Genome-Scale Predictions of the Transcription Factor Binding Sites of Cys 2 His 2 Zinc Finger Proteins in Yeast June 17 th, 2005
|
|
|
- Herbert Warren
- 5 years ago
- Views:
Transcription
1 Genome-Scale Predictions of the Transcription Factor Binding Sites of Cys 2 His 2 Zinc Finger Proteins in Yeast June 17 th, 2005 John Brothers II 1,3 and Panayiotis V. Benos 1,2 1 Bioengineering and Bioinformatics Summer Institute, 2 Department of Computational Biology, University of Pittsburgh, Pittsburgh, PA 15213, 3 Bioinformatics, Department of Biological Sciences, Rochester Institute of Technology, Rochester, NY Introduction Cell expression and regulation of genes is still not very well understood except for a few deeply studied processes (Doniger et al., 2005). The challenge to discover a clear deterministic model for gene regulation is considered one of the greater challenges of modern computation biology and bioinformatics (Benos et al., 2002). With the combined experimental and computational data that has been discovered about regulatory motifs in Saccharomyces cerevisiae, it has become possible to pursue not only the identification of functioning transcription factor binding sites (Doniger et al., 2005), but to aim for that goal of discovering a deterministic model for the prediction of TFBSs in entire yeast genomes. By finding all of the possible TFBSs and evaluating the possible DNA interactions of the Cys 2 Hys 2 zinc finger protein domains (the domain most found on transcription factors) to discover which DNA sequences they bind to, we would be able to help expand and assess the accuracy of the model developed by Benos et al for the
2 prediction of binding sites and further improve the overall understanding of the processes behind cell regulation (Benos et al, 2002). Methodology: The DNA binding preferences of yeast Cys 2 His 2 zinc finger proteins, composed of only two or three fingers, will be determined by using position specific scoring matrices (PSSM) (Stormo, 2000). For Saccharomyces cerevisiae, a program entitled C2H2-enoLOGOS will be used to directly ascertain the PSSM models for any of the yeast Cys 2 His 2 zinc finger proteins with only two or three fingers (Workman et al., 2005). To determine the PSSM models for any other species of yeast, members of the Cys 2 Hys 2 zinc finger protein family will be extracted from the Pfam database ( By doing comparisons to the human or mouse EGR1 protein, the crucial amino acids that bind to positions 1, +3, and +6 on the DNA helix will be determined (see Figure 1). Those amino acid identities will be used to determine the position-specific base Figure 1. The DNA-binding of the EGR1 protein. preferences of the protein using the three contact model developed by Benos et all (Benos et al., 2002). Once the PSSM models for all 2- and 3-finger S. cerevisiae proteins have been calculated, the orthologues of these proteins in other yeast species will be
3 determined using Basic Local Alignment Search Tools (BLAST). For this experiment, a protein of another yeast species will be considered an orthologous protein to a S. cerevisiae protein if said protein is the top hit in the BLAST search with the amino acid sequence of the S. cerevisiae protein. Those proteins that are found to have no orthologues in other yeast species will be excluded from further analysis. Since yeast species usually have short 5 UTRs, the 1.5kb upstream of the translation start site should contain all of the important DNA regulatory elements, therefore for this experiment, that section upstream of the yeast gene will be considered the promoter region. All of the promoter regions of all the yeast genes of the multiple yeast genomes will then be identified. Those sequences will be derived from publicly available datasets or from published genomes. The promoters of the orthologous genes will then be searched for possible binding sites of each of the zinc finger proteins using a strategy similar to the FOOTER algorithm, which is where the position of the site and the PSSM score are used to decide whether a possible site is conserved between two species. Since there are multiple species and genomes available, the FOOTER algorithm may be expanded to work with multiple species comparisons. The predictions made using the above techniques will help to discover and assess further the accuracy of the Cys 2 Hys 2 zinc finger model developed by Benos et al (Benos et al., 2002). The final determination of the accuracy of this developing model for the genome-scale predictions of the binding sites of the Cys 2 Hys 2 zinc finger proteins in yeast will be assessed by evaluation of the
4 above predictions with publicly available gene microarray data and chromatin immunoprecipitation (ChIP) data (Lee et al., 2002). Other ways to assess the computed predictions may also be used. Expected Results 1. To improve the understanding of the gene regulation through the mathematical modeling of the transcription-factor binding sites using zinc finger proteins. 2. To expand the FOOTER algorithm in such a way that it can be used to do multiple comparisons between more than two species of yeast. 3. To assess and improve the current model for predicting genome-scale binding sites used by Benos, et al. References Benos, P.V., A.S. Lapedes, and G.D. Stormo Probabilistic code for DNA recognition by proteins of the EGR family. J Mol Biol 323: Doniger, S.W., J. Huh, and J.C. Fay Identification of functional transcription factor binding sites using closely related Saccharomyces species. Genome Res 15: Lee, T.I., N.J. Rinaldi, F. Robert, D.T. Odom, Z. Bar-Joseph, G.K. Gerber, N.M. Hannett, C.T. Harbison, C.M. Thompson, I. Simon, J. Zeitlinger, E.G. Jennings, H.L. Murray, D.B. Gordon, B. Ren, J.J.
5 Wyrick, J.B. Tagne, T.L. Volkert, E. Fraenkel, D.K. Gifford, and R.A. Young Transcriptional regulatory networks in Saccharomyces cerevisiae. Science 298: Stormo, G.D DNA binding sites: representation and discovery. Bioinformatics 16: Workman, C.T., Y. Yin, D.L. Corcoran, T. Ideker, G.D. Stormo, and P.V. Benos EnoLOGOS: a versatile web tool for energy normalized sequence logos. Nucleic Acids Res 000:
Le proteine regolative variano nei vari tipi cellulari e in funzione degli stimoli ambientali
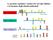 Le proteine regolative variano nei vari tipi cellulari e in funzione degli stimoli ambientali Tipo cellulare 1 Tipo cellulare 2 Tipo cellulare 3 DNA-protein Crosslink Lisi Frammentazione Immunopurificazione
Le proteine regolative variano nei vari tipi cellulari e in funzione degli stimoli ambientali Tipo cellulare 1 Tipo cellulare 2 Tipo cellulare 3 DNA-protein Crosslink Lisi Frammentazione Immunopurificazione
Introduction to Transcription Factor Binding Sites (TFBS) Cells control the expression of genes using Transcription Factors.
