml.
26 FIG. Surrounding DNA sequences of the bla CTX-M genes for the forty two isolates. Row 1, E. coli 1, and 1; row, E. coli,,, 1, 1, 1, 1, 0,,,,,,,,,,,, K. oxytoca, K. pneumoniae,,,,,, 0, 1,,,,, P. mirabilis, and ; row, E. coli,, 1, 1, 1, 0, and P. mirabilis.
27 ISEcp1 tnpa ISEcp1 tnpa ISEcp1 tnpa IRR IRR 0-bp -bp IRR -bp bla CTX-M-1 bla CTX-M-1 bla CTX-M-1 IS0 IRL tnpa
Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR
 Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR 1 The problem We wish to clone a yet unknown gene from a known
Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR 1 The problem We wish to clone a yet unknown gene from a known
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
Supplementary. Table 1: Oligonucleotides and Plasmids. complementary to positions from 77 of the SRα '- GCT CTA GAG AAC TTG AAG TAC AGA CTG C
 Supplementary Table 1: Oligonucleotides and Plasmids 913954 5'- GCT CTA GAG AAC TTG AAG TAC AGA CTG C 913955 5'- CCC AAG CTT ACA GTG TGG CCA TTC TGC TG 223396 5'- CGA CGC GTA CAG TGT GGC CAT TCT GCT G
Supplementary Table 1: Oligonucleotides and Plasmids 913954 5'- GCT CTA GAG AAC TTG AAG TAC AGA CTG C 913955 5'- CCC AAG CTT ACA GTG TGG CCA TTC TGC TG 223396 5'- CGA CGC GTA CAG TGT GGC CAT TCT GCT G
Supplemental Data Supplemental Figure 1.
 Supplemental Data Supplemental Figure 1. Silique arrangement in the wild-type, jhs, and complemented lines. Wild-type (WT) (A), the jhs1 mutant (B,C), and the jhs1 mutant complemented with JHS1 (Com) (D)
Supplemental Data Supplemental Figure 1. Silique arrangement in the wild-type, jhs, and complemented lines. Wild-type (WT) (A), the jhs1 mutant (B,C), and the jhs1 mutant complemented with JHS1 (Com) (D)
Arabidopsis actin depolymerizing factor AtADF4 mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB
 Arabidopsis actin depolymerizing factor mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB Files in this Data Supplement: Supplemental Table S1 Supplemental Table
Arabidopsis actin depolymerizing factor mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB Files in this Data Supplement: Supplemental Table S1 Supplemental Table
Table S1. Bacterial strains (Related to Results and Experimental Procedures)
 Table S1. Bacterial strains (Related to Results and Experimental Procedures) Strain number Relevant genotype Source or reference 1045 AB1157 Graham Walker (Donnelly and Walker, 1989) 2458 3084 (MG1655)
Table S1. Bacterial strains (Related to Results and Experimental Procedures) Strain number Relevant genotype Source or reference 1045 AB1157 Graham Walker (Donnelly and Walker, 1989) 2458 3084 (MG1655)
Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH).
 Bisulfite Treatment of DNA Dilute DNA sample to 2µg DNA in 50µl ddh 2 O. Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH). Incubate in a 37ºC water bath for 30 minutes. To 55µl samples
Bisulfite Treatment of DNA Dilute DNA sample to 2µg DNA in 50µl ddh 2 O. Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH). Incubate in a 37ºC water bath for 30 minutes. To 55µl samples
Cat. # Product Size DS130 DynaExpress TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1
 Product Name: Kit Component TA PCR Cloning Kit (ptakn-2) Cat. # Product Size DS130 TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1 2 Ligation Buffer
Product Name: Kit Component TA PCR Cloning Kit (ptakn-2) Cat. # Product Size DS130 TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1 2 Ligation Buffer
ACCEPTED. Laboratory Medicine, Kosin University College of Medicine, , 34 Amnam-Dong,
 AAC Accepts, published online ahead of print on 21 May 2007 Antimicrob. Agents Chemother. doi:10.1128/aac.00279-07 Copyright 2007, American Society for Microbiology and/or the Listed Authors/Institutions.
AAC Accepts, published online ahead of print on 21 May 2007 Antimicrob. Agents Chemother. doi:10.1128/aac.00279-07 Copyright 2007, American Society for Microbiology and/or the Listed Authors/Institutions.
Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis
 1 2 3 4 5 6 7 8 9 10 11 12 Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis Information Research). Exons
1 2 3 4 5 6 7 8 9 10 11 12 Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis Information Research). Exons
Hes6. PPARα. PPARγ HNF4 CD36
 SUPPLEMENTARY INFORMATION Supplementary Table Positions and Sequences of ChIP primers -63 AGGTCACTGCCA -79 AGGTCTGCTGTG Hes6-0067 GGGCAaAGTTCA ACOT -395 GGGGCAgAGTTCA PPARα -309 GGCTCAaAGTTCAaGTTCA CPTa
SUPPLEMENTARY INFORMATION Supplementary Table Positions and Sequences of ChIP primers -63 AGGTCACTGCCA -79 AGGTCTGCTGTG Hes6-0067 GGGCAaAGTTCA ACOT -395 GGGGCAgAGTTCA PPARα -309 GGCTCAaAGTTCAaGTTCA CPTa
Y-chromosomal haplogroup typing Using SBE reaction
 Schematic of multiplex PCR followed by SBE reaction Multiplex PCR Exo SAP purification SBE reaction 5 A 3 ddatp ddgtp 3 T 5 A G 3 T 5 3 5 G C 5 3 3 C 5 ddttp ddctp 5 T 3 T C 3 A 5 3 A 5 5 C 3 3 G 5 3 G
Schematic of multiplex PCR followed by SBE reaction Multiplex PCR Exo SAP purification SBE reaction 5 A 3 ddatp ddgtp 3 T 5 A G 3 T 5 3 5 G C 5 3 3 C 5 ddttp ddctp 5 T 3 T C 3 A 5 3 A 5 5 C 3 3 G 5 3 G
strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular
![strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular](/thumbs/72/66361212.jpg) Additional file 2 Identification of AOX1 in P. pastoris GS115 with a Mut s phenotype Results and Discussion The HBsAg producing strain was originally identified as a Mut s (methanol utilization slow) strain
