Detailed Characterization of an Apparently Unspliced,
|
|
|
- Justin Phelps
- 6 years ago
- Views:
Transcription
1 JOURNAL OF VIROLOGY, Sept. 1982, p X/82/ $02.00/0 Vol. 43. No. 3 Detailed Characterization of an Apparently Unspliced, Herpes Simplex Virus Type 1 Another Gene Mapping in the Interior of KENNETH G. DRAPER, RAYMOND J. FRINK, AND EDWARD K. WAGNER* Department of Molecular Biology and Biochemistry, Unii ersity of California, Inrine, Irvline, Cailifornia Received 19 February 1982/Accepted 20 May 1982 We precisely localized the coding region and determined the nucleotide sequence of a 1.2-kilobase herpes simplex virus type 1 mrna which underlies the 3' region of the 5.2-kilobase mrna mapping in HindlIl fragment K. This mrna, which lacks readily detectable splices, has its own promoter by the criteria of identification of putative herpes simplex virus type 1 control sequences and in vitro transcription by a Manley polymerase system. Previously, we characterized the major mrna species of herpes simplex virus type 1 (HSV-1) mapping in Hindlll fragments K (0.527 to 0.592) and L (0.592 to 0.647) and EcoRI fragment 1 (0.633 to 0.721) (1, 9, lla). Many of these mrnas overlap each other. As a good example, consider the 6.9-, 5.2-, 1.7-, and 1.2- kilobase (kb) mrnas mapping between and in HindIll fragments K and L. The 5.2-kb (3) mrna has its 5' end in the interior of the 6.9-kb (y) mrna juxtaposed to the 3' end of the 1.7-kb (y) mrna (see Fig. 1A). The 6.9-kb mrna, then, is a result of an inefficient transcription termination at the end of the "gene" for the 1.7-kb mrna. We previously showed that the promoter for the 5.2-kb mrna maps just upstream of the 5' end of this mrna and shares certain features with the promoter of another ( HSV-1 mrna, thymidine kinase (tk). The 5' end of the 1.2-kb ( mrna underlying the 3' end of the 6.9- and 5.2-kb mrnas could be generated in the same way or could result from using the 5.2-kb mrna promoter and splicing. As shown in this note, the region just upstream of the 5' end of the 1.2-kb mrna has sequence similarities to the other HSV-1 promoters previously characterized. Further, transcription of this mrna can be initiated by using a Manley uninfected cell lysate transcription system (10, 13), indicating that this mrna is under its own promoter control and lacks detectable splicing. We located the 5' end of the 1.2-kb (1) mrna to be approximately 300 bases upstream of the HindIll site at and the common 3' end of all three mrnas to be 900 bases downstream of this site (1). A high-resolution restriction map of the region in question is shown in Fig. 1B. Here, Hinfl, Sall, AvaI, HindIll, PvuII, and DdeI sites are indicated. We used two HSV-1 DNA clones for these studies, BamHI-HindIII fragment O-K (0.576 to 0.592) and HindIII-BamHI fragment L-0 (0.592 to 0.602). Details of our cloning procedures were described elsewhere (1, 8). We located the 5' end of the 1.2-kb mrna by SI nuclease digestion of hybrids between the Sall- AvaI fragment 5' end labeled at the AvaI site and infected-cell polyadenylated mrna (viral mrna). The basic details of isolation of viral mrna via the use of the Palmiter Mg't precipitation of polyribosomes has been described previously (1, 18). We also have described our methods for end labeling viral DNA fragments (9, 10, 15). In this case, BamHI-HindIII fragment O-K was digested with AvalI, 5' end labeled, and redigested with Sall, and the labeled fragment was strand separated on a nondenaturing acrylamide gel as described by Maxam and Gilbert (15). S1 nuclease digestion of hybrids gave two distinct bands (Fig. 2A); one was 370 bases long, owing to protection of the full length of the DNA by the 6.9- and 5.2-kb mrnas, and one was 250 bases long, owing to protection by the 1.2-kb mrna. The relative intensity of the two bands (Fig. 2A) provides a good measure of the abundance of the 1.2-kb mrna relative to the two larger species. We precisely located the 5' end of the mrna to be 251 bases upstream from the AvaI site (data not shown) by running the Si nuclease-protected fragment against a Maxam-Gilbert sequence ladder. The 5' end was located in the sequence TGTACT at the A residue (see sequence data below). We similarly 1123 located the 3' end of the three colinear mrnas to be approximately 100 bases downstream (3') of the rightmost DdeI site by using strandseparated DNA from HindIII-BamHI fragment L-0 (0.592 to 0.602) 3'-labeled at the DdeI site. Details of 3' end labeling have been described
2 1124 NOTES A Kpn PVuI Pvut[ Pvull Kpn X Ba Ba Sa X Bgl So H Ba looon, p , F---* /3~ ~~~00d /3~~~ ,000d 40,000d B (5') (On H en Hint iihinf So Hinf Hinf H PvuIE DDE DDE Hinf Hinf Hinf 11 H 4- Ava 1 TATA" (ATG) (tga (TGA) J. VIROL. FIG. 1. (A) Schematic localization of mrna species mapping between and on the P arrangement of the HSV-1 genome. Values are based on S1 nuclease analysis and in vitro translational studies (1). Arrows indicate direction of transcription and approximate 5' and 3' termini of mrna species. The total length of the mrnas and their relative times of appearance are shown above the arrows, and the size of the polypeptide encoded by each species is shown below the bracket. (B) High-resolution map of the HSV-1 region (0.587 to 0.602) encoding the 1,200-base P mrna, with flanking regions. Specific sites are based on results described in the text. Note (i) the location of the 5' and 3' termini of the 1,200-base mrna, (ii) the presence of the canonical TATA box, and (iii) the putative site of the initiation (ATG) and termination (TGA) codons of the 40,000-dalton polypeptide encoded by this mrna. The restriction sites used to derive the sequence found in Fig. 3 are shown; enzymes used were Hinfl (Hinf), Sall (Sa), AilaIl HindllI (H), PFill, and Ddel (DDE). AM I.. AMN. FIG. 2. (A) S1 nuclease analysis of mrna species mapping proximal to the 5' terminus of the 1,200-base 3 HSV-1 clone BamHI-HindlIl fragment O-K (0.576 to 0.592) was mrna of HSV-1. Five micrograms of the digested with AvI, 5' labeled with [32 P]ATP, redigested with Sall, and electrophoresed on a 6% acrylamide gel by the method of Maxam and Gilbert (15). The 370-base piece representing the HSV-1 region of to was further purified, after denaturation, by electrophoresis on a nondenaturing 5% acrylamide (1:50 cross-link) gel. Samples containing the resultant 370-base single strand of DNA and 10,ug of viral polyadenylated RNA were hybridized in 50 p.1 of hybridization buffer (80% formamide, 0.4 M Na+, 0.1 M HEPES [N-2-hydroxyethylpiperazine-N'-2-ethanesulfonic acid] [ph 8.0], M EDTA) for 16 h. Si nuclease analysis was done essentially by the method of Berk and Sharp as described previously (4-6. 9). After nuclease digestion, samples were
3 VOL. 43, previously (9; data not shown). The DNA fragment ranging between the second DdeI site and the BwlnHI site at was separated from a digest by using a nondenaturing 6% acrylamide fragment separation gel as described by Maxam and Gilbert (15). The precise localization of the 5' end of the 1.2-kb (a) mrna allowed us to examine the nucleotide sequence upstream of it (see below) and suggested that this mrna is unspliced, since a TATA box and a CAT box sequence could be identified. Uninfected-cell RNA polymerase II recognizes the promoter for the 1 tk gene, since this gene is expressed in biochemically transformed cells and in micro-injected amphibian oocytes (17, 24). The early (1) alkaline exonuclease of HSV-1 is similarly expressible in amphibian oocytes (19), and we suggested that the ability of uninfected-cell polymerase to recognize 3 promoters is general, since we found that the Manley polymerase recognized major 1 promoters mapping in Hindlll fragments K and L (10). The situation with the Manley system is not absolutely clear, since several workers have reported that multiple transcription initiation sites are seen when HSV- 1 BaonHI fragment P (0.298 to 0.318). which encodes tk, is used as a template. Our rather more clear-cut results suggested that we can use NOTES 1125 the uninfected system to identify early HSV-1 promoters. We used three templates in a transcription runoff experiment (10, 21) to determine whether accurate initiation at the 5' end of the 1.2-kb mrna occurred. We used a commercial system, in a 50-,ul total volume, with Qx-[32P]UTP (400 Ci/mmol; Amersham Corp.) and 4,ug of template made by appropriately digesting the BamnHI-HindIII fragment O-K (0.576 to 0.592) in the pbr322 vector. Use of template formed by digestion with BarnHI and HinidIII gave a radioactive band 370 bases long (Fig. 2B1 upper panel), a size expected from the position of the 5' end of the 1.2-kb mrna (371 bases upstream of the HindIII site; see below). This same band was seen when the template was cut with both Sall and BarnHI-HindIII suggesting that any recognition sites for polymerase are between the Sall site (120 bases upstream of the 5' end of the mrna; see below) and the 5' end of the mrna. When the template was formed by digestion of the template with AalI, BaitHI, and Hinidlll, a product 250 bases long was seen. Several other size bands were also seen, notably one about 150 bases long and one 600 bases long; these sizes were unaffected by the enzymes used to cut the template, and we suggest they were due to the presence of promoters on the pbr portion of the suspended in 30,ul of 98% formamide-1( mm HEPES (ph 8.0). heat denatured at 95 C for 2 min. and electrophoresed in a 5% acrylamide (1:40 cross-link). 