Roseobacter denitrificans genome annotation using Manatee
|
|
|
- Franklin Harris
- 6 years ago
- Views:
Transcription
1 Roseobacter denitrificans genome annotation using Manatee Chaitanya R. Acharya Computational Biosciences Arizona State University May 3, 2006
2 Acknowledgements Thanks to my project advisor Dr. Robert Blankenship, who proposed this internship project and provided guidance throughout. Thanks to my team: Sumedha Gholba, Hector Ramos, Wesley Swingley, Heather Matthies and Michael Lince, for their help and encouragement throughout. Thanks to Dr. Jeffrey Touchman and his crew (at TGen) for their help in understanding the sequence deformities. Thanks to my committee: Dr. Rosemary Renaut, Dr. Blankenship and Dr. Martin Wojciechowski, for the knowledge they have shared throughout my program and for review of this applied project. Thanks to Loretta Goldberg, for her help in putting together this presentation.
3 This internship project is presented in partial fulfillment of the requirements of the Professional Science Master s in Computational Biosciences, Arizona State University Research conducted from June 2005 August 2005
4 Project Overview
5 Annotation Putative genes are located and identified on a newly sequenced genome Homology search is also used to understand the physical and functional characteristics of the gene products Identify the role played by that specific gene product, if it is a protein, in a metabolic pathway (Metabolic profile) The elements of the annotation process include gene finding, homology searches, functional assignment, ORF management and finally making this data available to the public.
6 Phototrophic Genome Project This internship project is a part of phototrophic genome project Obtaining genomic sequences of four representative bacterial species: Heliobacteria (Heliobacterium modesticaldum), Aerobic Phototrophic bacterium (Roseobacter denitrificans), Alpha Proteobacteria (Rhodocista centenaria) and Cyanobacteria (Acaryachloris marina). Please visit
7 16S rrna diversity analysis Courtesy
8 Roseobacter denitrificans a model aerobic phototrophic bacteria Depend of the respiration of organic compounds for growth Cessation of photosynthetic pigment synthesis upon illumination Requires a respiratory terminal electron acceptor (invariably oxygen) Questions are raised on the evolution and genetic regulation of photosynthesis
9 Significance Roseobacter denitrificans genome The evolutionary genesis of photosynthetic genes True evolutionary positions of aerobic phototrophic bacteria would be clarified by whole genome comparisons Pathways of carbon dioxide fixation and production Constructing the metabolic profile is very important which could be tested in biochemical and molecular biology experiments Light and oxygen signal transduction in gene expression Study both oxygen- and light-responsive pathways by cloning, gene-disruption methods and over-expressing genes for biochemical and biophysical analysis of purified proteins
10 Why Roseobacter denitrificans? Readily cultivated in the laboratory Electron transfer pathways establish that this is the model aerobic phototrophic bacterium Only bacterium that is capable of anaerobic growth (nitrate as terminal electron acceptor) Marine bacterium High GC content (~59%) and relatively small genome size (~4 Mb)
11 Sequencing R. denitrificans: TGen s role Sequencing R. denitrificans was accomplished at Translatory Genomics Institute (TGen) R. denitrificans contains a primary circular chromosome of 4,133,097 base pairs and four plasmid containing a total of 4,403 predicted coding sequences
12 TIGR s role
13 Annotation Engine (AE) at TIGR First executes the annotation in a format that promotes consistency of data types across all genomes Allows straightforward reincorporation of annotation data back into the Comprehensive Microbial Resource (CMR) data management system Aids in quality control of the sequence Preliminary annotation is displayed on the web
14 Components of AE Production of output from TIGR's automated annotation pipeline - includes search results and automatically generated annotation in a MySQL database and associated files The manual annotation tool Manatee - an open source web based interface for interacting with and editing annotation data
15 Elements of AE Gene Finding using the Glimmer system Predicting candidate genes and the proteins they code for Functional assignment of all predicted proteins Search against an internal non-identical amino acid (NIAA) database Data supplied to the AE user or annotator Data is available as a MySQL database Providing annotation tool, Manatee. TIGR's manual annotation tool
16 Glimmer System Identifies probable open reading frames (ORFs) The Glimmer software system developed by Salzberg and Delcher et al. is used to find genes in many prokaryotic genomes such as bacterial, viral and archaeal genomes. The algorithm at the core of the Glimmer is an Interpolated Markov Model (IMM), which is a special kind of Markov chain. A Markov chain calculated the statistical information about any sequence by computing the conditional probability P(x S) that a nucleotide x appears after a sequence S. Second- or fifth- or eighth-order Markov chains work well because they are computing statistics based on codons, dicodons and tricodons respectively.
17 Glimmering STEP 1: Choosing a Training set Homology Search on the Genomic Seq BLAST-Extend- Repraze (BER) STEP 2: Identify ORFs
18 Glimmer Algorithm
19 Visualizing genes Proteins searched against HMMs using hmmpfam program. Two-fold homology search is performed to identify probable ORFs. ORFs visualized in sixframe translations maps HMMs used here are Pfam HMM and TIGRFAM
20 Translation Maps Green regions: High scoring ORFs Red regions: Low scoring ORFs
21 Testing Overlaps Mapping the ORFs Scoring of the overlap region
22 Manatee Manatee is a web-based gene evaluation and genome annotation tool that can view, modify, and store annotation for prokaryotic and eukaryotic genomes Manatee consists of a suite of programs, including Gene Ontology (GO) classifications, Blast-Extend Repraze (BER), Blast search data, paralogous families, and annotation suggestions generated from the automated analysis. It is an open-source initiative that was developed for two main reasons: 1) to help biologists annotate their genomes using a powerful, stand-alone web application with a robustly designed relational annotation database, and 2) to invite developers from all over the world to enhance Manatee s ability to completely accomplish biological goals.
23 ORF Management
24 Determining Functional Roles of Predicted Proteins Determined by homology searching and/or experimental characterization of proteins High identities (at least >35%) prove that the sequences share their functions. All the functional assignments made after sequence alignments should be considered putative (suggested) until confirmed by experimental characterization.
25 Determining Functional Roles of Predicted Proteins Characterized proteins are stored in Characterized Table within the database; they are designated a confidence status They are all color coded: Green color = full experimental characterization Red color = characterized by Swiss-Prot (by an automated process) Sky blue color = partial characterization Olive color = trusted to be characterized Blue-green color = only a fragment/domain has been characterized Fuzzy gray color = void Gray color = sequence exists in the Omnium (database that underlies TIGR s CMR)
26 Non-identical amino acid (NIAA) database This file is composed of protein sequences from many protein translations of all ORFs searched against hidden Markov models (HMM) built from the multiple amino acid sequence alignments. Each HMM is associated with a noise cutoff score and a trusted cutoff score. ORFs are considered to be members of the HMM model if they score higher than the trusted cutoff. TIGR classifies TIGR and Pfam HMMs into fifteen isologies. Each isology defines a specific database match
27 BLAST-Extend Repraze (BER) The BER alignments are stored in a mini-database and the closest matches are displayed in a table. They are assigned scores. Table displayed in Gene Curation Page (GCP) of an ORF as BER Skim. The matches are color coded (as mentioned earlier) depending on their characterization
28 BER Skim
29 Evidence Picture Green colored matches are all characterized Clusters of Orthologous groups (COGs) also play an important role in predicted protein identification.
30 One look at a match There are two parts: 1. Top gold box 2. Bottom alignment
31 One look at a match
32 Alignments Codons read from top to bottom Asterisk indicates a stop codon Negative numbers indicate downstream sequence from the putative start site Start sites are all color coded There are three start sites- ATG (Methionine), TTG (Leucine) and GTG (Valine) Solid line indicates identical amino acid, broken line indicates similar amino acid
33 Identifying a start site Courtesy
34 Genome View
35 Editing Start site Green nucleotides represent ribosome binding sites, purple nucleotides represent amino acids of the query gene and blue nucleotides represent amino acids of one of the genes in the region other than the query gene. Start site edited based on the Genome View
36 Gene Curation Page Has ORF descriptors The annotator has the option of populating six fields: Common name (com_name), Gene Symbol (gen_sym), Enzyme Commission number (ec_num), comments, TIGR roles (role_id) and GO terms HMMs are all shown along with the BER Skim Annotation begins here...
