Bayesian Variable Selection and Data Integration for Biological Regulatory Networks
|
|
|
- Anastasia Banks
- 6 years ago
- Views:
Transcription
1 Bayesian Variable Selection and Data Integration for Biological Regulatory Networks Shane T. Jensen Department of Statistics The Wharton School, University of Pennsylvania Gary Chen and Christian Stoeckert, Jr Department of Bioengineering and Department of Genetics University of Pennsylvania Shane T. Jensen 1 March 5, 2008
2 Motivation Genes are long sequences of DNA that are transcribed to eventually become a protein Near-identical genetic material can lead to many different cell types and species A critical aspect of cellular function is how genes are regulated and which genes are regulated together Shane T. Jensen 2 March 5, 2008
3 Gene Regulatory Networks Genes are regulated by transcription factor (TF) proteins that bind directly to the DNA sequence near to a gene The bound protein affects the amount of transcription, thereby affecting the amount of protein produced The collection of TFs and their target genes is often called the gene regulatory network Goal is to elucidate regulatory network: which genes are targeted for regulation by a particuler TF? Shane T. Jensen 3 March 5, 2008
4 Different Data Types Gene expression data: microarray chips used amounts of mrna present for each gene across many conditions ChIP binding data: antibodies used to identify areas of genome physically bound by a particular TF Promoter element data: binding sites for a TF discovered by a sequence search algorithm Shane T. Jensen 4 March 5, 2008
5 Gene Expression Data Gene expression: measure of whether gene is turned on or turned off at a specific time Genes with similar expression across time or in different conditions may be coregulated Detect groups of genes that have correlated gene expression across many conditions Shane T. Jensen 5 March 5, 2008
6 ChIP Binding Data Chromatin Immunoprecipitation Experiments Antibodies used to pull out parts of genomic sequence that are physically bound to a particular TF Genes in close proximity to a TF binding site are possibly regulatory targets of that TF Shane T. Jensen 6 March 5, 2008
7 Promoter Element Data Some known promoter elements: the set of sequence binding sites recognized by a particular TF Promoter elements highly conserved but not identical: A C G T atgacgtctagcatcgaaatcgacgacgatcgacgactagctactctacgatcg aaaacatcgattgacgtttggtcgtaactttggcacgatcagcgatcgatcact aacagctatgacgtcgaaatcgaacatcgagacggacggcaacgtctacgatcg aaaacatcagctagcagcactagctaggattgacgtttggtcgtaactttggct aattatgctacgtgacgtacacgtacgtgacggactaagtcagctagcgtagct aattatgctacgtacgcggctcgctacactgacggagcatcaggtatttgacgt aaaaggcatcagctagcagcactagctaggtgacctggtcgtaactttggct aattatgctacgtggcgtacacgtacgtgacggactaagtcagctagcgtagct Matrix used to scan genomic sequences for putative promoter elements, which are then used to predict regulated genes Shane T. Jensen 7 March 5, 2008
8 Problem with Standard Methods These data sources, when used by themselves, provide only partial information for regulation: expression data gives only evidence of co-expression, not necessarily co-regulation ChIP binding data gives only evidence of physical TF binding, but binding is not necessarily functional promoter element data gives only possibility of TF binding site, but site may not be functional Need a principled approach to combine these complementary, but heterogeneous, sources of information Shane T. Jensen 8 March 5, 2008
9 Available Data Data: expression, ChIP binding, and promoter element data for 106 TFs in Yeast gene expression data across T different experiments g it = log-expression of gene i in experiment t f jt = log-expression of TF j in experiment t ChIP binding data for each gene i and TF j b ij = probability that TF j physically binds near gene i promoter element data for each gene i and TF j m ij = probability that gene i has a binding site for TF j Shane T. Jensen 9 March 5, 2008
10 Regulatory Indicators Regulatory network is formulated as unknown indicators: C ij =1 C ij =0 if gene i is actually regulated by TF j otherwise These C ij variables give the edges that connect TFs and their target genes on a regulatory graph C will be inferred using a Bayesian hierarchical model principled framework for combining heterogeneous data sources by using informed prior distributions Shane T. Jensen 10 March 5, 2008
11 Likelihood Model First model level involves target gene expression g it as a linear function of TF expression: g it = α i + j β j C ij f jt + ɛ it Error term is normally distributed: ɛ it Normal(0,σ 2 ) Regulation indicators C ij perform variable selection : only TFs j with C ij =1involved in expression of target gene i Biological reality: often the simultaneous action of multiple TFs are needed to change target gene expression Shane T. Jensen 11 March 5, 2008
12 Likelihood Model II We allow for synergistic relationships between pairs of TFs by also including interaction terms in our model: g it = α i + j β j C ij f jt + j k γ jk C ij C ik f jt f kt + ɛ it Sign of each interaction coefficient γ jk is unrestricted, so we are allowing for both synergistic and antagonistic relationships between pairs of TFs Non-informative priors used for parameters: α, β, γ, σ 2 Shane T. Jensen 12 March 5, 2008
13 Informed Prior Distribution Second model level is an informed prior distribution for our unknown regulation indicators C ij that involves both ChIP binding data b ij and promoter element data m ij : p(c ij m ij,b ij ) [ b C ij ij (1 b ij) 1 C ij ] wj [ ] m C ij ij (1 m ij) 1 C 1 wj ij Weight w j balances prior ChIP-binding information b ij vs prior promoter element information m ij Weights w j are TF-specific and reflect relative quality of ChIP binding data vs. promoter element data for TF j each w j treated as unknown variable with uniform prior Shane T. Jensen 13 March 5, 2008
14 Network Sparsity The probabilities from both ChIP binding data and promoter element data are mostly near zero: Density ChIP binding probs Sequence motif probs Values of b or m Prior implication that the network is quite sparse: each TF regulates only a small proportion of genes Shane T. Jensen 14 March 5, 2008
15 Implementation Get draws from joint posterior distribution using a Gibbs sampling strategy. 1. Sampling α, β, γ, σ 2 given C, w, g, f, b, m standard random effects model 2. Sampling each C ij given α, β, γ, σ 2, w, g, f, b, m easy 0-1 posterior probability calculation for each C ij 3. Sampling each w j given C, α, β, γ, σ 2, g, f, b, m grid sampler over the (0,1) range Shane T. Jensen 15 March 5, 2008
16 Inference Inference 1: posterior samples of C ij used to infer target genes for each TF j gene i is a target of TF j P(C ij =1 Y) > 0.5 Inference 2: posterior samples of interaction coefs γ jk used to find TF pairs with significant relationship Inference 3: posterior samples of weights w j used to infer quality of ChIP vs. promoter element data for different TFs Shane T. Jensen 16 March 5, 2008
