Material and Methods Bacterial strains. Construction of C4750 strain derivatives. Mutagenesis assays.
|
|
|
- Lewis Jennings
- 6 years ago
- Views:
Transcription
1 Material and Methods Bacterial strains. 787 E. coli strains have been used in this study: (i-iii) 547 were isolated from fecal samples, (iii and iv) 177 were pathogenic isolates, (v) 62 were isolated from secondary environments, and the MG1655 E. coli K12 laboratory strain was included. (i) 298 commensal strains sampled mostly from wild mammals and birds were collected in: Mexico (10 from reptiles, 49 from birds and 198 from mammals), Venezuela (2 from mammals), Antarctic (2 from birds and 10 from mammals) and Australia (27 from mammals). (ii) 188 human commensals were collected in Croatia (61), France (70) and Mali (57). (iii) 61 commensal strains (29 human and 32 animal) and 11 urinary tract infections (UTI) strains from the E. coli reference (ECOR) collection have been studied (1). (iv) 166 pathogenic strains were isolated from humans in France, 80 during the course of extra-intestinal (UTI) and intra-intestinal [13 enteroinvasive E. coli (EIEC) and 73 Shigella] infections. (v) 62 strains were collected from secondary environments in Mexico (15 air, 14 mud, and 33 water). Construction of C4750 strain derivatives. C4750 is a human commensal strain isolated in Croatia. It belongs to B1 E. coli phylogenetic group (2). This strain was used as a model for genetic analysis because it has strong MAC phenotype (see Table 2 in an article). C4750 mutant derivatives were constructed by P1-mediated transduction of the following alleles: rpos359::tn10 and hfq-1::ωcm R (provided by M. Winkler, University of Texas, Huston, Texas, USA); rssb::cm R (provided by Henge-Aaronis, Free University, Berlin Germany); cyaa ilv::kan R, lexa1(ind - ) malb::tn9 and pola1 zgj::tn10 (provided by R. D Ari, Jacques Monod Institute, Paris, France); crp::cm R (provided by J. Blazquez, Hospital Ramon y Cajal, Madrid, Spain); reca306 srl::tn10 and polb 1::ΩSm R -Sp R (provided by D. Touati, Jacques Monod Institute, Paris, France); uvra::cm R (provided by N. Goosen, Leiden University, Leiden, Netherlands); recb268::tn10 (provided by R.G. Lloyd, University of Nottingham, Nottingham, UK); and muts201::tn5 (our strain collection). Plasmids used in this study were: pacyc184 and its derivatives pmq341 and pmq339 carrying muts and mutl genes, respectively (provided by M.G. Marinus, University of Massachusetts Medical School, Worcester, MA, USA). Mutagenesis assays. The frequency of mutation to antibiotic resistance was measured by using a modification of a previously described method (3) to 10 3 cells from an overnight culture were inoculated on nitro-cellulose filters (NC-45, Schleicher and Schuell) laid on fresh
2 869 plates (NaCl 5g/l, Bactotryptone 10g/l, yeast extract 5g/l, agar 15g/l). Each plate contained 3 filters [instead of one per plate as in (3)]. Plates were incubated at 37 for 1 to 7 days. Anaerobic growth conditions were created in jars using «GENbox anaer» generator (BioMérieux). Bacterial colonies were resuspended in 1 ml of 869 and incubated for 1 hour at 37 to allow for antibiotic resistance expression. Appropriate dilutions were then spread on 869 and 869 antibiotic plates. The antibiotic resistant mutants were scored after 24 h [instead of 48 h as in (3)] of incubation at 37 C. The frequency of mutation was calculated from at least 3 independent experiments, each with three independent cultures. The antibiotics were added as follows: 5-fluoro-uracil 5 µg/ml, mecillinam 10 µg/ml, nalidixic acid 40 µg/ml, phosphomycin 30 µg/ml, rifampicin 100 µg/ml and streptomycin 100 µg/ml. We have also used eleven lacz alleles that allow detection of all base substitutions and five frameshifts (4, 5). A set of C4750 strains mutants were constructed by their co-transduction of lacz and laci42::tn10 alleles (provided by D. Touati, Jacques Monod Institute, Paris, France) from eleven F episomes to C4750 chromosome. The rich medium used in our experiments contained traces of lactose or a lactose-like carbon source, which rendered exact measurements of mutagenesis impossible. Therefore, we measured mutagenesis in colonies that have been incubated seven days on agarose plates (without addition of any other substance) as follows cells from overnight cultures were washed three times in 10-2 M MgSO 4, inoculated on nitrocellulose filters (NC-45, Schleicher and Schuell) and laid on fresh 1.5% agarose plates. Plates were incubated at 37 for 1 to 7 days and mutants scored on M9 lactose medium (after 48h of incubation at 37 ). In parallel, dilutions were spread on 869 and 869 rifampicin plates. Sequencing C4750 rpob Rif R alleles. The part of rpob gene, known to contain almost all rpob mutations responsible for the rifampicin resistance (6), was amplified using PCR. The PCR primer sequences were: 1525-up: GGC GAT CTG GAT ACC CTG ATG C and 2198-down: CGG AGT CAA CGG CAA CAG CAC. Sequencing was performed by the «Centre d Etude du Polymorphisme Humain», Paris, using Big Dye Terminator Cycle Sequencing Ready Reaction kit with AmpliTaq DNA polymerase, FS (Perkin-Elmer, Applied Biosystems Division) and analyzed in an automated ABI PRISM 377 XL DNA sequencer (Perkin-Elmer, Applied Biosystems Division) following the manufacturer s instructions. Competition between different C4750 rpob alleles. Four different Rif R mutants (isolated on D1), each having a G:C->A:T mutation in a different position in the rpob gene, were
3 used in competition experiments that have been performed as follows: The «instant colonies» having about 10 9 cells, with 1/10 2, 1/10 3 and 1/10 4 Rif R /Rif S ratios, have been reconstructed. After seven days of aging in colonies, none of the Rif R mutant populations exceeded the initial input frequency. Simulation of the selection of stress-induced mutator alleles. To study the selection acting on alleles modifying the mutation rate under conditions encountered by bacteria in aging colonies, we have used modified individual-based model as previously described (7). We studied the response of clonal bacterial population (of 10 9 cells) to directional selection. The adaptive peak was characterized by 17 beneficial mutations (8, 5, 2 and 1 mutations of fitness effect 2%, 2.5%, 5% and 10% appearing with frequencies 4.34, 4.9, 4 and 2 x 10-9, respectively) following an exponential distribution with a mean mutation rate of 4 x 10-9 and a mean fitness effect of 3% [similar to the one estimated by Gerrish and Lenski (8)]. The rate of lethal mutations was fixed to 10-5 and that of deleterious mutations to 1.7 x 10-4 (with a 1% effect on fitness per deleterious mutation) (9). All fitness effects were multiplicative. The population grew exponentially from a single cell to 10 9 cells. On the average, every 30 generations, the population was submitted to a stress for a period of (on the average) 10 generations. During this phase of stress, the population size decreased by 0.05% for every generation and inducible mutator alleles were induced leading to an increased mutation rate. Using this model we studied the time needed for the population to reach the adaptive peak and the probability of fixation of a constitutive or inducible mutator allele emerging at a frequency of 5 x Because mutations that would improve survival to stress would inevitably increase the selection of inducible mutators, to be conservative, we chose not to take them into account in our model. Mutations that would specifically improve survival to general stresses (e.g., starvation) may sometimes exist in nature. However, it is likely that most of such mutations have already been fixed in bacterial genome, which have regularly encountered starvation for millions of years. If some of these mutations are not fixed, it should be due to some tradeoff, (i.e., these mutations being counter-selected during non-stressful conditions) and therefore they would not contribute to the long-term adaptation considered here. Supplementary Data
