Debugging the Bug. Lessons learned from developing integrating EcoCyc and ijr904. Jeremy Zucker Dana-Farber Cancer Institute Harvard Medical School
|
|
|
- Magnus Henry
- 6 years ago
- Views:
Transcription
1 Debugging the Bug Lessons learned from developing integrating EcoCyc and ijr904 Jeremy Zucker Dana-Farber Cancer Institute Harvard Medical School
2 Automated Metabolic Flux Analysis Pipeline Genome Pathway Tools Nutrients & Objective Transporter prediction Pathway/ Genome Database PGDB query Model Definition (SBML) FBA & MOMA Network Debugging Flux prediction
3 The team The system The questions The approach The lessons learned The results The next steps The project
4 The Team SRI: Markus and Ingrid Palsson lab: Adam Feist and Jennifer Reed Northwestern: Chris Henry York University: Gavin Thomas BioPAX: Alan Ruttenberg and Jeremy Zucker
5 The System: E. coli K12 MG1655 Model Year(s) No. of metabolic reactions No. of metabolites Majewski and Domach Varma and Palsson Pramanik and (317) 289 (305) Keasling 1998 Edwards and Palsson Covert and Palssonc Reed and Palsson
6 The System: E. coli K12 MG1655 KB: EcoCyc 9.5 Goal: Representation of metabolic knowledge Maturity: 13+ years Size: 4480 genes 1075 enzyme rxns 977 compounds Model: ijr904 Goal: Constraintbased metabolic flux model Maturity: 13+ years Size: 904 genes 929 enzyme rxns 626 compounds
7 The Goal Map ijr904 compounds to EcoCyc compounds Map ijr904 reactions to EcoCyc reactions Get a working Flux balance model for EcoCyc Create FBA enabled BioCyc PGDB's Create a disciplined approach to data integration.
8 The approach Automated mapping of compounds Semi-automated mapping of compounds Manual mapping of compounds Automated mapping of reactions Semi-automated mapping of reactions Manual mapping of reactions
9 Lessons Learned Standard is better than best A good representation is the key to good problem solving --Patrick Winston 1+1 is equal but not identical to 2 When it comes to data cleaning, there is no such thing as a free lunch --Sir Tim Berners-Lee You don't have to do it all by yourself Matt Temple
10 Standard is better than best Utilize W3C Web Ontology Language (OWL) Utilize open source description logic reasoners (Pellet, FaCT++) Simplify DL syntax with lisp macros. Utilize BioPAX/Pathway tools schema where possible, extend when not possible.
11 What is OWL? 3 flavors: OWL-Lite, OWL-DL, OWL-Full Description Logic. Very similar to Framebased systems Description logic reasoners are sound and complete A standard in flux (OWL 1.1 coming out soon)
12 A good representation is the key to good problem solving -- Patrick Winston CAS and KEGG ID's are supposed to be unique identifiers, so make the unificationxref property inverse functional Compounds are classes Reactions can be defined in terms of compounds
13 1+1is equal but not identical to 2 When are two reactions equal? Same EC number? When does one reaction subsume another? Same enzymes catalyze? When are two genes equal? dut vs dfp When are two compounds equal? Same CAS number? Same KEGG ID? structural isomers: Glucose vs. Fructose enantiomers: D-Glucose vs L-Glucose Beta-D-glucose vs Alpha-D-glucose
14 When it comes to data integration, there's no such thing as a free lunch Bugs in --Sir CAS Tim and KEGG Berners-Lee ID Xrefs Merged reactions hisd Reactions in comments Multiple functions of the same gene Missing compound mappings
15 How do I use a DL-reasoner? Pellet: Java, accessible from ABCL: (define-ontology simple-classhierarchy () (class!a :partial) (class!b :partial) (class!c :partial) (sub-class-of!b!a) (sub-class-of!c!b) ) (describe-entity!c (kb simpleclass-hierarchy)) Direct Superclasses: ex:b All Superclasses: ex:b, ex:a Axioms: FaCT++: C++, with lisp syntax interface (equal_c trans-rxn-157 (and (atmost 2 left) (atleast 1 left phospho-enol-pyruvate ) (atleast 1 left glc ) (atmost 2 right) (atleast 1 right glc-6-p ) (atleast 1 right pyruvate )))
16 Reasoning with FaCT++ ecocyc:ribonucleoside-dip-reducti-rxn (and (atmost 3 left) (atleast 1 left ecocyc:ox-thioredoxin ) (atleast 1 left ecocyc:water ) (atleast 1 left ecocyc:deoxy-ribonucleoside-diphosphates ) (atmost 2 right) (atleast 1 right ecocyc:red-thioredoxin ) (atleast 1 right ecocyc:ribonucleoside-diphosphates ))) (equal_c ijr904:rndr3 (and (atleast 1 right ijr904:cdp ) (atleast 1 right ijr904:trdrd ) (atmost 2 right) (atleast 1 left ijr904:dcdp ) (atleast 1 left ijr904:h2o ) (atleast 1 left ijr904:trdox ) (atmost 3 left))) (equal_c ijr904:trdrd ecocyc:red-thioredoxin ) (equal_c ijr904:trdox ecocyc:ox-thioredoxin ) (equal_c ijr904:dcdp ecocyc:dcdp ) (equal_c ijr904:h2o ecocyc:water ) (implies ecocyc:dcdp ecocyc:deoxy-ribonucleoside-diphosphates ) (implies ecocyc:cdp ecocyc:ribonucleoside-diphosphates ) => (:SUB "ijr904:rndr3" :SUPER ("ecocyc:ribonucleoside-dip-reducti-rxn")))
17 The results (so far) 589 out of 773 ijr904 compounds manually mapped to EcoCyc 41 errors in EcoCyc compound crossreferences 80 errors in ijr904 compound crossreferences 637 out of 937 ijr904 reactions automatically mapped to EcoCyc. 83 reaction subsumptions
18 Bugs in Network structure revealed by Forward and Backward chaining Unproduced metabolite Precursor metabolite Missing essential compound Biomas Essential compounds
19 Nutrient analysis: Forward-chaining Fired Reaction Known Nutrient set Unfired Reaction Missing essential compound Biomas Essential compounds
20 Metabolic flux analysis assumptions From annotated genomes to metabolic flux models and kinetic parameter fitting Daniel Segrè et al., Omics, 7: (2003).