17 Comparison of Predictions Primary goal is prediction of target genes based on estimated posterior probability P(C ij =1 Y) > 0.5 Can compare to several other current approaches: 1. MA-Networker: Gao et.al GRAM: Bar-Joseph et.al ReMoDiscovery: Lemmens et.al Two external measures used for validation 1. similarity of MIPS functions between target genes 2. response of target genes to TF knockout Shane T. Jensen 17 March 5, 2008
18 MIPS functional categories Each gene in Yeast has an assigned MIPS functional category from Munich information center for protein sequences Gene targets with similar functions are more likely be in same biological pathway, which validates the inference that they are regulated by a common transcription factor Calculated fraction of inferred target genes that shared similar functional categories for each TF, and then averaged across all TFs Shane T. Jensen 18 March 5, 2008
19 Fraction of Target Genes with Similar Functional Category Our Model Previous Methods Thresholded Data All 3 Exp+ChIP Exp Only MA Networker GRAM ReMoDiscovery Binding Expression Gene targets from our full model have slightly higher functional similarity than other methods All integration methods better than single data source Shane T. Jensen 19 March 5, 2008
20 Knockout Experiments Knockout experiments are gold standard for regulatory activity of individual TFs Knockout strain of yeast was created with a specific TF removed from the genome. Gene targets of knocked-out TF should show large response between wild-type and knock-out strains Calculated t-statistic of response to TF knockout for inferred target genes for 4 available knockout expts Shane T. Jensen 20 March 5, 2008
21 T-statistic for Knockout Response GCN4 knockout experiment SWI4 knockout experiment Our Model Previous Methods Thresholded Data Our Model Previous Methods Thresholded Data All 3 ExpChIP Exp MANet GRAM ReMo Bind Exp All 3 ExpChIP Exp MANet GRAM ReMo Bind Exp YAP1 knockout experiment SWI5 knockout experiment Our Model Previous Methods Thresholded Data Our Model Previous Methods Thresholded Data All 3 ExpChIP Exp MANet GRAM ReMo Bind Exp All 3 ExpChIP Exp MANet GRAM ReMo Bind Exp Our gene targets show greater response to TF knockout across all 4 knockout experiments Shane T. Jensen 21 March 5, 2008
22 Inference for Weight Variables Posterior distributions of w j variables for same 39 TFs: K K K K ABF1 ACE2 BAS1 CAD1 CBF1 FKH1 FKH2 GAL4 GCN4 GCR1 GCR2 HAP2 HAP3 HAP4 HSF1 INO2 LEU3 MBP1 MCM1 MET31 MSN4 NDD1 PDR1 PHO4 PUT3 RAP1 RCS1 REB1 RLM11 RME1 ROX1 SKN7 SMP1 STB1 STE12 SWI4 SWI5 SWI6 YAP1 Centered substantially higher than 0.5: suggests that ChIP binding data is generally superior to promoter element data Shane T. Jensen 22 March 5, 2008
23 Interactions between TFs Many recent papers have focused on combinatorial relationships between TFs Which pairs of TFs bind to same set of target genes? We can address this question by examining the posterior distribution of each interaction effect γ jk Positive γ jk s suggest a synergistic relationship, whereas negative γ jk s suggest an antagonistic relationship In our Yeast application, we found that 84 TF pairs have significant γ jk coefficients Shane T. Jensen 23 March 5, 2008
24 Interactions between TFs Many predicted interactions are known and involved in several important pathways Nodes = TFs and edges = significant interactions Shane T. Jensen 24 March 5, 2008
25 Mouse Application Also applied our model to one Mouse TF, C/EBP-β, which has all three data types available We identified 14/16 validated C/EBP-β targets More targets missed when using only single data source Our model also potentially reduces false positives: we predict 38 target genes compared to 72 predicted from expression data alone or 779 from ChIP data alone Estimated weight of w =0.92 for favoring ChIP binding data over promoter element data promoter element data useful in some instances, but generally less discriminative power than ChIP data Shane T. Jensen 25 March 5, 2008
26 Summary Combining multiple data sources (expression, ChIP binding and promoter element data) leads to improved predictions Bayesian hierarchical model is a natural framework for integrating heterogenous data sources Most Bayesian variable selection approaches use non-informative priors for selection indicators Our approach uses informed priors for our selection indicators based on addditional data sources Shane T. Jensen 26 March 5, 2008
27 Summary II Fully probabilistic approach: no reliance pre-clustering of data or dependence on arbitrary parameter cutoffs Flexibility for genes to belong to multiple regulatory clusters and pairs of transcription factors to interact Variable weight methodology achieves appropriate balance of priors: we confirm common belief that promoter element data is less reliable, but useful in some cases Shane T. Jensen 27 March 5, 2008
28 References Chen, G., Jensen, S.T. and Stoeckert, C. (2007). "Clustering of Genes into Regulons using Integrated Modeling." Genome Biology 8:R4 Jensen, S.T., Chen, G., and Stoeckert, C. (2007). "Bayesian Variable Selection and Data Integration for Biological Regulatory Networks." Annals of Applied Statistics 1: Shane T. Jensen 28 March 5, 2008
Predicting eukaryotic transcriptional cooperativity by Bayesian network integration of genome-wide data
 Published online 6 August 2009 Nucleic Acids Research, 2009, Vol. 37, No. 18 5943 5958 doi:10.1093/nar/gkp625 Predicting eukaryotic transcriptional cooperativity by Bayesian network integration of genome-wide
Published online 6 August 2009 Nucleic Acids Research, 2009, Vol. 37, No. 18 5943 5958 doi:10.1093/nar/gkp625 Predicting eukaryotic transcriptional cooperativity by Bayesian network integration of genome-wide
Technical University of Denmark
 1 of 13 Technical University of Denmark Written exam, 15 December 2007 Course name: Introduction to Systems Biology Course no. 27041 Aids allowed: Open Book Exam Provide your answers and calculations on
1 of 13 Technical University of Denmark Written exam, 15 December 2007 Course name: Introduction to Systems Biology Course no. 27041 Aids allowed: Open Book Exam Provide your answers and calculations on
Machine learning applications in genomics: practical issues & challenges. Yuzhen Ye School of Informatics and Computing, Indiana University
 Machine learning applications in genomics: practical issues & challenges Yuzhen Ye School of Informatics and Computing, Indiana University Reference Machine learning applications in genetics and genomics
Machine learning applications in genomics: practical issues & challenges Yuzhen Ye School of Informatics and Computing, Indiana University Reference Machine learning applications in genetics and genomics
Network System Inference
 Network System Inference Francis J. Doyle III University of California, Santa Barbara Douglas Lauffenburger Massachusetts Institute of Technology WTEC Systems Biology Final Workshop March 11, 2005 What
Network System Inference Francis J. Doyle III University of California, Santa Barbara Douglas Lauffenburger Massachusetts Institute of Technology WTEC Systems Biology Final Workshop March 11, 2005 What
