Gentamicin resistance in clinical isolates of Escherichia coll'encoded by genes of veterinary origin
|
|
|
- Kristina Fox
- 5 years ago
- Views:
Transcription
1 J. Med. Microbiol. Vol. 40 (1994), The Pathological Society of Great Britain and Ireland Gentamicin resistance in clinical isolates of Escherichia coll'encoded by genes of veterinary origin A. P. JOHNSON, LOUISE BURNS, N. WOODFORD, E. J. THRELFALL", JAY NAIDOOP, E. M. COOKEP and R. C. GEORGE Antibiotic Reference Unit, Laboratory of Hospital Infection and * Laboratory of Enteric Pathogens, Central Public Health Laboratory, Colindale Avenue, London NW9 5HT Summary. Seven (27 %) of 26 gentamicinresistant human clinical isolates of Escherichia coli were resistant to the veterinary aminoglycoside antibiotic apramycin. A gentamicinresistant Klebsiella pneumoniae isolate from a patient infected with gentamicin/apramycinresistant E. coli was also resistant to apramycin. DNA hybridisation studies showed that all gentamkin/ apramycinresistant isolates contained a gene encoding the enzyme 3Naminoglycoside acetyltransferase type IV (AAC[3]IV) that mediates resistance to gentamicin and apramycin in bacteria isolated from animals. Seven of the eight gentamicin/apramycinresistant isolates were also resistant to the veterinary antihelminthic agent hygromycin B, a phenomenon observed previously in gentamicin/apramycinresistant Enterobacteriaceae isolated from animals. Resistance to gentamicin/apramycin and hygromycin B was cotransferable in six of the isolates. Restriction enzyme analysis of plasmids in apramycinresistant transconjugants derived from and K. pneumoniae isolates from the same patient were virtually identical, suggesting that intergeneric transfer of plasmids encoding apramycin resistance had occurred in vivo. These findings support the view that resistance to gentamicin and apramycin in clinical isolates of results from the spread of resistant organisms from animals to man, with subsequent interstrain or interspecies spread, or both, of resistance genes on transferable plasmids. Introduction Apramycin is an aminoglycoside antibiotic which has been used extensively in veterinary medicine since 1980.l However, it has not been used therapeutically in man. Although early studies indicated that resistance to apramycin was rare in bacteria from either human or veterinary source^,^^ resistant organisms, particularly Escherichia coli and Salmonella spp., have been isolated from the faeces of farm animals treated with apramycin.' Investigation of the mechanism of resistance in isolates from farm animals in the USA showed that the organisms produced a novel aminoglycosidemodifying enzyme subsequently designated (3)Naminoglycoside acetyltransferase type IV (AAC [3]IV).5 This enzyme acetylated not only apramycin but other aminoglycosides including gentamicin and tobramycin, which are used to treat serious infections in man.j Subsequent studies have shown a similar mechanism of resistance in apramycinresistant Salmonella spp. and isolated from farm animals in the UK1*637 and France.' Received 8 June 1993; accepted 17 Sept t Present address : Unilever Research, Port Sunlight Laboratory, Wirral L63 3JW. $ Present address : Public Health Laboratory Service HQ, Colindale Avenue, London NW9 5DF. Downloaded from by IP: The observation that apramycin usage in veterinary medicine may result in carriage and faecal excretion by farm animals of enteric bacteria crossresistant to gentamicin, gave rise to concern that these resistant bacteria might spread to man. Indeed, in 1986, Threlfall and coworkers reported the isolation of S. typhimurium phage type 204c resistant to apramycin and gentamicin from both cattle and man.6 Subsequently, other species of Enterobacteriaceae resistant to apramycin and gentamicin due to production of AAC(3)IV have been isolated from patients in hospitals in Belgi~m,~ Spain'' and the UK.ll In view of this, a collection of gentamicinresistant Enterobacteriaceae obtained from patients in the UK were assessed for resistance to apramycin, and resistant organisms detected were studied with regard to the mechanism and genetics of their resistance. Materials and methods Apramycinresistant isolates Twentysix nonfaecal clinical isolates of gentamicinresistant referred to the Laboratory of Hospital Infection during were screened for resistance to apramycin by an agar incorporation
2 222 A. P. JOHNSON ET AL. Table I. Clinical isolates of gentamicin/apramycinresistant Enterobacteriaceae Species Isolate no. Patient Location Source 0 Serogroup Resistance profile K. pneumoniae El 79 K180 E444 E545 E548 E550 E597 E635 A A B C D E F G Taunton Taunton Edinburgh Southampton Biliary drainage tube Biliary drainage tube Not known 83 Gm, Apr, Tob, Hyg NT Gm, Apr, Tob, Hyg, Amp Gm, Apr, Tob, Hyg, Amp, Tet, Trm Gm, Apr, Tob, Hyg, Amp, Tet, Trm Gm, Apr, Tob, Hyg, Tet Gm, Apr, Tob, Hyg, Tet Gm, Apr, Tob, Hyg, Amp Gm, Apr, Tob, Hyg, Amp, Trm Gm, gentamicin ; Apr, apramycin; Tob, tobramycin; Hyg, hygromycin B ; Amp, ampicillin ; Tet, tetracycline; Trm, trimethoprim ; NT, nont ypable, breakpoint method with apramycin at a concentration of 16 mg/l. Seven isolates were found to be resistant. A gentamicinresistant isolate of Klebsiellapneumoniae obtained simultaneously with gentamicin/apramycinresistant from one patient was also found to be resistant to apramycin. The sources of the eight apramycinresistant isolates together with their serotypes and antibiograms are given in table I. Determination of minimum inhibitory concentrations (MICS) MICs of several antimicrobial agents, including apramycin and hygromycin B (kindly provided by Lilly Research Laboratories), were determined by an agar dilution method in Isosensitest agar supplemented with lysed horse blood 2 YO v/v. Serial twofold dilutions of each antimicrobial agent were incorporated into the medium and plates were inoculated by a multipoint inoculator (Diamed Diagnostics) with an inoculum of cfu/spot. Plasmid analysis Plasmids were extracted by the method of Kado and Liu12 and separated by electrophoresis in agarose % w/v gels. The molecular sizes of plasmids were estimated by comparison with plasmids of known size. In some experiments, extracted plasmids were digested with restriction endonucleases under conditions specified by the enzyme manufacturer (Boehringer Mannheim), and the resulting fragments were separated by agarose gel electrophoresis. The sizes of restriction fragments were determined by comparison with fragments of linear DNA of known size (DNA mo1.wt markers I and 11; Boehringer Mannheim). Transfer of apramycin resistance Conjugation experiments were performed overnight in stationary broth culture at 37 C. The recipient organism was strain 14R525 which is plasmidfree and resistant to nalidixic acid. Transconjugants were detected by plating the mixture of donor and recipient organisms on nutrient agar containing nalidixic acid 50 mg/l and apramycin 16 mg/l. IP: Probe for gene encoding AAC(3)IV The DNA probe specific for the gene encoding production of AAC(3)IV consisted of a 740bp Sac1 fragment of plasmid pwp Plasmid pwp70 1 was purified on a Qiagen column (Qiagen pack 500, Diagen) and digested overnight with Sac I. The 740bp fragment was separated by electrophoresis through an agarose 1 YO w/v gel, extracted from the gel by Prepa Gene (BioRad), and labelled with digoxigenin from a commercially available kit (Nonradioactive DNA Labelling and Detection Kit, Boehringer Mannheim). Preparation of DNA for hybridisation studies For dotblot assays, DNA extracted from bacteria as described previo~slyl~ was denatured by heating at 95 C for 10 min, cooled on ice and spotted on to nylon membranes (Hybond, Amersham), and allowed to dry in air. The membranes were baked at 80 C for 2 h then stored at room temperature until required. For Southern blot analysis of plasmid DNA, plasmids were extracted and subjected to gel electrophoresis as described above, then transferred to nylon membranes with vacuum blotting equipment (Vacugene, Pharmacia LKB). The membranes were baked at 80 C for 2 h to fix the DNA and then stored at room temperature. Results Ant im icro bial resis tance The MICs of apramycin, gentamicin, tobramycin and hygromycin B for the eight clinical isolates are shown in table 11. All eight isolates were resistant to apramycin (MIC 2 mg/l), gentamicin (MIC 16 mg/l) and tobramycin (MIC > mg/l) but were sensitive to amikacin (MIC < 2 mg/l). All of the clinical isolates had MICs of hygromycin B of > mg/l with the exception of isolate E635 which had an MIC of hygromycin B of 64 mg/l. One isolate (E550) lost resistance to aminoglycosides during subculture in the laboratory. This sensitive variant was designated E550S. The apramycinsensitive variant E550S, strain 14R525