37 Annotation Protocol Genome View check for overlap HMM evidence picture BER SKIM Edit start site Check links Naming Characterized match (p-value < 1050 or 35% identity) PFAM Gene Symbol EC number Comment TIGR roles Gene Ontology terms (Can be skipped) Submit Data Report Frame shifts, start errors & overlaps
38 Overlaps in Genome View Overlaps have to be checked for before being discarded Look at the sequence and check for any frame shift mutations
39 Demo Please go to:
40 Observations R. denitrificans lacks key Calvin cycle enzymes ribulose bisphosphate carboxylase (RubisCO), phosphoribulokinase (PRK), and other proteins typically coded by the Calvin cycle operons in closely related anaerobic purple bacteria. Suggests evidence for a scattered loss of ancestral carbon-fixation in the α-proteobacterial tree. Some putative genes related to the C4 sequestration, Crassulacean acid metabolism (CAM), and anaplerotic carbon-fixation enzymes such as pyruvate-orthophosphate dikinase and phosphoenolpyruvate (PEP) carboxylase, are present. Hypothesized that APBs fix CO 2 heterotrophically using a C4 sequestration pathway supplemented by additional CO 2 provided by CO oxidation and heterotrophic respiration.
41 Future Directions Unclear how R. denitrificans adapted to the changing atmospheric conditions, and how RubisCO evolved in the APBs such as R. denitrificans. Functional characterization of proteins in labs would allow scientists to further probe the organism and understand various other metabolic pathways associated with the overall metabolic profile of R. denitrificans. More comparative sequence (or genome) analysis of the sequenced APBs
42 References [Alts 90] Altschul S F, Gish W, Miller W, Myers E W, Lipman D J;Basic Local Search Alignment Tool; Journal of Molecular Biology 1990; 215: [Alts 97] Altschul S F, Madden T L, Schaffer A A, Zhang J, Zhang Z, Miller W, Lipman D J; Gapped BLAST and PSI_BLAST: a new generation of protein database search programs; Nucleic Acid Res 1997; 25: [Arth 99] Arthur L. Delcher, Douglas Harmon, Simon Kasif, Owen White and Steven L. Salzberg; Improved microbial gene identification with GLIMMER; Nucleic Acid Res 1999; 27: [Arth 05] Arthur L. Delcher; GLIMMER Release Notes, Version 3.01 (Beta) 10 October, 2005 [Bate 90] Bateman A., et al.; The Pfam protein families database; Nucleic Acid Res. 1990; 28(1): [Bea 02] Beatty JT; On the natural selection and evolution of the aerobic phototrophic bacteria; Photosynth Res 2002; 73: [Bla 02] Blankenship RE (2002) Molecular Mechanisms of Photosynthesis, Blackwell Science, Oxford, UK.
43 References [Bla 98] Blankenship RE and Hartman H (1998); The origin and evolution of oxygenic photosynthesis. Trends Biochem Sci 23: Ed 99] Eddy S; Profile hidden Markov models; Bioinformatics; 14(9): [Gis 93] Gish W, et al.; Identification of protein coding regions by database [Haf 01] similarity search; Nat. Gent 1993; 3(3): Haft D, et al.; TIGRFAMs: A protein family resource for the functional identification of proteins; Nucleic Acids Res 2001; 29(1):41-3 [Jean 98] Jeanmougin F, Thompson J D, Gouy M, Higgins D G, and Gibson, T J; Multiple Sequence Alignments with Clustal X; Trends Biochem Sci 1998; 23:403-5 [Sal 98] Salzberg S., et al.; Microbial gene identification using Interpolated Markov Models; Nucleic Acid Res 1998; 26(2): [Smi 81] Smith T F, et al.; Identification of common molecular subsequences; J Mol Biol 1981; 147(1): [Tigr 04] TIGR; Michelle Gwinn; Prokaryotic Annotation Overview, October 2004 [Tigr 04] TIGR; Michelle Gwinn; A guide to Manatee, October 2004
44 References [Tigr 05] TIGR; Michelle Gwinn, William Nelson, Robert Dodson, Steven Salzberg, Owen White; Small Genome Annotation and Data Management at TIGR [Tigr 05] TIGR; Domain Based Paralogous Protein Families; Annotation Workshop July13, 2005 [TGen] Translatory Genomics Institute; [Wes 06] Wesley D S, et al; A ubiquitous pathway marine phototroph with a novel carbon-fixation pathway; submitted to PNAS, 2006 [Man 05] Manatee web site; ; edit version active from June 2005 August 2005; [1] Phototrophic Genome Project web site;
Small Genome Annotation and Data Management at TIGR
 Small Genome Annotation and Data Management at TIGR Michelle Gwinn, William Nelson, Robert Dodson, Steven Salzberg, Owen White Abstract TIGR has developed, and continues to refine, a comprehensive, efficient
Small Genome Annotation and Data Management at TIGR Michelle Gwinn, William Nelson, Robert Dodson, Steven Salzberg, Owen White Abstract TIGR has developed, and continues to refine, a comprehensive, efficient
Glossary of Commonly used Annotation Terms
 Glossary of Commonly used Annotation Terms Akela a general use server for the annotation group as well as other groups throughout TIGR. Annotation Notebook a link from the gene list page that is associated
Glossary of Commonly used Annotation Terms Akela a general use server for the annotation group as well as other groups throughout TIGR. Annotation Notebook a link from the gene list page that is associated
Gene Prediction Chengwei Luo, Amanda McCook, Nadeem Bulsara, Phillip Lee, Neha Gupta, and Divya Anjan Kumar
 Gene Prediction Chengwei Luo, Amanda McCook, Nadeem Bulsara, Phillip Lee, Neha Gupta, and Divya Anjan Kumar Gene Prediction Introduction Protein-coding gene prediction RNA gene prediction Modification
Gene Prediction Chengwei Luo, Amanda McCook, Nadeem Bulsara, Phillip Lee, Neha Gupta, and Divya Anjan Kumar Gene Prediction Introduction Protein-coding gene prediction RNA gene prediction Modification
Gene Prediction Chengwei Luo, Amanda McCook, Nadeem Bulsara, Phillip Lee, Neha Gupta, and Divya Anjan Kumar
 Gene Prediction Chengwei Luo, Amanda McCook, Nadeem Bulsara, Phillip Lee, Neha Gupta, and Divya Anjan Kumar Gene Prediction Introduction Protein-coding gene prediction RNA gene prediction Modification
Gene Prediction Chengwei Luo, Amanda McCook, Nadeem Bulsara, Phillip Lee, Neha Gupta, and Divya Anjan Kumar Gene Prediction Introduction Protein-coding gene prediction RNA gene prediction Modification
IGS Annota*on Engine and Manatee. Michelle Gwinn Giglio Pathway Tools Workshop October 2010
 IGS Annota*on Engine and Manatee Michelle Gwinn Giglio Pathway Tools Workshop October 2010 IGS Annota*on Engine A free service to anyone with a prokaryo*c sequence they wish to annotate that provides:
IGS Annota*on Engine and Manatee Michelle Gwinn Giglio Pathway Tools Workshop October 2010 IGS Annota*on Engine A free service to anyone with a prokaryo*c sequence they wish to annotate that provides:
Motif Discovery from Large Number of Sequences: a Case Study with Disease Resistance Genes in Arabidopsis thaliana
 Motif Discovery from Large Number of Sequences: a Case Study with Disease Resistance Genes in Arabidopsis thaliana Irfan Gunduz, Sihui Zhao, Mehmet Dalkilic and Sun Kim Indiana University, School of Informatics
Motif Discovery from Large Number of Sequences: a Case Study with Disease Resistance Genes in Arabidopsis thaliana Irfan Gunduz, Sihui Zhao, Mehmet Dalkilic and Sun Kim Indiana University, School of Informatics
BIOINFORMATICS TO ANALYZE AND COMPARE GENOMES
 BIOINFORMATICS TO ANALYZE AND COMPARE GENOMES We sequenced and assembled a genome, but this is only a long stretch of ATCG What should we do now? 1. find genes What are the starting and end points for
BIOINFORMATICS TO ANALYZE AND COMPARE GENOMES We sequenced and assembled a genome, but this is only a long stretch of ATCG What should we do now? 1. find genes What are the starting and end points for
Textbook Reading Guidelines
 Understanding Bioinformatics by Marketa Zvelebil and Jeremy Baum Last updated: May 1, 2009 Textbook Reading Guidelines Preface: Read the whole preface, and especially: For the students with Life Science
Understanding Bioinformatics by Marketa Zvelebil and Jeremy Baum Last updated: May 1, 2009 Textbook Reading Guidelines Preface: Read the whole preface, and especially: For the students with Life Science
Textbook Reading Guidelines
 Understanding Bioinformatics by Marketa Zvelebil and Jeremy Baum Last updated: January 16, 2013 Textbook Reading Guidelines Preface: Read the whole preface, and especially: For the students with Life Science
Understanding Bioinformatics by Marketa Zvelebil and Jeremy Baum Last updated: January 16, 2013 Textbook Reading Guidelines Preface: Read the whole preface, and especially: For the students with Life Science
Two Mark question and Answers
 1. Define Bioinformatics Two Mark question and Answers Bioinformatics is the field of science in which biology, computer science, and information technology merge into a single discipline. There are three
1. Define Bioinformatics Two Mark question and Answers Bioinformatics is the field of science in which biology, computer science, and information technology merge into a single discipline. There are three
Product Applications for the Sequence Analysis Collection
 Product Applications for the Sequence Analysis Collection Pipeline Pilot Contents Introduction... 1 Pipeline Pilot and Bioinformatics... 2 Sequence Searching with Profile HMM...2 Integrating Data in a
Product Applications for the Sequence Analysis Collection Pipeline Pilot Contents Introduction... 1 Pipeline Pilot and Bioinformatics... 2 Sequence Searching with Profile HMM...2 Integrating Data in a
Following text taken from Suresh Kumar. Bioinformatics Web - Comprehensive educational resource on Bioinformatics. 6th May.2005
 Bioinformatics is the recording, annotation, storage, analysis, and searching/retrieval of nucleic acid sequence (genes and RNAs), protein sequence and structural information. This includes databases of
Bioinformatics is the recording, annotation, storage, analysis, and searching/retrieval of nucleic acid sequence (genes and RNAs), protein sequence and structural information. This includes databases of
Genome Sequence Assembly
 Genome Sequence Assembly Learning Goals: Introduce the field of bioinformatics Familiarize the student with performing sequence alignments Understand the assembly process in genome sequencing Introduction:
Genome Sequence Assembly Learning Goals: Introduce the field of bioinformatics Familiarize the student with performing sequence alignments Understand the assembly process in genome sequencing Introduction:
FUNCTIONAL BIOINFORMATICS
 Molecular Biology-2018 1 FUNCTIONAL BIOINFORMATICS PREDICTING THE FUNCTION OF AN UNKNOWN PROTEIN Suppose you have found the amino acid sequence of an unknown protein and wish to find its potential function.
Molecular Biology-2018 1 FUNCTIONAL BIOINFORMATICS PREDICTING THE FUNCTION OF AN UNKNOWN PROTEIN Suppose you have found the amino acid sequence of an unknown protein and wish to find its potential function.
Gene Prediction Background & Strategy Faction 2 February 22, 2017
 Gene Prediction Background & Strategy Faction 2 February 22, 2017 Group Members: Michelle Kim Khushbu Patel Krithika Xinrui Zhou Chen Lin Sujun Zhao Hannah Hatchell rohini mopuri Jack Cartee Introduction
Gene Prediction Background & Strategy Faction 2 February 22, 2017 Group Members: Michelle Kim Khushbu Patel Krithika Xinrui Zhou Chen Lin Sujun Zhao Hannah Hatchell rohini mopuri Jack Cartee Introduction
TIGR THE INSTITUTE FOR GENOMIC RESEARCH
 Introduction to Genome Annotation: Overview of What You Will Learn This Week C. Robin Buell May 21, 2007 Types of Annotation Structural Annotation: Defining genes, boundaries, sequence motifs e.g. ORF,
Introduction to Genome Annotation: Overview of What You Will Learn This Week C. Robin Buell May 21, 2007 Types of Annotation Structural Annotation: Defining genes, boundaries, sequence motifs e.g. ORF,
Bacterial Genome Annotation
 Bacterial Genome Annotation Bacterial Genome Annotation For an annotation you want to predict from the sequence, all of... protein-coding genes their stop-start the resulting protein the function the control
Bacterial Genome Annotation Bacterial Genome Annotation For an annotation you want to predict from the sequence, all of... protein-coding genes their stop-start the resulting protein the function the control
ab initio and Evidence-Based Gene Finding
 ab initio and Evidence-Based Gene Finding A basic introduction to annotation Outline What is annotation? ab initio gene finding Genome databases on the web Basics of the UCSC browser Evidence-based gene
ab initio and Evidence-Based Gene Finding A basic introduction to annotation Outline What is annotation? ab initio gene finding Genome databases on the web Basics of the UCSC browser Evidence-based gene
Identifying Regulatory Regions using Multiple Sequence Alignments
 Identifying Regulatory Regions using Multiple Sequence Alignments Prerequisites: BLAST Exercise: Detecting and Interpreting Genetic Homology. Resources: ClustalW is available at http://www.ebi.ac.uk/tools/clustalw2/index.html
Identifying Regulatory Regions using Multiple Sequence Alignments Prerequisites: BLAST Exercise: Detecting and Interpreting Genetic Homology. Resources: ClustalW is available at http://www.ebi.ac.uk/tools/clustalw2/index.html
Genes and gene finding