Lecture 7: April 7, 2005
 Analysis of Gene Expression Data Spring Semester, 2005 Lecture 7: April 7, 2005 Lecturer: R.Shamir and C.Linhart Scribe: A.Mosseri, E.Hirsh and Z.Bronstein 1 7.1 Promoter Analysis 7.1.1 Introduction to
Analysis of Gene Expression Data Spring Semester, 2005 Lecture 7: April 7, 2005 Lecturer: R.Shamir and C.Linhart Scribe: A.Mosseri, E.Hirsh and Z.Bronstein 1 7.1 Promoter Analysis 7.1.1 Introduction to
Identifying Signaling Pathways. BMI/CS 776 Spring 2016 Anthony Gitter
 Identifying Signaling Pathways BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2016 Anthony Gitter gitter@biostat.wisc.edu Goals for lecture Challenges of integrating high-throughput assays Connecting relevant
Identifying Signaling Pathways BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2016 Anthony Gitter gitter@biostat.wisc.edu Goals for lecture Challenges of integrating high-throughput assays Connecting relevant
Whole Transcriptome Analysis of Illumina RNA- Seq Data. Ryan Peters Field Application Specialist
 Whole Transcriptome Analysis of Illumina RNA- Seq Data Ryan Peters Field Application Specialist Partek GS in your NGS Pipeline Your Start-to-Finish Solution for Analysis of Next Generation Sequencing Data
Whole Transcriptome Analysis of Illumina RNA- Seq Data Ryan Peters Field Application Specialist Partek GS in your NGS Pipeline Your Start-to-Finish Solution for Analysis of Next Generation Sequencing Data
Bayesian Networks as framework for data integration
 Bayesian Networks as framework for data integration Jun Zhu, Ph. D. Department of Genomics and Genetic Sciences Icahn Institute of Genomics and Multiscale Biology Icahn Medical School at Mount Sinai New
Bayesian Networks as framework for data integration Jun Zhu, Ph. D. Department of Genomics and Genetic Sciences Icahn Institute of Genomics and Multiscale Biology Icahn Medical School at Mount Sinai New
Supplementary materials
 Supplementary materials Calculation of the growth rate for each gene In the growth rate dataset, each gene has many different growth rates under different conditions. The average growth rate for gene i
Supplementary materials Calculation of the growth rate for each gene In the growth rate dataset, each gene has many different growth rates under different conditions. The average growth rate for gene i
Introduction to gene expression microarray data analysis
 Introduction to gene expression microarray data analysis Outline Brief introduction: Technology and data. Statistical challenges in data analysis. Preprocessing data normalization and transformation. Useful
Introduction to gene expression microarray data analysis Outline Brief introduction: Technology and data. Statistical challenges in data analysis. Preprocessing data normalization and transformation. Useful
Machine Learning in Computational Biology CSC 2431
 Machine Learning in Computational Biology CSC 2431 Lecture 9: Combining biological datasets Instructor: Anna Goldenberg What kind of data integration is there? What kind of data integration is there? SNPs
Machine Learning in Computational Biology CSC 2431 Lecture 9: Combining biological datasets Instructor: Anna Goldenberg What kind of data integration is there? What kind of data integration is there? SNPs
Systematic comparison of CRISPR/Cas9 and RNAi screens for essential genes
 CORRECTION NOTICE Nat. Biotechnol. doi:10.1038/nbt. 3567 Systematic comparison of CRISPR/Cas9 and RNAi screens for essential genes David W Morgens, Richard M Deans, Amy Li & Michael C Bassik In the version
CORRECTION NOTICE Nat. Biotechnol. doi:10.1038/nbt. 3567 Systematic comparison of CRISPR/Cas9 and RNAi screens for essential genes David W Morgens, Richard M Deans, Amy Li & Michael C Bassik In the version
Module networks: identifying regulatory modules and their condition-specific regulators from gene expression data
 Module networks: identifying regulatory modules and their condition-specific regulators from gene expression data Eran Segal 1,6, Michael Shapira 2, Aviv Regev 3,5,6, Dana Pe er 4,6, David Botstein 2,
Module networks: identifying regulatory modules and their condition-specific regulators from gene expression data Eran Segal 1,6, Michael Shapira 2, Aviv Regev 3,5,6, Dana Pe er 4,6, David Botstein 2,
On polyclonality of intestinal tumors
 Michael A. University of Wisconsin Chaos and Complex Systems April 2006 Thanks Linda Clipson W.F. Dove Rich Halberg Stephen Stanhope Ruth Sullivan Andrew Thliveris Outline Bio Three statistical questions
Michael A. University of Wisconsin Chaos and Complex Systems April 2006 Thanks Linda Clipson W.F. Dove Rich Halberg Stephen Stanhope Ruth Sullivan Andrew Thliveris Outline Bio Three statistical questions
Analysing the Immune System with Fisher Features
 Analysing the Immune System with John Department of Computer Science University College London WITMSE, Helsinki, September 2016 Experiment β chain CDR3 TCR repertoire sequenced from CD4 spleen cells. unimmunised
Analysing the Immune System with John Department of Computer Science University College London WITMSE, Helsinki, September 2016 Experiment β chain CDR3 TCR repertoire sequenced from CD4 spleen cells. unimmunised
Genomic models in bayz
 Genomic models in bayz Luc Janss, Dec 2010 In the new bayz version the genotype data is now restricted to be 2-allelic markers (SNPs), while the modeling option have been made more general. This implements
Genomic models in bayz Luc Janss, Dec 2010 In the new bayz version the genotype data is now restricted to be 2-allelic markers (SNPs), while the modeling option have been made more general. This implements
A Greedy Algorithm for Minimizing the Number of Primers in Multiple PCR Experiments
 A Greedy Algorithm for Minimizing the Number of Primers in Multiple PCR Experiments Koichiro Doi Hiroshi Imai doi@is.s.u-tokyo.ac.jp imai@is.s.u-tokyo.ac.jp Department of Information Science, Faculty of
A Greedy Algorithm for Minimizing the Number of Primers in Multiple PCR Experiments Koichiro Doi Hiroshi Imai doi@is.s.u-tokyo.ac.jp imai@is.s.u-tokyo.ac.jp Department of Information Science, Faculty of
Introduction to genome biology
 Introduction to genome biology Lisa Stubbs We ve found most genes; but what about the rest of the genome? Genome size* 12 Mb 95 Mb 170 Mb 1500 Mb 2700 Mb 3200 Mb #coding genes ~7000 ~20000 ~14000 ~26000
Introduction to genome biology Lisa Stubbs We ve found most genes; but what about the rest of the genome? Genome size* 12 Mb 95 Mb 170 Mb 1500 Mb 2700 Mb 3200 Mb #coding genes ~7000 ~20000 ~14000 ~26000
Microarray Gene Expression Analysis at CNIO
 Microarray Gene Expression Analysis at CNIO Orlando Domínguez Genomics Unit Biotechnology Program, CNIO 8 May 2013 Workflow, from samples to Gene Expression data Experimental design user/gu/ubio Samples
Microarray Gene Expression Analysis at CNIO Orlando Domínguez Genomics Unit Biotechnology Program, CNIO 8 May 2013 Workflow, from samples to Gene Expression data Experimental design user/gu/ubio Samples
Introduction to Bioinformatics. Fabian Hoti 6.10.
