Automated Imaging and Dual-Mask Analysis of γh2ax Foci to Determine DNA Damage on an Individual Cell Basis
|
|
|
- Charleen Phillips
- 6 years ago
- Views:
Transcription
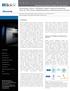
1 A p p l i c a t i o n N o t e Automated Imaging and Dual-Mask Analysis of γh2ax Foci to Determine DNA Damage on an Individual Cell Basis Brad Larson, BioTek Instruments, Inc., Winooski, VT USA Asha Sinha and Sachin Katyal, University of Manitoba, Winnipeg, Manitoba, Canada Key Words: Gamma H2AX Gamma H2A.X Histone H2AX Serine 139 Double Strand DNA Break DNA Damage DNA Repair Genotoxicity BioTek Instruments, Inc. P.O. Box 998, Highland Park, Winooski, Vermont USA Phone: Outside the USA: customercare@biotek.com Copyright 2016 Introduction In mammalian cells, one of the most serious types of DNA damage is a double-stranded break (DSB), where the DNA double helix is completely severed. DSBs are associated with cell death and tumorigenesis, so they are of keen interest in understanding cancerous mechanisms as well as developing therapeutic treatments. One of the most characterized early markers of DNA DSBs is the phosphorylation of the histone 2AX (H2AX) to γ-h2ax. Here, a positive feedback loop is created between γ-h2ax and phosphatidyl inositol 3' kinase-related kinases, where repair systems are recruited to the damage, creating nucleation sites, or foci, that correspond to the level of DSB. When the foci are bound to fluorescently labeled antibodies, overall γ-h2ax levels can be visualized and quantified via immunofluorescence microscopy. The conventional, manual method of evaluating foci via a γ-h2ax assay offers limited throughput and is subject to operator variability. Incorporating automated imaging and analysis into the assay workflow can help to enhance throughput, assay robustness and accuracy, while also increasing overall laboratory efficiency. Additionally, while manual methods allow for overall population assessment captured within an image or set of images, they are not conducive to analysis of individual foci within the population. Here, we demonstrate an automated γ-h2ax assay workflow using a novel cell imaging multimode reader with advanced data analysis capabilities to process and image multiple samples simultaneously, including dual-mask capabilities to assess real-time population-level data as well as individual foci data, for a complete data set. This automated assay format provides an accurate, robust method to assess DNA damage in mammalian cells. Materials and Methods Materials Cells U251 (human glial) cells (Catalog No ) were purchased from Sigma Aldrich (Saint Louis, MO). Assay and Experimental Components The known cytotoxin, camptothecin (Catalog No ) was purchased from EMD Millipore (Billerica, MA). The Alexa Fluor 647 anti-h2a.x- Phosphorylated (Ser139) antibody (Catalog No ) was purchased from BioLegend (San Diego, CA). Cytation 5 Cell Imaging Multi-Mode Reader Cytation 5 is a modular multi-mode microplate reader combined with automated digital microscopy. Filter- and monochromator-based microplate reading are available, along with laserbased excitation for Alpha assays. The microscopy module provides up to 60x magnification in fluorescence, brightfield, color brightfield and phase contrast. With special emphasis on live-cell assays, Cytation 5 features shaking, temperature control to 65 C, CO 2 /O 2 gas control and dual injectors for kinetic assays. The instrument was used to image the stained DNA using the GFP imaging channel. Integrated Gen5 Microplate Reader and Imager Software controls Cytation 5, and also automates image capture, analysis and processing. Methods Gamma H2AX Assay Performance Cultured U251 cells were treated with concentrations of camptothecin ranging from 1-0 µm. Immunostaining was then performed using the parameters listed in Table 1.
2 Step Number 1 Gamma H2AX Immunostaining Procedure Description Fix cells with 4% paraformaldehyde Time/Iteration 10 minutes 2 Wash with PBS, ph 7.4 2x 3 Permeabilize with 0.5% Triton-X 100 in PBS, ph minutes 4 Wash in PBS, ph 7.4 2x 5 6 Dispense 50 µl/well of 1º antibody (1:1000; γh2ax) in 3% BSA plus PBS Wash with 0.1% Triton- X 100 in PBS, ph ºC 7 Wash with PBS, ph 7.4 2x 8 Dispense 150 µl/well of Hoechst (0.2 µg/ml) in PBS, ph 7.4 1x 1 4 ºC Table 1. Immunostaining procedure using Alexa Fluor 647 anti- H2AX-Phosphorylated (Ser139) antibody. Following completion of the staining procedure, the plate was then imaged by the Cytation 5. Table 2 lists the settings used to perform automated image capture of each sample well. First Imaging Channel Second Imaging Channel Objective Montage Montage Overlap Exposure Imaging Parameters DAPI CY5 10x 3 Row by 3 Column Auto for Stitching Auto Exposure based upon Positive (1 µm Camptothecin) and Negative (0 µm Camptothecin) Control Wells Table 2. Automated gamma H2AX imaging parameters. Imaging Pre-Processing Parameters First Imaging Channel Table 3. Image pre-processing parameters. Primary Cellular Analysis Parameters Imaging Channel CY5 Threshold 5000 Split touching objects Fill holes in masks Min. Object Size 0 µm Max. Object Size 8 µm Include primary edge objects Analyze entire image Unchecked Advanced Detection Options Flattening Size Image Smoothing Strength Evaluate On Primary Mask Table 4. Foci analysis parameters. DAPI Rolling Ball Diameter Auto (136 µm) Image Smoothing Strength 0 Second Image Channel CY5 Rolling Ball Diameter 2 µm Image Smoothing Strength 0 Primary Foci Analysis Primary mask cellular analysis criteria (Table 4) were applied to automatically place object masks around individual foci. The average area of an individual foci was then used in conjunction with results from the dual-mask analysis to calculate the number of foci per nuclei on an individual foci basis. 2 µm (Rolling Ball diameter) 0 Cycles of 3x3 average filter 5% of Lowest Pixels Use Threshold Mask Image Pre-Processing Image pre-processing was applied to remove excessive background signal from the images using the criteria listed in Table 3. As the individual foci are small in diameter, and may be in close proximity within the nucleus, an optimized subset, or area, of the image that is analyzed to distinguish between background and true signal was required. A greatly reduced value of 2 µm provided the most optimal final images for subsequent analysis. Dual Mask Individual Nuclei Foci Analysis Primary mask cellular analysis criteria (Table 5) were applied to automatically place object masks around nuclei in each captured image. Secondary mask cellular analysis criteria (Table 6) were then also applied to place linked additional masks around individual foci. The area of the primary mask was minimally expanded such that areas within the primary mask meeting the criteria to identify foci would still fall within the nucleus. Threshold in Mask was also selected to allow individual secondary masks to be placed anywhere within the initial primary mask meeting the included criteria. 2
3 Primary Cellular Analysis Parameters Imaging Channel DAPI Threshold Auto (35) Split touching objects Fill holes in masks Min. Object Size 10 µm Max. Object Size 100 µm Include primary edge objects Analyze entire image Unchecked Advanced Detection Options Flattening Size Image Smoothing Strength Evaluate On Primary Mask Table 5. Primary mask nuclei analysis parameters. Auto 300 µm (Rolling Ball diameter) 2 Cycles of 3x3 average filter 5% of Lowest Pixels Use Threshold Mask Secondary Cellular Analysis Parameters Imaging Channel Measure Within a Secondary Mask CY5 Include Primary and Secondary Area in Analysis Expand Primary Mask 1 µm Threshold 5000 Method Fill Holes in the Mask Table 6. Secondary mask foci analysis parameters. Threshold in Mask Unchecked Results and Discussion Automated 96-Well γh2ax Assay Imaging and Pre-Processing Automated γ-h2ax assay imaging, performed in 96-well format, was validated using U251 cells exposed to various camptothecin concentrations. Cytation 5 automatically imaged the samples using a 3x3 image montage to ensure that a statistically relevant number of nuclei were analyzed per image. Auto exposure was used with each imaging channel (Table 2) and optimized using positive and negative control wells to ensure that the integrated Gen5 Microplate Reader and Imager Software settings accurately imaged the nuclei and labeled foci in each image. Following image capture, Gen5 automatically pre-processed the samples to remove excess background signal and properly prepare each image for analysis. In the case of assays where the signal of interest emanates from small punctate areas within the well, this step is critical, particularly when large numbers of puncta fall within small areas of the image. This is observed in Figure 1A where cells were exposed to a high (1 µm) concentration of camptothecin. While individual labeled foci can be seen in certain nuclei, others containing higher foci numbers appear where the nuclei is completely covered with the fluorescent signal. This phenomenon precludes the possibility of performing accurate analysis of the DNA damage within these nuclei. A. B. Figure 1. Image background signal removal via pre-processing. Zoomed 10x images (A) before; and (B) after pre-processing criteria (Table 3) applied using Gen5 Software. 3
