Design Principles in Synthetic Biology
|
|
|
- Liliana Poole
- 6 years ago
- Views:
Transcription
1 Design Principles in Synthetic Biology Chris Myers 1, Nathan Barker 2, Hiroyuki Kuwahara 3, Curtis Madsen 1, Nam Nguyen 1, Michael Samoilov 4, and Adam Arkin 4 1 University of Utah 2 Southern Utah University 3 Microsoft Research, Trento, Italy 4 University of California, Berkeley Design Principles in Biological Systems April 24, 2008
2 Synthetic Biology Increasing number of labs are designing more ambitious and mission critical synthetic biology projects. These projects construct synthetic genetic circuits from DNA. These synthetic genetic circuits can potentially result in: A better understanding of how microorganisms function by examining differences in vivo compared to in silico (Sprinzak/Elowitz). More efficient pathways for the production of antimalarial drugs (Dae et al.). Bacteria that can metabolize toxic chemicals (Brazil et al.). Bacteria that can hunt and kill tumors (Anderson et al.).
3 Genetic Design Automation (GDA) Electronic Design Automation (EDA) tools have facilitated the design of ever more complex integrated circuits each year. Crucial to the success of synthetic biology is an improvement in methods and tools for Genetic Design Automation (GDA). Existing GDA tools require biologists to design at the molecular level. Roughly equivalent to designing electronic circuits at the layout level. Analysis of genetic circuits is also performed at this very low level. A GDA tool that supports higher levels of abstraction is essential.
4 Overview This talk describes our research to develop such a GDA tool. This tool has helped us examine design principles for synthetic biology. As a case study, will describe the design of a genetic Muller C-element.
5 Current State of GDA Tools MIT has created a registry of standard biological parts used to design synthetic genetic circuits ( Methods and tools are needed to assist in the design and analysis of synthetic genetic circuits using these parts. BioJADE provides a schematic capture interface to the MIT parts registry. Systems Biology Markup Language (SBML) has been proposed as a standard representation for the simulation of biological systems. Many simulation tools have been developed that accept models in the SBML format (BioPathwise, BioSPICE, CellDesigner, SimBiology, etc.).
6 Systems Biology Markup Language (SBML) SBML models biological systems at the molecular level. A typical SBML model is composed of a number of chemical species (i.e., proteins, genes, etc.) and reactions that transform these species. This is a very low level representation which is roughly equivalent to the layout level for electronic circuits. Designing and simulating genetic circuits at this level of detail is extremely tedious and time-consuming. Therefore, there is a need for higher-level abstractions for modeling, analysis, and design of genetic circuits.
7 BioSim Genetic Circuit Perform Experiments Insert into Host Plasmid Construct Plasmid Experimental Biological DNA Data Knowledge Sequence Learn Model GCM Synthesis Simulation Data Abstraction/ Simulation SBML Model
8 BioSim: Analysis Genetic Circuit Perform Experiments Insert into Host Plasmid Construct Plasmid Experimental Biological DNA Data Knowledge Sequence Learn Model GCM Synthesis Simulation Data Abstraction/ Simulation SBML Model
9 BioSim: Design Genetic Circuit Perform Experiments Insert into Host Plasmid Construct Plasmid Experimental Biological DNA Data Knowledge Sequence Learn Model GCM Synthesis Simulation Data Abstraction/ Simulation SBML Model
10 BioSim: Design Genetic Circuit Perform Experiments Insert into Host Plasmid Construct Plasmid Experimental Biological DNA Data Knowledge Sequence Learn Model GCM Synthesis Simulation Data Abstraction/ Simulation SBML Model
11 BioSim: Genetic Circuit Model Genetic Circuit Perform Experiments Insert into Host Plasmid Construct Plasmid Experimental Biological DNA Data Knowledge Sequence Learn Model GCM Synthesis Simulation Data Abstraction/ Simulation SBML Model
12 Phage λ Virus
13 Phage λ Decision Circuit
14 Phage λ Decision Circuit
15 Genetic Circuits CI Dimer DNA Dimerization CI Protein Repression Activation Pre O E O R ci Operator Sites CII Protein Degradation Translation mrna Transcription Pr RNAP RNAP RNAP cii Promoters Genes
16 Genetic Circuits CI Dimer DNA Dimerization CI Protein Repression Activation Pre O E O R ci Operator Sites CII Protein Degradation Translation mrna Transcription Pr RNAP RNAP RNAP cii Promoters Genes
17 Genetic Circuits CI Dimer DNA Dimerization CI Protein Repression Activation Pre O E O R ci Operator Sites CII Protein Degradation Translation mrna Transcription Pr RNAP RNAP RNAP cii Promoters Genes
18 Genetic Circuits CI Dimer DNA Dimerization CI Protein Repression Activation Pre O E O R ci Operator Sites CII Protein Degradation Translation mrna Transcription RNAP RNAP RNAP RNAP cii Promoters Genes
19 Genetic Circuits CI Dimer DNA Dimerization CI Protein Repression Activation Pre O E O R ci Operator Sites CII Protein Degradation Translation mrna Transcription RNAP RNAP RNAP cii Promoters Genes
20 Genetic Circuits CI Dimer DNA Dimerization CI Protein Repression Activation Pre O E O R ci Operator Sites CII Protein Degradation Translation mrna Transcription RNAP RNAP RNAP cii Promoters Genes
21 Genetic Circuits CI Dimer DNA Dimerization CI Protein Repression Activation Pre O E O R ci Operator Sites CII Protein Degradation Translation mrna Transcription Pr RNAP RNAP RNAP cii Promoters Genes
22 Genetic Circuits CI Dimer DNA Dimerization CI Protein Repression Activation Pre O E O R ci Operator Sites CII Protein Degradation Translation mrna Transcription Pr RNAP RNAP RNAP cii Promoters Genes
23 Genetic Circuits CI Dimer DNA Dimerization CI Protein Repression Activation Pre O E O R ci Operator Sites CII Protein Degradation Translation mrna Transcription Pr RNAP RNAP RNAP cii Promoters Genes
24 Genetic Circuits CI Dimer DNA Dimerization CI Protein Repression Activation Pre O E O R ci Operator Sites CII Protein Degradation Translation mrna Transcription Pr RNAP RNAP RNAP cii Promoters Genes
25 Genetic Circuits CI Dimer DNA Dimerization CI Protein Repression Activation Pre O E O R ci Operator Sites CII Protein Degradation Translation mrna Transcription Pr RNAP RNAP RNAP cii Promoters Genes
26 Genetic Circuits CI Dimer DNA Dimerization ci CI Protein Repression Activation Pre O E O R Operator Sites CII Protein Transcription Pr RNAP RNAP RNAP cii Degradation Translation mrna Promoters Genes
27 Genetic Circuits CI Dimer DNA Dimerization ci CI Protein Repression Activation Pre O E O R Operator Sites CII Protein Transcription Pr RNAP RNAP RNAP cii Degradation Translation mrna Promoters Genes
28 Genetic Circuits CI Dimer DNA Dimerization CI Protein Repression Activation Pre O E O R ci Operator Sites CII Protein Degradation Translation mrna Transcription Pr RNAP RNAP RNAP cii Promoters Genes
29 Genetic Circuits CI Dimer DNA Dimerization CI Protein Repression Activation Pre O E O R ci Operator Sites CII Protein Degradation Translation mrna Transcription Pr RNAP RNAP RNAP cii Promoters Genes
30 Genetic Circuits CI Dimer DNA Dimerization CI Protein Repression Activation Pre O E O R ci Operator Sites CII Protein Degradation Translation mrna Transcription Pr RNAP RNAP RNAP cii Promoters Genes
31 Genetic Circuits CI Dimer DNA Dimerization CI Protein Repression Activation Pre O E O R ci Operator Sites CII Protein Degradation Translation mrna Transcription Pr RNAP RNAP RNAP cii Promoters Genes
32 Genetic Circuits CI Dimer DNA Dimerization CI Protein Repression Activation Pre O E O R ci Operator Sites CII Protein Degradation Translation mrna Transcription Pr RNAP RNAP RNAP cii Promoters Genes
33 Logical Representation CI CII
34 Graphical Representation CI Pre Pr CII
35 Genetic Circuit Model (GCM) Provides a higher level of abstraction than SBML. Includes only important species and their influences upon each other. A GCM is a tuple S,P,G,I,S d where: S is a finite set of species; P is a finite set of promoters; G : P 2 S maps promoters to sets of species; I S P {a,r} is a finite set of influences; S d S is a set of species that influence as dimers.