 Genes and gene finding Ben Langmead Department of Computer Science You are free to use these slides. If you do, please sign the guestbook (www.langmead-lab.org/teaching-materials), or email me (ben.langmead@gmail.com)
Genes and gene finding Ben Langmead Department of Computer Science You are free to use these slides. If you do, please sign the guestbook (www.langmead-lab.org/teaching-materials), or email me (ben.langmead@gmail.com)
Self-test Quiz for Chapter 12 (From DNA to Protein: Genotype to Phenotype)
 Self-test Quiz for Chapter 12 (From DNA to Protein: Genotype to Phenotype) Question#1: One-Gene, One-Polypeptide The figure below shows the results of feeding trials with one auxotroph strain of Neurospora
Self-test Quiz for Chapter 12 (From DNA to Protein: Genotype to Phenotype) Question#1: One-Gene, One-Polypeptide The figure below shows the results of feeding trials with one auxotroph strain of Neurospora
How to Use This Presentation
 How to Use This Presentation To View the presentation as a slideshow with effects select View on the menu bar and click on Slide Show. To advance through the presentation, click the right-arrow key or
How to Use This Presentation To View the presentation as a slideshow with effects select View on the menu bar and click on Slide Show. To advance through the presentation, click the right-arrow key or
Bioinformatics: Sequence Analysis. COMP 571 Luay Nakhleh, Rice University
 Bioinformatics: Sequence Analysis COMP 571 Luay Nakhleh, Rice University Course Information Instructor: Luay Nakhleh (nakhleh@rice.edu); office hours by appointment (office: DH 3119) TA: Leo Elworth (DH
Bioinformatics: Sequence Analysis COMP 571 Luay Nakhleh, Rice University Course Information Instructor: Luay Nakhleh (nakhleh@rice.edu); office hours by appointment (office: DH 3119) TA: Leo Elworth (DH
Sequence Analysis Lab Protocol
 Sequence Analysis Lab Protocol You will need this handout of instructions The sequence of your plasmid from the ABI The Accession number for Lambda DNA J02459 The Accession number for puc 18 is L09136
Sequence Analysis Lab Protocol You will need this handout of instructions The sequence of your plasmid from the ABI The Accession number for Lambda DNA J02459 The Accession number for puc 18 is L09136
Gene Identification in silico
 Gene Identification in silico Nita Parekh, IIIT Hyderabad Presented at National Seminar on Bioinformatics and Functional Genomics, at Bioinformatics centre, Pondicherry University, Feb 15 17, 2006. Introduction
Gene Identification in silico Nita Parekh, IIIT Hyderabad Presented at National Seminar on Bioinformatics and Functional Genomics, at Bioinformatics centre, Pondicherry University, Feb 15 17, 2006. Introduction
Bioinformatics & Protein Structural Analysis. Bioinformatics & Protein Structural Analysis. Learning Objective. Proteomics
 The molecular structures of proteins are complex and can be defined at various levels. These structures can also be predicted from their amino-acid sequences. Protein structure prediction is one of the
The molecular structures of proteins are complex and can be defined at various levels. These structures can also be predicted from their amino-acid sequences. Protein structure prediction is one of the
9/19/13. cdna libraries, EST clusters, gene prediction and functional annotation. Biosciences 741: Genomics Fall, 2013 Week 3
 cdna libraries, EST clusters, gene prediction and functional annotation Biosciences 741: Genomics Fall, 2013 Week 3 1 2 3 4 5 6 Figure 2.14 Relationship between gene structure, cdna, and EST sequences
cdna libraries, EST clusters, gene prediction and functional annotation Biosciences 741: Genomics Fall, 2013 Week 3 1 2 3 4 5 6 Figure 2.14 Relationship between gene structure, cdna, and EST sequences
I AM NOT A METAGENOMIC EXPERT. I am merely the MESSENGER. Blaise T.F. Alako, PhD EBI Ambassador
 I AM NOT A METAGENOMIC EXPERT I am merely the MESSENGER Blaise T.F. Alako, PhD EBI Ambassador blaise@ebi.ac.uk Hubert Denise Alex Mitchell Peter Sterk Sarah Hunter http://www.ebi.ac.uk/metagenomics Blaise
I AM NOT A METAGENOMIC EXPERT I am merely the MESSENGER Blaise T.F. Alako, PhD EBI Ambassador blaise@ebi.ac.uk Hubert Denise Alex Mitchell Peter Sterk Sarah Hunter http://www.ebi.ac.uk/metagenomics Blaise
Lecture 10. Ab initio gene finding
 Lecture 10 Ab initio gene finding Uses of probabilistic sequence Segmentation models/hmms Multiple alignment using profile HMMs Prediction of sequence function (gene family models) ** Gene finding ** Review
Lecture 10 Ab initio gene finding Uses of probabilistic sequence Segmentation models/hmms Multiple alignment using profile HMMs Prediction of sequence function (gene family models) ** Gene finding ** Review
Ab Initio SERVER PROTOTYPE FOR PREDICTION OF PHOSPHORYLATION SITES IN PROTEINS*
 COMPUTATIONAL METHODS IN SCIENCE AND TECHNOLOGY 9(1-2) 93-100 (2003/2004) Ab Initio SERVER PROTOTYPE FOR PREDICTION OF PHOSPHORYLATION SITES IN PROTEINS* DARIUSZ PLEWCZYNSKI AND LESZEK RYCHLEWSKI BiolnfoBank
COMPUTATIONAL METHODS IN SCIENCE AND TECHNOLOGY 9(1-2) 93-100 (2003/2004) Ab Initio SERVER PROTOTYPE FOR PREDICTION OF PHOSPHORYLATION SITES IN PROTEINS* DARIUSZ PLEWCZYNSKI AND LESZEK RYCHLEWSKI BiolnfoBank
CHAPTER 21 LECTURE SLIDES
 CHAPTER 21 LECTURE SLIDES Prepared by Brenda Leady University of Toledo To run the animations you must be in Slideshow View. Use the buttons on the animation to play, pause, and turn audio/text on or off.
CHAPTER 21 LECTURE SLIDES Prepared by Brenda Leady University of Toledo To run the animations you must be in Slideshow View. Use the buttons on the animation to play, pause, and turn audio/text on or off.
This is the accepted version of this conference paper: Buckingham, Lawrence and Hogan, James and Mann, Scott
 QUT Digital Repository: http://eprints.qut.edu.au/ This is the accepted version of this conference paper: Buckingham, Lawrence and Hogan, James and Mann, Scott and Wirges, Sally (2010) BLAST Atlas : a
QUT Digital Repository: http://eprints.qut.edu.au/ This is the accepted version of this conference paper: Buckingham, Lawrence and Hogan, James and Mann, Scott and Wirges, Sally (2010) BLAST Atlas : a
Annotation and the analysis of annotation terms. Brian J. Knaus USDA Forest Service Pacific Northwest Research Station
 Annotation and the analysis of annotation terms. Brian J. Knaus USDA Forest Service Pacific Northwest Research Station 1 Library preparation Sequencing Hypothesis testing Bioinformatics 2 Why annotate?
Annotation and the analysis of annotation terms. Brian J. Knaus USDA Forest Service Pacific Northwest Research Station 1 Library preparation Sequencing Hypothesis testing Bioinformatics 2 Why annotate?