21 First, analyze the steady state-behavior v 5 v 5 v 1 A v 2 v 3 B 2B C E v 7 v 4 2D v 6 v 8 D F R 9 v 1 A v 2 v 3 B 2B C E v 7 v 4 2D v 6 v 8 D F v 9 v 5 All metabolic capabilities in steady states are composed of elementary flux modes, which are the minimal sets of enzymes that can each generate valid steady states. --Trends Biotechnol 1999 Feb;17(2):53-60 v 2 B E v 2D 1 A v 6 v D 8 v 3 2B C v 7 v 4 F v 9
22 Wild-type FBA Prediction 10 R 5 B E A R B R 4 2D R 1 10 R R D 20 R 3 C R 7 F R 9
23 Mutant FBA Prediction R 5 B E A R 2 2B 2D R 1 10 R R 8 6 S T O P R 4 D 10 R 3 10 C 10 R 7 F R 9
24 Mutant MOMA prediction R 5 B E A R B R 4 2D R R 8 R 6 S T O P D 6.7 R C 6.7 R 7 F 3.3 R 9
25 Classifying the bugs bugs in the data (syntax) Mass balance: Thermodynamics (Gibbs) Missing chemical formulas Missing/incorrect gene (Errors in Genome annotation) Missing/incorrect reactions (Orphaned enzymes) Missing pathways (pathway hole filling) bugs in the data representation (semantics) Identity problem: Palsson integration, Synonyms, fuzzy names insufficient expressivity: Polymerization, generalized reactions, Transport protein database, SBML Provenance (multiple source problem) Bugs in the model assumptions with respect to experiment Growth: FBA vs. MOMA vs. ROOM Internal fluxes: Isotope labelling Epistasis Minimal nutrient sets Expression, Protein levels, and Metabolite concentrations
26 Diagnose and debug the bug Fixing bugs in the data (syntax) Mass balance: balance-p diagnosis uses atomic balances for C S N and P Provenance used to fix the problem with stoichiometry Thermodynamics: include Gibbs free energy Diagnosis: Can be calculated empirically Missing gene: Diagnosis: Network--filling Orphaned enzymes Missing/incorrect gene: (Errors in Genome annotation) Diagnosis: Gene-enzyme-reaction predicate in conjuction with knockout experiments Missing/incorrect reactions (Orphaned enzymes) Diagnosis: Nutrient analysis, backward chaining Missing pathways Bugs in the data representation (semantics) Identity problem: Palsson integration, Synonyms, fuzzy names insufficient expressivity: Polymerization, generalized reactions, Transport protein database, SBML Provenance (multiple source problem) Bugs in the model assumptions with respect to experiment Growth: FBA vs. MOMA vs. ROOM Internal fluxes: Isotope labelling Epistasis Minimal nutrient sets Expression, Protein levels, and Metabolite concentrations
27 Debugging bugs bugs in the data (syntax) Mass balance: Thermodynamics (Gibbs) Missing chemical formulas Missing/incorrect gene (Errors in Genome annotation) Missing/incorrect reactions (Orphaned enzymes) Missing pathways (pathway hole filling) bugs in the data representation (semantics) Identity problem: Palsson integration, Synonyms, fuzzy names insufficient expressivity: Polymerization, generalized reactions, Transport protein database, SBML Provenance (multiple source problem) Orphaned enzymes Errors in genome annotation Enzyme genomics project
28
29 Bug class Diagnosis Fix
MetaFlux in Pathway Tools (Short Tutorial)
 MetaFlux in Pathway Tools (Short Tutorial) Mario Latendresse SRI International 22 September 2015 Outline 1 Introduction to Flux Balance Analysis What is Flux Balance Analysis (FBA)? Overview of MetaFlux
MetaFlux in Pathway Tools (Short Tutorial) Mario Latendresse SRI International 22 September 2015 Outline 1 Introduction to Flux Balance Analysis What is Flux Balance Analysis (FBA)? Overview of MetaFlux
Basic modeling approaches for biological systems -II. Mahesh Bule
 Basic modeling approaches for biological systems -II Mahesh Bule Modeling Metabolism Metabolism is general term for two kind of reactions Catabolic reactions (breakdown of complex compounds to get energy
Basic modeling approaches for biological systems -II Mahesh Bule Modeling Metabolism Metabolism is general term for two kind of reactions Catabolic reactions (breakdown of complex compounds to get energy
Multiple Gap-Filling of Flux-Balance Models
 Multiple Gap-Filling of Flux-Balance Models Mario Latendresse and Peter D. Karp Bioinformatics Research Group SRI International, Menlo Park, USA COBRA 2011, Reykjavik, Iceland, June 25 Outline 1 The Problem
Multiple Gap-Filling of Flux-Balance Models Mario Latendresse and Peter D. Karp Bioinformatics Research Group SRI International, Menlo Park, USA COBRA 2011, Reykjavik, Iceland, June 25 Outline 1 The Problem
NIH Public Access Author Manuscript Nat Biotechnol. Author manuscript; available in PMC 2011 June 6.
 NIH Public Access Author Manuscript Published in final edited form as: Nat Biotechnol. 2010 March ; 28(3): 245 248. doi:10.1038/nbt.1614. What is flux balance analysis? Jeffrey D. Orth, University of California
NIH Public Access Author Manuscript Published in final edited form as: Nat Biotechnol. 2010 March ; 28(3): 245 248. doi:10.1038/nbt.1614. What is flux balance analysis? Jeffrey D. Orth, University of California
Pathway Tools Schema and Semantic Inference Layer: Pathways and the Overview. SRI International Bioinformatics
 Pathway Tools Schema and Semantic Inference Layer: Pathways and the Overview 1 Outline Pathways Representation of Pathways Querying Pathways Programmatically How Pathway Diagrams are Generated Future Work:
Pathway Tools Schema and Semantic Inference Layer: Pathways and the Overview 1 Outline Pathways Representation of Pathways Querying Pathways Programmatically How Pathway Diagrams are Generated Future Work:
Using Pathway Tools & Matlab for Flux Balance Analysis. Kent Peterson 25 Aug. 2009
 Using Pathway Tools & Matlab for Flux Balance Analysis Kent Peterson 25 Aug. 2009 Summary Flux Balance Analysis (FBA) Workflow: from Ptools to Matlab, and back again A simple example What next? 2 Let s
Using Pathway Tools & Matlab for Flux Balance Analysis Kent Peterson 25 Aug. 2009 Summary Flux Balance Analysis (FBA) Workflow: from Ptools to Matlab, and back again A simple example What next? 2 Let s
Kyoto Encyclopedia of Genes and Genomes (KEGG)
 NPTEL Biotechnology -Systems Biology Kyoto Encyclopedia of Genes and Genomes (KEGG) Dr. M. Vijayalakshmi School of Chemical and Biotechnology SASTRA University Joint Initiative of IITs and IISc Funded
NPTEL Biotechnology -Systems Biology Kyoto Encyclopedia of Genes and Genomes (KEGG) Dr. M. Vijayalakshmi School of Chemical and Biotechnology SASTRA University Joint Initiative of IITs and IISc Funded
A Formal Ontology for the Cell-Cycle Domain
 A Formal Ontology for the Cell-Cycle Domain Erick ANTEZANA Dept. of Plant Systems Biology. Flanders Interuniversity Institute for Biotechnology/Ghent University. Ghent BELGIUM. erant@psb.ugent.be Overview
A Formal Ontology for the Cell-Cycle Domain Erick ANTEZANA Dept. of Plant Systems Biology. Flanders Interuniversity Institute for Biotechnology/Ghent University. Ghent BELGIUM. erant@psb.ugent.be Overview
Inference in Metabolic Network Models using Flux Balance Analysis
 Inference in Metabolic Network Models using Flux Balance Analysis BMI/CS 776 www.biostat.wisc.edu/bmi776/ Mark Craven craven@biostat.wisc.edu Spring 2011 Goals for Lecture the key concepts to understand
Inference in Metabolic Network Models using Flux Balance Analysis BMI/CS 776 www.biostat.wisc.edu/bmi776/ Mark Craven craven@biostat.wisc.edu Spring 2011 Goals for Lecture the key concepts to understand
Metabolism and metabolic networks. Metabolism is the means by which cells acquire energy and building blocks for cellular material
 Metabolism and metabolic networks Metabolism is the means by which cells acquire energy and building blocks for cellular material Metabolism is organized into sequences of biochemical reactions, metabolic
Metabolism and metabolic networks Metabolism is the means by which cells acquire energy and building blocks for cellular material Metabolism is organized into sequences of biochemical reactions, metabolic
Metabolic modeling (4cr)
 582605 Metabolic modeling (4cr) Lecturer: prof. Juho Rousu Course assistant: Markus Heinonen Lectures: Tuesdays and Fridays, 14.15-16, B119 Exercises: 16.03.-24.04. Tuesdays 16.15-18, C221 Course topics:
582605 Metabolic modeling (4cr) Lecturer: prof. Juho Rousu Course assistant: Markus Heinonen Lectures: Tuesdays and Fridays, 14.15-16, B119 Exercises: 16.03.-24.04. Tuesdays 16.15-18, C221 Course topics:
Constraint-Based Workshops
 Constraint-Based Workshops 10. Flux Coupling (again) & Metabolic Engineering February 28 th, 2008 Constraint-Based Methods Optimal Solutions 1. FBA 2. Flux Variability Flux Dependencies 1. Robustness 2.