36 GCM Graphical Representation A bipartite graph with species and promoters as the two types of nodes. Species are connected to promoters using influences I, and promoters are connected to species using function G. To simplify presentation, graphs shown using only species as nodes, edges are inferred using I and G, and edges are labeled with the promoter that links the species.
37 Influences on the Same Promoter A B A B C P1 P1 C P1 c
38 Influences on the Same Promoter A B A B C P1 P1 C P1 c A B C
39 Influences on Different Promoters A C A B P1 C P2 B P1 c C P2 c
40 Influences on Different Promoters A C A B P1 C P2 B P1 c C P2 c A B C
41 GCM Parameters Parameter Sym Structure Value Units Initial species count n s species 0 molecule Dimerization equilibrium K d species.05 Degradation rate k d species molecule 1 sec Initial promoter count n g promoter 2 molecule Stoichiometry of production n p promoter 10 molecule Degree of cooperativity n c promoter 2 molecule RNAP binding equilibrium K o promoter.033 Open complex production rate k o promoter.05 Basal production rate k b promoter.0001 Activated production rate k a promoter.25 Repression binding equilibrium K r influence.5 Activation binding equilibrium K a influence molecule 1 sec 1 sec 1 sec 1 molecule nc 1 molecule (nc+1)
42 GCM versus SBML Representation CI Pre Pr CII
43 SBML Example
44 SBML Example
45 SBML Example
46 SBML Example
47 SBML Example
48 SBML Example
49 SBML Example
50 SBML Example
51 Synthesizing SBML from a GCM Representation Create degradation reactions Create open complex formation reactions Create dimerization reactions Create repression reactions Create activation reactions
52 Degradation Reactions
53 Open Complex Formation Reactions
54 Dimerization Reactions
55 Repression Reactions
56 Activation Reactions
57 Complete SBML Model
58 Classical Chemical Kinetics Uses ordinary differential equations (ODE) to represent the system to be analyzed, and it assumes: A system is well-stirred. Number of molecules in a cell is high. Concentrations can be viewed as continuous variables. Reactions occur continuously and deterministically. Genetic circuits involve small molecule counts. Gene expression can have substantial fluctuations. ODEs do not capture non-deterministic behavior.
59 Stochastic Chemical Kinetics To more accurately predict the temporal behavior of genetic circuits, stochastic chemical kinetics formalism can be used. Probabilistically predicts the dynamics of biochemical systems. Describes the time evolution of a system as a discrete-state jump Markov process governed by the chemical master equation (CME). Can simulate it using Gillespie s Stochastic Simulation Algorithm (SSA). It exactly tracks the quantities of each molecular species, and treats each reaction as a separate random event. Only practical for small systems with no major time-scale separations. Abstraction is essential for efficient analysis of any realistic system.
60 Automatic Abstraction Reaction Model Reaction-based Abstraction Abstracted Reaction Model State-based Abstraction SAC Model Markov Chain Analysis Stochastic Simulation Results Begins with a reaction-based model in SBML. Next, it automatically abstracts this model leveraging the quasi-steady state assumption, whenever possible. Finally, it encodes chemical species concentrations into Boolean (or n-ary) levels to produce a stochastic asynchronous circuit model. It can now be analyzed using Markov chain analysis.
61 Automatic Abstraction Reaction Model Reaction-based Abstraction Abstracted Reaction Model State-based Abstraction SAC Model Markov Chain Analysis Stochastic Simulation Results Begins with a reaction-based model in SBML. Next, it automatically abstracts this model leveraging the quasi-steady state assumption, whenever possible. Finally, it encodes chemical species concentrations into Boolean (or n-ary) levels to produce a stochastic asynchronous circuit model. It can now be analyzed using Markov chain analysis.
62 Automatic Abstraction Reaction Model Reaction-based Abstraction Abstracted Reaction Model State-based Abstraction SAC Model Markov Chain Analysis Stochastic Simulation Results Begins with a reaction-based model in SBML. Next, it automatically abstracts this model leveraging the quasi-steady state assumption, whenever possible. Finally, it encodes chemical species concentrations into Boolean (or n-ary) levels to produce a stochastic asynchronous circuit model. It can now be analyzed using Markov chain analysis.
63 Automatic Abstraction Reaction Model Reaction-based Abstraction Abstracted Reaction Model State-based Abstraction SAC Model Markov Chain Analysis Stochastic Simulation Results Begins with a reaction-based model in SBML. Next, it automatically abstracts this model leveraging the quasi-steady state assumption, whenever possible. Finally, it encodes chemical species concentrations into Boolean (or n-ary) levels to produce a stochastic asynchronous circuit model. It can now be analyzed using Markov chain analysis.
64 Automatic Abstraction Reaction Model Reaction-based Abstraction Abstracted Reaction Model State-based Abstraction SAC Model Markov Chain Analysis Stochastic Simulation Results Begins with a reaction-based model in SBML. Next, it automatically abstracts this model leveraging the quasi-steady state assumption, whenever possible. Finally, it encodes chemical species concentrations into Boolean (or n-ary) levels to produce a stochastic asynchronous circuit model. It can now be analyzed using Markov chain analysis.
65 Automatic Abstraction Reaction Model Reaction-based Abstraction Abstracted Reaction Model State-based Abstraction SAC Model Markov Chain Analysis Stochastic Simulation Results Begins with a reaction-based model in SBML. Next, it automatically abstracts this model leveraging the quasi-steady state assumption, whenever possible. Finally, it encodes chemical species concentrations into Boolean (or n-ary) levels to produce a stochastic asynchronous circuit model. It can now be analyzed using Markov chain analysis.
66 Automatic Abstraction Reaction Model Reaction-based Abstraction Abstracted Reaction Model State-based Abstraction SAC Model Markov Chain Analysis Stochastic Simulation Results Begins with a reaction-based model in SBML. Next, it automatically abstracts this model leveraging the quasi-steady state assumption, whenever possible. Finally, it encodes chemical species concentrations into Boolean (or n-ary) levels to produce a stochastic asynchronous circuit model. It can now be analyzed using Markov chain analysis.
67 Dimerization Reduction
68 Dimerization Reduction
69 Operator Site Reduction (PR)
70 Operator Site Reduction (PR)
71 Operator Site Reduction (PRE)
72 Operator Site Reduction (PRE)
73 Similar Reaction Combination
74 Modifier Constant Propagation
75 Final SBML Model 10 species and 10 reactions reduced to 2 species and 4 reactions
76 BioSim: Genetic Circuit Editor
77 BioSim: SBML Editor
78 BioSim: Simulator
79 BioSim: Parameter Editor
80 BioSim: Graph Editor
81 GCM Advantages Greatly increases the speed of model development and reduces the number of errors in the resulting models. Allows efficient exploration of the effects of parameter variation. Constrains SBML model such that it can be more easily abstracted resulting in substantial improvement in simulation time.
82 Genetic Muller C-Element A B C C A B C C 1 0 C 1 1 1
83 Toggle Switch C-Element (Genetic Circuit) A D X A B A B D E X F Y Z S R Q C P1 B P2 D P7 E d x E X e x P3 y F f F Z Y P8 f P4 z C Y Z P5 c y P6 z
84 Toggle Switch C-Element (GCM) A D X P1 B d E x X Y P2 D e x P3 y F E P7 f F Z P8 f P4 z C Y Z P5 c y P6 z
85 Toggle Switch C-Element (SBML)
86 Toggle Switch C-Element (Abstracted) Reduced from 34 species and 31 reactions to 9 species and 15 reactions.
87 Toggle Switch C-Element (Simulation) Simulation time improved from 312 seconds to 20 seconds.
88 Majority Gate C-Element (Genetic Circuit) A B X Y D E C Z A X Y D P1 x y P5 d B Z X D E C P2 z x P4 d P7 e P8 c D Y Z D P3 y z P6 d
89 Majority Gate C-Element (GCM) A X Y D P1 x y P5 d B Z X D E C P2 z x P4 d P7 e P8 c D Y Z D P3 y z P6 d
90 Majority Gate C-Element (Simulation)
91 Speed-Independent C-Element (Genetic Circuit) A B S1 S2 S4 S3 C A S4 X S1 S2 S3 P1 s4 x P4 s1 P5 s2 P6 s3 B S4 Y S1 S3 S2 Z P2 s4 y P7 s1 P8 s2 z S4 S3 Z C S2 S4 P3 s3 z P9 c P10 s4
92 Speed-Independent C-Element (GCM) A S4 X S1 S2 S3 P1 s4 x P4 s1 P5 s2 P6 s3 B S4 Y S1 S3 S2 Z P2 s4 y P7 s1 P8 s2 z S4 S3 Z C S2 S4 P3 s3 z P9 c P10 s4
93 Speed-Independent C-Element (Simulation)
94 Ordinary Differential Equation Analysis Use Law of Mass Action to derive an ODE model. Study behavior of our model at steady state. Analyze nullclines to characterize the gate.