3 GENTAMICIN RESISTANCE FROM VETERINARY BACTERIA 223 Table 11. Antimicrobial resistance and DNA hybridisation reactions of apramycinresistant clinical isolates and transconjugants. Organism El79 El79 Tc K180 K180 Tc E444 E444 Tc E548 E548 Tc E550 E550 Tc E550S E597 E597 Tc E545 E635 14R525 MIC (mg/l) of Hybridisation of probe* apramycin gentamicin tobramycin hygromycin B Wholecell DNAt Plasmid DNA$ Tc, transconjugant. * 740bp Sac1 fragment of plasmid ~WP701.l~ t Dot blots. $ Southern blots of plasmid DNA. and four other apramycinsensitive clinical isolates of had MICs of hygromycin B in the range 64 mg/l. There was interisolate variation with regard to susceptibility to tetracycline, ampicillin and trimethoprim (table I) but all the isolates were susceptible to ciprofloxacin, chloramphenicol, cefuroxime, cefotaxime and ceftazidime. Plasmid content of clinical isolates With the exception of isolates El79 and K180, all the gentamicin/apramycinresistant isolates exhibited distinct plasmid profiles ; the plasmids from isolates from the same geographical area varied with regard to both number and molecular sizes. Isolates El79 and K180, which were isolated from the same clinical source, each contained a single plasmid of c. 90 kb. All the other isolates contained either three or four plasmids which ranged in size from 1.5 kb to c. 90 kb. Transfer of apramycin resistance Apramycinresistant transconjugants were obtained from six of the eight clinical isolates (E179, K180, E444, E548, E550 and E597) (table 11). All the apramycinresistant transconjugants were resistant to gentamicin, tobramycin and hygromycin B (table 11). Plasmid analysis showed that transconjugants obtained from three isolates (E179, K180 and E597) contained a single high mol. wt plasmid of c. 90 kb. Transconjugants from isolate E548 contained two plasmids of c. 90 kb and 4.5 kb, whereas transconjugants from isolate E550 contained two plasmids of c. 90 kb and 68 kb. In contrast, a series of transconjugants from isolate E444 contained two or IP: three plasmids, 5070 kb in size, many of which did not align with plasmids present in the donor, presumably reflecting molecular reorganisation of the plasmids during conjugation. For further study of this isolate, one transconjugant was chosen which contained two plasmids of c. 50 kb and 60 kb respectively, the larger of which aligned with a plasmid present in the donor. Restriction enzyme digestion analysis showed that the single plasmids present in the transconjugants derived from isolates El79 and K180 were virtually identical. EcoRI digests of both plasmids each comprised a series of 14 similar fragments, from 1.2 to 23 kb in size (figure). The plasmid in the transconjugant derived from isolate El79 differed from the plasmid in the transconjugant from isolate K180 in that the latter plasmid contained a unique EcoRI fragment of c. 4 kb (figure). A similar type of result was obtained with ClaI. In contrast, the EcoRI and ClaI digestion profiles of the single plasmid in the transconjugant from isolate E597 appeared quite distinct (data not shown). Hy br idisa t ion studies In dotblot assays, the DNA probe specific for the gene encoding AAC(3)IV hybridised with DNA extracted from each of the apramycinresistant clinical isolates. DNA extracted from five isolates of sensitive to apramycin and the apramycinsensitive variant of isolate E550 (E550S) failed to hybridise with the probe. When Southern blots of plasmid preparations of the apramycinresistant clinical isolates were tested, a single plasmid in each of six isolates (E179, K180, E545, E548, E550 and E635) hybridised with the
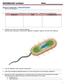
4 224 A. P. JOHNSON ET AL. Figure. EcoRI digests of plasmids of apramycinresistant transconjugants from isolates El79 and K180. Lane 1, molecular size (kb) markers ; 2, transconjugant derived from El 79 ; 3, transconjugant derived from K 180. probe. The size of the plasmids that hybridised with the probe ranged from c. 70 to 90 kb. With two isolates (E444 and E597), hybridisation of the probe with plasmid DNA was not observed. In subsequent experiments, Southern blots of plasmids from isolates exhibiting transferable resistance together with plasmids from their respective transconjugants were examined. A single plasmid of c. 90 kb that hybridised with the probe was observed in isolates E179, K180, E548, E550 and their respective transconjugants. As before, no hybridisation was observed with plasmids from isolates E444 and E597, but a plasmid of c. 70 kb in the transconjugant from isolate E444, and a plasmid of c. 90 kb in the transconjugant from isolate E597 did hybridise with the probe. Discussion Studies of gent amicin/ apramycinresist an t Enterobacteriaceae isolated from animals have shown that resistance is due to production of the enzyme AAC(3)IV.58 Molecular analysis of an apramycinresistant Salmonella sp. of animal origin13 further showed that the gene encoding AAC(3)IV is closely linked to a gene encoding resistance to the aminocyclitol antibiotic hygromycin B, which has been used as an antihelminthic agent in farm animals. Therefore, it was interesting to note that each of the eight clinical isolates of gentamicin/apramycinresistant Enterobacteriaceae described in the present study hybridised with a DNA probe specific for the gene encoding AAC(3)IV, and that seven of the eight isolates were also resistant to hygromycin B. Resistance to apramycin was transferable in six of the eight clinical isolates. A DNA probe specific for the gene encoding the enzyme AAC(3)IV hybridised with individual plasmids present in four isolates with transferable resistance, and also with plasmids present in their respective transconjugants. In contrast, although the probe reacted with wholecell DNA of two further isolates with transferable resistance, no hybridisation was observed with extracted plasmid DNA, suggesting a chromosomal location for the gene. The observation that the probe did hybridise with plasmid DNA in transconjugants derived from the latter two isolates suggests transposition of the gene from the chromosome to a plasmid during conjugation. Several workers have reported insertion sequences associated with the genes encoding resistance to apramycin and hygromycin B,l09l3 which is compatible with these observations. With the six isolates in which apramycin resistance was found to be transferable, resistant transconjugants always acquired resistance to hygromycin B. In transconjugants produced by three donor isolates, only a single plasmid was transferred suggesting that the genes for resistance to apramycin and hygromycin B were linked on the same plasmid. These observations are in agreement with the previously reported finding that the genes for resistance to apramycin and hygromycin B are in the same operon and would thus be transferred jointly," but with transconjugants from other donors such linkage could not be inferred as more than one plasmid was transferred. However, one isolate (E550), which transferred more than one plasmid in mating experiments, gave rise to a gentamicin/apramycinsensitive variant on subculture. This variant, which had lost a single plasmid, also became sensitive to hygromycin B, indicating that both resistance genes were encoded by this plasmid. As apramycin and hygromycin B have never been used in human medicine, the most likely explanation for the emergence of resistance to gentamicin and apramycin due to production of AAC(3)IV in human isolates of Enterobacteriaceae, and in particular E. coli, is that the genetic determinant of resistance has been acquired either directly or indirectly from gentamicin/apramycinresistant bacteria of veterinary origin. For example, apramycinresistant S. typhimurium phage type 204c originally isolated from cattle, was also subsequently isolated from man6 Although infection or colonisation of man with such organisms may be transient, the genes encoding apramycin resistance may be transferred by conjugation to other strains of the Enterobacteriaceae found in the normal human intestinal flora. Such intergeneric transfer of a plasmid encoding resistance to apramycin has recently been reported in cattle.' In the present study, restriction enzyme analysis of the plasmids encoding resistance to apramycin and hygromycin B in the E. coli (E179) and K. pneumoniae (K180) isolates from IP:
5 GENTAMICIN RESISTANCE FROM VETERINARY BACTERIA 225 the same patient were very similar, suggesting that intergeneric plasmid transfer had occurred in a human host. An alternative explanation for the emergence of gentamicin/apramycin resistance resulting from the production of AAC(3)IV in bacteria in man, is that the resistance trait evolved independently from that seen in bacteria from animals, under the selection pressure of gentamicin and tobramycin usage in clinical medicine. According to this hypothesis, it is reasonable to expect that other species of Enterobacteriaceae besides and Salmonella spp. would also exhibit resistance to gentamicin and apramycin resulting from the production of AAC(3)IV. However, this does not seem to be the case, as shown by analysis of isolates of Klebsiella and Enterobacter spp. referred to the Laboratory of Hospital Infection from UK hospitals. Between 1987 and 1991, only two (0.6%) of 306 gentamicinresistant Klebsiella spp. (not including isolate K180) and one (0.7%) of 144 gentamicinresistant Enterobacter spp. were also resistant to apramycin (R. C. George, unpublished observations). Similarly, Lovering et ai.l5 reported that while apramycin resistance was detected among two of 18 gentamicinresistant isolates of, it was not detected among 72 isolates of other genera including Klebsiella, Citrobacter, Enterobacter, Serratia, Proteus and Procidencia spp. Thus, apramycin resistance among Enterobacteriaceae isolated from man is found predominantly among '9 11* l5 and Salmonella ~pp.,~ which are species known to spread from animals to man. We have insufficient clinical and epidemiological data to determine whether the patients infected with the gentamicin/apramycinresistant E. coli reported here acquired the infecting organisms