 Introduction to Bioinformatics Fabian Hoti 6.10. Analysis of Microarray Data Introduction Different types of microarrays Experiment Design Data Normalization Feature selection/extraction Clustering Introduction
Introduction to Bioinformatics Fabian Hoti 6.10. Analysis of Microarray Data Introduction Different types of microarrays Experiment Design Data Normalization Feature selection/extraction Clustering Introduction
3. human genomics clone genes associated with genetic disorders. 4. many projects generate ordered clones that cover genome
 Lectures 30 and 31 Genome analysis I. Genome analysis A. two general areas 1. structural 2. functional B. genome projects a status report 1. 1 st sequenced: several viral genomes 2. mitochondria and chloroplasts
Lectures 30 and 31 Genome analysis I. Genome analysis A. two general areas 1. structural 2. functional B. genome projects a status report 1. 1 st sequenced: several viral genomes 2. mitochondria and chloroplasts
EECS730: Introduction to Bioinformatics
 EECS730: Introduction to Bioinformatics Lecture 14: Microarray Some slides were adapted from Dr. Luke Huan (University of Kansas), Dr. Shaojie Zhang (University of Central Florida), and Dr. Dong Xu and
EECS730: Introduction to Bioinformatics Lecture 14: Microarray Some slides were adapted from Dr. Luke Huan (University of Kansas), Dr. Shaojie Zhang (University of Central Florida), and Dr. Dong Xu and
DNA Microarrays and Computational Analysis of DNA Microarray. Data in Cancer Research
 DNA Microarrays and Computational Analysis of DNA Microarray Data in Cancer Research Mario Medvedovic, Jonathan Wiest Abstract 1. Introduction 2. Applications of microarrays 3. Analysis of gene expression
DNA Microarrays and Computational Analysis of DNA Microarray Data in Cancer Research Mario Medvedovic, Jonathan Wiest Abstract 1. Introduction 2. Applications of microarrays 3. Analysis of gene expression
Einführung in die Genetik
 Einführung in die Genetik Prof. Dr. Kay Schneitz (EBio Pflanzen) http://plantdev.bio.wzw.tum.de schneitz@wzw.tum.de Prof. Dr. Claus Schwechheimer (PlaSysBiol) http://wzw.tum.de/sysbiol claus.schwechheimer@wzw.tum.de
Einführung in die Genetik Prof. Dr. Kay Schneitz (EBio Pflanzen) http://plantdev.bio.wzw.tum.de schneitz@wzw.tum.de Prof. Dr. Claus Schwechheimer (PlaSysBiol) http://wzw.tum.de/sysbiol claus.schwechheimer@wzw.tum.de
Characterization of Allele-Specific Copy Number in Tumor Genomes
 Characterization of Allele-Specific Copy Number in Tumor Genomes Hao Chen 2 Haipeng Xing 1 Nancy R. Zhang 2 1 Department of Statistics Stonybrook University of New York 2 Department of Statistics Stanford
Characterization of Allele-Specific Copy Number in Tumor Genomes Hao Chen 2 Haipeng Xing 1 Nancy R. Zhang 2 1 Department of Statistics Stonybrook University of New York 2 Department of Statistics Stanford
2/23/16. Protein-Protein Interactions. Protein Interactions. Protein-Protein Interactions: The Interactome
 Protein-Protein Interactions Protein Interactions A Protein may interact with: Other proteins Nucleic Acids Small molecules Protein-Protein Interactions: The Interactome Experimental methods: Mass Spec,
Protein-Protein Interactions Protein Interactions A Protein may interact with: Other proteins Nucleic Acids Small molecules Protein-Protein Interactions: The Interactome Experimental methods: Mass Spec,
Mapping strategies for sequence reads
 Mapping strategies for sequence reads Ernest Turro University of Cambridge 21 Oct 2013 Quantification A basic aim in genomics is working out the contents of a biological sample. 1. What distinct elements
Mapping strategies for sequence reads Ernest Turro University of Cambridge 21 Oct 2013 Quantification A basic aim in genomics is working out the contents of a biological sample. 1. What distinct elements
MATH 5610, Computational Biology
 MATH 5610, Computational Biology Lecture 2 Intro to Molecular Biology (cont) Stephen Billups University of Colorado at Denver MATH 5610, Computational Biology p.1/24 Announcements Error on syllabus Class
MATH 5610, Computational Biology Lecture 2 Intro to Molecular Biology (cont) Stephen Billups University of Colorado at Denver MATH 5610, Computational Biology p.1/24 Announcements Error on syllabus Class
Analysis of Microarray Data
 Analysis of Microarray Data Lecture 3: Visualization and Functional Analysis George Bell, Ph.D. Senior Bioinformatics Scientist Bioinformatics and Research Computing Whitehead Institute Outline Review
Analysis of Microarray Data Lecture 3: Visualization and Functional Analysis George Bell, Ph.D. Senior Bioinformatics Scientist Bioinformatics and Research Computing Whitehead Institute Outline Review
Optimizing Synthetic DNA for Metabolic Engineering Applications. Howard Salis Penn State University
 Optimizing Synthetic DNA for Metabolic Engineering Applications Howard Salis Penn State University Synthetic Biology Specify a function Build a genetic system (a DNA molecule) Genetic Pseudocode call producequorumsignal(luxi
Optimizing Synthetic DNA for Metabolic Engineering Applications Howard Salis Penn State University Synthetic Biology Specify a function Build a genetic system (a DNA molecule) Genetic Pseudocode call producequorumsignal(luxi
The Next Generation of Transcription Factor Binding Site Prediction
 The Next Generation of Transcription Factor Binding Site Prediction Anthony Mathelier*, Wyeth W. Wasserman* Centre for Molecular Medicine and Therapeutics at the Child and Family Research Institute, Department
The Next Generation of Transcription Factor Binding Site Prediction Anthony Mathelier*, Wyeth W. Wasserman* Centre for Molecular Medicine and Therapeutics at the Child and Family Research Institute, Department
Recent technology allow production of microarrays composed of 70-mers (essentially a hybrid of the two techniques)
 Microarrays and Transcript Profiling Gene expression patterns are traditionally studied using Northern blots (DNA-RNA hybridization assays). This approach involves separation of total or polya + RNA on
Microarrays and Transcript Profiling Gene expression patterns are traditionally studied using Northern blots (DNA-RNA hybridization assays). This approach involves separation of total or polya + RNA on
CS 5984: Application of Basic Clustering Algorithms to Find Expression Modules in Cancer
 CS 5984: Application of Basic Clustering Algorithms to Find Expression Modules in Cancer T. M. Murali January 31, 2006 Innovative Application of Hierarchical Clustering A module map showing conditional
CS 5984: Application of Basic Clustering Algorithms to Find Expression Modules in Cancer T. M. Murali January 31, 2006 Innovative Application of Hierarchical Clustering A module map showing conditional
Chapter 24: Promoters and Enhancers
 Chapter 24: Promoters and Enhancers A typical gene transcribed by RNA polymerase II has a promoter that usually extends upstream from the site where transcription is initiated the (#1) of transcription
Chapter 24: Promoters and Enhancers A typical gene transcribed by RNA polymerase II has a promoter that usually extends upstream from the site where transcription is initiated the (#1) of transcription
 http://genemapping.org/ Epistasis in Association Studies David Evans Law of Independent Assortment Biological Epistasis Bateson (99) a masking effect whereby a variant or allele at one locus prevents
http://genemapping.org/ Epistasis in Association Studies David Evans Law of Independent Assortment Biological Epistasis Bateson (99) a masking effect whereby a variant or allele at one locus prevents
Introduction to Microarray Data Analysis and Gene Networks. Alvis Brazma European Bioinformatics Institute
 Introduction to Microarray Data Analysis and Gene Networks Alvis Brazma European Bioinformatics Institute A brief outline of this course What is gene expression, why it s important Microarrays and how
Introduction to Microarray Data Analysis and Gene Networks Alvis Brazma European Bioinformatics Institute A brief outline of this course What is gene expression, why it s important Microarrays and how
Einführung in die Genetik
 Einführung in die Genetik Prof. Dr. Kay Schneitz (EBio Pflanzen) http://plantdev.bio.wzw.tum.de schneitz@wzw.tum.de Twitter: @PlantDevTUM, #genetiktum FB: Plant Development TUM Prof. Dr. Claus Schwechheimer
Einführung in die Genetik Prof. Dr. Kay Schneitz (EBio Pflanzen) http://plantdev.bio.wzw.tum.de schneitz@wzw.tum.de Twitter: @PlantDevTUM, #genetiktum FB: Plant Development TUM Prof. Dr. Claus Schwechheimer
Bioinformatics of Transcriptional Regulation
 Bioinformatics of Transcriptional Regulation Carl Herrmann IPMB & DKFZ c.herrmann@dkfz.de Wechselwirkung von Maßnahmen und Auswirkungen Einflussmöglichkeiten in einem Dialog From genes to active compounds
Bioinformatics of Transcriptional Regulation Carl Herrmann IPMB & DKFZ c.herrmann@dkfz.de Wechselwirkung von Maßnahmen und Auswirkungen Einflussmöglichkeiten in einem Dialog From genes to active compounds
CHAPTER 21 LECTURE SLIDES
 CHAPTER 21 LECTURE SLIDES Prepared by Brenda Leady University of Toledo To run the animations you must be in Slideshow View. Use the buttons on the animation to play, pause, and turn audio/text on or off.
CHAPTER 21 LECTURE SLIDES Prepared by Brenda Leady University of Toledo To run the animations you must be in Slideshow View. Use the buttons on the animation to play, pause, and turn audio/text on or off.