8 M urea gel. The results of such an analysis are shown in the column marked Sal-Ava*. Size standard (S.S.) was prepared by digesting 5 p.g of pbr322 with Hinfl and EcoRI restriction endonucleases and subsequent 5' labeling with 132P]ATP. Marker DNA waxs also denatured before electrophoresis. Numbers represent sizes of DNA fragments in nucleotides. Exposure was overnight without intensifying screens. (B) In vitro transcription, using the HSV-1 clone Ba,n1HI-HindlIl fragment O-K as a template. Four micrograms of the HSV-1 clone BainHI-Hindlll fragment O-K was suitably digested and used as a template in transcriptional runoff experiments. using a Manley HeLa cell lysate system (13). Incubations in a 50-,ul total volume were performed as described previously (10). RNA was isolated and analyzed by electrophoresis at 600 V for 2.5 h on denaturing 5% acrylamide (40:1). 8 M urea gels (1.8 by 40 by 200 mm). Column 1, Template DNA digested with BatinHI and Hindlll: column 2. template DNA digested with BamnHI. Sall, and Hindlll; column 3, template DNA digested with BamfHll Ailail. and Hindlll. Size markers are based on comigration with 5' end-labeled fragments produced by Hinfl digestion of pbr322. The 250- and 370-base RNA transcription products are indicated. Exposure was for 5 days with intensifying screens (22). (C) Analysis of HSV-1 DNA sequence proximal to the 5' end of the 1.2-kb l mrna. Ten micrograms of the HSV-1 clone BatiHI-HindIll fragment O-K was digested with Sall and Hindlll. The 490-base DNA fragment was 5' end labeled, strand separated. and sequenced from both ends as descried by Maxam and Gilbert (15). Chains interrupted at G, G+A, C+T, and C residues were fractionated on an 8% acrylamide gel (80 by 30) cm by 0.5 mm). The sequence shown represents a portion of the strand which was labeled at the Still site (t).588) and proceeds through the 5' end of the 1.2-kb mrna toward the Hindlll site. Note the presence of the ATATAA sequence 29 bases 5' to the transcription initiation site of the mrna. (D) Hybridization of unlabeled in vitro transcription product to the coding strand of HSV-1 DNA. In vitro transcriptions were performed as described above, except equimolar concentrations (0.5 mm) of all four base triphosphates were used, and no labeled nucleotide was present. Template was 4 FLg of cloned HSV-1 DNA from BamiiHl-Hindlll fragment O-K digested with BainHI and Hindlll. Thirty microliters of dilution buffer (10) was substituted for the HeLa cell lysate in the ''no enzyme" reactions. Transcription products were purified from template DNA as described in the text. After extensive dialysis, each set of transcription products was hybridized with 5-pLg equivalents of the singlestranded, 5'-labeled, 370-base HSV-1 (Avial-Sall) DNA fragment described in the legend to Fig. 2A. Hybridizations were carried out in 10 1.d of aqueous buffer (0.2 M Na+-0.1 M HEPES IpH M EDTA) in sealed capillary tubes at 65 C for 16 h. Reactions were quenched by dilution into 400 [LI of SI buffer. and SI nuclease digestion was performed (1, 9). Products of the digestion were denatured and analyzed on denaturing gels as shown in panel B. 5' End-labeled fragments of Hinfl-digested pbr322 were used as size standards. Exposure was for 4 days with intensifying screens (22).
4 1126 NOTES J. VIROL. template, such as those which have been described previously (10). We confirmed the synthesis of an RNA product with a 5' end very near the location of the 1.2-kb mrna by using Si nuclease analysis of in vitro transcription products (Fig. 2D). The Manley polymerase lysate was incubated with BamHI-HindIII-digested template and unlabeled base triphosphates (0.5 mm each); a second incubation with no enzymes was carried out as a control. Radioactive carrier RNA (100,000 32p cpm; 10 [ig) was added to the products of reaction, and the RNA was separated from the template by 36 h of centrifugation in a 1-ml CaCl gradient of a starting density of 1.6 g/ cm3. Centrifugation was carried out in an SW60 Ti rotor at 50,000 rpm at 17 C. These conditions were found adequate to remove essentially all of the template DNA from the RNA pellet in separate experiments. The pelleted RNA was redissolved in water, dialyzed versus 0.1 M NaCl-0.01 M Tris (ph 7.4) M EDTA, and then hybridized under aqueous conditions with strand-separated HSV-1 DNA 5' end labeled at the AvaI site as described above and in the legend to Fig. 2D. Hybridization for 18 h yielded several Si-resistant bands, caused by the structure of the radiolabeled DNA probe, as well as some undigested material, as shown by the bands of 220, 240, and 370 bases in both the control and experimental tracks. However, a band of 250 bases was consistently seen in the enzyme incubation but was missing in the control. We concluded that this was owing to synthesis of RNA initiating 250 bases upstream of the AvaI site (1). The in vitro transcription data indicated that the 1.2-kb mrna has a functional promoter within 120 bases upstream of its 5' end. We carried out Maxam-Gilbert DNA sequence analysis of the DNA encoding this mrna. We sequenced both strands of DNA end labeled at the restriction sites indicated in Fig. lb. The methodology in which strand-separated DNA was used was exactly as described by Maxam and Gilbert and previously by us (10, 15). An example of the sequence data proceeding downstream from the Sall site at is shown in Fig. 2C. The sequence is of the same sense as the mrna, and the sequence ATATAA starting 29 bases upstream of the 5' end of the mrna is indicated. The full mrna sense sequence of the DNA beginning 313 bases upstream of the 5' end of the 1.2-kb mrna and going 1,244 bases downstream is shown in Fig. 3. The 5' end of the mrna is, as noted, 29 bases downstream from a putative TATA box sequence. The sequence TCAC is seen 90 bases upstream. In the 5.2-kb I mrna, the sequence ACATC is seen at -90 bases (10), and in tk, the sequence TCATT is seen at -88 bases (16, 23). We suggest that this is a CAT box sequence (2, 3, 14). More significantly, the region between -110 and -97 is mainly A's and C's, which was also seen in the two other 3 mrnas characterized, but not with 13y or y mrnas. Further comparative studies in progress will indicate how significant and general this finding is. The only translation start signal in the mrna sequence is seen at position 151. The sequence around this translation start (GCCATGG) is a favored one for eucaryotic translation starts (12). The reading frame defined by this start signal is open for 1,017 bases, defining a 339 amino acid polypeptide whose predicted composition would give a molecular weight of 37,970, a value in good agreement with our in vitro translation value of 40,000. The 3' end of the 6.9-, 5.2-, and 1.2-kb mrnas is very near the sequence ATAATAAA. The sequence ATAAAA is found 69 bases downstream of this sequence (data not shown), and the sequence TAATTTTATT is downstream another 60 bases from that (data not shown). Since this region encodes the 3' end of mrnas on both strands, we must regard it as an efficient polyadenylation region. The sequence AATAAAA has been implicated as the 3' stop signal of both HSV-1 tk (16, 23) and other eucaryotic mrnas (20). Whether the departure from this nominal sequence seen in the present case is significant is as yet unknown, but ATrich regions are seen in the area encoding the 3' ends of several HSV-1 mrnas around the HindlIl site at (unpublished data), so the general character of the polyadenylation signal seems well established. In light of the fact that a similar size polypeptide immunologically cross-reacting with a 140,000-dalton HSV-2 polypeptide is encoded by the analogous region of HSV-2 and can be detected in some HSV-2-transformed cells (11), it is interesting to ask what the relationship is between the HSV-2 polypeptides and the one encoded by HSV-1. The following facts are clear: in HSV-1, there is no open reading frame upstream of the 5' end of the 1.2-kb mrna that would reasonably allow an in-phase fusion protein to be synthesized from the 5.2-kb mrna. The frame that is used as the phase for the reading of the 1.2-kb mrna is terminated many times upstream of the translation initiator in the mrna. Similar results have been reported by Clements and McLauchlan (7). Another area of difference between HSV-1 and HSV-2 is that the 1.2-kb mrna in HSV-2 appears to be a major mrna (11), whereas the protein analogous to the 140,000-dalton product of the major 5.2-kb a HSV-1 mrna is not readily seen, and then only late. Therefore, some interesting differences in
