Proteomics Standards Research Group
|
|
|
- Arron Boone
- 6 years ago
- Views:
Transcription
1 A B R F Proteomics Standards Research Group Association of Biomolecular Resource Facilities Proteomics Standards Research Group (sprg)
2 Phosphorylation of Proteins Important post-translational modification Major role in regulation of cellular processes. Analysis of phosphorylation sites is a major challenge for proteomics labs. Many techniques and approaches available. What works best?
3 PRG 2003 Study J Biomolecular Tech 14, , digested proteins (5 pmole and 0.2 fmole) 2 synthetic phospho-peptides (1 pmole each) Results: 54 Data sets returned and 67 Analyses; 8 Labs Identified 1 Pi-peptide; 8 Labs Identified the other Pi-peptide; IMAC users did not fare better.
4 sprg 2007 Study Mixture of Proteins Catalase - 10 pmol Troponin T - 25 pmol Osteopontin - 50 pmol Ovalbumin - 50 pmol Cystatin pmol α-s1-casein pmol α-s2-casein pmol Which phosphorylated residues were found? Which techniques were most successful?
5 sprg 2007 Study Some known sites were detected by >50% of labs. Other known sites were identified by few or no labs. Enrichment methods did not make a big difference. Lack of statistical/spectral methods for identifying Pi-peptides. Examination of spectra underlying reported identifications indicated that identifications were not always made based on clear-cut spectral evidence. This study demonstrates the difficulty confronting the creation of a standard mix of phosphoproteins.
6 sprg 2010 Study Goal Goal: A readily available phospho-peptide standard useful for: Learning Pi-peptide detection before running that once in a lifetime research sample Identifying Phosphorylation Sites Evaluating Current and New techniques Developing Your Own Techniques Benchmarking Goal: What are the best practices being used?
7 sprg 2010 Study Design 23 synthetic Pi-peptides Each present at 5 pmole Based on sprg 2009 Study Included 2 peptides from PRG 2003 Phosphorylations sites: Ser, Thr and Tyr Single Pi: 14 Double Pi: 5 Triple Pi: 3 Quadruple Pi: 1 Positional isomers. Digest of six proteins Each present at 5 pmole Recombinant human or bovine HPLC purified Each protein was digested separately with Trypsin Naturally occurring phosphopeptides present in the background digests
8 6 Proteins Selected as Digest Background Tryptic Peptides Selected from the 6 Proteins Each Protein Was Reduced, Alkylated, Digested & Tested Phosphorylated Versions of Select Tryptic Peptides Were Synthesized, Purified, Quantified and Tested Digested Proteins and Pi-Peptides Were Combined, Dried, Aliquotted & Tested (5 pmol Each) sprg 2010 Sample Mailed to Requesters Participants Analyzed the Sample and Returned Results by SurveyMonkey The sprg Analyzed the Results and Presents to Captive Audience (You Are Here!)
9 Who Volunteered?
10 Please indicate your ABRF membership status ABRF Member 1 ABRF Corporate Sponsor Non-member
11 Which best describes your geographic location? Asia Africa Europe North America
12 Austria China Denmark Germany Ireland Israel Italy no response Singapore South Africa South Korea Spain Switzerland The Netherlands United States 18 Please indicate your country
13 Academic Biotech/Pharma Government Manufacturer Other Which best describes your lab?
14 Core or resource facility status? 12 Core/Resource Facility only 18 Non-core research lab Conduct both core functions and non-core lab research 9
15 How long have you been involved in proteomics? < 1 year 1-2 years 3-4 years 5-10 years >10 years
16 What do you consider your level of experience in proteomics? Novice Experienced Expert
17 How long have you been involved in phosphoproteomics? < 1 year 1-2 years 3-4 years 5-10 years >10 years 1
18 30 What do you consider your level of experience in phosphoproteomics? Novice Experienced Expert
19 Overview of Results
20 Identification Rates By Peptide sprg2010 PRG 2003 Protein MW, Da Sequence # of times ID % ID # of times ID % ID ALBU_HUMAN ALBU_HUMAN AEFAEV#SK (1) LVNEV#TEFAK (1) 34 81% 40 95% NQO2_HUMAN PRDX1_HUMAN PRDX1_HUMAN PRDX1_HUMAN PRDX1_HUMAN PRDX1_HUMAN NVAVDEL#SR (1) ADEGI#SFR (1) DISLSD#YK (1) DI#SLSD#YK (2) DISL#SD#YK (2) DI#SL#SD#YK (3) 35 83% 39 93% 34 81% 25 60% 20 48% 10 24% PRDX1_HUMAN SYHC_HUMAN LVQAFQF#TDK (1) DQGGELL#SLR (1) 40 95% 38 90% SYHC_HUMAN UBIQ_HUMAN UBIQ_HUMAN UBIQ_HUMAN UBIQ_HUMAN UBIQ_HUMAN UBIQ_HUMAN UBIQ_HUMAN UBIQ_HUMAN UBIQ_HUMAN UBIQ_HUMAN PDIA1_BOVIN PDIA1_BOVIN IF#SIVEQR (1) ES#TLHLVLR (1) E#S#TLHLVLR (2) E#STLHLVLR (1) TLSD#YNIQK (1) #TLSDYNIQK (1) #TL#SDYNIQK (2) TITLEVEP#SD#TIENVK (2) #TITLEVEP#SD#TIENVK (3) #TI#TLEVEP#SD#TIENVK (4) #TL#SD#YNIQK (3) SV#SDYEGK (1) THILLFLPK#SVSDYEGK (1) 38 90% 25 60% 15 36% 23 55% 31 74% 31 74% 25 60% 26 62% 10 24% 9 21% 16 38% 17 40% 26 62% 8 15% 8 15%
21 Sequence Identification Rates By Peptide # of p sites AEFAEVpSK (ALBU) % LVNEVpTEFAK (ALBU) % NVAVDELpSR (NQO2) % ADEGIpSFR (PRDX1) % DISLSDpYK (PRDX1) % DIpSLSDpYK (PRDX1) % DISLpSDpYK (PRDX1) % DIpSLpSDpYK (PRDX1) % LVQAFQFpTDK (PRDX1) % DQGGELLpSLR (SYHC) % IFpSIVEQR (SYHC) % # of times ID % ID Sequence # of p sites # of times ID % ID ESpTLHLVLR (UBIQ) % EpSpTLHLVLR (UBIQ) % EpSTLHLVLR (UBIQ) % TLSDpYNIQK (UBIQ) % ptlsdyniqk (UBIQ) % ptlpsdyniqk (UBIQ) % TITLEVEPpSDpTIENVK (UBIQ) % ptitleveppsdptienvk (UBIQ) % ptiptleveppsdptienvk (UBIQ) % ptlpsdpyniqk (UBIQ) % SVpSDYEGK (PDIA1) % THILLFLPKpSVSDYEGK (PDIA1) %
22 % ID rate of each phospeptide sequence 100% 90% 80% 81% 95% 83% 93% 81% 95% 90% 90% 74% 74% Blue - 1 Phospho Red - 2 Phospho Green - 3 Phospho Purple - 4 Phospho 70% 60% 60% 60% 55% 60% 62% 62% 50% 48% 40% 36% 38% 40% 30% 20% 24% 24% 21% 10% 0%
23 Success of Detection by # of Phosphorylations per Peptide 1.2 Phosphopeptide Identification Success Rate, Median Monophosphorylated Doubly Phosphorylated Triply Phosphorylated Quadruply Phosphorylated
24 70% 30% 83% 91% 70% 83% 61% 83% 48%48% 87% 52% 57% 91% 48% 26% 83% 22% 74% 43% 65% 74% 57% 87% 74% 83% 48% 43% 48% 43% 78% 70% 87% 61%61% 48% 96% 70%70% 52%52% 26% 0% 10% 20% 30% 40% 50% 60% 70% 80% 90% 100% Lab Number % of Correct Phosphopeptide Identifcations
25 IMAC: Enrichment Options Immobilized Metal Ion Affinity Chromatography Chelated Fe 3+ or other metal ions on a matrix retains Phosphopeptides Methylation of Asp and Glu recommended TiO2: Phosphopeptides are retained by an Ti or Zr matrix; Methylation of Asp and Glu recommended ERLIC: Electrostatic Repulsion-Hydrophilic Interaction Chromatography Weak Anion Exchange Resin (PolyWax LP) run under Low ph and High Organic conditions Phosphopeptides are reatined by the column