4 By performing the pre-processing step in Gen5, with the incorporation of a 2 µm diameter in which to analyze each portion of the image, superfluous fluorescent signal not associated with the labeled foci is eliminated from the image (Figure 1B). Individual labeled foci are then visualized in each nuclei and accurately analyzed. Following image capture Gen5 automatically stitched together the nine individual image tiles in the original configuration of the 3x3 montage. This allows for analysis of the entire imaged area of the well. Figure 2. Image stitching. Final stitched image of the nine tiles included in the 3x3 montage. The final images accurately portrayed the extent of DNA damage from each test condition, where low numbers of labeled foci per nuclei were seen in untreated cells (Figure 3A), while those treated with camptothecin showed increasing numbers of nuclei affected as well as an increasing number of labeled foci per nuclei (Figures 3B-D). A. 0 µm Camptothecin (untreated) B µm Camptothecin C. 0.1 µm Camptothecin D. 1 µm Camptothecin Figure 3. Final processed images following camptothecin treatment and immunofluorescent staining. Zoomed images captured using a 10x objective and 3x3 montage of U251 cells exposed to (A) 0 µm (untreated) camptothecin; (B) 0.01 µm camptothecin; (C) 0.1 µm camptothecin; or (D) 1 µm camptothecin. Blue: Hoechst stained nuclei; Red: CY5 signal from labeled foci. 4
5 Automated γh2ax Assay Analysis Individual Foci Identification Primary cellular analysis criteria (Table 4) were applied to all images captured with Gen5 Software using the fluorescent signal from the CY5 channel to place object masks around individual labeled foci (Figure 4). Minimum and maximum object sizes were set low due to the small size of each labeled foci. The background flattening size, or rolling ball diameter, was also set to 2 µm as was seen in the image pre-processing step, to aid in accurate identification of each foci. The area of each individual foci was reported and averaged per image. A value of 1.95 µm was determined as the average area of a labeled foci. This value was then used in the second cellular analysis step to calculate the number of labeled foci within each individual nuclei. Figure 4. Automated cellular analysis for specific identification of labeled foci. Object masks placed using signal from antibody labeled foci. Images captured using a 10x objective, 3x3 image montage, and DAPI and CY5 imaging channels. Dual-Mask Identification of Labeled Foci per Nuclei Primary and secondary cellular analysis criteria (Tables 5 and 6) were applied to all images captured using Gen5. Primary cellular analysis automatically masked each nuclei (Figure 5A), while secondary analysis automatically placed individual masks around nuclear foci specifically within the original primary masks (Figure 5B). A. B. Figure 5. Automated γh2ax dual-mask analysis based on user-programmed parameters. Primary cellular analysis object masks showing (A) nuclei and secondary masks showing (B) labeled foci, respectively. Images captured using a 10x objective, 3x3 image montage, in addition to DAPI and CY5 imaging channels. By using primary cellular analysis to initially mask nuclei, the foci identified were properly linked to each nuclei; allowing for analysis on a single nuclei level. Subpopulation analysis was then applied to identify nuclei in each image exhibiting positive DNA damage. The total signal from the CY5 channel within objects identified using secondary analysis was used to apply threshold criteria. As foci appear and increase in number per nuclei the total fluorescence in the CY5 channel within the object masks placed around the foci also increases. Therefore, a value greater than 100,000 RFU was set to eliminate nuclei exhibiting low levels of fluorescence in the CY5 channel not emanating from true labeled foci. Using this criteria, nuclei containing distinct labeled foci meet the set threshold parameter for being γh2ax positive cells (Figure 6). 5
6 A. B. Figure 6. γh2ax positive nuclei. (A) Nuclei meeting γh2ax positive subpopulation criteria; and (B) labeled foci within positive nuclei. Positive γh2ax nuclei in purple and negative nuclei in yellow. Labeled Foci per Individual Nuclei Calculation Upon completion of the gh2ax assay procedure and image preparation, Gen5 was used to calculate the commonly used assay metric labeled foci number per nuclei on an individual nuclei basis. This was generated using the following formula and object metric of interest (Table 7), in addition to the average foci area value from the initial foci analysis (1.95 µm 2 ), to automatically calculate the number of foci per nuclei: M 1 /1.95 Secondary Mask Calculated Metric Area_2[Tsf[Cy5] Description Total foci area value within the secondary masks of an identified nuclei Data Reduction Designation M 1 Table 7. Number of foci per nuclei metrics. When the total area is divided by the average area of a labeled foci, the final foci count within nuclei is yielded. Dual-Mask γh2ax Assay Validation Data The results in Figure 7A illustrate that the total coverage area per nuclei of labeled foci increases in direct correlation to the camptothecin treatment level. Then by applying the formula explained above, the number of labeled foci per nuclei is calculated (Figure 7B). This value agrees with expected results as the number of foci in an individual nucleus starts at an insignificant level in wells containing untreated cells, indicating little to no DNA damage, and increases directly with camptothecin treatment. A. B. 6 Figure 7. Labeled foci analyses using U251 cells exposed to various camptothecin concentrations. (A) Total foci coverage area per nuclei, and (B) calculated foci number in individual nuclei. Twelve replicates tested at each compound concentration.
7 Because the foci count is calculated from the total foci coverage area for each nuclei counted in the image, using the dual-mask analysis, it is also possible to quickly assess results for an individual nuclei by clicking on that object. The linked data is then highlighted (Figure 8). Therefore, if an interesting phenomenon is observed in a particular object, all relevant data can be immediately scrutinized. Figure 8. Observation of single object data. Objects of interest highlighted in red. Relevant data such as labeled foci area coverage and calculated foci count highlighted in the data table to the right. Finally, using the data from the dual mask individual nuclei foci analysis, subpopulation parameters were applied to the primary objects, representing txd in each image using the criteria listed in Table 8. The criteria used was the combined total signal within object masks placed around foci interior to previously identified nuclei. The established condition separates the nuclei exhibiting DNA damage, as witnessed by a minimal total signal from identified labeled foci, from undamaged nuclei. Subpopulation Parameters Criteria Description Condition Integral_2 (Secondary Mask CY5 Integral Signal) Table 8. Gamma H2AX positive cell criteria. Total CY5 fluorescence from identified foci within an individual nuclei > Using the subpopulation parameter listed in Table 8, the number of gh2ax positive nuclei per test condition was also automatically determined (Figure 9A). By comparing this number to the total nuclei in the image, a normalized gh2ax positive nuclei percentage was then calculated (Figure 9B). A. B. 7 Figure 9. γh2ax positive (A) nuclei count and (B) ratio to total nuclei counted. U251 cells exposed to various concentrations of camptothecin. Twelve replicates tested at each compound concentration.
8 From the graphs in Figure 9 it is also evident that the number and ratio of nuclei meeting the established subpopulation criteria increase when cells are exposed to higher concentrations of camptothecin. Results for both foci count per nuclei and percent γh2ax positive nuclei exhibit similar trends across all camptothecin concentrations tested, confirming the ability of the automated imaging procedure and dual-mask analysis method to deliver accurate, robust data on an individual nuclei and population basis. Conclusions Incorporating a cell imaging multi-mode reader to automate the γ-h2ax assay provides increased throughput to aid in laboratory efficiency. At the same time, dual-mask capabilities allow population level analyses as well as single nuclei scrutiny, and all analyses are automatically performed without the need for separate software. Automated imaging and analyses also eliminates human subjectivity and analysis errors that could potentially skew results. This creates a robust, user-friendly and accurate method to detect DNA damage in mammalian cells. 8 AN063016_19, Rev. 08/08/16
Brightfield and Fluorescence Imaging using 3D PrimeSurface Ultra-Low Attachment Microplates
 A p p l i c a t i o n N o t e Brightfield and Fluorescence Imaging using 3D PrimeSurface Ultra-Low Attachment Microplates Brad Larson, BioTek Instruments, Inc., Winooski, VT USA Anju Dang, S-BIO, Hudson,
A p p l i c a t i o n N o t e Brightfield and Fluorescence Imaging using 3D PrimeSurface Ultra-Low Attachment Microplates Brad Larson, BioTek Instruments, Inc., Winooski, VT USA Anju Dang, S-BIO, Hudson,
Use of Phase Contrast Imaging to Track Morphological Cellular Changes due to Apoptotic Activity
 A p p l i c a t i o n N o t e Use of Phase Contrast Imaging to Track Morphological Cellular Changes due to Apoptotic Activity Brad Larson and Peter Banks, Applications Department, BioTek Instruments, Inc.,
A p p l i c a t i o n N o t e Use of Phase Contrast Imaging to Track Morphological Cellular Changes due to Apoptotic Activity Brad Larson and Peter Banks, Applications Department, BioTek Instruments, Inc.,
Automated, Kinetic Imaging of Cell Migration and Invasion Assays using Corning FluoroBlok Permeable Supports
 A p p l i c a t i o n N o t e Automated, Kinetic Imaging of Cell Migration and Invasion Assays using Corning FluoroBlok Permeable Supports Brad Larson, Principal Scientist, Applications Department, BioTek
A p p l i c a t i o n N o t e Automated, Kinetic Imaging of Cell Migration and Invasion Assays using Corning FluoroBlok Permeable Supports Brad Larson, Principal Scientist, Applications Department, BioTek
Imaging of BacMam Transfected U-2 OS Cells
 A p p l i c a t i o n N o t e Imaging of BacMam Transfected U-2 OS Cells Optimization of Transfection Conditions Using the Cytation 3 Multi- Mode Reader and Gen5 Data Analysis Software Paul Held Ph. D.