either directly or indirectly from veterinary sources or whether the isolates were components of the patients' own bacterial flora that had acquired resistance genes from other organisms (e.g., Salmonella spp.) of veterinary origin. Although some of the isolates belonged to serogroups 08, 075 and 0117, which occur commonly in animals (Dr C. Wray, personal communication), these serogroups are also common among human clinical isolates (Dr T. Cheasty, personal communication). The seven clinical isolates of gentamicin/ apramycinresistant reported here belonged to various serogroups, showed distinct plasmid and antibioticresistance profiles and were from various geographical locations, suggesting that they were not related epidemiologically. Although the number of isolates described here is small, resistance to apramycin is not routinely investigated in clinical laboratories, and therefore, the extent of the problem is unknown. However, it should be noted that in a recent study of gentamicinresistant isolated in a hospital in Liverpool, 26 YO of the isolates (a figure similar to that noted in the present study) were resistant to apramycin.'l The incidence of resistance to gentamicin remains relatively low in bacteraemia isolates of in the UK (12 l6 but the fact that clinical isolates of bacteria resistant to gentamicin and apramycin resulting from the production of AAC(3)IV have also been detected in Belgiumg and Spain'' suggests that the problem may be widespread. Clearly, further work is needed to study the epidemiology of the problem, both in human clinical and veterinary settings. We thank Dr W. Piepersberg, University of Munich, for providing plasmid pwp701 and Dr T. Cheasty, Laboratory of Enteric Pathogens, Central Public Health Laboratory, for typing the isolates. We also thank Dr C. Wray, Central Veterinary Laboratory, Weybridge for helpful discussions. An abstract describing part of this work has been published in the Proceedings of the 5th European Congress of Clinical Microbiology and Infectious Diseases (Oslo, September 1991). References 1. Wray C, Hedges RW, Shannon KP, Bradley DE. Apramycin and gentamicin resistance in Escherichia coli and salmonellas isolated from farm animals. J Hyg 1986; 97: Wick WE, Welles JS. Nebramycin, a new broadspectrum antibiotic complex. In : Hobby GL (ed) Antimicrobial agents and chemotherapy1967. Ann Arbor Michigan, American Society for Microbiology : Ryden R, Moore BJ. The in vitro of apramycin, a new aminocyclitol antibiotic. J Antimicrob Chemother 1977; 3: ChaslusDancla E, Lafont JP. Resistance to gentamicin and apramycin in Escherichia coli from calves in France. Vet Rec 1985; 117: 9G Davies J, O'Connor S. Enzymatic modification of aminoglycoside antibiotics : 3Naminoacetyltransferase with broad specificity that determines resistance to the novel aminoglycoside apramycin. Antimicrob Agents Chemother 1978 ; 14: Threlfall EJ, Rowe B, Ferguson JL, Ward LR. Characterization of plasmids conferring resistance to gentamicin and apramycin in strains of Salmonella typhimurium phage type 204c isolated in Britain. J Hyg 1986; 97: Hunter JEB, Shelley JC, Walton JR, Hart CA, Bennett M. Apramycin resistance plasmids in Escherichia coli : possible IP: transfer to Salmonella typhimurium in calves. Epidemiol Infect 1992; 108: ChaslusDancla E, Martel JL, Carlier C, Lafont JP, CourvalinP. Emergence of aminoglycoside 3Nacetyltransferase IV in Escherichia coli and Salmonella typhimurium isolated from animals in France. Antimicrob Agents Chemother 1986; 29: ChaslusDancla E, Glupczynski Y, Gerbaud G, Lagorce M, Lafont JP, Courvalin P. Detection of apramycin resistant Enterobacteriaceae in hospital isolates. FEMS Microbiol Lett 1989; : Salauze D, Otal I, GomezLus R, Davies J. Aminoglycoside acetyltransferase 3IV (aacc4) and hygromycin B 4 1 phosphotransferase (hphb) in bacteria isolated from human and animal sources. Antimicrob Agents Chemother 1990; 34: Hunter JEB, Hart CA, Shelley JC, Walton JR, Bennett M. Human isolates of apramycinresistant Escherichia coli which contain the genes for the AAC(3)IV enzyme. Epidemiol Infect 1993; 110: Kado CI, Liu ST. Rapid procedure for detection and isolation of large and small plasmids. J Bacteriol 1981; 145: Brau B, Pilz U, Piepersberg W. Genes for gentamicin(3)nacetyltransferases I11 and IV: Nucleotide sequence of the AAC(3)IV gene and possible involvement of an IS 140 element in its expression. Mol Gen Genet 1984; 193:
6 226 A. P. JOHNSON ET AL. 14. Moxon ER, Deich RA, Connelly C. Cloning of chromosomal DNA from Haemophilus injuenzae. Its use for studying the expression of type 6 capsule and virulence. J Clin Znuest 1984; 73: Lovering AM, Bywater MJ, Holt HA, Champion HM, Reeves DS. Resistance of bacterial pathogens to four amino 36: glycosides and six other antibacterial and prevalence of aminoglycoside modifying enzymes, in 20 UK centres. J Antimicrob Chemother 1988; 22: George RC, Norbury PB, James D. Surveillance of antibiotic resistance in England and Wales. J Med Microbiol 1992; IP:
CHAPTER 2A HOW DO YOU BEGIN TO CLONE A GENE? CHAPTER 2A STUDENT GUIDE 2013 Amgen Foundation. All rights reserved.
 CHAPTER 2A HOW DO YOU BEGIN TO CLONE A GENE? 35 INTRODUCTION In the Program Introduction, you learned that the increase in diabetes in the United States has resulted in a great demand for its treatment,
CHAPTER 2A HOW DO YOU BEGIN TO CLONE A GENE? 35 INTRODUCTION In the Program Introduction, you learned that the increase in diabetes in the United States has resulted in a great demand for its treatment,
Molecular Genetics Techniques. BIT 220 Chapter 20
 Molecular Genetics Techniques BIT 220 Chapter 20 What is Cloning? Recombinant DNA technologies 1. Producing Recombinant DNA molecule Incorporate gene of interest into plasmid (cloning vector) 2. Recombinant
Molecular Genetics Techniques BIT 220 Chapter 20 What is Cloning? Recombinant DNA technologies 1. Producing Recombinant DNA molecule Incorporate gene of interest into plasmid (cloning vector) 2. Recombinant
An estimate of the physical distance between two linked markers in Haemophilus influenzae
 J. Biosci., Vol. 13, No. 3, September 1988, pp. 223 228. Printed in India. An estimate of the physical distance between two linked markers in Haemophilus influenzae Ε. Β. SAMIWALA, VASUDHA P. JOSHI and
J. Biosci., Vol. 13, No. 3, September 1988, pp. 223 228. Printed in India. An estimate of the physical distance between two linked markers in Haemophilus influenzae Ε. Β. SAMIWALA, VASUDHA P. JOSHI and
Determination of Pseudomonas aeruginosa by Biochemical Test Methods Test, a Modified Biochemical Test for
 Japan. J. Microbiol. Vol. 14 (4), 279-284, 1970 Determination of Pseudomonas aeruginosa II. Acylamidase by Biochemical Test Methods the Identification Test, a Modified Biochemical Test for of Pseudomonas
Japan. J. Microbiol. Vol. 14 (4), 279-284, 1970 Determination of Pseudomonas aeruginosa II. Acylamidase by Biochemical Test Methods the Identification Test, a Modified Biochemical Test for of Pseudomonas
Lecture Four. Molecular Approaches I: Nucleic Acids
 Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
The role of PHE s AMRHAI Reference Unit
 The role of PHE s AMRHAI Reference Unit Professor Neil Woodford Antimicrobial Resistance & Healthcare Associated Infections (AMRHAI) Reference Unit Crown copyright What does AMRHAI offer? Susceptibility
The role of PHE s AMRHAI Reference Unit Professor Neil Woodford Antimicrobial Resistance & Healthcare Associated Infections (AMRHAI) Reference Unit Crown copyright What does AMRHAI offer? Susceptibility
2Departament Patologt'a i
 Epidemiol. Infect. (1994), 112. 253-261 253 Copyright (C 1994 Cambridge University Press Insertion sequence IS200 fingerprinting of Salmonella typhi: an assessment of epidemiological applicability E. J.
Epidemiol. Infect. (1994), 112. 253-261 253 Copyright (C 1994 Cambridge University Press Insertion sequence IS200 fingerprinting of Salmonella typhi: an assessment of epidemiological applicability E. J.
Title: Description of the First Escherichia coli Clinical Isolate Harboring Colistin-
 AAC Accepted Manuscript Posted Online 24 October 2016 Antimicrob. Agents Chemother. doi:10.1128/aac.01945-16 Copyright 2016, American Society for Microbiology. All Rights Reserved. 1 2 Title: Description
AAC Accepted Manuscript Posted Online 24 October 2016 Antimicrob. Agents Chemother. doi:10.1128/aac.01945-16 Copyright 2016, American Society for Microbiology. All Rights Reserved. 1 2 Title: Description
2054, Chap. 13, page 1
 2054, Chap. 13, page 1 I. Microbial Recombination and Plasmids (Chapter 13) A. recombination = process of combining genetic material from 2 organisms to produce a genotype different from either parent
2054, Chap. 13, page 1 I. Microbial Recombination and Plasmids (Chapter 13) A. recombination = process of combining genetic material from 2 organisms to produce a genotype different from either parent
Recombinant DNA Technology. The Role of Recombinant DNA Technology in Biotechnology. yeast. Biotechnology. Recombinant DNA technology.
 PowerPoint Lecture Presentations prepared by Mindy Miller-Kittrell, North Carolina State University C H A P T E R 8 Recombinant DNA Technology The Role of Recombinant DNA Technology in Biotechnology Biotechnology?
PowerPoint Lecture Presentations prepared by Mindy Miller-Kittrell, North Carolina State University C H A P T E R 8 Recombinant DNA Technology The Role of Recombinant DNA Technology in Biotechnology Biotechnology?