95 ODE Analysis: Nullclines for Toggle C-Element Toggle, Inputs low 120 dy=0 dz= Y Z Stable
96 ODE Analysis: Nullclines for Toggle C-Element Toggle, Inputs Mixed 120 dy=0 dz=0 100 Y Stable Unstable Stable Z
97 ODE Analysis: Nullclines for Toggle C-Element 120 Stable Toggle, Inputs High dy=0 dz= Y Z
98 ODE Analysis: Nullclines for Toggle C-Element Toggle, Inputs Mixed 120 dy=0 dz=0 100 Y Stable Unstable Stable Z
99 Stochastic Simulation: State Change from Low to High Toggle, Inputs Mixed 120 dy=0 dz=0 100 Y Stable? 20 Unstable Stable Z
100 Stochastic Simulation: State Change from Low to High Low to High maj heat high maj light high tog heat high tog light high si heat high si light high Failure Rate Time (s)
101 Stochastic Simulation: State Change from High to Low Toggle, Inputs Mixed 120 dy=0 dz=0 100 Y Stable 40 20? Unstable Stable Z
102 Stochastic Simulation: State Change from High to Low High to Low maj heat low maj light low tog heat low tog light low si heat low si light low Failure Rate Time (s)
103 Effect of Gene Count Failure Rate Low to High maj heat high maj light high tog heat high tog light high si heat high si light high Number of Genes
104 Effect of Cooperativity Low to High maj heat high maj light high tog heat high tog light high si heat high si light high Failure Rate Cooperativity
105 Effect of Repression Strength Low to High maj heat high maj light high tog heat high tog light high si heat high si light high Failure Rate Repression
106 Effect of Decay Rates 1 Low to High Failure Rate maj heat high maj light high tog heat high tog light high si heat high si light high Decay Rate
107 Effect of Dual Rail Toggle, Inputs Mixed 120 dy=0 dz=0 100 Y Stable? 20 Unstable Stable Z
108 Effect of Dual Rail Low to High single tog heat high single tog light high dual tog heat high dual tog light high 0.02 Failure Rate Time (s)
109 Effect of Dual Rail Toggle, Inputs Mixed 120 dy=0 dz=0 100 Y Stable 40 20? Unstable Stable Z
110 Effect of Dual Rail High to Low single tog heat low single tog light low dual tog heat low dual tog light low Failure Rate Time (s)
111 Design Principles in Synthetic Biology Speed-independence does not necessarily imply better robustness. Higher gene counts improve production rates, higher equilibrium values, and more robust operation. Cooperativity of at least two is required to produce the necessary non-linearity for state-holding. Repressors should bind efficiently. Decay rates cannot be too high. Dual-rail outputs are essential.
112 Future Work: Modular Design More levels of hierarchy are needed in the GCM format. We plan to create structural constructs that allow us to connect GCM s for separate modules through species ports. Allow design at the logical and higher levels of abstraction.
113 Biologically Inspired Circuit Design Human inner ear performs the equivalent of one billion floating point operations per second and consumes only 14 µw while a game console with similar performance burns about 50 W (Sarpeshkar, 2006). We believe this difference is due to over designing components in order to achieve an extremely low probability of failure in every device. Future silicon and nano-devices will be much less reliable. For Moore s law to continue, future design methods should support the design of reliable systems using unreliable components. Biological systems constructed from very noisy and unreliable devices. GDA tools may be useful for future integrated circuit technologies. Biological systems tend to be more asynchronous and analog in nature, so future engineered circuits will likely need to be also.
114 Acknowledgments Nathan Barker Hiroyuki Kuwahara Nam Nguyen Curtis Madsen Michael Samoilov Adam Arkin This work is supported by the National Science Foundation under Grants No and CCF
Synthetic Biology: A New Application Area for Design Automation Research
 Synthetic Biology: A New Application Area for Design Automation Research Chris Myers 1, Nathan Barker 2, Hiroyuki Kuwahara 3, Curtis Madsen 1, Nam Nguyen 4, Chris Winstead 5 1 University of Utah 2 Southern
Synthetic Biology: A New Application Area for Design Automation Research Chris Myers 1, Nathan Barker 2, Hiroyuki Kuwahara 3, Curtis Madsen 1, Nam Nguyen 4, Chris Winstead 5 1 University of Utah 2 Southern
Engineering Genetic Circuits
 Engineering Genetic Circuits I use the book and slides of Chris J. Myers Lecture 0: Preface Chris J. Myers (Lecture 0: Preface) Engineering Genetic Circuits 1 / 19 Samuel Florman Engineering is the art
Engineering Genetic Circuits I use the book and slides of Chris J. Myers Lecture 0: Preface Chris J. Myers (Lecture 0: Preface) Engineering Genetic Circuits 1 / 19 Samuel Florman Engineering is the art
Digitally Programmed Cells
 Digitally Programmed Cells Ron Weiss PI: Tom Knight MIT Artificial Intelligence Laboratory Goal Process-Control Cellular Computers -- Microbial Robotics Unique features: small, self-replicating, energy-efficient
Digitally Programmed Cells Ron Weiss PI: Tom Knight MIT Artificial Intelligence Laboratory Goal Process-Control Cellular Computers -- Microbial Robotics Unique features: small, self-replicating, energy-efficient
Cell-free TX-TL systems for synthetic biology
 Cell-free TX-TL systems for synthetic biology Vincent Noireaux, University of Minnesota Caltech workshop, August 2013 1 Outline: Introduction - Motivations. Cell-free TX-TL. Synthetic gene circuits and
Cell-free TX-TL systems for synthetic biology Vincent Noireaux, University of Minnesota Caltech workshop, August 2013 1 Outline: Introduction - Motivations. Cell-free TX-TL. Synthetic gene circuits and
Synthetic Biology of Genetic Circuit
 Synthetic Biology of Genetic Circuit 1 Mallar, K. N. and 2 Prasenjit, D. 1 M.Sc. (Agri.), Dept. of Biotechnology, AAU, Jorhat, Assam 2 M.Sc. (Agri.), Dept. of Biotechnology, UAS, Dharwad, Karnataka Correspondence
Synthetic Biology of Genetic Circuit 1 Mallar, K. N. and 2 Prasenjit, D. 1 M.Sc. (Agri.), Dept. of Biotechnology, AAU, Jorhat, Assam 2 M.Sc. (Agri.), Dept. of Biotechnology, UAS, Dharwad, Karnataka Correspondence
Genetic Circuit Building Blocks for Cellular Computation, Communications, and Signal Processing
 Genetic Circuit Building Blocks for Cellular Computation, Communications, and Signal Processing Ron Weiss (rweiss@princeton.edu), Subhayu Basu (basu@princeton.edu), Sara Hooshangi (sara@princeton.edu),
Genetic Circuit Building Blocks for Cellular Computation, Communications, and Signal Processing Ron Weiss (rweiss@princeton.edu), Subhayu Basu (basu@princeton.edu), Sara Hooshangi (sara@princeton.edu),
Network System Inference
 Network System Inference Francis J. Doyle III University of California, Santa Barbara Douglas Lauffenburger Massachusetts Institute of Technology WTEC Systems Biology Final Workshop March 11, 2005 What
Network System Inference Francis J. Doyle III University of California, Santa Barbara Douglas Lauffenburger Massachusetts Institute of Technology WTEC Systems Biology Final Workshop March 11, 2005 What
Proposed Data Model for the Next Version of the Synthetic Biology Open Language
 pubs.acs.org/synthbio Proposed Data Model for the Next Version of the Synthetic Biology Open Language Nicholas Roehner,, * Ernst Oberortner, Matthew Pocock, Jacob Beal, Kevin Clancy, Curtis Madsen, Goksel
pubs.acs.org/synthbio Proposed Data Model for the Next Version of the Synthetic Biology Open Language Nicholas Roehner,, * Ernst Oberortner, Matthew Pocock, Jacob Beal, Kevin Clancy, Curtis Madsen, Goksel
Technical University of Denmark
 1 of 13 Technical University of Denmark Written exam, 15 December 2007 Course name: Introduction to Systems Biology Course no. 27041 Aids allowed: Open Book Exam Provide your answers and calculations on
1 of 13 Technical University of Denmark Written exam, 15 December 2007 Course name: Introduction to Systems Biology Course no. 27041 Aids allowed: Open Book Exam Provide your answers and calculations on
What is the Difference between a Human and a Chimp? T. M. Murali
 What is the Difference between a Human and a Chimp? T. M. Murali murali@cs.vt.edu http://bioinformatics.cs.vt.edu/ murali Introduction to Computational Biology and Bioinformatics (CS 3824) March 25 and
What is the Difference between a Human and a Chimp? T. M. Murali murali@cs.vt.edu http://bioinformatics.cs.vt.edu/ murali Introduction to Computational Biology and Bioinformatics (CS 3824) March 25 and
A systems approach to biology
 A systems approach to biology SB200 Lecture 2 18 September 2008 Jeremy Gunawardena jeremy@hms.harvard.edu I do not hold formal office hours. Please send me an e-mail if you have questions or would like
A systems approach to biology SB200 Lecture 2 18 September 2008 Jeremy Gunawardena jeremy@hms.harvard.edu I do not hold formal office hours. Please send me an e-mail if you have questions or would like
7.1 The lac Operon 7-1