A p p l i c a t i o n N o t e Imaging of BacMam Transfected U-2 OS Cells Optimization of Transfection Conditions Using the Cytation 3 Multi- Mode Reader and Gen5 Data Analysis Software Paul Held Ph. D.
Long-Term Hepatotoxicity Studies using Cultured Human ipsc- Derived Hepatocytes
 A p p l i c a t i o n N o t e Long-Term Hepatotoxicity Studies using Cultured Human ipsc- Derived Hepatocytes Brad Larson and Peter Banks, BioTek Instruments, Inc., Winooski, VT USA Coby Carlson and Michael
A p p l i c a t i o n N o t e Long-Term Hepatotoxicity Studies using Cultured Human ipsc- Derived Hepatocytes Brad Larson and Peter Banks, BioTek Instruments, Inc., Winooski, VT USA Coby Carlson and Michael
Monitoring Cell Cycle Progression in Cancer Cells
 A p p l i c a t i o n N o t e Monitoring Cell Cycle Progression in Cancer Cells Using Nuclear Staining to Assess Cellular DNA Content Paul Held, Ph.D., Laboratory Manager, Applications Department, BioTek
A p p l i c a t i o n N o t e Monitoring Cell Cycle Progression in Cancer Cells Using Nuclear Staining to Assess Cellular DNA Content Paul Held, Ph.D., Laboratory Manager, Applications Department, BioTek
Automated Imaging and Analysis of a Novel Comet Assay to Enable High Throughput Genotoxicity Testing
 A p p l i c a t i o n N o t e Automated Imaging and Analysis of a Novel Comet Assay to Enable High Throughput Genotoxicity Testing Brad Larson, BioTek Instruments, Inc., Winooski, VT Clare Whittaker and
A p p l i c a t i o n N o t e Automated Imaging and Analysis of a Novel Comet Assay to Enable High Throughput Genotoxicity Testing Brad Larson, BioTek Instruments, Inc., Winooski, VT Clare Whittaker and
Validation of High-Throughput Wound Healing Assay using 3D Cell Patterning and Automated, Kinetic Imaging
 A p p l i c a t i o n N o t e Validation of High-Throughput Wound Healing Assay using 3D Cell Patterning and Automated, Kinetic Imaging Brad Larson and Peter Banks, BioTek Instruments, Inc., Winooski,
A p p l i c a t i o n N o t e Validation of High-Throughput Wound Healing Assay using 3D Cell Patterning and Automated, Kinetic Imaging Brad Larson and Peter Banks, BioTek Instruments, Inc., Winooski,
Automated, Multi-Parameter, Kinetic Methods to Quantify Cell Death
 A p p l i c a t i o n N o t e Automated, Multi-Parameter, Kinetic Methods to Quantify Cell Death Brad Larson, Principal Scientist, BioTek Instruments, Inc., Winooski, VT USA Abstract Automated, multi-parameter
A p p l i c a t i o n N o t e Automated, Multi-Parameter, Kinetic Methods to Quantify Cell Death Brad Larson, Principal Scientist, BioTek Instruments, Inc., Winooski, VT USA Abstract Automated, multi-parameter
Live Cell Imaging of RNA Expression
 A p p l i c a t i o n N o t e Live Cell Imaging of RNA Expression Using the Cytation 3 Cell Imaging Multi-Mode Reader to Image SmartFlare RNA Probes Paul Held Ph. D, Laboratory Manager, Applications Dept.,
A p p l i c a t i o n N o t e Live Cell Imaging of RNA Expression Using the Cytation 3 Cell Imaging Multi-Mode Reader to Image SmartFlare RNA Probes Paul Held Ph. D, Laboratory Manager, Applications Dept.,
Automated Digital Microscopy
 A p p l i c a t i o n G u i d e Peter Banks, Ph.D. and Peter J. Brescia, Applications Department, BioTek Instruments, Inc., Winooski, VT Table of Contents Introduction ----------------------------------------------------------------------------------------------------------------------
A p p l i c a t i o n G u i d e Peter Banks, Ph.D. and Peter J. Brescia, Applications Department, BioTek Instruments, Inc., Winooski, VT Table of Contents Introduction ----------------------------------------------------------------------------------------------------------------------
Comparison of Oridonin Cytotoxicity in U-2 OS and HepG2 Cells
 A p p l i c a t i o n N o t e Comparison of Oridonin Cytotoxicity in U-2 OS and HepG2 Cells Using the BioSpa 8 to Manage Repeated Reagent Additions to Live Cells Paul Held, PhD, Laboratory Manager, Applications
A p p l i c a t i o n N o t e Comparison of Oridonin Cytotoxicity in U-2 OS and HepG2 Cells Using the BioSpa 8 to Manage Repeated Reagent Additions to Live Cells Paul Held, PhD, Laboratory Manager, Applications
Automated Imaging Assay for Characterizing Ca 2+ Flux with R-GECO Biosensor
 A p p l i c a t i o n N o t e Automated Imaging Assay for Characterizing Ca 2+ Flux with R-GECO Biosensor Joe Clayton, Ph.D., and Peter Banks, Ph.D., BioTek Instruments, Inc., Winooski, VT USA Abstract
A p p l i c a t i o n N o t e Automated Imaging Assay for Characterizing Ca 2+ Flux with R-GECO Biosensor Joe Clayton, Ph.D., and Peter Banks, Ph.D., BioTek Instruments, Inc., Winooski, VT USA Abstract
EPIGENTEK. EpiQuik In Situ DNA Damage Assay Kit. Base Catalog # P-6001 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE
 EpiQuik In Situ DNA Damage Assay Kit Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik In Situ DNA Damage Assay Kit is suitable for specifically measuring DNA damage in situ through
EpiQuik In Situ DNA Damage Assay Kit Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik In Situ DNA Damage Assay Kit is suitable for specifically measuring DNA damage in situ through
Performance of a Label-Free Image-Based 2D Scratch Wound Healing Assay to monitor Cell Migration and its Inhibition
 A p p l i c a t i o n N o t e Performance of a Label-Free Image-Based 2D Scratch Wound Healing Assay to monitor Cell Migration and its Inhibition Leonie Rieger and Brad Larson, Applications Department,
A p p l i c a t i o n N o t e Performance of a Label-Free Image-Based 2D Scratch Wound Healing Assay to monitor Cell Migration and its Inhibition Leonie Rieger and Brad Larson, Applications Department,
Utility of Variable Bandwidth Monochromators for Quantification of Fluorescent Probes in Produced Effluent Water
 A p p l i c a t i o n N o t e Utility of Variable Bandwidth Monochromators for Quantification of Fluorescent Probes in Produced Effluent Water Brad Larson and Peter Banks, BioTek Instruments, Inc., Winooski,
A p p l i c a t i o n N o t e Utility of Variable Bandwidth Monochromators for Quantification of Fluorescent Probes in Produced Effluent Water Brad Larson and Peter Banks, BioTek Instruments, Inc., Winooski,
High Contrast Brightfield
 Tech Note High Contrast Brightfield Enabling Microplate-based Automated Label-free Cell Counting Paul Held, Joe Clayton, and Peter Banks Introduction Cell counting is a basic technique for cell culture
Tech Note High Contrast Brightfield Enabling Microplate-based Automated Label-free Cell Counting Paul Held, Joe Clayton, and Peter Banks Introduction Cell counting is a basic technique for cell culture
Introduction. Figure 1. Oris Cell Migration Assay Principle
 Optimizing Performance of the Membrane-free, Oris Cell Migration Assay for High Throughput Screening using the BioTek Synergy HT Multi-Mode Microplate Reader Keren I. Hulkower, Renee L. Herber, and Scott
Optimizing Performance of the Membrane-free, Oris Cell Migration Assay for High Throughput Screening using the BioTek Synergy HT Multi-Mode Microplate Reader Keren I. Hulkower, Renee L. Herber, and Scott
Investigation of Cell Migration using a High Density Cell Exclusion Assay and Automated Microplate Imager
 A p p l i c a t i o n N o t e Investigation of Cell Migration using a High Density Cell Exclusion Assay and Automated Microplate Imager Peter J. Brescia and Peter Banks, Applications Department, BioTek
A p p l i c a t i o n N o t e Investigation of Cell Migration using a High Density Cell Exclusion Assay and Automated Microplate Imager Peter J. Brescia and Peter Banks, Applications Department, BioTek
Determination of Endogenous Cellular Kinase Activity Through the Use of an Automated, High Throughput AlphaScreen - Based Assay
 A p p l i c a t i o n N o t e Determination of Endogenous Cellular Kinase Activity Through the Use of an Automated, High Throughput AlphaScreen - Based Assay Brad Larson and Peter Banks, Applications Department,