Constraint-Based Workshops 10. Flux Coupling (again) & Metabolic Engineering February 28 th, 2008 Constraint-Based Methods Optimal Solutions 1. FBA 2. Flux Variability Flux Dependencies 1. Robustness 2.
From DNA Sequences to Genome-Scale Metabolic Models (to Network Projects in CellNetAnalyzer)
 From DNA Sequences to Genome-Scale Metabolic Models (to Network Projects in CellNetAnalyzer) Sven Thiele Max Planck Institute for Dynamics of Complex Technical Systems, Germany Introduction A genome-scale
From DNA Sequences to Genome-Scale Metabolic Models (to Network Projects in CellNetAnalyzer) Sven Thiele Max Planck Institute for Dynamics of Complex Technical Systems, Germany Introduction A genome-scale
Pathway analysis. Martina Kutmon Department of Bioinformatics Maastricht University
 Pathway analysis Martina Kutmon Department of Bioinformatics Maastricht University Who are we? Department of Bioinformatics @ Maastricht University Martina Kutmon PhD student, 3rd year Anwesha Bohler PhD
Pathway analysis Martina Kutmon Department of Bioinformatics Maastricht University Who are we? Department of Bioinformatics @ Maastricht University Martina Kutmon PhD student, 3rd year Anwesha Bohler PhD
All commands and actions are indicated as highlighted yellow steps CellNetAnalyzer and MatLab commands are in Courier Font
 Steady State Metabolic Modeling Homework In this lab we will use the application CellNetAnalyzer (CNA) to model metabolic pathways in E. coli and M. tuberculosis. This lab is a revised version from a lab
Steady State Metabolic Modeling Homework In this lab we will use the application CellNetAnalyzer (CNA) to model metabolic pathways in E. coli and M. tuberculosis. This lab is a revised version from a lab
FROM DISCOVERY TO INSIGHT
 Agilent Pathway Architect FROM DISCOVERY TO INSIGHT METABOLOMICS PROTEOMICS PATHWAY ARCHITECT GENOMICS TRANSCRIPTOMICS AGILENT PATHWAY ARCHITECT SPEED DISCOVERY TO UNDERSTANDING Today s scientists face
Agilent Pathway Architect FROM DISCOVERY TO INSIGHT METABOLOMICS PROTEOMICS PATHWAY ARCHITECT GENOMICS TRANSCRIPTOMICS AGILENT PATHWAY ARCHITECT SPEED DISCOVERY TO UNDERSTANDING Today s scientists face
Metabolic Networks. Ulf Leser and Michael Weidlich
 Metabolic Networks Ulf Leser and Michael Weidlich This Lecture Introduction Systems biology & modelling Metabolism & metabolic networks Network reconstruction Strategy & workflow Mathematical representation
Metabolic Networks Ulf Leser and Michael Weidlich This Lecture Introduction Systems biology & modelling Metabolism & metabolic networks Network reconstruction Strategy & workflow Mathematical representation
Description of WWW interface to: GSMN-TB: a web-based genome scale network model of Mycobacterium. tuberculosis metabolism
 Description of WWW interface to: GSMN-TB: a web-based genome scale network model of Mycobacterium tuberculosis metabolism Dany JV Beste 1 *, Tracy Hooper 1 *, Graham Stewart 1, Bhushan Bonde 1, Claudio
Description of WWW interface to: GSMN-TB: a web-based genome scale network model of Mycobacterium tuberculosis metabolism Dany JV Beste 1 *, Tracy Hooper 1 *, Graham Stewart 1, Bhushan Bonde 1, Claudio
Qualitative causal analyses of biosimulation models
 Qualitative causal analyses of biosimulation models Maxwell L. Neal 1, John H. Gennari 1, Daniel L. Cook 1,2 1 Department of Biomedical Informatics and Medical Education 2 Department of Physiology and
Qualitative causal analyses of biosimulation models Maxwell L. Neal 1, John H. Gennari 1, Daniel L. Cook 1,2 1 Department of Biomedical Informatics and Medical Education 2 Department of Physiology and
An Algorithm to Assemble Gene-Protein-Reaction Associations for Genome-Scale Metabolic Model Reconstruction
 An Algorithm to Assemble Gene-Protein-Reaction Associations for Genome-Scale Metabolic Model Reconstruction João Cardoso 1,2,PauloVilaça 1,2,Simão Soares 1, and Miguel Rocha 2 1 SilicoLife Lda., Spinpark,
An Algorithm to Assemble Gene-Protein-Reaction Associations for Genome-Scale Metabolic Model Reconstruction João Cardoso 1,2,PauloVilaça 1,2,Simão Soares 1, and Miguel Rocha 2 1 SilicoLife Lda., Spinpark,
An Algorithm to Assemble Gene-Protein-Reaction Associations for Genome-Scale Metabolic Model Reconstruction
 An Algorithm to Assemble Gene-Protein-Reaction Associations for Genome-Scale Metabolic Model Reconstruction João Cardoso 1,2, Paulo Vilaça 1,2, Simão Soares 1,andMiguelRocha 2 1 SilicoLife Lda., Spinpark,
An Algorithm to Assemble Gene-Protein-Reaction Associations for Genome-Scale Metabolic Model Reconstruction João Cardoso 1,2, Paulo Vilaça 1,2, Simão Soares 1,andMiguelRocha 2 1 SilicoLife Lda., Spinpark,
A study on the robustness of strain optimization algorithms
 A study on the robustness of strain optimization algorithms Paulo Vilaça, Paulo Maia, and Miguel Rocha Abstract In recent years, there have been considerable advances in the use of genome-scale metabolic
A study on the robustness of strain optimization algorithms Paulo Vilaça, Paulo Maia, and Miguel Rocha Abstract In recent years, there have been considerable advances in the use of genome-scale metabolic
Design Principles in Synthetic Biology
 Design Principles in Synthetic Biology Chris Myers 1, Nathan Barker 2, Hiroyuki Kuwahara 3, Curtis Madsen 1, Nam Nguyen 1, Michael Samoilov 4, and Adam Arkin 4 1 University of Utah 2 Southern Utah University
Design Principles in Synthetic Biology Chris Myers 1, Nathan Barker 2, Hiroyuki Kuwahara 3, Curtis Madsen 1, Nam Nguyen 1, Michael Samoilov 4, and Adam Arkin 4 1 University of Utah 2 Southern Utah University
Object Groups. SRI International Bioinformatics
 Object Groups 1 SRI International Bioinformatics Object Groups Collect and save lists of genes, metabolites, pathways Transform, filter, and analyze them Share groups with colleagues Use groups in conjunction
Object Groups 1 SRI International Bioinformatics Object Groups Collect and save lists of genes, metabolites, pathways Transform, filter, and analyze them Share groups with colleagues Use groups in conjunction
Deposited research article Using Topology of the Metabolic Network to Predict Viability of Mutant Strains Zeba Wunderlich and Leonid Mirny*
 This information has not been peer-reviewed. Responsibility for the findings rests solely with the author(s). Deposited research article Using Topology of the Metabolic Network to Predict Viability of
This information has not been peer-reviewed. Responsibility for the findings rests solely with the author(s). Deposited research article Using Topology of the Metabolic Network to Predict Viability of
DRAGON DATABASE OF GENES ASSOCIATED WITH PROSTATE CANCER (DDPC) Monique Maqungo
 DRAGON DATABASE OF GENES ASSOCIATED WITH PROSTATE CANCER (DDPC) Monique Maqungo South African National Bioinformatics Institute University of the Western Cape RELEVEANCE OF DATA SHARING! Fragmented data
DRAGON DATABASE OF GENES ASSOCIATED WITH PROSTATE CANCER (DDPC) Monique Maqungo South African National Bioinformatics Institute University of the Western Cape RELEVEANCE OF DATA SHARING! Fragmented data
OPTIMALITY CRITERIA FOR THE PREDICTION OF METABOLIC FLUXES IN YEAST MUTANTS