26 Enrichment Identification by Peptide vs. Method of Enrichment #TI#TLEVER#SD#TIENVK #TITLEVER#SD#TIENVK Peptide - #TI#TLEVEP#SD#TIENVK Peptide - #TITLEVEP#SD#TIENVK Peptide - #TL#SD#YNIQK Peptide - #TL#SDYNIQK Peptide - #TLSDYNIQK TiO2 None IMAC & TiO2 IMAC ERLIC Peptide - ADEGI#SFR Peptide - AEFAEV#SK Peptide - DI#SL#SD#YK Peptide - DI#SLSD#YK Peptide - DISL#SD#YK TiO2 None IMAC & TiO2 IMAC ERLIC Peptide - DISLSD#YK Peptide - DQGGELL#SLR Peptide - E#S#TLHLVLR Peptide - E#STLHLVLR Peptide - ES#TLHLVLR TiO2 None IMAC & TiO2 IMAC ERLIC Peptide - IF#SIVEQR Peptide - LVNEV#TEFAK Peptide - LVQAFQF#TDK Peptide - NVAVDEL#SR Peptide - SV#SDYEGK TiO2 None IMAC & TiO2 IMAC ERLIC Peptide - THILLFLPK#SVSDYEGK Peptide - TITLEVEP#SD#TIENVK Peptide - TLSD#YNIQK TiO2 None IMAC & TiO2 IMAC ERLIC TIO2 ERLIC None IMAC IMAC & TIO Entry # Entry #
27 Method of Enrichment vs. # of Peptides Correct ERLIC IMAC IMAC/TiO2 None TiO2 ERLIC IMAC IMAC and TiO2 Enrichment - None TiO RG RG 32677V 69464V RG RG # correct TIO2 ERLIC None IMAC IMAC & TIO2
28 Pi-Peptides Detected by Enrichment and # of Pi s per Peptide ERLIC IMAC IMAC/TiO2 1 Pi 2 Pi 3 Pi 4 Pi None TiO2
29 # correct CID CID & ETD CID & HCD CID CID & ETD CID & HCD RG V RG RG CID & HCD & ETD CID & MRM CID & PSD CID & PSD CID & HCD & ETD CID & MRM MS3 MS^E Not Specified RG 32677V N.S CID MS^E 24107
30 Years of Experience vs. # of Peptides Correct 0 yrs 1 yrs 3 yrs 5 yrs 10 yrs Years Experience - 0 Years Experience - 1 Years Experience - 3 Years Experience - 5 Years Experience RG RG V V RG 85285RG # correct TIO2 ERLIC None IMAC IMAC & TIO2 = SD = Average
31 Participant Comments on the sprg 2010 Study
32 How much time did it take for you to prepare and analyze this sample? < 1 hour 1-4 hours 4-8 hours > 8 hours Other < 1 hour 1-4 hours 4-8 hours > 8 hours Other
33 How much time did it take for you to analyze the data? < 1 hour 1-4 hours 4-8 hours > 8 hours Other no response < 1 hour 1-4 hours 4-8 hours > 8 hours Other no response
34 How much time did it take you to complete this survey? < 1 hour 1 hour 2 hours 3 hours 4 hours Other 1
35 Do you think this type of study has been useful? 1 Yes No 38
36 Do you think this type of standard would be useful as a positive control prior to "real world" phospho-peptide experimentation? Yes No Other no response 36
37 What changes would make this phosphopeptide mixture more useful? Make study materials available after the study Make clearer statements of reagent concentration and source Interactive study group for learning and discussion Include more PTMs Vary phosphopeptide concentrations Increase background complexity Decrease complexity
38 How would you rate this study's level of difficulty? Easy Just Right Challenging Too Difficult
39 Have you participated in previous ABRF studies? Yes No no response
40 Based on this study, would you consider participating in future ABRF studies? Yes No Other no response 35
41 What type for study would you like to see the ABRF sprg conduct in the future? quantification glycoproteomics complex samples aceetylation de novo sequencing FDR estimation 3 4
42 sprg 2010 Study: Acknowledgements Mathias Madalinski of the Mechtler Lab (ICP Austria) for peptide synthesis, purification and analysis. Jim Makusky (NIH) as The Anonymizer. Sigma for the purified proteins.
43 To the Fearless 43 Participants: Thank You! Please Visit Our Poster # RG-8 ABRF-sPRG2010 Study: Development of a Phosphopeptide Standard for Proteomics
44 Please Visit Our Website Click on Research Groups then sprg Questions, Ideas or Interested in Joining Us? Contact Alexander Ivanov (New Chair) or Jim Farmar (Old Chair)
45 sprg Members 2010 Christopher Colangelo - Yale University Jim Farmar - Univ. of Virginia Alexander Ivanov (Chair) - Harvard School of Public Health Christopher Kinsinger - National Cancer Institute Jeffrey Kowalak (EB Liaison) - NIMH Karl Mechtler Research Institute of Molecular Pathology, Austria Brett Phinney - UC Davis Genome Center Manfred Raida - Experimental Therapeutic Center Susan Weintraub - Univ. of Texas Health Science Center at San Antonio
46
47 Pi-Peptides Detected by Enrichment and # of Pi s per Peptide Enrichment - ERLIC Enrichment - IMAC Enrichment - IMAC & TiO2 Enrichment - None Enrichment - TiO2 Y axis = # phosphorylations X axis = peptide Z = entry number
48 Peptide TLSD#YNIQK TITLEVEP#SD#TIENVK THILLFLPK#SVSDYEGK SV#SDYEGK NVAVDEL#SR LVQAFQF#TDK LVNEV#TEFAK IF#SIVEQR ES#TLHLVLR E#STLHLVLR E#S#TLHLVLR DQGGELL#SLR DISLSD#YK DISL#SD#YK DI#SLSD#YK DI#SL#SD#YK AEFAEV#SK ADEGI#SFR #TLSDYNIQK #TL#SDYNIQK #TL#SD#YNIQK #TITLEVEP#SD#TIENVK #TI#TLEVEP#SD#TIENVK TLSD#YNIQK TITLEVEP#SD#TIENVK THILLFLPK#SVSDYEGK SV#SDYEGK NVAVDEL#SR LVQAFQF#TDK LVNEV#TEFAK IF#SIVEQR ES#TLHLVLR E#STLHLVLR E#S#TLHLVLR DQGGELL#SLR DISLSD#YK DISL#SD#YK DI#SLSD#YK DI#SL#SD#YK AEFAEV#SK ADEGI#SFR #TLSDYNIQK #TL#SDYNIQK #TL#SD#YNIQK #TITLEVEP#SD#TIENVK #TI#TLEVEP#SD#TIENVK TLSD#YNIQK TITLEVEP#SD#TIENVK THILLFLPK#SVSDYEGK SV#SDYEGK NVAVDEL#SR LVQAFQF#TDK LVNEV#TEFAK IF#SIVEQR ES#TLHLVLR E#STLHLVLR E#S#TLHLVLR DQGGELL#SLR DISLSD#YK DISL#SD#YK DI#SLSD#YK DI#SL#SD#YK AEFAEV#SK ADEGI#SFR #TLSDYNIQK #TL#SDYNIQK #TL#SD#YNIQK #TITLEVEP#SD#TIENVK #TI#TLEVEP#SD#TIENVK TLSD#YNIQK TITLEVEP#SD#TIENVK THILLFLPK#SVSDYEGK SV#SDYEGK NVAVDEL#SR LVQAFQF#TDK LVNEV#TEFAK IF#SIVEQR ES#TLHLVLR E#STLHLVLR E#S#TLHLVLR DQGGELL#SLR DISLSD#YK DISL#SD#YK DI#SLSD#YK DI#SL#SD#YK AEFAEV#SK ADEGI#SFR #TLSDYNIQK #TL#SDYNIQK #TL#SD#YNIQK #TITLEVEP#SD#TIENVK #TI#TLEVEP#SD#TIENVK TLSD#YNIQK TITLEVEP#SD#TIENVK THILLFLPK#SVSDYEGK SV#SDYEGK NVAVDEL#SR LVQAFQF#TDK LVNEV#TEFAK IF#SIVEQR ES#TLHLVLR E#STLHLVLR E#S#TLHLVLR DQGGELL#SLR DISLSD#YK DISL#SD#YK DI#SLSD#YK DI#SL#SD#YK AEFAEV#SK ADEGI#SFR #TLSDYNIQK #TL#SDYNIQK #TL#SD#YNIQK #TITLEVEP#SD#TIENVK #TI#TLEVEP#SD#TIENVK Method of Enrichment vs. # of Phosphorylations and By Peptide # phosphorylations - 1 # phosphorylations - 2 # phosphorylations - 3 # phosphorylations - 4 Entry # Entry # ERLIC IMAC IMAC/TiO2 None TiO2
49 Did you or your lab receive more than one study sample? 1 4 Yes No no repsonse 34
50 Did you have knowledge of the results from another lab at anytime during this study? 1 Yes No 38
The 2012 PRG study: Assessing longitudinal variability in routine peptide LC-MS/MS analysis
 The 2012 PRG study: Assessing longitudinal variability in routine peptide LC-MS/MS analysis Thanks! 2012 PRG Maureen K. Bunger (Chair) - North Carolina Central University Tracy M Andacht - Centers for
The 2012 PRG study: Assessing longitudinal variability in routine peptide LC-MS/MS analysis Thanks! 2012 PRG Maureen K. Bunger (Chair) - North Carolina Central University Tracy M Andacht - Centers for
Mass Spectrometry and Proteomics - Lecture 6 - Matthias Trost Newcastle University
 Mass Spectrometry and Proteomics - Lecture 6 - Matthias Trost Newcastle University matthias.trost@ncl.ac.uk Previously Quantitation techniques Label-free TMT SILAC Peptide identification and FDR 194 Lecture
Mass Spectrometry and Proteomics - Lecture 6 - Matthias Trost Newcastle University matthias.trost@ncl.ac.uk Previously Quantitation techniques Label-free TMT SILAC Peptide identification and FDR 194 Lecture
Liver Mitochondria Proteomics Employing High-Resolution MS Technology
 Liver Mitochondria Proteomics Employing High-Resolution MS Technology Jenny Ho, 1 Loïc Dayon, 2 John Corthésy, 2 Umberto De Marchi, 2 Antonio Núñez, 2 Andreas Wiederkehr, 2 Rosa Viner, 3 Michael Blank,