Applied bioinformatics in genomics
 Applied bioinformatics in genomics Productive bioinformatics in a genome sequencing center Heiko Liesegang Warschau 2005 The omics pyramid: 1. 2. 3. 4. 5. Genome sequencing Genome annotation Transcriptomics
Applied bioinformatics in genomics Productive bioinformatics in a genome sequencing center Heiko Liesegang Warschau 2005 The omics pyramid: 1. 2. 3. 4. 5. Genome sequencing Genome annotation Transcriptomics
Sequence Based Function Annotation. Qi Sun Bioinformatics Facility Biotechnology Resource Center Cornell University
 Sequence Based Function Annotation Qi Sun Bioinformatics Facility Biotechnology Resource Center Cornell University Usage scenarios for sequence based function annotation Function prediction of newly cloned
Sequence Based Function Annotation Qi Sun Bioinformatics Facility Biotechnology Resource Center Cornell University Usage scenarios for sequence based function annotation Function prediction of newly cloned
MB 668 Microbial Bioinformatics and Genome Evolution. 4 credits Spring, 2017
 MB 668 Microbial Bioinformatics and Genome Evolution 4 credits Spring, 2017 Instructors: T. Sharpton, R. Mueller and S. Giovannoni Thomas Sharpton (Microbiology): thomas.sharpton@oregonstate.edu Office
MB 668 Microbial Bioinformatics and Genome Evolution 4 credits Spring, 2017 Instructors: T. Sharpton, R. Mueller and S. Giovannoni Thomas Sharpton (Microbiology): thomas.sharpton@oregonstate.edu Office
UCSC Genome Browser. Introduction to ab initio and evidence-based gene finding
 UCSC Genome Browser Introduction to ab initio and evidence-based gene finding Wilson Leung 06/2006 Outline Introduction to annotation ab initio gene finding Basics of the UCSC Browser Evidence-based gene
UCSC Genome Browser Introduction to ab initio and evidence-based gene finding Wilson Leung 06/2006 Outline Introduction to annotation ab initio gene finding Basics of the UCSC Browser Evidence-based gene
Gene Prediction Group
 Group Ben, Jasreet, Jeff, Jia, Kunal TACCTGAAAAAGCACATAATACTTATGCGTATCCGCCCTAAACACTGCCTTCTTTCTCAA AGAAGATGTCGCCGCTTTTCAACCGAACGATGTGTTCTTCGCCGTTTTCTCGGTAGTGCA TATCGATGATTCACGTTTCGGCAGTGCAGGCACCGGCGCATATTCAGGATACCGGACGCT
Group Ben, Jasreet, Jeff, Jia, Kunal TACCTGAAAAAGCACATAATACTTATGCGTATCCGCCCTAAACACTGCCTTCTTTCTCAA AGAAGATGTCGCCGCTTTTCAACCGAACGATGTGTTCTTCGCCGTTTTCTCGGTAGTGCA TATCGATGATTCACGTTTCGGCAGTGCAGGCACCGGCGCATATTCAGGATACCGGACGCT
Prokaryotic Annotation Pipeline SOP HGSC, Baylor College of Medicine
 1 Abstract A prokaryotic annotation pipeline was developed to automatically annotate draft and complete bacterial genomes. The protein coding genes in the genomes are predicted by the combination of Glimmer
1 Abstract A prokaryotic annotation pipeline was developed to automatically annotate draft and complete bacterial genomes. The protein coding genes in the genomes are predicted by the combination of Glimmer
Chapter 7. Motif finding (week 11) Chapter 8. Sequence binning (week 11)
 Course organization Introduction ( Week 1) Part I: Algorithms for Sequence Analysis (Week 1-11) Chapter 1-3, Models and theories» Probability theory and Statistics (Week 2)» Algorithm complexity analysis
Course organization Introduction ( Week 1) Part I: Algorithms for Sequence Analysis (Week 1-11) Chapter 1-3, Models and theories» Probability theory and Statistics (Week 2)» Algorithm complexity analysis
DNA REPLICATION & BIOTECHNOLOGY Biology Study Review
 DNA REPLICATION & BIOTECHNOLOGY Biology Study Review DNA DNA is found in, in the nucleus. It controls cellular activity by regulating the production of, which includes It is a very long molecule made up
DNA REPLICATION & BIOTECHNOLOGY Biology Study Review DNA DNA is found in, in the nucleus. It controls cellular activity by regulating the production of, which includes It is a very long molecule made up
MATH 5610, Computational Biology
 MATH 5610, Computational Biology Lecture 2 Intro to Molecular Biology (cont) Stephen Billups University of Colorado at Denver MATH 5610, Computational Biology p.1/24 Announcements Error on syllabus Class
MATH 5610, Computational Biology Lecture 2 Intro to Molecular Biology (cont) Stephen Billups University of Colorado at Denver MATH 5610, Computational Biology p.1/24 Announcements Error on syllabus Class
The study of the structure, function, and interaction of cellular proteins is called. A) bioinformatics B) haplotypics C) genomics D) proteomics
 Human Biology, 12e (Mader / Windelspecht) Chapter 21 DNA Which of the following is not a component of a DNA molecule? A) a nitrogen-containing base B) deoxyribose sugar C) phosphate D) phospholipid Messenger
Human Biology, 12e (Mader / Windelspecht) Chapter 21 DNA Which of the following is not a component of a DNA molecule? A) a nitrogen-containing base B) deoxyribose sugar C) phosphate D) phospholipid Messenger
7.1 Techniques for Producing and Analyzing DNA. SBI4U Ms. Ho-Lau
 7.1 Techniques for Producing and Analyzing DNA SBI4U Ms. Ho-Lau What is Biotechnology? From Merriam-Webster: the manipulation of living organisms or their components to produce useful usually commercial
7.1 Techniques for Producing and Analyzing DNA SBI4U Ms. Ho-Lau What is Biotechnology? From Merriam-Webster: the manipulation of living organisms or their components to produce useful usually commercial
M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION
 M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION Chapter 7 Microbial Genetics Lecture prepared by Mindy Miller-Kittrell, University of Tennessee, Knoxville The Structure and Replication
M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION Chapter 7 Microbial Genetics Lecture prepared by Mindy Miller-Kittrell, University of Tennessee, Knoxville The Structure and Replication
Genome annotation & EST
 Genome annotation & EST What is genome annotation? The process of taking the raw DNA sequence produced by the genome sequence projects and adding the layers of analysis and interpretation necessary
Genome annotation & EST What is genome annotation? The process of taking the raw DNA sequence produced by the genome sequence projects and adding the layers of analysis and interpretation necessary
Bioinformatics Translation Exercise
 Bioinformatics Translation Exercise Purpose: The following activity is an introduction to the Biology Workbench, a site that allows users to make use of the growing number of tools for bioinformatics analysis.
Bioinformatics Translation Exercise Purpose: The following activity is an introduction to the Biology Workbench, a site that allows users to make use of the growing number of tools for bioinformatics analysis.
Videos. Lesson Overview. Fermentation
 Lesson Overview Fermentation Videos Bozeman Transcription and Translation: https://youtu.be/h3b9arupxzg Drawing transcription and translation: https://youtu.be/6yqplgnjr4q Objectives 29a) I can contrast
Lesson Overview Fermentation Videos Bozeman Transcription and Translation: https://youtu.be/h3b9arupxzg Drawing transcription and translation: https://youtu.be/6yqplgnjr4q Objectives 29a) I can contrast
Year III Pharm.D Dr. V. Chitra
 Year III Pharm.D Dr. V. Chitra 1 Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Only one strand of DNA serves
Year III Pharm.D Dr. V. Chitra 1 Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Only one strand of DNA serves
ProGen: GPHMM for prokaryotic genomes
 ProGen: GPHMM for prokaryotic genomes Sharad Akshar Punuganti May 10, 2011 Abstract ProGen is an implementation of a Generalized Pair Hidden Markov Model (GPHMM), a model which can be used to perform both
ProGen: GPHMM for prokaryotic genomes Sharad Akshar Punuganti May 10, 2011 Abstract ProGen is an implementation of a Generalized Pair Hidden Markov Model (GPHMM), a model which can be used to perform both
Outline. Introduction to ab initio and evidence-based gene finding. Prokaryotic gene predictions
 Outline Introduction to ab initio and evidence-based gene finding Overview of computational gene predictions Different types of eukaryotic gene predictors Common types of gene prediction errors Wilson
Outline Introduction to ab initio and evidence-based gene finding Overview of computational gene predictions Different types of eukaryotic gene predictors Common types of gene prediction errors Wilson
Outline. Gene Finding Questions. Recap: Prokaryotic gene finding Eukaryotic gene finding The human gene complement Regulation
 Tues, Nov 29: Gene Finding 1 Online FCE s: Thru Dec 12 Thurs, Dec 1: Gene Finding 2 Tues, Dec 6: PS5 due Project presentations 1 (see course web site for schedule) Thurs, Dec 8 Final papers due Project
Tues, Nov 29: Gene Finding 1 Online FCE s: Thru Dec 12 Thurs, Dec 1: Gene Finding 2 Tues, Dec 6: PS5 due Project presentations 1 (see course web site for schedule) Thurs, Dec 8 Final papers due Project
4/6/2015. Bacterial Growth and Nutrition. Nutrients + Oxygen. Temperature. Temperature
 Bacterial Growth and Nutrition ph Nutrients + Oxygen Temperature Temperature 1 Environmental Oxygen Requirements -- can support or hinder growth 1. Aerobic need high oxygen concentration to grow 2. Anaerobic
Bacterial Growth and Nutrition ph Nutrients + Oxygen Temperature Temperature 1 Environmental Oxygen Requirements -- can support or hinder growth 1. Aerobic need high oxygen concentration to grow 2. Anaerobic
Molecular Basis of Inheritance
 Molecular Basis of Inheritance Question 1: Group the following as nitrogenous bases and nucleosides: Adenine, Cytidine, Thymine, Guanosine, Uracil and Cytosine. Answer Nitrogenous bases present in the
Molecular Basis of Inheritance Question 1: Group the following as nitrogenous bases and nucleosides: Adenine, Cytidine, Thymine, Guanosine, Uracil and Cytosine. Answer Nitrogenous bases present in the
Just the Facts: A Basic Introduction to the Science Underlying NCBI Resources
 National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools News About NCBI Site Map
National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools News About NCBI Site Map
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature12212 Supplementary Discussion Contamination Assessment We evaluated the amount of human contamination in our viral DNA preparations by identifying sequences
SUPPLEMENTARY INFORMATION doi:10.1038/nature12212 Supplementary Discussion Contamination Assessment We evaluated the amount of human contamination in our viral DNA preparations by identifying sequences
Section 10.3 Outline 10.3 How Is the Base Sequence of a Messenger RNA Molecule Translated into Protein?