Additional file 2 Identification of AOX1 in P. pastoris GS115 with a Mut s phenotype Results and Discussion The HBsAg producing strain was originally identified as a Mut s (methanol utilization slow) strain
Supplemental material
 Supplemental material Diversity of O-antigen repeat-unit structures can account for the substantial sequence variation of Wzx translocases Yaoqin Hong and Peter R. Reeves School of Molecular Bioscience,
Supplemental material Diversity of O-antigen repeat-unit structures can account for the substantial sequence variation of Wzx translocases Yaoqin Hong and Peter R. Reeves School of Molecular Bioscience,
PCR analysis was performed to show the presence and the integrity of the var1csa and var-
 Supplementary information: Methods: Table S1: Primer Name Nucleotide sequence (5-3 ) DBL3-F tcc ccg cgg agt gaa aca tca tgt gac tg DBL3-R gac tag ttt ctt tca ata aat cac tcg c DBL5-F cgc cct agg tgc ttc
Supplementary information: Methods: Table S1: Primer Name Nucleotide sequence (5-3 ) DBL3-F tcc ccg cgg agt gaa aca tca tgt gac tg DBL3-R gac tag ttt ctt tca ata aat cac tcg c DBL5-F cgc cct agg tgc ttc
Supplementary Figure 1A A404 Cells +/- Retinoic Acid
 Supplementary Figure 1A A44 Cells +/- Retinoic Acid 1 1 H3 Lys4 di-methylation SM-actin VEC cfos (-) RA (+) RA 14 1 1 8 6 4 H3 Lys79 di-methylation SM-actin VEC cfos (-) RA (+) RA Supplementary Figure
Supplementary Figure 1A A44 Cells +/- Retinoic Acid 1 1 H3 Lys4 di-methylation SM-actin VEC cfos (-) RA (+) RA 14 1 1 8 6 4 H3 Lys79 di-methylation SM-actin VEC cfos (-) RA (+) RA Supplementary Figure
PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells
 Supplementary Information for: PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells Ju Hye Jang 1, Hyun Kim 2, Mi Jung Jang 2, Ju Hyun Cho 1,2,* 1 Research Institute
Supplementary Information for: PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells Ju Hye Jang 1, Hyun Kim 2, Mi Jung Jang 2, Ju Hyun Cho 1,2,* 1 Research Institute
Supporting Information. Copyright Wiley-VCH Verlag GmbH & Co. KGaA, Weinheim, 2006
 Supporting Information Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Supporting Information for Expanding the Genetic
Supporting Information Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Supporting Information for Expanding the Genetic
Supporting information for Biochemistry, 1995, 34(34), , DOI: /bi00034a013
 Supporting information for Biochemistry, 1995, 34(34), 10807 10815, DOI: 10.1021/bi00034a013 LESNIK 10807-1081 Terms & Conditions Electronic Supporting Information files are available without a subscription
Supporting information for Biochemistry, 1995, 34(34), 10807 10815, DOI: 10.1021/bi00034a013 LESNIK 10807-1081 Terms & Conditions Electronic Supporting Information files are available without a subscription
Genetic support of Extended- Spectrum ß-Lactamases
 Genetic support of Extended- Spectrum ß-Lactamases Laurent Poirel Dept of Microbiology (Pr Nordmann) Bicêtre Hospital. South-Paris Medical School. France ESCMID Conference on ESBL 29-31 May 2006 TEM-like
Genetic support of Extended- Spectrum ß-Lactamases Laurent Poirel Dept of Microbiology (Pr Nordmann) Bicêtre Hospital. South-Paris Medical School. France ESCMID Conference on ESBL 29-31 May 2006 TEM-like
Chromosome-Encoded AmpC and CTX-M Extended-Spectrum -Lactamases in Clinical Isolates of Proteus mirabilis from Korea
 ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Apr. 2011, p. 1414 1419 Vol. 55, No. 4 0066-4804/11/$12.00 doi:10.1128/aac.01835-09 Copyright 2011, American Society for Microbiology. All Rights Reserved. Chromosome-Encoded
ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Apr. 2011, p. 1414 1419 Vol. 55, No. 4 0066-4804/11/$12.00 doi:10.1128/aac.01835-09 Copyright 2011, American Society for Microbiology. All Rights Reserved. Chromosome-Encoded
Disease and selection in the human genome 3
 Disease and selection in the human genome 3 Ka/Ks revisited Please sit in row K or forward RBFD: human populations, adaptation and immunity Neandertal Museum, Mettman Germany Sequence genome Measure expression
Disease and selection in the human genome 3 Ka/Ks revisited Please sit in row K or forward RBFD: human populations, adaptation and immunity Neandertal Museum, Mettman Germany Sequence genome Measure expression
Legends for supplementary figures 1-3
 High throughput resistance profiling of Plasmodium falciparum infections based on custom dual indexing and Illumina next generation sequencing-technology Sidsel Nag 1,2 *, Marlene D. Dalgaard 3, Poul-Erik
High throughput resistance profiling of Plasmodium falciparum infections based on custom dual indexing and Illumina next generation sequencing-technology Sidsel Nag 1,2 *, Marlene D. Dalgaard 3, Poul-Erik
Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC
 Supplementary Appendixes Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC ACG TAG CTC CGG CTG GA-3 for vimentin, /5AmMC6/TCC CTC GCG CGT GGC TTC CGC
Supplementary Appendixes Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC ACG TAG CTC CGG CTG GA-3 for vimentin, /5AmMC6/TCC CTC GCG CGT GGC TTC CGC
II 0.95 DM2 (RPP1) DM3 (At3g61540) b
 Table S2. F 2 Segregation Ratios at 16 C, Related to Figure 2 Cross n c Phenotype Model e 2 Locus A Locus B Normal F 1 -like Enhanced d Uk-1/Uk-3 149 64 36 49 DM2 (RPP1) DM1 (SSI4) a Bla-1/Hh-0 F 3 111
Table S2. F 2 Segregation Ratios at 16 C, Related to Figure 2 Cross n c Phenotype Model e 2 Locus A Locus B Normal F 1 -like Enhanced d Uk-1/Uk-3 149 64 36 49 DM2 (RPP1) DM1 (SSI4) a Bla-1/Hh-0 F 3 111
Materials Protein synthesis kit. This kit consists of 24 amino acids, 24 transfer RNAs, four messenger RNAs and one ribosome (see below).