 Identification of Functional Transcription Factor Binding Sites using Closely Related Saccharomyces species Scott W. Doniger 1, Juyong Huh 2, and Justin C. Fay 1,2 1 Computation Biology Program and 2 Department
Identification of Functional Transcription Factor Binding Sites using Closely Related Saccharomyces species Scott W. Doniger 1, Juyong Huh 2, and Justin C. Fay 1,2 1 Computation Biology Program and 2 Department
WebMOTIFS: Automated discovery, filtering, and scoring of DNA sequence motifs using multiple programs and Bayesian approaches
 WebMOTIFS: Automated discovery, filtering, and scoring of DNA sequence motifs using multiple programs and Bayesian approaches Katherine A. Romer 1, Guy-Richard Kayombya 1, Ernest Fraenkel 2,3 1 Department
WebMOTIFS: Automated discovery, filtering, and scoring of DNA sequence motifs using multiple programs and Bayesian approaches Katherine A. Romer 1, Guy-Richard Kayombya 1, Ernest Fraenkel 2,3 1 Department
Structural Analysis of the EGR Family of Transcription Factors: Templates for Predicting Protein DNA Interactions
 Introduction Structural Analysis of the EGR Family of Transcription Factors: Templates for Predicting Protein DNA Interactions Jamie Duke, Rochester Institute of Technology Mentor: Carlos Camacho, University
Introduction Structural Analysis of the EGR Family of Transcription Factors: Templates for Predicting Protein DNA Interactions Jamie Duke, Rochester Institute of Technology Mentor: Carlos Camacho, University
ECS 234: Genomic Data Integration ECS 234
 : Genomic Data Integration Heterogeneous Data Integration DNA Sequence Microarray Proteomics >gi 12004594 gb AF217406.1 Saccharomyces cerevisiae uridine nucleosidase (URH1) gene, complete cds ATGGAATCTGCTGATTTTTTTACCTCACGAAACTTATTAAAACAGATAATTTCCCTCATCTGCAAGGTTG
: Genomic Data Integration Heterogeneous Data Integration DNA Sequence Microarray Proteomics >gi 12004594 gb AF217406.1 Saccharomyces cerevisiae uridine nucleosidase (URH1) gene, complete cds ATGGAATCTGCTGATTTTTTTACCTCACGAAACTTATTAAAACAGATAATTTCCCTCATCTGCAAGGTTG
Transcriptional Regulatory Code of a Eukaryotic Genome
 Supplementary Information for Transcriptional Regulatory Code of a Eukaryotic Genome Christopher T. Harbison 1,2*, D. Benjamin Gordon 1*, Tong Ihn Lee 1, Nicola J. Rinaldi 1,2, Kenzie Macisaac 3, Timothy
Supplementary Information for Transcriptional Regulatory Code of a Eukaryotic Genome Christopher T. Harbison 1,2*, D. Benjamin Gordon 1*, Tong Ihn Lee 1, Nicola J. Rinaldi 1,2, Kenzie Macisaac 3, Timothy
A new framework for identifying combinatorial regulation of. transcription factors: a case study of the yeast cell cycle
 A new framework for identifying combinatorial regulation of transcription factors: a case study of the yeast cell cycle Junbai Wang 1 * 1. Department of Biological Sciences, Columbia University, 1212 Amsterdam
A new framework for identifying combinatorial regulation of transcription factors: a case study of the yeast cell cycle Junbai Wang 1 * 1. Department of Biological Sciences, Columbia University, 1212 Amsterdam
Chapter 10. Bioinformatics. Samuel Kaski, Janne Nikkilä, Merja Oja, Leo Lahti, Jarkko Venna, Eerika Savia, Janne Sinkkonen, Jaakko Peltonen
 Chapter 10 Bioinformatics Samuel Kaski, Janne Nikkilä, Merja Oja, Leo Lahti, Jarkko Venna, Eerika Savia, Janne Sinkkonen, Jaakko Peltonen 133 134 Bioinformatics 10.1 Introduction Bioinformatics refers
Chapter 10 Bioinformatics Samuel Kaski, Janne Nikkilä, Merja Oja, Leo Lahti, Jarkko Venna, Eerika Savia, Janne Sinkkonen, Jaakko Peltonen 133 134 Bioinformatics 10.1 Introduction Bioinformatics refers
Characterizing DNA binding sites high throughput approaches Biol4230 Tues, April 24, 2018 Bill Pearson Pinn 6-057
 Characterizing DNA binding sites high throughput approaches Biol4230 Tues, April 24, 2018 Bill Pearson wrp@virginia.edu 4-2818 Pinn 6-057 Reviewing sites: affinity and specificity representation binding
Characterizing DNA binding sites high throughput approaches Biol4230 Tues, April 24, 2018 Bill Pearson wrp@virginia.edu 4-2818 Pinn 6-057 Reviewing sites: affinity and specificity representation binding
Probabilistic Code for DNA Recognition by Proteins of the EGR Family
 doi:10.1016/s0022-2836(02)00917-8 available online at http://www.idealibrary.com on Bw J. Mol. Biol. (2002) 323, 701 727 Probabilistic Code for DNA Recognition by Proteins of the EGR Family Panayiotis
doi:10.1016/s0022-2836(02)00917-8 available online at http://www.idealibrary.com on Bw J. Mol. Biol. (2002) 323, 701 727 Probabilistic Code for DNA Recognition by Proteins of the EGR Family Panayiotis
Sequence Motif Analysis
 Sequence Motif Analysis Lecture in M.Sc. Biomedizin, Module: Proteinbiochemie und Bioinformatik Jonas Ibn-Salem Andrade group Johannes Gutenberg University Mainz Institute of Molecular Biology March 7,
Sequence Motif Analysis Lecture in M.Sc. Biomedizin, Module: Proteinbiochemie und Bioinformatik Jonas Ibn-Salem Andrade group Johannes Gutenberg University Mainz Institute of Molecular Biology March 7,
In 1996, the genome of Saccharomyces cerevisiae was completed due to the work of
 Summary: Kellis, M. et al. Nature 423,241-253. Background In 1996, the genome of Saccharomyces cerevisiae was completed due to the work of approximately 600 scientists world-wide. This group of researchers
Summary: Kellis, M. et al. Nature 423,241-253. Background In 1996, the genome of Saccharomyces cerevisiae was completed due to the work of approximately 600 scientists world-wide. This group of researchers
AS A SERVICE TO THE RESEARCH COMMUNITY, GENOME BIOLOGY PROVIDES A 'PREPRINT' DEPOSITORY
 This information has not been peer-reviewed. Responsibility for the findings rests solely with the author(s). Deposited research article All motifs are not created equal: structural properties of transcription
This information has not been peer-reviewed. Responsibility for the findings rests solely with the author(s). Deposited research article All motifs are not created equal: structural properties of transcription
Representation in Supervised Machine Learning Application to Biological Problems
 Representation in Supervised Machine Learning Application to Biological Problems Frank Lab Howard Hughes Medical Institute & Columbia University 2010 Robert Howard Langlois Hughes Medical Institute What