5 VOL. 43, 1982 NOTES TAAAGGAACTGGA ACGCACGTTTAGGGGAAGCGCCTCCTGGAG GTGATGAACAGCTCGACGCCAAGCAGGGTT CCGTGGCGCAGGCGCTCCCGTGCCTAGAGC CCACCCACCCCCTCCGGCGATTCAAGACCG CGTTTGACTACGACCAGAAGTTGCTGATCG ACCTGTGTGCGGACCGCGCCCCCTACGTCG TCGACCATAGCCAATCCATGACCCTGTATG TCACGGAGAAGGCGGACGGGACCCTCCCAG CCTCCACCCTGGTCCGCCTTCTGGTCCACG CATATAAGCGCGGACTAAAAACAGGGATGT ACTACTGCAAGGTTCCGAAGGCGACCAACA GCGGGGTCTTTGGCGGCGACGACCCAATTG TCTGCACGGCTGCGCGCTGTGACCGACAAA CCCCCTCCGCGCCAGGCCCGCCGCCATCGT CGTCGCCGTCCCACGCGCTCCCCCGCTGCC ATG GAT TCC GCG GCC CCA GCC TCT CCC CCG CTC TGG GCC CAT ACG GGC CAT AGC GCG ACG GCG GAC CTA GCG ATC CAG ATT CCA AAG TGC +241 CCC +310 CTT +379 GGC +448 CTG +517 GAA +586 CGC +655 GTG +724 CTC +793 CCT GAC CCC 315 AAC CGC GAG CTC GGC GGC 525 GTC GCA GAG TAC 660 CGG GAA GTT TTG CAT CAG 255 GAG AGG TAC TTC 330 TGG CTG GAA ACC 390 AGC TTT TAC CGC 465 CTC TCC GGC CTG CAC TCG CGC GTG 600 GTG GCC GGC ACC 675 TGC GCC TCC GTT 735 CCG CCA TCG CCC 810 CCG GGA CGA GGC TAC ACC TCC CAG TGT CCC GAC ATT AAC CAC CTG CGC TCC CTC AGC ATC GAG CTT GTT TTC GTG GGG GAC GAG GAG GAC GTC TCC AAG CTT TCC GAG TTC CTC TTC GCT TTC CTG TCG GCC GCC GAC GAC CTG GTT ACG GAA AAC TTT GAG CAG AAG GAC ATT CTC CAC TAC TAC GTG GAG CAG GAA TGC ATC TAC AAC ATC ATC CAG CTG GTG CTT TTC CAC AAC AAC GAC CAG GCG CGC ATC AAC CAC CCG GCC ATC CGC GCC CAG GTG GAC TGG CTG GAA GCG CGG CCG GAA AAG CAT TCT CAT GAT CCT CAT CGA GGG CAT CTT TTT TGC CGC CTA CCT TCG CAC CAA CAA CCT TCT GCG GGT CAC CTG CCG GTC AAA CGA CGT GCA CAC GAC GGC CTC GTG TTA CAT CTA CAA CAA CTA CCT AGG CGG OCA CGC CAA GCC CCC GCC CGA CCG CGT GTA CGG GCT GTC CGC CAA GCG GTC GAG ATC GAG ATC GGA TTT ATC CGA TCC CAG GCa CCG ACG GAC AGC CAT ATC CTG AGC CCG GCa GCG CTG GCG GCC ATC GAA AAC TAC ' 1068 GTG CGA TTC AGC GCG GAT CGC CTG TTG GGC CTT ATC CAC ATG AAG CCA CTG TTT TCC GCC CCA CCC CCC GAC GCC AGC TTT CCG CTG AGC CTC ATa TCC ACC GAC AAA CAC ACC AAT TTT TTC GAG TGT CGC AGC ACC TCC TAC GCC GGG GCG GTC GTC AAC GAT CTG TGA GTGTCGCGGCGCGCTTCTACCCGTGTTTGC CCATAATAAACCTCT GAACCAAACTTTGGGTCTCATTGTGATTC FIG. 3. Nucleotide sequence of the noncoding strand of HSV-1 DNA encoding the 1,200-base 3 mrna and its 5' and 3' flanking regions. DNA sequence analysis was performed by the procedure of Maxam and Gilbert (15). Cloned DNA was end labeled with [32P]ATP, and isolated strands of DNA were then sequenced, using gels (30 by 80 cm by 0.5 mm). All sequences were done at least in duplicate, and both DNA strands were sequenced. The sequence from 313 bases upstream of the 5' end of the mrna to 1,244 bases downstream is shown for the mrna sense strand. As discussed in the text, the putative CAT box signal is at position -90, the AC-rich region is at -112 through -104, and the TATA box is between -29 and -24. The translation termination codon TAA is seen at -313 and defines reading frame 1. Other terminator sequences in this frame are seen at positions 201, 285, 312, 471, 693, and 855. Translation termination codons in frame 2 are seen at positions -290, -216, -156, -102, and -15. The frame is opened with the AUG (ATG) codon at position 151 and closed again at positions 1168, 1204, and This is the reading frame for the encoded polypeptide whose molecular weight is 37,970, based on its calculated amino acid composition. The third potential reading frame has translation terminators at positions -269, -266, -176, -113, -26, +80, +218, +965, +1040, and The restriction sites indicated in Fig. 1B are as follows: Hinfl sites (GANTC) are at positions -192, +154, +228, +1004, and +1240; the Sall site (GTCGAC) is at -121, the AvaI site (CPyCGPuG) is at position +247; the Hindlll site (AAGCTT) is at position + 367; the PvuII site (CAGCTG) is at position + 550; and Ddel sites (CTNAG) are at positions +964 and
6 1128 NOTES temporal control may be operating in the two infectious cycles. This work was supported by Public Health Service grant CA11861 from the National Institutes of Health. We thank R. Costa and L. Hall for assistance and discussion and J. Wagner for editing and preparing this manuscript. LITERATURE CITED 1. Anderson, K. P., R. J. Frink, G. B. Devi, B. H. Gaylord, R. H. Costa, and E. K. Wagner Detailed charaicterization of the mrna mapping in the HioidlIl fragment K region of the herpes simplex virus type genome. J. Virol. 37: Benoist, C., and P. Chambon In vivo sequence requirements of the SV40 early promoter region. NatuLre (London) 290: Benoist, C., K. O'Hare, R. Breathnach, and P. Chambon The ovalbumin gene-sequence of putative control regions. Nucleic Acids Res. 8: Berk, A., and P. Sharp Sizing and matpping of early adenovirus mrnas by gel electrophoresis of S1 endonuclease-digested hybrids. Cell 12: Berk, A., and P. Sharp Spliced early mrnas of simian virus 40. Proc. Natl. Acad. Sci. U.S.A. 75: Berk, A., and P. Sharp Structure ot the adenoviruls 2 early mrnas. Cell 14: Clements, J., and J. McLauchlan Control regions involved in the expression of two 3' co-terminal eairly mrnas. p. 57. Ini A. S. Kaplan, M. Lat Placa. F. Raipp, and B. Roizman (ed.). International workshop on herpesviruses. Esculapio Publishing Co., Bologna. Italy. 8. Costa, R. H., B. G. Devi, K. P. Anderson, B. H. Gaylord, and E. K. Wagner Characterization of a maijor late herpes simplex virus type 1 mrna. J. Virol. 38: Frink, R. J., K. P. Anderson, and E. K. Wagner Herpes simplex virus type I Hio1dIlI fragment L encodes spliced and complementary mrna species. J. Virol. 39: Frink, R., K. Draper, and E. Wagner Uninfected cell polymerase efficiently transcribes early but not late herpes simplex virus type I mrna. Proc. Natl. Acad. Sci. U.S.A. 78: Galloway, D. A., L. C. Goldstein, and J. B. Lewis Identification of proteins encoded by a fragment of herpes simplex virus type 2 DNA that has transforming activity. J. Virol. 42: J. VIROL. 1la.Hall, L. M., K. G. Draper, R. J. Frink, R. H. Costa, and E. K. Wagner Herpes simplex virus mrna species mapping in EcoRI fragment I. J. Virol. 43: Kozak, M Possible role of flanking nucleotides in recognition of the AUG initiator codon by eukar-yotic ribosomes. Nucleic Acids Res. 9: Manley, J., A. Fire, A. Cano, P. Sharp, and M. Gefter DNA-dependent transcription of adenovirus genes in a soluble whole cell extract. Proc. Nattl. Acad. Sci. U.S.A. 77: Mathis, D., and P. Chambon The SV40 early region TATA box is required for accurate in vitro initiation of transcription. Nature (London) 290: Maxam, A., and W. Gilbert Sequencing end-labeled DNA with base-specific chemical cleavages. Methods Enzymol. 65: McKnight, S The nucleotide sequence and trainscript map of the herpes simplex viruls thymidine kinase gene. Nucleic Acids Res. 8: McKnight, S., and E. Gavis Expression of the herpes thymidine kinase gene in Xenopus aiievis oocytes: an assay for the study of deletion mutants constructed in vitro. Nucleic Acids Res. 8: Palmiter, R. D Mg'- precipitation of ribonucleoprotein complexes. Expedient techniques ftr the isolation of undegraded polysomes and messenger ribonucleic aicid. Biochemistry 13: Preston, C Synthesis of HSV exonuclease in Xenopus laevis oocytes, p. 58. In A. S. Kaplain. M. La Plaicat, F. Rapp, and B. Roizman (ed.), International workshop on herpesviruses. Esculapio Publishing Co., Bologna. Italy. 20. Proudfoot, N., and G. Brownlee ' Non-coding region sequences in eucaryotic mrna. Natture (London) 263: Rio, D., A. Robbins, R. Myers, and R. Tjian. 1980(. Regulation of SV40) early transcription in vitro bv al purified tumor antigen. Proc. Natl. Acad. Sci. U.S.A. 77: Swanstrom, R., and P. Shank X-ray intensifying screens greatly enhance the detection by autoradiography of the radioactive isotopes 32P and 1251 Anal. Biochem. 86: Wagner, M., J. Sharp, and W. Summers Nucleotide sequence of the thymidine kinase gene of herpes simplex virus type 1. Proc. NatI. Acad. Sci. U.S.A. 78: Wigler, M., S. Silverstein, L.-S. Lee, A. Pellicer, Y.-C. Cheng, and R. Axel Transfer of purified herpes virus thymidine kinase gene to cultured motuse cells. Cell 11:
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR
 Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR 1 The problem We wish to clone a yet unknown gene from a known
Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR 1 The problem We wish to clone a yet unknown gene from a known
Supporting information for Biochemistry, 1995, 34(34), , DOI: /bi00034a013
 Supporting information for Biochemistry, 1995, 34(34), 10807 10815, DOI: 10.1021/bi00034a013 LESNIK 10807-1081 Terms & Conditions Electronic Supporting Information files are available without a subscription
Supporting information for Biochemistry, 1995, 34(34), 10807 10815, DOI: 10.1021/bi00034a013 LESNIK 10807-1081 Terms & Conditions Electronic Supporting Information files are available without a subscription
Supplementary. Table 1: Oligonucleotides and Plasmids. complementary to positions from 77 of the SRα '- GCT CTA GAG AAC TTG AAG TAC AGA CTG C
 Supplementary Table 1: Oligonucleotides and Plasmids 913954 5'- GCT CTA GAG AAC TTG AAG TAC AGA CTG C 913955 5'- CCC AAG CTT ACA GTG TGG CCA TTC TGC TG 223396 5'- CGA CGC GTA CAG TGT GGC CAT TCT GCT G
Supplementary Table 1: Oligonucleotides and Plasmids 913954 5'- GCT CTA GAG AAC TTG AAG TAC AGA CTG C 913955 5'- CCC AAG CTT ACA GTG TGG CCA TTC TGC TG 223396 5'- CGA CGC GTA CAG TGT GGC CAT TCT GCT G
Materials Protein synthesis kit. This kit consists of 24 amino acids, 24 transfer RNAs, four messenger RNAs and one ribosome (see below).