 Protein Synthesis Instructions The purpose of today s lab is to: Understand how a cell manufactures proteins from amino acids, using information stored in the genetic code. Assemble models of four very
Protein Synthesis Instructions The purpose of today s lab is to: Understand how a cell manufactures proteins from amino acids, using information stored in the genetic code. Assemble models of four very
ORFs and genes. Please sit in row K or forward
 ORFs and genes Please sit in row K or forward https://www.flickr.com/photos/teseum/3231682806/in/photostream/ Question: why do some strains of Vibrio cause cholera and others don t? Methods Mechanisms
ORFs and genes Please sit in row K or forward https://www.flickr.com/photos/teseum/3231682806/in/photostream/ Question: why do some strains of Vibrio cause cholera and others don t? Methods Mechanisms
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/10/494/eaan6284/dc1 Supplementary Materials for Activation of master virulence regulator PhoP in acidic ph requires the Salmonella-specific protein UgtL Jeongjoon
www.sciencesignaling.org/cgi/content/full/10/494/eaan6284/dc1 Supplementary Materials for Activation of master virulence regulator PhoP in acidic ph requires the Salmonella-specific protein UgtL Jeongjoon
Supplementary Information. Construction of Lasso Peptide Fusion Proteins
 Supplementary Information Construction of Lasso Peptide Fusion Proteins Chuhan Zong 1, Mikhail O. Maksimov 2, A. James Link 2,3 * Departments of 1 Chemistry, 2 Chemical and Biological Engineering, and
Supplementary Information Construction of Lasso Peptide Fusion Proteins Chuhan Zong 1, Mikhail O. Maksimov 2, A. James Link 2,3 * Departments of 1 Chemistry, 2 Chemical and Biological Engineering, and
ΔPDD1 x ΔPDD1. ΔPDD1 x wild type. 70 kd Pdd1. Pdd3
 Supplemental Fig. S1 ΔPDD1 x wild type ΔPDD1 x ΔPDD1 70 kd Pdd1 50 kd 37 kd Pdd3 Supplemental Fig. S1. ΔPDD1 strains express no detectable Pdd1 protein. Western blot analysis of whole-protein extracts
Supplemental Fig. S1 ΔPDD1 x wild type ΔPDD1 x ΔPDD1 70 kd Pdd1 50 kd 37 kd Pdd3 Supplemental Fig. S1. ΔPDD1 strains express no detectable Pdd1 protein. Western blot analysis of whole-protein extracts
CTX-M-27. Escherichia coli 312 Clinical and Laboratory Standards Institute double-disk synergy test 15 (4.8 ) extendedspectrum
 2011 25 b- CTX-M-27 1) 1, 2) 1, 3) 1) 2) 3) 22 7 30 23 1 14 2007 12 2008 2 7 Escherichia coli 312 Clinical and Laboratory Standards Institute double-disk synergy test 15 (4.8 ) extendedspectrum b-lactamase
2011 25 b- CTX-M-27 1) 1, 2) 1, 3) 1) 2) 3) 22 7 30 23 1 14 2007 12 2008 2 7 Escherichia coli 312 Clinical and Laboratory Standards Institute double-disk synergy test 15 (4.8 ) extendedspectrum b-lactamase
Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were
 1 Supplemental methods 2 3 4 5 6 7 8 9 1 11 12 13 14 15 16 17 18 19 21 22 23 Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were monitored by quantitative reverse-transcription
1 Supplemental methods 2 3 4 5 6 7 8 9 1 11 12 13 14 15 16 17 18 19 21 22 23 Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were monitored by quantitative reverse-transcription
When Carbapenem-Hydrolyzing ß-Lactamase KPC. attributed to outer-membrane protein deficiency coupled with plasmid-mediated
 AAC Accepts, published online ahead of print on 18 July 2011 Antimicrob. Agents Chemother. doi:10.1128/aac.00719-11 Copyright 2011, American Society for Microbiology and/or the Listed Authors/Institutions.
AAC Accepts, published online ahead of print on 18 July 2011 Antimicrob. Agents Chemother. doi:10.1128/aac.00719-11 Copyright 2011, American Society for Microbiology and/or the Listed Authors/Institutions.
Detection and characterization of extended spectrum β-lactamase producing Escherichia coli from poultry of eastern India
 Detection and characterization of extended spectrum β-lactamase producing Escherichia coli from poultry of eastern India Dr. Samiran Bandyopadhyay Scientist Indian Veterinary Research Institute Eastern
Detection and characterization of extended spectrum β-lactamase producing Escherichia coli from poultry of eastern India Dr. Samiran Bandyopadhyay Scientist Indian Veterinary Research Institute Eastern
Multiplexing Genome-scale Engineering
 Multiplexing Genome-scale Engineering Harris Wang, Ph.D. Department of Systems Biology Department of Pathology & Cell Biology http://wanglab.c2b2.columbia.edu Rise of Genomics An Expanding Toolbox Esvelt
Multiplexing Genome-scale Engineering Harris Wang, Ph.D. Department of Systems Biology Department of Pathology & Cell Biology http://wanglab.c2b2.columbia.edu Rise of Genomics An Expanding Toolbox Esvelt
Gene synthesis by circular assembly amplification
 Gene synthesis by circular assembly amplification Duhee Bang & George M Church Supplementary figures and text: Supplementary Figure 1. Dpo4 gene (1.05kb) construction by various methods. Supplementary
Gene synthesis by circular assembly amplification Duhee Bang & George M Church Supplementary figures and text: Supplementary Figure 1. Dpo4 gene (1.05kb) construction by various methods. Supplementary
Resistance, Yonsei University College of Medicine, Seoul, Korea; and 2 Department of
 AAC Accepts, published online ahead of print on March 0 Antimicrob. Agents Chemother. doi:./aac.0- Copyright 0, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved.
AAC Accepts, published online ahead of print on March 0 Antimicrob. Agents Chemother. doi:./aac.0- Copyright 0, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved.
ORIGINAL ARTICLE /j x
 ORIGINAL ARTICLE 10.1111/j.1469-0691.2006.01645.x Development and clinical validation of a molecular diagnostic assay to detect CTX-M-type b-lactamases in Enterobacteriaceae J. D. D. Pitout 1,2,3, N. Hamilton
ORIGINAL ARTICLE 10.1111/j.1469-0691.2006.01645.x Development and clinical validation of a molecular diagnostic assay to detect CTX-M-type b-lactamases in Enterobacteriaceae J. D. D. Pitout 1,2,3, N. Hamilton
Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system
 Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system Dr. Tim Welsink Molecular Biology Transient Gene Expression OUTLINE Short
Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system Dr. Tim Welsink Molecular Biology Transient Gene Expression OUTLINE Short
Supplemental Data. mir156-regulated SPL Transcription. Factors Define an Endogenous Flowering. Pathway in Arabidopsis thaliana
 Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
Prevalence and molecular characterization of clinical isolates of Escherichia coli expressing an AmpC phenotype
 J Antimicrob Chemother 2010; 65: 460 464 doi:10.1093/jac/dkp484 Advance publication 22 January 2010 Prevalence and molecular characterization of clinical isolates of Escherichia coli expressing an AmpC
J Antimicrob Chemother 2010; 65: 460 464 doi:10.1093/jac/dkp484 Advance publication 22 January 2010 Prevalence and molecular characterization of clinical isolates of Escherichia coli expressing an AmpC
JAC A nosocomial outbreak of Pseudomonas aeruginosa isolates expressing the extended-spectrum β-lactamase GES-2 in South Africa
 Journal of Antimicrobial Chemotherapy (2002) 49, 561 565 JAC A nosocomial outbreak of Pseudomonas aeruginosa isolates expressing the extended-spectrum β-lactamase GES-2 in South Africa Laurent Poirel a,
Journal of Antimicrobial Chemotherapy (2002) 49, 561 565 JAC A nosocomial outbreak of Pseudomonas aeruginosa isolates expressing the extended-spectrum β-lactamase GES-2 in South Africa Laurent Poirel a,
SUMMARY. Key words: antibioticresistance, Enterobacteriaceae, ESBL, CTX-M,
 SUMMARY Key words: antibioticresistance, Enterobacteriaceae, ESBL, CTX-M, The doctoral thesis entitled Prevalence of Enterobacteriaceae producing extendedspectrum beta-lactamases (ESBL) isolated from broilers
SUMMARY Key words: antibioticresistance, Enterobacteriaceae, ESBL, CTX-M, The doctoral thesis entitled Prevalence of Enterobacteriaceae producing extendedspectrum beta-lactamases (ESBL) isolated from broilers
MacBlunt PCR Cloning Kit Manual
 MacBlunt PCR Cloning Kit Manual Shipping and Storage MacBlunt PCR Cloning Kits are shipped on dry ice. Each kit contains a box with cloning reagents and an attached bag with Eco-Blue Competent Cells (optional).