4 MAC conferring resistance to different antibiotics. All strains have been tested for their capacity to generate mutations conferring resistance to rifampicin. In addition, 10 natural isolates were randomly chosen among strong MAC mutators (based on test with rifampicin), and mutations conferring resistance to 5-fluorouracil, mecillinam, phosphomycin, streptomycine and nalidixic acid was measured in aging colonies. None responded to the same level as for rifampicin, but the increase was significant for 5-fluorouracil (D7/D1=3.4; Mann-Whitney: p=0.004), mecillinam (D7/D1=1.9; Mann-Whitney: p=0.01) and phosphomycin (D7/D1=4.8, Mann-Whitney: p=0.009). No increase between D1 and D7 was observed when mutagenesis to streptomycin and nalidixic acid resistance was measured. The frequency of mutants resistant to those two antibiotics even decreased at D7 relative to D1 for most of the natural isolates, suggesting that the majority of mutations conferring resistance to nalidixic acid and streptomycin (residing in gyrb and rpsl genes, respectively), may be deleterious or lethal to bacteria in aging colonies. laci papillation assay. The appearance of mutants was also monitored on minimal media plates using a laci papillation assay (10). Out of 392 randomly chosen natural isolates, 29% showed a laci-papillating phenotype. The increase in MAC to Rif R (on rich-media plates as described above) was 16.2 and 5.2-fold for laci papillators and non-papillators, respectively (Mann-Whitney p=0.0003). Thus, there was a positive correlation between MACs in the two mutation detection systems (in rpob and laci genes) under two different experimental conditions. Growth and survival of the C4750 derivatives in aging colonies. D7/D1 ratios of the median number of viable cells/colony for C4750 derivatives were as follows: 1.7 for parental strain, 0.6 for rpos, 0.7 for hfq, 1.2 for rssb, 0.5 for cyaa, 2.6 for crp,1.7 for reca, 0.6 for lexa1, 2.2 for polb, 0.3 for uvra, 3.6 for pola, 4.2 for recb, 1.0 for muts, 1.7 for parental strain carrying pacyc184 plasmid, 1.7 for parental strain carrying muts + plasmid and 1.3 for parental strain carrying mutl + plasmid. These data were obtained from at least 3 independent experiments (each with three independent cultures) for each genotype. The results showed that there is no clear correlation between the change of the number of viable cells/colony and the MAC (see Table 2 in an article). Selection of stress-inducible and constitutive mutator alleles. The simulations showed that the selection acting on a stress-inducible mutator allele (increasing X fold the mutation rate under stress) is only a little bit less efficient than the one acting on a constitutive mutator allele of
5 the same averaged mutagenic effect (X ) [X = K X+(1-K); K being the fraction of generations spent under stress]. This small difference is not due to the net production of deleterious mutations, because both mutator mechanisms generate the same amount of deleterious mutations in the long run (data not shown). However, during the strong burst of mutagenesis, beneficial mutations are more likely to co-occur with deleterious mutations (data not shown) and therefore to be lost. Hence, stress inducible mutators will produce less selectable beneficial mutations than their constitutive equivalents. The difference in selection of two types of mutator alleles may also be the consequence of the frequency of stresses. We observed that the selection of stress inducible mutators is less efficient when stresses are not frequent (data not shown). Since over-production of mutations in stress-inducible mutator genotype is conditional on the presence of stress, the increase in the generation of beneficial mutations by stress-inducible mutators, will be delayed compared to constitutive mutator. References 1. H. Ochman, R. K. Selander, J. Bacteriol. 157, (1984). 2. P. Duriez et al., Microbiology 147, (2001). 3. F. Taddei, I. Matic, M. Radman, Proc. Natl. Acad. Sci. USA 92, (1995). 4. C. G. Cupples, M. Cabrera, C. Cruz, J. H. Miller, Genetics 125, (1990). 5. C. G. Cupples, J. H. Miller, Proc. Natl. Acad. Sci. USA 86, (1989). 6. D. Jin, C. Gross, J. Mol. Biol. 202, (1988). 7. O. Tenaillon, B. Toupance, H. Le Nagard, F. Taddei, B. Godelle, Genetics 152, (1999). 8. P. J. Gerrish, R. E. Lenski, Genetica , (1998). 9. T. T. Kibota, M. Lynch, Nature 381, (1996). 10. Y. Nghiem, M. Cabrera, C. G. Cupples, J. H. Miller, Proc. Natl. Acad. Sci. USA 85, (1988).
The plasmid shown to the right has an oriv and orit at the positions indicated, and is known to replicate bidirectionally.
 Name Microbial Genetics, BIO 410/510 2008 Exam II The plasmid shown to the right has an oriv and orit at the positions indicated, and is known to replicate bidirectionally. 1.) Indicate where replication
Name Microbial Genetics, BIO 410/510 2008 Exam II The plasmid shown to the right has an oriv and orit at the positions indicated, and is known to replicate bidirectionally. 1.) Indicate where replication
Lac Operon contains three structural genes and is controlled by the lac repressor: (1) LacY protein transports lactose into the cell.
 Regulation of gene expression a. Expression of most genes can be turned off and on, usually by controlling the initiation of transcription. b. Lactose degradation in E. coli (Negative Control) Lac Operon
Regulation of gene expression a. Expression of most genes can be turned off and on, usually by controlling the initiation of transcription. b. Lactose degradation in E. coli (Negative Control) Lac Operon
SUPPLEMENTARY MATERIALS AND METHODS. E. coli strains, plasmids, and growth conditions. Escherichia coli strain P90C (1)
 SUPPLEMENTARY MATERIALS AND METHODS E. coli strains, plasmids, and growth conditions. Escherichia coli strain P90C (1) dinb::kan (lab stock) derivative was used as wild-type. MG1655 alka tag dinb (2) is
SUPPLEMENTARY MATERIALS AND METHODS E. coli strains, plasmids, and growth conditions. Escherichia coli strain P90C (1) dinb::kan (lab stock) derivative was used as wild-type. MG1655 alka tag dinb (2) is
# ml too many to count # ml 161/173 # ml 4/1
 Biol 322 Fall 2012 Study Sheet for Quiz #2 Quiz #2 is scheduled for Thursday Nov 10 th and will be worth 30-40 pts. This quiz will cover: Mutagenesis Lab: Parts 1 & 2 Bacterial Genetics: F X F- Cross All
Biol 322 Fall 2012 Study Sheet for Quiz #2 Quiz #2 is scheduled for Thursday Nov 10 th and will be worth 30-40 pts. This quiz will cover: Mutagenesis Lab: Parts 1 & 2 Bacterial Genetics: F X F- Cross All
5.) Name and describe one gene product in E.coli that is associated with performing each step in the recombination process. (6pts)
 Student ID# Bacterial Genetics, BIO 4443/6443 Spring Semester 2001 Exam II 1.) What is the primary difference between conjugative plasmids and mobilizable plasmids? What genes are typically found on conjugative
Student ID# Bacterial Genetics, BIO 4443/6443 Spring Semester 2001 Exam II 1.) What is the primary difference between conjugative plasmids and mobilizable plasmids? What genes are typically found on conjugative
The Mosaic Nature of Genomes
 The Mosaic Nature of Genomes n DNA sequence is not static Mutations of single bases Large deletions Large insertions of sequence n Transferred from other species n New functions useful in particular situations
The Mosaic Nature of Genomes n DNA sequence is not static Mutations of single bases Large deletions Large insertions of sequence n Transferred from other species n New functions useful in particular situations
Section 3. Synthetic combinations of various. termination defective rho, nusg and. nusa mutants: Spectrum of efficiency of
 Section 3 Synthetic combinations of various termination defective rho, nusg and nusa mutants: Spectrum of efficiency of Rho-dependent transcription termination Introduction As described in Sections 1 and
Section 3 Synthetic combinations of various termination defective rho, nusg and nusa mutants: Spectrum of efficiency of Rho-dependent transcription termination Introduction As described in Sections 1 and
October 20th, Bacteria are not Lamarckian
 October 20th, 1993 Bacteria are not Lamarckian Antoine Danchin Institut Pasteur Unité de Régulation de l'expression Génétique Département de Biochimie et Génétique Moléculaire 28 rue du Dr Roux - 75724
October 20th, 1993 Bacteria are not Lamarckian Antoine Danchin Institut Pasteur Unité de Régulation de l'expression Génétique Département de Biochimie et Génétique Moléculaire 28 rue du Dr Roux - 75724
Confirming the Phenotypes of E. coli Strains
 Confirming the Phenotypes of E. coli Strains INTRODUCTION Before undertaking any experiments, we need to confirm that the phenotypes of the E. coli strains we intend to use in the planned experiments correspond
Confirming the Phenotypes of E. coli Strains INTRODUCTION Before undertaking any experiments, we need to confirm that the phenotypes of the E. coli strains we intend to use in the planned experiments correspond
What determines if a mutation is deleterious, neutral, or beneficial?