 Protein Synthesis Instructions The purpose of today s lab is to: Understand how a cell manufactures proteins from amino acids, using information stored in the genetic code. Assemble models of four very
Protein Synthesis Instructions The purpose of today s lab is to: Understand how a cell manufactures proteins from amino acids, using information stored in the genetic code. Assemble models of four very
ORFs and genes. Please sit in row K or forward
 ORFs and genes Please sit in row K or forward https://www.flickr.com/photos/teseum/3231682806/in/photostream/ Question: why do some strains of Vibrio cause cholera and others don t? Methods Mechanisms
ORFs and genes Please sit in row K or forward https://www.flickr.com/photos/teseum/3231682806/in/photostream/ Question: why do some strains of Vibrio cause cholera and others don t? Methods Mechanisms
Disease and selection in the human genome 3
 Disease and selection in the human genome 3 Ka/Ks revisited Please sit in row K or forward RBFD: human populations, adaptation and immunity Neandertal Museum, Mettman Germany Sequence genome Measure expression
Disease and selection in the human genome 3 Ka/Ks revisited Please sit in row K or forward RBFD: human populations, adaptation and immunity Neandertal Museum, Mettman Germany Sequence genome Measure expression
RPA-AB RPA-C Supplemental Figure S1: SDS-PAGE stained with Coomassie Blue after protein purification.
 RPA-AB RPA-C (a) (b) (c) (d) (e) (f) Supplemental Figure S: SDS-PAGE stained with Coomassie Blue after protein purification. (a) RPA; (b) RPA-AB; (c) RPA-CDE; (d) RPA-CDE core; (e) RPA-DE; and (f) RPA-C
RPA-AB RPA-C (a) (b) (c) (d) (e) (f) Supplemental Figure S: SDS-PAGE stained with Coomassie Blue after protein purification. (a) RPA; (b) RPA-AB; (c) RPA-CDE; (d) RPA-CDE core; (e) RPA-DE; and (f) RPA-C
Arabidopsis actin depolymerizing factor AtADF4 mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB
 Arabidopsis actin depolymerizing factor mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB Files in this Data Supplement: Supplemental Table S1 Supplemental Table
Arabidopsis actin depolymerizing factor mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB Files in this Data Supplement: Supplemental Table S1 Supplemental Table
PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells
 Supplementary Information for: PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells Ju Hye Jang 1, Hyun Kim 2, Mi Jung Jang 2, Ju Hyun Cho 1,2,* 1 Research Institute
Supplementary Information for: PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells Ju Hye Jang 1, Hyun Kim 2, Mi Jung Jang 2, Ju Hyun Cho 1,2,* 1 Research Institute
Supplemental Data Supplemental Figure 1.
 Supplemental Data Supplemental Figure 1. Silique arrangement in the wild-type, jhs, and complemented lines. Wild-type (WT) (A), the jhs1 mutant (B,C), and the jhs1 mutant complemented with JHS1 (Com) (D)
Supplemental Data Supplemental Figure 1. Silique arrangement in the wild-type, jhs, and complemented lines. Wild-type (WT) (A), the jhs1 mutant (B,C), and the jhs1 mutant complemented with JHS1 (Com) (D)
Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC
 Supplementary Appendixes Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC ACG TAG CTC CGG CTG GA-3 for vimentin, /5AmMC6/TCC CTC GCG CGT GGC TTC CGC
Supplementary Appendixes Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC ACG TAG CTC CGG CTG GA-3 for vimentin, /5AmMC6/TCC CTC GCG CGT GGC TTC CGC
Hes6. PPARα. PPARγ HNF4 CD36
 SUPPLEMENTARY INFORMATION Supplementary Table Positions and Sequences of ChIP primers -63 AGGTCACTGCCA -79 AGGTCTGCTGTG Hes6-0067 GGGCAaAGTTCA ACOT -395 GGGGCAgAGTTCA PPARα -309 GGCTCAaAGTTCAaGTTCA CPTa
SUPPLEMENTARY INFORMATION Supplementary Table Positions and Sequences of ChIP primers -63 AGGTCACTGCCA -79 AGGTCTGCTGTG Hes6-0067 GGGCAaAGTTCA ACOT -395 GGGGCAgAGTTCA PPARα -309 GGCTCAaAGTTCAaGTTCA CPTa
Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis
 1 2 3 4 5 6 7 8 9 10 11 12 Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis Information Research). Exons
1 2 3 4 5 6 7 8 9 10 11 12 Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis Information Research). Exons
Y-chromosomal haplogroup typing Using SBE reaction
 Schematic of multiplex PCR followed by SBE reaction Multiplex PCR Exo SAP purification SBE reaction 5 A 3 ddatp ddgtp 3 T 5 A G 3 T 5 3 5 G C 5 3 3 C 5 ddttp ddctp 5 T 3 T C 3 A 5 3 A 5 5 C 3 3 G 5 3 G
Schematic of multiplex PCR followed by SBE reaction Multiplex PCR Exo SAP purification SBE reaction 5 A 3 ddatp ddgtp 3 T 5 A G 3 T 5 3 5 G C 5 3 3 C 5 ddttp ddctp 5 T 3 T C 3 A 5 3 A 5 5 C 3 3 G 5 3 G
Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH).
 Bisulfite Treatment of DNA Dilute DNA sample to 2µg DNA in 50µl ddh 2 O. Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH). Incubate in a 37ºC water bath for 30 minutes. To 55µl samples
Bisulfite Treatment of DNA Dilute DNA sample to 2µg DNA in 50µl ddh 2 O. Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH). Incubate in a 37ºC water bath for 30 minutes. To 55µl samples
Supplemental Table 1. Mutant ADAMTS3 alleles detected in HEK293T clone 4C2. WT CCTGTCACTTTGGTTGATAGC MVLLSLWLIAAALVEVR
 Supplemental Dataset Supplemental Table 1. Mutant ADAMTS3 alleles detected in HEK293T clone 4C2. DNA sequence Amino acid sequence WT CCTGTCACTTTGGTTGATAGC MVLLSLWLIAAALVEVR Allele 1 CCTGTC------------------GATAGC
Supplemental Dataset Supplemental Table 1. Mutant ADAMTS3 alleles detected in HEK293T clone 4C2. DNA sequence Amino acid sequence WT CCTGTCACTTTGGTTGATAGC MVLLSLWLIAAALVEVR Allele 1 CCTGTC------------------GATAGC
Cat. # Product Size DS130 DynaExpress TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1
 Product Name: Kit Component TA PCR Cloning Kit (ptakn-2) Cat. # Product Size DS130 TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1 2 Ligation Buffer
Product Name: Kit Component TA PCR Cloning Kit (ptakn-2) Cat. # Product Size DS130 TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1 2 Ligation Buffer
PCR analysis was performed to show the presence and the integrity of the var1csa and var-
 Supplementary information: Methods: Table S1: Primer Name Nucleotide sequence (5-3 ) DBL3-F tcc ccg cgg agt gaa aca tca tgt gac tg DBL3-R gac tag ttt ctt tca ata aat cac tcg c DBL5-F cgc cct agg tgc ttc
Supplementary information: Methods: Table S1: Primer Name Nucleotide sequence (5-3 ) DBL3-F tcc ccg cgg agt gaa aca tca tgt gac tg DBL3-R gac tag ttt ctt tca ata aat cac tcg c DBL5-F cgc cct agg tgc ttc
Supplementary Figure 1A A404 Cells +/- Retinoic Acid
 Supplementary Figure 1A A44 Cells +/- Retinoic Acid 1 1 H3 Lys4 di-methylation SM-actin VEC cfos (-) RA (+) RA 14 1 1 8 6 4 H3 Lys79 di-methylation SM-actin VEC cfos (-) RA (+) RA Supplementary Figure
Supplementary Figure 1A A44 Cells +/- Retinoic Acid 1 1 H3 Lys4 di-methylation SM-actin VEC cfos (-) RA (+) RA 14 1 1 8 6 4 H3 Lys79 di-methylation SM-actin VEC cfos (-) RA (+) RA Supplementary Figure
Supporting Online Information
 Supporting Online Information Isolation of Human Genomic DNA Sequences with Expanded Nucleobase Selectivity Preeti Rathi, Sara Maurer, Grzegorz Kubik and Daniel Summerer* Department of Chemistry and Chemical
Supporting Online Information Isolation of Human Genomic DNA Sequences with Expanded Nucleobase Selectivity Preeti Rathi, Sara Maurer, Grzegorz Kubik and Daniel Summerer* Department of Chemistry and Chemical
SAY IT WITH DNA: Protein Synthesis Activity by Larry Flammer
 TEACHER S GUIDE SAY IT WITH DNA: Protein Synthesis Activity by Larry Flammer SYNOPSIS This activity uses the metaphor of decoding a secret message for the Protein Synthesis process. Students teach themselves
TEACHER S GUIDE SAY IT WITH DNA: Protein Synthesis Activity by Larry Flammer SYNOPSIS This activity uses the metaphor of decoding a secret message for the Protein Synthesis process. Students teach themselves
NAME:... MODEL ANSWER... STUDENT NUMBER:... Maximum marks: 50. Internal Examiner: Hugh Murrell, Computer Science, UKZN
 COMP710, Bioinformatics with Julia, Test One, Thursday the 20 th of April, 2017, 09h30-11h30 1 NAME:...... MODEL ANSWER... STUDENT NUMBER:...... Maximum marks: 50 Internal Examiner: Hugh Murrell, Computer
COMP710, Bioinformatics with Julia, Test One, Thursday the 20 th of April, 2017, 09h30-11h30 1 NAME:...... MODEL ANSWER... STUDENT NUMBER:...... Maximum marks: 50 Internal Examiner: Hugh Murrell, Computer
Supplemental Data. mir156-regulated SPL Transcription. Factors Define an Endogenous Flowering. Pathway in Arabidopsis thaliana
 Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
Lecture 11: Gene Prediction