MacBlunt PCR Cloning Kit Manual Shipping and Storage MacBlunt PCR Cloning Kits are shipped on dry ice. Each kit contains a box with cloning reagents and an attached bag with Eco-Blue Competent Cells (optional).
Supplemental Table 1. Mutant ADAMTS3 alleles detected in HEK293T clone 4C2. WT CCTGTCACTTTGGTTGATAGC MVLLSLWLIAAALVEVR
 Supplemental Dataset Supplemental Table 1. Mutant ADAMTS3 alleles detected in HEK293T clone 4C2. DNA sequence Amino acid sequence WT CCTGTCACTTTGGTTGATAGC MVLLSLWLIAAALVEVR Allele 1 CCTGTC------------------GATAGC
Supplemental Dataset Supplemental Table 1. Mutant ADAMTS3 alleles detected in HEK293T clone 4C2. DNA sequence Amino acid sequence WT CCTGTCACTTTGGTTGATAGC MVLLSLWLIAAALVEVR Allele 1 CCTGTC------------------GATAGC
การสอบสวนโรคต ดเช อในโรงพยาบาล
 การสอบสวนโรคต ดเช อในโรงพยาบาล MRSA Model in Rajavithi Hospital by Chanwit Tribuddharat, M.D., Ph.D. Department of Microbiology Faculty of Medicine Siriraj Hospital Mahidol University E-mail sictb@mahidol.ac.th
การสอบสวนโรคต ดเช อในโรงพยาบาล MRSA Model in Rajavithi Hospital by Chanwit Tribuddharat, M.D., Ph.D. Department of Microbiology Faculty of Medicine Siriraj Hospital Mahidol University E-mail sictb@mahidol.ac.th
Supporting Information
 Supporting Information Table S1. Oligonucleotide sequences used in this work Oligo DNA A B C D CpG-A CpG-B CpG-C CpG-D Sequence 5 ACA TTC CTA AGT CTG AAA CAT TAC AGC TTG CTA CAC GAG AAG AGC CGC CAT AGT
Supporting Information Table S1. Oligonucleotide sequences used in this work Oligo DNA A B C D CpG-A CpG-B CpG-C CpG-D Sequence 5 ACA TTC CTA AGT CTG AAA CAT TAC AGC TTG CTA CAC GAG AAG AGC CGC CAT AGT
SAY IT WITH DNA: Protein Synthesis Activity by Larry Flammer
 TEACHER S GUIDE SAY IT WITH DNA: Protein Synthesis Activity by Larry Flammer SYNOPSIS This activity uses the metaphor of decoding a secret message for the Protein Synthesis process. Students teach themselves
TEACHER S GUIDE SAY IT WITH DNA: Protein Synthesis Activity by Larry Flammer SYNOPSIS This activity uses the metaphor of decoding a secret message for the Protein Synthesis process. Students teach themselves
JOURNAL OF INTERNATIONAL ACADEMIC RESEARCH FOR MULTIDISCIPLINARY Impact Factor 1.393, ISSN: , Volume 2, Issue 8, September 2014
 EVALUATION OF CHROMAGAR-CTX FOR THE DETECTION OF CTX-M-ESBL- PRODUCING GRAM NEGATIVE BACTERIAL ISOLATES RASHA H. ELSHERIF* LAMIAA A. MADKOUR** REHAM A. DWEDAR*** *Clinical Pathology Department, Faculty
EVALUATION OF CHROMAGAR-CTX FOR THE DETECTION OF CTX-M-ESBL- PRODUCING GRAM NEGATIVE BACTERIAL ISOLATES RASHA H. ELSHERIF* LAMIAA A. MADKOUR** REHAM A. DWEDAR*** *Clinical Pathology Department, Faculty
SUPPLEMENTARY MATERIALS AND METHODS. E. coli strains, plasmids, and growth conditions. Escherichia coli strain P90C (1)
 SUPPLEMENTARY MATERIALS AND METHODS E. coli strains, plasmids, and growth conditions. Escherichia coli strain P90C (1) dinb::kan (lab stock) derivative was used as wild-type. MG1655 alka tag dinb (2) is
SUPPLEMENTARY MATERIALS AND METHODS E. coli strains, plasmids, and growth conditions. Escherichia coli strain P90C (1) dinb::kan (lab stock) derivative was used as wild-type. MG1655 alka tag dinb (2) is
Overexpression Normal expression Overexpression Normal expression. 26 (21.1%) N (%) P-value a N (%)
 SUPPLEMENTARY TABLES Table S1. Alteration of ZNF322A protein expression levels in relation to clinicopathological parameters in 123 Asian and 74 Caucasian lung cancer patients. Asian patients Caucasian
SUPPLEMENTARY TABLES Table S1. Alteration of ZNF322A protein expression levels in relation to clinicopathological parameters in 123 Asian and 74 Caucasian lung cancer patients. Asian patients Caucasian
An engineered tryptophan zipper-type peptide as a molecular recognition scaffold
 SUPPLEMENTARY MATERIAL An engineered tryptophan zipper-type peptide as a molecular recognition scaffold Zihao Cheng and Robert E. Campbell* Supplementary Methods Library construction for FRET-based screening
SUPPLEMENTARY MATERIAL An engineered tryptophan zipper-type peptide as a molecular recognition scaffold Zihao Cheng and Robert E. Campbell* Supplementary Methods Library construction for FRET-based screening
Project 07/111 Final Report October 31, Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines
 Project 07/111 Final Report October 31, 2007. Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines Project Leader: Dr Douglas C. Hodgins (519-824-4120 Ex 54758, fax 519-824-5930)
Project 07/111 Final Report October 31, 2007. Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines Project Leader: Dr Douglas C. Hodgins (519-824-4120 Ex 54758, fax 519-824-5930)
OXA-type beta-lactamases among extended-spectrum cephalosporin-resistant Pseudomonas aeruginosa isolates in a university hospital in southern Taiwan
 OXA-type J Microbiol ESBLs Immunol in P. Infect aeruginosa 2006;39:130-134 OXA-type beta-lactamases among extended-spectrum cephalosporin-resistant Pseudomonas aeruginosa isolates in a university hospital
OXA-type J Microbiol ESBLs Immunol in P. Infect aeruginosa 2006;39:130-134 OXA-type beta-lactamases among extended-spectrum cephalosporin-resistant Pseudomonas aeruginosa isolates in a university hospital
G+C content. 1 Introduction. 2 Chromosomes Topology & Counts. 3 Genome size. 4 Replichores and gene orientation. 5 Chirochores.