 BIO 184 - PAL Problem Set Lecture 6 (Brooker Chapter 18) Mutations Section A. Types of mutations Define and give an example the following terms: allele; phenotype; genotype; Define and give an example
BIO 184 - PAL Problem Set Lecture 6 (Brooker Chapter 18) Mutations Section A. Types of mutations Define and give an example the following terms: allele; phenotype; genotype; Define and give an example
Inside the Burch Lab: E. Coli and Triclosan Resistance. By: Pamela Lammonds
 Inside the Burch Lab: E. Coli and Triclosan Resistance By: Pamela Lammonds Purpose and Goals of Research Concerns over infectious disease have risen in the past few years. In response to this concern,
Inside the Burch Lab: E. Coli and Triclosan Resistance By: Pamela Lammonds Purpose and Goals of Research Concerns over infectious disease have risen in the past few years. In response to this concern,
Generation of Non-typeable Haemophilus influenzae Directed Gene Deletion Mutants Jeroen D. Langereis *
 Generation of Non-typeable Haemophilus influenzae Directed Gene Deletion Mutants Jeroen D. Langereis * Laboratory of Pediatric Infectious Diseases, Department of Pediatrics and Laboratory of Medical Immunology,
Generation of Non-typeable Haemophilus influenzae Directed Gene Deletion Mutants Jeroen D. Langereis * Laboratory of Pediatric Infectious Diseases, Department of Pediatrics and Laboratory of Medical Immunology,
Materials and methods. by University of Washington Yeast Resource Center) from several promoters, including
 Supporting online material for Elowitz et al. report Materials and methods Strains and plasmids. Plasmids expressing CFP or YFP (wild-type codons, developed by University of Washington Yeast Resource Center)
Supporting online material for Elowitz et al. report Materials and methods Strains and plasmids. Plasmids expressing CFP or YFP (wild-type codons, developed by University of Washington Yeast Resource Center)
Supplementary figures
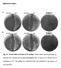 Supplementary figures A ΔrelAΔlon ΔrelA ΔrelAΔlon Δlon ppgpp 0 Δlon B ΔrelAΔlon ΔrelA ΔrelAΔlon Δlon ppgpp 0 Δlon Fig. S1. Growth defect of strains in LB medium. Strains whose relevant genotypes are indicated
Supplementary figures A ΔrelAΔlon ΔrelA ΔrelAΔlon Δlon ppgpp 0 Δlon B ΔrelAΔlon ΔrelA ΔrelAΔlon Δlon ppgpp 0 Δlon Fig. S1. Growth defect of strains in LB medium. Strains whose relevant genotypes are indicated
Biology 322 Fall 2010 Transfer of genetic information in the bacterium Escherichia coli: Part I
 Biology 322 Fall 2010 Transfer of genetic information in the bacterium Escherichia coli: Part I REQUIRED Reading Assignments: Superbugs on the Hoof http://fire.biol.wwu.edu/trent/trent/superbugs.pdf Triple
Biology 322 Fall 2010 Transfer of genetic information in the bacterium Escherichia coli: Part I REQUIRED Reading Assignments: Superbugs on the Hoof http://fire.biol.wwu.edu/trent/trent/superbugs.pdf Triple
Advantages of genetic analysis in bacteria/phage. Selections vs Screens. Mutations. Mutations lecture: March 4, 2009
 Mutations lecture: March 4, 2009 1. Genetic analysis of bacteria: the whys, hows, and whats 2. Luria-Delbrück and beyond: we still care! 3. Analysis of essential genes! RNA pol, merodiploids and amber
Mutations lecture: March 4, 2009 1. Genetic analysis of bacteria: the whys, hows, and whats 2. Luria-Delbrück and beyond: we still care! 3. Analysis of essential genes! RNA pol, merodiploids and amber
encodes a sigma factor to modify the recognition of the E.coli RNA polymerase (Several other answers would also be acceptable for each phage)
 Name Student ID# Bacterial Genetics, BIO 4443/6443 Spring Semester 2001 Final Exam 1.) Different bacteriophage utilize different mechanisms to ensure that their own genes (and not host s genes) are transcribed
Name Student ID# Bacterial Genetics, BIO 4443/6443 Spring Semester 2001 Final Exam 1.) Different bacteriophage utilize different mechanisms to ensure that their own genes (and not host s genes) are transcribed
Unit 2: Metabolism and Survival Sub-Topic (2.7) Genetic Control of Metabolism (2.8) Ethical considerations in the use of microorganisms
 Unit 2: Metabolism and Survival Sub-Topic (2.7) Genetic Control of Metabolism (2.8) Ethical considerations in the use of microorganisms Duncanrig Secondary JHM&MHC 2015 Page 1 of 18 On completion of this
Unit 2: Metabolism and Survival Sub-Topic (2.7) Genetic Control of Metabolism (2.8) Ethical considerations in the use of microorganisms Duncanrig Secondary JHM&MHC 2015 Page 1 of 18 On completion of this
TD-P Revision 3.0 Creation Date: 6/9/2016 Revision Date: 8/16/2018
 Protocol TD-P Revision 3.0 Creation Date: 6/9/2016 Revision Date: 8/16/2018 Electrotransformation of Agrobacterium tumefaciens Modified from Methods in Molecular Biology Vol. 47 Introduction Agrobacterium
Protocol TD-P Revision 3.0 Creation Date: 6/9/2016 Revision Date: 8/16/2018 Electrotransformation of Agrobacterium tumefaciens Modified from Methods in Molecular Biology Vol. 47 Introduction Agrobacterium
HI-Control BL21(DE3) & HI-Control 10G Chemically Competent Cells
 HI-Control BL21(DE3) & HI-Control 10G Chemically Competent Cells FOR RESEARCH USE ONLY. NOT FOR HUMAN OR DIAGNOSTIC USE MA156 Rev.31OCT2016 Table of Contents Components & Storage Conditions... 3 HI-Control
HI-Control BL21(DE3) & HI-Control 10G Chemically Competent Cells FOR RESEARCH USE ONLY. NOT FOR HUMAN OR DIAGNOSTIC USE MA156 Rev.31OCT2016 Table of Contents Components & Storage Conditions... 3 HI-Control
Protocol CRISPR Genome Editing In Cell Lines
 Protocol CRISPR Genome Editing In Cell Lines Protocol 2: HDR donor plasmid applications (gene knockout, gene mutagenesis, gene tagging, Safe Harbor ORF knock-in) Notes: 1. sgrna validation: GeneCopoeia
Protocol CRISPR Genome Editing In Cell Lines Protocol 2: HDR donor plasmid applications (gene knockout, gene mutagenesis, gene tagging, Safe Harbor ORF knock-in) Notes: 1. sgrna validation: GeneCopoeia
Some types of Mutagenesis
 Mutagenesis What Is a Mutation? Genetic information is encoded by the sequence of the nucleotide bases in DNA of the gene. The four nucleotides are: adenine (A), thymine (T), guanine (G), and cytosine
Mutagenesis What Is a Mutation? Genetic information is encoded by the sequence of the nucleotide bases in DNA of the gene. The four nucleotides are: adenine (A), thymine (T), guanine (G), and cytosine
Biology 201 (Genetics) Exam #3 120 points 20 November Read the question carefully before answering. Think before you write.