 7.1 The lac Operon The lac operon was the first operon discovered It contains 3 genes coding for E. coli proteins that permit the bacteria to use the sugar lactose Galactoside permease (lacy) which transports
7.1 The lac Operon The lac operon was the first operon discovered It contains 3 genes coding for E. coli proteins that permit the bacteria to use the sugar lactose Galactoside permease (lacy) which transports
Synthetic Biology. Molecular Mechanisms. Concepts. Applications
 Synthetic Biology Molecular Mechanisms Concepts Applications 09.05.07 The central dogma - genetic machinery http://old.mb.au.dk/graphics/dogma.jpg Transcription (in E. Coli) DNA RNA Translation RNA Protein
Synthetic Biology Molecular Mechanisms Concepts Applications 09.05.07 The central dogma - genetic machinery http://old.mb.au.dk/graphics/dogma.jpg Transcription (in E. Coli) DNA RNA Translation RNA Protein
number Done by Corrected by Doctor Hamed Al Zoubi
 number 3 Done by Neda a Baniata Corrected by Waseem Abu Obeida Doctor Hamed Al Zoubi Note: it is important to refer to slides. Bacterial genetics *The main concepts we will talk about in this lecture:
number 3 Done by Neda a Baniata Corrected by Waseem Abu Obeida Doctor Hamed Al Zoubi Note: it is important to refer to slides. Bacterial genetics *The main concepts we will talk about in this lecture:
V 1 Introduction! Fri, Oct 24, 2014! Bioinformatics 3 Volkhard Helms!
 V 1 Introduction! Fri, Oct 24, 2014! Bioinformatics 3 Volkhard Helms! How Does a Cell Work?! A cell is a crowded environment! => many different proteins,! metabolites, compartments,! On a microscopic level!
V 1 Introduction! Fri, Oct 24, 2014! Bioinformatics 3 Volkhard Helms! How Does a Cell Work?! A cell is a crowded environment! => many different proteins,! metabolites, compartments,! On a microscopic level!
Learning Petri Net Models of Non-Linear Gene Interactions
 Learning Petri Net Models of Non-Linear Gene Interactions Abstract Understanding how an individual s genetic make-up influences their risk of disease is a problem of paramount importance. Although machine
Learning Petri Net Models of Non-Linear Gene Interactions Abstract Understanding how an individual s genetic make-up influences their risk of disease is a problem of paramount importance. Although machine
Optimizing Synthetic DNA for Metabolic Engineering Applications. Howard Salis Penn State University
 Optimizing Synthetic DNA for Metabolic Engineering Applications Howard Salis Penn State University Synthetic Biology Specify a function Build a genetic system (a DNA molecule) Genetic Pseudocode call producequorumsignal(luxi
Optimizing Synthetic DNA for Metabolic Engineering Applications Howard Salis Penn State University Synthetic Biology Specify a function Build a genetic system (a DNA molecule) Genetic Pseudocode call producequorumsignal(luxi
RNA and Protein Synthesis
 RNA and Protein Synthesis CTE: Agriculture and Natural Resources: C5.3 Understand various cell actions, such as osmosis and cell division. C5.4 Compare and contrast plant and animal cells, bacteria, and
RNA and Protein Synthesis CTE: Agriculture and Natural Resources: C5.3 Understand various cell actions, such as osmosis and cell division. C5.4 Compare and contrast plant and animal cells, bacteria, and
Synthetic Biology. Sustainable Energy. Therapeutics Industrial Enzymes. Agriculture. Accelerating Discoveries, Expanding Possibilities. Design.
 Synthetic Biology Accelerating Discoveries, Expanding Possibilities Sustainable Energy Therapeutics Industrial Enzymes Agriculture Design Build Generate Solutions to Advance Synthetic Biology Research
Synthetic Biology Accelerating Discoveries, Expanding Possibilities Sustainable Energy Therapeutics Industrial Enzymes Agriculture Design Build Generate Solutions to Advance Synthetic Biology Research
DNA Structure and Properties Basic Properties Predicting Melting Temperature. Dinesh Yadav
 DNA Structure and Properties Basic Properties Predicting Melting Temperature Dinesh Yadav Nucleic Acid Structure Question: Is this RNA or DNA? Molecules of Life, pp. 15 2 Nucleic Acid Bases Molecules of
DNA Structure and Properties Basic Properties Predicting Melting Temperature Dinesh Yadav Nucleic Acid Structure Question: Is this RNA or DNA? Molecules of Life, pp. 15 2 Nucleic Acid Bases Molecules of
THESIS A KINETIC MODEL DEVELOPMENT OF THE M13 BACTERIOPHAGE LIFE CYCLE. Submitted by. Steven William Smeal
 THESIS A KINETIC MODEL DEVELOPMENT OF THE M13 BACTERIOPHAGE LIFE CYCLE Submitted by Steven William Smeal Department of Chemical and Biological Engineering In partial fulfillment of the requirements For
THESIS A KINETIC MODEL DEVELOPMENT OF THE M13 BACTERIOPHAGE LIFE CYCLE Submitted by Steven William Smeal Department of Chemical and Biological Engineering In partial fulfillment of the requirements For
Cellular Gate Technology
 Cellular Gate Technology Thomas F. Knight, Jr. Gerald Jay Sussman MIT Artificial Intelligence Laboratory July 1997 A Scientist discovers that which exists. An Engineer creates that which never was. Theodore
Cellular Gate Technology Thomas F. Knight, Jr. Gerald Jay Sussman MIT Artificial Intelligence Laboratory July 1997 A Scientist discovers that which exists. An Engineer creates that which never was. Theodore
If Dna Has The Instructions For Building Proteins Why Is Mrna Needed
 If Dna Has The Instructions For Building Proteins Why Is Mrna Needed if a strand of DNA has the sequence CGGTATATC, then the complementary each strand of DNA contains the info needed to produce the complementary
If Dna Has The Instructions For Building Proteins Why Is Mrna Needed if a strand of DNA has the sequence CGGTATATC, then the complementary each strand of DNA contains the info needed to produce the complementary
M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION
 M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION Chapter 7 Microbial Genetics Lecture prepared by Mindy Miller-Kittrell, University of Tennessee, Knoxville The Structure and Replication
M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION Chapter 7 Microbial Genetics Lecture prepared by Mindy Miller-Kittrell, University of Tennessee, Knoxville The Structure and Replication
Developmental Biology BY1101 P. Murphy
 Developmental Biology BY1101 P. Murphy Lecture 7 Cellular differentiation and the regulation of gene expression. In this lecture we looked at two main questions: How is gene expression regulated? (revision
Developmental Biology BY1101 P. Murphy Lecture 7 Cellular differentiation and the regulation of gene expression. In this lecture we looked at two main questions: How is gene expression regulated? (revision
The Structure and Genetic Map of Lambda phage
 NPTEL Biotechnology - Systems Biology The Structure and Genetic Map of Lambda phage Dr. M. Vijayalakshmi School of Chemical and Biotechnology SASTRA University Joint Initiative of IITs and IISc Funded
NPTEL Biotechnology - Systems Biology The Structure and Genetic Map of Lambda phage Dr. M. Vijayalakshmi School of Chemical and Biotechnology SASTRA University Joint Initiative of IITs and IISc Funded
DNA Circuits for Analog Computing
 DNA Circuits for Analog Computing Tianqi Song Department of Computer Science Duke University 1 Outline Motivation What is DNA computing? Why are we interested in DNA computing? What has been done? Why
DNA Circuits for Analog Computing Tianqi Song Department of Computer Science Duke University 1 Outline Motivation What is DNA computing? Why are we interested in DNA computing? What has been done? Why
Genetics Lecture 21 Recombinant DNA
 Genetics Lecture 21 Recombinant DNA Recombinant DNA In 1971, a paper published by Kathleen Danna and Daniel Nathans marked the beginning of the recombinant DNA era. The paper described the isolation of
Genetics Lecture 21 Recombinant DNA Recombinant DNA In 1971, a paper published by Kathleen Danna and Daniel Nathans marked the beginning of the recombinant DNA era. The paper described the isolation of
Tackling Cancer Biology with Fundamental Physics
 The MIT Physical Science Oncology Center Tackling Cancer Biology with Fundamental Physics by Alexander van Oudenaarden 52 ) van oudenaarden mit physics annual 2010 s physics makes increasingly important
The MIT Physical Science Oncology Center Tackling Cancer Biology with Fundamental Physics by Alexander van Oudenaarden 52 ) van oudenaarden mit physics annual 2010 s physics makes increasingly important
Independent Study Guide The Blueprint of Life, from DNA to Protein (Chapter 7)
 Independent Study Guide The Blueprint of Life, from DNA to Protein (Chapter 7) I. General Principles (Chapter 7 introduction) a. Morse code distinct series of dots and dashes encode the 26 letters of the
Independent Study Guide The Blueprint of Life, from DNA to Protein (Chapter 7) I. General Principles (Chapter 7 introduction) a. Morse code distinct series of dots and dashes encode the 26 letters of the
Lecture 18. 2) Check to see whether the mutation is recessive and trans-acting (most will be).