 1 Introduction 2 Chromosomes Topology & Counts 3 Genome size 4 Replichores and gene orientation 5 Chirochores 6 7 Codon usage 121 marc.bailly-bechet@univ-lyon1.fr Bacterial genome structures Introduction
1 Introduction 2 Chromosomes Topology & Counts 3 Genome size 4 Replichores and gene orientation 5 Chirochores 6 7 Codon usage 121 marc.bailly-bechet@univ-lyon1.fr Bacterial genome structures Introduction
for Programmed Chemo-enzymatic Synthesis of Antigenic Oligosaccharides
 Supporting Information Design of α-transglucosidases of Controlled Specificity for Programmed Chemo-enzymatic Synthesis of Antigenic Oligosaccharides Elise Champion ±,,,, Isabelle André ±,,, Claire Moulis
Supporting Information Design of α-transglucosidases of Controlled Specificity for Programmed Chemo-enzymatic Synthesis of Antigenic Oligosaccharides Elise Champion ±,,,, Isabelle André ±,,, Claire Moulis
Dierks Supplementary Fig. S1
 Dierks Supplementary Fig. S1 ITK SYK PH TH K42R wt K42R (kinase deficient) R29C E42K Y323F R29C E42K Y323F (reduced phospholipid binding) (enhanced phospholipid binding) (reduced Cbl binding) E42K Y323F
Dierks Supplementary Fig. S1 ITK SYK PH TH K42R wt K42R (kinase deficient) R29C E42K Y323F R29C E42K Y323F (reduced phospholipid binding) (enhanced phospholipid binding) (reduced Cbl binding) E42K Y323F
Supplemental Data. Bennett et al. (2010). Plant Cell /tpc
 BRN1 ---------MSSSNGGVPPGFRFHPTDEELLHYYLKKKISYEKFEMEVIKEVDLNKIEPWDLQDRCKIGSTPQNEWYFFSHKDRKYPTGS 81 BRN2 --------MGSSSNGGVPPGFRFHPTDEELLHYYLKKKISYQKFEMEVIREVDLNKLEPWDLQERCKIGSTPQNEWYFFSHKDRKYPTGS 82 SMB
BRN1 ---------MSSSNGGVPPGFRFHPTDEELLHYYLKKKISYEKFEMEVIKEVDLNKIEPWDLQDRCKIGSTPQNEWYFFSHKDRKYPTGS 81 BRN2 --------MGSSSNGGVPPGFRFHPTDEELLHYYLKKKISYQKFEMEVIREVDLNKLEPWDLQERCKIGSTPQNEWYFFSHKDRKYPTGS 82 SMB
Detection and molecular characterization of extended spectrum of beta lactamase (ESBL) producing Escherichia coli
 ISSN: 2319-7706 Volume 2 Number 8 (2013) pp. 196-205 http://www.ijcmas.com Original Research Article Detection and molecular characterization of extended spectrum of beta lactamase (ESBL) producing Escherichia
ISSN: 2319-7706 Volume 2 Number 8 (2013) pp. 196-205 http://www.ijcmas.com Original Research Article Detection and molecular characterization of extended spectrum of beta lactamase (ESBL) producing Escherichia
ACCEPTED. Department of Microbiology and Clinical Microbiology, Cerrahpasa Faculty of Medicine, University
 JCM Accepts, published online ahead of print on January 00 J. Clin. Microbiol. doi:./jcm.01-0 Copyright 00, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved.
JCM Accepts, published online ahead of print on January 00 J. Clin. Microbiol. doi:./jcm.01-0 Copyright 00, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved.
Supplemental Information. Human Senataxin Resolves RNA/DNA Hybrids. Formed at Transcriptional Pause Sites. to Promote Xrn2-Dependent Termination
 Supplemental Information Molecular Cell, Volume 42 Human Senataxin Resolves RNA/DNA Hybrids Formed at Transcriptional Pause Sites to Promote Xrn2-Dependent Termination Konstantina Skourti-Stathaki, Nicholas
Supplemental Information Molecular Cell, Volume 42 Human Senataxin Resolves RNA/DNA Hybrids Formed at Transcriptional Pause Sites to Promote Xrn2-Dependent Termination Konstantina Skourti-Stathaki, Nicholas
Supplementary Figures
 Supplementary Figures Supplementary Fig. 1 Characterization of GSCs. a. Immunostaining of primary GSC spheres from GSC lines. Nestin (neural progenitor marker, red), TLX (green). Merged images of nestin,
Supplementary Figures Supplementary Fig. 1 Characterization of GSCs. a. Immunostaining of primary GSC spheres from GSC lines. Nestin (neural progenitor marker, red), TLX (green). Merged images of nestin,
MCB421 FALL2005 EXAM#3 ANSWERS Page 1 of 12. ANSWER: Both transposon types form small duplications of adjacent host DNA sequences.
 Page 1 of 12 (10pts) 1. There are two mechanisms for transposition used by bacterial transposable elements: replicative (Tn3) and non-replicative (Tn5 and Tn10). Compare and contrast the two mechanisms
Page 1 of 12 (10pts) 1. There are two mechanisms for transposition used by bacterial transposable elements: replicative (Tn3) and non-replicative (Tn5 and Tn10). Compare and contrast the two mechanisms
RPA-AB RPA-C Supplemental Figure S1: SDS-PAGE stained with Coomassie Blue after protein purification.
 RPA-AB RPA-C (a) (b) (c) (d) (e) (f) Supplemental Figure S: SDS-PAGE stained with Coomassie Blue after protein purification. (a) RPA; (b) RPA-AB; (c) RPA-CDE; (d) RPA-CDE core; (e) RPA-DE; and (f) RPA-C
RPA-AB RPA-C (a) (b) (c) (d) (e) (f) Supplemental Figure S: SDS-PAGE stained with Coomassie Blue after protein purification. (a) RPA; (b) RPA-AB; (c) RPA-CDE; (d) RPA-CDE core; (e) RPA-DE; and (f) RPA-C
Characterization of Clinical Isolates of Enterobacteriaceae from Italy by the BD Phoenix Extended-Spectrum -Lactamase Detection Method
 JOURNAL OF CLINICAL MICROBIOLOGY, Apr. 2003, p. 1463 1468 Vol. 41, No. 4 0095-1137/03/$08.00 0 DOI: 10.1128/JCM.41.4.1463 1468.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved.