A p p l i c a t i o n N o t e Determination of Endogenous Cellular Kinase Activity Through the Use of an Automated, High Throughput AlphaScreen - Based Assay Brad Larson and Peter Banks, Applications Department,
OxiSelect DNA Double Strand Break (DSB) Staining Kit
 Product Manual OxiSelect DNA Double Strand Break (DSB) Staining Kit Catalog Number STA-321 100 assays FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction DNA double-strand breaks (DSBs)
Product Manual OxiSelect DNA Double Strand Break (DSB) Staining Kit Catalog Number STA-321 100 assays FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction DNA double-strand breaks (DSBs)
Quantification of MMP Activity and Inhibition in a 3D Tumor Invasion Model
 A p p l i c a t i o n N o t e Quantification of MMP Activity and Inhibition in a 3D Tumor Invasion Model Using a Novel Fluorescence Detection Technology and Digital Widefield Microscopy Brad Larson and
A p p l i c a t i o n N o t e Quantification of MMP Activity and Inhibition in a 3D Tumor Invasion Model Using a Novel Fluorescence Detection Technology and Digital Widefield Microscopy Brad Larson and
Normalization of Agilent Seahorse XF Data by In-situ Cell Counting Using a BioTek Cytation 5
 Normalization of Agilent Seahorse XF Data by In-situ Cell Counting Using a BioTek Cytation Application Note Authors Yoonseok Kam 1, Ned Jastromb 1, Joe Clayton, Paul Held, and Brian P. Dranka 1 1 Agilent
Normalization of Agilent Seahorse XF Data by In-situ Cell Counting Using a BioTek Cytation Application Note Authors Yoonseok Kam 1, Ned Jastromb 1, Joe Clayton, Paul Held, and Brian P. Dranka 1 1 Agilent
Cost-Effective Automated Hepatotoxicity Testing using in vitro 3D Bioprinting
 A p p l i c a t i o n N o t e Cost-Effective Automated Hepatotoxicity Testing using in vitro 3D Bioprinting Brad Larson, Principal Scientist, BioTek Instruments, Inc., Winooski, VT USA Alexa Rosenberg,
A p p l i c a t i o n N o t e Cost-Effective Automated Hepatotoxicity Testing using in vitro 3D Bioprinting Brad Larson, Principal Scientist, BioTek Instruments, Inc., Winooski, VT USA Alexa Rosenberg,
Add Live, Dead and Total Dyes. Overnight Variable 30 min. 10 min 5 min. 30 min
 Cell Viability Analysis using Calcein AM, Propidium Iodide and Hoechst Application Description Celigo Application Plate Type Major Steps Cell viability analysis using Calcein AM, Propidium Iodide and Hoechst
Cell Viability Analysis using Calcein AM, Propidium Iodide and Hoechst Application Description Celigo Application Plate Type Major Steps Cell viability analysis using Calcein AM, Propidium Iodide and Hoechst
Determination of Cell Cycle Time
 A p p l i c a t i o n N o t e Determination of Cell Cycle Time Using Protein and DNA Staining to Monitor Cell Stage in Synchronized Cells Paul Held, Ph.D., Laboratory Manager, Applications Department,
A p p l i c a t i o n N o t e Determination of Cell Cycle Time Using Protein and DNA Staining to Monitor Cell Stage in Synchronized Cells Paul Held, Ph.D., Laboratory Manager, Applications Department,
An Automated, Cell-based Platform for the Rapid Detection of Novel Androgen Receptor Modulators
 A p p l i c a t i o n N o t e An Automated, Cell-based Platform for the Rapid Detection of Novel Androgen Receptor Modulators Brad Larson and Peter Banks, BioTek Instruments, Inc., Winooski, VT Bruce Sherf,
A p p l i c a t i o n N o t e An Automated, Cell-based Platform for the Rapid Detection of Novel Androgen Receptor Modulators Brad Larson and Peter Banks, BioTek Instruments, Inc., Winooski, VT Bruce Sherf,
An Automated Walk-Away System to Perform Differentiation of 3D Mesenchymal Stem Cell Spheroids
 A p p l i c a t i o n N o t e An Automated Walk-Away System to Perform Differentiation of 3D Mesenchymal Stem Cell Spheroids Brad Larson, BioTek Instruments, Inc., Winooski, VT USA Glauco Souza, Nano3D
A p p l i c a t i o n N o t e An Automated Walk-Away System to Perform Differentiation of 3D Mesenchymal Stem Cell Spheroids Brad Larson, BioTek Instruments, Inc., Winooski, VT USA Glauco Souza, Nano3D
Assessment of HDAC1 Inhibition Using an Automated, Bioluminescent Histone Deacetylase I/II Assay
 A p p l i c a t i o n N o t e Assessment of HDAC1 Inhibition Using an Automated, Bioluminescent Histone Deacetylase I/II Assay Brad Larson, BioTek Instruments, Inc., Winooski, VT Tracy Worzella, Promega
A p p l i c a t i o n N o t e Assessment of HDAC1 Inhibition Using an Automated, Bioluminescent Histone Deacetylase I/II Assay Brad Larson, BioTek Instruments, Inc., Winooski, VT Tracy Worzella, Promega
EdU Cell Proliferation Kit. User Manual
 EdU Cell Proliferation Kit User Manual Ordering information: (for detailed kit content see Table 2) EdU Click Kits: Product number EdU Used fluorescent dye BCK-EdU488 BCK-EdU555 BCK-EdU594 BCK-EdU647 5
EdU Cell Proliferation Kit User Manual Ordering information: (for detailed kit content see Table 2) EdU Click Kits: Product number EdU Used fluorescent dye BCK-EdU488 BCK-EdU555 BCK-EdU594 BCK-EdU647 5
Automated Monitoring of Protein Expression and Metastatic Cell Migration using 3D Bioprinted Colorectal Cancer Cells
 A p p l i c a t i o n N o t e Automated Monitoring of Protein Expression and Metastatic Cell Migration using 3D Bioprinted Colorectal Cancer Cells Brad Larson, Leonie Rieger, BioTek Instruments, Inc.,
A p p l i c a t i o n N o t e Automated Monitoring of Protein Expression and Metastatic Cell Migration using 3D Bioprinted Colorectal Cancer Cells Brad Larson, Leonie Rieger, BioTek Instruments, Inc.,
HCS Mitotic Index Kit
 HCS Mitotic Index Kit Catalog no. H10293 Table 1. Contents and storage information. Material Amount Concentration Storage Stability phospho-h3 rabbit polyclonal antibody (Component A) 25 μl Not applicable
HCS Mitotic Index Kit Catalog no. H10293 Table 1. Contents and storage information. Material Amount Concentration Storage Stability phospho-h3 rabbit polyclonal antibody (Component A) 25 μl Not applicable
A Cost-effective Workflow for High-Throughput Screening of G- Protein Coupled Receptors (GPCRs)
 A p p l i c a t i o n N o t e A Cost-effective Workflow for High-Throughput Screening of G- Protein Coupled Receptors (GPCRs) Monitoring Receptor Mediated Calcium Flux Paul Held and Peter Banks, BioTek
A p p l i c a t i o n N o t e A Cost-effective Workflow for High-Throughput Screening of G- Protein Coupled Receptors (GPCRs) Monitoring Receptor Mediated Calcium Flux Paul Held and Peter Banks, BioTek
An Automated DELFIA ADCC Assay Method using a CD16.NK-92 Cell Line
 A p p l i c a t i o n N o t e An Automated DELFIA ADCC Assay Method using a CD16.NK-92 Cell Line Brad Larson, Senior Scientist, Applications Department, BioTek Instruments, Inc., Winooski, VT Introduction
A p p l i c a t i o n N o t e An Automated DELFIA ADCC Assay Method using a CD16.NK-92 Cell Line Brad Larson, Senior Scientist, Applications Department, BioTek Instruments, Inc., Winooski, VT Introduction
Monitoring Saccharomyces cerevisiea Growth with Brightfield Microscopy in Real Time
 A p p l i c a t i o n N o t e Monitoring Saccharomyces cerevisiea Growth with Brightfield Microscopy in Real Time Using the Lionheart FX and the ONIX 2 system to Image and Analyze Yeast Cell Growth Paul
A p p l i c a t i o n N o t e Monitoring Saccharomyces cerevisiea Growth with Brightfield Microscopy in Real Time Using the Lionheart FX and the ONIX 2 system to Image and Analyze Yeast Cell Growth Paul
Cell Synchrony Determination Using Microscopic Imaging
 A p p l i c a t i o n N o t e Cell Synchrony Determination Using Microscopic Imaging Use of Nuclear Staining to Assess Cell Cycle Stage by Nuclear DNA Content Paul Held Ph. D., Laboratory Manager, Applications
A p p l i c a t i o n N o t e Cell Synchrony Determination Using Microscopic Imaging Use of Nuclear Staining to Assess Cell Cycle Stage by Nuclear DNA Content Paul Held Ph. D., Laboratory Manager, Applications
Total Histone H3 Acetylation Detection Fast Kit (Fluorometric)
 Total Histone H3 Acetylation Detection Fast Kit (Fluorometric) Catalog Number KA1539 48 assays Version: 02 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Intended Use...