 Lecture 18 In the preceding examples of bacterial gene regulation, we have used known regulatory mechanisms to see how mutations in different elements of the system would behave in dominance tests and
Lecture 18 In the preceding examples of bacterial gene regulation, we have used known regulatory mechanisms to see how mutations in different elements of the system would behave in dominance tests and
Genetics Lecture Notes Lectures 17 19
 Genetics Lecture Notes 7.03 2005 Lectures 17 19 Lecture 17 Gene Regulation We are now going to look at ways that genetics can be used to study gene regulation. The issue is how cells adjust the expression
Genetics Lecture Notes 7.03 2005 Lectures 17 19 Lecture 17 Gene Regulation We are now going to look at ways that genetics can be used to study gene regulation. The issue is how cells adjust the expression
Lecture Four. Molecular Approaches I: Nucleic Acids
 Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
Bio-inspired Models of Computation. An Introduction
 Bio-inspired Models of Computation An Introduction Introduction (1) Natural Computing is the study of models of computation inspired by the functioning of biological systems Natural Computing is not Bioinformatics
Bio-inspired Models of Computation An Introduction Introduction (1) Natural Computing is the study of models of computation inspired by the functioning of biological systems Natural Computing is not Bioinformatics
Molecular Genetics Student Objectives
 Molecular Genetics Student Objectives Exam 1: Enduring understanding 3.A: Heritable information provides for continuity of life. Essential knowledge 3.A.1: DNA, and in some cases RNA, is the primary source
Molecular Genetics Student Objectives Exam 1: Enduring understanding 3.A: Heritable information provides for continuity of life. Essential knowledge 3.A.1: DNA, and in some cases RNA, is the primary source
Chapter 8 DNA Recognition in Prokaryotes by Helix-Turn-Helix Motifs
 Chapter 8 DNA Recognition in Prokaryotes by Helix-Turn-Helix Motifs 1. Helix-turn-helix proteins 2. Zinc finger proteins 3. Leucine zipper proteins 4. Beta-scaffold factors 5. Others λ-repressor AND CRO
Chapter 8 DNA Recognition in Prokaryotes by Helix-Turn-Helix Motifs 1. Helix-turn-helix proteins 2. Zinc finger proteins 3. Leucine zipper proteins 4. Beta-scaffold factors 5. Others λ-repressor AND CRO
AP Biology Gene Expression/Biotechnology REVIEW
 AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
A PETRI NET MODEL FOR SIMULATION OF CONTAINER TERMINALS OPERATIONS
 Advanced OR and AI Methods in Transportation A PETRI NET MODEL FOR SIMULATION OF CONTAINER TERMINALS OPERATIONS Guido MAIONE 1, Michele OTTOMANELLI 1 Abstract. In this paper a model to simulate the operation
Advanced OR and AI Methods in Transportation A PETRI NET MODEL FOR SIMULATION OF CONTAINER TERMINALS OPERATIONS Guido MAIONE 1, Michele OTTOMANELLI 1 Abstract. In this paper a model to simulate the operation
Sequence Analysis Lab Protocol
 Sequence Analysis Lab Protocol You will need this handout of instructions The sequence of your plasmid from the ABI The Accession number for Lambda DNA J02459 The Accession number for puc 18 is L09136
Sequence Analysis Lab Protocol You will need this handout of instructions The sequence of your plasmid from the ABI The Accession number for Lambda DNA J02459 The Accession number for puc 18 is L09136
Prokaryotic Transcription
 Prokaryotic Transcription Contents 1 The Lactose Intolerance of Bacteria 2 The Lac Operon 3 Lac Operon Simulation 4 LacZ as a reporter gene The Lactose Intolerance of Bacteria The standard growth kinetics
Prokaryotic Transcription Contents 1 The Lactose Intolerance of Bacteria 2 The Lac Operon 3 Lac Operon Simulation 4 LacZ as a reporter gene The Lactose Intolerance of Bacteria The standard growth kinetics
REGULATION OF GENE EXPRESSION
 REGULATION OF GENE EXPRESSION Each cell of a living organism contains thousands of genes. But all genes do not function at a time. Genes function according to requirements of the cell. Genes control the
REGULATION OF GENE EXPRESSION Each cell of a living organism contains thousands of genes. But all genes do not function at a time. Genes function according to requirements of the cell. Genes control the
Metabolic Networks. Ulf Leser and Michael Weidlich
 Metabolic Networks Ulf Leser and Michael Weidlich This Lecture Introduction Systems biology & modelling Metabolism & metabolic networks Network reconstruction Strategy & workflow Mathematical representation
Metabolic Networks Ulf Leser and Michael Weidlich This Lecture Introduction Systems biology & modelling Metabolism & metabolic networks Network reconstruction Strategy & workflow Mathematical representation
Building blocks of a biochemical CPU based on DNA transcription logic
 Building blocks of a biochemical CPU based on DNA transcription logic Mario Lauria Dept of Computer Science and Engineering lauria@cis.ohio-state.edu Muthu M Pugalanthiran Dept. of Electrical Engineering
Building blocks of a biochemical CPU based on DNA transcription logic Mario Lauria Dept of Computer Science and Engineering lauria@cis.ohio-state.edu Muthu M Pugalanthiran Dept. of Electrical Engineering
Protein Synthesis. Lab Exercise 12. Introduction. Contents. Objectives
 Lab Exercise Protein Synthesis Contents Objectives 1 Introduction 1 Activity.1 Overview of Process 2 Activity.2 Transcription 2 Activity.3 Translation 3 Resutls Section 4 Introduction Having information
Lab Exercise Protein Synthesis Contents Objectives 1 Introduction 1 Activity.1 Overview of Process 2 Activity.2 Transcription 2 Activity.3 Translation 3 Resutls Section 4 Introduction Having information
Transcription in Eukaryotes
 Transcription in Eukaryotes Biology I Hayder A Giha Transcription Transcription is a DNA-directed synthesis of RNA, which is the first step in gene expression. Gene expression, is transformation of the
Transcription in Eukaryotes Biology I Hayder A Giha Transcription Transcription is a DNA-directed synthesis of RNA, which is the first step in gene expression. Gene expression, is transformation of the
Genetic Engineering for Biofuels Production
 Genetic Engineering for Biofuels Production WSE 573 Spring 2013 Greeley Beck INTRODUCTION Alternative transportation fuels are needed in the United States because of oil supply insecurity, oil price increases,
Genetic Engineering for Biofuels Production WSE 573 Spring 2013 Greeley Beck INTRODUCTION Alternative transportation fuels are needed in the United States because of oil supply insecurity, oil price increases,
A Parallel Implementation of the Modus Ponens Inference Rule in a DNA Strand-Displacement System