JOURNAL OF CLINICAL MICROBIOLOGY, Apr. 2003, p. 1463 1468 Vol. 41, No. 4 0095-1137/03/$08.00 0 DOI: 10.1128/JCM.41.4.1463 1468.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved.
NAME:... MODEL ANSWER... STUDENT NUMBER:... Maximum marks: 50. Internal Examiner: Hugh Murrell, Computer Science, UKZN
 COMP710, Bioinformatics with Julia, Test One, Thursday the 20 th of April, 2017, 09h30-11h30 1 NAME:...... MODEL ANSWER... STUDENT NUMBER:...... Maximum marks: 50 Internal Examiner: Hugh Murrell, Computer
COMP710, Bioinformatics with Julia, Test One, Thursday the 20 th of April, 2017, 09h30-11h30 1 NAME:...... MODEL ANSWER... STUDENT NUMBER:...... Maximum marks: 50 Internal Examiner: Hugh Murrell, Computer
The biomérieux solution. VITEK2 : A challenge with ESBL ESBL. Karen Bush
 International Newsletter n 4 December 2003 Through the IDENTIFYING RESISTANCE Newsletter, biomérieux s ambition is to contribute to the awareness and progress in the field of resistance to antibiotics.
International Newsletter n 4 December 2003 Through the IDENTIFYING RESISTANCE Newsletter, biomérieux s ambition is to contribute to the awareness and progress in the field of resistance to antibiotics.
SUPPLEMENTAL TABLE S1. Additional descriptions of plasmid constructions and the oligonucleotides used Plasmid or Oligonucleotide
 SUPPLEMENTAL TABLE S1. Additional descriptions of plasmid constructions and the oligonucleotides used Plasmid or Oligonucleotide former/ working Description a designation Plasmids pes213a b pes213-tn5
SUPPLEMENTAL TABLE S1. Additional descriptions of plasmid constructions and the oligonucleotides used Plasmid or Oligonucleotide former/ working Description a designation Plasmids pes213a b pes213-tn5
Emergence and persistence of integron structures harbouring VIM genes in the Children s Memorial Health Institute, Warsaw, Poland,
 Journal of Antimicrobial Chemotherapy (2009) 63, 269 273 doi:10.1093/jac/dkn512 Advance Access publication 18 December 2008 Emergence and persistence of integron structures harbouring VIM genes in the
Journal of Antimicrobial Chemotherapy (2009) 63, 269 273 doi:10.1093/jac/dkn512 Advance Access publication 18 December 2008 Emergence and persistence of integron structures harbouring VIM genes in the
Occurrence of viruses in apple and pear orchards in Latvia.
 INTERNATIONAL SCIENTIFIC CONFERENCE Sustainable Fruit Growing: From Plant to Product Occurrence of viruses in apple and pear orchards in Latvia. Neda P pola, Anna K le, Inga Moro ko Bi evska. The economically
INTERNATIONAL SCIENTIFIC CONFERENCE Sustainable Fruit Growing: From Plant to Product Occurrence of viruses in apple and pear orchards in Latvia. Neda P pola, Anna K le, Inga Moro ko Bi evska. The economically
S4B fluorescence (AU)
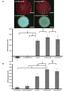 A S4B fluorescence (AU) S4B fluorescence (AU) dsbb csgba csgd dsbb csgba bcsa 5000 * NS NS 4000 * 3000 2000 1000 0 ΔcsgBAΔbcsA ΔcsgDΔdsbBΔbcsA ΔcsgBA ΔdsbBΔcsgBA ΔcsgDΔdsbB B -1000 4000 * * NS 3500 * 3000
A S4B fluorescence (AU) S4B fluorescence (AU) dsbb csgba csgd dsbb csgba bcsa 5000 * NS NS 4000 * 3000 2000 1000 0 ΔcsgBAΔbcsA ΔcsgDΔdsbBΔbcsA ΔcsgBA ΔdsbBΔcsgBA ΔcsgDΔdsbB B -1000 4000 * * NS 3500 * 3000
Distribution of Extended-Spectrum -Lactamases in Clinical Isolates of Enterobacteriaceae in Vietnam
 ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Dec. 2002, p. 3739 3743 Vol. 46, No. 12 0066-4804/02/$04.00 0 DOI: 10.1128/AAC.46.12.3739 3743.2002 Copyright 2002, American Society for Microbiology. All Rights
ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Dec. 2002, p. 3739 3743 Vol. 46, No. 12 0066-4804/02/$04.00 0 DOI: 10.1128/AAC.46.12.3739 3743.2002 Copyright 2002, American Society for Microbiology. All Rights
Lecture 11: Gene Prediction
 Lecture 11: Gene Prediction Study Chapter 6.11-6.14 1 Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and end positions of genes in a genome Where are
Lecture 11: Gene Prediction Study Chapter 6.11-6.14 1 Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and end positions of genes in a genome Where are
Antimicrobial Agents and Chemotherapy New Data Letter
 AAC Accepted Manuscript Posted Online 15 August 2016 Antimicrob. Agents Chemother. doi:10.1128/aac.01519-16 Copyright 2016, American Society for Microbiology. All Rights Reserved. 1 Antimicrobial Agents
AAC Accepted Manuscript Posted Online 15 August 2016 Antimicrob. Agents Chemother. doi:10.1128/aac.01519-16 Copyright 2016, American Society for Microbiology. All Rights Reserved. 1 Antimicrobial Agents
Keywords: qepa, qnrs1, rmtb, bla LAP-1, ISCR3C, gene co-transferance. Introduction
 Available online at www.annclinlabsci.org Annals of Clinical & Laboratory Science, vol. 39, no. 1, 2009 55 Accumulation of Plasmid-Mediated Fluoroquinolone Resistance Genes, qepa and qnrs1, in Enterobacter
Available online at www.annclinlabsci.org Annals of Clinical & Laboratory Science, vol. 39, no. 1, 2009 55 Accumulation of Plasmid-Mediated Fluoroquinolone Resistance Genes, qepa and qnrs1, in Enterobacter
PCR-based Markers and Cut Flower Longevity in Carnation
 PCRbased Markers and Cut Flower Longevity in Carnation Laura De Benedetti, Luca Braglia, Simona Bruna, Gianluca Burchi *, Antonio Mercuri and Tito Schiva Istituto Sperimentale per la Floricoltura, Corso
PCRbased Markers and Cut Flower Longevity in Carnation Laura De Benedetti, Luca Braglia, Simona Bruna, Gianluca Burchi *, Antonio Mercuri and Tito Schiva Istituto Sperimentale per la Floricoltura, Corso
11th Meeting of the Science Working Group. Lima, Peru, October 2012 SWG-11-JM-11
 11th Meeting of the Science Working Group Lima, Peru, 15-19 October 2012 Russian population genetics studies of jack mackerel in the South Pacific P.K.Afanasiev M.A.Rabchun A.I.Glubokov Introduction. In
11th Meeting of the Science Working Group Lima, Peru, 15-19 October 2012 Russian population genetics studies of jack mackerel in the South Pacific P.K.Afanasiev M.A.Rabchun A.I.Glubokov Introduction. In
Supporting Online Information
 Supporting Online Information Isolation of Human Genomic DNA Sequences with Expanded Nucleobase Selectivity Preeti Rathi, Sara Maurer, Grzegorz Kubik and Daniel Summerer* Department of Chemistry and Chemical
Supporting Online Information Isolation of Human Genomic DNA Sequences with Expanded Nucleobase Selectivity Preeti Rathi, Sara Maurer, Grzegorz Kubik and Daniel Summerer* Department of Chemistry and Chemical
Search for and Analysis of Single Nucleotide Polymorphisms (SNPs) in Rice (Oryza sativa, Oryza rufipogon) and Establishment of SNP Markers
 DNA Research 9, 163 171 (2002) Search for and Analysis of Single Nucleotide Polymorphisms (SNPs) in Rice (Oryza sativa, Oryza rufipogon) and Establishment of SNP Markers Shinobu Nasu, Junko Suzuki, Rieko
DNA Research 9, 163 171 (2002) Search for and Analysis of Single Nucleotide Polymorphisms (SNPs) in Rice (Oryza sativa, Oryza rufipogon) and Establishment of SNP Markers Shinobu Nasu, Junko Suzuki, Rieko
Supplemental Table 1. Primers used for PCR.