Liver Mitochondria Proteomics Employing High-Resolution MS Technology Jenny Ho, 1 Loïc Dayon, 2 John Corthésy, 2 Umberto De Marchi, 2 Antonio Núñez, 2 Andreas Wiederkehr, 2 Rosa Viner, 3 Michael Blank,
Titansphere Phos-TiO Kit For the Selective Enrichment of Phosphopeptide
 Titansphere Phos-TiO Kit For the Selective Enrichment of Phosphopeptide Protein Phosphorylation is one of the most important post-translational modifications on a variety of cellular functions including
Titansphere Phos-TiO Kit For the Selective Enrichment of Phosphopeptide Protein Phosphorylation is one of the most important post-translational modifications on a variety of cellular functions including
Agilent AssayMAP Bravo Platform AUTOMATED PROTEIN AND PEPTIDE SAMPLE PREPARATION FOR MASS SPEC ANALYSIS
 Agilent AssayMAP Bravo Platform AUTOMATED PROTEIN AND PEPTIDE SAMPLE PREPARATION FOR MASS SPEC ANALYSIS GET FAST, ACCURATE, REPRODUCIBLE RESULTS The Agilent AssayMAP platform is an easy to use yet flexible
Agilent AssayMAP Bravo Platform AUTOMATED PROTEIN AND PEPTIDE SAMPLE PREPARATION FOR MASS SPEC ANALYSIS GET FAST, ACCURATE, REPRODUCIBLE RESULTS The Agilent AssayMAP platform is an easy to use yet flexible
IPRG 2015 (PROTEOME INFORMATICS RESEARCH GROUP) DIFFERENTIAL ABUNDANCE IN LABEL-FREE PROTEOMICS. Olga Vitek
 IPRG 21 (PROTEOME INFORMATICS RESEARCH GROUP) DIFFERENTIAL ABUNDANCE IN LABEL-FREE PROTEOMICS Olga Vitek IPRG 21 iprg committee Henry Lam - Hong Kong University of Science and Technology (Co-chair) Eugene
IPRG 21 (PROTEOME INFORMATICS RESEARCH GROUP) DIFFERENTIAL ABUNDANCE IN LABEL-FREE PROTEOMICS Olga Vitek IPRG 21 iprg committee Henry Lam - Hong Kong University of Science and Technology (Co-chair) Eugene
Peptide enrichment and fractionation
 Protein sample preparation Successful analysis of low-abundance proteins and/or the identification of posttranslationally modified peptides often require several steps: enrichment, fractionation, and/or
Protein sample preparation Successful analysis of low-abundance proteins and/or the identification of posttranslationally modified peptides often require several steps: enrichment, fractionation, and/or
iprg-2016 Proteome Informatics Research Group Study: Inferring Proteoforms from Bottom-up Proteomics Data
 iprg-2016 Proteome Informatics Research Group Study: Inferring Proteoforms from Bottom-up Proteomics Data The ABRF iprg Research Group October 10, 2016 Dear iprg 2016 Study Participant, Thank you for participating
iprg-2016 Proteome Informatics Research Group Study: Inferring Proteoforms from Bottom-up Proteomics Data The ABRF iprg Research Group October 10, 2016 Dear iprg 2016 Study Participant, Thank you for participating
Quantification of Isotope Encoded Proteins in 2D Gels
 Quantification of Isotope Encoded Proteins in 2D Gels Using Surface Enhanced Resonance Raman Giselle M. Knudsen 1, Brandon M. Davis 2, Shirshendu K. Deb 1, Yvette Loethen 2, Ravindra Gudihal 1, Pradeep
Quantification of Isotope Encoded Proteins in 2D Gels Using Surface Enhanced Resonance Raman Giselle M. Knudsen 1, Brandon M. Davis 2, Shirshendu K. Deb 1, Yvette Loethen 2, Ravindra Gudihal 1, Pradeep
MBios 478: Mass Spectrometry Applications [Dr. Wyrick] Slide #1. Lecture 25: Mass Spectrometry Applications
![MBios 478: Mass Spectrometry Applications [Dr. Wyrick] Slide #1. Lecture 25: Mass Spectrometry Applications MBios 478: Mass Spectrometry Applications [Dr. Wyrick] Slide #1. Lecture 25: Mass Spectrometry Applications](/thumbs/74/71368870.jpg) MBios 478: Mass Spectrometry Applications [Dr. Wyrick] Slide #1 Lecture 25: Mass Spectrometry Applications Measuring Protein Abundance o ICAT o DIGE Identifying Post-Translational Modifications Protein-protein
MBios 478: Mass Spectrometry Applications [Dr. Wyrick] Slide #1 Lecture 25: Mass Spectrometry Applications Measuring Protein Abundance o ICAT o DIGE Identifying Post-Translational Modifications Protein-protein
Phosphorylated Compounds by HPLC
 Phosphorylated Compounds by HPLC Featuring ERLIC, HILIC and other Ion-Exchange Techniques Table of Contents Phospholipids by HILIC Page 2 Crude Phosphorothioate by Anion Exchange 2 Phosphorylation Variants
Phosphorylated Compounds by HPLC Featuring ERLIC, HILIC and other Ion-Exchange Techniques Table of Contents Phospholipids by HILIC Page 2 Crude Phosphorothioate by Anion Exchange 2 Phosphorylation Variants
Lebedev A, 1 Damoc E, 2 Makarov A, 2 Samguina T 1 1. Moscow State University, Moscow, Russia; 2 ThermoFisher Scientific, Bremen, Germany
 Winning The Last Battle Against Edman Degradation: Reliable Leucine/Iso-leucine Differentiation In Peptide Sequencing Using an Orbitrap Fusion Mass Spectrometer Lebedev A, 1 Damoc E, 2 Makarov A, 2 Samguina
Winning The Last Battle Against Edman Degradation: Reliable Leucine/Iso-leucine Differentiation In Peptide Sequencing Using an Orbitrap Fusion Mass Spectrometer Lebedev A, 1 Damoc E, 2 Makarov A, 2 Samguina
Supplementary Figure 1. Processing and quality control for recombinant proteins.
 Supplementary Figure 1 Processing and quality control for recombinant proteins. (a) Schematic representation of processing of a recombinant protein library. Recombinant proteins were generated individually.
Supplementary Figure 1 Processing and quality control for recombinant proteins. (a) Schematic representation of processing of a recombinant protein library. Recombinant proteins were generated individually.
Small, Standardized Protein Database Provides Rapid and Statistically Significant Peptide Identifications for Targeted Searches Using Percolator
 Small, Standardized Protein Database Provides Rapid and Statistically Significant Peptide Identifications for Targeted es Using Percolator Shadab Ahmad 1, Amol Prakash 1, David Sarracino 1, Bryan Krastins
Small, Standardized Protein Database Provides Rapid and Statistically Significant Peptide Identifications for Targeted es Using Percolator Shadab Ahmad 1, Amol Prakash 1, David Sarracino 1, Bryan Krastins
Automated platform of µlc-ms/ms using SAX trap column for
 Analytical and Bioanalytical Chemistry Electronic Supplementary Material Automated platform of µlc-ms/ms using SAX trap column for highly efficient phosphopeptide analysis Xionghua Sun, Xiaogang Jiang
Analytical and Bioanalytical Chemistry Electronic Supplementary Material Automated platform of µlc-ms/ms using SAX trap column for highly efficient phosphopeptide analysis Xionghua Sun, Xiaogang Jiang
Overview. Tools for Protein Sample Preparation, 2-D Electrophoresis, and Imaging and Analysis
 Expression Proteomics // Tools for Protein Separation and Analysis www.expressionproteomics.com 1 2 3 4 Overview Tools for Protein Sample Preparation, 2-D Electrophoresis, and Imaging and Analysis overview
Expression Proteomics // Tools for Protein Separation and Analysis www.expressionproteomics.com 1 2 3 4 Overview Tools for Protein Sample Preparation, 2-D Electrophoresis, and Imaging and Analysis overview
Molecular & Cellular Proteomics 8: , 2009.