Total Histone H3 Acetylation Detection Fast Kit (Fluorometric) Catalog Number KA1539 48 assays Version: 02 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Intended Use...
Assay Name: Antibody-Dependent Drug Uptake Assay
 Assay Name: Antibody-Dependent Drug Uptake Assay Assay ID: Celigo_02_0019 Table of Contents Experiment: Antibody-Dependent Drug Uptake Assay... 2 Celigo Setup...2 Assay Protocol and Plate Setup...3 Results...5
Assay Name: Antibody-Dependent Drug Uptake Assay Assay ID: Celigo_02_0019 Table of Contents Experiment: Antibody-Dependent Drug Uptake Assay... 2 Celigo Setup...2 Assay Protocol and Plate Setup...3 Results...5
The Synergy 2, A Novel Approach to Microplate Multi- Detection for HTS and Drug Discovery
 The Synergy 2, A ovel Approach to Microplate Multi- Detection for HTS and Drug Discovery Paul Held and Xavier Amouretti, BioTek Instruments, Inc., Winooski, VT 05404 Abstract The Synergy 2 is a new type
The Synergy 2, A ovel Approach to Microplate Multi- Detection for HTS and Drug Discovery Paul Held and Xavier Amouretti, BioTek Instruments, Inc., Winooski, VT 05404 Abstract The Synergy 2 is a new type
Validation of a Novel Tumoroid-Based Cell Culture Model to Perform 3D in vitro Cell Signaling Analyses
 A p p l i c a t i o n N o t e Validation of a Novel Tumoroid-Based Cell Culture Model to Perform 3D in vitro Cell Signaling Analyses Comparison of Results Generated with an in vitro Cell Signaling Assay
A p p l i c a t i o n N o t e Validation of a Novel Tumoroid-Based Cell Culture Model to Perform 3D in vitro Cell Signaling Analyses Comparison of Results Generated with an in vitro Cell Signaling Assay
The Impact of a 3-Dimensional Human Liver Microtissue Model on Long-term Hepatotoxicity Studies
 A p p l i c a t i o n N o t e The Impact of a 3-Dimensional Human Liver Microtissue Model on Long-term Hepatotoxicity Studies Brad Larson and Peter Banks, BioTek Instruments, Inc., Winooski, VT Stewart
A p p l i c a t i o n N o t e The Impact of a 3-Dimensional Human Liver Microtissue Model on Long-term Hepatotoxicity Studies Brad Larson and Peter Banks, BioTek Instruments, Inc., Winooski, VT Stewart
An Image-Based Method to Detect and Quantify T Cell Mediated Cytotoxicity of 2D and 3D Target Cell Models
 A p p l i c a t i o n N o t e An Image-Based Method to Detect and Quantify T Cell Mediated Cytotoxicity of 2D and 3D Target Cell Models Brad Larson, Principal Scientist, BioTek Instruments, Inc., Winooski,
A p p l i c a t i o n N o t e An Image-Based Method to Detect and Quantify T Cell Mediated Cytotoxicity of 2D and 3D Target Cell Models Brad Larson, Principal Scientist, BioTek Instruments, Inc., Winooski,
SUPPLEMENTARY INFORMATION FIGURE LEGENDS
 SUPPLEMENTARY INFORMATION FIGURE LEGENDS Fig. S1. Radiation-induced phosphorylation of Rad50 at a specific site. A. Rad50 is an in vitro substrate for ATM. A series of Rad50-GSTs covering the entire molecule
SUPPLEMENTARY INFORMATION FIGURE LEGENDS Fig. S1. Radiation-induced phosphorylation of Rad50 at a specific site. A. Rad50 is an in vitro substrate for ATM. A series of Rad50-GSTs covering the entire molecule
Cell Health and Viability Assays Real-time automated measurements of cell health and viability inside your incubator
 INCUCYTE LIVE-CELL ANALYSIS SYSTEM Cell Health and Viability Assays Real-time automated measurements of cell health and viability inside your incubator See what your cells are doing and when they do it
INCUCYTE LIVE-CELL ANALYSIS SYSTEM Cell Health and Viability Assays Real-time automated measurements of cell health and viability inside your incubator See what your cells are doing and when they do it
Assay Name: HPC proliferation measurement using Ki-67 cellular marker
 Assay Name: HPC proliferation measurement using Ki-67 cellular marker Assay ID: Celigo_02_0014 Table of Contents Experiment: HPC proliferation measurement using Ki-67 cellular marker... 2 Celigo Setup...2
Assay Name: HPC proliferation measurement using Ki-67 cellular marker Assay ID: Celigo_02_0014 Table of Contents Experiment: HPC proliferation measurement using Ki-67 cellular marker... 2 Celigo Setup...2
SUPPLEMENTARY INFORMATION
 Figure S1: Activation of the ATM pathway by I-PpoI. A. HEK293T cells were either untransfected, vector transfected, transfected with an I-PpoI expression vector, or subjected to 2Gy γ-irradiation. 24 hrs
Figure S1: Activation of the ATM pathway by I-PpoI. A. HEK293T cells were either untransfected, vector transfected, transfected with an I-PpoI expression vector, or subjected to 2Gy γ-irradiation. 24 hrs
WHEN MULTIMODE MEETS IMAGING YOU GAIN A WHOLE NEW PERSPECTIVE. EnSight Multimode Plate Reader
 WHEN MULTIMODE MEETS IMAGING YOU GAIN A WHOLE NEW PERSPECTIVE EnSight Multimode Plate Reader TRANSLATING RELEVANT RESULTS INTO REAL INSIGHTS 2 THE WAY TO GREATER CONFIDENCE IN YOUR RESULTS Today s leading
WHEN MULTIMODE MEETS IMAGING YOU GAIN A WHOLE NEW PERSPECTIVE EnSight Multimode Plate Reader TRANSLATING RELEVANT RESULTS INTO REAL INSIGHTS 2 THE WAY TO GREATER CONFIDENCE IN YOUR RESULTS Today s leading
A Cell-based DELFIA Assay for Measuring Phosphorylation of Proteins
 A Cell-based DELFIA Assay for Measuring Phosphorylation of Proteins Sofia Vikström, Christel Gripenberg-Lerche, Pertti Hurskainen and Ilkka Hemmilä PerkinElmer Life and Analytical Sciences, Wallac Oy,
A Cell-based DELFIA Assay for Measuring Phosphorylation of Proteins Sofia Vikström, Christel Gripenberg-Lerche, Pertti Hurskainen and Ilkka Hemmilä PerkinElmer Life and Analytical Sciences, Wallac Oy,
Synergy NEO HTS Multi-Mode Microplate Reader and Life Technologies Reagents
 A p p l i c a t i o n G u i d e Synergy NEO HTS Multi-Mode Microplate Reader and Life Technologies Reagents High Performance Multi-Detection Capabilities that Span the Small Molecule Drug Discovery Process
A p p l i c a t i o n G u i d e Synergy NEO HTS Multi-Mode Microplate Reader and Life Technologies Reagents High Performance Multi-Detection Capabilities that Span the Small Molecule Drug Discovery Process
< Supporting Information >
 SUPPORTING INFORMATION 1 < Supporting Information > Discovery of autophagy modulators through the construction of high-content screening platform via monitoring of lipid droplets Sanghee Lee, Eunha Kim,
SUPPORTING INFORMATION 1 < Supporting Information > Discovery of autophagy modulators through the construction of high-content screening platform via monitoring of lipid droplets Sanghee Lee, Eunha Kim,
EpiQuik Global Histone H3 Phosphorylation (Ser28) Assay Kit (Colorimetric)