 A Parallel Implementation of the Modus Ponens Inference Rule in a DNA Strand-Displacement System Jack K. Horner P.O. Box 266 Los Alamos NM 87544 USA PDPTA 2013 Abstract Computation implemented in DNA reactions
A Parallel Implementation of the Modus Ponens Inference Rule in a DNA Strand-Displacement System Jack K. Horner P.O. Box 266 Los Alamos NM 87544 USA PDPTA 2013 Abstract Computation implemented in DNA reactions
Superposition and Synthetic Genetic Devices: Framework and Model System to Investigate Linearity in Escherichia coli
 Superposition and Synthetic Genetic Devices: Framework and Model System to Investigate Linearity in Escherichia coli Meghdad Hajimorad Electrical Engineering and Computer Sciences University of California
Superposition and Synthetic Genetic Devices: Framework and Model System to Investigate Linearity in Escherichia coli Meghdad Hajimorad Electrical Engineering and Computer Sciences University of California
encodes a sigma factor to modify the recognition of the E.coli RNA polymerase (Several other answers would also be acceptable for each phage)
 Name Student ID# Bacterial Genetics, BIO 4443/6443 Spring Semester 2001 Final Exam 1.) Different bacteriophage utilize different mechanisms to ensure that their own genes (and not host s genes) are transcribed
Name Student ID# Bacterial Genetics, BIO 4443/6443 Spring Semester 2001 Final Exam 1.) Different bacteriophage utilize different mechanisms to ensure that their own genes (and not host s genes) are transcribed
Solutions to Quiz II
 MIT Department of Biology 7.014 Introductory Biology, Spring 2005 Solutions to 7.014 Quiz II Class Average = 79 Median = 82 Grade Range % A 90-100 27 B 75-89 37 C 59 74 25 D 41 58 7 F 0 40 2 Question 1
MIT Department of Biology 7.014 Introductory Biology, Spring 2005 Solutions to 7.014 Quiz II Class Average = 79 Median = 82 Grade Range % A 90-100 27 B 75-89 37 C 59 74 25 D 41 58 7 F 0 40 2 Question 1
Modeling Stem Cell Induction Processes
 Filipe Grácio 1,2, Joaquim Cabral 1, Bruce Tidor 2,3,4 * 1 Institute for Biotechnology and Bioengineering (IBB), Centre for Biological and Chemical Engineering, Instituto Superior Técnico, Lisboa, Portugal,
Filipe Grácio 1,2, Joaquim Cabral 1, Bruce Tidor 2,3,4 * 1 Institute for Biotechnology and Bioengineering (IBB), Centre for Biological and Chemical Engineering, Instituto Superior Técnico, Lisboa, Portugal,
Regulation of gene expression. (Lehninger pg )
 Regulation of gene expression (Lehninger pg. 1072-1085) Today s lecture Gene expression Constitutive, inducible, repressible genes Specificity factors, activators, repressors Negative and positive gene
Regulation of gene expression (Lehninger pg. 1072-1085) Today s lecture Gene expression Constitutive, inducible, repressible genes Specificity factors, activators, repressors Negative and positive gene
A versatile framework for microbial engineering using synthetic noncoding
 FOCUS ON synthetic biology REVIEWS A versatile framework for microbial engineering using synthetic noncoding RNAs Lei S. Qi 1 3 and Adam P. Arkin 3 5 Abstract Synthetic non-coding RNAs have emerged as
FOCUS ON synthetic biology REVIEWS A versatile framework for microbial engineering using synthetic noncoding RNAs Lei S. Qi 1 3 and Adam P. Arkin 3 5 Abstract Synthetic non-coding RNAs have emerged as
Lecture 3 (FW) January 28, 2009 Cloning of DNA; PCR amplification Reading assignment: Cloning, ; ; 330 PCR, ; 329.
 Lecture 3 (FW) January 28, 2009 Cloning of DNA; PCR amplification Reading assignment: Cloning, 240-245; 286-87; 330 PCR, 270-274; 329. Take Home Lesson(s) from Lecture 2: 1. DNA is a double helix of complementary
Lecture 3 (FW) January 28, 2009 Cloning of DNA; PCR amplification Reading assignment: Cloning, 240-245; 286-87; 330 PCR, 270-274; 329. Take Home Lesson(s) from Lecture 2: 1. DNA is a double helix of complementary
Bayesian Networks as framework for data integration
 Bayesian Networks as framework for data integration Jun Zhu, Ph. D. Department of Genomics and Genetic Sciences Icahn Institute of Genomics and Multiscale Biology Icahn Medical School at Mount Sinai New
Bayesian Networks as framework for data integration Jun Zhu, Ph. D. Department of Genomics and Genetic Sciences Icahn Institute of Genomics and Multiscale Biology Icahn Medical School at Mount Sinai New
O C. 5 th C. 3 rd C. the national health museum
 Elements of Molecular Biology Cells Cells is a basic unit of all living organisms. It stores all information to replicate itself Nucleus, chromosomes, genes, All living things are made of cells Prokaryote,
Elements of Molecular Biology Cells Cells is a basic unit of all living organisms. It stores all information to replicate itself Nucleus, chromosomes, genes, All living things are made of cells Prokaryote,
Modeling of Protein Production Process by Finite Automata (FA)
 Modeling of Protein Production Process by Finite Automata (FA) ISSAC BARJIS 1, JOE W. YEOL 2, YEONG SOON RYU 3 Physics and Biomedical Sciences 1, Mechanical Engineering Technology 3 City University of
Modeling of Protein Production Process by Finite Automata (FA) ISSAC BARJIS 1, JOE W. YEOL 2, YEONG SOON RYU 3 Physics and Biomedical Sciences 1, Mechanical Engineering Technology 3 City University of
Patterns of Regulation from mrna and Protein Time Series
 Metabolic Engineering 2, 210217 (2000), available online at http:www.idealibrary.com on Patterns of Regulation from mrna and Protein Time Series Lingchong You and John Yin 1 Department of Chemical Engineering,
Metabolic Engineering 2, 210217 (2000), available online at http:www.idealibrary.com on Patterns of Regulation from mrna and Protein Time Series Lingchong You and John Yin 1 Department of Chemical Engineering,
CEng 713 Evolutionary Computation, Lecture Notes
 CEng 713 Evolutionary Computation, Lecture Notes Introduction to Evolutionary Computation Evolutionary Computation Elements of Evolution: Reproduction Random variation Competition Selection of contending
CEng 713 Evolutionary Computation, Lecture Notes Introduction to Evolutionary Computation Evolutionary Computation Elements of Evolution: Reproduction Random variation Competition Selection of contending
Name Class Date. Practice Test
 Name Class Date 12 DNA Practice Test Multiple Choice Write the letter that best answers the question or completes the statement on the line provided. 1. What do bacteriophages infect? a. mice. c. viruses.
Name Class Date 12 DNA Practice Test Multiple Choice Write the letter that best answers the question or completes the statement on the line provided. 1. What do bacteriophages infect? a. mice. c. viruses.