 Lecture 11: Gene Prediction Study Chapter 6.11-6.14 1 Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and end positions of genes in a genome Where are
Lecture 11: Gene Prediction Study Chapter 6.11-6.14 1 Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and end positions of genes in a genome Where are
Dierks Supplementary Fig. S1
 Dierks Supplementary Fig. S1 ITK SYK PH TH K42R wt K42R (kinase deficient) R29C E42K Y323F R29C E42K Y323F (reduced phospholipid binding) (enhanced phospholipid binding) (reduced Cbl binding) E42K Y323F
Dierks Supplementary Fig. S1 ITK SYK PH TH K42R wt K42R (kinase deficient) R29C E42K Y323F R29C E42K Y323F (reduced phospholipid binding) (enhanced phospholipid binding) (reduced Cbl binding) E42K Y323F
Multiplexing Genome-scale Engineering
 Multiplexing Genome-scale Engineering Harris Wang, Ph.D. Department of Systems Biology Department of Pathology & Cell Biology http://wanglab.c2b2.columbia.edu Rise of Genomics An Expanding Toolbox Esvelt
Multiplexing Genome-scale Engineering Harris Wang, Ph.D. Department of Systems Biology Department of Pathology & Cell Biology http://wanglab.c2b2.columbia.edu Rise of Genomics An Expanding Toolbox Esvelt
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/10/494/eaan6284/dc1 Supplementary Materials for Activation of master virulence regulator PhoP in acidic ph requires the Salmonella-specific protein UgtL Jeongjoon
www.sciencesignaling.org/cgi/content/full/10/494/eaan6284/dc1 Supplementary Materials for Activation of master virulence regulator PhoP in acidic ph requires the Salmonella-specific protein UgtL Jeongjoon
2
 1 2 3 4 5 6 7 Supplemental Table 1. Magnaporthe oryzae strains generated in this study. Strain background Genotype Strain name Description Guy-11 H1:RFP H1:RFP Strain expressing Histone H1- encoding gene
1 2 3 4 5 6 7 Supplemental Table 1. Magnaporthe oryzae strains generated in this study. Strain background Genotype Strain name Description Guy-11 H1:RFP H1:RFP Strain expressing Histone H1- encoding gene
Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were
 1 Supplemental methods 2 3 4 5 6 7 8 9 1 11 12 13 14 15 16 17 18 19 21 22 23 Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were monitored by quantitative reverse-transcription
1 Supplemental methods 2 3 4 5 6 7 8 9 1 11 12 13 14 15 16 17 18 19 21 22 23 Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were monitored by quantitative reverse-transcription
Lecture 19A. DNA computing
 Lecture 19A. DNA computing What exactly is DNA (deoxyribonucleic acid)? DNA is the material that contains codes for the many physical characteristics of every living creature. Your cells use different
Lecture 19A. DNA computing What exactly is DNA (deoxyribonucleic acid)? DNA is the material that contains codes for the many physical characteristics of every living creature. Your cells use different
ΔPDD1 x ΔPDD1. ΔPDD1 x wild type. 70 kd Pdd1. Pdd3
 Supplemental Fig. S1 ΔPDD1 x wild type ΔPDD1 x ΔPDD1 70 kd Pdd1 50 kd 37 kd Pdd3 Supplemental Fig. S1. ΔPDD1 strains express no detectable Pdd1 protein. Western blot analysis of whole-protein extracts
Supplemental Fig. S1 ΔPDD1 x wild type ΔPDD1 x ΔPDD1 70 kd Pdd1 50 kd 37 kd Pdd3 Supplemental Fig. S1. ΔPDD1 strains express no detectable Pdd1 protein. Western blot analysis of whole-protein extracts
strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular
![strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular](/thumbs/72/66361212.jpg) Additional file 2 Identification of AOX1 in P. pastoris GS115 with a Mut s phenotype Results and Discussion The HBsAg producing strain was originally identified as a Mut s (methanol utilization slow) strain
Additional file 2 Identification of AOX1 in P. pastoris GS115 with a Mut s phenotype Results and Discussion The HBsAg producing strain was originally identified as a Mut s (methanol utilization slow) strain
DNA sentences. How are proteins coded for by DNA? Materials. Teacher instructions. Student instructions. Reflection
 DNA sentences How are proteins coded for by DNA? Deoxyribonucleic acid (DNA) is the molecule of life. DNA is one of the most recognizable nucleic acids, a double-stranded helix. The process by which DNA
DNA sentences How are proteins coded for by DNA? Deoxyribonucleic acid (DNA) is the molecule of life. DNA is one of the most recognizable nucleic acids, a double-stranded helix. The process by which DNA
Supplementary Information. Construction of Lasso Peptide Fusion Proteins
 Supplementary Information Construction of Lasso Peptide Fusion Proteins Chuhan Zong 1, Mikhail O. Maksimov 2, A. James Link 2,3 * Departments of 1 Chemistry, 2 Chemical and Biological Engineering, and
Supplementary Information Construction of Lasso Peptide Fusion Proteins Chuhan Zong 1, Mikhail O. Maksimov 2, A. James Link 2,3 * Departments of 1 Chemistry, 2 Chemical and Biological Engineering, and
Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of
 Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of the cell line) were immunostained for HA, acetylated
Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of the cell line) were immunostained for HA, acetylated
Supplementary Figures
 Supplementary Figures Supplementary Fig. 1 Characterization of GSCs. a. Immunostaining of primary GSC spheres from GSC lines. Nestin (neural progenitor marker, red), TLX (green). Merged images of nestin,
Supplementary Figures Supplementary Fig. 1 Characterization of GSCs. a. Immunostaining of primary GSC spheres from GSC lines. Nestin (neural progenitor marker, red), TLX (green). Merged images of nestin,
Supporting Information. Copyright Wiley-VCH Verlag GmbH & Co. KGaA, Weinheim, 2006
 Supporting Information Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Supporting Information for Expanding the Genetic
Supporting Information Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Supporting Information for Expanding the Genetic
Project 07/111 Final Report October 31, Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines
 Project 07/111 Final Report October 31, 2007. Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines Project Leader: Dr Douglas C. Hodgins (519-824-4120 Ex 54758, fax 519-824-5930)
Project 07/111 Final Report October 31, 2007. Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines Project Leader: Dr Douglas C. Hodgins (519-824-4120 Ex 54758, fax 519-824-5930)
Gene synthesis by circular assembly amplification
 Gene synthesis by circular assembly amplification Duhee Bang & George M Church Supplementary figures and text: Supplementary Figure 1. Dpo4 gene (1.05kb) construction by various methods. Supplementary
Gene synthesis by circular assembly amplification Duhee Bang & George M Church Supplementary figures and text: Supplementary Figure 1. Dpo4 gene (1.05kb) construction by various methods. Supplementary
MacBlunt PCR Cloning Kit Manual
 MacBlunt PCR Cloning Kit Manual Shipping and Storage MacBlunt PCR Cloning Kits are shipped on dry ice. Each kit contains a box with cloning reagents and an attached bag with Eco-Blue Competent Cells (optional).
MacBlunt PCR Cloning Kit Manual Shipping and Storage MacBlunt PCR Cloning Kits are shipped on dry ice. Each kit contains a box with cloning reagents and an attached bag with Eco-Blue Competent Cells (optional).
Table S1. Bacterial strains (Related to Results and Experimental Procedures)
 Table S1. Bacterial strains (Related to Results and Experimental Procedures) Strain number Relevant genotype Source or reference 1045 AB1157 Graham Walker (Donnelly and Walker, 1989) 2458 3084 (MG1655)
Table S1. Bacterial strains (Related to Results and Experimental Procedures) Strain number Relevant genotype Source or reference 1045 AB1157 Graham Walker (Donnelly and Walker, 1989) 2458 3084 (MG1655)
Supporting Information
 Supporting Information Transfection of DNA Cages into Mammalian Cells Email: a.turberfield@physics.ox.ac.uk Table of Contents Supporting Figure 1 DNA tetrahedra used in transfection experiments 2 Supporting
Supporting Information Transfection of DNA Cages into Mammalian Cells Email: a.turberfield@physics.ox.ac.uk Table of Contents Supporting Figure 1 DNA tetrahedra used in transfection experiments 2 Supporting
Chapter 13 Chromatin Structure and its Effects on Transcription
 Chapter 13 Chromatin Structure and its Effects on Transcription Students must be positive that they understand standard PCR. There is a resource on the web for this purpose. Warn them before this class.
Chapter 13 Chromatin Structure and its Effects on Transcription Students must be positive that they understand standard PCR. There is a resource on the web for this purpose. Warn them before this class.