SXT-related integrating conjugative element and IncC plasmids in Vibrio cholerae O1 strains in Eastern Africa
 Journal of Antimicrobial Chemotherapy Advance Access published January 20, 2009 Journal of Antimicrobial Chemotherapy doi:10.1093/jac/dkn542 SXT-related integrating conjugative element and IncC plasmids
Journal of Antimicrobial Chemotherapy Advance Access published January 20, 2009 Journal of Antimicrobial Chemotherapy doi:10.1093/jac/dkn542 SXT-related integrating conjugative element and IncC plasmids
Supporting Information-Tables
 Supporting Information-Tables Table S1. Bacterial strains and plasmids used in this work Bacterial strains Description Source of reference Streptococcus pneumoniae 1 Cp1015 non-capsulated and βl susceptible
Supporting Information-Tables Table S1. Bacterial strains and plasmids used in this work Bacterial strains Description Source of reference Streptococcus pneumoniae 1 Cp1015 non-capsulated and βl susceptible
BACTERIAL CONJUGATION. To demonstrate the technical procedure to monitor the conjugational transfer of genetic material from one cell to another.
 BACTERIAL CONJUGATION I. OBJECTIVES To demonstrate the technical procedure to monitor the conjugational transfer of genetic material from one cell to another. To learn about the various genetic elements
BACTERIAL CONJUGATION I. OBJECTIVES To demonstrate the technical procedure to monitor the conjugational transfer of genetic material from one cell to another. To learn about the various genetic elements
Detection of aac(6 0 )-Ib-cr in KPC-producing Klebsiella pneumoniae isolates from Tel Aviv, Israel
 Journal of Antimicrobial Chemotherapy (2009) 64, 718 722 doi:10.1093/jac/dkp272 Advance Access publication 4 August 2009 Detection of aac(6 0 )-Ib-cr in KPC-producing Klebsiella pneumoniae isolates from
Journal of Antimicrobial Chemotherapy (2009) 64, 718 722 doi:10.1093/jac/dkp272 Advance Access publication 4 August 2009 Detection of aac(6 0 )-Ib-cr in KPC-producing Klebsiella pneumoniae isolates from
M. Ben-David 1, O. Hammer 1, A.Scinderman 1, Y. Gluckman-Yavo 1, M. Fridman 1, D. Gohman 1, G. Ingber 1 and E. Zahavy 2
 441 Rapid Gram Negative Antimicrobial Susceptibility Testing Directly from Positive Blood Culture based on a Unique Spectral Intensity Ratio Analysis via Single Fluorescence Membrane Dye Staining and Flow
441 Rapid Gram Negative Antimicrobial Susceptibility Testing Directly from Positive Blood Culture based on a Unique Spectral Intensity Ratio Analysis via Single Fluorescence Membrane Dye Staining and Flow
DEC PCR KIT. SSI Diagnostica
 DEC PCR KIT SSI Diagnostica 2 DEC PCR Kit for PCR Detection of Diarrhoeagenic E. coli (DEC) for in vitro diagnostic use Application The DEC PCR Kit is for in vitro diagnostic PCR detection of diarrhoeagenic
DEC PCR KIT SSI Diagnostica 2 DEC PCR Kit for PCR Detection of Diarrhoeagenic E. coli (DEC) for in vitro diagnostic use Application The DEC PCR Kit is for in vitro diagnostic PCR detection of diarrhoeagenic
Recombinant DNA Technology
 Recombinant DNA Technology Common General Cloning Strategy Target DNA from donor organism extracted, cut with restriction endonuclease and ligated into a cloning vector cut with compatible restriction
Recombinant DNA Technology Common General Cloning Strategy Target DNA from donor organism extracted, cut with restriction endonuclease and ligated into a cloning vector cut with compatible restriction
Reading Lecture 3: 24-25, 45, Lecture 4: 66-71, Lecture 3. Vectors. Definition Properties Types. Transformation
 Lecture 3 Reading Lecture 3: 24-25, 45, 55-66 Lecture 4: 66-71, 75-79 Vectors Definition Properties Types Transformation 56 VECTORS- Definition Vectors are carriers of a DNA fragment of interest Insert
Lecture 3 Reading Lecture 3: 24-25, 45, 55-66 Lecture 4: 66-71, 75-79 Vectors Definition Properties Types Transformation 56 VECTORS- Definition Vectors are carriers of a DNA fragment of interest Insert
Suggest a technique that could be used to provide molecular evidence that all English Elm trees form a clone. ... [1]
![Suggest a technique that could be used to provide molecular evidence that all English Elm trees form a clone. ... [1] Suggest a technique that could be used to provide molecular evidence that all English Elm trees form a clone. ... [1]](/thumbs/77/74807071.jpg) 1 Molecular evidence E Ulmus procera, form a genetically isolated clone. English Elms developed from a variety of elm brought to Britain from Rome in the first century A.D. Although English Elm trees make
1 Molecular evidence E Ulmus procera, form a genetically isolated clone. English Elms developed from a variety of elm brought to Britain from Rome in the first century A.D. Although English Elm trees make
Prevalence and molecular characterization of clinical isolates of Escherichia coli expressing an AmpC phenotype
 J Antimicrob Chemother 2010; 65: 460 464 doi:10.1093/jac/dkp484 Advance publication 22 January 2010 Prevalence and molecular characterization of clinical isolates of Escherichia coli expressing an AmpC
J Antimicrob Chemother 2010; 65: 460 464 doi:10.1093/jac/dkp484 Advance publication 22 January 2010 Prevalence and molecular characterization of clinical isolates of Escherichia coli expressing an AmpC
Lab 5/5a Transformation of E. coli with a Recombinant Plasmid
 Lab 5/5a Transformation of E. coli with a Recombinant Plasmid Lab 2 Pre Lab Readiness Familiarity and Proper use of micropipettes Remember the 1 st and 2 nd stops Aseptic Technique Antibiotic Resistance
Lab 5/5a Transformation of E. coli with a Recombinant Plasmid Lab 2 Pre Lab Readiness Familiarity and Proper use of micropipettes Remember the 1 st and 2 nd stops Aseptic Technique Antibiotic Resistance
Mutagenesis for Studying Gene Function Spring, 2007 Guangyi Wang, Ph.D. POST103B
 Mutagenesis for Studying Gene Function Spring, 2007 Guangyi Wang, Ph.D. POST103B guangyi@hawaii.edu http://www.soest.hawaii.edu/marinefungi/ocn403webpage.htm Overview of Last Lecture DNA microarray hybridization
Mutagenesis for Studying Gene Function Spring, 2007 Guangyi Wang, Ph.D. POST103B guangyi@hawaii.edu http://www.soest.hawaii.edu/marinefungi/ocn403webpage.htm Overview of Last Lecture DNA microarray hybridization
Hosted by Paul Webber page 1
 A Webber Training Teleclass April 8, 24 Bacterial Typing Methods and Their Value in Infection Control Giles Edwards Consultant Microbiologist Stobhill Hospital, Glasgow, UK Scottish MRSA Reference Laboratory
A Webber Training Teleclass April 8, 24 Bacterial Typing Methods and Their Value in Infection Control Giles Edwards Consultant Microbiologist Stobhill Hospital, Glasgow, UK Scottish MRSA Reference Laboratory
Relationship between Adhesion to Intestinal Caco-2 Cells and Multidrug Resistance in Klebsiella pneumoniae Clinical Isolates
 JOURNAL OF CLINICAL MICROBIOLOGY, June 1997, p. 1499 1503 Vol. 35, No. 6 0095-1137/97/$04.00 0 Copyright 1997, American Society for Microbiology Relationship between Adhesion to Intestinal Caco-2 Cells
JOURNAL OF CLINICAL MICROBIOLOGY, June 1997, p. 1499 1503 Vol. 35, No. 6 0095-1137/97/$04.00 0 Copyright 1997, American Society for Microbiology Relationship between Adhesion to Intestinal Caco-2 Cells
Genetic Engineering & Recombinant DNA
 Genetic Engineering & Recombinant DNA Chapter 10 Copyright The McGraw-Hill Companies, Inc) Permission required for reproduction or display. Applications of Genetic Engineering Basic science vs. Applied
Genetic Engineering & Recombinant DNA Chapter 10 Copyright The McGraw-Hill Companies, Inc) Permission required for reproduction or display. Applications of Genetic Engineering Basic science vs. Applied
Genetic Background Page 1 PHAGE P22
 Genetic Background Page 1 PHAGE P22 Growth of P22. P22 is a temperate phage that infects Salmonella by binding to the O-antigen, part of the lipopolysaccharide on the outer membrane. After infection, P22
Genetic Background Page 1 PHAGE P22 Growth of P22. P22 is a temperate phage that infects Salmonella by binding to the O-antigen, part of the lipopolysaccharide on the outer membrane. After infection, P22
Biotechnology. Chapter 20. Biology Eighth Edition Neil Campbell and Jane Reece. PowerPoint Lecture Presentations for
 Chapter 20 Biotechnology PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from Joan Sharp Copyright
Chapter 20 Biotechnology PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from Joan Sharp Copyright
BIOTECHNOLOGY OLD BIOTECHNOLOGY (TRADITIONAL BIOTECHNOLOGY) MODERN BIOTECHNOLOGY RECOMBINANT DNA TECHNOLOGY.