 Name KEY Section Biology 201 (Genetics) Exam #3 120 points 20 November 2006 Read the question carefully before answering. Think before you write. You will have up to 50 minutes to take this exam. After
Name KEY Section Biology 201 (Genetics) Exam #3 120 points 20 November 2006 Read the question carefully before answering. Think before you write. You will have up to 50 minutes to take this exam. After
Chapter VI: Mutagenesis to Restore Chimera Function
 99 Chapter VI: Mutagenesis to Restore Chimera Function Introduction Identifying characteristics of functional and nonfunctional chimeras is one way to address the underlying reasons for why chimeric proteins
99 Chapter VI: Mutagenesis to Restore Chimera Function Introduction Identifying characteristics of functional and nonfunctional chimeras is one way to address the underlying reasons for why chimeric proteins
Supplementary Information. reduction in replication fidelity
 Supplementary Information β-lactam antibiotics promote bacterial mutagenesis via an RpoS-mediated reduction in replication fidelity Authors: A. Gutierrez 1, L. Laureti 1, S. Crussard 1, H. Abida 1, A.
Supplementary Information β-lactam antibiotics promote bacterial mutagenesis via an RpoS-mediated reduction in replication fidelity Authors: A. Gutierrez 1, L. Laureti 1, S. Crussard 1, H. Abida 1, A.
Designing and creating your gene knockout Background The rada gene was identified as a gene, that when mutated, caused cells to become hypersensitive
 Designing and creating your gene knockout Background The rada gene was identified as a gene, that when mutated, caused cells to become hypersensitive to ionizing radiation. However, why these mutants are
Designing and creating your gene knockout Background The rada gene was identified as a gene, that when mutated, caused cells to become hypersensitive to ionizing radiation. However, why these mutants are
Supplementary Information
 Supplementary Information Supplementary Figures Figure S1. Study of mgtl translation in vitro. (A) Detection of 5 LR RNA using wild-type and anti-sd (91-95) substituted templates in a transcription-translation
Supplementary Information Supplementary Figures Figure S1. Study of mgtl translation in vitro. (A) Detection of 5 LR RNA using wild-type and anti-sd (91-95) substituted templates in a transcription-translation
4 Mutant Hunts - To Select or to Screen (Perhaps Even by Brute Force)
 Genetic Techniques for Biological Research Corinne A. Michels Copyright q 2002 John Wiley & Sons, Ltd ISBNs: 0-471-89921-6 (Hardback); 0-470-84662-3 (Electronic) 4 Mutant Hunts - To Select or to Screen
Genetic Techniques for Biological Research Corinne A. Michels Copyright q 2002 John Wiley & Sons, Ltd ISBNs: 0-471-89921-6 (Hardback); 0-470-84662-3 (Electronic) 4 Mutant Hunts - To Select or to Screen
2014 Pearson Education, Inc. CH 8: Recombinant DNA Technology
 CH 8: Recombinant DNA Technology Biotechnology the use of microorganisms to make practical products Recombinant DNA = DNA from 2 different sources What is Recombinant DNA Technology? modifying genomes
CH 8: Recombinant DNA Technology Biotechnology the use of microorganisms to make practical products Recombinant DNA = DNA from 2 different sources What is Recombinant DNA Technology? modifying genomes
MCB 421 Exam #1 (A) Fall There are 9 questions and 1 supplement on last page. Answer all 9 questions. Be sure your name is on each page
 MCB 421 Exam #1 (A) Fall 2006 There are 9 questions and 1 supplement on last page. Answer all 9 questions. Be sure your name is on each page 1). (8 points) A Luria-Delbruck fluctuation test was done to
MCB 421 Exam #1 (A) Fall 2006 There are 9 questions and 1 supplement on last page. Answer all 9 questions. Be sure your name is on each page 1). (8 points) A Luria-Delbruck fluctuation test was done to
BACTERIAL GENETICS. How does the DNA in the bacterial cell replicate
 BACTERIAL GENETICS Bacterial genetics is the study of gene structure and function in bacteria. Genetics itself is concerned with determining the number, location, and character of the genes of an organism.
BACTERIAL GENETICS Bacterial genetics is the study of gene structure and function in bacteria. Genetics itself is concerned with determining the number, location, and character of the genes of an organism.
Regulation of enzyme synthesis
 Regulation of enzyme synthesis The lac operon is an example of an inducible operon - it is normally off, but when a molecule called an inducer is present, the operon turns on. The trp operon is an example
Regulation of enzyme synthesis The lac operon is an example of an inducible operon - it is normally off, but when a molecule called an inducer is present, the operon turns on. The trp operon is an example
Association of gyra mutation in Mycobacterium tuberculosis isolates with phenotypic ofloxacin resistance detected by resazurin microtiter assay
 Association of gyra mutation in Mycobacterium tuberculosis isolates with phenotypic ofloxacin resistance detected by resazurin microtiter assay Dr Asho Ali King Abdul Aziz University Jeddah, Saudi Arabia
Association of gyra mutation in Mycobacterium tuberculosis isolates with phenotypic ofloxacin resistance detected by resazurin microtiter assay Dr Asho Ali King Abdul Aziz University Jeddah, Saudi Arabia
Nature Biotechnology: doi: /nbt Supplementary Figure 1
 Supplementary Figure 1 Negative selection analysis of sgrnas targeting all Brd4 exons comparing day 2 to day 10 time points. Systematic evaluation of 64 Brd4 sgrnas in negative selection experiments, targeting
Supplementary Figure 1 Negative selection analysis of sgrnas targeting all Brd4 exons comparing day 2 to day 10 time points. Systematic evaluation of 64 Brd4 sgrnas in negative selection experiments, targeting
Supplemental Data. The Survival Advantage of Olfaction. in a Competitive Environment. Supplemental Experimental Procedures
 Supplemental Data The Survival Advantage of Olfaction in a Competitive Environment Kenta Asahina, Viktoryia Pavlenkovich, and Leslie B. Vosshall Supplemental Experimental Procedures Experimental animals
Supplemental Data The Survival Advantage of Olfaction in a Competitive Environment Kenta Asahina, Viktoryia Pavlenkovich, and Leslie B. Vosshall Supplemental Experimental Procedures Experimental animals
CH 8: Recombinant DNA Technology
 CH 8: Recombinant DNA Technology Biotechnology the use of microorganisms to make practical products Recombinant DNA = DNA from 2 different sources What is Recombinant DNA Technology? modifying genomes
CH 8: Recombinant DNA Technology Biotechnology the use of microorganisms to make practical products Recombinant DNA = DNA from 2 different sources What is Recombinant DNA Technology? modifying genomes
Protocols for cell lines using CRISPR/CAS
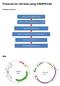 Protocols for cell lines using CRISPR/CAS Procedure overview Map Preparation of CRISPR/CAS plasmids Expression vectors for guide RNA (U6-gRNA) and Cas9 gene (CMV-p-Cas9) are ampicillin-resist ant and stable
Protocols for cell lines using CRISPR/CAS Procedure overview Map Preparation of CRISPR/CAS plasmids Expression vectors for guide RNA (U6-gRNA) and Cas9 gene (CMV-p-Cas9) are ampicillin-resist ant and stable
Module 6 Microbial Genetics. Chapter 8
 Module 6 Microbial Genetics Chapter 8 Structure and function of the genetic material Genetics science of o Study of what genes are, how they determine the characteristics of an organism, how they carry
Module 6 Microbial Genetics Chapter 8 Structure and function of the genetic material Genetics science of o Study of what genes are, how they determine the characteristics of an organism, how they carry
Game Plan. Lecture. Lab. Biofilms Review of basic genetics Bacterial gene structure Gene regulation Mutations
 Game Plan Lecture Biofilms Review of basic genetics Bacterial gene structure Gene regulation Mutations Lab Review serial dilutions and Calculate generation times Effects on growth: temp, ph, O2 Pre-labs
Game Plan Lecture Biofilms Review of basic genetics Bacterial gene structure Gene regulation Mutations Lab Review serial dilutions and Calculate generation times Effects on growth: temp, ph, O2 Pre-labs
Recombinant DNA Technology. The Role of Recombinant DNA Technology in Biotechnology. yeast. Biotechnology. Recombinant DNA technology.
 PowerPoint Lecture Presentations prepared by Mindy Miller-Kittrell, North Carolina State University C H A P T E R 8 Recombinant DNA Technology The Role of Recombinant DNA Technology in Biotechnology Biotechnology?