 Supplemental Table 1. Primers used for PCR. Gene Type Primer Sequence Genotyping and semi-quantitative RT-PCR F 5 -TTG CCC GAT CAC CAT CTG TA-3 rwa1-1 R 5 -TGT AGC GAT CAA GGC CTG ATC TAA-3 LB 5 -TAG CAT
Supplemental Table 1. Primers used for PCR. Gene Type Primer Sequence Genotyping and semi-quantitative RT-PCR F 5 -TTG CCC GAT CAC CAT CTG TA-3 rwa1-1 R 5 -TGT AGC GAT CAA GGC CTG ATC TAA-3 LB 5 -TAG CAT
Investigation of Klebsiella pneumoniae Isolates Producing SHV-12 and SHV-11 β-lactamases in Korean Hospitals
 J. Microbiol. Biotechnol. (2009), 19(2), 000 000 doi: 10.4014/jmb.0808.472 First published online 3 June 2009 Investigation of Klebsiella pneumoniae Isolates Producing SHV-12 and SHV-11 β-lactamases in
J. Microbiol. Biotechnol. (2009), 19(2), 000 000 doi: 10.4014/jmb.0808.472 First published online 3 June 2009 Investigation of Klebsiella pneumoniae Isolates Producing SHV-12 and SHV-11 β-lactamases in
Supporting Information
 Supporting Information Barderas et al. 10.1073/pnas.0801221105 SI Text: Docking of gastrin to Constructed scfv Models Interactive predocking of the 4-WL-5 motif into the central pocket observed in the
Supporting Information Barderas et al. 10.1073/pnas.0801221105 SI Text: Docking of gastrin to Constructed scfv Models Interactive predocking of the 4-WL-5 motif into the central pocket observed in the
SUPPORTING INFORMATION
 SUPPORTING INFORMATION Investigation of the Biosynthesis of the Lasso Peptide Chaxapeptin Using an E. coli-based Production System Helena Martin-Gómez, Uwe Linne, Fernando Albericio, Judit Tulla-Puche,*
SUPPORTING INFORMATION Investigation of the Biosynthesis of the Lasso Peptide Chaxapeptin Using an E. coli-based Production System Helena Martin-Gómez, Uwe Linne, Fernando Albericio, Judit Tulla-Puche,*
Multiplex PCR Study of Plasmid-Mediated AmpC Beta-Lactamase Genes in Clinical Isolates of Escherichia coli
 J Med Bacteriol. Vol. 5, No. 5, 6 (216): pp.21-28 jmb.tums.ac.ir Multiplex PCR Study of Plasmid-Mediated AmpC Beta-Lactamase Genes in Clinical Isolates of Escherichia coli Maryam Dehghani 1, Azam Haddadi
J Med Bacteriol. Vol. 5, No. 5, 6 (216): pp.21-28 jmb.tums.ac.ir Multiplex PCR Study of Plasmid-Mediated AmpC Beta-Lactamase Genes in Clinical Isolates of Escherichia coli Maryam Dehghani 1, Azam Haddadi
2
 1 2 3 4 5 6 7 Supplemental Table 1. Magnaporthe oryzae strains generated in this study. Strain background Genotype Strain name Description Guy-11 H1:RFP H1:RFP Strain expressing Histone H1- encoding gene
1 2 3 4 5 6 7 Supplemental Table 1. Magnaporthe oryzae strains generated in this study. Strain background Genotype Strain name Description Guy-11 H1:RFP H1:RFP Strain expressing Histone H1- encoding gene
Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of
 Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of the cell line) were immunostained for HA, acetylated
Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of the cell line) were immunostained for HA, acetylated
Primer Design Workshop. École d'été en géné-que des champignons 2012 Dr. Will Hintz University of Victoria
 Primer Design Workshop École d'été en géné-que des champignons 2012 Dr. Will Hintz University of Victoria Scenario You have discovered the presence of a novel endophy5c organism living inside the cells
Primer Design Workshop École d'été en géné-que des champignons 2012 Dr. Will Hintz University of Victoria Scenario You have discovered the presence of a novel endophy5c organism living inside the cells
Evaluation of a Double Synergy Differential Test (DSDT) for differential detection of ESBL and AmpC-type
 NEW MICROBIOLOGICA, 35, 221-225, 2012 Evaluation of a Double Synergy Differential Test (DSDT) for differential detection of ESBL and AmpC-type β-lactamases in Escherichia coli, Klebsiella pneumoniae and
NEW MICROBIOLOGICA, 35, 221-225, 2012 Evaluation of a Double Synergy Differential Test (DSDT) for differential detection of ESBL and AmpC-type β-lactamases in Escherichia coli, Klebsiella pneumoniae and
Extended Spectrum β-lactamases: Critical Tools of Bacterial Resistance
 Review Article Mahidol University Journal of Pharmaceutical Science 2012; 39 (1), 1-8 Extended Spectrum β-lactamases: Critical Tools of Bacterial Resistance Department of Microbiology, Faculty of Pharmacy,
Review Article Mahidol University Journal of Pharmaceutical Science 2012; 39 (1), 1-8 Extended Spectrum β-lactamases: Critical Tools of Bacterial Resistance Department of Microbiology, Faculty of Pharmacy,
hcd1tg/hj1tg/ ApoE-/- hcd1tg/hj1tg/ ApoE+/+
 ApoE+/+ ApoE-/- ApoE-/- H&E (1x) Supplementary Figure 1. No obvious pathology is observed in the colon of diseased ApoE-/me. Colon samples were fixed in 1% formalin and laid out in Swiss rolls for paraffin
ApoE+/+ ApoE-/- ApoE-/- H&E (1x) Supplementary Figure 1. No obvious pathology is observed in the colon of diseased ApoE-/me. Colon samples were fixed in 1% formalin and laid out in Swiss rolls for paraffin