Improving the Accuracy of Base Calls and Error Predictions for GS 20 DNA Sequence Data
 Improving the Accuracy of Base Calls and Error Predictions for GS 20 DNA Sequence Data Justin S. Hogg Department of Computational Biology University of Pittsburgh Pittsburgh, PA 15213 jsh32@pitt.edu Abstract
Improving the Accuracy of Base Calls and Error Predictions for GS 20 DNA Sequence Data Justin S. Hogg Department of Computational Biology University of Pittsburgh Pittsburgh, PA 15213 jsh32@pitt.edu Abstract
WebMOTIFS: Automated discovery, filtering, and scoring of DNA sequence motifs using multiple programs and Bayesian approaches
 WebMOTIFS: Automated discovery, filtering, and scoring of DNA sequence motifs using multiple programs and Bayesian approaches Katherine A. Romer 1, Guy-Richard Kayombya 1, Ernest Fraenkel 2,3 1 Department
WebMOTIFS: Automated discovery, filtering, and scoring of DNA sequence motifs using multiple programs and Bayesian approaches Katherine A. Romer 1, Guy-Richard Kayombya 1, Ernest Fraenkel 2,3 1 Department
Meta-analysis discovery of. tissue-specific DNA sequence motifs. from mammalian gene expression data
 Meta-analysis discovery of tissue-specific DNA sequence motifs from mammalian gene expression data Bertrand R. Huber 1,3 and Martha L. Bulyk 1,2,3 1 Division of Genetics, Department of Medicine, 2 Department
Meta-analysis discovery of tissue-specific DNA sequence motifs from mammalian gene expression data Bertrand R. Huber 1,3 and Martha L. Bulyk 1,2,3 1 Division of Genetics, Department of Medicine, 2 Department
Protein-Protein-Interaction Networks. Ulf Leser, Samira Jaeger
 Protein-Protein-Interaction Networks Ulf Leser, Samira Jaeger SHK Stelle frei Ab 1.9.2015, 2 Jahre, 41h/Monat Verbundprojekt MaptTorNet: Pankreatische endokrine Tumore Insb. statistische Aufbereitung und
Protein-Protein-Interaction Networks Ulf Leser, Samira Jaeger SHK Stelle frei Ab 1.9.2015, 2 Jahre, 41h/Monat Verbundprojekt MaptTorNet: Pankreatische endokrine Tumore Insb. statistische Aufbereitung und
Microarray Technique. Some background. M. Nath
 Microarray Technique Some background M. Nath Outline Introduction Spotting Array Technique GeneChip Technique Data analysis Applications Conclusion Now Blind Guess? Functional Pathway Microarray Technique
Microarray Technique Some background M. Nath Outline Introduction Spotting Array Technique GeneChip Technique Data analysis Applications Conclusion Now Blind Guess? Functional Pathway Microarray Technique
Gene Expression and Heritable Phenotype. CBS520 Eric Nabity
 Gene Expression and Heritable Phenotype CBS520 Eric Nabity DNA is Just the Beginning DNA was determined to be the genetic material, and the structure was identified as a (double stranded) double helix.
Gene Expression and Heritable Phenotype CBS520 Eric Nabity DNA is Just the Beginning DNA was determined to be the genetic material, and the structure was identified as a (double stranded) double helix.
Generative Models for Networks and Applications to E-Commerce
 Generative Models for Networks and Applications to E-Commerce Patrick J. Wolfe (with David C. Parkes and R. Kang-Xing Jin) Division of Engineering and Applied Sciences Department of Statistics Harvard
Generative Models for Networks and Applications to E-Commerce Patrick J. Wolfe (with David C. Parkes and R. Kang-Xing Jin) Division of Engineering and Applied Sciences Department of Statistics Harvard
7 Gene Isolation and Analysis of Multiple
 Genetic Techniques for Biological Research Corinne A. Michels Copyright q 2002 John Wiley & Sons, Ltd ISBNs: 0-471-89921-6 (Hardback); 0-470-84662-3 (Electronic) 7 Gene Isolation and Analysis of Multiple
Genetic Techniques for Biological Research Corinne A. Michels Copyright q 2002 John Wiley & Sons, Ltd ISBNs: 0-471-89921-6 (Hardback); 0-470-84662-3 (Electronic) 7 Gene Isolation and Analysis of Multiple
Exploration and Analysis of DNA Microarray Data
 Exploration and Analysis of DNA Microarray Data Dhammika Amaratunga Senior Research Fellow in Nonclinical Biostatistics Johnson & Johnson Pharmaceutical Research & Development Javier Cabrera Associate
Exploration and Analysis of DNA Microarray Data Dhammika Amaratunga Senior Research Fellow in Nonclinical Biostatistics Johnson & Johnson Pharmaceutical Research & Development Javier Cabrera Associate
DNA Microarray Data Oligonucleotide Arrays
 DNA Microarray Data Oligonucleotide Arrays Sandrine Dudoit, Robert Gentleman, Rafael Irizarry, and Yee Hwa Yang Bioconductor Short Course 2003 Copyright 2002, all rights reserved Biological question Experimental
DNA Microarray Data Oligonucleotide Arrays Sandrine Dudoit, Robert Gentleman, Rafael Irizarry, and Yee Hwa Yang Bioconductor Short Course 2003 Copyright 2002, all rights reserved Biological question Experimental
Le proteine regolative variano nei vari tipi cellulari e in funzione degli stimoli ambientali
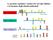 Le proteine regolative variano nei vari tipi cellulari e in funzione degli stimoli ambientali Tipo cellulare 1 Tipo cellulare 2 Tipo cellulare 3 DNA-protein Crosslink Lisi Frammentazione Immunopurificazione
Le proteine regolative variano nei vari tipi cellulari e in funzione degli stimoli ambientali Tipo cellulare 1 Tipo cellulare 2 Tipo cellulare 3 DNA-protein Crosslink Lisi Frammentazione Immunopurificazione
SIMS2003. Instructors:Rus Yukhananov, Alex Loguinov BWH, Harvard Medical School. Introduction to Microarray Technology.