Representation in Supervised Machine Learning Application to Biological Problems Frank Lab Howard Hughes Medical Institute & Columbia University 2010 Robert Howard Langlois Hughes Medical Institute What
Learning Methods for DNA Binding in Computational Biology
 Learning Methods for DNA Binding in Computational Biology Mark Kon Dustin Holloway Yue Fan Chaitanya Sai Charles DeLisi Boston University IJCNN Orlando August 16, 2007 Outline Background on Transcription
Learning Methods for DNA Binding in Computational Biology Mark Kon Dustin Holloway Yue Fan Chaitanya Sai Charles DeLisi Boston University IJCNN Orlando August 16, 2007 Outline Background on Transcription
Introduction to Bioinformatics
 Introduction to Bioinformatics Changhui (Charles) Yan Old Main 401 F http://www.cs.usu.edu www.cs.usu.edu/~cyan 1 How Old Is The Discipline? "The term bioinformatics is a relatively recent invention, not
Introduction to Bioinformatics Changhui (Charles) Yan Old Main 401 F http://www.cs.usu.edu www.cs.usu.edu/~cyan 1 How Old Is The Discipline? "The term bioinformatics is a relatively recent invention, not
Bio5488 Practice Midterm (2018) 1. Next-gen sequencing
 1. Next-gen sequencing 1. You have found a new strain of yeast that makes fantastic wine. You d like to sequence this strain to ascertain the differences from S. cerevisiae. To accurately call a base pair,
1. Next-gen sequencing 1. You have found a new strain of yeast that makes fantastic wine. You d like to sequence this strain to ascertain the differences from S. cerevisiae. To accurately call a base pair,
Textbook Reading Guidelines
 Understanding Bioinformatics by Marketa Zvelebil and Jeremy Baum Last updated: May 1, 2009 Textbook Reading Guidelines Preface: Read the whole preface, and especially: For the students with Life Science
Understanding Bioinformatics by Marketa Zvelebil and Jeremy Baum Last updated: May 1, 2009 Textbook Reading Guidelines Preface: Read the whole preface, and especially: For the students with Life Science
The spatial distribution of cis regulatory elements in yeast promoters and its implications for transcriptional regulation
 RESEARCH ARTICLE Open Access The spatial distribution of cis regulatory elements in yeast promoters and its implications for transcriptional regulation Zhenguo Lin 1, Wei-Sheng Wu 2, Han Liang 1,4, Yong
RESEARCH ARTICLE Open Access The spatial distribution of cis regulatory elements in yeast promoters and its implications for transcriptional regulation Zhenguo Lin 1, Wei-Sheng Wu 2, Han Liang 1,4, Yong
Biology 644: Bioinformatics
 Processes Activation Repression Initiation Elongation.... Processes Splicing Editing Degradation Translation.... Transcription Translation DNA Regulators DNA-Binding Transcription Factors Chromatin Remodelers....
Processes Activation Repression Initiation Elongation.... Processes Splicing Editing Degradation Translation.... Transcription Translation DNA Regulators DNA-Binding Transcription Factors Chromatin Remodelers....
Identification of individual motifs on the genome scale. Some slides are from Mayukh Bhaowal
 Identification of individual motifs on the genome scale Some slides are from Mayukh Bhaowal Two papers Nature 423, 241-254 (15 May 2003) Sequencing and comparison of yeast species to identify genes and
Identification of individual motifs on the genome scale Some slides are from Mayukh Bhaowal Two papers Nature 423, 241-254 (15 May 2003) Sequencing and comparison of yeast species to identify genes and
Grundlagen der Bioinformatik Summer Lecturer: Prof. Daniel Huson
 Grundlagen der Bioinformatik, SoSe 11, D. Huson, April 11, 2011 1 1 Introduction Grundlagen der Bioinformatik Summer 2011 Lecturer: Prof. Daniel Huson Office hours: Thursdays 17-18h (Sand 14, C310a) 1.1
Grundlagen der Bioinformatik, SoSe 11, D. Huson, April 11, 2011 1 1 Introduction Grundlagen der Bioinformatik Summer 2011 Lecturer: Prof. Daniel Huson Office hours: Thursdays 17-18h (Sand 14, C310a) 1.1
Transcription Gene regulation
 Transcription Gene regulation The machine that transcribes a gene is composed of perhaps 50 proteins, including RNA polymerase, the enzyme that converts DNA code into RNA code. A crew of transcription
Transcription Gene regulation The machine that transcribes a gene is composed of perhaps 50 proteins, including RNA polymerase, the enzyme that converts DNA code into RNA code. A crew of transcription
Massimo Vergassola CNRS, Institut Pasteur
 Massimo Vergassola CNRS, Institut Pasteur Dr. Massimo Vergassola, Institute Pasteur, Paris (KITP Bio Seminar 11/23/04) 1 ! "# $!% Bcd,nos,tor, dl,cad, Hb,Kr,tll,... Eve,ftz,h,... En,wg,dsh,... Dr. Massimo
Massimo Vergassola CNRS, Institut Pasteur Dr. Massimo Vergassola, Institute Pasteur, Paris (KITP Bio Seminar 11/23/04) 1 ! "# $!% Bcd,nos,tor, dl,cad, Hb,Kr,tll,... Eve,ftz,h,... En,wg,dsh,... Dr. Massimo
NGS Approaches to Epigenomics
 I519 Introduction to Bioinformatics, 2013 NGS Approaches to Epigenomics Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Background: chromatin structure & DNA methylation Epigenomic
I519 Introduction to Bioinformatics, 2013 NGS Approaches to Epigenomics Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Background: chromatin structure & DNA methylation Epigenomic
Introduction to Bioinformatics
 Introduction to Bioinformatics Dortmund, 16.-20.07.2007 Lectures: Sven Rahmann Exercises: Udo Feldkamp, Michael Wurst 1 Goals of this course Learn about Software tools Databases Methods (Algorithms) in
Introduction to Bioinformatics Dortmund, 16.-20.07.2007 Lectures: Sven Rahmann Exercises: Udo Feldkamp, Michael Wurst 1 Goals of this course Learn about Software tools Databases Methods (Algorithms) in
Sequence Databases and database scanning
 Sequence Databases and database scanning Marjolein Thunnissen Lund, 2012 Types of databases: Primary sequence databases (proteins and nucleic acids). Composite protein sequence databases. Secondary databases.
Sequence Databases and database scanning Marjolein Thunnissen Lund, 2012 Types of databases: Primary sequence databases (proteins and nucleic acids). Composite protein sequence databases. Secondary databases.