 BIOTECHNOLOGY Biotechnology can be defined as the use of micro-organisms, plant or animal cells or their components or enzymes from organisms to produce products and processes (services) useful to human
BIOTECHNOLOGY Biotechnology can be defined as the use of micro-organisms, plant or animal cells or their components or enzymes from organisms to produce products and processes (services) useful to human
BS 50 Genetics and Genomics Week of Nov 29
 BS 50 Genetics and Genomics Week of Nov 29 Additional Practice Problems for Section Problem 1. A linear piece of DNA is digested with restriction enzymes EcoRI and HinDIII, and the products are separated
BS 50 Genetics and Genomics Week of Nov 29 Additional Practice Problems for Section Problem 1. A linear piece of DNA is digested with restriction enzymes EcoRI and HinDIII, and the products are separated
Recombinants and Transformation
 Jesse Ruben Partner Roman Verner BMB 442 Recombinants and Transformation Introduction The goal of this experiment was to take two antibiotic resistance genes for ampicillin and kanamycin from plasmids
Jesse Ruben Partner Roman Verner BMB 442 Recombinants and Transformation Introduction The goal of this experiment was to take two antibiotic resistance genes for ampicillin and kanamycin from plasmids
Adsorption of Lipid-containing Bacteriophages PR4 and PRDl to Pili Determined by a P-1 Incompatibility Group Plasmid
 Journal of General Microbiology (I 977), 98,6 19423 Printed in Great Britain 619 Adsorption of Lipid-containing Bacteriophages PR4 and PRDl to Pili Determined by a P-1 Incompatibility Group Plasmid By
Journal of General Microbiology (I 977), 98,6 19423 Printed in Great Britain 619 Adsorption of Lipid-containing Bacteriophages PR4 and PRDl to Pili Determined by a P-1 Incompatibility Group Plasmid By
About Transformation
 About Transformation In 1928, Frederick Griffith was working on this problem of finding a vaccine against pneumonia caused by the bacteria Streptococcus pneumoniae. Here s what he found: In Experiment
About Transformation In 1928, Frederick Griffith was working on this problem of finding a vaccine against pneumonia caused by the bacteria Streptococcus pneumoniae. Here s what he found: In Experiment
Spostiamo ora la nostra attenzione sui batteri, e batteriofagi
 Spostiamo ora la nostra attenzione sui batteri, e batteriofagi Bacteria Mutate Spontaneously and Grow at an Exponential Rate. Useful for genetics studies, development of genetic engineering Teoria dell'adattamento
Spostiamo ora la nostra attenzione sui batteri, e batteriofagi Bacteria Mutate Spontaneously and Grow at an Exponential Rate. Useful for genetics studies, development of genetic engineering Teoria dell'adattamento
2054, Chap. 14, page 1
 2054, Chap. 14, page 1 I. Recombinant DNA technology (Chapter 14) A. recombinant DNA technology = collection of methods used to perform genetic engineering 1. genetic engineering = deliberate modification
2054, Chap. 14, page 1 I. Recombinant DNA technology (Chapter 14) A. recombinant DNA technology = collection of methods used to perform genetic engineering 1. genetic engineering = deliberate modification
Supplementary Information for
 Supplementary Information for Direct, rapid antimicrobial susceptibility test from positive blood cultures based on microscopic imaging analysis J. Choi 1, H. Y. Jeong 2,3, G. Y. Lee 2,4, S. Han 1, S.
Supplementary Information for Direct, rapid antimicrobial susceptibility test from positive blood cultures based on microscopic imaging analysis J. Choi 1, H. Y. Jeong 2,3, G. Y. Lee 2,4, S. Han 1, S.
Chapter 9. Biotechnology and DNA Technology
 Chapter 9 Biotechnology and DNA Technology SLOs Compare and contrast biotechnology, recombinant DNA technology, and genetic engineering. Identify the roles of a clone and a vector in making recombined
Chapter 9 Biotechnology and DNA Technology SLOs Compare and contrast biotechnology, recombinant DNA technology, and genetic engineering. Identify the roles of a clone and a vector in making recombined
Supplementary appendix
 Supplementary appendix This appendix formed part of the original submission and has been peer reviewed. We post it as supplied by the authors. Supplement to: Quan J, Li X, Chen Y, et al. Prevalence of
Supplementary appendix This appendix formed part of the original submission and has been peer reviewed. We post it as supplied by the authors. Supplement to: Quan J, Li X, Chen Y, et al. Prevalence of
PRODUCT INFORMATION. Composition of SOC medium supplied :
 Product Name : Competent Cell BL21(DE3)pLysS Code No. : DS260 Size : 100 μl 10 Competency : > 5 10 7 cfu/μg (puc19) Supplied product : SOC medium, 1 ml 10 This product is for research use only Description
Product Name : Competent Cell BL21(DE3)pLysS Code No. : DS260 Size : 100 μl 10 Competency : > 5 10 7 cfu/μg (puc19) Supplied product : SOC medium, 1 ml 10 This product is for research use only Description
Chapter 9 Antimicrobial Susceptibility Testing (Agar Disk Diffusion Method)
 Chapter 9 Antimicrobial Susceptibility Testing (Agar Disk Diffusion Method) The disk diffusion method presented in this chapter has been carefully standardized by the National Committee for Clinical Laboratory
Chapter 9 Antimicrobial Susceptibility Testing (Agar Disk Diffusion Method) The disk diffusion method presented in this chapter has been carefully standardized by the National Committee for Clinical Laboratory
Chapter 9 Genetic Engineering
 Chapter 9 Genetic Engineering Biotechnology: use of microbes to make a protein product Recombinant DNA Technology: Insertion or modification of genes to produce desired proteins Genetic engineering: manipulation
Chapter 9 Genetic Engineering Biotechnology: use of microbes to make a protein product Recombinant DNA Technology: Insertion or modification of genes to produce desired proteins Genetic engineering: manipulation
MOLECULAR GENETICS: TRANSFORMATION AND CLONING adapted by Dr. D. L. Vogelien
 Introduction MOLECULAR GENETICS: TRANSFORMATION AND CLONING adapted by Dr. D. L. Vogelien The field of molecular genetics has resulted in a number of practical applications that have been of tremendous
Introduction MOLECULAR GENETICS: TRANSFORMATION AND CLONING adapted by Dr. D. L. Vogelien The field of molecular genetics has resulted in a number of practical applications that have been of tremendous
Supplementary Material
 Supplementary Material Gene Inactivation Study on gntk, a Putative C-methyltransferase Gene in Gentamicin Biosynthesis from Micromonospora echinospora Suman Karki Jin-Yong Kim Si-Hyung Park Hyung-Jin Kwon
Supplementary Material Gene Inactivation Study on gntk, a Putative C-methyltransferase Gene in Gentamicin Biosynthesis from Micromonospora echinospora Suman Karki Jin-Yong Kim Si-Hyung Park Hyung-Jin Kwon
Evaluation of a Rapid Bauer-Kirby Antibiotic Susceptibility
 ANTIMICROBIAL AGENTS AND CHEMoTHERAPY, Mar. 1975. p. 250-255 Copyright 0 1975 American Society for Microbiology Vol. 7, No. 3 Printed in USA. Evaluation of a Rapid Bauer-Kirby Antibiotic Susceptibility
ANTIMICROBIAL AGENTS AND CHEMoTHERAPY, Mar. 1975. p. 250-255 Copyright 0 1975 American Society for Microbiology Vol. 7, No. 3 Printed in USA. Evaluation of a Rapid Bauer-Kirby Antibiotic Susceptibility
The plasmid shown to the right has an oriv and orit at the positions indicated, and is known to replicate bidirectionally.
 Name Microbial Genetics, BIO 410/510 2008 Exam II The plasmid shown to the right has an oriv and orit at the positions indicated, and is known to replicate bidirectionally. 1.) Indicate where replication
Name Microbial Genetics, BIO 410/510 2008 Exam II The plasmid shown to the right has an oriv and orit at the positions indicated, and is known to replicate bidirectionally. 1.) Indicate where replication
Comparative in-vitro activity of cefaclor against urinary tract isolates of Escherichia coli
 Journal of Antimicrobial Chemotherapy (1996) 38, 59-66 Comparative in-vitro activity of cefaclor against urinary tract isolates of Escherichia coli C. A. Webster,. Curran and K. J. Towner* Department of
Journal of Antimicrobial Chemotherapy (1996) 38, 59-66 Comparative in-vitro activity of cefaclor against urinary tract isolates of Escherichia coli C. A. Webster,. Curran and K. J. Towner* Department of
Prevention of Transmission of the Superbug Carbapenem-Resistant Enterobacteriaceae (CRE) during Gastrointestinal Endoscopy
 Prevention of Transmission of the Superbug Carbapenem-Resistant Enterobacteriaceae (CRE) during Gastrointestinal Endoscopy Sponsored by: Presentation by: Lawrence F. Muscarella, Ph.D. President LFM Healthcare
Prevention of Transmission of the Superbug Carbapenem-Resistant Enterobacteriaceae (CRE) during Gastrointestinal Endoscopy Sponsored by: Presentation by: Lawrence F. Muscarella, Ph.D. President LFM Healthcare
Lectures of Dr.Mohammad Alfaham. The Bacterial Genetics
 Lectures of Dr.Mohammad Alfaham The Bacterial Genetics is the total collection of genes carried by a bacterium both on its chromosome and on its extrachromosomal genetic elements (plasmids) A Gene A gene
Lectures of Dr.Mohammad Alfaham The Bacterial Genetics is the total collection of genes carried by a bacterium both on its chromosome and on its extrachromosomal genetic elements (plasmids) A Gene A gene
Genetics and Genomics in Medicine Chapter 3. Questions & Answers
 Genetics and Genomics in Medicine Chapter 3 Multiple Choice Questions Questions & Answers Question 3.1 Which of the following statements, if any, is false? a) Amplifying DNA means making many identical
Genetics and Genomics in Medicine Chapter 3 Multiple Choice Questions Questions & Answers Question 3.1 Which of the following statements, if any, is false? a) Amplifying DNA means making many identical
Taxonomy. Classification of microorganisms 3/12/2017. Is the study of classification. Chapter 10 BIO 220
 Taxonomy Is the study of classification Organisms are classified based on relatedness to each other Chapter 10 BIO 220 Fig. 10.1 1 Species Binomial nomenclature for species identification A eukaryotic
Taxonomy Is the study of classification Organisms are classified based on relatedness to each other Chapter 10 BIO 220 Fig. 10.1 1 Species Binomial nomenclature for species identification A eukaryotic
Section A: Prokaryotes Types and Structure 1. What is microbiology?