 123 OPTIMALITY CRITERIA FOR THE PREDICTION OF METABOLIC FLUXES IN YEAST MUTANTS EVAN S. SNITKIN 1 DANIEL SEGRÈ 1,2 esnitkin@bu.edu dsegre@bu.edu 1 Graduate Program in Bioinformatics, Boston University,
123 OPTIMALITY CRITERIA FOR THE PREDICTION OF METABOLIC FLUXES IN YEAST MUTANTS EVAN S. SNITKIN 1 DANIEL SEGRÈ 1,2 esnitkin@bu.edu dsegre@bu.edu 1 Graduate Program in Bioinformatics, Boston University,
An update on Genomic CDS, a complex ontology for pharmacogenomics and clinical decision support
 An update on Genomic CDS, a complex ontology for pharmacogenomics and clinical decision support José Antonio Minarro-Giménez 1, Matthias Samwald 1 1 Section for Medical Expert and Knowledge-Based Systems;
An update on Genomic CDS, a complex ontology for pharmacogenomics and clinical decision support José Antonio Minarro-Giménez 1, Matthias Samwald 1 1 Section for Medical Expert and Knowledge-Based Systems;
Statement of Tasks and Intent of Sponsor
 Industrialization of Biology: A Roadmap to Accelerate Advanced Manufacturing of Chemicals Statement of Tasks and Intent of Sponsor Friedrich Srienc Program Director Biotechnology, Biochemical, and Biomass
Industrialization of Biology: A Roadmap to Accelerate Advanced Manufacturing of Chemicals Statement of Tasks and Intent of Sponsor Friedrich Srienc Program Director Biotechnology, Biochemical, and Biomass
Agilent Software Tools for Mass Spectrometry Based Multi-omics Studies
 Agilent Software Tools for Mass Spectrometry Based Multi-omics Studies Technical Overview Introduction The central dogma for biological information flow is expressed as a series of chemical conversions
Agilent Software Tools for Mass Spectrometry Based Multi-omics Studies Technical Overview Introduction The central dogma for biological information flow is expressed as a series of chemical conversions
Toward a Richer Representation of Sequence Variation in the Sequence Ontology Michael Bada 1 and Karen Eilbeck 2 1
 Toward a Richer Representation of Sequence Variation in the Sequence Ontology Michael Bada 1 and Karen Eilbeck 2 1 University of Colorado Anschutz Medical Campus, Department of Pharmacology, MS 8303, RC-1
Toward a Richer Representation of Sequence Variation in the Sequence Ontology Michael Bada 1 and Karen Eilbeck 2 1 University of Colorado Anschutz Medical Campus, Department of Pharmacology, MS 8303, RC-1
AGILENT S BIOINFORMATICS ANALYSIS SOFTWARE
 ACCELERATING PROGRESS IS IN OUR GENES AGILENT S BIOINFORMATICS ANALYSIS SOFTWARE GENESPRING GENE EXPRESSION (GX) MASS PROFILER PROFESSIONAL (MPP) PATHWAY ARCHITECT (PA) See Deeper. Reach Further. BIOINFORMATICS
ACCELERATING PROGRESS IS IN OUR GENES AGILENT S BIOINFORMATICS ANALYSIS SOFTWARE GENESPRING GENE EXPRESSION (GX) MASS PROFILER PROFESSIONAL (MPP) PATHWAY ARCHITECT (PA) See Deeper. Reach Further. BIOINFORMATICS
BIOINFORMATICS ORIGINAL PAPER doi: /bioinformatics/bts508
 Vol. 28 no. 20 2012, pages 2648 2653 BIOINFORMATICS ORIGINAL PAPER doi:10.1093/bioinformatics/bts508 Systems biology Advance Access publication August 24, 2012 Qualitative translation of relations from
Vol. 28 no. 20 2012, pages 2648 2653 BIOINFORMATICS ORIGINAL PAPER doi:10.1093/bioinformatics/bts508 Systems biology Advance Access publication August 24, 2012 Qualitative translation of relations from
The SEED Family
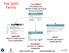 The SEED Family www.nmpdr.org www.theseed.org Number of known sequences How much has been sequenced? 100 Environmentalbacteria sequencing l genome s First bacterial genome Year 1,000 bacteria l genome
The SEED Family www.nmpdr.org www.theseed.org Number of known sequences How much has been sequenced? 100 Environmentalbacteria sequencing l genome s First bacterial genome Year 1,000 bacteria l genome
Outline and learning objectives. From Proteomics to Systems Biology. Integration of omics - information
 From to Systems Biology Outline and learning objectives Omics science provides global analysis tools to study entire systems How to obtain omics - What can we learn Limitations Integration of omics - In-class
From to Systems Biology Outline and learning objectives Omics science provides global analysis tools to study entire systems How to obtain omics - What can we learn Limitations Integration of omics - In-class
Euna Jeong. Curriculum Vitae
 Euna Jeong Curriculum Vitae Institute of Medical Science University of Tokyo 4-6-1 Shirokanedai, Minato-ku, Tokyo 108-8639 JAPAN eajeong@ims.u-tokyo.ac.jp Phone: +81-3-5449-5615 Fax: +81-3-5449-5442 Research
Euna Jeong Curriculum Vitae Institute of Medical Science University of Tokyo 4-6-1 Shirokanedai, Minato-ku, Tokyo 108-8639 JAPAN eajeong@ims.u-tokyo.ac.jp Phone: +81-3-5449-5615 Fax: +81-3-5449-5442 Research
Engineering Genetic Circuits
 Engineering Genetic Circuits I use the book and slides of Chris J. Myers Lecture 0: Preface Chris J. Myers (Lecture 0: Preface) Engineering Genetic Circuits 1 / 19 Samuel Florman Engineering is the art
Engineering Genetic Circuits I use the book and slides of Chris J. Myers Lecture 0: Preface Chris J. Myers (Lecture 0: Preface) Engineering Genetic Circuits 1 / 19 Samuel Florman Engineering is the art
Big Data in Agriculture Challenges. Pascal Neveu INRA Montpellier
 Big Data in Agriculture Challenges INRA Montpellier The rise of Big Data in agriculture More data production from heterogeneous sources 2 The rise of Big Data in agriculture More and more data services
Big Data in Agriculture Challenges INRA Montpellier The rise of Big Data in agriculture More data production from heterogeneous sources 2 The rise of Big Data in agriculture More and more data services
Bioinformatics & Protein Structural Analysis. Bioinformatics & Protein Structural Analysis. Learning Objective. Proteomics
 The molecular structures of proteins are complex and can be defined at various levels. These structures can also be predicted from their amino-acid sequences. Protein structure prediction is one of the
The molecular structures of proteins are complex and can be defined at various levels. These structures can also be predicted from their amino-acid sequences. Protein structure prediction is one of the
Inter-Agency Conference on Metabolic Engineering
 Proceedings of the Inter-Agency Conference on Metabolic Engineering 2006 Metabolic Engineering Working Group February 14, 2006 WORLD TECHNOLOGY EVALUATION CENTER, INC. The World Technology Evaluation Center,
Proceedings of the Inter-Agency Conference on Metabolic Engineering 2006 Metabolic Engineering Working Group February 14, 2006 WORLD TECHNOLOGY EVALUATION CENTER, INC. The World Technology Evaluation Center,
CS 5984: Topics and Schedule
 CS 5984: and Schedule T. M. Murali January 19, 2006 T. M. Murali January 19, 2006 CS 5984: and Schedule Continuum of Models in Systems Biology From Building with a scaffold: emerging strategies for high-
CS 5984: and Schedule T. M. Murali January 19, 2006 T. M. Murali January 19, 2006 CS 5984: and Schedule Continuum of Models in Systems Biology From Building with a scaffold: emerging strategies for high-
What is Bioinformatics?