 SIMS2003 Instructors:Rus Yukhananov, Alex Loguinov BWH, Harvard Medical School Introduction to Microarray Technology. Lecture 1 I. EXPERIMENTAL DETAILS II. ARRAY CONSTRUCTION III. IMAGE ANALYSIS Lecture
SIMS2003 Instructors:Rus Yukhananov, Alex Loguinov BWH, Harvard Medical School Introduction to Microarray Technology. Lecture 1 I. EXPERIMENTAL DETAILS II. ARRAY CONSTRUCTION III. IMAGE ANALYSIS Lecture
Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017
 Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017 Agenda What is Functional Genomics? RNA Transcription/Gene Expression Measuring Gene
Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017 Agenda What is Functional Genomics? RNA Transcription/Gene Expression Measuring Gene
3.1.4 DNA Microarray Technology
 3.1.4 DNA Microarray Technology Scientists have discovered that one of the differences between healthy and cancer is which genes are turned on in each. Scientists can compare the gene expression patterns
3.1.4 DNA Microarray Technology Scientists have discovered that one of the differences between healthy and cancer is which genes are turned on in each. Scientists can compare the gene expression patterns
DNA Microarray Technology
 CHAPTER 1 DNA Microarray Technology All living organisms are composed of cells. As a functional unit, each cell can make copies of itself, and this process depends on a proper replication of the genetic
CHAPTER 1 DNA Microarray Technology All living organisms are composed of cells. As a functional unit, each cell can make copies of itself, and this process depends on a proper replication of the genetic
ALGORITHMS IN BIO INFORMATICS. Chapman & Hall/CRC Mathematical and Computational Biology Series A PRACTICAL INTRODUCTION. CRC Press WING-KIN SUNG
 Chapman & Hall/CRC Mathematical and Computational Biology Series ALGORITHMS IN BIO INFORMATICS A PRACTICAL INTRODUCTION WING-KIN SUNG CRC Press Taylor & Francis Group Boca Raton London New York CRC Press
Chapman & Hall/CRC Mathematical and Computational Biology Series ALGORITHMS IN BIO INFORMATICS A PRACTICAL INTRODUCTION WING-KIN SUNG CRC Press Taylor & Francis Group Boca Raton London New York CRC Press
Transcription Gene regulation
 Transcription Gene regulation The machine that transcribes a gene is composed of perhaps 50 proteins, including RNA polymerase, the enzyme that converts DNA code into RNA code. A crew of transcription
Transcription Gene regulation The machine that transcribes a gene is composed of perhaps 50 proteins, including RNA polymerase, the enzyme that converts DNA code into RNA code. A crew of transcription
Calculation of Spot Reliability Evaluation Scores (SRED) for DNA Microarray Data
 Protocol Calculation of Spot Reliability Evaluation Scores (SRED) for DNA Microarray Data Kazuro Shimokawa, Rimantas Kodzius, Yonehiro Matsumura, and Yoshihide Hayashizaki This protocol was adapted from
Protocol Calculation of Spot Reliability Evaluation Scores (SRED) for DNA Microarray Data Kazuro Shimokawa, Rimantas Kodzius, Yonehiro Matsumura, and Yoshihide Hayashizaki This protocol was adapted from
Roche Molecular Biochemicals Technical Note No. LC 10/2000
 Roche Molecular Biochemicals Technical Note No. LC 10/2000 LightCycler Overview of LightCycler Quantification Methods 1. General Introduction Introduction Content Definitions This Technical Note will introduce
Roche Molecular Biochemicals Technical Note No. LC 10/2000 LightCycler Overview of LightCycler Quantification Methods 1. General Introduction Introduction Content Definitions This Technical Note will introduce
computational analysis of cell-to-cell heterogeneity in single-cell rna-sequencing data reveals hidden subpopulations of cells
 computational analysis of cell-to-cell heterogeneity in single-cell rna-sequencing data reveals hidden subpopulations of cells Buettner et al., (2015) Nature Biotechnology, 1 32. doi:10.1038/nbt.3102 Saket
computational analysis of cell-to-cell heterogeneity in single-cell rna-sequencing data reveals hidden subpopulations of cells Buettner et al., (2015) Nature Biotechnology, 1 32. doi:10.1038/nbt.3102 Saket
Green Fluorescent Protein (GFP) Purification. Hydrophobic Interaction Chromatography
 Green Fluorescent Protein (GFP) Purification Hydrophobic Interaction Chromatography What is the GFP gene? GFP is a green fluorescent protein that is normally found in jellyfish. It has been engineered
Green Fluorescent Protein (GFP) Purification Hydrophobic Interaction Chromatography What is the GFP gene? GFP is a green fluorescent protein that is normally found in jellyfish. It has been engineered
Gene Expression Data Analysis (I)
 Gene Expression Data Analysis (I) Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Bioinformatics tasks Biological question Experiment design Microarray experiment
Gene Expression Data Analysis (I) Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Bioinformatics tasks Biological question Experiment design Microarray experiment
Decoding Chromatin States with Epigenome Data Advanced Topics in Computa8onal Genomics
 Decoding Chromatin States with Epigenome Data 02-715 Advanced Topics in Computa8onal Genomics HMMs for Decoding Chromatin States Epigene8c modifica8ons of the genome have been associated with Establishing
Decoding Chromatin States with Epigenome Data 02-715 Advanced Topics in Computa8onal Genomics HMMs for Decoding Chromatin States Epigene8c modifica8ons of the genome have been associated with Establishing
Ana Teresa Freitas 2016/2017
 Finding Regulatory Motifs in DNA Sequences Ana Teresa Freitas 2016/2017 Combinatorial Gene Regulation A recent microarray experiment showed that when gene X is knocked out, 20 other genes are not expressed
Finding Regulatory Motifs in DNA Sequences Ana Teresa Freitas 2016/2017 Combinatorial Gene Regulation A recent microarray experiment showed that when gene X is knocked out, 20 other genes are not expressed
Offshoring and the Functional Structure of Labour Demand in Advanced Economies
 Offshoring and the Functional Structure of Labour Demand in Advanced Economies A. Jiang, S. Miroudot, G. J. De Vries Discussant: Catia Montagna Motivation Due to declining communication and coordination
Offshoring and the Functional Structure of Labour Demand in Advanced Economies A. Jiang, S. Miroudot, G. J. De Vries Discussant: Catia Montagna Motivation Due to declining communication and coordination
Methods of Biomaterials Testing Lesson 3-5. Biochemical Methods - Molecular Biology -
 Methods of Biomaterials Testing Lesson 3-5 Biochemical Methods - Molecular Biology - Chromosomes in the Cell Nucleus DNA in the Chromosome Deoxyribonucleic Acid (DNA) DNA has double-helix structure The
Methods of Biomaterials Testing Lesson 3-5 Biochemical Methods - Molecular Biology - Chromosomes in the Cell Nucleus DNA in the Chromosome Deoxyribonucleic Acid (DNA) DNA has double-helix structure The
Name_BS50 Exam 3 Key (Fall 2005) Page 2 of 5
 Name_BS50 Exam 3 Key (Fall 2005) Page 2 of 5 Question 1. (14 points) Several Hfr strains derived from the same F + strain were crossed separately to an F - strain, giving the results indicated in the table
Name_BS50 Exam 3 Key (Fall 2005) Page 2 of 5 Question 1. (14 points) Several Hfr strains derived from the same F + strain were crossed separately to an F - strain, giving the results indicated in the table
V 1 Introduction! Fri, Oct 24, 2014! Bioinformatics 3 Volkhard Helms!
 V 1 Introduction! Fri, Oct 24, 2014! Bioinformatics 3 Volkhard Helms! How Does a Cell Work?! A cell is a crowded environment! => many different proteins,! metabolites, compartments,! On a microscopic level!
V 1 Introduction! Fri, Oct 24, 2014! Bioinformatics 3 Volkhard Helms! How Does a Cell Work?! A cell is a crowded environment! => many different proteins,! metabolites, compartments,! On a microscopic level!
Transcription factor binding site identification using the Self-Organizing Map
 Bioinformatics Advance Access published January 12, 2005 Bioinfor matics Oxford University Press 2005; all rights reserved. Transcription factor binding site identification using the Self-Organizing Map
Bioinformatics Advance Access published January 12, 2005 Bioinfor matics Oxford University Press 2005; all rights reserved. Transcription factor binding site identification using the Self-Organizing Map
Supporting Information
 Supporting Information Ho et al. 1.173/pnas.81288816 SI Methods Sequences of shrna hairpins: Brg shrna #1: ccggcggctcaagaaggaagttgaactcgagttcaacttccttcttgacgnttttg (TRCN71383; Open Biosystems). Brg shrna
Supporting Information Ho et al. 1.173/pnas.81288816 SI Methods Sequences of shrna hairpins: Brg shrna #1: ccggcggctcaagaaggaagttgaactcgagttcaacttccttcttgacgnttttg (TRCN71383; Open Biosystems). Brg shrna
less sensitive than RNA-seq but more robust analysis pipelines expensive but quantitiatve standard but typically not high throughput
 Chapter 11: Gene Expression The availability of an annotated genome sequence enables massively parallel analysis of gene expression. The expression of all genes in an organism can be measured in one experiment.