 Felix S. Oppermann, Florian Gnad, Jesper V. Olsen, Renate Hornberger, Zolta n Greff, Gyorgy Keri, Matthias Mann, and Henrik Daub. Molecular & Cellular Proteomics 8:1751 1764, 2009. Presented by Yi-Ting
Felix S. Oppermann, Florian Gnad, Jesper V. Olsen, Renate Hornberger, Zolta n Greff, Gyorgy Keri, Matthias Mann, and Henrik Daub. Molecular & Cellular Proteomics 8:1751 1764, 2009. Presented by Yi-Ting
Targeted Phosphoproteomics Analysis of Immunoaffinity Enriched Tyrosine Phosphorylation in Mouse Tissue
 Targeted Phosphoproteomics Analysis of Immunoaffinity Enriched Tyrosine Phosphorylation in Mouse Tissue Application Note Authors Ravi K. Krovvidi Agilent Technologies India Pvt. Ltd, Bangalore, India Charles
Targeted Phosphoproteomics Analysis of Immunoaffinity Enriched Tyrosine Phosphorylation in Mouse Tissue Application Note Authors Ravi K. Krovvidi Agilent Technologies India Pvt. Ltd, Bangalore, India Charles
Improving Retention Time Precision and Chromatography of Early Eluting Peptides with Acetonitrile/Water Blends as Solvent B
 Improving Retention Time Precision and Chromatography of Early Eluting Peptides with Acetonitrile/Water Blends as Solvent B Stephan Meding, Aran Paulus, and Remco Swart ¹Thermo Fisher Scientific, Germering,
Improving Retention Time Precision and Chromatography of Early Eluting Peptides with Acetonitrile/Water Blends as Solvent B Stephan Meding, Aran Paulus, and Remco Swart ¹Thermo Fisher Scientific, Germering,
Quantification of Host Cell Protein Impurities Using the Agilent 1290 Infinity II LC Coupled with the 6495B Triple Quadrupole LC/MS System
 Application Note Biotherapeutics Quantification of Host Cell Protein Impurities Using the Agilent 9 Infinity II LC Coupled with the 6495B Triple Quadrupole LC/MS System Authors Linfeng Wu and Yanan Yang
Application Note Biotherapeutics Quantification of Host Cell Protein Impurities Using the Agilent 9 Infinity II LC Coupled with the 6495B Triple Quadrupole LC/MS System Authors Linfeng Wu and Yanan Yang
ipep User s Guide Proteomics
 Proteomics ipep User s Guide ipep has been designed to aid in designing sample preparation techniques around the detection of specific sites or motifs in proteins and proteomes. The first application,
Proteomics ipep User s Guide ipep has been designed to aid in designing sample preparation techniques around the detection of specific sites or motifs in proteins and proteomes. The first application,
Faster, easier, flexible proteomics solutions
 Agilent HPLC-Chip LC/MS Faster, easier, flexible proteomics solutions Our measure is your success. products applications soft ware services Phospho- Anaysis Intact Glycan & Glycoprotein Agilent s HPLC-Chip
Agilent HPLC-Chip LC/MS Faster, easier, flexible proteomics solutions Our measure is your success. products applications soft ware services Phospho- Anaysis Intact Glycan & Glycoprotein Agilent s HPLC-Chip
RockerBox. Filtering massive Mascot search results at the.dat level
 RockerBox Filtering massive Mascot search results at the.dat level Challenges Big experiments High amount of data Large raw and.dat files (> 2GB) How to handle our results?? The 2.2 peptide summary could
RockerBox Filtering massive Mascot search results at the.dat level Challenges Big experiments High amount of data Large raw and.dat files (> 2GB) How to handle our results?? The 2.2 peptide summary could
Strategies for Quantitative Proteomics. Atelier "Protéomique Quantitative" La Grande Motte, France - June 26, 2007
 Strategies for Quantitative Proteomics Atelier "Protéomique Quantitative", France - June 26, 2007 Bruno Domon, Ph.D. Institut of Molecular Systems Biology ETH Zurich Zürich, Switzerland OUTLINE Introduction
Strategies for Quantitative Proteomics Atelier "Protéomique Quantitative", France - June 26, 2007 Bruno Domon, Ph.D. Institut of Molecular Systems Biology ETH Zurich Zürich, Switzerland OUTLINE Introduction
Thermo Scientific Solutions for Intact-Protein Analysis. Better, Faster Decisions for. Biotherapeutic Development
 Thermo Scientific Solutions for Intact-Protein Analysis Better, Faster Decisions for Biotherapeutic Development Quickly and Accurately Assess Product Quality and Safety Therapeutic proteins and monoclonal
Thermo Scientific Solutions for Intact-Protein Analysis Better, Faster Decisions for Biotherapeutic Development Quickly and Accurately Assess Product Quality and Safety Therapeutic proteins and monoclonal
Nano LC at 20 nl/min Made Easy: A Splitless Pump Combined with Fingertight UHPLC Nano Column to Boost LC-MS Sensitivity in Proteomics
 Nano LC at 20 nl/min Made Easy: A Splitless Pump Combined with Fingertight UHPLC Nano Column to Boost LC-MS Sensitivity in Proteomics Rieux L 1., De Pra M 1., Köcher T 2., Mechtler K 2., Swart R 1 1 Thermo
Nano LC at 20 nl/min Made Easy: A Splitless Pump Combined with Fingertight UHPLC Nano Column to Boost LC-MS Sensitivity in Proteomics Rieux L 1., De Pra M 1., Köcher T 2., Mechtler K 2., Swart R 1 1 Thermo
Thermo Scientific EASY-Spray Technology. Plug-and-spray with. State-of-the-art performance
 Thermo Scientific EASY-Spray Technology Plug-and-spray with State-of-the-art performance Effortless nano electrospray ionization Integrated design EASY-Spray source Nano-flow LC-MS relies critically on
Thermo Scientific EASY-Spray Technology Plug-and-spray with State-of-the-art performance Effortless nano electrospray ionization Integrated design EASY-Spray source Nano-flow LC-MS relies critically on
Algorithm for Matching Additional Spectra
 Improved Methods for Comprehensive Sample Analysis Using Protein Prospector Peter R. Baker 1, Katalin F. Medzihradszky 1 and Alma L. Burlingame 1 1 Mass Spectrometry Facility, Dept. of Pharmaceutical Chemistry,
Improved Methods for Comprehensive Sample Analysis Using Protein Prospector Peter R. Baker 1, Katalin F. Medzihradszky 1 and Alma L. Burlingame 1 1 Mass Spectrometry Facility, Dept. of Pharmaceutical Chemistry,
Modification Site Localization Scoring Integrated into a Search Engine
 Modification Site Localization Scoring Integrated into a Search Engine Peter R. Baker 1, Jonathan C. Trinidad 1, Katalin F. Medzihradszky 1, Alma L. Burlingame 1 and Robert J. Chalkley 1 1 Mass Spectrometry
Modification Site Localization Scoring Integrated into a Search Engine Peter R. Baker 1, Jonathan C. Trinidad 1, Katalin F. Medzihradszky 1, Alma L. Burlingame 1 and Robert J. Chalkley 1 1 Mass Spectrometry
Metabolomics Research Group
 Metabolomics Research Group Current Members: Amrita K Cheema: Georgetown University John M Asara: BIDMC/Harvard Medical School Thomas Neubert : NYU (EB Liaison) Chris Turck: Max Planck Institute (Chair)
Metabolomics Research Group Current Members: Amrita K Cheema: Georgetown University John M Asara: BIDMC/Harvard Medical School Thomas Neubert : NYU (EB Liaison) Chris Turck: Max Planck Institute (Chair)
CCRD Proteomics facility RIH-Brown University
 CCRD Proteomics facility RIH-Brown University Experimental design TO Publication Nagib Ahsan, PhD Where is the CCRD Proteomics Facility COBRE CCRD Proteomics Core Facility 1 Hoppin St, Providence, 02903
CCRD Proteomics facility RIH-Brown University Experimental design TO Publication Nagib Ahsan, PhD Where is the CCRD Proteomics Facility COBRE CCRD Proteomics Core Facility 1 Hoppin St, Providence, 02903
Proteomics and some of its Mass Spectrometric Applications
 Proteomics and some of its Mass Spectrometric Applications What? Large scale screening of proteins, their expression, modifications and interactions by using high-throughput approaches 2 1 Why? The number
Proteomics and some of its Mass Spectrometric Applications What? Large scale screening of proteins, their expression, modifications and interactions by using high-throughput approaches 2 1 Why? The number
Kinetics Review. Tonight at 7 PM Phys 204 We will do two problems on the board (additional ones than in the problem sets)
 Quiz 1 Kinetics Review Tonight at 7 PM Phys 204 We will do two problems on the board (additional ones than in the problem sets) I will post the problems with solutions on Toolkit for those that can t make
Quiz 1 Kinetics Review Tonight at 7 PM Phys 204 We will do two problems on the board (additional ones than in the problem sets) I will post the problems with solutions on Toolkit for those that can t make
Understanding life WITH NEXT GENERATION PROTEOMICS SOLUTIONS
 Innovative services and products for highly multiplexed protein discovery and quantification to help you understand the biological processes that shape life. Understanding life WITH NEXT GENERATION PROTEOMICS
Innovative services and products for highly multiplexed protein discovery and quantification to help you understand the biological processes that shape life. Understanding life WITH NEXT GENERATION PROTEOMICS
Nature inspired design of motif specific antibody scaffolds *
 Nature inspired design of motif specific antibody scaffolds * Nature Biotechnology, October 2013, Vol. 31, #10 * Scientific information in the following slides is from this article, if not stated otherwise
Nature inspired design of motif specific antibody scaffolds * Nature Biotechnology, October 2013, Vol. 31, #10 * Scientific information in the following slides is from this article, if not stated otherwise
ProteinPilot Report for ProteinPilot Software
 ProteinPilot Report for ProteinPilot Software Detailed Analysis of Protein Identification / Quantitation Results Automatically Sean L Seymour, Christie Hunter SCIEX, USA Powerful mass spectrometers like
ProteinPilot Report for ProteinPilot Software Detailed Analysis of Protein Identification / Quantitation Results Automatically Sean L Seymour, Christie Hunter SCIEX, USA Powerful mass spectrometers like
Detecting Challenging Post Translational Modifications (PTMs) using CESI-MS
 Detecting Challenging Post Translational Modifications (PTMs) using CESI-MS Sensitive workflow for the detection of PTMs in biological samples from small sample volumes Klaus Faserl, 1 Bettina Sarg, 1
Detecting Challenging Post Translational Modifications (PTMs) using CESI-MS Sensitive workflow for the detection of PTMs in biological samples from small sample volumes Klaus Faserl, 1 Bettina Sarg, 1
Proteomics. Proteomics is the study of all proteins within organism. Challenges
 Proteomics Proteomics is the study of all proteins within organism. Challenges 1. The proteome is larger than the genome due to alternative splicing and protein modification. As we have said before we
Proteomics Proteomics is the study of all proteins within organism. Challenges 1. The proteome is larger than the genome due to alternative splicing and protein modification. As we have said before we
ProteinChip IMAC30 Array (Immobilized Metal Affinity Capture) Instruction Manual. Catalog #C
 ProteinChip IMAC30 Array (Immobilized Metal Affinity Capture) Instruction Manual Catalog #C57-30078 For technical support, call your local Bio-Rad office, or in the US, call 1-800-4BIORAD (1-800-424-6723).