PowerPoint Lecture Presentations prepared by Mindy Miller-Kittrell, North Carolina State University C H A P T E R 8 Recombinant DNA Technology The Role of Recombinant DNA Technology in Biotechnology Biotechnology?
Genetics Lecture Notes Lectures 13 16
 Genetics Lecture Notes 7.03 2005 Lectures 13 16 Lecture 13 Transposable elements Transposons are usually from 10 3 to 10 4 base pairs in length, depending on the transposon type. The key property of transposons
Genetics Lecture Notes 7.03 2005 Lectures 13 16 Lecture 13 Transposable elements Transposons are usually from 10 3 to 10 4 base pairs in length, depending on the transposon type. The key property of transposons
Table S1. Bacterial strains (Related to Results and Experimental Procedures)
 Table S1. Bacterial strains (Related to Results and Experimental Procedures) Strain number Relevant genotype Source or reference 1045 AB1157 Graham Walker (Donnelly and Walker, 1989) 2458 3084 (MG1655)
Table S1. Bacterial strains (Related to Results and Experimental Procedures) Strain number Relevant genotype Source or reference 1045 AB1157 Graham Walker (Donnelly and Walker, 1989) 2458 3084 (MG1655)
Proofreading deficiency of Pol I increases the levels of spontaneous. rpob mutations in E. coli
 Published in final edited form as: Mutat Res. 2011 July 1; 712(1-2): 28 32. Proofreading deficiency of Pol I increases the levels of spontaneous rpob mutations in E. coli K. Makiela-Dzbenska a, P. Jonczyk
Published in final edited form as: Mutat Res. 2011 July 1; 712(1-2): 28 32. Proofreading deficiency of Pol I increases the levels of spontaneous rpob mutations in E. coli K. Makiela-Dzbenska a, P. Jonczyk
PCR analysis was performed to show the presence and the integrity of the var1csa and var-
 Supplementary information: Methods: Table S1: Primer Name Nucleotide sequence (5-3 ) DBL3-F tcc ccg cgg agt gaa aca tca tgt gac tg DBL3-R gac tag ttt ctt tca ata aat cac tcg c DBL5-F cgc cct agg tgc ttc
Supplementary information: Methods: Table S1: Primer Name Nucleotide sequence (5-3 ) DBL3-F tcc ccg cgg agt gaa aca tca tgt gac tg DBL3-R gac tag ttt ctt tca ata aat cac tcg c DBL5-F cgc cct agg tgc ttc
HOUR EXAM I BIOLOGY 422 FALL, In the spirit of the honor code, I pledge that I have neither given nor received help on this exam.
 Name First Last (Please Print) PID Number - HOUR EXAM I BIOLOGY 422 FALL, 2011 In the spirit of the honor code, I pledge that I have neither given nor received help on this exam. 1 Signature 2 3 4 5 6
Name First Last (Please Print) PID Number - HOUR EXAM I BIOLOGY 422 FALL, 2011 In the spirit of the honor code, I pledge that I have neither given nor received help on this exam. 1 Signature 2 3 4 5 6
FROM EXPERIMENTS IN BACTERIAL GENETICS AND GENE TECHNIQUE
 Uppsala 2001-04-01 REPORT FROM EXPERIMENTS IN BACTERIAL GENETICS AND GENE TECHNIQUE Laboratory assistants: Maria Jönsson Amera Gibreel Students: Contents ASSIGNMENT:... 3 INTRODUCTION:... 3 MATERIAL AND
Uppsala 2001-04-01 REPORT FROM EXPERIMENTS IN BACTERIAL GENETICS AND GENE TECHNIQUE Laboratory assistants: Maria Jönsson Amera Gibreel Students: Contents ASSIGNMENT:... 3 INTRODUCTION:... 3 MATERIAL AND
7.02 Microbial Genetics in Lab Quiz. Fall, September 27, 2001 ANSWER KEY
 7.02 Microbial Genetics in Lab Quiz Fall, 2001 September 27, 2001 ANSWER KEY This quiz contains 4 questions worth a total of 48 points. Be sure to write your name, Bench letter and Undergraduate TA s (UTA)
7.02 Microbial Genetics in Lab Quiz Fall, 2001 September 27, 2001 ANSWER KEY This quiz contains 4 questions worth a total of 48 points. Be sure to write your name, Bench letter and Undergraduate TA s (UTA)
A set of lacz mutations in Escherichia coli that allow rapid detection of each of the six base substitutions
 Proc. Natd. cad. Sci. US Vol. 86, pp. 5345-5349, July 1989 Biochemistry set of lacz mutations in Escherichia coli that allow rapid detection of each of the six base substitutions CLIRE. CUPPLES* ND JEFFREY
Proc. Natd. cad. Sci. US Vol. 86, pp. 5345-5349, July 1989 Biochemistry set of lacz mutations in Escherichia coli that allow rapid detection of each of the six base substitutions CLIRE. CUPPLES* ND JEFFREY
Engineering D66N mutant using quick change site directed mutagenesis. Harkewal Singh 09/01/2010
 Engineering D66N mutant using quick change site directed mutagenesis Harkewal Singh 09/01/2010 1 1- What is quick change site directed mutagenesis? 2- An overview of the kit contents. 3- A brief information
Engineering D66N mutant using quick change site directed mutagenesis Harkewal Singh 09/01/2010 1 1- What is quick change site directed mutagenesis? 2- An overview of the kit contents. 3- A brief information
Understanding the Cellular Mechanism of the Excess Microsporocytes I (EMSI) Gene. Andrew ElBardissi, The Pennsylvania State University
 Understanding the Cellular Mechanism of the Excess Microsporocytes I (EMSI) Gene Andrew ElBardissi, The Pennsylvania State University Abstract: Hong Ma, The Pennsylvania State University The Excess Microsporocytes
Understanding the Cellular Mechanism of the Excess Microsporocytes I (EMSI) Gene Andrew ElBardissi, The Pennsylvania State University Abstract: Hong Ma, The Pennsylvania State University The Excess Microsporocytes
Game Plan. Lecture. Lab. Growth Curve. Biofilms Review of basic genetics Bacterial gene structure Gene regulation Mutations
 Game Plan Lecture Biofilms Review of basic genetics Bacterial gene structure Gene regulation Mutations Lab Growth Curve Pre-labs Effects on growth: temp, ph, O2 What is a biofilm Biofilm = an organized
Game Plan Lecture Biofilms Review of basic genetics Bacterial gene structure Gene regulation Mutations Lab Growth Curve Pre-labs Effects on growth: temp, ph, O2 What is a biofilm Biofilm = an organized
Bacterial Mutation Types Mechanisms And Mutant Detection
 Bacterial Mutation Types Mechanisms And Mutant Detection 1 / 5 2 / 5 3 / 5 Bacterial Mutation Types Mechanisms And Substitution of a nucleotide and Deletion or addition of them is. two mechanisms of mutation.
Bacterial Mutation Types Mechanisms And Mutant Detection 1 / 5 2 / 5 3 / 5 Bacterial Mutation Types Mechanisms And Substitution of a nucleotide and Deletion or addition of them is. two mechanisms of mutation.
Biology (Microbiology): Exam #3
 NAME: PLEDGE: Biology 50-384 (Microbiology): Exam #3 1. You have isolated a series of mutants that have altered patterns of ß-galactosidase and lactose permease activity (i.e. they are different that the
NAME: PLEDGE: Biology 50-384 (Microbiology): Exam #3 1. You have isolated a series of mutants that have altered patterns of ß-galactosidase and lactose permease activity (i.e. they are different that the
GENETICS - CLUTCH CH.5 GENETICS OF BACTERIA AND VIRUSES.