 EpiQuik Global Histone H3 Phosphorylation (Ser28) Assay Kit (Colorimetric) Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Global Histone H3 Phosphorylation (Ser28) Assay Kit (Colorimetric)
EpiQuik Global Histone H3 Phosphorylation (Ser28) Assay Kit (Colorimetric) Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Global Histone H3 Phosphorylation (Ser28) Assay Kit (Colorimetric)
Multiplex Fluorescence Assays for Adherence Cells without Trypsinization
 Multiplex Fluorescence Assays for Adherence Cells without Trypsinization The combination of a bright field and three fluorescent channels allows the Celigo to perform many multiplexed assays. A gating
Multiplex Fluorescence Assays for Adherence Cells without Trypsinization The combination of a bright field and three fluorescent channels allows the Celigo to perform many multiplexed assays. A gating
Automation of HTRF elf4e Kinase Assay Using a 3D Tumoroid- Based Cell Model
 A p p l i c a t i o n N o t e Automation of HTRF elf4e Kinase Assay Using a 3D Tumoroid- Based Cell Model Incorporation of the MultiFlo FX Microplate Dispenser to Create 3D Cell Cultures using the RAFT
A p p l i c a t i o n N o t e Automation of HTRF elf4e Kinase Assay Using a 3D Tumoroid- Based Cell Model Incorporation of the MultiFlo FX Microplate Dispenser to Create 3D Cell Cultures using the RAFT
Development of an AlphaLISA MEK1 Kinase Assay Using Full-Length ERK2 Substrate
 TECHNICAL NOTE Development of an AlphaLISA MEK1 Kinase Assay Using Full-Length ERK2 Substrate AlphaLISA Technology Author: Jeanine Hinterneder, PhD PerkinElmer, Inc. Hopkinton, MA Introduction The mitogen-activated
TECHNICAL NOTE Development of an AlphaLISA MEK1 Kinase Assay Using Full-Length ERK2 Substrate AlphaLISA Technology Author: Jeanine Hinterneder, PhD PerkinElmer, Inc. Hopkinton, MA Introduction The mitogen-activated
Application Note. Imaging and Evaluation of DNA Damage Employing the γh2ax-assay with CELLAVISTA and the YT-software. Abstract AN-F225-XVIII-01
 1 Imaging and Evaluation of DNA Damage Employing the γh2ax-assay with CELLAVISTA and the YT-software Lückerath F 1, CHRISTMANN T 1, GEISEN R 1, LÜKE J 1, THIES F 1 AND WERDELMANN B 1 [1] SYNENTEC GMBH
1 Imaging and Evaluation of DNA Damage Employing the γh2ax-assay with CELLAVISTA and the YT-software Lückerath F 1, CHRISTMANN T 1, GEISEN R 1, LÜKE J 1, THIES F 1 AND WERDELMANN B 1 [1] SYNENTEC GMBH
The Nuclear Area Factor (NAF): a measure for cell apoptosis using microscopy and image analysis
 The Nuclear Area Factor (NAF): a measure for cell apoptosis using microscopy and image analysis Mark A. DeCoster Department of Biomedical Engineering and Institute for Micromanufacturing, Louisiana Tech
The Nuclear Area Factor (NAF): a measure for cell apoptosis using microscopy and image analysis Mark A. DeCoster Department of Biomedical Engineering and Institute for Micromanufacturing, Louisiana Tech
Optimization of a Multi-Mode Detection Model for Measuring Real-time Cellular Respiration and Mitochondrial Function using Fluorophoric Biosensors
 A p p l i c a t i o n N o t e Optimization of a Multi-Mode Detection Model for Measuring Real-time Cellular Respiration and Mitochondrial Function using Fluorophoric Biosensors Wendy Goodrich, Applications
A p p l i c a t i o n N o t e Optimization of a Multi-Mode Detection Model for Measuring Real-time Cellular Respiration and Mitochondrial Function using Fluorophoric Biosensors Wendy Goodrich, Applications
Adapta Assay Setup Guide on the BioTek Instruments Synergy 2 Multi-Mode Microplate Reader
 Page 1 of 19 Adapta Assay Setup Guide on the BioTek Instruments Synergy 2 Multi-Mode Microplate Reader NOTE: The BioTek Instruments Synergy 2 Multi-Mode Microplate Reader was tested for compatibility with
Page 1 of 19 Adapta Assay Setup Guide on the BioTek Instruments Synergy 2 Multi-Mode Microplate Reader NOTE: The BioTek Instruments Synergy 2 Multi-Mode Microplate Reader was tested for compatibility with
Cholesterol Uptake Cell-Based Assay Kit
 Cholesterol Uptake Cell-Based Assay Kit Item No. 600440 www.caymanchem.com Customer Service 800.364.9897 Technical Support 888.526.5351 1180 E. Ellsworth Rd Ann Arbor, MI USA TABLE OF CONTENTS GENERAL
Cholesterol Uptake Cell-Based Assay Kit Item No. 600440 www.caymanchem.com Customer Service 800.364.9897 Technical Support 888.526.5351 1180 E. Ellsworth Rd Ann Arbor, MI USA TABLE OF CONTENTS GENERAL
UV Fluorescence Polarization as a Means to Investigate Protein Conformational and Mass Change
 A p p l i c a t i o n N o t e UV Fluorescence Polarization as a Means to Investigate Protein Conformational and Mass Change Using Intrinsic Tryptophan Fluorescence in Conjunction with UV-capable Polarizers
A p p l i c a t i o n N o t e UV Fluorescence Polarization as a Means to Investigate Protein Conformational and Mass Change Using Intrinsic Tryptophan Fluorescence in Conjunction with UV-capable Polarizers
FIGURE S1. Representative images to illustrate RAD51 foci induction in FANCD2 wild-type (wt) and
 Supplementary Figures FIGURE S1. Representative images to illustrate RAD51 foci induction in FANCD2 wild-type (wt) and mutant (mut) cells. Higher magnification is shown in Figure 1A. Arrows indicate cisplatin-induced
Supplementary Figures FIGURE S1. Representative images to illustrate RAD51 foci induction in FANCD2 wild-type (wt) and mutant (mut) cells. Higher magnification is shown in Figure 1A. Arrows indicate cisplatin-induced
Supplementary Information
 Supplementary Information stability is regulated by CK2-dependent interaction with R2TP complex Patrick von Morgen 1,2, Kamila Burdova 1, Thomas G. Flower 3, Nicola J. O'Reilly 4, Simon J. Boulton 5, Stephen
Supplementary Information stability is regulated by CK2-dependent interaction with R2TP complex Patrick von Morgen 1,2, Kamila Burdova 1, Thomas G. Flower 3, Nicola J. O'Reilly 4, Simon J. Boulton 5, Stephen
EarlyTox Cell Integrity Kit
 EarlyTox Cell Integrity Kit The EarlyTox Cell Integrity Kit from Molecular Devices is an optimized set of reagents that simplifies the measurement of live and dead cells in a single well. The assay uses
EarlyTox Cell Integrity Kit The EarlyTox Cell Integrity Kit from Molecular Devices is an optimized set of reagents that simplifies the measurement of live and dead cells in a single well. The assay uses
Synergy, Cytation and FLx800 Multi-Mode Microplate Readers
 Fluorescence Filter, Cube and Mirror List Synergy, Cytation and FLx800 Multi-Mode Microplate Readers Use this Guide to find part numbers and other details for filters, filter cubes and dichroic mirrors