ProteinChip IMAC30 Array (Immobilized Metal Affinity Capture) Instruction Manual Catalog #C57-30078 For technical support, call your local Bio-Rad office, or in the US, call 1-800-4BIORAD (1-800-424-6723).
A Fast and Robust Linear ph Gradient Separation Platform for Monoclonal Antibody (MAb) Charge Variant Analysis
 A Fast and Robust Linear ph Gradient Separation Platform for Monoclonal Antibody (MAb) Charge Variant Analysis Shanhua Lin, Julia Baek, and Chris Pohl Thermo Fisher Scientific, Sunnyvale, CA, USA Overview
A Fast and Robust Linear ph Gradient Separation Platform for Monoclonal Antibody (MAb) Charge Variant Analysis Shanhua Lin, Julia Baek, and Chris Pohl Thermo Fisher Scientific, Sunnyvale, CA, USA Overview
Basic protein and peptide science for proteomics. Henrik Johansson
 Basic protein and peptide science for proteomics Henrik Johansson Proteins are the main actors in the cell Membranes Transport and storage Chemical factories DNA Building proteins Structure Proteins mediate
Basic protein and peptide science for proteomics Henrik Johansson Proteins are the main actors in the cell Membranes Transport and storage Chemical factories DNA Building proteins Structure Proteins mediate
Exam MOL3007 Functional Genomics
 Faculty of Medicine Department of Cancer Research and Molecular Medicine Exam MOL3007 Functional Genomics Thursday December 20 th 9.00-13.00 ECTS credits: 7.5 Number of pages (included front-page): 5 Supporting
Faculty of Medicine Department of Cancer Research and Molecular Medicine Exam MOL3007 Functional Genomics Thursday December 20 th 9.00-13.00 ECTS credits: 7.5 Number of pages (included front-page): 5 Supporting
Phosphoproteomics and Lung Cancer Research
 Int. J. Mol. Sci. 2012, 13, 12287-12314; doi:10.3390/ijms131012287 Review OPEN ACCESS International Journal of Molecular Sciences ISSN 1422-0067 www.mdpi.com/journal/ijms Phosphoproteomics and Lung Cancer
Int. J. Mol. Sci. 2012, 13, 12287-12314; doi:10.3390/ijms131012287 Review OPEN ACCESS International Journal of Molecular Sciences ISSN 1422-0067 www.mdpi.com/journal/ijms Phosphoproteomics and Lung Cancer
Phosphoproteomics. PhD Henri Blomster
 hosphoproteomics hd Henri Blomster Human Genome roject Budding yeast 6,000 genes Nematode 15,000 genes Human 25,000 genes AAachment of chemical groups Covalent aaachment of polypepedes rotein cleavage
hosphoproteomics hd Henri Blomster Human Genome roject Budding yeast 6,000 genes Nematode 15,000 genes Human 25,000 genes AAachment of chemical groups Covalent aaachment of polypepedes rotein cleavage
CHROMATOGRAPHY OF DIFFICULT PROTEINS. Andrew J. Alpert, Ph.D. PolyLC Inc. Columbia, MD U.S.A.
 CHROMATOGRAPHY OF DIFFICULT PROTEINS Andrew J. Alpert, Ph.D. PolyLC Inc. Columbia, MD U.S.A. PROTEOMICS ANALYSES A butterfly and a caterpillar have the same genome but different proteomes. Techniques for
CHROMATOGRAPHY OF DIFFICULT PROTEINS Andrew J. Alpert, Ph.D. PolyLC Inc. Columbia, MD U.S.A. PROTEOMICS ANALYSES A butterfly and a caterpillar have the same genome but different proteomes. Techniques for
Agilent Technologies. Jim Hollenhorst, Ph.D. Senior Director of Technology. FAPESP/Agilent Workshop August 30,
 Agilent Technologies Jim Hollenhorst, Ph.D. Senior Director of Technology 1 A Brief History of Agilent 1939 1998: 1999: 2005: 2006 2010: 2013: 2013: 2014: The Hewlett-Packard years 60-year heritage of
Agilent Technologies Jim Hollenhorst, Ph.D. Senior Director of Technology 1 A Brief History of Agilent 1939 1998: 1999: 2005: 2006 2010: 2013: 2013: 2014: The Hewlett-Packard years 60-year heritage of
Isolation of Tryptic Phosphopeptides by ERLIC (Electrostatic Repulsion-Hydrophilic Interaction Chromatography)
 Edited, HPLC 2008 poster P-2412-W Isolation of Tryptic Phosphopeptides by ERLIC (Electrostatic Repulsion-Hydrophilic Interaction Chromatography) Andrew Alpert 1, Goran Mitulović 2, and Karl Mechtler 2
Edited, HPLC 2008 poster P-2412-W Isolation of Tryptic Phosphopeptides by ERLIC (Electrostatic Repulsion-Hydrophilic Interaction Chromatography) Andrew Alpert 1, Goran Mitulović 2, and Karl Mechtler 2
Proteomics and Cancer
 Proteomics and Cancer Japan Society for the Promotion of Science (JSPS) Science Dialogue Program at Niitsu Senior High School Niitsu, Niigata September 4th 2006 Vladimir Valera, M.D, PhD JSPS Postdoctoral
Proteomics and Cancer Japan Society for the Promotion of Science (JSPS) Science Dialogue Program at Niitsu Senior High School Niitsu, Niigata September 4th 2006 Vladimir Valera, M.D, PhD JSPS Postdoctoral
Isolation of Tryptic Phosphopeptides by ERLIC (Electrostatic Repulsion-Hydrophilic Interaction Chromatography)
 HPLC 2008 poster P-2412-W Isolation of Tryptic Phosphopeptides by ERLIC (Electrostatic Repulsion-Hydrophilic Interaction Chromatography) Andrew Alpert 1, Goran Mitulović 2, and Karl Mechtler 2 1 PolyLC
HPLC 2008 poster P-2412-W Isolation of Tryptic Phosphopeptides by ERLIC (Electrostatic Repulsion-Hydrophilic Interaction Chromatography) Andrew Alpert 1, Goran Mitulović 2, and Karl Mechtler 2 1 PolyLC
Introduction. Three different cases for different dosage forms and pharmacopeias, respectively, must be distinguished (Table 1).