Identifying Regulatory Regions using Multiple Sequence Alignments
 Identifying Regulatory Regions using Multiple Sequence Alignments Prerequisites: BLAST Exercise: Detecting and Interpreting Genetic Homology. Resources: ClustalW is available at http://www.ebi.ac.uk/tools/clustalw2/index.html
Identifying Regulatory Regions using Multiple Sequence Alignments Prerequisites: BLAST Exercise: Detecting and Interpreting Genetic Homology. Resources: ClustalW is available at http://www.ebi.ac.uk/tools/clustalw2/index.html
Uncovering differentially expressed pathways with protein interaction and gene expression data
 The Second International Symposium on Optimization and Systems Biology (OSB 08) Lijiang, China, October 31 November 3, 2008 Copyright 2008 ORSC & APORC, pp. 74 82 Uncovering differentially expressed pathways
The Second International Symposium on Optimization and Systems Biology (OSB 08) Lijiang, China, October 31 November 3, 2008 Copyright 2008 ORSC & APORC, pp. 74 82 Uncovering differentially expressed pathways
Introduction to BIOINFORMATICS
 COURSE OF BIOINFORMATICS a.a. 2016-2017 Introduction to BIOINFORMATICS What is Bioinformatics? (I) The sinergy between biology and informatics What is Bioinformatics? (II) From: http://www.bioteach.ubc.ca/bioinfo2010/
COURSE OF BIOINFORMATICS a.a. 2016-2017 Introduction to BIOINFORMATICS What is Bioinformatics? (I) The sinergy between biology and informatics What is Bioinformatics? (II) From: http://www.bioteach.ubc.ca/bioinfo2010/
BIOINFORMATICS Introduction
 BIOINFORMATICS Introduction Mark Gerstein, Yale University bioinfo.mbb.yale.edu/mbb452a 1 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu What is Bioinformatics? (Molecular) Bio -informatics One idea
BIOINFORMATICS Introduction Mark Gerstein, Yale University bioinfo.mbb.yale.edu/mbb452a 1 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu What is Bioinformatics? (Molecular) Bio -informatics One idea
Analysis of ChIP-seq data with R / Bioconductor
 Analysis of ChIP-seq data with R / Bioconductor Martin Morgan Bioconductor / Fred Hutchinson Cancer Research Center Seattle, WA, USA 8-10 June 2009 ChIP-seq Chromatin immunopreciptation to enrich sample
Analysis of ChIP-seq data with R / Bioconductor Martin Morgan Bioconductor / Fred Hutchinson Cancer Research Center Seattle, WA, USA 8-10 June 2009 ChIP-seq Chromatin immunopreciptation to enrich sample
OrthoGibbs & PhyloScan
 OrthoGibbs & PhyloScan A comparative genomics approach to locating transcription factor binding sites Lee Newberg, Wadsworth Center, 6/19/2007 Acknowledgments Team: C. Steven Carmack (Wadsworth) Sean P.
OrthoGibbs & PhyloScan A comparative genomics approach to locating transcription factor binding sites Lee Newberg, Wadsworth Center, 6/19/2007 Acknowledgments Team: C. Steven Carmack (Wadsworth) Sean P.
Inferring protein DNA dependencies using motif alignments and mutual information
 BIOINFORMATICS Vol. 23 ISMB/ECCB 2007, pages i297 i304 doi:10.1093/bioinformatics/btm215 Inferring protein DNA dependencies using motif alignments and mutual information Shaun Mahony 1, *, Philip E. Auron
BIOINFORMATICS Vol. 23 ISMB/ECCB 2007, pages i297 i304 doi:10.1093/bioinformatics/btm215 Inferring protein DNA dependencies using motif alignments and mutual information Shaun Mahony 1, *, Philip E. Auron
ChIP-Seq Data Analysis. J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015
 ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA
ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA
ChIP-Seq Tools. J Fass UCD Genome Center Bioinformatics Core Wednesday September 16, 2015
 ChIP-Seq Tools J Fass UCD Genome Center Bioinformatics Core Wednesday September 16, 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA or
ChIP-Seq Tools J Fass UCD Genome Center Bioinformatics Core Wednesday September 16, 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA or
ChIP-Seq Data Analysis. J Fass UCD Genome Center Bioinformatics Core Wednesday December 17, 2014
 ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday December 17, 2014 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind
ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday December 17, 2014 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind
BIOINFORMATICS ORIGINAL PAPER
 BIOINFORMATICS ORIGINAL PAPER Vol. 28 no. 5 2012, pages 701 708 doi:10.1093/bioinformatics/bts002 Systems biology Advance Access publication January 11, 2012 De novo motif discovery facilitates identification
BIOINFORMATICS ORIGINAL PAPER Vol. 28 no. 5 2012, pages 701 708 doi:10.1093/bioinformatics/bts002 Systems biology Advance Access publication January 11, 2012 De novo motif discovery facilitates identification
Introduction to Bioinformatics Online Course: IBT
 Introduction to Bioinformatics Online Course: IBT Multiple Sequence Alignment Building Multiple Sequence Alignment Lec5: Interpreting your MSA Using Logos Using Logos - Logos are a terrific way to generate
Introduction to Bioinformatics Online Course: IBT Multiple Sequence Alignment Building Multiple Sequence Alignment Lec5: Interpreting your MSA Using Logos Using Logos - Logos are a terrific way to generate
Lecture 22 Eukaryotic Genes and Genomes III
 Lecture 22 Eukaryotic Genes and Genomes III In the last three lectures we have thought a lot about analyzing a regulatory system in S. cerevisiae, namely Gal regulation that involved a hand full of genes.
Lecture 22 Eukaryotic Genes and Genomes III In the last three lectures we have thought a lot about analyzing a regulatory system in S. cerevisiae, namely Gal regulation that involved a hand full of genes.
Improvement of TRANSFAC Matrices Using Multiple Local Alignment of Transcription Factor Binding Site Sequences
 68 Genome Informatics 16(1): 68 72 (2005) Improvement of TRANSFAC Matrices Using Multiple Local Alignment of Transcription Factor Binding Site Sequences Yutao Fu 1 Zhiping Weng 1,2 bibin@bu.edu zhiping@bu.edu
68 Genome Informatics 16(1): 68 72 (2005) Improvement of TRANSFAC Matrices Using Multiple Local Alignment of Transcription Factor Binding Site Sequences Yutao Fu 1 Zhiping Weng 1,2 bibin@bu.edu zhiping@bu.edu
Applying Machine Learning Strategy in Transcription Factor DNA Bindings Site Prediction
 Applying Machine Learning Strategy in Transcription Factor DNA Bindings Site Prediction Ziliang Qian Key Laboratory of Systems Biology Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences,
Applying Machine Learning Strategy in Transcription Factor DNA Bindings Site Prediction Ziliang Qian Key Laboratory of Systems Biology Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences,
CS313 Exercise 1 Cover Page Fall 2017
 CS313 Exercise 1 Cover Page Fall 2017 Due by the start of class on Monday, September 18, 2017. Name(s): In the TIME column, please estimate the time you spent on the parts of this exercise. Please try