2054, Chap. 14, page 1
 2054, Chap. 14, page 1 I. Recombinant DNA technology (Chapter 14) A. recombinant DNA technology = collection of methods used to perform genetic engineering 1. genetic engineering = deliberate modification
2054, Chap. 14, page 1 I. Recombinant DNA technology (Chapter 14) A. recombinant DNA technology = collection of methods used to perform genetic engineering 1. genetic engineering = deliberate modification
Ch. 10 Notes DNA: Transcription and Translation
 Ch. 10 Notes DNA: Transcription and Translation GOALS Compare the structure of RNA with that of DNA Summarize the process of transcription Relate the role of codons to the sequence of amino acids that
Ch. 10 Notes DNA: Transcription and Translation GOALS Compare the structure of RNA with that of DNA Summarize the process of transcription Relate the role of codons to the sequence of amino acids that
DNA Transcription. Visualizing Transcription. The Transcription Process
 DNA Transcription By: Suzanne Clancy, Ph.D. 2008 Nature Education Citation: Clancy, S. (2008) DNA transcription. Nature Education 1(1) If DNA is a book, then how is it read? Learn more about the DNA transcription
DNA Transcription By: Suzanne Clancy, Ph.D. 2008 Nature Education Citation: Clancy, S. (2008) DNA transcription. Nature Education 1(1) If DNA is a book, then how is it read? Learn more about the DNA transcription
Green Fluorescent Protein (GFP) Purification. Hydrophobic Interaction Chromatography
 Green Fluorescent Protein (GFP) Purification Hydrophobic Interaction Chromatography What is the GFP gene? GFP is a green fluorescent protein that is normally found in jellyfish. It has been engineered
Green Fluorescent Protein (GFP) Purification Hydrophobic Interaction Chromatography What is the GFP gene? GFP is a green fluorescent protein that is normally found in jellyfish. It has been engineered
Rapporto di Ricerca CS
 UNIVERSITÀ CA FOSCARI DI VENEZIA Dipartimento di Informatica Technical Report Series in Computer Science Rapporto di Ricerca CS-2006-8 Novembre 2006 Andrea Marin Biological Pathways representation using
UNIVERSITÀ CA FOSCARI DI VENEZIA Dipartimento di Informatica Technical Report Series in Computer Science Rapporto di Ricerca CS-2006-8 Novembre 2006 Andrea Marin Biological Pathways representation using
Genetic drift. 1. The Nature of Genetic Drift
 Genetic drift. The Nature of Genetic Drift To date, we have assumed that populations are infinite in size. This assumption enabled us to easily calculate the expected frequencies of alleles and genotypes
Genetic drift. The Nature of Genetic Drift To date, we have assumed that populations are infinite in size. This assumption enabled us to easily calculate the expected frequencies of alleles and genotypes
DNA is the genetic material. DNA structure. Chapter 7: DNA Replication, Transcription & Translation; Mutations & Ames test
 DNA is the genetic material Chapter 7: DNA Replication, Transcription & Translation; Mutations & Ames test Dr. Amy Rogers Bio 139 General Microbiology Hereditary information is carried by DNA Griffith/Avery
DNA is the genetic material Chapter 7: DNA Replication, Transcription & Translation; Mutations & Ames test Dr. Amy Rogers Bio 139 General Microbiology Hereditary information is carried by DNA Griffith/Avery
Biochemistry study of the molecular basis of life
 Biochemistry : An Introduction Biochemistry study of the molecular basis of life n Study of the chemistry of living organisms Studies organic molecules & organic reactions in living organisms n Living
Biochemistry : An Introduction Biochemistry study of the molecular basis of life n Study of the chemistry of living organisms Studies organic molecules & organic reactions in living organisms n Living
2/23/16. Protein-Protein Interactions. Protein Interactions. Protein-Protein Interactions: The Interactome
 Protein-Protein Interactions Protein Interactions A Protein may interact with: Other proteins Nucleic Acids Small molecules Protein-Protein Interactions: The Interactome Experimental methods: Mass Spec,
Protein-Protein Interactions Protein Interactions A Protein may interact with: Other proteins Nucleic Acids Small molecules Protein-Protein Interactions: The Interactome Experimental methods: Mass Spec,
Programmed Cell Applications. Programming Cells
 The Device Physics of Cellular Logic Gates Ron Weiss Department of Electrical Engineering Princeton University SC-1 Programming Cells Environment actuators sensors Biochemical logic circuit.5µm Understand
The Device Physics of Cellular Logic Gates Ron Weiss Department of Electrical Engineering Princeton University SC-1 Programming Cells Environment actuators sensors Biochemical logic circuit.5µm Understand
Contents... vii. List of Figures... xii. List of Tables... xiv. Abbreviatons... xv. Summary... xvii. 1. Introduction In vitro evolution...
 vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
Introduction to Microarray Data Analysis and Gene Networks. Alvis Brazma European Bioinformatics Institute
 Introduction to Microarray Data Analysis and Gene Networks Alvis Brazma European Bioinformatics Institute A brief outline of this course What is gene expression, why it s important Microarrays and how
Introduction to Microarray Data Analysis and Gene Networks Alvis Brazma European Bioinformatics Institute A brief outline of this course What is gene expression, why it s important Microarrays and how
7 Gene Isolation and Analysis of Multiple
 Genetic Techniques for Biological Research Corinne A. Michels Copyright q 2002 John Wiley & Sons, Ltd ISBNs: 0-471-89921-6 (Hardback); 0-470-84662-3 (Electronic) 7 Gene Isolation and Analysis of Multiple
Genetic Techniques for Biological Research Corinne A. Michels Copyright q 2002 John Wiley & Sons, Ltd ISBNs: 0-471-89921-6 (Hardback); 0-470-84662-3 (Electronic) 7 Gene Isolation and Analysis of Multiple
Prokaryotic Transcription
 Prokaryotic Transcription Transcription Basics DNA is the genetic material Nucleic acid Capable of self-replication and synthesis of RNA RNA is the middle man Nucleic acid Structure and base sequence are
Prokaryotic Transcription Transcription Basics DNA is the genetic material Nucleic acid Capable of self-replication and synthesis of RNA RNA is the middle man Nucleic acid Structure and base sequence are
Answers to Module 1. An obligate aerobe is an organism that has an absolute requirement of oxygen for growth.
 Answers to Module 1 Short Answers 1) What is an obligate aerobe? An obligate aerobe is an organism that has an absolute requirement of oxygen for growth. What about facultative anaerobe? 2) Distinguish
Answers to Module 1 Short Answers 1) What is an obligate aerobe? An obligate aerobe is an organism that has an absolute requirement of oxygen for growth. What about facultative anaerobe? 2) Distinguish
Viruses 11/30/2015. Chapter 19. Key Concepts in Chapter 19
 Chapter 19 Viruses Dr. Wendy Sera Houston Community College Biology 1406 Key Concepts in Chapter 19 1. A virus consists of a nucleic acid surrounded by a protein coat. 2. Viruses replicate only in host
Chapter 19 Viruses Dr. Wendy Sera Houston Community College Biology 1406 Key Concepts in Chapter 19 1. A virus consists of a nucleic acid surrounded by a protein coat. 2. Viruses replicate only in host
Chapter 13 - Concept Mapping
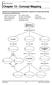 Chapter 13 - Concept Mapping Using the terms and phrases provided below, complete the concept map showing the discovery of DNA structure. amount of base pairs five-carbon sugar purine DNA polymerases Franklin
Chapter 13 - Concept Mapping Using the terms and phrases provided below, complete the concept map showing the discovery of DNA structure. amount of base pairs five-carbon sugar purine DNA polymerases Franklin
Fundamentals of Bioinformatics: computation, biology, computational biology
 Fundamentals of Bioinformatics: computation, biology, computational biology Vasilis J. Promponas Bioinformatics Research Laboratory Department of Biological Sciences University of Cyprus A short self-introduction
Fundamentals of Bioinformatics: computation, biology, computational biology Vasilis J. Promponas Bioinformatics Research Laboratory Department of Biological Sciences University of Cyprus A short self-introduction
Cell-free synthetic biology provides an experimental
 pubs.acs.org/synthbio An E. coli Cell-Free Expression Toolbox: Application to Synthetic Gene Circuits and Artificial Cells Jonghyeon Shin and Vincent Noireaux* School of Physics and Astronomy, University
pubs.acs.org/synthbio An E. coli Cell-Free Expression Toolbox: Application to Synthetic Gene Circuits and Artificial Cells Jonghyeon Shin and Vincent Noireaux* School of Physics and Astronomy, University
From the Quest for Fire to GATTACA: Systems and Synthetic Biology as the Next Wave of Technology Disruption in the Anthropocene
 From the Quest for Fire to GATTACA: Systems and Synthetic Biology as the Next Wave of Technology Disruption in the Anthropocene Dr. George Poste Chief Scientist, Complex Adaptive Systems Initiative and
From the Quest for Fire to GATTACA: Systems and Synthetic Biology as the Next Wave of Technology Disruption in the Anthropocene Dr. George Poste Chief Scientist, Complex Adaptive Systems Initiative and
Product Applications for the Sequence Analysis Collection
 Product Applications for the Sequence Analysis Collection Pipeline Pilot Contents Introduction... 1 Pipeline Pilot and Bioinformatics... 2 Sequence Searching with Profile HMM...2 Integrating Data in a
Product Applications for the Sequence Analysis Collection Pipeline Pilot Contents Introduction... 1 Pipeline Pilot and Bioinformatics... 2 Sequence Searching with Profile HMM...2 Integrating Data in a
The Integrated Biomedical Sciences Graduate Program
 The Integrated Biomedical Sciences Graduate Program at the university of notre dame Cutting-edge biomedical research and training that transcends traditional departmental and disciplinary boundaries to
The Integrated Biomedical Sciences Graduate Program at the university of notre dame Cutting-edge biomedical research and training that transcends traditional departmental and disciplinary boundaries to
2012 GENERAL [5 points]
![2012 GENERAL [5 points] 2012 GENERAL [5 points]](/thumbs/75/72452386.jpg) GENERAL [5 points] 2012 Mark all processes that are part of the 'standard dogma of molecular' [ ] DNA replication [ ] transcription [ ] translation [ ] reverse transposition [ ] DNA restriction [ ] DNA
GENERAL [5 points] 2012 Mark all processes that are part of the 'standard dogma of molecular' [ ] DNA replication [ ] transcription [ ] translation [ ] reverse transposition [ ] DNA restriction [ ] DNA
8/21/2014. From Gene to Protein
 From Gene to Protein Chapter 17 Objectives Describe the contributions made by Garrod, Beadle, and Tatum to our understanding of the relationship between genes and enzymes Briefly explain how information
From Gene to Protein Chapter 17 Objectives Describe the contributions made by Garrod, Beadle, and Tatum to our understanding of the relationship between genes and enzymes Briefly explain how information
Bio-Reagent Services. Custom Gene Services. Gateway to Smooth Molecular Biology! Your Innovation Partner in Drug Discovery!