evaluated with UAS CLB eliciting UAS CIT -N Libraries increase in the
 Supplementary Figures Supplementary Figure 1: Promoter scaffold library assemblies. Many ensembless of libraries were evaluated in this work. As a legend, the box outline color in top half of the figure
Supplementary Figures Supplementary Figure 1: Promoter scaffold library assemblies. Many ensembless of libraries were evaluated in this work. As a legend, the box outline color in top half of the figure
H. Wu, B.-G. Liu, J.-H. Liu, Y.-S. Pan, L. Yuan and G.-Z. Hu
 Phenotypic and molecular characterization of CTX-M-14 extended-spectrum β-lactamase and plasmid-mediated ACT-like AmpC β-lactamase produced by Klebsiella pneumoniae isolates from chickens in Henan Province,
Phenotypic and molecular characterization of CTX-M-14 extended-spectrum β-lactamase and plasmid-mediated ACT-like AmpC β-lactamase produced by Klebsiella pneumoniae isolates from chickens in Henan Province,
ORIGINAL ARTICLE /j x
 ORIGINAL ARTICLE 10.1111/j.1469-0691.2008.02030.x Metallo-b-lactamase-producing Pseudomonas aeruginosa isolated from a large tertiary centre in Kenya J. D. D. Pitout 1,2,3, G. Revathi 4, B. L Chow 1, B.
ORIGINAL ARTICLE 10.1111/j.1469-0691.2008.02030.x Metallo-b-lactamase-producing Pseudomonas aeruginosa isolated from a large tertiary centre in Kenya J. D. D. Pitout 1,2,3, G. Revathi 4, B. L Chow 1, B.
Anti-Pim-1 (Cat#3247), anti-met (Cat#3127), anti-ron (Cat#2654), Anti-EGFR
 Supplementary Methods Antibodies Anti-Pim-1 (Cat#3247), anti-met (Cat#3127), anti-ron (Cat#2654), Anti-EGFR (Cat#2646), anti-igf1r (Cat#3018), anti-insr (Cat#3020), anti-akt (pan, Cat#4691), anti-phospho-akt
Supplementary Methods Antibodies Anti-Pim-1 (Cat#3247), anti-met (Cat#3127), anti-ron (Cat#2654), Anti-EGFR (Cat#2646), anti-igf1r (Cat#3018), anti-insr (Cat#3020), anti-akt (pan, Cat#4691), anti-phospho-akt
Received 23 December 2010/Returned for modification 11 January 2011/Accepted 7 February 2011
 JOURNAL OF CLINICAL MICROBIOLOGY, Apr. 2011, p. 1608 1613 Vol. 49, No. 4 0095-1137/11/$12.00 doi:10.1128/jcm.02607-10 Copyright 2011, American Society for Microbiology. All Rights Reserved. Evaluation
JOURNAL OF CLINICAL MICROBIOLOGY, Apr. 2011, p. 1608 1613 Vol. 49, No. 4 0095-1137/11/$12.00 doi:10.1128/jcm.02607-10 Copyright 2011, American Society for Microbiology. All Rights Reserved. Evaluation
Received 24 June 2008/Returned for modification 12 August 2008/Accepted 14 September 2008
 ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Dec. 2008, p. 4268 4273 Vol. 52, No. 12 0066-4804/08/$08.00 0 doi:10.1128/aac.00830-08 Copyright 2008, American Society for Microbiology. All Rights Reserved. High
ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Dec. 2008, p. 4268 4273 Vol. 52, No. 12 0066-4804/08/$08.00 0 doi:10.1128/aac.00830-08 Copyright 2008, American Society for Microbiology. All Rights Reserved. High
REVIEW. Genetic support of extended-spectrum b-lactamases L. Poirel, T. Naas and P. Nordmann
 REVIEW Genetic support of extended-spectrum b-lactamases L. Poirel, T. Naas and P. Nordmann Service de Bactériologie-Virologie, Hôpital de Bicêtre, South-Paris Medical School, University Paris XI, Le Kremlin-Bicêtre,
REVIEW Genetic support of extended-spectrum b-lactamases L. Poirel, T. Naas and P. Nordmann Service de Bactériologie-Virologie, Hôpital de Bicêtre, South-Paris Medical School, University Paris XI, Le Kremlin-Bicêtre,
Ongoing epidemic of bla VIM-1 -positive Klebsiella pneumoniae in Athens, Greece: a prospective survey
 Journal of Antimicrobial Chemotherapy (2008) 61, 59 63 doi:10.1093/jac/dkm443 Advance Access publication 13 November 2007 Ongoing epidemic of bla VIM-1 -positive Klebsiella pneumoniae in Athens, Greece:
Journal of Antimicrobial Chemotherapy (2008) 61, 59 63 doi:10.1093/jac/dkm443 Advance Access publication 13 November 2007 Ongoing epidemic of bla VIM-1 -positive Klebsiella pneumoniae in Athens, Greece:
Key words: ESBLs; Ceftriaxone; Escherichia coli; MICs; Bacteremia
 JCM Accepts, published online ahead of print on 16 April 2014 J. Clin. Microbiol. doi:10.1128/jcm.00716-14 Copyright 2014, American Society for Microbiology. All Rights Reserved. 1 1 2 Title: Determining
JCM Accepts, published online ahead of print on 16 April 2014 J. Clin. Microbiol. doi:10.1128/jcm.00716-14 Copyright 2014, American Society for Microbiology. All Rights Reserved. 1 1 2 Title: Determining
Occurrence of TEM & SHV gene in extended spectrum b-lactamases (ESBLs) producing Klebsiella sp. isolated from a tertiary care hospital
 Indian J Med Res 125, February 2007, pp 173-178 Occurrence of TEM & SHV gene in extended spectrum b-lactamases (ESBLs) producing Klebsiella sp. isolated from a tertiary care hospital Prabha Lal, Arti Kapil,
Indian J Med Res 125, February 2007, pp 173-178 Occurrence of TEM & SHV gene in extended spectrum b-lactamases (ESBLs) producing Klebsiella sp. isolated from a tertiary care hospital Prabha Lal, Arti Kapil,