 Section A: Prokaryotes Types and Structure 1. What is microbiology? 2. Compare and contrast characteristics of each bacterial type: Eubacteria and Archaebacteria. Eubacteria Both Archaebacteria 3. Label
Section A: Prokaryotes Types and Structure 1. What is microbiology? 2. Compare and contrast characteristics of each bacterial type: Eubacteria and Archaebacteria. Eubacteria Both Archaebacteria 3. Label
Plasmid-Mediated Quinolone Resistance in Clinical Isolates of Escherichia coli from Shanghai, China
 ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, July 2003, p. 2242 2248 Vol. 47, No. 7 0066-4804/03/$08.00 0 DOI: 10.1128/AAC.47.7.2242 2248.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved.
ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, July 2003, p. 2242 2248 Vol. 47, No. 7 0066-4804/03/$08.00 0 DOI: 10.1128/AAC.47.7.2242 2248.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved.
Molecular Cell Biology - Problem Drill 11: Recombinant DNA
 Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
3-Lactamase in Enterobacteria
 ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Feb. 1984, p. 268-272 0066-4804/84/020268-05$02.00/0 Copyright 1984, American Society for Microbiology Vol. 25, No. 2 Detection of PSE-2 3-Lactamase in Enterobacteria
ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Feb. 1984, p. 268-272 0066-4804/84/020268-05$02.00/0 Copyright 1984, American Society for Microbiology Vol. 25, No. 2 Detection of PSE-2 3-Lactamase in Enterobacteria
Regulation of enzyme synthesis
 Regulation of enzyme synthesis The lac operon is an example of an inducible operon - it is normally off, but when a molecule called an inducer is present, the operon turns on. The trp operon is an example
Regulation of enzyme synthesis The lac operon is an example of an inducible operon - it is normally off, but when a molecule called an inducer is present, the operon turns on. The trp operon is an example
NCERT. 2. An enzyme catalysing the removal of nucleotides from the ends of DNA is: a. endonuclease b. exonuclease c. DNA ligase d.
 BIOTECHNOLOGY PRINCIPLES AND PROCESSES 75 CHAPTER 11 BIOTECHNOLOGY: PRINCIPLES AND PROCESSES 1. Rising of dough is due to: MULTIPLE-CHOICE QUESTIONS a. Multiplication of yeast b. Production of CO 2 c.
BIOTECHNOLOGY PRINCIPLES AND PROCESSES 75 CHAPTER 11 BIOTECHNOLOGY: PRINCIPLES AND PROCESSES 1. Rising of dough is due to: MULTIPLE-CHOICE QUESTIONS a. Multiplication of yeast b. Production of CO 2 c.
Quantitation and Purification of Acquired Plasmid DNA Coding for Antimicrobial Drug Resistance
 Cantaurus, Vol. 7, 23-27, May 1999 McPherson College Division of Science and Technology Quantitation and Purification of Acquired Plasmid DNA Coding for Antimicrobial Drug Resistance J. Roy Johnson Jr.
Cantaurus, Vol. 7, 23-27, May 1999 McPherson College Division of Science and Technology Quantitation and Purification of Acquired Plasmid DNA Coding for Antimicrobial Drug Resistance J. Roy Johnson Jr.
GeNei TM Transformation Teaching Kit Manual
 Teaching Kit Manual Cat No. New Cat No. KT07 107385 KT07A 106220 Revision No.: 00060505 CONTENTS Page No. Objective 3 Principle 3 Kit Description 6 Materials Provided 7 Procedure 9 Observation & Interpretation
Teaching Kit Manual Cat No. New Cat No. KT07 107385 KT07A 106220 Revision No.: 00060505 CONTENTS Page No. Objective 3 Principle 3 Kit Description 6 Materials Provided 7 Procedure 9 Observation & Interpretation
Guidelines for Laboratory Verification of Performance of the FilmArray Blood Culture Identification (BCID) Panel
 Guidelines for Laboratory Verification of Performance of the FilmArray Blood Culture Identification (BCID) Panel Purpose The Clinical Laboratory Improvement Amendments (CLIA), passed in 1988, establishes
Guidelines for Laboratory Verification of Performance of the FilmArray Blood Culture Identification (BCID) Panel Purpose The Clinical Laboratory Improvement Amendments (CLIA), passed in 1988, establishes
Ezy MIC Strip FEATURES AND ADVANTAGES
 Imipenem with & without EDTA Ezy MIC Strips (IPM+EDTA/IPM) (Imipenem + EDTA: 1-64 mcg/ml) (Imipenem : 4-256 mcg/ml) Antimicrobial Susceptibility Testing For In Vitro Diagnostic use EM078 Not for Medicinal
Imipenem with & without EDTA Ezy MIC Strips (IPM+EDTA/IPM) (Imipenem + EDTA: 1-64 mcg/ml) (Imipenem : 4-256 mcg/ml) Antimicrobial Susceptibility Testing For In Vitro Diagnostic use EM078 Not for Medicinal
Abstract. Mary Jane Ferraro, PhD, MPH Jana M. Swenson, MMSc
 January 2009 Vol. 29 No. 2 Replaces M07-A7 Vol. 26 No. 2 Methods for Dilution Antimicrobial Susceptibility Tests for Bacteria That Grow Aerobically; Approved Standard Eighth Edition This document addresses
January 2009 Vol. 29 No. 2 Replaces M07-A7 Vol. 26 No. 2 Methods for Dilution Antimicrobial Susceptibility Tests for Bacteria That Grow Aerobically; Approved Standard Eighth Edition This document addresses
Ongoing epidemic of bla VIM-1 -positive Klebsiella pneumoniae in Athens, Greece: a prospective survey
 Journal of Antimicrobial Chemotherapy (2008) 61, 59 63 doi:10.1093/jac/dkm443 Advance Access publication 13 November 2007 Ongoing epidemic of bla VIM-1 -positive Klebsiella pneumoniae in Athens, Greece:
Journal of Antimicrobial Chemotherapy (2008) 61, 59 63 doi:10.1093/jac/dkm443 Advance Access publication 13 November 2007 Ongoing epidemic of bla VIM-1 -positive Klebsiella pneumoniae in Athens, Greece:
EU Reference Laboratory for E. coli Department of Veterinary Public Health and Food Safety Unit of Foodborne Zoonoses Istituto Superiore di Sanità
 Detection of Enterotoxigenic Escherichia coli in food by Real Time PCR amplification of the lt, sth, and stp genes, encoding the heat-labile and heat-stable enterotoxins 1. Aim and field of application
Detection of Enterotoxigenic Escherichia coli in food by Real Time PCR amplification of the lt, sth, and stp genes, encoding the heat-labile and heat-stable enterotoxins 1. Aim and field of application
Chapter 11. Restriction mapping. Objectives
 Restriction mapping Restriction endonucleases (REs) are part of bacterial defense systems. REs recognize and cleave specific sites in DNA molecules. REs are an indispensable tool in molecular biology for
Restriction mapping Restriction endonucleases (REs) are part of bacterial defense systems. REs recognize and cleave specific sites in DNA molecules. REs are an indispensable tool in molecular biology for
Learning Objectives :
 Learning Objectives : Understand the basic differences between genomic and cdna libraries Understand how genomic libraries are constructed Understand the purpose for having overlapping DNA fragments in
Learning Objectives : Understand the basic differences between genomic and cdna libraries Understand how genomic libraries are constructed Understand the purpose for having overlapping DNA fragments in
Site-directed mutagenesis of proteins
 IFM/Kemi Linköpings Universitet August 2013/LGM Labmanual Site-directed mutagenesis of proteins Figur 1: Flow-chart of the site-directed mutagenesis lab exercise 2 Site-specific mutagenesis Introduction
IFM/Kemi Linköpings Universitet August 2013/LGM Labmanual Site-directed mutagenesis of proteins Figur 1: Flow-chart of the site-directed mutagenesis lab exercise 2 Site-specific mutagenesis Introduction
HiPer Plasmid DNA Cloning Teaching Kit
 HiPer Plasmid DNA Cloning Teaching Kit Product Code: HTBM022 Number of experiments that can be performed: 5 Duration of Experiment: 4 days Day 1- Preparation of media and revival of E. coli Host Day2-
HiPer Plasmid DNA Cloning Teaching Kit Product Code: HTBM022 Number of experiments that can be performed: 5 Duration of Experiment: 4 days Day 1- Preparation of media and revival of E. coli Host Day2-
EM021. Co-Trimoxazole Ezy MIC TM Strip (COT)( mcg/ml) (Trimethoprim/ Sulphamethoxazole) Antimicrobial Susceptibility Testing
 Co-Trimoxazole Ezy MIC TM Strip (COT)(0.002-32 mcg/ml) (Trimethoprim/ Sulphamethoxazole) Antimicrobial Susceptibility Testing For In Vitro Diagnostic use EM021 Not for Medicinal Use It is a unique MIC
Co-Trimoxazole Ezy MIC TM Strip (COT)(0.002-32 mcg/ml) (Trimethoprim/ Sulphamethoxazole) Antimicrobial Susceptibility Testing For In Vitro Diagnostic use EM021 Not for Medicinal Use It is a unique MIC
Antibiotic Resistance in Providencia stuartii Isolated In
 JOURNAL OF CLINICAL MICROBIOLOGY, June 1981, p. 1099-1104 0095-1137/81/061099-06$02.00/0 Vol. 13, No. 6 Antibiotic Resistance in Providencia stuartii Isolated In Hospitals PATRICK J. McHALE,' CONOR T.