 !! www.clutchprep.com CONCEPT: WORKING WITH MICROORGANISMS Bacteria are easy to with in a laboratory setting They are fast dividing, take up little space, and are easily grown in a lab - Plating is when
!! www.clutchprep.com CONCEPT: WORKING WITH MICROORGANISMS Bacteria are easy to with in a laboratory setting They are fast dividing, take up little space, and are easily grown in a lab - Plating is when
ANSWERS TO Problem set questions from Exam 2 Unit Mutations, Bacterial Genetics, and Bacterial Gene Regulation
 ANSWERS TO Problem set questions from Exam 2 Unit Mutations, Bacterial Genetics, and Bacterial Gene Regulation Central Dogma, Mutagens and Mutations 1. The three stop codons in the genetic code are 5 UAG3,
ANSWERS TO Problem set questions from Exam 2 Unit Mutations, Bacterial Genetics, and Bacterial Gene Regulation Central Dogma, Mutagens and Mutations 1. The three stop codons in the genetic code are 5 UAG3,
MCB421 FALL2005 EXAM#3 ANSWERS Page 1 of 12. ANSWER: Both transposon types form small duplications of adjacent host DNA sequences.
 Page 1 of 12 (10pts) 1. There are two mechanisms for transposition used by bacterial transposable elements: replicative (Tn3) and non-replicative (Tn5 and Tn10). Compare and contrast the two mechanisms
Page 1 of 12 (10pts) 1. There are two mechanisms for transposition used by bacterial transposable elements: replicative (Tn3) and non-replicative (Tn5 and Tn10). Compare and contrast the two mechanisms
BACTERIAL CONJUGATION. To demonstrate the technical procedure to monitor the conjugational transfer of genetic material from one cell to another.
 BACTERIAL CONJUGATION I. OBJECTIVES To demonstrate the technical procedure to monitor the conjugational transfer of genetic material from one cell to another. To learn about the various genetic elements
BACTERIAL CONJUGATION I. OBJECTIVES To demonstrate the technical procedure to monitor the conjugational transfer of genetic material from one cell to another. To learn about the various genetic elements
Z Competent E. coli Transformation
 467PR G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Z Competent E. coli Transformation (Cat. # GZ 4, GZ 5) think proteins! think G-Biosciences
467PR G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Z Competent E. coli Transformation (Cat. # GZ 4, GZ 5) think proteins! think G-Biosciences
StrataPrep Plasmid Miniprep Kit
 StrataPrep Plasmid Miniprep Kit INSTRUCTION MANUAL Catalog #400761 and #400763 Revision A For In Vitro Use Only 400761-12 LIMITED PRODUCT WARRANTY This warranty limits our liability to replacement of this
StrataPrep Plasmid Miniprep Kit INSTRUCTION MANUAL Catalog #400761 and #400763 Revision A For In Vitro Use Only 400761-12 LIMITED PRODUCT WARRANTY This warranty limits our liability to replacement of this
Bacterial Genetics, BIO 4443/6443 Fall Semester 2002 Exam I. Name Answer Key
 Name Answer Key Student pid# Bacterial Genetics, BIO 4443/6443 Fall Semester 2002 Exam I 1.) What is a sigma factor? Why does the cell contain multiple sigma factors? (5pts) Sigma Is a subunit of the RNA
Name Answer Key Student pid# Bacterial Genetics, BIO 4443/6443 Fall Semester 2002 Exam I 1.) What is a sigma factor? Why does the cell contain multiple sigma factors? (5pts) Sigma Is a subunit of the RNA
Molecular Genetics Techniques. BIT 220 Chapter 20
 Molecular Genetics Techniques BIT 220 Chapter 20 What is Cloning? Recombinant DNA technologies 1. Producing Recombinant DNA molecule Incorporate gene of interest into plasmid (cloning vector) 2. Recombinant
Molecular Genetics Techniques BIT 220 Chapter 20 What is Cloning? Recombinant DNA technologies 1. Producing Recombinant DNA molecule Incorporate gene of interest into plasmid (cloning vector) 2. Recombinant
Influence of tlp5, che1p, tlp4a, and che4stas Promoters on Chemotaxis in Azospirillum brasilense
 University of Tennessee, Knoxville Trace: Tennessee Research and Creative Exchange University of Tennessee Honors Thesis Projects University of Tennessee Honors Program 5-2015 Influence of tlp5, che1p,
University of Tennessee, Knoxville Trace: Tennessee Research and Creative Exchange University of Tennessee Honors Thesis Projects University of Tennessee Honors Program 5-2015 Influence of tlp5, che1p,
Lab 5/5a Transformation of E. coli with a Recombinant Plasmid
 Lab 5/5a Transformation of E. coli with a Recombinant Plasmid Lab 2 Pre Lab Readiness Familiarity and Proper use of micropipettes Remember the 1 st and 2 nd stops Aseptic Technique Antibiotic Resistance
Lab 5/5a Transformation of E. coli with a Recombinant Plasmid Lab 2 Pre Lab Readiness Familiarity and Proper use of micropipettes Remember the 1 st and 2 nd stops Aseptic Technique Antibiotic Resistance
pgm-t Cloning Kit For direct cloning of PCR products generated by Taq DNA polymerases For research use only Cat. # : GVT202 Size : 20 Reactions
 pgm-t Cloning Kit For direct cloning of PCR products generated by Taq DNA polymerases Cat. # : GVT202 Size : 20 Reactions Store at -20 For research use only 1 pgm-t Cloning Kit Cat. No.: GVT202 Kit Contents
pgm-t Cloning Kit For direct cloning of PCR products generated by Taq DNA polymerases Cat. # : GVT202 Size : 20 Reactions Store at -20 For research use only 1 pgm-t Cloning Kit Cat. No.: GVT202 Kit Contents
7.03 Problem Set 3 Due before 5 PM on Wednesday, October 18 Hand in answers in recitation section or in the box outside of
 7.03 Problem Set 3 Due before 5 PM on Wednesday, October 18 Hand in answers in recitation section or in the box outside of 68-120 1. The following DNA sequence fragment comes from the middle of a bacterial
7.03 Problem Set 3 Due before 5 PM on Wednesday, October 18 Hand in answers in recitation section or in the box outside of 68-120 1. The following DNA sequence fragment comes from the middle of a bacterial
Chapter 9 Microbial Genetics
 Chapter 9 Microbial Genetics You are expected to know details of 1) DNA replication 2) RNA synthesis (transcription) 3) Protein synthesis (translation) Genome & Genes A genome is all the genetic information
Chapter 9 Microbial Genetics You are expected to know details of 1) DNA replication 2) RNA synthesis (transcription) 3) Protein synthesis (translation) Genome & Genes A genome is all the genetic information
CopyCutter EPI400 Electrocompetent E. coli CopyCutter EPI400 Chemically Competent E. coli CopyCutter Induction Solution
 CopyCutter EPI400 Electrocompetent E. coli CopyCutter EPI400 Chemically Competent E. coli CopyCutter Induction Solution Cat. Nos. C400EL10, C400CH10, and CIS40025 Available exclusively thru Lucigen. lucigen.com/epibio
CopyCutter EPI400 Electrocompetent E. coli CopyCutter EPI400 Chemically Competent E. coli CopyCutter Induction Solution Cat. Nos. C400EL10, C400CH10, and CIS40025 Available exclusively thru Lucigen. lucigen.com/epibio
Transformation of Escherichia coli With a Chimeric Plasmid
 Transformation of Escherichia coli With a Chimeric Plasmid Now that we have generated recombinant molecules, we must next amplify them by inserting them into an acceptable host so that they may be analyzed