Legends for supplementary figures 1-3
 High throughput resistance profiling of Plasmodium falciparum infections based on custom dual indexing and Illumina next generation sequencing-technology Sidsel Nag 1,2 *, Marlene D. Dalgaard 3, Poul-Erik
High throughput resistance profiling of Plasmodium falciparum infections based on custom dual indexing and Illumina next generation sequencing-technology Sidsel Nag 1,2 *, Marlene D. Dalgaard 3, Poul-Erik
Supporting Information
 Supporting Information Table S1. Oligonucleotide sequences used in this work Oligo DNA A B C D CpG-A CpG-B CpG-C CpG-D Sequence 5 ACA TTC CTA AGT CTG AAA CAT TAC AGC TTG CTA CAC GAG AAG AGC CGC CAT AGT
Supporting Information Table S1. Oligonucleotide sequences used in this work Oligo DNA A B C D CpG-A CpG-B CpG-C CpG-D Sequence 5 ACA TTC CTA AGT CTG AAA CAT TAC AGC TTG CTA CAC GAG AAG AGC CGC CAT AGT
Supplemental Information. Human Senataxin Resolves RNA/DNA Hybrids. Formed at Transcriptional Pause Sites. to Promote Xrn2-Dependent Termination
 Supplemental Information Molecular Cell, Volume 42 Human Senataxin Resolves RNA/DNA Hybrids Formed at Transcriptional Pause Sites to Promote Xrn2-Dependent Termination Konstantina Skourti-Stathaki, Nicholas
Supplemental Information Molecular Cell, Volume 42 Human Senataxin Resolves RNA/DNA Hybrids Formed at Transcriptional Pause Sites to Promote Xrn2-Dependent Termination Konstantina Skourti-Stathaki, Nicholas
G+C content. 1 Introduction. 2 Chromosomes Topology & Counts. 3 Genome size. 4 Replichores and gene orientation. 5 Chirochores.
 1 Introduction 2 Chromosomes Topology & Counts 3 Genome size 4 Replichores and gene orientation 5 Chirochores 6 7 Codon usage 121 marc.bailly-bechet@univ-lyon1.fr Bacterial genome structures Introduction
1 Introduction 2 Chromosomes Topology & Counts 3 Genome size 4 Replichores and gene orientation 5 Chirochores 6 7 Codon usage 121 marc.bailly-bechet@univ-lyon1.fr Bacterial genome structures Introduction
Overexpression Normal expression Overexpression Normal expression. 26 (21.1%) N (%) P-value a N (%)
 SUPPLEMENTARY TABLES Table S1. Alteration of ZNF322A protein expression levels in relation to clinicopathological parameters in 123 Asian and 74 Caucasian lung cancer patients. Asian patients Caucasian
SUPPLEMENTARY TABLES Table S1. Alteration of ZNF322A protein expression levels in relation to clinicopathological parameters in 123 Asian and 74 Caucasian lung cancer patients. Asian patients Caucasian
Genomics and Gene Recognition Genes and Blue Genes
 Genomics and Gene Recognition Genes and Blue Genes November 1, 2004 Prokaryotic Gene Structure prokaryotes are simplest free-living organisms studying prokaryotes can give us a sense what is the minimum
Genomics and Gene Recognition Genes and Blue Genes November 1, 2004 Prokaryotic Gene Structure prokaryotes are simplest free-living organisms studying prokaryotes can give us a sense what is the minimum
Supplemental material
 Supplemental material Diversity of O-antigen repeat-unit structures can account for the substantial sequence variation of Wzx translocases Yaoqin Hong and Peter R. Reeves School of Molecular Bioscience,
Supplemental material Diversity of O-antigen repeat-unit structures can account for the substantial sequence variation of Wzx translocases Yaoqin Hong and Peter R. Reeves School of Molecular Bioscience,
Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system
 Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system Dr. Tim Welsink Molecular Biology Transient Gene Expression OUTLINE Short
Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system Dr. Tim Welsink Molecular Biology Transient Gene Expression OUTLINE Short
SUPPORTING INFORMATION
 SUPPORTING INFORMATION Investigation of the Biosynthesis of the Lasso Peptide Chaxapeptin Using an E. coli-based Production System Helena Martin-Gómez, Uwe Linne, Fernando Albericio, Judit Tulla-Puche,*
SUPPORTING INFORMATION Investigation of the Biosynthesis of the Lasso Peptide Chaxapeptin Using an E. coli-based Production System Helena Martin-Gómez, Uwe Linne, Fernando Albericio, Judit Tulla-Puche,*
Det matematisk-naturvitenskapelige fakultet
 UNIVERSITETET I OSLO Det matematisk-naturvitenskapelige fakultet Exam in: MBV4010 Arbeidsmetoder i molekylærbiologi og biokjemi I MBV4010 Methods in molecular biology and biochemistry I Day of exam: Friday
UNIVERSITETET I OSLO Det matematisk-naturvitenskapelige fakultet Exam in: MBV4010 Arbeidsmetoder i molekylærbiologi og biokjemi I MBV4010 Methods in molecular biology and biochemistry I Day of exam: Friday
An engineered tryptophan zipper-type peptide as a molecular recognition scaffold
 SUPPLEMENTARY MATERIAL An engineered tryptophan zipper-type peptide as a molecular recognition scaffold Zihao Cheng and Robert E. Campbell* Supplementary Methods Library construction for FRET-based screening
SUPPLEMENTARY MATERIAL An engineered tryptophan zipper-type peptide as a molecular recognition scaffold Zihao Cheng and Robert E. Campbell* Supplementary Methods Library construction for FRET-based screening
Supporting Information
 Supporting Information Barderas et al. 10.1073/pnas.0801221105 SI Text: Docking of gastrin to Constructed scfv Models Interactive predocking of the 4-WL-5 motif into the central pocket observed in the
Supporting Information Barderas et al. 10.1073/pnas.0801221105 SI Text: Docking of gastrin to Constructed scfv Models Interactive predocking of the 4-WL-5 motif into the central pocket observed in the
evaluated with UAS CLB eliciting UAS CIT -N Libraries increase in the
 Supplementary Figures Supplementary Figure 1: Promoter scaffold library assemblies. Many ensembless of libraries were evaluated in this work. As a legend, the box outline color in top half of the figure
Supplementary Figures Supplementary Figure 1: Promoter scaffold library assemblies. Many ensembless of libraries were evaluated in this work. As a legend, the box outline color in top half of the figure
II 0.95 DM2 (RPP1) DM3 (At3g61540) b
 Table S2. F 2 Segregation Ratios at 16 C, Related to Figure 2 Cross n c Phenotype Model e 2 Locus A Locus B Normal F 1 -like Enhanced d Uk-1/Uk-3 149 64 36 49 DM2 (RPP1) DM1 (SSI4) a Bla-1/Hh-0 F 3 111
Table S2. F 2 Segregation Ratios at 16 C, Related to Figure 2 Cross n c Phenotype Model e 2 Locus A Locus B Normal F 1 -like Enhanced d Uk-1/Uk-3 149 64 36 49 DM2 (RPP1) DM1 (SSI4) a Bla-1/Hh-0 F 3 111
BioInformatics and Computational Molecular Biology. Course Website
 BioInformatics and Computational Molecular Biology Course Website http://bioinformatics.uchc.edu What is Bioinformatics Bioinformatics upgrades the information content of biological measurements. Discovery
BioInformatics and Computational Molecular Biology Course Website http://bioinformatics.uchc.edu What is Bioinformatics Bioinformatics upgrades the information content of biological measurements. Discovery
Supplemental Data. Bennett et al. (2010). Plant Cell /tpc
 BRN1 ---------MSSSNGGVPPGFRFHPTDEELLHYYLKKKISYEKFEMEVIKEVDLNKIEPWDLQDRCKIGSTPQNEWYFFSHKDRKYPTGS 81 BRN2 --------MGSSSNGGVPPGFRFHPTDEELLHYYLKKKISYQKFEMEVIREVDLNKLEPWDLQERCKIGSTPQNEWYFFSHKDRKYPTGS 82 SMB
BRN1 ---------MSSSNGGVPPGFRFHPTDEELLHYYLKKKISYEKFEMEVIKEVDLNKIEPWDLQDRCKIGSTPQNEWYFFSHKDRKYPTGS 81 BRN2 --------MGSSSNGGVPPGFRFHPTDEELLHYYLKKKISYQKFEMEVIREVDLNKLEPWDLQERCKIGSTPQNEWYFFSHKDRKYPTGS 82 SMB
Supplemental Table 1. Primers used for PCR.