 What is Bioinformatics? Bioinformatics is the field of science in which biology, computer science, and information technology merge to form a single discipline. - NCBI The ultimate goal of the field is
What is Bioinformatics? Bioinformatics is the field of science in which biology, computer science, and information technology merge to form a single discipline. - NCBI The ultimate goal of the field is
Product Applications for the Sequence Analysis Collection
 Product Applications for the Sequence Analysis Collection Pipeline Pilot Contents Introduction... 1 Pipeline Pilot and Bioinformatics... 2 Sequence Searching with Profile HMM...2 Integrating Data in a
Product Applications for the Sequence Analysis Collection Pipeline Pilot Contents Introduction... 1 Pipeline Pilot and Bioinformatics... 2 Sequence Searching with Profile HMM...2 Integrating Data in a
Glycomics Project Overview
 Wright State University CORE Scholar Kno.e.sis Publications The Ohio Center of Excellence in Knowledge- Enabled Computing (Kno.e.sis) 2007 Glycomics Project Overview Satya S. Sahoo Wright State University
Wright State University CORE Scholar Kno.e.sis Publications The Ohio Center of Excellence in Knowledge- Enabled Computing (Kno.e.sis) 2007 Glycomics Project Overview Satya S. Sahoo Wright State University
Using genome-context data to identify specific types of functional associations in pathway/genome databases
 BIOINFORMATICS Vol. 23 ISMB/ECCB 2007, pages i205 i211 doi:10.1093/bioinformatics/btm213 Using genome-context data to identify specific types of functional associations in pathway/genome databases Michelle
BIOINFORMATICS Vol. 23 ISMB/ECCB 2007, pages i205 i211 doi:10.1093/bioinformatics/btm213 Using genome-context data to identify specific types of functional associations in pathway/genome databases Michelle
Using semantic web technology to accelerate plant breeding.
 Using semantic web technology to accelerate plant breeding. Pierre-Yves Chibon 1,2,3, Benoît Carrères 1, Heleena de Weerd 1, Richard G. F. Visser 1,2,3, and Richard Finkers 1,3 1 Wageningen UR Plant Breeding,
Using semantic web technology to accelerate plant breeding. Pierre-Yves Chibon 1,2,3, Benoît Carrères 1, Heleena de Weerd 1, Richard G. F. Visser 1,2,3, and Richard Finkers 1,3 1 Wageningen UR Plant Breeding,
COMPARATIVE DETERMINATION OF BIOMASS COMPOSITION IN DIFFERENTIALLY ACTIVE METABOLIC STATES
 COMPARATIVE DETERMINATION OF BIOMASS COMPOSITION IN DIFFERENTIALLY ACTIVE METABOLIC STATES HSUAN-CHAO CHIU DANIEL SEGRÈ, hcchiu@bu.edu dsegre@bu.edu Graduate Program in Bioinformatics, Boston University,
COMPARATIVE DETERMINATION OF BIOMASS COMPOSITION IN DIFFERENTIALLY ACTIVE METABOLIC STATES HSUAN-CHAO CHIU DANIEL SEGRÈ, hcchiu@bu.edu dsegre@bu.edu Graduate Program in Bioinformatics, Boston University,
What Can Semantics do for Bioinformatics?
 Wright State University CORE Scholar Kno.e.sis Publications The Ohio Center of Excellence in Knowledge- Enabled Computing (Kno.e.sis) 10-29-2003 What Can Semantics do for Bioinformatics? Amit P. Sheth
Wright State University CORE Scholar Kno.e.sis Publications The Ohio Center of Excellence in Knowledge- Enabled Computing (Kno.e.sis) 10-29-2003 What Can Semantics do for Bioinformatics? Amit P. Sheth
Integrating quantitative proteomics and metabolomics with a genome-scale metabolic network model
 BIOINFORMATICS Vol. 26 ISMB 2010, pages i255 i260 doi:10.1093/bioinformatics/btq183 Integrating quantitative proteomics and metabolomics with a genome-scale metabolic network model Keren Yizhak 1,, Tomer
BIOINFORMATICS Vol. 26 ISMB 2010, pages i255 i260 doi:10.1093/bioinformatics/btq183 Integrating quantitative proteomics and metabolomics with a genome-scale metabolic network model Keren Yizhak 1,, Tomer
From Proteomics to Systems Biology. Integration of omics - information
 From Proteomics to Systems Biology Integration of omics - information Outline and learning objectives Omics science provides global analysis tools to study entire systems How to obtain omics - data What
From Proteomics to Systems Biology Integration of omics - information Outline and learning objectives Omics science provides global analysis tools to study entire systems How to obtain omics - data What
Pathway Analysis. Min Kim Bioinformatics Core Facility 2/28/2018
 Pathway Analysis Min Kim Bioinformatics Core Facility 2/28/2018 Outline 1. Background 2. Databases: KEGG, Reactome, Biocarta, Gene Ontology, MSigDB, MetaCyc, SMPDB, IPA. 3. Statistical Methods: Overlap
Pathway Analysis Min Kim Bioinformatics Core Facility 2/28/2018 Outline 1. Background 2. Databases: KEGG, Reactome, Biocarta, Gene Ontology, MSigDB, MetaCyc, SMPDB, IPA. 3. Statistical Methods: Overlap
Constraining the metabolic genotype phenotype relationship using a phylogeny of in silico methods
 Constraining the metabolic genotype phenotype relationship using a phylogeny of in silico methods Nathan E. Lewis 1, Harish Nagarajan 2 and Bernhard O. Palsson 1 Abstract Reconstructed microbial metabolic
Constraining the metabolic genotype phenotype relationship using a phylogeny of in silico methods Nathan E. Lewis 1, Harish Nagarajan 2 and Bernhard O. Palsson 1 Abstract Reconstructed microbial metabolic
Ad hoc improvement in Biotechnology thanks to ad hoc application of Computer Science
 University of Verona Department of Biotechnology Department of Computer Science Ad hoc improvement in Biotechnology thanks to ad hoc application of Computer Science Elisa Salvetti Giovanna E. Felis Diego
University of Verona Department of Biotechnology Department of Computer Science Ad hoc improvement in Biotechnology thanks to ad hoc application of Computer Science Elisa Salvetti Giovanna E. Felis Diego
Expanding BioPAX format by integrating Gene Regulation. Biological information. EMBnet.journal 16.1 RESEARCH PAPERS 39
 EMBnet.journal 16.1 RESEARCH PAPERS 39 Expanding BioPAX format by integrating Gene Regulation Irma Martínez-Flores, Verónica Jiménez-Jacinto, Alejandra C. López-Fuentes and Julio Collado-Vides Programa
EMBnet.journal 16.1 RESEARCH PAPERS 39 Expanding BioPAX format by integrating Gene Regulation Irma Martínez-Flores, Verónica Jiménez-Jacinto, Alejandra C. López-Fuentes and Julio Collado-Vides Programa
Ontologies and the Dynamics of Organisational Environments: An Example of a Group Memory System for the Management of Group Competencies
 Proceedings of I-KNOW 03 Graz, Austria, July 2-4, 2003 Ontologies and the Dynamics of Organisational Environments: An Example of a Group Memory System for the Management of Group Competencies José Braga