Fluorescence Filter, Cube and Mirror List Synergy, Cytation and FLx800 Multi-Mode Microplate Readers Use this Guide to find part numbers and other details for filters, filter cubes and dichroic mirrors
Sarker et al. Supplementary Material. Subcellular Fractionation
 Supplementary Material Subcellular Fractionation Transfected 293T cells were harvested with phosphate buffered saline (PBS) and centrifuged at 2000 rpm (500g) for 3 min. The pellet was washed, re-centrifuged
Supplementary Material Subcellular Fractionation Transfected 293T cells were harvested with phosphate buffered saline (PBS) and centrifuged at 2000 rpm (500g) for 3 min. The pellet was washed, re-centrifuged
Four different active promoter genes were chosen, ATXN7L2, PSRC1, CELSR2 and
 SUPPLEMENTARY MATERIALS AND METHODS Chromatin Immunoprecipitation for qpcr analysis Four different active promoter genes were chosen, ATXN7L2, PSRC1, CELSR2 and IL24, all located on chromosome 1. Primer
SUPPLEMENTARY MATERIALS AND METHODS Chromatin Immunoprecipitation for qpcr analysis Four different active promoter genes were chosen, ATXN7L2, PSRC1, CELSR2 and IL24, all located on chromosome 1. Primer
ATPlite Assay Performance in Human Primary Cells
 A P P L I C AT I O N N O T E Cell Viability Assays Author: Verena Brucklacher-Waldert Crescendo Biologics Cambridge, UK Assay Performance in Human Primary Cells Introduction In vitro assays using primary
A P P L I C AT I O N N O T E Cell Viability Assays Author: Verena Brucklacher-Waldert Crescendo Biologics Cambridge, UK Assay Performance in Human Primary Cells Introduction In vitro assays using primary
Performance of cell viability and cytotoxicity assays on the IN Cell Analyzer 3000
 GE Healthcare Application Note 28-4070-51 AA IN Cell Analyzer 3000 Performance of cell viability and cytotoxicity assays on the IN Cell Analyzer 3000 Key words: cell-based assay viability cytotoxicity
GE Healthcare Application Note 28-4070-51 AA IN Cell Analyzer 3000 Performance of cell viability and cytotoxicity assays on the IN Cell Analyzer 3000 Key words: cell-based assay viability cytotoxicity
Validation and Comparison of Single-Step Flash and Dual-Spectral Luciferase Reporter Gene Assays using the Synergy Line of Microplate Readers
 A p p l i c a t i o n N o t e Validation and Comparison of Single-Step Flash and Dual-Spectral Luciferase Reporter Gene Assays using the Synergy Line of Microplate Readers Peter J. Brescia and Peter Banks,
A p p l i c a t i o n N o t e Validation and Comparison of Single-Step Flash and Dual-Spectral Luciferase Reporter Gene Assays using the Synergy Line of Microplate Readers Peter J. Brescia and Peter Banks,
Quantitation of ssdna using OliGreen Fluorescent Stain
 Quantitation of ssdna using OliGreen Fluorescent Stain Several different techniques require the use of short synthetic oligonucleotide molecules, often referred to as primers. In each case, the use of
Quantitation of ssdna using OliGreen Fluorescent Stain Several different techniques require the use of short synthetic oligonucleotide molecules, often referred to as primers. In each case, the use of
Automated, High Throughput, HTRF -Based Detection of Histone Methyltransferase and Demethylase Enzyme Activity
 A p p l i c a t i o n N o t e Automated, High Throughput, HTRF -Based Detection of Histone Methyltransferase and Demethylase Enzyme Activity Brad Larson and Peter Banks, Applications Department, BioTek
A p p l i c a t i o n N o t e Automated, High Throughput, HTRF -Based Detection of Histone Methyltransferase and Demethylase Enzyme Activity Brad Larson and Peter Banks, Applications Department, BioTek
Peter Brescia, Applications Scientist and Peter Banks, Scientific Director, BioTek Instruments, Inc.
 Guidelines for the Provision of Residual Volumes of Assay Buffer in Microplate Wash Assays Setting aspiration manifold z-heights in ELx45 / EL46 software to provide predetermined residual volumes in the
Guidelines for the Provision of Residual Volumes of Assay Buffer in Microplate Wash Assays Setting aspiration manifold z-heights in ELx45 / EL46 software to provide predetermined residual volumes in the
EPIGENTEK. EpiQuik Global Acetyl Histone H3- K14 Quantification Kit (Fluorometric) Base Catalog # P-4013 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE
 EpiQuik Global Acetyl Histone H3- K14 Quantification Kit (Fluorometric) Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Global Acetyl Histone H3-K14 Quantification Kit (Fluorometric)
EpiQuik Global Acetyl Histone H3- K14 Quantification Kit (Fluorometric) Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Global Acetyl Histone H3-K14 Quantification Kit (Fluorometric)
High-throughput physical phenotyping of cell differentiation
 Supplementary file High-throughput physical phenotyping of cell differentiation Jonathan Lin1,*, Donghyuk Kim1,*, Henry Tse2, Peter Tseng3, Lillian Peng1, Manjima Dhar1, Saravanan Karumbayaram4 and Dino
Supplementary file High-throughput physical phenotyping of cell differentiation Jonathan Lin1,*, Donghyuk Kim1,*, Henry Tse2, Peter Tseng3, Lillian Peng1, Manjima Dhar1, Saravanan Karumbayaram4 and Dino
EGFR (Phospho-Ser695)
 Assay Biotechnology Company www.assaybiotech.com Tel: 1-877-883-7988 Fax: 1-877-610-9758 EGFR (Phospho-Ser695) Colorimetric Cell-Based ELISA Kit Catalog #: OKAG02090 Please read the provided manual entirely
Assay Biotechnology Company www.assaybiotech.com Tel: 1-877-883-7988 Fax: 1-877-610-9758 EGFR (Phospho-Ser695) Colorimetric Cell-Based ELISA Kit Catalog #: OKAG02090 Please read the provided manual entirely
Segments of the obstructed intestinal loops were fixed in 4% paraformaldehyde
 Supplementary text Supplementary materials and methods Histopathological examination Segments of the obstructed intestinal loops were fixed in 4% paraformaldehyde (PFA) and embedded in paraffin wax with
Supplementary text Supplementary materials and methods Histopathological examination Segments of the obstructed intestinal loops were fixed in 4% paraformaldehyde (PFA) and embedded in paraffin wax with
IncuCyte Live-Cell Immunocytochemistry Assay
 IncuCyte Live-Cell Immunocytochemistry Assay For cell surface marker analysis This protocol describes a solution for measuring immunocytochemistry in live-cells expressing a surface antigen of interest.
IncuCyte Live-Cell Immunocytochemistry Assay For cell surface marker analysis This protocol describes a solution for measuring immunocytochemistry in live-cells expressing a surface antigen of interest.