 Supplemental Table 1. Primers used for PCR. Gene Type Primer Sequence Genotyping and semi-quantitative RT-PCR F 5 -TTG CCC GAT CAC CAT CTG TA-3 rwa1-1 R 5 -TGT AGC GAT CAA GGC CTG ATC TAA-3 LB 5 -TAG CAT
Supplemental Table 1. Primers used for PCR. Gene Type Primer Sequence Genotyping and semi-quantitative RT-PCR F 5 -TTG CCC GAT CAC CAT CTG TA-3 rwa1-1 R 5 -TGT AGC GAT CAA GGC CTG ATC TAA-3 LB 5 -TAG CAT
PCR-based Markers and Cut Flower Longevity in Carnation
 PCRbased Markers and Cut Flower Longevity in Carnation Laura De Benedetti, Luca Braglia, Simona Bruna, Gianluca Burchi *, Antonio Mercuri and Tito Schiva Istituto Sperimentale per la Floricoltura, Corso
PCRbased Markers and Cut Flower Longevity in Carnation Laura De Benedetti, Luca Braglia, Simona Bruna, Gianluca Burchi *, Antonio Mercuri and Tito Schiva Istituto Sperimentale per la Floricoltura, Corso
Complexity of the Ruminococcus flavefaciens FD-1 cellulosome reflects an expansion of family-related protein-protein interactions
 Complexity of the Ruminococcus flavefaciens FD-1 cellulosome reflects an expansion of family-related protein-protein interactions Vered Israeli-Ruimy 1,*, Pedro Bule 2,*, Sadanari Jindou 3, Bareket Dassa
Complexity of the Ruminococcus flavefaciens FD-1 cellulosome reflects an expansion of family-related protein-protein interactions Vered Israeli-Ruimy 1,*, Pedro Bule 2,*, Sadanari Jindou 3, Bareket Dassa
Codon Bias with PRISM. 2IM24/25, Fall 2007
 Codon Bias with PRISM 2IM24/25, Fall 2007 from RNA to protein mrna vs. trna aminoacid trna anticodon mrna codon codon-anticodon matching Watson-Crick base pairing A U and C G binding first two nucleotide
Codon Bias with PRISM 2IM24/25, Fall 2007 from RNA to protein mrna vs. trna aminoacid trna anticodon mrna codon codon-anticodon matching Watson-Crick base pairing A U and C G binding first two nucleotide
Anti-Pim-1 (Cat#3247), anti-met (Cat#3127), anti-ron (Cat#2654), Anti-EGFR
 Supplementary Methods Antibodies Anti-Pim-1 (Cat#3247), anti-met (Cat#3127), anti-ron (Cat#2654), Anti-EGFR (Cat#2646), anti-igf1r (Cat#3018), anti-insr (Cat#3020), anti-akt (pan, Cat#4691), anti-phospho-akt
Supplementary Methods Antibodies Anti-Pim-1 (Cat#3247), anti-met (Cat#3127), anti-ron (Cat#2654), Anti-EGFR (Cat#2646), anti-igf1r (Cat#3018), anti-insr (Cat#3020), anti-akt (pan, Cat#4691), anti-phospho-akt
Promoter 1. Promoter 2. Enhancer 2
 Essays 1. An animal normally has two copies of the Xan gene. The protein made by this gene helps the animal clear petroleum-based pollutants from its body. Tine Xan What we know about the Xan gene is shown
Essays 1. An animal normally has two copies of the Xan gene. The protein made by this gene helps the animal clear petroleum-based pollutants from its body. Tine Xan What we know about the Xan gene is shown
Protein Structure Analysis
 BINF 731 Protein Structure Analysis http://binf.gmu.edu/vaisman/binf731/ Iosif Vaisman COMPUTATIONAL BIOLOGY COMPUTATIONAL STRUCTURAL BIOLOGY COMPUTATIONAL MOLECULAR BIOLOGY BIOINFORMATICS STRUCTURAL BIOINFORMATICS
BINF 731 Protein Structure Analysis http://binf.gmu.edu/vaisman/binf731/ Iosif Vaisman COMPUTATIONAL BIOLOGY COMPUTATIONAL STRUCTURAL BIOLOGY COMPUTATIONAL MOLECULAR BIOLOGY BIOINFORMATICS STRUCTURAL BIOINFORMATICS
Sequencing of DNA lesions facilitated by site-specific excision via base. excision repair DNA glycosylases yielding ligatable gaps
 Supporting information Sequencing of DNA lesions facilitated by site-specific excision via base excision repair DNA glycosylases yielding ligatable gaps Jan Riedl, Aaron M. Fleming, and Cynthia J. Burrows*
Supporting information Sequencing of DNA lesions facilitated by site-specific excision via base excision repair DNA glycosylases yielding ligatable gaps Jan Riedl, Aaron M. Fleming, and Cynthia J. Burrows*
Expression of Recombinant Proteins
 Expression of Recombinant Proteins Uses of Cloned Genes sequencing reagents (eg, probes) protein production insufficient natural quantities modify/mutagenesis library screening Expression Vector Features
Expression of Recombinant Proteins Uses of Cloned Genes sequencing reagents (eg, probes) protein production insufficient natural quantities modify/mutagenesis library screening Expression Vector Features
SUPPLEMENTARY INFORMATION
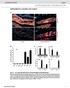 doi: 10.1038/nature07182 SUPPLEMENTAL FIGURES AND TABLES Fig. S1. myf5-expressing cells give rise to brown fat depots and skeletal muscle (a) Perirenal BAT from control (cre negative) and myf5-cre:r26r3-yfp
doi: 10.1038/nature07182 SUPPLEMENTAL FIGURES AND TABLES Fig. S1. myf5-expressing cells give rise to brown fat depots and skeletal muscle (a) Perirenal BAT from control (cre negative) and myf5-cre:r26r3-yfp
11th Meeting of the Science Working Group. Lima, Peru, October 2012 SWG-11-JM-11
 11th Meeting of the Science Working Group Lima, Peru, 15-19 October 2012 Russian population genetics studies of jack mackerel in the South Pacific P.K.Afanasiev M.A.Rabchun A.I.Glubokov Introduction. In
11th Meeting of the Science Working Group Lima, Peru, 15-19 October 2012 Russian population genetics studies of jack mackerel in the South Pacific P.K.Afanasiev M.A.Rabchun A.I.Glubokov Introduction. In
Homework. A bit about the nature of the atoms of interest. Project. The role of electronega<vity
 Homework Why cited articles are especially useful. citeulike science citation index When cutting and pasting less is more. Project Your protein: I will mail these out this weekend If you haven t gotten
Homework Why cited articles are especially useful. citeulike science citation index When cutting and pasting less is more. Project Your protein: I will mail these out this weekend If you haven t gotten
Supplemental Data. Distinct Pathways for snorna and mrna Termination
 Molecular Cell, Volume 24 Supplemental Data Distinct Pathways for snorna and mrna Termination Minkyu Kim, Lidia Vasiljeva, Oliver J. Rando, Alexander Zhelkovsky, Claire Moore, and Stephen Buratowski A
Molecular Cell, Volume 24 Supplemental Data Distinct Pathways for snorna and mrna Termination Minkyu Kim, Lidia Vasiljeva, Oliver J. Rando, Alexander Zhelkovsky, Claire Moore, and Stephen Buratowski A
PROTEIN SYNTHESIS Study Guide
 PART A. Read the following: PROTEIN SYNTHESIS Study Guide Protein synthesis is the process used by the body to make proteins. The first step of protein synthesis is called Transcription. It occurs in the
PART A. Read the following: PROTEIN SYNTHESIS Study Guide Protein synthesis is the process used by the body to make proteins. The first step of protein synthesis is called Transcription. It occurs in the
Supporting Information. Trifluoroacetophenone-Linked Nucleotides and DNA for Studying of DNA-protein Interactions by 19 F NMR Spectroscopy
 Supporting Information Trifluoroacetophenone-Linked Nucleotides and DNA for Studying of DNA-protein Interactions by 19 F NMR Spectroscopy Agata Olszewska, Radek Pohl and Michal Hocek # * Institute of Organic
Supporting Information Trifluoroacetophenone-Linked Nucleotides and DNA for Studying of DNA-protein Interactions by 19 F NMR Spectroscopy Agata Olszewska, Radek Pohl and Michal Hocek # * Institute of Organic
Primer Design Workshop. École d'été en géné-que des champignons 2012 Dr. Will Hintz University of Victoria
 Primer Design Workshop École d'été en géné-que des champignons 2012 Dr. Will Hintz University of Victoria Scenario You have discovered the presence of a novel endophy5c organism living inside the cells
Primer Design Workshop École d'été en géné-que des champignons 2012 Dr. Will Hintz University of Victoria Scenario You have discovered the presence of a novel endophy5c organism living inside the cells
S4B fluorescence (AU)
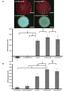 A S4B fluorescence (AU) S4B fluorescence (AU) dsbb csgba csgd dsbb csgba bcsa 5000 * NS NS 4000 * 3000 2000 1000 0 ΔcsgBAΔbcsA ΔcsgDΔdsbBΔbcsA ΔcsgBA ΔdsbBΔcsgBA ΔcsgDΔdsbB B -1000 4000 * * NS 3500 * 3000
A S4B fluorescence (AU) S4B fluorescence (AU) dsbb csgba csgd dsbb csgba bcsa 5000 * NS NS 4000 * 3000 2000 1000 0 ΔcsgBAΔbcsA ΔcsgDΔdsbBΔbcsA ΔcsgBA ΔdsbBΔcsgBA ΔcsgDΔdsbB B -1000 4000 * * NS 3500 * 3000
The B1 Protein Guides the Biosynthesis of a Lasso Peptide
 The B1 Protein Guides the Biosynthesis of a Lasso Peptide Shaozhou Zhu 1,2, Christopher D. Fage 1, Julian D. Hegemann 1, Andreas Mielcarek 1, Dushan Yan 1, Uwe Linne 1 & Mohamed A. Marahiel*,1 1 Department
The B1 Protein Guides the Biosynthesis of a Lasso Peptide Shaozhou Zhu 1,2, Christopher D. Fage 1, Julian D. Hegemann 1, Andreas Mielcarek 1, Dushan Yan 1, Uwe Linne 1 & Mohamed A. Marahiel*,1 1 Department
SUPPLEMENTAL TABLE S1. Additional descriptions of plasmid constructions and the oligonucleotides used Plasmid or Oligonucleotide
 SUPPLEMENTAL TABLE S1. Additional descriptions of plasmid constructions and the oligonucleotides used Plasmid or Oligonucleotide former/ working Description a designation Plasmids pes213a b pes213-tn5
SUPPLEMENTAL TABLE S1. Additional descriptions of plasmid constructions and the oligonucleotides used Plasmid or Oligonucleotide former/ working Description a designation Plasmids pes213a b pes213-tn5
Search for and Analysis of Single Nucleotide Polymorphisms (SNPs) in Rice (Oryza sativa, Oryza rufipogon) and Establishment of SNP Markers
 DNA Research 9, 163 171 (2002) Search for and Analysis of Single Nucleotide Polymorphisms (SNPs) in Rice (Oryza sativa, Oryza rufipogon) and Establishment of SNP Markers Shinobu Nasu, Junko Suzuki, Rieko
DNA Research 9, 163 171 (2002) Search for and Analysis of Single Nucleotide Polymorphisms (SNPs) in Rice (Oryza sativa, Oryza rufipogon) and Establishment of SNP Markers Shinobu Nasu, Junko Suzuki, Rieko
Supplementary Information
 Supplementary Information A general solution for opening double-stranded DNA for isothermal amplification Gangyi Chen, Juan Dong, Yi Yuan, Na Li, Xin Huang, Xin Cui* and Zhuo Tang* Supplementary Materials
Supplementary Information A general solution for opening double-stranded DNA for isothermal amplification Gangyi Chen, Juan Dong, Yi Yuan, Na Li, Xin Huang, Xin Cui* and Zhuo Tang* Supplementary Materials
Molecular Level of Genetics
 Molecular Level of Genetics Most of the molecules found in humans and other living organisms fall into one of four categories: 1. carbohydrates (sugars and starches) 2. lipids (fats, oils, and waxes) 3.