Proceedings of I-KNOW 03 Graz, Austria, July 2-4, 2003 Ontologies and the Dynamics of Organisational Environments: An Example of a Group Memory System for the Management of Group Competencies José Braga
Microarray Data Analysis in GeneSpring GX 11. Month ##, 200X
 Microarray Data Analysis in GeneSpring GX 11 Month ##, 200X Agenda Genome Browser GO GSEA Pathway Analysis Network building Find significant pathways Extract relations via NLP Data Visualization Options
Microarray Data Analysis in GeneSpring GX 11 Month ##, 200X Agenda Genome Browser GO GSEA Pathway Analysis Network building Find significant pathways Extract relations via NLP Data Visualization Options
Technical University of Denmark
 1 of 13 Technical University of Denmark Written exam, 15 December 2007 Course name: Introduction to Systems Biology Course no. 27041 Aids allowed: Open Book Exam Provide your answers and calculations on
1 of 13 Technical University of Denmark Written exam, 15 December 2007 Course name: Introduction to Systems Biology Course no. 27041 Aids allowed: Open Book Exam Provide your answers and calculations on
Introduction to Software Testing
 Introduction to Software Testing Introduction Chapter 1 introduces software testing by : describing the activities of a test engineer defining a number of key terms explaining the central notion of test
Introduction to Software Testing Introduction Chapter 1 introduces software testing by : describing the activities of a test engineer defining a number of key terms explaining the central notion of test
Genetics Lecture Notes Lectures 17 19
 Genetics Lecture Notes 7.03 2005 Lectures 17 19 Lecture 17 Gene Regulation We are now going to look at ways that genetics can be used to study gene regulation. The issue is how cells adjust the expression
Genetics Lecture Notes 7.03 2005 Lectures 17 19 Lecture 17 Gene Regulation We are now going to look at ways that genetics can be used to study gene regulation. The issue is how cells adjust the expression
The knowledge-driven exploration of integrated biomedical knowledge sources facilitates the generation of new hypotheses
 The knowledge-driven exploration of integrated biomedical knowledge sources facilitates the generation of new hypotheses Vinh Nguyen 1, Olivier Bodenreider 2, Todd Minning 3, and Amit Sheth 1 1 Kno.e.sis
The knowledge-driven exploration of integrated biomedical knowledge sources facilitates the generation of new hypotheses Vinh Nguyen 1, Olivier Bodenreider 2, Todd Minning 3, and Amit Sheth 1 1 Kno.e.sis
Adaptive Evolution Towards Predicted Optimal Growth in Escherichia coli K-12
 Adaptive Evolution Towards Predicted Optimal Growth in Escherichia coli K-12 Rafael U. Ibarra 1,*, Jeremy S. Edwards 2,*, and Bernhard O. Palsson 1,3 1 Department of Bioengineering University of California,
Adaptive Evolution Towards Predicted Optimal Growth in Escherichia coli K-12 Rafael U. Ibarra 1,*, Jeremy S. Edwards 2,*, and Bernhard O. Palsson 1,3 1 Department of Bioengineering University of California,
Functional profiling of metagenomic short reads: How complex are complex microbial communities?
 Functional profiling of metagenomic short reads: How complex are complex microbial communities? Rohita Sinha Senior Scientist (Bioinformatics), Viracor-Eurofins, Lee s summit, MO Understanding reality,
Functional profiling of metagenomic short reads: How complex are complex microbial communities? Rohita Sinha Senior Scientist (Bioinformatics), Viracor-Eurofins, Lee s summit, MO Understanding reality,
From Ontology for Genetic Interval (OGI) to Sequence Assembly Ontology Applying to Next Generation Sequencing
 From Ontology for Genetic Interval (OGI) to Sequence Assembly Ontology Applying to Next Generation Sequencing Yu Lin 1, Hiroshi Tarui 1, Peter Simons 2 1. Genome Resource and Analysis Unit, Genomics Laboratory,
From Ontology for Genetic Interval (OGI) to Sequence Assembly Ontology Applying to Next Generation Sequencing Yu Lin 1, Hiroshi Tarui 1, Peter Simons 2 1. Genome Resource and Analysis Unit, Genomics Laboratory,
Identifying Minimal Genomes and Essential Genes in Metabolic Model Using Flux Balance Analysis
 Identifying Minimal Genomes and Essential Genes in Metabolic Model Using Flux Balance Analysis Abdul Hakim Mohamed Salleh 1, Mohd Saberi Mohamad 1, Safaai Deris 1, and Rosli Md. Illias 2 1 Artificial Intelligence
Identifying Minimal Genomes and Essential Genes in Metabolic Model Using Flux Balance Analysis Abdul Hakim Mohamed Salleh 1, Mohd Saberi Mohamad 1, Safaai Deris 1, and Rosli Md. Illias 2 1 Artificial Intelligence
Introducing the Fair and Logical Trading Project p.1
 Introducing the Fair and Logical Trading Project David M. Eyers with Alan S. Abrahams, Steven O. Kimbrough, Jean M. Bacon, and Andrew J.I. Jones University of Cambridge, UK University of Pennsylvania,
Introducing the Fair and Logical Trading Project David M. Eyers with Alan S. Abrahams, Steven O. Kimbrough, Jean M. Bacon, and Andrew J.I. Jones University of Cambridge, UK University of Pennsylvania,
A New Database of Genetic and. Molecular Pathways. Minoru Kanehisa. sequencing projects have been. Mbp) and for several bacteria including
 Toward Pathway Engineering: A New Database of Genetic and Molecular Pathways Minoru Kanehisa Institute for Chemical Research, Kyoto University From Genome Sequences to Functions The Human Genome Project
Toward Pathway Engineering: A New Database of Genetic and Molecular Pathways Minoru Kanehisa Institute for Chemical Research, Kyoto University From Genome Sequences to Functions The Human Genome Project
Systems biology of genetic interactions in yeast metabolism
 Systems biology of genetic interactions in yeast metabolism Balázs Papp Evolutionary Systems Biology Group Biological Research Centre of the HAS Szeged, Hungary Genetic interaction (GI) refers to the nonindependence
Systems biology of genetic interactions in yeast metabolism Balázs Papp Evolutionary Systems Biology Group Biological Research Centre of the HAS Szeged, Hungary Genetic interaction (GI) refers to the nonindependence
Using genome context data to identify missing enzymes in PGDBs. SRI International Bioinformatics
 Using genome context data to identify missing enzymes in PGDBs 1 Outline Motivation Genome context methods used Principle behind PHFiller- Algorithm Bayesian classifier Validation Gold-standard dataset
Using genome context data to identify missing enzymes in PGDBs 1 Outline Motivation Genome context methods used Principle behind PHFiller- Algorithm Bayesian classifier Validation Gold-standard dataset
Can Semantic Web Technologies enable Translational Medicine? (Or Can Translational Medicine help enrich the Semantic Web?)