CS313 Exercise 1 Cover Page Fall 2017 Due by the start of class on Monday, September 18, 2017. Name(s): In the TIME column, please estimate the time you spent on the parts of this exercise. Please try
ACMES: fast multiple-genome searches for short repeat sequences with concurrent cross-species information retrieval
 Nucleic Acids Research, 2004, Vol. 32, Web Server issue W649 W653 DOI 10.1093/nar/gkh455 ACMES fast multiple-genome searches for short repeat sequences with concurrent cross-species information retrieval
Nucleic Acids Research, 2004, Vol. 32, Web Server issue W649 W653 DOI 10.1093/nar/gkh455 ACMES fast multiple-genome searches for short repeat sequences with concurrent cross-species information retrieval
Browser Exercises - I. Alignments and Comparative genomics
 Browser Exercises - I Alignments and Comparative genomics 1. Navigating to the Genome Browser (GBrowse) Note: For this exercise use http://www.tritrypdb.org a. Navigate to the Genome Browser (GBrowse)
Browser Exercises - I Alignments and Comparative genomics 1. Navigating to the Genome Browser (GBrowse) Note: For this exercise use http://www.tritrypdb.org a. Navigate to the Genome Browser (GBrowse)
Lecture 15. Promoters, TFs. #15_Sept26
 BCB 444/544 Lecture 15 More Review: RNA, Proteins, Promoters, TFs Next time: Profiles & Hidden Markov Models (HMMs) #15_Sept26 BCB 444/544 F07 ISU Dobbs #15 - RNA, Proteins, Promoters, TFs 9/26/07 1 Required
BCB 444/544 Lecture 15 More Review: RNA, Proteins, Promoters, TFs Next time: Profiles & Hidden Markov Models (HMMs) #15_Sept26 BCB 444/544 F07 ISU Dobbs #15 - RNA, Proteins, Promoters, TFs 9/26/07 1 Required
Discovering gene regulatory control using ChIP-chip and ChIP-seq. Part 1. An introduction to gene regulatory control, concepts and methodologies
 Discovering gene regulatory control using ChIP-chip and ChIP-seq Part 1 An introduction to gene regulatory control, concepts and methodologies Ian Simpson ian.simpson@.ed.ac.uk http://bit.ly/bio2links
Discovering gene regulatory control using ChIP-chip and ChIP-seq Part 1 An introduction to gene regulatory control, concepts and methodologies Ian Simpson ian.simpson@.ed.ac.uk http://bit.ly/bio2links
GENES AND CHROMOSOMES V. Lecture 7. Biology Department Concordia University. Dr. S. Azam BIOL 266/
 1 GENES AND CHROMOSOMES V Lecture 7 BIOL 266/4 2014-15 Dr. S. Azam Biology Department Concordia University 2 CELL NUCLEUS AND THE CONTROL OF GENE EXPRESSION An Overview of Gene Regulation in Eukaryotes
1 GENES AND CHROMOSOMES V Lecture 7 BIOL 266/4 2014-15 Dr. S. Azam Biology Department Concordia University 2 CELL NUCLEUS AND THE CONTROL OF GENE EXPRESSION An Overview of Gene Regulation in Eukaryotes
Worksheet for Bioinformatics
 Worksheet for Bioinformatics ACTIVITY: Learn to use biological databases and sequence analysis tools Exercise 1 Biological Databases Objective: To use public biological databases to search for latest research
Worksheet for Bioinformatics ACTIVITY: Learn to use biological databases and sequence analysis tools Exercise 1 Biological Databases Objective: To use public biological databases to search for latest research
Motif Search CMSC 423
 Motif Search CMSC 423 Central Dogma of Biology proteins Translation mrna (T U) Transcription Genome DNA = double-stranded, linear molecule each strand is string over {A,C,G,T} strands are complements of
Motif Search CMSC 423 Central Dogma of Biology proteins Translation mrna (T U) Transcription Genome DNA = double-stranded, linear molecule each strand is string over {A,C,G,T} strands are complements of
Protein Sequence Analysis. BME 110: CompBio Tools Todd Lowe April 19, 2007 (Slide Presentation: Carol Rohl)
 Protein Sequence Analysis BME 110: CompBio Tools Todd Lowe April 19, 2007 (Slide Presentation: Carol Rohl) Linear Sequence Analysis What can you learn from a (single) protein sequence? Calculate it s physical
Protein Sequence Analysis BME 110: CompBio Tools Todd Lowe April 19, 2007 (Slide Presentation: Carol Rohl) Linear Sequence Analysis What can you learn from a (single) protein sequence? Calculate it s physical
MEMOFinder: combining de novo motif prediction methods with a database of known motifs
 MEMOFinder: combining de novo motif prediction methods with a database of known motifs Bartek Wilczy«ski, Miªosz Dar»ynkiewicz and Jerzy Tiuryn Institute of Informatics, University of Warsaw September
MEMOFinder: combining de novo motif prediction methods with a database of known motifs Bartek Wilczy«ski, Miªosz Dar»ynkiewicz and Jerzy Tiuryn Institute of Informatics, University of Warsaw September
Applications of HMMs in Epigenomics
 I529: Machine Learning in Bioinformatics (Spring 2013) Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Contents Background:
I529: Machine Learning in Bioinformatics (Spring 2013) Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Contents Background:
Measuring Protein-DNA interactions
 Measuring Protein-DNA interactions How is Biological Complexity Achieved? Mediated by Transcription Factors (TFs) 2 Transcription Factors are genetic switches 3 Regulation of Gene Expression by Transcription
Measuring Protein-DNA interactions How is Biological Complexity Achieved? Mediated by Transcription Factors (TFs) 2 Transcription Factors are genetic switches 3 Regulation of Gene Expression by Transcription
FUNCTIONAL BIOINFORMATICS
 Molecular Biology-2018 1 FUNCTIONAL BIOINFORMATICS PREDICTING THE FUNCTION OF AN UNKNOWN PROTEIN Suppose you have found the amino acid sequence of an unknown protein and wish to find its potential function.
Molecular Biology-2018 1 FUNCTIONAL BIOINFORMATICS PREDICTING THE FUNCTION OF AN UNKNOWN PROTEIN Suppose you have found the amino acid sequence of an unknown protein and wish to find its potential function.
A niched Pareto genetic algorithm for finding variable length regulatory motifs in DNA sequences
 3 Biotech (2012) 2:141 148 DOI 10.1007/s13205-011-0040-6 ORIGINAL ARTICLE A niched Pareto genetic algorithm for finding variable length regulatory motifs in DNA sequences Shripal Vijayvargiya Pratyoosh
3 Biotech (2012) 2:141 148 DOI 10.1007/s13205-011-0040-6 ORIGINAL ARTICLE A niched Pareto genetic algorithm for finding variable length regulatory motifs in DNA sequences Shripal Vijayvargiya Pratyoosh
Sequence Analysis. II: Sequence Patterns and Matrices. George Bell, Ph.D. WIBR Bioinformatics and Research Computing
 Sequence Analysis II: Sequence Patterns and Matrices George Bell, Ph.D. WIBR Bioinformatics and Research Computing Sequence Patterns and Matrices Multiple sequence alignments Sequence patterns Sequence
Sequence Analysis II: Sequence Patterns and Matrices George Bell, Ph.D. WIBR Bioinformatics and Research Computing Sequence Patterns and Matrices Multiple sequence alignments Sequence patterns Sequence
FINAL COPY - 12-Feb DNA Familial Binding Profiles Made Easy: Comparison of Various Motif Alignment and Clustering Strategies.