Transformation of Escherichia coli With a Chimeric Plasmid Now that we have generated recombinant molecules, we must next amplify them by inserting them into an acceptable host so that they may be analyzed
Link between iron homeostasis, oxidative mutagenesis and antibiotic resistance evolution
 Thesis of the PhD dissertation Link between iron homeostasis, oxidative mutagenesis and antibiotic resistance evolution Orsolya Katinka Méhi Supervisor: Dr. Csaba Pál, senior research associate PhD School
Thesis of the PhD dissertation Link between iron homeostasis, oxidative mutagenesis and antibiotic resistance evolution Orsolya Katinka Méhi Supervisor: Dr. Csaba Pál, senior research associate PhD School
Biol 432L Midterm Oct 6, 2008 Name: 1. Midterm 1, Answer Key Oct. 26, 2009
 Biol 432L Midterm Oct 6, 2008 Name: 1 Midterm 1, Answer Key Oct. 26, 2009 Honor Pledge: I have neither given nor received any unauthorized help on this exam: Name Printed: Signature: 1a. (2 pts) Imagine
Biol 432L Midterm Oct 6, 2008 Name: 1 Midterm 1, Answer Key Oct. 26, 2009 Honor Pledge: I have neither given nor received any unauthorized help on this exam: Name Printed: Signature: 1a. (2 pts) Imagine
Biol/Chem 475 Spring 2007
 Biol/Chem 475 Spring 2007 Goal of lab: For most of the quarter, we will be exploring a gene family that was first discovered in fruitlfies and then found to be present in humans and worms and fish and
Biol/Chem 475 Spring 2007 Goal of lab: For most of the quarter, we will be exploring a gene family that was first discovered in fruitlfies and then found to be present in humans and worms and fish and
Supplemental Materials and Methods. Yeast strains. Strain genotypes are listed in Table S1. To generate MATainc::LEU2,
 Supplemental Materials and Methods Yeast strains. Strain genotypes are listed in Table S1. To generate MATainc::LEU2, the LEU2 marker was inserted by PCR-mediated recombination (BRACHMANN et al. 1998)
Supplemental Materials and Methods Yeast strains. Strain genotypes are listed in Table S1. To generate MATainc::LEU2, the LEU2 marker was inserted by PCR-mediated recombination (BRACHMANN et al. 1998)
An engineered tryptophan zipper-type peptide as a molecular recognition scaffold
 SUPPLEMENTARY MATERIAL An engineered tryptophan zipper-type peptide as a molecular recognition scaffold Zihao Cheng and Robert E. Campbell* Supplementary Methods Library construction for FRET-based screening
SUPPLEMENTARY MATERIAL An engineered tryptophan zipper-type peptide as a molecular recognition scaffold Zihao Cheng and Robert E. Campbell* Supplementary Methods Library construction for FRET-based screening
Lecture 1. Basic Definitions and Nucleic Acids. Basic Definitions you should already know
 Lecture 1. Basic Definitions and Nucleic Acids Basic Definitions you should already know apple DNA: Deoxyribonucleic Acid apple RNA: Ribonucleic Acid apple mrna: messenger RNA: contains the genetic information(coding
Lecture 1. Basic Definitions and Nucleic Acids Basic Definitions you should already know apple DNA: Deoxyribonucleic Acid apple RNA: Ribonucleic Acid apple mrna: messenger RNA: contains the genetic information(coding
Supporting Information: DNA-mediated signaling by proteins with 4Fe-4S clusters is necessary for genomic integrity
 Supporting Information: DNA-mediated signaling by proteins with 4Fe-4S clusters is necessary for genomic integrity Michael A. Grodick, 1 Helen M. Segal, 1 Theodore J. Zwang, 1 and Jacqueline K. Barton
Supporting Information: DNA-mediated signaling by proteins with 4Fe-4S clusters is necessary for genomic integrity Michael A. Grodick, 1 Helen M. Segal, 1 Theodore J. Zwang, 1 and Jacqueline K. Barton
4/3/2013. DNA Synthesis Replication of Bacterial DNA Replication of Bacterial DNA
 4/3/03 3 4 5 6 7 8 9 0 Chapter 8 Microbial Genetics Terminology Genetics: The study of what genes are, how they carry information, how information is expressed, and how genes are replicated Gene: A segment
4/3/03 3 4 5 6 7 8 9 0 Chapter 8 Microbial Genetics Terminology Genetics: The study of what genes are, how they carry information, how information is expressed, and how genes are replicated Gene: A segment
Haverford College Annual Progress Report: 2012 Formula Grant
 Haverford College Annual Progress Report: 2012 Formula Grant Reporting Period January 1, 2013 June 30, 2013 Formula Grant Overview Haverford College received $18,238 in formula funds for the grant award
Haverford College Annual Progress Report: 2012 Formula Grant Reporting Period January 1, 2013 June 30, 2013 Formula Grant Overview Haverford College received $18,238 in formula funds for the grant award
Supplemental Material: Rev1 promotes replication through UV lesions in conjunction with DNA
 Supplemental Material: Rev1 promotes replication through UV lesions in conjunction with DNA polymerases,, and, but not with DNA polymerase Jung-Hoon Yoon, Jeseong Park, Juan Conde, Maki Wakamiya, Louise
Supplemental Material: Rev1 promotes replication through UV lesions in conjunction with DNA polymerases,, and, but not with DNA polymerase Jung-Hoon Yoon, Jeseong Park, Juan Conde, Maki Wakamiya, Louise
BIOTECHNOLOGY. Sticky & blunt ends. Restriction endonucleases. Gene cloning an overview. DNA isolation & restriction
 BIOTECHNOLOGY RECOMBINANT DNA TECHNOLOGY Recombinant DNA technology involves sticking together bits of DNA from different sources. Made possible because DNA & the genetic code are universal. 2004 Biology
BIOTECHNOLOGY RECOMBINANT DNA TECHNOLOGY Recombinant DNA technology involves sticking together bits of DNA from different sources. Made possible because DNA & the genetic code are universal. 2004 Biology
BP-reaction (recombination of PCR products into entry vector) Created on: 06/01/10 Version: 1.1 No.: 1 Page 1 of 6
 Created on: 06/01/10 Version: 1.1 No.: 1 Page 1 of 6 1. Background This reaction generates bacteria that contain recombinant Gateway-compatible entry plasmid. The PCRamplified ORFs (SOPs ORF-PCR 1 (two
Created on: 06/01/10 Version: 1.1 No.: 1 Page 1 of 6 1. Background This reaction generates bacteria that contain recombinant Gateway-compatible entry plasmid. The PCRamplified ORFs (SOPs ORF-PCR 1 (two
Lecture Four. Molecular Approaches I: Nucleic Acids
 Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
Supplementary Figure 1 A) MMC provokes a rapid replication arrest. We used a strain with a dnaats allele to
 Supplementary Figure 1 A) MMC provokes a rapid replication arrest. We used a strain with a dnaats allele to block the initiation of replication when placed at 40 C. The ori/ter ratio was monitored by qpcr.
Supplementary Figure 1 A) MMC provokes a rapid replication arrest. We used a strain with a dnaats allele to block the initiation of replication when placed at 40 C. The ori/ter ratio was monitored by qpcr.
1a. What is the ratio of feathered to unfeathered shanks in the offspring of the above cross?