 Can Semantic Web Technologies enable Translational Medicine? (Or Can Translational Medicine help enrich the Semantic Web?) Vipul Kashyap 1, Tonya Hongsermeier 1, Samuel Aronson 2 1 Clinical Informatics
Can Semantic Web Technologies enable Translational Medicine? (Or Can Translational Medicine help enrich the Semantic Web?) Vipul Kashyap 1, Tonya Hongsermeier 1, Samuel Aronson 2 1 Clinical Informatics
Experience with Weka by Predictive Classification on Gene-Expre
 Experience with Weka by Predictive Classification on Gene-Expression Data Czech Technical University in Prague Faculty of Electrical Engineering Department of Cybernetics Intelligent Data Analysis lab
Experience with Weka by Predictive Classification on Gene-Expression Data Czech Technical University in Prague Faculty of Electrical Engineering Department of Cybernetics Intelligent Data Analysis lab
Imaging informatics computer assisted mammogram reading Clinical aka medical informatics CDSS combining bioinformatics for diagnosis, personalized
 1 2 3 Imaging informatics computer assisted mammogram reading Clinical aka medical informatics CDSS combining bioinformatics for diagnosis, personalized medicine, risk assessment etc Public Health Bio
1 2 3 Imaging informatics computer assisted mammogram reading Clinical aka medical informatics CDSS combining bioinformatics for diagnosis, personalized medicine, risk assessment etc Public Health Bio
METABOLIC CAPABILITIES OF Actinobacillus succinogenes FOR SUCCINIC ACID PRODUCTION
 Brazilian Journal of Chemical Engineering ISSN 14-6632 Printed in Brazil www.abeq.org.br/bjche Vol. 31, No. 4, pp. 859-865, October - December, 214 dx.doi.org/1.159/14-6632.214314s2997 METABOLIC CAPABILITIES
Brazilian Journal of Chemical Engineering ISSN 14-6632 Printed in Brazil www.abeq.org.br/bjche Vol. 31, No. 4, pp. 859-865, October - December, 214 dx.doi.org/1.159/14-6632.214314s2997 METABOLIC CAPABILITIES
Rapporto di Ricerca CS
 UNIVERSITÀ CA FOSCARI DI VENEZIA Dipartimento di Informatica Technical Report Series in Computer Science Rapporto di Ricerca CS-2006-8 Novembre 2006 Andrea Marin Biological Pathways representation using
UNIVERSITÀ CA FOSCARI DI VENEZIA Dipartimento di Informatica Technical Report Series in Computer Science Rapporto di Ricerca CS-2006-8 Novembre 2006 Andrea Marin Biological Pathways representation using
Semantic Web Technologies for Biomedicine. Natasha Noy Stanford Center for Biomedical Informatics Research Stanford University
 Semantic Web Technologies for Biomedicine Natasha Noy Stanford Center for Biomedical Informatics Research Stanford University Why Medicine and Biology? LOTS of data The data is in different database silos
Semantic Web Technologies for Biomedicine Natasha Noy Stanford Center for Biomedical Informatics Research Stanford University Why Medicine and Biology? LOTS of data The data is in different database silos
Discovery of Novel Metabolic Pathways in PGDBs
 Discovery of Novel Metabolic Pathways in PGDBs Luciana Ferrer Alexander Shearer Peter D. Karp Bioinformatics Research Group SRI International 1 Introduction We propose a computational method for the discovery
Discovery of Novel Metabolic Pathways in PGDBs Luciana Ferrer Alexander Shearer Peter D. Karp Bioinformatics Research Group SRI International 1 Introduction We propose a computational method for the discovery
OntoNaviERP: Ontology-supported Navigation in ERP Software Documentation
 OntoNaviERP: Ontology-supported Navigation in ERP Software Documentation 1,2 and Andreas Wechselberger 1 1 E-Business and Web Science Research Group, Bundeswehr University Munich, Germany 2 STI Innsbruck,
OntoNaviERP: Ontology-supported Navigation in ERP Software Documentation 1,2 and Andreas Wechselberger 1 1 E-Business and Web Science Research Group, Bundeswehr University Munich, Germany 2 STI Innsbruck,
Lecture 11: Constraint-Based Modelling: Variants
 Lecture 11: Constraint-Based Modelling: Variants AICTE STTP: Computational Systems Biology Karthik Raman Department of Biotechnology Indian Institute of Technology Madras https://home.iitm.ac.in/kraman/lab/
Lecture 11: Constraint-Based Modelling: Variants AICTE STTP: Computational Systems Biology Karthik Raman Department of Biotechnology Indian Institute of Technology Madras https://home.iitm.ac.in/kraman/lab/
IROA Metabolic Profiling Kits FAQ
 IROA Metabolic Profiling Kits FAQ Q: What does IROA stand for? A: The acronym refers to Isotope Ratio Outlier Analysis. Q: What does the IROA protocol involve? A: For the Basic IROA protocol, biomolecules
IROA Metabolic Profiling Kits FAQ Q: What does IROA stand for? A: The acronym refers to Isotope Ratio Outlier Analysis. Q: What does the IROA protocol involve? A: For the Basic IROA protocol, biomolecules
Getting Started in Biological Pathway Construction and Analysis Ganesh A. Viswanathan, Jeremy Seto, Sonali Patil, German Nudelman, Stuart C.
 Message from ISCB Getting Started in Biological Pathway Construction and Analysis Ganesh A. Viswanathan, Jeremy Seto, Sonali Patil, German Nudelman, Stuart C. Sealfon * Introduction Life depends on the
Message from ISCB Getting Started in Biological Pathway Construction and Analysis Ganesh A. Viswanathan, Jeremy Seto, Sonali Patil, German Nudelman, Stuart C. Sealfon * Introduction Life depends on the
Combinatorial complexity of pathway analysis in metabolic networks
 Combinatorial complexity of pathway analysis in metabolic networks Steffen Klamt 1 and Jörg Stelling Max Planck Institute for Dynamics of Complex Technical Systems, Sandtorstrasse 1, D-39106 Magdeburg,
Combinatorial complexity of pathway analysis in metabolic networks Steffen Klamt 1 and Jörg Stelling Max Planck Institute for Dynamics of Complex Technical Systems, Sandtorstrasse 1, D-39106 Magdeburg,
Ontologies - Useful tools in Life Sciences and Forensics
 Ontologies - Useful tools in Life Sciences and Forensics How today's Life Science Technologies can shape the Crime Sciences of tomorrow 04.07.2015 Dirk Labudde Mittweida Mittweida 2 Watson vs Watson Dr.
Ontologies - Useful tools in Life Sciences and Forensics How today's Life Science Technologies can shape the Crime Sciences of tomorrow 04.07.2015 Dirk Labudde Mittweida Mittweida 2 Watson vs Watson Dr.
Towards rational assessment of changes in metabolites from genetic modification:
 Towards rational assessment of changes in metabolites from genetic modification: Opportunities and challenges in Genomics, metabolomics, and metabolic network modeling Sue Rhee Carnegie Institution for
Towards rational assessment of changes in metabolites from genetic modification: Opportunities and challenges in Genomics, metabolomics, and metabolic network modeling Sue Rhee Carnegie Institution for
Knowledge discovery in biological data. Ivanisenko V.A Institute of Cytology and Genetics SB RAS Novosibirsk, Russia
 Knowledge discovery in biological data Ivanisenko V.A Institute of Cytology and Genetics SB RAS Novosibirsk, Russia Increase in body of biological data Publications are the main source of knowledge for
Knowledge discovery in biological data Ivanisenko V.A Institute of Cytology and Genetics SB RAS Novosibirsk, Russia Increase in body of biological data Publications are the main source of knowledge for
KEGG Kyoto Encyclopedia of Genes and Genomes
 KEGG Kyoto Encyclopedia of Genes and Genomes Objectives of KEGG Computerize current knowledge of biological systems in terms of pathway of interacting molecules or genes Maintain gene catalogs for sequenced
KEGG Kyoto Encyclopedia of Genes and Genomes Objectives of KEGG Computerize current knowledge of biological systems in terms of pathway of interacting molecules or genes Maintain gene catalogs for sequenced
MODEL 1. provides additional information about the performance of the CeSS algorithm.