 FINAL COPY - 12-Feb-2007 Accepted in PLoS Computational Biology DNA Familial Binding Profiles Made Easy: Comparison of Various Motif Alignment and Clustering Strategies Shaun Mahony 1,2 Philip E. Auron
FINAL COPY - 12-Feb-2007 Accepted in PLoS Computational Biology DNA Familial Binding Profiles Made Easy: Comparison of Various Motif Alignment and Clustering Strategies Shaun Mahony 1,2 Philip E. Auron
Genome Sequence Assembly
 Genome Sequence Assembly Learning Goals: Introduce the field of bioinformatics Familiarize the student with performing sequence alignments Understand the assembly process in genome sequencing Introduction:
Genome Sequence Assembly Learning Goals: Introduce the field of bioinformatics Familiarize the student with performing sequence alignments Understand the assembly process in genome sequencing Introduction:
Textbook Reading Guidelines
 Understanding Bioinformatics by Marketa Zvelebil and Jeremy Baum Last updated: January 16, 2013 Textbook Reading Guidelines Preface: Read the whole preface, and especially: For the students with Life Science
Understanding Bioinformatics by Marketa Zvelebil and Jeremy Baum Last updated: January 16, 2013 Textbook Reading Guidelines Preface: Read the whole preface, and especially: For the students with Life Science
Integrating Genomic Data to Predict Transcription
 Genome Informatics 16(1): 83 94 (2005) 83 Integrating Genomic Data to Predict Transcription Factor Binding Dustin dth128@bu. T. Holloway1 edu Mark mkon@bu. Kon2 edu Charles delisi@bu. DeLisi3 I Molecular
Genome Informatics 16(1): 83 94 (2005) 83 Integrating Genomic Data to Predict Transcription Factor Binding Dustin dth128@bu. T. Holloway1 edu Mark mkon@bu. Kon2 edu Charles delisi@bu. DeLisi3 I Molecular
ChIP. November 21, 2017
 ChIP November 21, 2017 functional signals: is DNA enough? what is the smallest number of letters used by a written language? DNA is only one part of the functional genome DNA is heavily bound by proteins,
ChIP November 21, 2017 functional signals: is DNA enough? what is the smallest number of letters used by a written language? DNA is only one part of the functional genome DNA is heavily bound by proteins,
Motif Discovery from Large Number of Sequences: a Case Study with Disease Resistance Genes in Arabidopsis thaliana
 Motif Discovery from Large Number of Sequences: a Case Study with Disease Resistance Genes in Arabidopsis thaliana Irfan Gunduz, Sihui Zhao, Mehmet Dalkilic and Sun Kim Indiana University, School of Informatics
Motif Discovery from Large Number of Sequences: a Case Study with Disease Resistance Genes in Arabidopsis thaliana Irfan Gunduz, Sihui Zhao, Mehmet Dalkilic and Sun Kim Indiana University, School of Informatics
Gene Identification in silico
 Gene Identification in silico Nita Parekh, IIIT Hyderabad Presented at National Seminar on Bioinformatics and Functional Genomics, at Bioinformatics centre, Pondicherry University, Feb 15 17, 2006. Introduction
Gene Identification in silico Nita Parekh, IIIT Hyderabad Presented at National Seminar on Bioinformatics and Functional Genomics, at Bioinformatics centre, Pondicherry University, Feb 15 17, 2006. Introduction
BMC Bioinformatics. Open Access. Abstract
 BMC Bioinformatics BioMed Central Research article Defining transcriptional networks through integrative modeling of mrna expression and transcription factor binding data Feng Gao 1, Barrett C Foat 1 and
BMC Bioinformatics BioMed Central Research article Defining transcriptional networks through integrative modeling of mrna expression and transcription factor binding data Feng Gao 1, Barrett C Foat 1 and
Introduction and History of Genome Modification. Adam Clore, PhD Director, Synthetic Biology Design
 Introduction and History of Genome Modification Adam Clore, PhD Director, Synthetic Biology Design Early Non-site Directed Genome Modification Homologous recombination in yeast TARGET GENE 5 Arm URA3 3
Introduction and History of Genome Modification Adam Clore, PhD Director, Synthetic Biology Design Early Non-site Directed Genome Modification Homologous recombination in yeast TARGET GENE 5 Arm URA3 3
This place covers: Methods or systems for genetic or protein-related data processing in computational molecular biology.
 G16B BIOINFORMATICS, i.e. INFORMATION AND COMMUNICATION TECHNOLOGY [ICT] SPECIALLY ADAPTED FOR GENETIC OR PROTEIN-RELATED DATA PROCESSING IN COMPUTATIONAL MOLECULAR BIOLOGY Methods or systems for genetic
G16B BIOINFORMATICS, i.e. INFORMATION AND COMMUNICATION TECHNOLOGY [ICT] SPECIALLY ADAPTED FOR GENETIC OR PROTEIN-RELATED DATA PROCESSING IN COMPUTATIONAL MOLECULAR BIOLOGY Methods or systems for genetic
Genome-Wide Survey of MicroRNA - Transcription Factor Feed-Forward Regulatory Circuits in Human. Supporting Information
 Genome-Wide Survey of MicroRNA - Transcription Factor Feed-Forward Regulatory Circuits in Human Angela Re #, Davide Corá #, Daniela Taverna and Michele Caselle # equal contribution * corresponding author,
Genome-Wide Survey of MicroRNA - Transcription Factor Feed-Forward Regulatory Circuits in Human Angela Re #, Davide Corá #, Daniela Taverna and Michele Caselle # equal contribution * corresponding author,
Following text taken from Suresh Kumar. Bioinformatics Web - Comprehensive educational resource on Bioinformatics. 6th May.2005
 Bioinformatics is the recording, annotation, storage, analysis, and searching/retrieval of nucleic acid sequence (genes and RNAs), protein sequence and structural information. This includes databases of
Bioinformatics is the recording, annotation, storage, analysis, and searching/retrieval of nucleic acid sequence (genes and RNAs), protein sequence and structural information. This includes databases of
Discovering gene regulatory control using ChIP-chip and ChIP-seq. An introduction to gene regulatory control, concepts and methodologies
 Discovering gene regulatory control using ChIP-chip and ChIP-seq An introduction to gene regulatory control, concepts and methodologies Ian Simpson ian.simpson@.ed.ac.uk bit.ly/bio2_2012 The Central Dogma
Discovering gene regulatory control using ChIP-chip and ChIP-seq An introduction to gene regulatory control, concepts and methodologies Ian Simpson ian.simpson@.ed.ac.uk bit.ly/bio2_2012 The Central Dogma
Integrating Genomic Data to Predict Transcription Factor Binding
 Genome Informatics 16(1): 83 94 (2005) 83 Integrating Genomic Data to Predict Transcription Factor Binding Dustin T. Holloway 1 Mark Kon 2 Charles DeLisi 3 dth128@bu.edu mkon@bu.edu delisi@bu.edu 1 Molecular
Genome Informatics 16(1): 83 94 (2005) 83 Integrating Genomic Data to Predict Transcription Factor Binding Dustin T. Holloway 1 Mark Kon 2 Charles DeLisi 3 dth128@bu.edu mkon@bu.edu delisi@bu.edu 1 Molecular
RSIR: Regularized Sliced Inverse Regression for Motif Discovery
 RSIR: Regularized Sliced Inverse Regression for Motif Discovery Wenxuan Zhong 1,, Peng Zeng 1,,PingMa 1,, Jun S. Liu 2, andyuzhu 2, Department of Statistics, Harvard University, Cambridge, MA 02138, USA,
RSIR: Regularized Sliced Inverse Regression for Motif Discovery Wenxuan Zhong 1,, Peng Zeng 1,,PingMa 1,, Jun S. Liu 2, andyuzhu 2, Department of Statistics, Harvard University, Cambridge, MA 02138, USA,
1 Najafabadi, H. S. et al. C2H2 zinc finger proteins greatly expand the human regulatory lexicon. Nat Biotechnol doi: /nbt.3128 (2015).