 1. Whether or not the shanks of chickens contains feathers is due to two independently assorting genes. Individuals have unfeathered shanks when they are homozygous for recessive genes at two loci; the
1. Whether or not the shanks of chickens contains feathers is due to two independently assorting genes. Individuals have unfeathered shanks when they are homozygous for recessive genes at two loci; the
COMPARISON OF CLONING AND SEQUENCING VS. TERMINAL RESTRICTION FRAGMENT ANALYSIS OF BACTERIA FROM HUMAN FECES. David Doherty and Danielle Elizondo
 COMPARISON OF CLONING AND SEQUENCING VS. TERMINAL RESTRICTION FRAGMENT ANALYSIS OF BACTERIA FROM HUMAN FECES. David Doherty and Danielle Elizondo October 24 th 2001 Abstract Extracted DNA from the Probiotic
COMPARISON OF CLONING AND SEQUENCING VS. TERMINAL RESTRICTION FRAGMENT ANALYSIS OF BACTERIA FROM HUMAN FECES. David Doherty and Danielle Elizondo October 24 th 2001 Abstract Extracted DNA from the Probiotic
Signature-Tagged Mutagenesis to Identify Virulence Genes in Salmonella choleraesuis
 Abstract Signature-Tagged Mutagenesis to Identify Virulence Genes in Salmonella choleraesuis Carol A. Lichtensteiger 1 and Eric R. Vimr 2 Department of Veterinary Pathobiology 1,2 and Veterinary Diagnostic
Abstract Signature-Tagged Mutagenesis to Identify Virulence Genes in Salmonella choleraesuis Carol A. Lichtensteiger 1 and Eric R. Vimr 2 Department of Veterinary Pathobiology 1,2 and Veterinary Diagnostic
Step 1: Digest vector with Reason for Step 1. Step 2: Digest T4 genomic DNA with Reason for Step 2: Step 3: Reason for Step 3:
 Biol/Chem 475 Spring 2007 Study Problems for Quiz 2 Quiz 2 (~50 pts) is scheduled for Monday May 14 It will cover all handouts and lab exercises to date except the handout/worksheet (yet to be distributed)
Biol/Chem 475 Spring 2007 Study Problems for Quiz 2 Quiz 2 (~50 pts) is scheduled for Monday May 14 It will cover all handouts and lab exercises to date except the handout/worksheet (yet to be distributed)
TransforMax EPI300 Electrocompetent E. coli TransforMax EPI300 Chemically Competent E. coli
 TransforMax EPI300 Electrocompetent E. coli TransforMax EPI300 Chemically Competent E. coli Cat. Nos. EC300102, EC300110, EC300150, and C300C105 Available exclusively thru Lucigen. lucigen.com/epibio www.lucigen.com
TransforMax EPI300 Electrocompetent E. coli TransforMax EPI300 Chemically Competent E. coli Cat. Nos. EC300102, EC300110, EC300150, and C300C105 Available exclusively thru Lucigen. lucigen.com/epibio www.lucigen.com
Pr oject Summar y. Rapid quantification of culturable and viable-but-nonculturable Escherichia coli O157:H7 in beef products using EMA-Real Time PCR
 Pr oject Summar y Rapid quantification of culturable and viable-but-nonculturable Escherichia coli O17:H7 in beef products using EMA-Real Time PCR Principal Investigator: Azlin Mustapha University of Missouri
Pr oject Summar y Rapid quantification of culturable and viable-but-nonculturable Escherichia coli O17:H7 in beef products using EMA-Real Time PCR Principal Investigator: Azlin Mustapha University of Missouri
SUPPLEMENTARY MATERIAL AND METHODS
 SUPPLEMENTARY MATERIAL AND METHODS Amplification of HEV ORF1, ORF2 and ORF3 genome regions Total RNA was extracted from 200 µl EDTA plasma using Cobas AmpliPrep total nucleic acid isolation kit (Roche,
SUPPLEMENTARY MATERIAL AND METHODS Amplification of HEV ORF1, ORF2 and ORF3 genome regions Total RNA was extracted from 200 µl EDTA plasma using Cobas AmpliPrep total nucleic acid isolation kit (Roche,
SCREENING AND PRESERVATION OF DNA LIBRARIES
 MODULE 4 LECTURE 5 SCREENING AND PRESERVATION OF DNA LIBRARIES 4-5.1. Introduction Library screening is the process of identification of the clones carrying the gene of interest. Screening relies on a
MODULE 4 LECTURE 5 SCREENING AND PRESERVATION OF DNA LIBRARIES 4-5.1. Introduction Library screening is the process of identification of the clones carrying the gene of interest. Screening relies on a
Genetic Background Page 1 PHAGE P22
 Genetic Background Page 1 PHAGE P22 Growth of P22. P22 is a temperate phage that infects Salmonella by binding to the O-antigen, part of the lipopolysaccharide on the outer membrane. After infection, P22
Genetic Background Page 1 PHAGE P22 Growth of P22. P22 is a temperate phage that infects Salmonella by binding to the O-antigen, part of the lipopolysaccharide on the outer membrane. After infection, P22
Mos1 insertion. MosTIC protocol-11/2006. repair template - Mos1 transposase expression - Mos1 excision - DSB formation. homolog arm.
 MosTIC (Mos1 excision induced Transgene Instructed gene Conversion) Valérie Robert (vrobert@biologie.ens.fr) and Jean-Louis Bessereau (jlbesse@biologie.ens.fr) (November 2006) Introduction: MosTIC (Robert
MosTIC (Mos1 excision induced Transgene Instructed gene Conversion) Valérie Robert (vrobert@biologie.ens.fr) and Jean-Louis Bessereau (jlbesse@biologie.ens.fr) (November 2006) Introduction: MosTIC (Robert
Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC
 Supplementary Appendixes Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC ACG TAG CTC CGG CTG GA-3 for vimentin, /5AmMC6/TCC CTC GCG CGT GGC TTC CGC
Supplementary Appendixes Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC ACG TAG CTC CGG CTG GA-3 for vimentin, /5AmMC6/TCC CTC GCG CGT GGC TTC CGC
Ames MPF TM Penta 1 Microplate Format Mutagenicity Assay
 Ames MPF TM Penta 1 Microplate Format Mutagenicity Assay Strains: S. typhimurium TA98, TA100, TA1535, TA1537 and E. coli WP2 uvra + E. coli WP2 [pkm101] Short Protocol For Research use only Version 2.0
Ames MPF TM Penta 1 Microplate Format Mutagenicity Assay Strains: S. typhimurium TA98, TA100, TA1535, TA1537 and E. coli WP2 uvra + E. coli WP2 [pkm101] Short Protocol For Research use only Version 2.0
Cat. # Product Size DS130 DynaExpress TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1
 Product Name: Kit Component TA PCR Cloning Kit (ptakn-2) Cat. # Product Size DS130 TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1 2 Ligation Buffer
Product Name: Kit Component TA PCR Cloning Kit (ptakn-2) Cat. # Product Size DS130 TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1 2 Ligation Buffer
Hetero-Stagger PCR Cloning Kit
 Product Name: Code No: Size: DynaExpress Hetero-Stagger PCR Cloning Kit DS150 20 reactions Kit Components: Box 1 (-20 ) phst-1 Vector, linearized Annealing Buffer Ligase Mixture phst Forward Sequence Primer
Product Name: Code No: Size: DynaExpress Hetero-Stagger PCR Cloning Kit DS150 20 reactions Kit Components: Box 1 (-20 ) phst-1 Vector, linearized Annealing Buffer Ligase Mixture phst Forward Sequence Primer
Bacterial Genetics. Stijn van der Veen
 Bacterial Genetics Stijn van der Veen Differentiating bacterial species Morphology (shape) Composition (cell envelope and other structures) Metabolism & growth characteristics Genetics Differentiating
Bacterial Genetics Stijn van der Veen Differentiating bacterial species Morphology (shape) Composition (cell envelope and other structures) Metabolism & growth characteristics Genetics Differentiating
Orthogonal Ribosome Biofirewall
 1 2 3 4 5 Supporting Information Orthogonal Ribosome Biofirewall Bin Jia,, Hao Qi,, Bing-Zhi Li,, Shuo pan,, Duo Liu,, Hong Liu,, Yizhi Cai, and Ying-Jin Yuan*, 6 7 8 9 10 11 12 Key Laboratory of Systems
1 2 3 4 5 Supporting Information Orthogonal Ribosome Biofirewall Bin Jia,, Hao Qi,, Bing-Zhi Li,, Shuo pan,, Duo Liu,, Hong Liu,, Yizhi Cai, and Ying-Jin Yuan*, 6 7 8 9 10 11 12 Key Laboratory of Systems
Enterococcus faecium. genesig Standard Kit. groes heat shock protein. 150 tests. Primerdesign Ltd. For general laboratory and research use only
 TM Primerdesign Ltd Enterococcus faecium groes heat shock protein genesig Standard Kit 150 tests For general laboratory and research use only 1 Introduction to Enterococcus faecium E. faecium is a Gram-positive,
TM Primerdesign Ltd Enterococcus faecium groes heat shock protein genesig Standard Kit 150 tests For general laboratory and research use only 1 Introduction to Enterococcus faecium E. faecium is a Gram-positive,
7.1 Techniques for Producing and Analyzing DNA. SBI4U Ms. Ho-Lau
 7.1 Techniques for Producing and Analyzing DNA SBI4U Ms. Ho-Lau What is Biotechnology? From Merriam-Webster: the manipulation of living organisms or their components to produce useful usually commercial
7.1 Techniques for Producing and Analyzing DNA SBI4U Ms. Ho-Lau What is Biotechnology? From Merriam-Webster: the manipulation of living organisms or their components to produce useful usually commercial