 MODEL 1 Sections 1 to 4 describe Model 1, a metabolic model of E. coli s Central Carbon Metabolism. Section 5 provides additional information about the performance of the CeSS algorithm. 1 Model structure:
MODEL 1 Sections 1 to 4 describe Model 1, a metabolic model of E. coli s Central Carbon Metabolism. Section 5 provides additional information about the performance of the CeSS algorithm. 1 Model structure:
ONTOLOGY BASED INFORMATION EXTRACTION FOR DISEASE INTELLIGENCE
 International Journal of Research in Computer Science eissn 2249-8265 Volume 2 Issue 6 (2012) pp. 7-19, A Unit of White Globe Publications doi: 10.7815/ijorcs.26.2012.051 ONTOLOGY BASED INFORMATION EXTRACTION
International Journal of Research in Computer Science eissn 2249-8265 Volume 2 Issue 6 (2012) pp. 7-19, A Unit of White Globe Publications doi: 10.7815/ijorcs.26.2012.051 ONTOLOGY BASED INFORMATION EXTRACTION
Jianlin Cheng, PhD Informatics Institute, Computer Science Department University of Missouri, Columbia Fall, 2011
 Jianlin Cheng, PhD Informatics Institute, Computer Science Department University of Missouri, Columbia Fall, 2011 Objectives Walk students through the complete process of sequencing, assembling and annotating
Jianlin Cheng, PhD Informatics Institute, Computer Science Department University of Missouri, Columbia Fall, 2011 Objectives Walk students through the complete process of sequencing, assembling and annotating
Upstream/Downstream Relation Detection of Signaling Molecules using Microarray Data
 Vol 1 no 1 2005 Pages 1 5 Upstream/Downstream Relation Detection of Signaling Molecules using Microarray Data Ozgun Babur 1 1 Center for Bioinformatics, Computer Engineering Department, Bilkent University,
Vol 1 no 1 2005 Pages 1 5 Upstream/Downstream Relation Detection of Signaling Molecules using Microarray Data Ozgun Babur 1 1 Center for Bioinformatics, Computer Engineering Department, Bilkent University,
Metabolic modelling in the development of cell factories by synthetic biology
 , http://dx.doi.org/10.5936/csbj.201210009 CSBJ Metabolic modelling in the development of cell factories by synthetic biology Paula Jouhten a,* Abstract: Cell factories are commonly microbial organisms
, http://dx.doi.org/10.5936/csbj.201210009 CSBJ Metabolic modelling in the development of cell factories by synthetic biology Paula Jouhten a,* Abstract: Cell factories are commonly microbial organisms
Ontologies examples and applications
 Web Science & Technologies University of Koblenz Landau, Germany Ontologies examples and applications 2 UMLS - Unified Medical Language System Framework consisting of several knowledge bases and according
Web Science & Technologies University of Koblenz Landau, Germany Ontologies examples and applications 2 UMLS - Unified Medical Language System Framework consisting of several knowledge bases and according
Project Manual Bio3055. Lung Cancer: Cytochrome P450 1A1
 Project Manual Bio3055 Lung Cancer: Cytochrome P450 1A1 Bednarski 2003 Funded by HHMI Lung Cancer: Cytochrome P450 1A1 Introduction: It is well known that smoking leads to an increased risk for lung cancer,
Project Manual Bio3055 Lung Cancer: Cytochrome P450 1A1 Bednarski 2003 Funded by HHMI Lung Cancer: Cytochrome P450 1A1 Introduction: It is well known that smoking leads to an increased risk for lung cancer,
Likelihood-Based Gene Annotations for Gap Filling and Quality Assessment in Genome-Scale Metabolic Models
 Likelihood-Based Gene Annotations for Gap Filling and Quality Assessment in Genome-Scale Metabolic Models Matthew N. Benedict 1, Michael B. Mundy 2, Christopher S. Henry 3, Nicholas Chia 2,4,5 *, Nathan
Likelihood-Based Gene Annotations for Gap Filling and Quality Assessment in Genome-Scale Metabolic Models Matthew N. Benedict 1, Michael B. Mundy 2, Christopher S. Henry 3, Nicholas Chia 2,4,5 *, Nathan
CSE 259: Logic in Computer Science (Spring 2017)
 CSE 259: Logic in Computer Science (Spring 2017) Final Study Guide Final: In class, Monday, May 1, 2:30PM to 4:20PM (1 hour & 50min) Comprehensive but with a bit more emphasis on the materials after mid-term
CSE 259: Logic in Computer Science (Spring 2017) Final Study Guide Final: In class, Monday, May 1, 2:30PM to 4:20PM (1 hour & 50min) Comprehensive but with a bit more emphasis on the materials after mid-term
Textbook Reading Guidelines
 Understanding Bioinformatics by Marketa Zvelebil and Jeremy Baum Last updated: May 1, 2009 Textbook Reading Guidelines Preface: Read the whole preface, and especially: For the students with Life Science
Understanding Bioinformatics by Marketa Zvelebil and Jeremy Baum Last updated: May 1, 2009 Textbook Reading Guidelines Preface: Read the whole preface, and especially: For the students with Life Science
Virtual Company Dossier: Vision and Concepts
 Virtual Company Dossier: Vision and Concepts Josef Makolm Federal Ministry of Finance Hintere Zollamtsstrasse 4 1030 Wien Koblenz, Austria phone: +43 mail: josef.makolm@bmf.gv.at Ansgar Mondorf University
Virtual Company Dossier: Vision and Concepts Josef Makolm Federal Ministry of Finance Hintere Zollamtsstrasse 4 1030 Wien Koblenz, Austria phone: +43 mail: josef.makolm@bmf.gv.at Ansgar Mondorf University
Major: Biochemistry. University of California Irvine. Bachelor of Biochemistry
 Major: Biochemistry University of California Irvine Bachelor of Biochemistry Course # Course Name Unit Course Description 93 Bio Sci 4 Cell biology, biochemistry, genetics, and the biology of organ systems.
Major: Biochemistry University of California Irvine Bachelor of Biochemistry Course # Course Name Unit Course Description 93 Bio Sci 4 Cell biology, biochemistry, genetics, and the biology of organ systems.
Ingenuity Pathway Analysis (IPA )
 Ingenuity Pathway Analysis (IPA ) For the analysis and interpretation of omics data IPA is a web-based software application for the analysis, integration, and interpretation of data derived from omics
Ingenuity Pathway Analysis (IPA ) For the analysis and interpretation of omics data IPA is a web-based software application for the analysis, integration, and interpretation of data derived from omics
Minimal metabolic pathway structure is consistent with associated biomolecular interactions
 Minimal metabolic pathway structure is consistent with associated biomolecular interactions Aarash Bordbar, Harish Nagarajan, Nathan E. Lewis, Haythem Latif, Ali Ebrahim, Stephen Federowicz, Jan Schellenberger,
Minimal metabolic pathway structure is consistent with associated biomolecular interactions Aarash Bordbar, Harish Nagarajan, Nathan E. Lewis, Haythem Latif, Ali Ebrahim, Stephen Federowicz, Jan Schellenberger,
Handling the heterogeneity of genomic and metabolic networks data within flexible workflows with the PADMet toolbox
 Handling the heterogeneity of genomic and metabolic networks data within flexible workflows with the PADMet toolbox Marie Chevallier, Meziane Aite, Jeanne Got, Guillaume Collet, Nicolas Loira, María Paz
Handling the heterogeneity of genomic and metabolic networks data within flexible workflows with the PADMet toolbox Marie Chevallier, Meziane Aite, Jeanne Got, Guillaume Collet, Nicolas Loira, María Paz
Shimadzu mul+- omics data analysis Tutorial 2017 June
 Shimadzu mul+- omics data analysis Tutorial 2017 June 2 What is the Shimadzu mul'- omics data analysis? 3 Shimadzu mul'- omics analysis visualiza'on 4 Shimadzu Mul'- omics analysis Gadgets 5 Installa'on
Shimadzu mul+- omics data analysis Tutorial 2017 June 2 What is the Shimadzu mul'- omics data analysis? 3 Shimadzu mul'- omics analysis visualiza'on 4 Shimadzu Mul'- omics analysis Gadgets 5 Installa'on