 F op-scoring motif Optimized motifs E Input sequences entral 1 bp region Dinucleotideshuffled seqs B D ll B1H-R predicted motifs Enriched B1H- R predicted motifs L!=!7! L!=!6! L!=5! L!=!4! L!=!3! L!=!2!
F op-scoring motif Optimized motifs E Input sequences entral 1 bp region Dinucleotideshuffled seqs B D ll B1H-R predicted motifs Enriched B1H- R predicted motifs L!=!7! L!=!6! L!=5! L!=!4! L!=!3! L!=!2!
2/19/13. Contents. Applications of HMMs in Epigenomics
 2/19/13 I529: Machine Learning in Bioinformatics (Spring 2013) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Background:
2/19/13 I529: Machine Learning in Bioinformatics (Spring 2013) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Background:
Gene expression analysis. Biosciences 741: Genomics Fall, 2013 Week 5. Gene expression analysis
 Gene expression analysis Biosciences 741: Genomics Fall, 2013 Week 5 Gene expression analysis From EST clusters to spotted cdna microarrays Long vs. short oligonucleotide microarrays vs. RT-PCR Methods
Gene expression analysis Biosciences 741: Genomics Fall, 2013 Week 5 Gene expression analysis From EST clusters to spotted cdna microarrays Long vs. short oligonucleotide microarrays vs. RT-PCR Methods
New Plant Breeding Techniques: Zn Finger Nucleases and Transcription Factors
 New Plant Breeding Techniques: Zn Finger Nucleases and Transcription Factors Andrew F. Roberts, Ph.D. Deputy Director, CERA September 19, 2013 Contents of the talk Old Plant Breeding Techniques and Biosafety
New Plant Breeding Techniques: Zn Finger Nucleases and Transcription Factors Andrew F. Roberts, Ph.D. Deputy Director, CERA September 19, 2013 Contents of the talk Old Plant Breeding Techniques and Biosafety
Gibbs Sampling and Centroids for Gene Regulation
 Gibbs Sampling and Centroids for Gene Regulation NY State Dept. of Health Wadsworth Center @ Albany Chapter American Statistical Association Acknowledgments Team: Sean P. Conlan (National Institutes of
Gibbs Sampling and Centroids for Gene Regulation NY State Dept. of Health Wadsworth Center @ Albany Chapter American Statistical Association Acknowledgments Team: Sean P. Conlan (National Institutes of
Sequence Analysis. BBSI 2006: Lecture #(χ+3) Takis Benos (2006) BBSI MAY P. Benos 1
 Sequence Analysis (part III) BBSI 2006: Lecture #(χ+3) Takis Benos (2006) BBSI 2006 31-MAY-2006 2006 P. Benos 1 Outline Sequence variation Distance measures Scoring matrices Pairwise alignments (global,
Sequence Analysis (part III) BBSI 2006: Lecture #(χ+3) Takis Benos (2006) BBSI 2006 31-MAY-2006 2006 P. Benos 1 Outline Sequence variation Distance measures Scoring matrices Pairwise alignments (global,
Protein function prediction using sequence motifs: A research proposal
 Protein function prediction using sequence motifs: A research proposal Asa Ben-Hur Abstract Protein function prediction, i.e. classification of protein sequences according to their biological function
Protein function prediction using sequence motifs: A research proposal Asa Ben-Hur Abstract Protein function prediction, i.e. classification of protein sequences according to their biological function
Bayesian Variable Selection and Data Integration for Biological Regulatory Networks
 Bayesian Variable Selection and Data Integration for Biological Regulatory Networks Shane T. Jensen Department of Statistics The Wharton School, University of Pennsylvania stjensen@wharton.upenn.edu Gary
Bayesian Variable Selection and Data Integration for Biological Regulatory Networks Shane T. Jensen Department of Statistics The Wharton School, University of Pennsylvania stjensen@wharton.upenn.edu Gary
Question 2: There are 5 retroelements (2 LINEs and 3 LTRs), 6 unclassified elements (XDMR and XDMR_DM), and 7 satellite sequences.
 Bio4342 Exercise 1 Answers: Detecting and Interpreting Genetic Homology (Answers prepared by Wilson Leung) Question 1: Low complexity DNA can be described as sequences that consist primarily of one or
Bio4342 Exercise 1 Answers: Detecting and Interpreting Genetic Homology (Answers prepared by Wilson Leung) Question 1: Low complexity DNA can be described as sequences that consist primarily of one or
Assessment of Algorithms for Inferring Positional Weight Matrix Motifs of Transcription Factor Binding Sites Using Protein Binding Microarray Data
 Assessment of Algorithms for Inferring Positional Weight Matrix Motifs of Transcription Factor Binding Sites Using Protein Binding Microarray Data Yaron Orenstein, Chaim Linhart, Ron Shamir* Blavatnik
Assessment of Algorithms for Inferring Positional Weight Matrix Motifs of Transcription Factor Binding Sites Using Protein Binding Microarray Data Yaron Orenstein, Chaim Linhart, Ron Shamir* Blavatnik
CS273B: Deep learning for Genomics and Biomedicine
 CS273B: Deep learning for Genomics and Biomedicine Lecture 2: Convolutional neural networks and applications to functional genomics 09/28/2016 Anshul Kundaje, James Zou, Serafim Batzoglou Outline Anatomy
CS273B: Deep learning for Genomics and Biomedicine Lecture 2: Convolutional neural networks and applications to functional genomics 09/28/2016 Anshul Kundaje, James Zou, Serafim Batzoglou Outline Anatomy
COMPUTER RESOURCES II:
 COMPUTER RESOURCES II: Using the computer to analyze data, using the internet, and accessing online databases Bio 210, Fall 2006 Linda S. Huang, Ph.D. University of Massachusetts Boston In the first computer
COMPUTER RESOURCES II: Using the computer to analyze data, using the internet, and accessing online databases Bio 210, Fall 2006 Linda S. Huang, Ph.D. University of Massachusetts Boston In the first computer
Transcription factor binding site prediction in vivo using DNA sequence and shape features
 Transcription factor binding site prediction in vivo using DNA sequence and shape features Anthony Mathelier, Lin Yang, Tsu-Pei Chiu, Remo Rohs, and Wyeth Wasserman anthony.mathelier@gmail.com @AMathelier
Transcription factor binding site prediction in vivo using DNA sequence and shape features Anthony Mathelier, Lin Yang, Tsu-Pei Chiu, Remo Rohs, and Wyeth Wasserman anthony.mathelier@gmail.com @AMathelier
Identifying Cooperativity Among Transcription Factors Controlling the Cell Cycle In Yeast