 An Integrated Solution for Calculating Results from Tests for Content Uniformity of Dosage Units According to the United States and European Pharmacopeia Barbara van Cann, Andreas Brunner, Fraser McLeod,
An Integrated Solution for Calculating Results from Tests for Content Uniformity of Dosage Units According to the United States and European Pharmacopeia Barbara van Cann, Andreas Brunner, Fraser McLeod,
Improved Peptide Maps Using Core-Shell Media: Explaining Better Biopharmaceutical Applications With Kinetex C18 Columns
 Improved Peptide Maps Using Core-Shell Media: Explaining Better Biopharmaceutical Applications With Kinetex C18 Columns Michael McGinley, Jeff Layne, and Vita Knudson Phenomenex, Inc., 411 Madrid Ave.,Torrance,
Improved Peptide Maps Using Core-Shell Media: Explaining Better Biopharmaceutical Applications With Kinetex C18 Columns Michael McGinley, Jeff Layne, and Vita Knudson Phenomenex, Inc., 411 Madrid Ave.,Torrance,
BIOC 463A Expt. 4: Column Chromatographic Methods Column Chromatography
 Column Chromatography Chromatography is the process use to separate molecules based on SOME physical property of the molecule: Mass (i.e. size) Charge Affinity for ligands or substrates Hydrophobic interactions
Column Chromatography Chromatography is the process use to separate molecules based on SOME physical property of the molecule: Mass (i.e. size) Charge Affinity for ligands or substrates Hydrophobic interactions
Strategies in proteomics
 Strategies in proteomics Systems biology - understand cellpathways, network, and complex interacting (includes Genomics, Proteomics, Metabolomics..) Biological processes - characterize protein complexes,
Strategies in proteomics Systems biology - understand cellpathways, network, and complex interacting (includes Genomics, Proteomics, Metabolomics..) Biological processes - characterize protein complexes,
for water and beverage analysis
 Thermo Scientific EQuan MAX Plus Systems Automated, high-throughput LC-MS solutions for water and beverage analysis Pesticides Pharmaceuticals Personal care products Endocrine disruptors Perfluorinated
Thermo Scientific EQuan MAX Plus Systems Automated, high-throughput LC-MS solutions for water and beverage analysis Pesticides Pharmaceuticals Personal care products Endocrine disruptors Perfluorinated
Instructions for Capillary Electrophoresis Peptide Analysis Kit
 Instructions for Capillary Electrophoresis Peptide Analysis Kit Catalog Number 148-4110 For Technical Service Call Your Local Bio-Rad Office or in the U.S. Call 1-800-4BIORAD (1-800-424-6723) Table of
Instructions for Capillary Electrophoresis Peptide Analysis Kit Catalog Number 148-4110 For Technical Service Call Your Local Bio-Rad Office or in the U.S. Call 1-800-4BIORAD (1-800-424-6723) Table of
An Introduction to the Association of Biomolecular Resource Facilities (ABRF) and Recent Results of a Laboratory Funding Survey
 An Introduction to the Association of Biomolecular Resource Facilities (ABRF) and Recent Results of a Laboratory Funding Survey Jack Simpson, Ph.D. member, Executive Board, ABRF Sr. Scientist, Advanced
An Introduction to the Association of Biomolecular Resource Facilities (ABRF) and Recent Results of a Laboratory Funding Survey Jack Simpson, Ph.D. member, Executive Board, ABRF Sr. Scientist, Advanced
Using HRAM Survey Analysis Combined with Rapid MS2 Data to Develop a Fragmentation Based Detection Workflow for Structure ID Acquisition
 Using HRAM Survey Analysis Combined with Rapid MS Data to Develop a Fragmentation Based Detection Workflow for Structure ID Acquisition Tim Stratton, 1 Caroline Ding, 1 Christoph Henrich, Hans Grensemann,
Using HRAM Survey Analysis Combined with Rapid MS Data to Develop a Fragmentation Based Detection Workflow for Structure ID Acquisition Tim Stratton, 1 Caroline Ding, 1 Christoph Henrich, Hans Grensemann,
GENReports: Market & Tech Analysis. > Gary Oosta, Ph.D.
 GENReports: Market & Tech Analysis > Enal Razvi, Ph.D. Managing Director Select Biosciences, Inc. enal@selectbio.us! > Gary Oosta, Ph.D. Biotechnology Analyst, Select Biosciences, Inc. Analysis,of,the,Point@of@Care,DiagnosCcs,(POCD),Marketplace,
GENReports: Market & Tech Analysis > Enal Razvi, Ph.D. Managing Director Select Biosciences, Inc. enal@selectbio.us! > Gary Oosta, Ph.D. Biotechnology Analyst, Select Biosciences, Inc. Analysis,of,the,Point@of@Care,DiagnosCcs,(POCD),Marketplace,
Purification: Step 1. Lecture 11 Protein and Peptide Chemistry. Cells: Break them open! Crude Extract
 Purification: Step 1 Lecture 11 Protein and Peptide Chemistry Cells: Break them open! Crude Extract Total contents of cell Margaret A. Daugherty Fall 2003 Big Problem: Crude extract is not the natural
Purification: Step 1 Lecture 11 Protein and Peptide Chemistry Cells: Break them open! Crude Extract Total contents of cell Margaret A. Daugherty Fall 2003 Big Problem: Crude extract is not the natural
Purification: Step 1. Protein and Peptide Chemistry. Lecture 11. Big Problem: Crude extract is not the natural environment. Cells: Break them open!
 Lecture 11 Protein and Peptide Chemistry Margaret A. Daugherty Fall 2003 Purification: Step 1 Cells: Break them open! Crude Extract Total contents of cell Big Problem: Crude extract is not the natural
Lecture 11 Protein and Peptide Chemistry Margaret A. Daugherty Fall 2003 Purification: Step 1 Cells: Break them open! Crude Extract Total contents of cell Big Problem: Crude extract is not the natural
Model 491 Prep Cell. Bio-Rad Tech Note Summaries. Size, %T Plasma protein 49 kd 12% 60 kd 7% 1773 Recombinant. proteins
 Model 491 Prep Cell Bio-Rad Tech Note Summaries 2-D Applications Bulletin Title # Component Preparative 2-D Purifies Proteins for Sequencing or Antibody Production Preparative 2-D Electrophoresis System
Model 491 Prep Cell Bio-Rad Tech Note Summaries 2-D Applications Bulletin Title # Component Preparative 2-D Purifies Proteins for Sequencing or Antibody Production Preparative 2-D Electrophoresis System
Protein analysis. Dr. Mamoun Ahram Summer semester, Resources This lecture Campbell and Farrell s Biochemistry, Chapters 5
 Protein analysis Dr. Mamoun Ahram Summer semester, 2015-2016 Resources This lecture Campbell and Farrell s Biochemistry, Chapters 5 Bases of protein separation Proteins can be purified on the basis Solubility
Protein analysis Dr. Mamoun Ahram Summer semester, 2015-2016 Resources This lecture Campbell and Farrell s Biochemistry, Chapters 5 Bases of protein separation Proteins can be purified on the basis Solubility
Thermo Scientific EASY-nLC 1200 System. Leading in simplicity. and performance
 Thermo Scientific EASY-nLC 1200 System Leading in simplicity and performance Peak performance made EASY Effortless ultra high performance for everybody A straightforward LC-MS solution Optimized and integrated
Thermo Scientific EASY-nLC 1200 System Leading in simplicity and performance Peak performance made EASY Effortless ultra high performance for everybody A straightforward LC-MS solution Optimized and integrated
Two-Dimensional LC Protein Separation on Monolithic Columns in a Fully Automated Workflow
 Technical Note Two-Dimensional LC Protein Separation on Monolithic Columns in a Fully Automated Workflow Introduction The complex nature of proteomics samples requires high-resolution separation methods
Technical Note Two-Dimensional LC Protein Separation on Monolithic Columns in a Fully Automated Workflow Introduction The complex nature of proteomics samples requires high-resolution separation methods
Cell Signaling Technology
 Cell Signaling Technology PTMScan Direct: Multipathway v2.0 Proteomics Service Group January 14, 2013 PhosphoScan Deliverables Project Overview Methods PTMScan Direct: Multipathway V2.0 (Tables 1,2) Qualitative
Cell Signaling Technology PTMScan Direct: Multipathway v2.0 Proteomics Service Group January 14, 2013 PhosphoScan Deliverables Project Overview Methods PTMScan Direct: Multipathway V2.0 (Tables 1,2) Qualitative
columns PepSwift and ProSwift Capillary Monolithic Reversed-Phase Columns
 columns PepSwift and ProSwift Capillary Monolithic Reversed-Phase Columns PepSwift and ProSwift monolithic columns are specially designed for high-resolution LC/MS analysis in protein identification, biomarker