 Section 10.3 Outline 10.3 How Is the Base Sequence of a Messenger RNA Molecule Translated into Protein? Messenger RNA Carries Information for Protein Synthesis from the DNA to Ribosomes Ribosomes Consist
Section 10.3 Outline 10.3 How Is the Base Sequence of a Messenger RNA Molecule Translated into Protein? Messenger RNA Carries Information for Protein Synthesis from the DNA to Ribosomes Ribosomes Consist
Exam 2 Key - Spring 2008 A#: Please see us if you have any questions!
 Page 1 of 5 Exam 2 Key - Spring 2008 A#: Please see us if you have any questions! 1. A mutation in which parts of two nonhomologous chromosomes change places is called a(n) A. translocation. B. transition.
Page 1 of 5 Exam 2 Key - Spring 2008 A#: Please see us if you have any questions! 1. A mutation in which parts of two nonhomologous chromosomes change places is called a(n) A. translocation. B. transition.
COMPUTER RESOURCES II:
 COMPUTER RESOURCES II: Using the computer to analyze data, using the internet, and accessing online databases Bio 210, Fall 2006 Linda S. Huang, Ph.D. University of Massachusetts Boston In the first computer
COMPUTER RESOURCES II: Using the computer to analyze data, using the internet, and accessing online databases Bio 210, Fall 2006 Linda S. Huang, Ph.D. University of Massachusetts Boston In the first computer
THE SYNTHESIS OF LIVING MATTER: ENERGY AND INFORMATION
 THE SYNTHESIS OF LIVING MATTER: ENERGY AND INFORMATION Lectures 10/16, 10/21. Respiration Make a "black box" diagram showing the inputs and outputs of the three main reaction blocks in aerobic respiration;
THE SYNTHESIS OF LIVING MATTER: ENERGY AND INFORMATION Lectures 10/16, 10/21. Respiration Make a "black box" diagram showing the inputs and outputs of the three main reaction blocks in aerobic respiration;
Comparative Genomics. Page 1. REMINDER: BMI 214 Industry Night. We ve already done some comparative genomics. Loose Definition. Human vs.
 Page 1 REMINDER: BMI 214 Industry Night Comparative Genomics Russ B. Altman BMI 214 CS 274 Location: Here (Thornton 102), on TV too. Time: 7:30-9:00 PM (May 21, 2002) Speakers: Francisco De La Vega, Applied
Page 1 REMINDER: BMI 214 Industry Night Comparative Genomics Russ B. Altman BMI 214 CS 274 Location: Here (Thornton 102), on TV too. Time: 7:30-9:00 PM (May 21, 2002) Speakers: Francisco De La Vega, Applied
Degenerate site - twofold degenerate site - fourfold degenerate site
 Genetic code Codon: triple base pairs defining each amino acid. Why genetic code is triple? double code represents 4 2 = 16 different information triple code: 4 3 = 64 (two much to represent 20 amino acids)
Genetic code Codon: triple base pairs defining each amino acid. Why genetic code is triple? double code represents 4 2 = 16 different information triple code: 4 3 = 64 (two much to represent 20 amino acids)
Videos. Bozeman Transcription and Translation: Drawing transcription and translation:
 Videos Bozeman Transcription and Translation: https://youtu.be/h3b9arupxzg Drawing transcription and translation: https://youtu.be/6yqplgnjr4q Objectives 29a) I can contrast RNA and DNA. 29b) I can explain
Videos Bozeman Transcription and Translation: https://youtu.be/h3b9arupxzg Drawing transcription and translation: https://youtu.be/6yqplgnjr4q Objectives 29a) I can contrast RNA and DNA. 29b) I can explain
BIOLOGY 161 EXAM 1 Friday, 6 October 2000 page 1
 BIOLOGY 161 EXAM 1 Friday, 6 October 2000 page 1 Part 1. #1. Please state in a phrase what are the 2 major objectives of this course according to the syllabus? objective #1 - objective #2 - #2. Smoking
BIOLOGY 161 EXAM 1 Friday, 6 October 2000 page 1 Part 1. #1. Please state in a phrase what are the 2 major objectives of this course according to the syllabus? objective #1 - objective #2 - #2. Smoking
GeneMarkS-2: Raising Standards of Accuracy in Gene Recognition
 GeneMarkS-2: Raising Standards of Accuracy in Gene Recognition Alexandre Lomsadze 1^, Shiyuyun Tang 2^, Karl Gemayel 3^ and Mark Borodovsky 1,2,3 ^ joint first authors 1 Wallace H. Coulter Department of
GeneMarkS-2: Raising Standards of Accuracy in Gene Recognition Alexandre Lomsadze 1^, Shiyuyun Tang 2^, Karl Gemayel 3^ and Mark Borodovsky 1,2,3 ^ joint first authors 1 Wallace H. Coulter Department of
BIOL 300 Foundations of Biology Summer 2017 Telleen Lecture Outline
 BIOL 300 Foundations of Biology Summer 2017 Telleen Lecture Outline RNA, the Genetic Code, Proteins I. How RNA differs from DNA A. The sugar ribose replaces deoxyribose. The presence of the oxygen on the
BIOL 300 Foundations of Biology Summer 2017 Telleen Lecture Outline RNA, the Genetic Code, Proteins I. How RNA differs from DNA A. The sugar ribose replaces deoxyribose. The presence of the oxygen on the
ENGR 213 Bioengineering Fundamentals April 25, A very coarse introduction to bioinformatics
 A very coarse introduction to bioinformatics In this exercise, you will get a quick primer on how DNA is used to manufacture proteins. You will learn a little bit about how the building blocks of these
A very coarse introduction to bioinformatics In this exercise, you will get a quick primer on how DNA is used to manufacture proteins. You will learn a little bit about how the building blocks of these
Biology 373, Molecular Biology LAB Time and Location of Class: Materials: Instructor: Dr. Kurt Toenjes Course Introduction and Objectives
 Biology 373, Molecular Biology LAB Time and Location of Class: T 2:00-5:30 or W 2:00-5:30 in SCI 123 Materials: Molecular Biology of the Cell, Fifth edition, Lab handouts Instructor: Dr. Kurt Toenjes,
Biology 373, Molecular Biology LAB Time and Location of Class: T 2:00-5:30 or W 2:00-5:30 in SCI 123 Materials: Molecular Biology of the Cell, Fifth edition, Lab handouts Instructor: Dr. Kurt Toenjes,
BIOINFORMATICS AND SYSTEM BIOLOGY (INTERNATIONAL PROGRAM)
 BIOINFORMATICS AND SYSTEM BIOLOGY (INTERNATIONAL PROGRAM) PROGRAM TITLE DEGREE TITLE Master of Science Program in Bioinformatics and System Biology (International Program) Master of Science (Bioinformatics
BIOINFORMATICS AND SYSTEM BIOLOGY (INTERNATIONAL PROGRAM) PROGRAM TITLE DEGREE TITLE Master of Science Program in Bioinformatics and System Biology (International Program) Master of Science (Bioinformatics
RNA ID missing Word ID missing Word DNA ID missing Word
 Table #1 Vocab Term RNA ID missing Word ID missing Word DNA ID missing Word Definition Define Base pairing rules of A=T and C=G are used for this process DNA duplicates, or makes a copy of, itself. Synthesis
Table #1 Vocab Term RNA ID missing Word ID missing Word DNA ID missing Word Definition Define Base pairing rules of A=T and C=G are used for this process DNA duplicates, or makes a copy of, itself. Synthesis
Genomic Annotation Lab Exercise By Jacob Jipp and Marian Kaehler Luther College, Department of Biology Genomics Education Partnership 2010
 Genomic Annotation Lab Exercise By Jacob Jipp and Marian Kaehler Luther College, Department of Biology Genomics Education Partnership 2010 Genomics is a new and expanding field with an increasing impact
Genomic Annotation Lab Exercise By Jacob Jipp and Marian Kaehler Luther College, Department of Biology Genomics Education Partnership 2010 Genomics is a new and expanding field with an increasing impact
AC Algorithms for Mining Biological Sequences (COMP 680)
 AC-04-18 Algorithms for Mining Biological Sequences (COMP 680) Instructor: Mathieu Blanchette School of Computer Science and McGill Centre for Bioinformatics, 332 Duff Building McGill University, Montreal,
AC-04-18 Algorithms for Mining Biological Sequences (COMP 680) Instructor: Mathieu Blanchette School of Computer Science and McGill Centre for Bioinformatics, 332 Duff Building McGill University, Montreal,
Functional profiling of metagenomic short reads: How complex are complex microbial communities?