Molecular Level of Genetics Most of the molecules found in humans and other living organisms fall into one of four categories: 1. carbohydrates (sugars and starches) 2. lipids (fats, oils, and waxes) 3.
 www.lessonplansinc.com Topic: Gene Mutations WS Summary: Students will learn about frame shift mutations and base substitution mutations. Goals & Objectives: Students will be able to demonstrate how mutations
www.lessonplansinc.com Topic: Gene Mutations WS Summary: Students will learn about frame shift mutations and base substitution mutations. Goals & Objectives: Students will be able to demonstrate how mutations
Table S1. Sequences of mutagenesis primers used to create altered rdpa- and sdpa genes
 Supplementary Table and Figures for Structural Basis for the Enantiospecificities of R- and S-Specific Phenoxypropionate/α-Ketoglutarate Dioxygenases by Tina A. Müller, Maria I. Zavodszky, Michael Feig,
Supplementary Table and Figures for Structural Basis for the Enantiospecificities of R- and S-Specific Phenoxypropionate/α-Ketoglutarate Dioxygenases by Tina A. Müller, Maria I. Zavodszky, Michael Feig,
Efficient genome replication of hepatitis B virus using adenovirus vector: a compact pregenomic RNA-expression unit
 Efficient genome replication of hepatitis B virus using adenovirus vector: a compact pregenomic RNA-expression unit Mariko Suzuki 1, Saki Kondo 1, Manabu Yamasaki 2, Norie Matsuda 2, Akio Nomoto 2, Tetsuro
Efficient genome replication of hepatitis B virus using adenovirus vector: a compact pregenomic RNA-expression unit Mariko Suzuki 1, Saki Kondo 1, Manabu Yamasaki 2, Norie Matsuda 2, Akio Nomoto 2, Tetsuro
hcd1tg/hj1tg/ ApoE-/- hcd1tg/hj1tg/ ApoE+/+
 ApoE+/+ ApoE-/- ApoE-/- H&E (1x) Supplementary Figure 1. No obvious pathology is observed in the colon of diseased ApoE-/me. Colon samples were fixed in 1% formalin and laid out in Swiss rolls for paraffin
ApoE+/+ ApoE-/- ApoE-/- H&E (1x) Supplementary Figure 1. No obvious pathology is observed in the colon of diseased ApoE-/me. Colon samples were fixed in 1% formalin and laid out in Swiss rolls for paraffin
Engineering D66N mutant using quick change site directed mutagenesis. Harkewal Singh 09/01/2010
 Engineering D66N mutant using quick change site directed mutagenesis Harkewal Singh 09/01/2010 1 1- What is quick change site directed mutagenesis? 2- An overview of the kit contents. 3- A brief information
Engineering D66N mutant using quick change site directed mutagenesis Harkewal Singh 09/01/2010 1 1- What is quick change site directed mutagenesis? 2- An overview of the kit contents. 3- A brief information
Lezione 10. Bioinformatica. Mauro Ceccanti e Alberto Paoluzzi
 Lezione 10 Bioinformatica Mauro Ceccanti e Alberto Paoluzzi Dip. Informatica e Automazione Università Roma Tre Dip. Medicina Clinica Università La Sapienza Lezione 10: Sintesi proteica Synthesis of proteins
Lezione 10 Bioinformatica Mauro Ceccanti e Alberto Paoluzzi Dip. Informatica e Automazione Università Roma Tre Dip. Medicina Clinica Università La Sapienza Lezione 10: Sintesi proteica Synthesis of proteins
Phosphate and R2D2 Restrict the Substrate Specificity of Dicer-2, an ATP- Driven Ribonuclease
 Supplemental Information Molecular Cell, Volume 42 hosphate and R2D2 Restrict the Substrate Specificity of Dicer-2, an AT- Driven Ribonuclease Elif Sarinay Cenik, Ryuya Fukunaga, Gang Lu, Robert Dutcher,
Supplemental Information Molecular Cell, Volume 42 hosphate and R2D2 Restrict the Substrate Specificity of Dicer-2, an AT- Driven Ribonuclease Elif Sarinay Cenik, Ryuya Fukunaga, Gang Lu, Robert Dutcher,
A green method of staining DNA in polyacrylamide gel electrophoresis based on fluorescent copper nanoclusters synthesized in situ
 Electronic Supplementary Material A green method of staining DNA in polyacrylamide gel electrophoresis based on fluorescent copper nanoclusters synthesized in situ Xiaoli Zhu 1, Hai Shi 1, Yalan Shen 1,
Electronic Supplementary Material A green method of staining DNA in polyacrylamide gel electrophoresis based on fluorescent copper nanoclusters synthesized in situ Xiaoli Zhu 1, Hai Shi 1, Yalan Shen 1,
SUPPORTING INFORMATION FILE
 Intrinsic and extrinsic connections of Tet3 dioxygenase with CXXC zinc finger modules Nan Liu, Mengxi Wang, Wen Deng, Christine S. Schmidt, Weihua Qin, Heinrich Leonhardt and Fabio Spada Department of
Intrinsic and extrinsic connections of Tet3 dioxygenase with CXXC zinc finger modules Nan Liu, Mengxi Wang, Wen Deng, Christine S. Schmidt, Weihua Qin, Heinrich Leonhardt and Fabio Spada Department of
Causes and Effects of N-Terminal Codon Bias in Bacterial Genes. Mikk Eelmets Journal Club
 Causes and Effects of N-Terminal Codon Bias in Bacterial Genes Mikk Eelmets Journal Club 21.2.214 Introduction Ribosomes were first observed in the mid-195s (Nobel Prize in 1974) Nobel Prize in 29 for
Causes and Effects of N-Terminal Codon Bias in Bacterial Genes Mikk Eelmets Journal Club 21.2.214 Introduction Ribosomes were first observed in the mid-195s (Nobel Prize in 1974) Nobel Prize in 29 for
Supplementary Information
 Supplementary Information Microbead-based biomimetic synthetic neighbors enhance survival and function of rat pancreatic β-cells Wei Li, a Samuel Lee, b Minglin Ma, a, f Soo Min Kim, b Patrick Guye, c
Supplementary Information Microbead-based biomimetic synthetic neighbors enhance survival and function of rat pancreatic β-cells Wei Li, a Samuel Lee, b Minglin Ma, a, f Soo Min Kim, b Patrick Guye, c
The Making of the Fittest: Natural Selection and Adaptation
 INTRODUCTION MOLECULAR GENETICS OF COLOR MUTATIONS IN ROCK POCKET MICE THE ROCK POCKET MOUSE The rock pocket mouse, Chaetodipus intermedius, is a small, nocturnal animal found in the deserts of the southwestern
INTRODUCTION MOLECULAR GENETICS OF COLOR MUTATIONS IN ROCK POCKET MICE THE ROCK POCKET MOUSE The rock pocket mouse, Chaetodipus intermedius, is a small, nocturnal animal found in the deserts of the southwestern
Supplemental Information. Target-Mediated Protection of Endogenous. MicroRNAs in C. elegans. Inventory of Supplementary Information
 Developmental Cell, Volume 20 Supplemental Information Target-Mediated Protection of Endogenous MicroRNAs in C. elegans Saibal Chatterjee, Monika Fasler, Ingo Büssing, and Helge Großhans Inventory of Supplementary
Developmental Cell, Volume 20 Supplemental Information Target-Mediated Protection of Endogenous MicroRNAs in C. elegans Saibal Chatterjee, Monika Fasler, Ingo Büssing, and Helge Großhans Inventory of Supplementary
Chapter 3: Information Storage and Transfer in Life
 Chapter 3: Information Storage and Transfer in Life The trapped scientist examples are great for conceptual purposes, but they do not accurately model how information in life changes because they do not
Chapter 3: Information Storage and Transfer in Life The trapped scientist examples are great for conceptual purposes, but they do not accurately model how information in life changes because they do not
A Genetically Encoded Toolbox for Glycocalyx Engineering: Tunable Control of Cell Adhesion,
 TITLE A Genetically Encoded Toolbox for Glycocalyx Engineering: Tunable Control of Cell Adhesion, Survival, and Cancer Cell Behaviors AUTHORS Carolyn R. Shurer *, Marshall J. Colville *, Vivek K. Gupta,
TITLE A Genetically Encoded Toolbox for Glycocalyx Engineering: Tunable Control of Cell Adhesion, Survival, and Cancer Cell Behaviors AUTHORS Carolyn R. Shurer *, Marshall J. Colville *, Vivek K. Gupta,
MCB421 FALL2005 EXAM#3 ANSWERS Page 1 of 12. ANSWER: Both transposon types form small duplications of adjacent host DNA sequences.
 Page 1 of 12 (10pts) 1. There are two mechanisms for transposition used by bacterial transposable elements: replicative (Tn3) and non-replicative (Tn5 and Tn10). Compare and contrast the two mechanisms
Page 1 of 12 (10pts) 1. There are two mechanisms for transposition used by bacterial transposable elements: replicative (Tn3) and non-replicative (Tn5 and Tn10). Compare and contrast the two mechanisms
Genes and Proteins. Objectives
 Genes and Proteins Lecture 15 Objectives At the end of this series of lectures, you should be able to: Define terms. Explain the central dogma of molecular biology. Describe the locations, reactants, and
Genes and Proteins Lecture 15 Objectives At the end of this series of lectures, you should be able to: Define terms. Explain the central dogma of molecular biology. Describe the locations, reactants, and