 Identifying Cooperativity Among Transcription Factors Controlling the Cell Cycle In Yeast Nilanjana Banerjee 1,2 and Michael Q. Zhang 1 Nucleic Acids Research, 2003, Vol.31, No.23 1 Cold Spring Harbor
Identifying Cooperativity Among Transcription Factors Controlling the Cell Cycle In Yeast Nilanjana Banerjee 1,2 and Michael Q. Zhang 1 Nucleic Acids Research, 2003, Vol.31, No.23 1 Cold Spring Harbor
VALLIAMMAI ENGINEERING COLLEGE
 VALLIAMMAI ENGINEERING COLLEGE SRM Nagar, Kattankulathur 603 203 DEPARTMENT OF COMPUTER SCIENCE AND ENGINEERING QUESTION BANK VII SEMESTER BM6005 BIO INFORMATICS Regulation 2013 Academic Year 2018-19 Prepared
VALLIAMMAI ENGINEERING COLLEGE SRM Nagar, Kattankulathur 603 203 DEPARTMENT OF COMPUTER SCIENCE AND ENGINEERING QUESTION BANK VII SEMESTER BM6005 BIO INFORMATICS Regulation 2013 Academic Year 2018-19 Prepared
Übung V. Einführung, Teil 1. Transktiptionelle Regulation TFBS
 Übung V Einführung, Teil 1 Transktiptionelle Regulation TFBS Transcription Factors These proteins promote transcription 1. Bind DNA 2. Activate Transcription These two functions usually reside on separate
Übung V Einführung, Teil 1 Transktiptionelle Regulation TFBS Transcription Factors These proteins promote transcription 1. Bind DNA 2. Activate Transcription These two functions usually reside on separate
Community-assisted genome annotation: The Pseudomonas example. Geoff Winsor, Simon Fraser University Burnaby (greater Vancouver), Canada
 Community-assisted genome annotation: The Pseudomonas example Geoff Winsor, Simon Fraser University Burnaby (greater Vancouver), Canada Overview Pseudomonas Community Annotation Project (PseudoCAP) Past
Community-assisted genome annotation: The Pseudomonas example Geoff Winsor, Simon Fraser University Burnaby (greater Vancouver), Canada Overview Pseudomonas Community Annotation Project (PseudoCAP) Past
Chapter 25: Regulating Eukaryotic Transcription The Ligand Responsive Activators
 Chapter 25: Regulating Eukaryotic Transcription The Ligand Responsive Activators At least 5 potential gene expression control points Superfamily of Gene Regulators Activation of gene structure Initiation
Chapter 25: Regulating Eukaryotic Transcription The Ligand Responsive Activators At least 5 potential gene expression control points Superfamily of Gene Regulators Activation of gene structure Initiation
DEVELOPING WEB TOOLS FOR DATA MINING AND ANALYSIS OF SAGE
 DEVELOPING WEB TOOLS FOR DATA MINING AND ANALYSIS OF SAGE Kristin Wheeler BBSI, University of Pittsburgh Grambling State University Panayiotis Benos,Ph.d Center for Computational Biology & Bioinformatics
DEVELOPING WEB TOOLS FOR DATA MINING AND ANALYSIS OF SAGE Kristin Wheeler BBSI, University of Pittsburgh Grambling State University Panayiotis Benos,Ph.d Center for Computational Biology & Bioinformatics
Computational Genomics
 Computational Genomics http://www.cs.cmu.edu/~02710 Ziv Bar-Joseph zivbj@cs.cmu.edu GHC 8006 Chakra Chennubhotla chakracs@pitt.edu Suite 3064, BST3 Topics Introduction (1 Week) Sequence analysis(4 weeks)
Computational Genomics http://www.cs.cmu.edu/~02710 Ziv Bar-Joseph zivbj@cs.cmu.edu GHC 8006 Chakra Chennubhotla chakracs@pitt.edu Suite 3064, BST3 Topics Introduction (1 Week) Sequence analysis(4 weeks)
Experiments on the Accuracy of Algorithms for Inferring the Structure of Genetic Regulatory Networks from Microarray Expression Levels 1
 Experiments on the Accuracy of Algorithms for Inferring the Structure of Genetic Regulatory Networks from Microarray Expression Levels 1 Frank C. Wimberly, Institute for Human and Machine Cognition, University
Experiments on the Accuracy of Algorithms for Inferring the Structure of Genetic Regulatory Networks from Microarray Expression Levels 1 Frank C. Wimberly, Institute for Human and Machine Cognition, University
In silico identification of transcriptional regulatory regions
 In silico identification of transcriptional regulatory regions Martti Tolvanen, IMT Bioinformatics, University of Tampere Eija Korpelainen, CSC Jarno Tuimala, CSC Introduction (Eija) Program Retrieval
In silico identification of transcriptional regulatory regions Martti Tolvanen, IMT Bioinformatics, University of Tampere Eija Korpelainen, CSC Jarno Tuimala, CSC Introduction (Eija) Program Retrieval
Introduction to Cellular Biology and Bioinformatics. Farzaneh Salari
 Introduction to Cellular Biology and Bioinformatics Farzaneh Salari Outline Bioinformatics Cellular Biology A Bioinformatics Problem What is bioinformatics? Computer Science Statistics Bioinformatics Mathematics...
Introduction to Cellular Biology and Bioinformatics Farzaneh Salari Outline Bioinformatics Cellular Biology A Bioinformatics Problem What is bioinformatics? Computer Science Statistics Bioinformatics Mathematics...
Sequence Analysis Lab Protocol
 Sequence Analysis Lab Protocol You will need this handout of instructions The sequence of your plasmid from the ABI The Accession number for Lambda DNA J02459 The Accession number for puc 18 is L09136
Sequence Analysis Lab Protocol You will need this handout of instructions The sequence of your plasmid from the ABI The Accession number for Lambda DNA J02459 The Accession number for puc 18 is L09136
KDM1B as a Link Between Histone Modification and DNA Methylation of Gene Imprinting During Gametogenesis
 KDM1B as a Link Between Histone Modification and DNA Methylation of Gene Imprinting During Gametogenesis BI 424 Advanced Molecular Genetics, Winter 2011 3/14/11 Abstract Genomic imprinting is a highly
KDM1B as a Link Between Histone Modification and DNA Methylation of Gene Imprinting During Gametogenesis BI 424 Advanced Molecular Genetics, Winter 2011 3/14/11 Abstract Genomic imprinting is a highly
Computational approaches to the discovery of regulatory elements in noncoding DNA. Michael Koldobskiy
 Computational approaches to the discovery of regulatory elements in noncoding DNA Michael Koldobskiy MB&B 452a December 13, 2002 INTRODUCTION Biological research in the post-genomic era has been charged
Computational approaches to the discovery of regulatory elements in noncoding DNA Michael Koldobskiy MB&B 452a December 13, 2002 INTRODUCTION Biological research in the post-genomic era has been charged