 Bio-Reagent Services Custom Gene Services Gateway to Smooth Molecular Biology! Gene Synthesis Mutagenesis Mutant Libraries Plasmid Preparation sirna and mirna Services Large-scale DNA Sequencing GenPool
Bio-Reagent Services Custom Gene Services Gateway to Smooth Molecular Biology! Gene Synthesis Mutagenesis Mutant Libraries Plasmid Preparation sirna and mirna Services Large-scale DNA Sequencing GenPool
Introducing Bioinformatics Concepts in CS1
 Introducing Bioinformatics Concepts in CS1 Stuart Hansen Computer Science Department University of Wisconsin - Parkside hansen@cs.uwp.edu Erica Eddy Computer Science Department University of Wisconsin
Introducing Bioinformatics Concepts in CS1 Stuart Hansen Computer Science Department University of Wisconsin - Parkside hansen@cs.uwp.edu Erica Eddy Computer Science Department University of Wisconsin
MOLECULAR BIOLOGY OF EUKARYOTES 2016 SYLLABUS
 03-442 Lectures: MWF 9:30-10:20 a.m. Doherty Hall 2105 03-742 Advanced Discussion Section: Time and place to be announced Probably Mon 4-6 p.m. or 6-8p.m.? Once we establish who is taking the advanced
03-442 Lectures: MWF 9:30-10:20 a.m. Doherty Hall 2105 03-742 Advanced Discussion Section: Time and place to be announced Probably Mon 4-6 p.m. or 6-8p.m.? Once we establish who is taking the advanced
Generative Models for Networks and Applications to E-Commerce
 Generative Models for Networks and Applications to E-Commerce Patrick J. Wolfe (with David C. Parkes and R. Kang-Xing Jin) Division of Engineering and Applied Sciences Department of Statistics Harvard
Generative Models for Networks and Applications to E-Commerce Patrick J. Wolfe (with David C. Parkes and R. Kang-Xing Jin) Division of Engineering and Applied Sciences Department of Statistics Harvard
Bi 8 Lecture 7. Ellen Rothenberg 26 January Reading: Ch. 3, pp ; panel 3-1
 Bi 8 Lecture 7 PROTEIN STRUCTURE, Functional analysis, and evolution Ellen Rothenberg 26 January 2016 Reading: Ch. 3, pp. 109-134; panel 3-1 (end with free amine) aromatic, hydrophobic small, hydrophilic
Bi 8 Lecture 7 PROTEIN STRUCTURE, Functional analysis, and evolution Ellen Rothenberg 26 January 2016 Reading: Ch. 3, pp. 109-134; panel 3-1 (end with free amine) aromatic, hydrophobic small, hydrophilic
Year III Pharm.D Dr. V. Chitra
 Year III Pharm.D Dr. V. Chitra 1 Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Only one strand of DNA serves
Year III Pharm.D Dr. V. Chitra 1 Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Only one strand of DNA serves
Identifying Signaling Pathways. BMI/CS 776 Spring 2016 Anthony Gitter
 Identifying Signaling Pathways BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2016 Anthony Gitter gitter@biostat.wisc.edu Goals for lecture Challenges of integrating high-throughput assays Connecting relevant
Identifying Signaling Pathways BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2016 Anthony Gitter gitter@biostat.wisc.edu Goals for lecture Challenges of integrating high-throughput assays Connecting relevant
The Genetic Code and Transcription. Chapter 12 Honors Genetics Ms. Susan Chabot
 The Genetic Code and Transcription Chapter 12 Honors Genetics Ms. Susan Chabot TRANSCRIPTION Copy SAME language DNA to RNA Nucleic Acid to Nucleic Acid TRANSLATION Copy DIFFERENT language RNA to Amino
The Genetic Code and Transcription Chapter 12 Honors Genetics Ms. Susan Chabot TRANSCRIPTION Copy SAME language DNA to RNA Nucleic Acid to Nucleic Acid TRANSLATION Copy DIFFERENT language RNA to Amino
Hierarchical Probabilistic Models for Operational-Level Course of Action Development
 Hierarchical Probabilistic Models for Operational-Level Course of Action Development Lucia Falzon, Lin Zhang and Mike Davies Information Technology Division Defence Science and Technology Organisation
Hierarchical Probabilistic Models for Operational-Level Course of Action Development Lucia Falzon, Lin Zhang and Mike Davies Information Technology Division Defence Science and Technology Organisation
Modeling cell fate specification during early embryonic development in mouse
 Université Libre de Bruxelles Modeling cell fate specification during early embryonic development in mouse Didier Gonze Unité de Chronobiologie Théorique Faculté des Sciences Université Libre de Bruxelles
Université Libre de Bruxelles Modeling cell fate specification during early embryonic development in mouse Didier Gonze Unité de Chronobiologie Théorique Faculté des Sciences Université Libre de Bruxelles
Genetic Engineering & Recombinant DNA
 Genetic Engineering & Recombinant DNA Chapter 10 Copyright The McGraw-Hill Companies, Inc) Permission required for reproduction or display. Applications of Genetic Engineering Basic science vs. Applied
Genetic Engineering & Recombinant DNA Chapter 10 Copyright The McGraw-Hill Companies, Inc) Permission required for reproduction or display. Applications of Genetic Engineering Basic science vs. Applied
Computational Biology I LSM5191
 Computational Biology I LSM5191 Lecture 5 Notes: Genetic manipulation & Molecular Biology techniques Broad Overview of: Enzymatic tools in Molecular Biology Gel electrophoresis Restriction mapping DNA
Computational Biology I LSM5191 Lecture 5 Notes: Genetic manipulation & Molecular Biology techniques Broad Overview of: Enzymatic tools in Molecular Biology Gel electrophoresis Restriction mapping DNA
Recombinant DNA Technology. The Role of Recombinant DNA Technology in Biotechnology. yeast. Biotechnology. Recombinant DNA technology.
 PowerPoint Lecture Presentations prepared by Mindy Miller-Kittrell, North Carolina State University C H A P T E R 8 Recombinant DNA Technology The Role of Recombinant DNA Technology in Biotechnology Biotechnology?
PowerPoint Lecture Presentations prepared by Mindy Miller-Kittrell, North Carolina State University C H A P T E R 8 Recombinant DNA Technology The Role of Recombinant DNA Technology in Biotechnology Biotechnology?
How Do You Clone a Gene?
 S-20 Edvo-Kit #S-20 How Do You Clone a Gene? Experiment Objective: The objective of this experiment is to gain an understanding of the structure of DNA, a genetically engineered clone, and how genes are
S-20 Edvo-Kit #S-20 How Do You Clone a Gene? Experiment Objective: The objective of this experiment is to gain an understanding of the structure of DNA, a genetically engineered clone, and how genes are
Neural Networks and Applications in Bioinformatics. Yuzhen Ye School of Informatics and Computing, Indiana University
 Neural Networks and Applications in Bioinformatics Yuzhen Ye School of Informatics and Computing, Indiana University Contents Biological problem: promoter modeling Basics of neural networks Perceptrons
Neural Networks and Applications in Bioinformatics Yuzhen Ye School of Informatics and Computing, Indiana University Contents Biological problem: promoter modeling Basics of neural networks Perceptrons
RNA Structure Prediction and Comparison. RNA Biology Background
 RN Structure Prediction and omparison Session 1 RN Biology Background Robert iegerich Faculty of Technology robert@techfak.ni-bielefeld.de October 13, 2013 Robert iegerich Overview of lecture topics The
RN Structure Prediction and omparison Session 1 RN Biology Background Robert iegerich Faculty of Technology robert@techfak.ni-bielefeld.de October 13, 2013 Robert iegerich Overview of lecture topics The