Chapter 11: Gene Expression The availability of an annotated genome sequence enables massively parallel analysis of gene expression. The expression of all genes in an organism can be measured in one experiment.
Procedia - Social and Behavioral Sciences 97 ( 2013 )
 Available online at www.sciencedirect.com ScienceDirect Procedia - Social and Behavioral Sciences 97 ( 2013 ) 602 611 Abstract The 9 th International Conference on Cognitive Science Filtering of background
Available online at www.sciencedirect.com ScienceDirect Procedia - Social and Behavioral Sciences 97 ( 2013 ) 602 611 Abstract The 9 th International Conference on Cognitive Science Filtering of background
Zool 3200: Cell Biology Exam 3 3/6/15
 Name: Trask Zool 3200: Cell Biology Exam 3 3/6/15 Answer each of the following questions in the space provided; circle the correct answer or answers for each multiple choice question and circle either
Name: Trask Zool 3200: Cell Biology Exam 3 3/6/15 Answer each of the following questions in the space provided; circle the correct answer or answers for each multiple choice question and circle either
ALSO: look at figure 5-11 showing exonintron structure of the beta globin gene
 S08 Biology 205 6/4/08 Reading Assignment Chapter 7: From DNA to Protein: How cells read the genome pg 237-243 on exons and introns (you are not responsible for the biochemistry of splicing: figures 7-15,16
S08 Biology 205 6/4/08 Reading Assignment Chapter 7: From DNA to Protein: How cells read the genome pg 237-243 on exons and introns (you are not responsible for the biochemistry of splicing: figures 7-15,16
Quantitative Real Time PCR USING SYBR GREEN
 Quantitative Real Time PCR USING SYBR GREEN SYBR Green SYBR Green is a cyanine dye that binds to double stranded DNA. When it is bound to D.S. DNA it has a much greater fluorescence than when bound to
Quantitative Real Time PCR USING SYBR GREEN SYBR Green SYBR Green is a cyanine dye that binds to double stranded DNA. When it is bound to D.S. DNA it has a much greater fluorescence than when bound to
Predicting Microarray Signals by Physical Modeling. Josh Deutsch. University of California. Santa Cruz
 Predicting Microarray Signals by Physical Modeling Josh Deutsch University of California Santa Cruz Predicting Microarray Signals by Physical Modeling p.1/39 Collaborators Shoudan Liang NASA Ames Onuttom
Predicting Microarray Signals by Physical Modeling Josh Deutsch University of California Santa Cruz Predicting Microarray Signals by Physical Modeling p.1/39 Collaborators Shoudan Liang NASA Ames Onuttom
Transcription factor binding site prediction in vivo using DNA sequence and shape features
 Transcription factor binding site prediction in vivo using DNA sequence and shape features Anthony Mathelier, Lin Yang, Tsu-Pei Chiu, Remo Rohs, and Wyeth Wasserman anthony.mathelier@gmail.com @AMathelier
Transcription factor binding site prediction in vivo using DNA sequence and shape features Anthony Mathelier, Lin Yang, Tsu-Pei Chiu, Remo Rohs, and Wyeth Wasserman anthony.mathelier@gmail.com @AMathelier
Combination of Neuro-Fuzzy Network Models with Biological Knowledge for Reconstructing Gene Regulatory Networks
 Journal of Bionic Engineering 8 (2011) 98 106 Combination of Neuro-Fuzzy Network Models with Biological Knowledge for Reconstructing Gene Regulatory Networks Guixia Liu 1, Lei Liu 1, Chunyu Liu 2, Ming
Journal of Bionic Engineering 8 (2011) 98 106 Combination of Neuro-Fuzzy Network Models with Biological Knowledge for Reconstructing Gene Regulatory Networks Guixia Liu 1, Lei Liu 1, Chunyu Liu 2, Ming
Reliable classification of two-class cancer data using evolutionary algorithms
 BioSystems 72 (23) 111 129 Reliable classification of two-class cancer data using evolutionary algorithms Kalyanmoy Deb, A. Raji Reddy Kanpur Genetic Algorithms Laboratory (KanGAL), Indian Institute of
BioSystems 72 (23) 111 129 Reliable classification of two-class cancer data using evolutionary algorithms Kalyanmoy Deb, A. Raji Reddy Kanpur Genetic Algorithms Laboratory (KanGAL), Indian Institute of
Lecture 10: Motif Finding Regulatory element detection using correlation with expression
 CS5238 Combinatorial methods in bioinformatics 2006/2007 Semester 1 Lecture 10: Motif Finding Lecturer: Wing-Kin Sung Scribe: Zhang Jingbo, Shrikant Kashyap 10.1 Regulatory element detection using correlation
CS5238 Combinatorial methods in bioinformatics 2006/2007 Semester 1 Lecture 10: Motif Finding Lecturer: Wing-Kin Sung Scribe: Zhang Jingbo, Shrikant Kashyap 10.1 Regulatory element detection using correlation
Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions. Supplementary Material
 Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions Joshua N. Burton 1, Andrew Adey 1, Rupali P. Patwardhan 1, Ruolan Qiu 1, Jacob O. Kitzman 1, Jay Shendure 1 1 Department
Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions Joshua N. Burton 1, Andrew Adey 1, Rupali P. Patwardhan 1, Ruolan Qiu 1, Jacob O. Kitzman 1, Jay Shendure 1 1 Department
DNA Transcription. Dr Aliwaini
 DNA Transcription 1 DNA Transcription-Introduction The synthesis of an RNA molecule from DNA is called Transcription. All eukaryotic cells have five major classes of RNA: ribosomal RNA (rrna), messenger
DNA Transcription 1 DNA Transcription-Introduction The synthesis of an RNA molecule from DNA is called Transcription. All eukaryotic cells have five major classes of RNA: ribosomal RNA (rrna), messenger
pint: probabilistic data integration for functional genomics
 pint: probabilistic data integration for functional genomics Olli-Pekka Huovilainen 1* and Leo Lahti 1,2 (1) Dpt. Information and Computer Science, Aalto University, Finland (2) Dpt. Veterinary Bioscience,
pint: probabilistic data integration for functional genomics Olli-Pekka Huovilainen 1* and Leo Lahti 1,2 (1) Dpt. Information and Computer Science, Aalto University, Finland (2) Dpt. Veterinary Bioscience,
Introduction to Molecular Biology
 Introduction to Molecular Biology Bioinformatics: Issues and Algorithms CSE 308-408 Fall 2007 Lecture 2-1- Important points to remember We will study: Problems from bioinformatics. Algorithms used to solve
Introduction to Molecular Biology Bioinformatics: Issues and Algorithms CSE 308-408 Fall 2007 Lecture 2-1- Important points to remember We will study: Problems from bioinformatics. Algorithms used to solve
CAP 5510: Introduction to Bioinformatics CGS 5166: Bioinformatics Tools. Giri Narasimhan
 CAP 5510: Introduction to Bioinformatics CGS 5166: Bioinformatics Tools Giri Narasimhan ECS 254; Phone: x3748 giri@cis.fiu.edu www.cis.fiu.edu/~giri/teach/bioinfs15.html Gene Expression q Process of transcription
CAP 5510: Introduction to Bioinformatics CGS 5166: Bioinformatics Tools Giri Narasimhan ECS 254; Phone: x3748 giri@cis.fiu.edu www.cis.fiu.edu/~giri/teach/bioinfs15.html Gene Expression q Process of transcription
Chapter 8 Lecture Outline. Transcription, Translation, and Bioinformatics
 Chapter 8 Lecture Outline Transcription, Translation, and Bioinformatics Replication, Transcription, Translation n Repetitive processes Build polymers of nucleotides or amino acids n All have 3 major steps