columns PepSwift and ProSwift Capillary Monolithic Reversed-Phase Columns PepSwift and ProSwift monolithic columns are specially designed for high-resolution LC/MS analysis in protein identification, biomarker
IEA Wind Co-operation. Name Hannele Holttinen IEA Wind Chair Workshop Winterwind 2011 Place Umeå, Sweden Date 9th Feb, 2011
 IEA Wind Co-operation Name Hannele Holttinen IEA Wind Chair Workshop Winterwind 2011 Place Umeå, Sweden Date 9th Feb, 2011 86% of the world wind capacity is in IEA Wind member countries Map IEA Wind Countries
IEA Wind Co-operation Name Hannele Holttinen IEA Wind Chair Workshop Winterwind 2011 Place Umeå, Sweden Date 9th Feb, 2011 86% of the world wind capacity is in IEA Wind member countries Map IEA Wind Countries
Application Note USD Purification of Mouse IgM from Cell Culture Supernatant by Cation Exchange Chromatography on CM Ceramic HyperD F Sorbent
 Application Note USD 241 Purification of Mouse IgM from Cell Culture Supernatant by Cation Exchange Chromatography on CM Ceramic HyperD F Sorbent What this Study Demonstrates T h i s s t u d y o n C a
Application Note USD 241 Purification of Mouse IgM from Cell Culture Supernatant by Cation Exchange Chromatography on CM Ceramic HyperD F Sorbent What this Study Demonstrates T h i s s t u d y o n C a
Fast protein separations with ion exchange columns
 Fast protein separations with ion exchange columns Koji Nakamura*, Yoshio Kato and Shuichi Okuzono Nanyo Research Lab, Tosoh Corporation, 4560 Kaisei-cho, Shunan, Yamaguchi 746-8501, Japan TP106 0807 ISPPP-05
Fast protein separations with ion exchange columns Koji Nakamura*, Yoshio Kato and Shuichi Okuzono Nanyo Research Lab, Tosoh Corporation, 4560 Kaisei-cho, Shunan, Yamaguchi 746-8501, Japan TP106 0807 ISPPP-05
Bringing custom solutions and contract manufacturing to life
 GE Healthcare Bringing custom solutions and contract manufacturing to life Imagined. Delivered. Supported. Custom and bulk solutions Tailored to your precise specifications, our custom products deliver
GE Healthcare Bringing custom solutions and contract manufacturing to life Imagined. Delivered. Supported. Custom and bulk solutions Tailored to your precise specifications, our custom products deliver
Development of a hybrid assay for the quantification of a mab drug in human serum at the low ng/ml levels
 Development of a hybrid assay for the quantification of a mab drug in human serum at the low ng/ml levels EBF Open Symposium 21-23 November 2018 marco.michi@aptuit.com Presentation Topics Current scenario
Development of a hybrid assay for the quantification of a mab drug in human serum at the low ng/ml levels EBF Open Symposium 21-23 November 2018 marco.michi@aptuit.com Presentation Topics Current scenario
Jet Steam Proteomics with Automated Sample Prep for Superior Peptide Quantitation
 Jet Steam Proteomics with Automated Sample Prep for Superior Peptide Quantitation Christine Miller Jason Russell June 2, 205 Overview Automated sample preparation Jet Stream Proteomics Peptide quantitation
Jet Steam Proteomics with Automated Sample Prep for Superior Peptide Quantitation Christine Miller Jason Russell June 2, 205 Overview Automated sample preparation Jet Stream Proteomics Peptide quantitation
Efficient Multi-Well Protein Purification Strategies
 Application Note PN 33576 Efficient Multi-Well Protein Purification Strategies Introduction Many tools and techniques are available today for protein purification. Development of a purification process
Application Note PN 33576 Efficient Multi-Well Protein Purification Strategies Introduction Many tools and techniques are available today for protein purification. Development of a purification process
New Approaches to Quantitative Proteomics Analysis
 New Approaches to Quantitative Proteomics Analysis Chris Hodgkins, Market Development Manager, SCIEX ANZ 2 nd November, 2017 Who is SCIEX? Founded by Dr. Barry French & others: University of Toronto Introduced
New Approaches to Quantitative Proteomics Analysis Chris Hodgkins, Market Development Manager, SCIEX ANZ 2 nd November, 2017 Who is SCIEX? Founded by Dr. Barry French & others: University of Toronto Introduced
Method Optimisation in Bottom-Up Analysis of Proteins
 Method Optimisation in Bottom-Up Analysis of Proteins M. Styles, 1 D. Smith, 2 J. Griffiths, 2 L. Pereira, 3 T. Edge 3 1 University of Manchester, UK; 2 Patterson Institute, Manchester, UK; 3 Thermo Fisher
Method Optimisation in Bottom-Up Analysis of Proteins M. Styles, 1 D. Smith, 2 J. Griffiths, 2 L. Pereira, 3 T. Edge 3 1 University of Manchester, UK; 2 Patterson Institute, Manchester, UK; 3 Thermo Fisher
Enrichment of EGFR/PI3K/AKT/PTEN Proteins for Research using Immunoprecipitation and with Mass Spectrometry-based Analysis
 Enrichment of /PI3K/AKT/ Proteins for Research using Immunoprecipitation and with Mass Spectrometry-based Analysis Bhavin Patel, Scott Meier, Kay Opperman, Paul Haney, Barbara Kaboord, John Rogers Thermo
Enrichment of /PI3K/AKT/ Proteins for Research using Immunoprecipitation and with Mass Spectrometry-based Analysis Bhavin Patel, Scott Meier, Kay Opperman, Paul Haney, Barbara Kaboord, John Rogers Thermo
Supplemental Materials
 Supplemental Materials MSGFDB Parameters For all MSGFDB searches, the following parameters were used: 30 ppm precursor mass tolerance, enable target-decoy search, enzyme specific scoring (for Arg-C, Asp-N,
Supplemental Materials MSGFDB Parameters For all MSGFDB searches, the following parameters were used: 30 ppm precursor mass tolerance, enable target-decoy search, enzyme specific scoring (for Arg-C, Asp-N,
Promix TM. Enter a New Era in Biomolecule Analysis with. Columns. Unsurpassed Selectivity and Peak Capacity for Peptides and Proteins
 Promix TM Enter a New Era in Biomolecule Analysis with Promix TM Columns Unsurpassed electivity and Peak Capacity for Peptides and Proteins Applications: Proteomics Peptide/Protein Analysis Peptide/Protein
Promix TM Enter a New Era in Biomolecule Analysis with Promix TM Columns Unsurpassed electivity and Peak Capacity for Peptides and Proteins Applications: Proteomics Peptide/Protein Analysis Peptide/Protein
Fast mass transfer Fast separations High throughput and improved productivity Long column lifetime Outstanding reproducibility Low carryover
 columns ProSwift Reversed-Phase Monolith Columns for Protein Analysis ProSwift reversed-phase columns use a unique monolith technology for fast, high-resolution HPLC and LC/MS separations of proteins.
columns ProSwift Reversed-Phase Monolith Columns for Protein Analysis ProSwift reversed-phase columns use a unique monolith technology for fast, high-resolution HPLC and LC/MS separations of proteins.
A Unique LC-MS Assay for Host Cell Proteins(HCPs) ) in Biologics
 A Unique LC-MS Assay for Host Cell Proteins(HCPs) ) in Biologics Catalin Doneanu,, Ph.D. Biopharmaceutical Sciences, Waters September 16, 2009 Mass Spec 2009 2009 Waters Corporation Host Cell Proteins
A Unique LC-MS Assay for Host Cell Proteins(HCPs) ) in Biologics Catalin Doneanu,, Ph.D. Biopharmaceutical Sciences, Waters September 16, 2009 Mass Spec 2009 2009 Waters Corporation Host Cell Proteins
[ MS IMAGING ] FULL SPECTRUM MOLECULAR IMAGING COMPLETE VISUALIZATION OF MOLECULAR DISTRIBUTIONS
![[ MS IMAGING ] FULL SPECTRUM MOLECULAR IMAGING COMPLETE VISUALIZATION OF MOLECULAR DISTRIBUTIONS [ MS IMAGING ] FULL SPECTRUM MOLECULAR IMAGING COMPLETE VISUALIZATION OF MOLECULAR DISTRIBUTIONS](/thumbs/93/113947637.jpg) [ MS IMAGING ] FULL SPECTRUM MOLECULAR IMAGING COMPLETE VISUALIZATION OF MOLECULAR DISTRIBUTIONS THE SPATIAL DISTRIBUTION OF MOLECULES, DETERMINED BY MS IMAGING, CAN PROVIDE A WEALTH OF INFORMATION REGARDING
[ MS IMAGING ] FULL SPECTRUM MOLECULAR IMAGING COMPLETE VISUALIZATION OF MOLECULAR DISTRIBUTIONS THE SPATIAL DISTRIBUTION OF MOLECULES, DETERMINED BY MS IMAGING, CAN PROVIDE A WEALTH OF INFORMATION REGARDING
Introduction to Protein Purification
 Introduction to Protein Purification 1 Day 1) Introduction to Protein Purification. Input for Purification Protocol Development - Guidelines for Protein Purification Day 2) Sample Preparation before Chromatography
Introduction to Protein Purification 1 Day 1) Introduction to Protein Purification. Input for Purification Protocol Development - Guidelines for Protein Purification Day 2) Sample Preparation before Chromatography
What are proteomics? And what can they tell us about seed maturation and germination?