 Functional profiling of metagenomic short reads: How complex are complex microbial communities? Rohita Sinha Senior Scientist (Bioinformatics), Viracor-Eurofins, Lee s summit, MO Understanding reality,
Functional profiling of metagenomic short reads: How complex are complex microbial communities? Rohita Sinha Senior Scientist (Bioinformatics), Viracor-Eurofins, Lee s summit, MO Understanding reality,
Tutorial for Stop codon reassignment in the wild
 Tutorial for Stop codon reassignment in the wild Learning Objectives This tutorial has two learning objectives: 1. Finding evidence of stop codon reassignment on DNA fragments. 2. Detecting and confirming
Tutorial for Stop codon reassignment in the wild Learning Objectives This tutorial has two learning objectives: 1. Finding evidence of stop codon reassignment on DNA fragments. 2. Detecting and confirming
Computational Biology 2. Pawan Dhar BII
 Computational Biology 2 Pawan Dhar BII Lecture 1 Introduction to terms, techniques and concepts in molecular biology Molecular biology - a primer Human body has 100 trillion cells each containing 3 billion
Computational Biology 2 Pawan Dhar BII Lecture 1 Introduction to terms, techniques and concepts in molecular biology Molecular biology - a primer Human body has 100 trillion cells each containing 3 billion
Bioinformatics Tools. Stuart M. Brown, Ph.D Dept of Cell Biology NYU School of Medicine
 Bioinformatics Tools Stuart M. Brown, Ph.D Dept of Cell Biology NYU School of Medicine Bioinformatics Tools Stuart M. Brown, Ph.D Dept of Cell Biology NYU School of Medicine Overview This lecture will
Bioinformatics Tools Stuart M. Brown, Ph.D Dept of Cell Biology NYU School of Medicine Bioinformatics Tools Stuart M. Brown, Ph.D Dept of Cell Biology NYU School of Medicine Overview This lecture will
Class XII Chapter 6 Molecular Basis of Inheritance Biology
 Question 1: Group the following as nitrogenous bases and nucleosides: Adenine, Cytidine, Thymine, Guanosine, Uracil and Cytosine. Nitrogenous bases present in the list are adenine, thymine, uracil, and
Question 1: Group the following as nitrogenous bases and nucleosides: Adenine, Cytidine, Thymine, Guanosine, Uracil and Cytosine. Nitrogenous bases present in the list are adenine, thymine, uracil, and
March 2008 Sample Questions Page 1 of 7. Sample Questions for the March Term Test. Biology 022
 March 2008 Sample Questions Page 1 of 7 Sample Questions for the March Term Test Biology 022 1. Given a preparation of isolated chloroplasts, which one of the following treatments would result in the thylakoid
March 2008 Sample Questions Page 1 of 7 Sample Questions for the March Term Test Biology 022 1. Given a preparation of isolated chloroplasts, which one of the following treatments would result in the thylakoid
Sequence Analysis. BBSI 2006: Lecture #(χ+3) Takis Benos (2006) BBSI MAY P. Benos 1
 Sequence Analysis (part III) BBSI 2006: Lecture #(χ+3) Takis Benos (2006) BBSI 2006 31-MAY-2006 2006 P. Benos 1 Outline Sequence variation Distance measures Scoring matrices Pairwise alignments (global,
Sequence Analysis (part III) BBSI 2006: Lecture #(χ+3) Takis Benos (2006) BBSI 2006 31-MAY-2006 2006 P. Benos 1 Outline Sequence variation Distance measures Scoring matrices Pairwise alignments (global,
Ch. 10 Notes DNA: Transcription and Translation
 Ch. 10 Notes DNA: Transcription and Translation GOALS Compare the structure of RNA with that of DNA Summarize the process of transcription Relate the role of codons to the sequence of amino acids that
Ch. 10 Notes DNA: Transcription and Translation GOALS Compare the structure of RNA with that of DNA Summarize the process of transcription Relate the role of codons to the sequence of amino acids that
Genomics and Gene Recognition Genes and Blue Genes
 Genomics and Gene Recognition Genes and Blue Genes November 1, 2004 Prokaryotic Gene Structure prokaryotes are simplest free-living organisms studying prokaryotes can give us a sense what is the minimum
Genomics and Gene Recognition Genes and Blue Genes November 1, 2004 Prokaryotic Gene Structure prokaryotes are simplest free-living organisms studying prokaryotes can give us a sense what is the minimum
M.Sc. Biochemistry (Colleges) onwards BHARATHIAR UNUIVERSITY, COIMBATORE
 Page 1 of 5 SCAA Dt. 24.04.2015 BHARATHIAR UNUIVERSITY, COIMBATORE 654 046. M.SC. BIOCHEMISTRY (Revised papers for the candidates admitted from the academic year 2015-16 onwards) Note: The revised syllabi
Page 1 of 5 SCAA Dt. 24.04.2015 BHARATHIAR UNUIVERSITY, COIMBATORE 654 046. M.SC. BIOCHEMISTRY (Revised papers for the candidates admitted from the academic year 2015-16 onwards) Note: The revised syllabi
GENETICS - CLUTCH CH.15 GENOMES AND GENOMICS.