JOURNAL OF CLINICAL MICROBIOLOGY, June 1981, p. 1099-1104 0095-1137/81/061099-06$02.00/0 Vol. 13, No. 6 Antibiotic Resistance in Providencia stuartii Isolated In Hospitals PATRICK J. McHALE,' CONOR T.
Shionogi Presents Results of the First Clinical Efficacy Trial and In Vitro Data on Cefiderocol (S ), a Siderophore Cephalosporin
 Shionogi Presents Results of the First Clinical Efficacy Trial and In Vitro Data on Cefiderocol (S-649266), a Siderophore Cephalosporin Osaka, Japan, April 22, 2017 - Shionogi & Co., Ltd. today announced
Shionogi Presents Results of the First Clinical Efficacy Trial and In Vitro Data on Cefiderocol (S-649266), a Siderophore Cephalosporin Osaka, Japan, April 22, 2017 - Shionogi & Co., Ltd. today announced
GenBuilder TM Plus Cloning Kit User Manual
 GenBuilder TM Plus Cloning Kit User Manual Cat.no L00744 Version 11242017 Ⅰ. Introduction... 2 I.1 Product Information... 2 I.2 Kit Contents and Storage... 2 I.3 GenBuilder Cloning Kit Workflow... 2 Ⅱ.
GenBuilder TM Plus Cloning Kit User Manual Cat.no L00744 Version 11242017 Ⅰ. Introduction... 2 I.1 Product Information... 2 I.2 Kit Contents and Storage... 2 I.3 GenBuilder Cloning Kit Workflow... 2 Ⅱ.
Cloning of a Gene Responsible for the Biosynthesis of Glutathione in Escherichia coli B
 APPLIED AND ENVIRONMENTAL MICROBIOLOGY, Dec. 1982, P. 1444-1448 Vol. 44, No. 6 0099-2240/82/121444-05$02.00/0 Copyright C 1982, American Society for Microbiology Cloning of a Gene Responsible for the Biosynthesis
APPLIED AND ENVIRONMENTAL MICROBIOLOGY, Dec. 1982, P. 1444-1448 Vol. 44, No. 6 0099-2240/82/121444-05$02.00/0 Copyright C 1982, American Society for Microbiology Cloning of a Gene Responsible for the Biosynthesis
Contents... vii. List of Figures... xii. List of Tables... xiv. Abbreviatons... xv. Summary... xvii. 1. Introduction In vitro evolution...
 vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
AP Biology. Chapter 20. Biotechnology: DNA Technology & Genomics. Biotechnology. The BIG Questions. Evolution & breeding of food plants
 What do you notice about these phrases? radar racecar Madam I m Adam Able was I ere I saw Elba a man, a plan, a canal, Panama Was it a bar or a bat I saw? Chapter 20. Biotechnology: DNA Technology & enomics
What do you notice about these phrases? radar racecar Madam I m Adam Able was I ere I saw Elba a man, a plan, a canal, Panama Was it a bar or a bat I saw? Chapter 20. Biotechnology: DNA Technology & enomics
Cloning and Characterization of E. meningoseptica Beta Lactamase
 Cloning and Characterization of E. meningoseptica Beta Lactamase Authors: Lindsey Purcell, Jessica Matts, Patricia Canaan* Department of Biochemistry and Molecular Biology Abstract Elizabethkingia meningoseptica
Cloning and Characterization of E. meningoseptica Beta Lactamase Authors: Lindsey Purcell, Jessica Matts, Patricia Canaan* Department of Biochemistry and Molecular Biology Abstract Elizabethkingia meningoseptica
1. Immunization. What is Immunization? 12/9/2016. Chapter 17: Immunization & Immune Testing. 1. Immunization 2. Diagnostic Immunology
 Chapter 17: Immunization & Immune Testing 1. Immunization 2. Diagnostic Immunology 1. Immunization Chapter Reading pp. 505-511 What is Immunization? A method of inducing artificial immunity by exposing
Chapter 17: Immunization & Immune Testing 1. Immunization 2. Diagnostic Immunology 1. Immunization Chapter Reading pp. 505-511 What is Immunization? A method of inducing artificial immunity by exposing
Chapter 17: Immunization & Immune Testing. 1. Immunization 2. Diagnostic Immunology
 Chapter 17: Immunization & Immune Testing 1. Immunization 2. Diagnostic Immunology 1. Immunization Chapter Reading pp. 505-511 What is Immunization? A method of inducing artificial immunity by exposing
Chapter 17: Immunization & Immune Testing 1. Immunization 2. Diagnostic Immunology 1. Immunization Chapter Reading pp. 505-511 What is Immunization? A method of inducing artificial immunity by exposing
CHAPTER 08: RECOMBINANT DNA TECHNOLOGY Pearson Education, Inc.
 CHAPTER 08: RECOMBINANT DNA TECHNOLOGY The Role of Recombinant DNA Technology in Biotechnology Biotechnology the use of microorganisms to make practical products Recombinant DNA technology Intentionally
CHAPTER 08: RECOMBINANT DNA TECHNOLOGY The Role of Recombinant DNA Technology in Biotechnology Biotechnology the use of microorganisms to make practical products Recombinant DNA technology Intentionally
HiPer Transformation Teaching Kit
 HiPer Transformation Teaching Kit Product Code: HTBM017 Number of experiments that can be performed: 10 Duration of Experiment: 4 days Day 1- Preparation of media and revival of E. coli Host Day 2- Inoculation
HiPer Transformation Teaching Kit Product Code: HTBM017 Number of experiments that can be performed: 10 Duration of Experiment: 4 days Day 1- Preparation of media and revival of E. coli Host Day 2- Inoculation
ABC. Methods for Determining Bactericidal Activity of Antimicrobial Agents; Approved Guideline. Volume 19 Number 18
 M26-A ISBN 1-56238-384-1 September 1999 ISSN 0273-3099 Methods for Determining Bactericidal Activity of Antimicrobial Agents; Approved Guideline Volume 19 Number 18 Arthur L. Barry, Ph.D. William A. Craig,
M26-A ISBN 1-56238-384-1 September 1999 ISSN 0273-3099 Methods for Determining Bactericidal Activity of Antimicrobial Agents; Approved Guideline Volume 19 Number 18 Arthur L. Barry, Ph.D. William A. Craig,
1a. What is the ratio of feathered to unfeathered shanks in the offspring of the above cross?
 Problem Set 5 answers 1. Whether or not the shanks of chickens contains feathers is due to two independently assorting genes. Individuals have unfeathered shanks when they are homozygous for recessive
Problem Set 5 answers 1. Whether or not the shanks of chickens contains feathers is due to two independently assorting genes. Individuals have unfeathered shanks when they are homozygous for recessive
ANTIMICROBIAL SUSCEPTIBILITY TESTING: ADVANCED
 ANTIMICROBIAL SUSCEPTIBILITY TESTING: ADVANCED Romney Humphries, Ph.D., DABMM Section Chief, Clinical Microbiology University of California Los Angeles Lost Angeles, CA Susan E. Sharp, Ph.D., DABMM, FAAM
ANTIMICROBIAL SUSCEPTIBILITY TESTING: ADVANCED Romney Humphries, Ph.D., DABMM Section Chief, Clinical Microbiology University of California Los Angeles Lost Angeles, CA Susan E. Sharp, Ph.D., DABMM, FAAM
Maurine A. Leverstein-van Hall,* Hetty E. M. Blok, Armand Paauw, Ad C. Fluit, Annet Troelstra, Ellen M. Mascini, Marc J. M. Bonten, and Jan Verhoef
 JOURNAL OF CLINICAL MICROBIOLOGY, Feb. 2006, p. 518 524 Vol. 44, No. 2 0095-1137/06/$08.00 0 doi:10.1128/jcm.44.2.518 524.2006 Copyright 2006, American Society for Microbiology. All Rights Reserved. Extensive
JOURNAL OF CLINICAL MICROBIOLOGY, Feb. 2006, p. 518 524 Vol. 44, No. 2 0095-1137/06/$08.00 0 doi:10.1128/jcm.44.2.518 524.2006 Copyright 2006, American Society for Microbiology. All Rights Reserved. Extensive
Chapter 8 Recombinant DNA Technology. 10/1/ MDufilho
 Chapter 8 Recombinant DNA Technology 10/1/2017 1 MDufilho The Role of Recombinant DNA Technology in Biotechnology Biotechnology? Recombinant deoxyribonucleic acid (DNA) technology Intentionally modifying
Chapter 8 Recombinant DNA Technology 10/1/2017 1 MDufilho The Role of Recombinant DNA Technology in Biotechnology Biotechnology? Recombinant deoxyribonucleic acid (DNA) technology Intentionally modifying
ONTARIO SCIENCE CENTRE. Teacher Guide. Way to Glow Program
 ONTARIO SCIENCE CENTRE Teacher Guide Way to Glow Program Table of Contents Bacterial transformation background information 3 Experimental procedure 5 Expected results 7 Post-program activity sheet 8 Post-program
ONTARIO SCIENCE CENTRE Teacher Guide Way to Glow Program Table of Contents Bacterial transformation background information 3 Experimental procedure 5 Expected results 7 Post-program activity sheet 8 Post-program
_ DNA absorbs light at 260 wave length and it s a UV range so we cant see DNA, we can see DNA only by staining it.