 Danseed Symposium Kobæk Strand, Skelskør 14 March, 2017 What are proteomics? And what can they tell us about seed maturation and germination? Ian Max Møller Department of Molecular Biology and Genetics
Danseed Symposium Kobæk Strand, Skelskør 14 March, 2017 What are proteomics? And what can they tell us about seed maturation and germination? Ian Max Møller Department of Molecular Biology and Genetics
ProteoMiner Protein Enrichment Technology
 Sample Preparation ProteoMiner Protein Enrichment Technology Digging Deeper in the Proteome Detect More Proteins With ProteoMiner Technology Than With Immunodepletion ProteoMiner Technology vs. an Agilent
Sample Preparation ProteoMiner Protein Enrichment Technology Digging Deeper in the Proteome Detect More Proteins With ProteoMiner Technology Than With Immunodepletion ProteoMiner Technology vs. an Agilent
Innovations for Protein Research. Protein Research. Powerful workflows built on solid science
 Innovations for Protein Research Protein Research Powerful workflows built on solid science Your partner in discovering innovative solutions for quantitative protein and biomarker research Applied Biosystems/MDS
Innovations for Protein Research Protein Research Powerful workflows built on solid science Your partner in discovering innovative solutions for quantitative protein and biomarker research Applied Biosystems/MDS
SUPPLEMENTARY INFORMATION
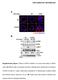 SUPPLEMENTARY INFORMATION Supplementary Figure 1 Effect of ROCK inhibition on lumen abnormality in MDCK cysts. (A) MDCK cells as indicated cultured in Matrigel were treated with and without Y27632 (10
SUPPLEMENTARY INFORMATION Supplementary Figure 1 Effect of ROCK inhibition on lumen abnormality in MDCK cysts. (A) MDCK cells as indicated cultured in Matrigel were treated with and without Y27632 (10
Charge Heterogeneity Analysis of Rituximab Innovator and Biosimilar mabs
 Charge Heterogeneity Analysis of Rituximab Innovator and Biosimilar mabs Application Note Author Suresh Babu C.V. Agilent Technologies India Pvt. Ltd, Bangalore, India Abstract This Application Note describes
Charge Heterogeneity Analysis of Rituximab Innovator and Biosimilar mabs Application Note Author Suresh Babu C.V. Agilent Technologies India Pvt. Ltd, Bangalore, India Abstract This Application Note describes
About OMICS Group Conferences
 About OMICS Group OMICS Group International is an amalgamation of Open Access publications and worldwide international science conferences and events. Established in the year 2007 with the sole aim of
About OMICS Group OMICS Group International is an amalgamation of Open Access publications and worldwide international science conferences and events. Established in the year 2007 with the sole aim of
Jet Stream Proteomics for Sensitive and Robust Standard Flow LC/MS
 Jet Stream Proteomics for Sensitive and Robust Standard Flow LC/MS Technical Overview Authors Yanan Yang, Vadiraj hat, and Christine Miller Agilent Technologies, Inc. Santa Clara, California Introduction
Jet Stream Proteomics for Sensitive and Robust Standard Flow LC/MS Technical Overview Authors Yanan Yang, Vadiraj hat, and Christine Miller Agilent Technologies, Inc. Santa Clara, California Introduction
Examining the components of your peptide sample with AccuPep QC. Lauren Lu, Ph.D. October 29, 2015, 9:00-10:00 AM EST
 Examining the components of your peptide sample with AccuPep QC Lauren Lu, Ph.D. October 29, 2015, 9:00-10:00 AM EST When do I need custom peptides? Custom peptides play an important role in many research
Examining the components of your peptide sample with AccuPep QC Lauren Lu, Ph.D. October 29, 2015, 9:00-10:00 AM EST When do I need custom peptides? Custom peptides play an important role in many research
Generating Quantitative Assays for Biomarker Development. Jeff Whiteaker, Ph.D. Paulovich Laboratory, Fred Hutchinson Cancer Research Center
 Generating Quantitative Assays for Biomarker Development Jeff Whiteaker, Ph.D. Paulovich Laboratory, Fred Hutchinson Cancer Research Center Reliable assays are available for
Generating Quantitative Assays for Biomarker Development Jeff Whiteaker, Ph.D. Paulovich Laboratory, Fred Hutchinson Cancer Research Center Reliable assays are available for
Thermo Scientific Orbitrap Fusion Lumos Tribrid Mass Spectrometer. Breakthrough Gains for Quantitative Biology. Sensitivity Transformed
 Thermo Scientific rbitrap Fusion Lumos Tribrid Mass Spectrometer Breakthrough Gains for Quantitative Biology Sensitivity Transformed The Most Advanced Instrument Built for the Most Challenging Experiments
Thermo Scientific rbitrap Fusion Lumos Tribrid Mass Spectrometer Breakthrough Gains for Quantitative Biology Sensitivity Transformed The Most Advanced Instrument Built for the Most Challenging Experiments
Identification of RNA Linkage Isomers by Anion-Exchange Purification with ESI-MS of Automatically-Desalted Phosphodiesterase-II Digests
 Identification of RNA Linkage Isomers by Anion-Exchange Purification with ESI-MS of Automatically-Desalted Phosphodiesterase-II Digests J.R. Thayer, 1 Nitin Puri, Chris Burnett, Mark Hail, 3 Srinivasa
Identification of RNA Linkage Isomers by Anion-Exchange Purification with ESI-MS of Automatically-Desalted Phosphodiesterase-II Digests J.R. Thayer, 1 Nitin Puri, Chris Burnett, Mark Hail, 3 Srinivasa
NPTEL VIDEO COURSE PROTEOMICS PROF. SANJEEVA SRIVASTAVA
 LECTURE-24 Quantitative proteomics: Stable isotope labeling by amino acids in cell culture (SILAC) TRANSCRIPT Welcome to the proteomics course. In today s lecture we will talk about Quantitative proteomics:
LECTURE-24 Quantitative proteomics: Stable isotope labeling by amino acids in cell culture (SILAC) TRANSCRIPT Welcome to the proteomics course. In today s lecture we will talk about Quantitative proteomics:
PYRIDOXAL PHOSHATE AS A TAG TO IDENTIFY ENZYMES WITHIN THE PLP-OME. A Thesis KAYLA J. MESSER
 PYRIDOXAL PHOSHATE AS A TAG TO IDENTIFY ENZYMES WITHIN THE PLP-OME A Thesis by KAYLA J. MESSER Submitted to the Office of Graduate Studies of Texas A&M University in partial fulfillment of the requirements
PYRIDOXAL PHOSHATE AS A TAG TO IDENTIFY ENZYMES WITHIN THE PLP-OME A Thesis by KAYLA J. MESSER Submitted to the Office of Graduate Studies of Texas A&M University in partial fulfillment of the requirements
Custom Antibody Services. Antibodies Designed Just for You. HuCAL Recombinant Monoclonal Antibody Generation Service
 Custom Antibody Services Antibodies Designed Just for You HuCAL Recombinant Monoclonal Antibody Generation Service Antibodies Designed Just for You What if there was a technology that would allow you to
Custom Antibody Services Antibodies Designed Just for You HuCAL Recombinant Monoclonal Antibody Generation Service Antibodies Designed Just for You What if there was a technology that would allow you to
Synthesis of diazo ketone solid supports and their application towards the enrichment of phosphorylated peptides
 Supplementary Material (ESI) for rganic & Biomolecular Chemistry Synthesis of diazo ketone solid supports and their application towards the enrichment of phosphorylated peptides Samantha M. Frawley 1,
Supplementary Material (ESI) for rganic & Biomolecular Chemistry Synthesis of diazo ketone solid supports and their application towards the enrichment of phosphorylated peptides Samantha M. Frawley 1,
ApplicationNOTE THE USE OF A NANOLOCKSPRAY ELECTROSPRAY INTERFACE FOR EXACT MASS PROTEOMICS STUDIES
 OVERVIEW This application note describes the implementation of a dual sprayer NanoLockSpray TM source on the Waters Micromass Q-Tof micro TM mass spectrometer This source consists of a dual sprayer arrangement;
OVERVIEW This application note describes the implementation of a dual sprayer NanoLockSpray TM source on the Waters Micromass Q-Tof micro TM mass spectrometer This source consists of a dual sprayer arrangement;
ABRF-PRG07: Advanced Quantitative Proteomics Study
 ARTICLE ABRF-PRG07: Advanced Quantitative Proteomics Study Arnold M. Falick, 1, * William S. Lane, 2 Kathryn S. Lilley, 3 Michael J. MacCoss, 4 Brett S. Phinney, 5 Nicholas E. Sherman, 6 Susan T. Weintraub,
ARTICLE ABRF-PRG07: Advanced Quantitative Proteomics Study Arnold M. Falick, 1, * William S. Lane, 2 Kathryn S. Lilley, 3 Michael J. MacCoss, 4 Brett S. Phinney, 5 Nicholas E. Sherman, 6 Susan T. Weintraub,
Lecture 5: 8/31. CHAPTER 5 Techniques in Protein Biochemistry
 Lecture 5: 8/31 CHAPTER 5 Techniques in Protein Biochemistry Chapter 5 Outline The proteome is the entire set of proteins expressed and modified by a cell under a particular set of biochemical conditions.
Lecture 5: 8/31 CHAPTER 5 Techniques in Protein Biochemistry Chapter 5 Outline The proteome is the entire set of proteins expressed and modified by a cell under a particular set of biochemical conditions.