 !! www.clutchprep.com CONCEPT: OVERVIEW OF GENOMICS Genomics is the study of genomes in their entirety Bioinformatics is the analysis of the information content of genomes - Genes, regulatory sequences,
!! www.clutchprep.com CONCEPT: OVERVIEW OF GENOMICS Genomics is the study of genomes in their entirety Bioinformatics is the analysis of the information content of genomes - Genes, regulatory sequences,
Grundlagen der Bioinformatik Summer Lecturer: Prof. Daniel Huson
 Grundlagen der Bioinformatik, SoSe 11, D. Huson, April 11, 2011 1 1 Introduction Grundlagen der Bioinformatik Summer 2011 Lecturer: Prof. Daniel Huson Office hours: Thursdays 17-18h (Sand 14, C310a) 1.1
Grundlagen der Bioinformatik, SoSe 11, D. Huson, April 11, 2011 1 1 Introduction Grundlagen der Bioinformatik Summer 2011 Lecturer: Prof. Daniel Huson Office hours: Thursdays 17-18h (Sand 14, C310a) 1.1
Computational Gene Finding
 Computational Gene Finding Dong Xu Digital Biology Laboratory Computer Science Department Christopher S. Life Sciences Center University of Missouri, Columbia E-mail: xudong@missouri.edu http://digbio.missouri.edu
Computational Gene Finding Dong Xu Digital Biology Laboratory Computer Science Department Christopher S. Life Sciences Center University of Missouri, Columbia E-mail: xudong@missouri.edu http://digbio.missouri.edu
Tutorial. Whole Metagenome Functional Analysis (beta) Sample to Insight. November 21, 2017
 Whole Metagenome Functional Analysis (beta) November 21, 2017 Sample to Insight QIAGEN Aarhus Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 www.qiagenbioinformatics.com AdvancedGenomicsSupport@qiagen.com
Whole Metagenome Functional Analysis (beta) November 21, 2017 Sample to Insight QIAGEN Aarhus Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 www.qiagenbioinformatics.com AdvancedGenomicsSupport@qiagen.com
2. Materials and Methods
 Identification of cancer-relevant Variations in a Novel Human Genome Sequence Robert Bruggner, Amir Ghazvinian 1, & Lekan Wang 1 CS229 Final Report, Fall 2009 1. Introduction Cancer affects people of all
Identification of cancer-relevant Variations in a Novel Human Genome Sequence Robert Bruggner, Amir Ghazvinian 1, & Lekan Wang 1 CS229 Final Report, Fall 2009 1. Introduction Cancer affects people of all
Genome annotation. Erwin Datema (2011) Sandra Smit (2012, 2013)
 Genome annotation Erwin Datema (2011) Sandra Smit (2012, 2013) Genome annotation AGACAAAGATCCGCTAAATTAAATCTGGACTTCACATATTGAAGTGATATCACACGTTTCTCTAAT AATCTCCTCACAATATTATGTTTGGGATGAACTTGTCGTGATTTGCCATTGTAGCAATCACTTGAA
Genome annotation Erwin Datema (2011) Sandra Smit (2012, 2013) Genome annotation AGACAAAGATCCGCTAAATTAAATCTGGACTTCACATATTGAAGTGATATCACACGTTTCTCTAAT AATCTCCTCACAATATTATGTTTGGGATGAACTTGTCGTGATTTGCCATTGTAGCAATCACTTGAA
Sequence Based Function Annotation
 Sequence Based Function Annotation Qi Sun Bioinformatics Facility Biotechnology Resource Center Cornell University Sequence Based Function Annotation 1. Given a sequence, how to predict its biological
Sequence Based Function Annotation Qi Sun Bioinformatics Facility Biotechnology Resource Center Cornell University Sequence Based Function Annotation 1. Given a sequence, how to predict its biological
Genes & Gene Finding
 Genes & Gene Finding Ben Langmead Department of Computer Science Please sign guestbook (www.langmead-lab.org/teaching-materials) to tell me briefly how you are using the slides. For original Keynote files,
Genes & Gene Finding Ben Langmead Department of Computer Science Please sign guestbook (www.langmead-lab.org/teaching-materials) to tell me briefly how you are using the slides. For original Keynote files,
Q2 (1 point). How many carbon atoms does a glucose molecule contain?
 Q1 (1 point). Name three amino acids that are typically found at the surface of integrated membrane proteins.. Q2 (1 point). How many carbon atoms does a glucose molecule contain? Q3 (1 point). What do
Q1 (1 point). Name three amino acids that are typically found at the surface of integrated membrane proteins.. Q2 (1 point). How many carbon atoms does a glucose molecule contain? Q3 (1 point). What do
Computational gene finding. Devika Subramanian Comp 470
 Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) The biological context Lec 1 Lec 2 Lec 3 Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) The biological context Lec 1 Lec 2 Lec 3 Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
An Evolutionary Optimization for Multiple Sequence Alignment
 195 An Evolutionary Optimization for Multiple Sequence Alignment 1 K. Lohitha Lakshmi, 2 P. Rajesh 1 M tech Scholar Department of Computer Science, VVIT Nambur, Guntur,A.P. 2 Assistant Prof Department
195 An Evolutionary Optimization for Multiple Sequence Alignment 1 K. Lohitha Lakshmi, 2 P. Rajesh 1 M tech Scholar Department of Computer Science, VVIT Nambur, Guntur,A.P. 2 Assistant Prof Department
From assembled genome to annotated genome
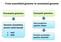 From assembled genome to annotated genome Procaryotic genomes Eucaryotic genomes Genome annotation servers (web based) 1. RAST 2. NCBI Gene prediction pipeline: Maker Function annotation pipeline: Blast2GO
From assembled genome to annotated genome Procaryotic genomes Eucaryotic genomes Genome annotation servers (web based) 1. RAST 2. NCBI Gene prediction pipeline: Maker Function annotation pipeline: Blast2GO
BIOLOGY. Bacteria and Archaea CAMPBELL. Reece Urry Cain Wasserman Minorsky Jackson. Lecture Presentation by Nicole Tunbridge and Kathleen Fitzpatrick
 CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson 27 Bacteria and Archaea Lecture Presentation by Nicole Tunbridge and Kathleen Fitzpatrick Masters of Adaptation Utah s Great Salt
CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson 27 Bacteria and Archaea Lecture Presentation by Nicole Tunbridge and Kathleen Fitzpatrick Masters of Adaptation Utah s Great Salt
Please sign below if you wish to have your grades posted by the last five digits of your SSN
 BIO 226R EXAM II (Sample) PRINT YOUR NAME SSN Please sign below if you wish to have your grades posted by the last five digits of your SSN Signature BIO 226R Exam II has 6 pages, and 27 questions. There
BIO 226R EXAM II (Sample) PRINT YOUR NAME SSN Please sign below if you wish to have your grades posted by the last five digits of your SSN Signature BIO 226R Exam II has 6 pages, and 27 questions. There
Profile HMMs. 2/10/05 CAP5510/CGS5166 (Lec 10) 1 START STATE 1 STATE 2 STATE 3 STATE 4 STATE 5 STATE 6 END
 Profile HMMs START STATE 1 STATE 2 STATE 3 STATE 4 STATE 5 STATE 6 END 2/10/05 CAP5510/CGS5166 (Lec 10) 1 Profile HMMs with InDels Insertions Deletions Insertions & Deletions DELETE 1 DELETE 2 DELETE 3
Profile HMMs START STATE 1 STATE 2 STATE 3 STATE 4 STATE 5 STATE 6 END 2/10/05 CAP5510/CGS5166 (Lec 10) 1 Profile HMMs with InDels Insertions Deletions Insertions & Deletions DELETE 1 DELETE 2 DELETE 3
Lecture 7 Motif Databases and Gene Finding
 Introduction to Bioinformatics for Medical Research Gideon Greenspan gdg@cs.technion.ac.il Lecture 7 Motif Databases and Gene Finding Motif Databases & Gene Finding Motifs Recap Motif Databases TRANSFAC
Introduction to Bioinformatics for Medical Research Gideon Greenspan gdg@cs.technion.ac.il Lecture 7 Motif Databases and Gene Finding Motif Databases & Gene Finding Motifs Recap Motif Databases TRANSFAC
Chapter 17. From Gene to Protein. Slide 1. Slide 2. Slide 3. Gene Expression. Which of the following is the best example of gene expression? Why?
 Slide 1 Chapter 17 From Gene to Protein PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
Slide 1 Chapter 17 From Gene to Protein PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
Teaching Principles of Enzyme Structure, Evolution, and Catalysis Using Bioinformatics
 KBM Journal of Science Education (2010) 1 (1): 7-12 doi: 10.5147/kbmjse/2010/0013 Teaching Principles of Enzyme Structure, Evolution, and Catalysis Using Bioinformatics Pablo Sobrado Department of Biochemistry,
KBM Journal of Science Education (2010) 1 (1): 7-12 doi: 10.5147/kbmjse/2010/0013 Teaching Principles of Enzyme Structure, Evolution, and Catalysis Using Bioinformatics Pablo Sobrado Department of Biochemistry,