Chapter 8 Lecture Outline Transcription, Translation, and Bioinformatics Replication, Transcription, Translation n Repetitive processes Build polymers of nucleotides or amino acids n All have 3 major steps
Exploring Similarities of Conserved Domains/Motifs
 Exploring Similarities of Conserved Domains/Motifs Sotiria Palioura Abstract Traditionally, proteins are represented as amino acid sequences. There are, though, other (potentially more exciting) representations;
Exploring Similarities of Conserved Domains/Motifs Sotiria Palioura Abstract Traditionally, proteins are represented as amino acid sequences. There are, though, other (potentially more exciting) representations;
Gene Regulatory Network Reconstruction Using Dynamic Bayesian Networks
 The University of Southern Mississippi The Aquila Digital Community Dissertations Spring 5-2013 Gene Regulatory Network Reconstruction Using Dynamic Bayesian Networks Haoni Li University of Southern Mississippi
The University of Southern Mississippi The Aquila Digital Community Dissertations Spring 5-2013 Gene Regulatory Network Reconstruction Using Dynamic Bayesian Networks Haoni Li University of Southern Mississippi
Functional Bioinformatics of Microarray Data: From Expression to Regulation
 Functional Bioinformatics of Microarray Data: From Expression to Regulation YVES MOREAU, FRANK DE SMET, GERT THIJS, STUDENT MEMBER, IEEE, KATHLEEN MARCHAL, AND BART DE MOOR, SENIOR MEMBER, IEEE Invited
Functional Bioinformatics of Microarray Data: From Expression to Regulation YVES MOREAU, FRANK DE SMET, GERT THIJS, STUDENT MEMBER, IEEE, KATHLEEN MARCHAL, AND BART DE MOOR, SENIOR MEMBER, IEEE Invited
Creation of a PAM matrix
 Rationale for substitution matrices Substitution matrices are a way of keeping track of the structural, physical and chemical properties of the amino acids in proteins, in such a fashion that less detrimental
Rationale for substitution matrices Substitution matrices are a way of keeping track of the structural, physical and chemical properties of the amino acids in proteins, in such a fashion that less detrimental
Multiple Testing in RNA-Seq experiments
 Multiple Testing in RNA-Seq experiments O. Muralidharan et al. 2012. Detecting mutations in mixed sample sequencing data using empirical Bayes. Bernd Klaus Institut für Medizinische Informatik, Statistik
Multiple Testing in RNA-Seq experiments O. Muralidharan et al. 2012. Detecting mutations in mixed sample sequencing data using empirical Bayes. Bernd Klaus Institut für Medizinische Informatik, Statistik
Global analysis of gene transcription regulation in prokaryotes
 Cell. Mol. Life Sci. DOI 10.1007/s00018-006-6184-6 Birkhäuser Verlag, Basel, 2006 Cellular and Molecular Life Sciences Review Global analysis of gene transcription regulation in prokaryotes D. Zhou* and
Cell. Mol. Life Sci. DOI 10.1007/s00018-006-6184-6 Birkhäuser Verlag, Basel, 2006 Cellular and Molecular Life Sciences Review Global analysis of gene transcription regulation in prokaryotes D. Zhou* and
Lecture 11: Gene Prediction
 Lecture 11: Gene Prediction Study Chapter 6.11-6.14 1 Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and end positions of genes in a genome Where are
Lecture 11: Gene Prediction Study Chapter 6.11-6.14 1 Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and end positions of genes in a genome Where are
DO NOT OPEN UNTIL TOLD TO START
 DO NOT OPEN UNTIL TOLD TO START BIO 312, Section 1: Fall 2012 December 4 th, 2012 Exam 3 Name (print neatly) Signature 7 digit student ID INSTRUCTIONS: 1. There are 12 pages to the exam. Make sure you
DO NOT OPEN UNTIL TOLD TO START BIO 312, Section 1: Fall 2012 December 4 th, 2012 Exam 3 Name (print neatly) Signature 7 digit student ID INSTRUCTIONS: 1. There are 12 pages to the exam. Make sure you
Designing Complex Omics Experiments
 Designing Complex Omics Experiments Xiangqin Cui Section on Statistical Genetics Department of Biostatistics University of Alabama at Birmingham 6/15/2015 Some slides are from previous lectures given by
Designing Complex Omics Experiments Xiangqin Cui Section on Statistical Genetics Department of Biostatistics University of Alabama at Birmingham 6/15/2015 Some slides are from previous lectures given by
Year III Pharm.D Dr. V. Chitra
 Year III Pharm.D Dr. V. Chitra 1 Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Only one strand of DNA serves
Year III Pharm.D Dr. V. Chitra 1 Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Only one strand of DNA serves
Discovery of Transcription Factor Binding Sites with Deep Convolutional Neural Networks
 Discovery of Transcription Factor Binding Sites with Deep Convolutional Neural Networks Reesab Pathak Dept. of Computer Science Stanford University rpathak@stanford.edu Abstract Transcription factors are
Discovery of Transcription Factor Binding Sites with Deep Convolutional Neural Networks Reesab Pathak Dept. of Computer Science Stanford University rpathak@stanford.edu Abstract Transcription factors are
Transcription in Eukaryotes
 Transcription in Eukaryotes Biology I Hayder A Giha Transcription Transcription is a DNA-directed synthesis of RNA, which is the first step in gene expression. Gene expression, is transformation of the
Transcription in Eukaryotes Biology I Hayder A Giha Transcription Transcription is a DNA-directed synthesis of RNA, which is the first step in gene expression. Gene expression, is transformation of the
Preprocessing Affymetrix GeneChip Data. Affymetrix GeneChip Design. Terminology TGTGATGGTGGGGAATGGGTCAGAAGGCCTCCGATGCGCCGATTGAGAAT
 Preprocessing Affymetrix GeneChip Data Credit for some of today s materials: Ben Bolstad, Leslie Cope, Laurent Gautier, Terry Speed and Zhijin Wu Affymetrix GeneChip Design 5 3 Reference sequence TGTGATGGTGGGGAATGGGTCAGAAGGCCTCCGATGCGCCGATTGAGAAT
Preprocessing Affymetrix GeneChip Data Credit for some of today s materials: Ben Bolstad, Leslie Cope, Laurent Gautier, Terry Speed and Zhijin Wu Affymetrix GeneChip Design 5 3 Reference sequence TGTGATGGTGGGGAATGGGTCAGAAGGCCTCCGATGCGCCGATTGAGAAT
Technical tips Session 5
 Technical tips Session 5 Chromatine Immunoprecipitation (ChIP): This is a powerful in vivo method to quantitate interaction of proteins associated with specific regions of the genome. It involves the immunoprecipitation
Technical tips Session 5 Chromatine Immunoprecipitation (ChIP): This is a powerful in vivo method to quantitate interaction of proteins associated with specific regions of the genome. It involves the immunoprecipitation
DNA Microarrays Introduction Part 2. Todd Lowe BME/BIO 210 April 11, 2007
 DNA Microarrays Introduction Part 2 Todd Lowe BME/BIO 210 April 11, 2007 Reading Assigned For Friday, please read two papers and be prepared to discuss in detail: Comprehensive Identification of Cell Cycle-related
DNA Microarrays Introduction Part 2 Todd Lowe BME/BIO 210 April 11, 2007 Reading Assigned For Friday, please read two papers and be prepared to discuss in detail: Comprehensive Identification of Cell Cycle-related