 * GEL ELECTROPHORESIS : its a technique aim to separate DNA in agel based on size, in this technique we add a sample of DNA in a wells in the gel, then we turn on the electricity, the DNA will travel in
* GEL ELECTROPHORESIS : its a technique aim to separate DNA in agel based on size, in this technique we add a sample of DNA in a wells in the gel, then we turn on the electricity, the DNA will travel in
Validation of the Automated Reading and Incubation System with Sensititre Plates for Antimicrobial Susceptibility Testing
 JOURNAL OF CLINICAL MICROBIOLOGY, May 2003, p. 1951 1956 Vol. 41, No. 5 0095-1137/03/$08.00 0 DOI: 10.1128/JCM.41.5.1951 1956.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved.
JOURNAL OF CLINICAL MICROBIOLOGY, May 2003, p. 1951 1956 Vol. 41, No. 5 0095-1137/03/$08.00 0 DOI: 10.1128/JCM.41.5.1951 1956.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved.
Guide-it Indel Identification Kit User Manual
 Clontech Laboratories, Inc. Guide-it Indel Identification Kit User Manual Cat. No. 631444 (120114) Clontech Laboratories, Inc. A Takara Bio Company 1290 Terra Bella Avenue, Mountain View, CA 94043, USA
Clontech Laboratories, Inc. Guide-it Indel Identification Kit User Manual Cat. No. 631444 (120114) Clontech Laboratories, Inc. A Takara Bio Company 1290 Terra Bella Avenue, Mountain View, CA 94043, USA
Aminoglycoside-Modifying Enzymes That Retain Activity
 ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, May 1980, p. 798-802 0066-4804/80/05-0798/05$02.00/0 Vol. 17, No. 5 5-epi-Sisomicin and 5-epi-Gentamicin B: Substrates for Aminoglycoside-Modifying Enzymes That Retain
ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, May 1980, p. 798-802 0066-4804/80/05-0798/05$02.00/0 Vol. 17, No. 5 5-epi-Sisomicin and 5-epi-Gentamicin B: Substrates for Aminoglycoside-Modifying Enzymes That Retain
The demonstration that wild-type T-DNA coding region can be replaced by any DNA sequence without any effect on its transfer from A.
 The demonstration that wild-type T-DNA coding region can be replaced by any DNA sequence without any effect on its transfer from A. tumefaciens to the plant inspired the promise that A. tumefaciens might
The demonstration that wild-type T-DNA coding region can be replaced by any DNA sequence without any effect on its transfer from A. tumefaciens to the plant inspired the promise that A. tumefaciens might
1. What is the structure and function of DNA? Describe in words or a drawing the structure of a DNA molecule. Be as detailed as possible.
 INTRODUCTION In the Program Introduction, you learned that the increase in diabetes in the United States has resulted in a great demand for its treatment, insulin. You also learned that the best way to
INTRODUCTION In the Program Introduction, you learned that the increase in diabetes in the United States has resulted in a great demand for its treatment, insulin. You also learned that the best way to
Plasmids-Mediated Antibiotic Resistance in E. coli
 Plasmids-Mediated Antibiotic Resistance in E. coli (Received: 08.12.1998) Nariman A. H. Aly*; Amany Tharwat** and Wesam F. El-Baz** *Microbial Genetics Dept., Genetic Engineering and Biotechnology Div.,
Plasmids-Mediated Antibiotic Resistance in E. coli (Received: 08.12.1998) Nariman A. H. Aly*; Amany Tharwat** and Wesam F. El-Baz** *Microbial Genetics Dept., Genetic Engineering and Biotechnology Div.,
Department of the Environment, Food and Rural Affairs
 Department of the Environment, Food and Rural Affairs Advisory Committee on Releases to the Environment Format 2: Release of genetically modified organisms other than higher plants PART C: SUMMARY NOTIFICATION
Department of the Environment, Food and Rural Affairs Advisory Committee on Releases to the Environment Format 2: Release of genetically modified organisms other than higher plants PART C: SUMMARY NOTIFICATION
CHAPTER 20 DNA TECHNOLOGY AND GENOMICS. Section A: DNA Cloning
 Section A: DNA Cloning 1. DNA technology makes it possible to clone genes for basic research and commercial applications: an overview 2. Restriction enzymes are used to make recombinant DNA 3. Genes can
Section A: DNA Cloning 1. DNA technology makes it possible to clone genes for basic research and commercial applications: an overview 2. Restriction enzymes are used to make recombinant DNA 3. Genes can
Effect of Storage of Mueller-Hinton Agar Plates on
 APPUED MICROBIOLOGY, Sept. 1970, p. 293-297 Copyright 1970 American Society for Microbiology Vol. 20, No. 3 Printed in U.S.A. Effect of Storage of Mueller-Hinton Agar Plates on Zone Sizes for Antimicrobial
APPUED MICROBIOLOGY, Sept. 1970, p. 293-297 Copyright 1970 American Society for Microbiology Vol. 20, No. 3 Printed in U.S.A. Effect of Storage of Mueller-Hinton Agar Plates on Zone Sizes for Antimicrobial
Polymerase Chain Reaction
 Polymerase Chain Reaction Amplify your insert or verify its presence 3H Taq platinum PCR mix primers Ultrapure Water PCR tubes PCR machine A. Insert amplification For insert amplification, use the Taq
Polymerase Chain Reaction Amplify your insert or verify its presence 3H Taq platinum PCR mix primers Ultrapure Water PCR tubes PCR machine A. Insert amplification For insert amplification, use the Taq
AP Laboratory: Microbes in Action Bacterial Transformation & Gel Electrophoresis
 AP Laboratory: Microbes in Action Name: Bacterial Transformation & Gel Electrophoresis Introduction In this laboratory you will use some basic tools of molecular biology to gain an understanding of some
AP Laboratory: Microbes in Action Name: Bacterial Transformation & Gel Electrophoresis Introduction In this laboratory you will use some basic tools of molecular biology to gain an understanding of some
BIOTECHNOLOGY : PRINCIPLES AND PROCESSES Biotechnologydeals with techniques of using live organisms or enzymes from organisms to produce products and
 BIOTECHNOLOGY : PRINCIPLES AND PROCESSES Biotechnologydeals with techniques of using live organisms or enzymes from organisms to produce products and processes useful to humans. Traditional form based
BIOTECHNOLOGY : PRINCIPLES AND PROCESSES Biotechnologydeals with techniques of using live organisms or enzymes from organisms to produce products and processes useful to humans. Traditional form based
4/26/2015. Cut DNA either: Cut DNA either:
 Ch.20 Enzymes that cut DNA at specific sequences (restriction sites) resulting in segments of DNA (restriction fragments) Typically 4-8 bp in length & often palindromic Isolated from bacteria (Hundreds
Ch.20 Enzymes that cut DNA at specific sequences (restriction sites) resulting in segments of DNA (restriction fragments) Typically 4-8 bp in length & often palindromic Isolated from bacteria (Hundreds
F or many years, the isolation of urinary pathogens has
 608 ORIGINAL ARTICLE A comparison of the performance of commercially available chromogenic agars for the isolation and presumptive identification of organisms from urine D Fallon, G Ackland, N Andrews,
608 ORIGINAL ARTICLE A comparison of the performance of commercially available chromogenic agars for the isolation and presumptive identification of organisms from urine D Fallon, G Ackland, N Andrews,
COMMITTEE FOR VETERINARY MEDICINAL PRODUCTS NOTE FOR GUIDANCE 1 : DNA VACCINES NON-AMPLIFIABLE IN EUKARYOTIC CELLS FOR VETERINARY USE
 The European Agency for the Evaluation of Medicinal Products Evaluation of Medicines for Veterinary Use CVMP/IWP/07/98-FINAL COMMITTEE FOR VETERINARY MEDICINAL PRODUCTS NOTE FOR GUIDANCE 1 : DNA VACCINES
The European Agency for the Evaluation of Medicinal Products Evaluation of Medicines for Veterinary Use CVMP/IWP/07/98-FINAL COMMITTEE FOR VETERINARY MEDICINAL PRODUCTS NOTE FOR GUIDANCE 1 : DNA VACCINES