Versatile, Cost-Effective Automation of Avian Influenza and Mycoplasma Gallispeticum-Synovaie ELISAs
 Versatile, Cost-Effective Automation of Avian Influenza and Mycoplasma Gallispeticum-Synovaie ELISAs Wendy Goodrich, Applications Scientist, BioTek Instruments, Inc. Matt Durgin, PAS Technical Service
Versatile, Cost-Effective Automation of Avian Influenza and Mycoplasma Gallispeticum-Synovaie ELISAs Wendy Goodrich, Applications Scientist, BioTek Instruments, Inc. Matt Durgin, PAS Technical Service
Cellomics ToxInsight Phospho-ATM Activation Cartridge
 INSTRUCTIONS Cellomics ToxInsight Phospho-ATM Activation Cartridge High-Content Imaging Reagents Number Description 8408702 Phospho-ATM Activation Cartridge, sufficient materials for 5 96 wells Contents:
INSTRUCTIONS Cellomics ToxInsight Phospho-ATM Activation Cartridge High-Content Imaging Reagents Number Description 8408702 Phospho-ATM Activation Cartridge, sufficient materials for 5 96 wells Contents:
ThiolTracker Violet (Glutathione Detection Reagent)
 ThiolTracker Violet (Glutathione Detection Reagent) Catalog nos. T10095, T10096 Table 1. Contents and storage information. Material Amount Concentration Storage* Stability ThiolTracker Violet 180 assays
ThiolTracker Violet (Glutathione Detection Reagent) Catalog nos. T10095, T10096 Table 1. Contents and storage information. Material Amount Concentration Storage* Stability ThiolTracker Violet 180 assays
Neural Stem Cell Characterization Kit
 Neural Stem Cell Characterization Kit Catalog No. SCR019 FOR RESEARCH USE ONLY Not for use in diagnostic procedures USA & Canada Phone: +1(800) 437-7500 Fax: +1 (951) 676-9209 Europe +44 (0) 23 8026 2233
Neural Stem Cell Characterization Kit Catalog No. SCR019 FOR RESEARCH USE ONLY Not for use in diagnostic procedures USA & Canada Phone: +1(800) 437-7500 Fax: +1 (951) 676-9209 Europe +44 (0) 23 8026 2233
ab CytoPainter ER Staining Kit Red Fluorescence
 ab139482 CytoPainter ER Staining Kit Red Fluorescence Instructions for Use Designed to detect Human endoplasmic reticulum by microscopy. This product is for research use only and is not intended for diagnostic
ab139482 CytoPainter ER Staining Kit Red Fluorescence Instructions for Use Designed to detect Human endoplasmic reticulum by microscopy. This product is for research use only and is not intended for diagnostic
The BioFlux 200 System Using Well Plate Microfluidics for Live Cell Assays Product Overview and Tutorial
 The BioFlux 200 System Using Well Plate Microfluidics for Live Cell Assays Product Overview and Tutorial Introduction to the BioFlux System Enables live-cell assays with precisely-controlled shear flow
The BioFlux 200 System Using Well Plate Microfluidics for Live Cell Assays Product Overview and Tutorial Introduction to the BioFlux System Enables live-cell assays with precisely-controlled shear flow
Fluorescence Microscopy. Terms and concepts to know: 10/11/2011. Visible spectrum (of light) and energy
 Fluorescence Microscopy Louisiana Tech University Ruston, Louisiana Microscopy Workshop Dr. Mark DeCoster Associate Professor Biomedical Engineering 1 Terms and concepts to know: Signal to Noise Excitation
Fluorescence Microscopy Louisiana Tech University Ruston, Louisiana Microscopy Workshop Dr. Mark DeCoster Associate Professor Biomedical Engineering 1 Terms and concepts to know: Signal to Noise Excitation
Limit of Detection. Theoretical Lowest Concentration
 Nucleic Acid Quantitation Detection Limits Today s biomedical testing has resulted in sample sizes becoming smaller and smaller, driving the need to measure samples with ever-lower detection limits. The
Nucleic Acid Quantitation Detection Limits Today s biomedical testing has resulted in sample sizes becoming smaller and smaller, driving the need to measure samples with ever-lower detection limits. The
TECHNICAL NOTE THE NEXT GENERATION OF MICROPLATES
 TECHNICAL NOTE TN0002: Optimiser Microplate System (ELISA) Setup Guide on the BioTek FLx800 Fluorescence Microplate Reader THE NEXT GENERATION OF MICROPLATES Better Immunoassays through Innovative Microfluidics
TECHNICAL NOTE TN0002: Optimiser Microplate System (ELISA) Setup Guide on the BioTek FLx800 Fluorescence Microplate Reader THE NEXT GENERATION OF MICROPLATES Better Immunoassays through Innovative Microfluidics
Automating a Direct, Cell-Based, Target-Compound Interaction Assay for G9a Histone Methyltransferase and Bromodomain Proteins
 A p p l i c a t i o n N o t e Automating a Direct, Cell-Based, Target-Compound Interaction Assay for G9a Histone Methyltransferase and Bromodomain Proteins Brad Larson and Peter Banks, BioTek Instruments,
A p p l i c a t i o n N o t e Automating a Direct, Cell-Based, Target-Compound Interaction Assay for G9a Histone Methyltransferase and Bromodomain Proteins Brad Larson and Peter Banks, BioTek Instruments,
The CQ1 Confocal Quantitative Image Cytometer and its Application to Biological Measurement
 The CQ1 Confocal Quantitative Image Cytometer and its Application to Biological Measurement Hirofumi Sakashita *1 Koji Ohashi *1 Kazuo Ozawa *2 ohei Tsubouchi *1 The CQ1 confocal quantitative image cytometer,
The CQ1 Confocal Quantitative Image Cytometer and its Application to Biological Measurement Hirofumi Sakashita *1 Koji Ohashi *1 Kazuo Ozawa *2 ohei Tsubouchi *1 The CQ1 confocal quantitative image cytometer,
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/NCHEM.1805 Visualization and Selective Chemical Targeting of RNA G-quadruplex Structures in the Cytoplasm of Human Cells Giulia Biffi 1, Marco Di Antonio 2, David Tannahill 1 and Shankar Balasubramanian
DOI: 10.1038/NCHEM.1805 Visualization and Selective Chemical Targeting of RNA G-quadruplex Structures in the Cytoplasm of Human Cells Giulia Biffi 1, Marco Di Antonio 2, David Tannahill 1 and Shankar Balasubramanian
T H E J O U R N A L O F C E L L B I O L O G Y
 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Monteiro et al., http://www.jcb.org/cgi/content/full/jcb.201306162/dc1 Figure S1. 3D deconvolution microscopy analysis of WASH and exocyst
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Monteiro et al., http://www.jcb.org/cgi/content/full/jcb.201306162/dc1 Figure S1. 3D deconvolution microscopy analysis of WASH and exocyst
Optimizing Dual-Glo Luciferase Assays with the Synergy HT Multi-Detection Microplate Reader
 Optimizing Dual-Glo Luciferase Assays with the Synergy HT Multi-Detection Microplate Reader Introduction Today s biological science and drug discovery research often involves the measurement of large numbers
Optimizing Dual-Glo Luciferase Assays with the Synergy HT Multi-Detection Microplate Reader Introduction Today s biological science and drug discovery research often involves the measurement of large numbers
MitoBiogenesis In-Cell ELISA Kit (Colorimetric)
 PROTOCOL MitoBiogenesis In-Cell ELISA Kit (Colorimetric) DESCRIPTION 1850 Millrace Drive, Suite 3A Eugene, Oregon 97403 MS643 Rev.2 For identifying inhibitors and activators of mitochondrial biogenesis
PROTOCOL MitoBiogenesis In-Cell ELISA Kit (Colorimetric) DESCRIPTION 1850 Millrace Drive, Suite 3A Eugene, Oregon 97403 MS643 Rev.2 For identifying inhibitors and activators of mitochondrial biogenesis
MF-ChemiBIS. Today s most comprehensive solution for your bio-imaging needs and applications. Documenting Nature
 MF-ChemiBIS Today s most comprehensive solution for your bio-imaging needs and applications Documenting Nature MF-ChemiBIS Excellence in bio-imaging The DNR Advantage As pioneers in bio-imaging technologies
MF-ChemiBIS Today s most comprehensive solution for your bio-imaging needs and applications Documenting Nature MF-ChemiBIS Excellence in bio-imaging The DNR Advantage As pioneers in bio-imaging technologies
Cellular Energy Flux in Real Time
 Cellular Energy Flux in Real Time Cytation 3 Imaging Reader MitoXpress Metabolism & Mitochondrial Function Assays MitoXpress Metabolism & Mitochondrial Function Assays are available from Optimization of
Cellular Energy Flux in Real Time Cytation 3 Imaging Reader MitoXpress Metabolism & Mitochondrial Function Assays MitoXpress Metabolism & Mitochondrial Function Assays are available from Optimization of
Detection and study of the formation and repair of DNA double-strand breaks after irradiation
 Laboratory of Radiation Biology (LRB) Detection and study of the formation and repair of DNA double-strand breaks after irradiation Students: Ioana-Cezara Bucataru Bianca-Gabriela Zota Katarína Žirová
Laboratory of Radiation Biology (LRB) Detection and study of the formation and repair of DNA double-strand breaks after irradiation Students: Ioana-Cezara Bucataru Bianca-Gabriela Zota Katarína Žirová
0.5% Triton X-100 for 5 min at room temperature. Fixed and permeabilized cells were
 1 Supplementary Methods Immunohistochemistry EBC-1 cells were fixed in 4% paraformaldehyde for 15 min at room temperature, followed by 0.5% Triton X-100 for 5 min at room temperature. Fixed and permeabilized
1 Supplementary Methods Immunohistochemistry EBC-1 cells were fixed in 4% paraformaldehyde for 15 min at room temperature, followed by 0.5% Triton X-100 for 5 min at room temperature. Fixed and permeabilized
USER GUIDE. Introduction. Catalog No. C10730, C10731
 USER GUIDE CellEvent Caspase-3/7 Red Detection Reagent Catalog No. C10730, C10731 Pub. No. MAN0016090 Rev. B.0 Table 1. Contents and storage Material C10730 Amount C10731 Concentration Storage* CellEvent
USER GUIDE CellEvent Caspase-3/7 Red Detection Reagent Catalog No. C10730, C10731 Pub. No. MAN0016090 Rev. B.0 Table 1. Contents and storage Material C10730 Amount C10731 Concentration Storage* CellEvent
Automated, High Throughput Methods for the Quantification of Endogenous Cellular Kinase Activity using HTRF Technology
 A p p l i c a t i o n N o t e Automated, High Throughput Methods for the Quantification of Endogenous Cellular Kinase Activity using HTRF Technology Brad Larson and Peter Banks, Applications Department,
A p p l i c a t i o n N o t e Automated, High Throughput Methods for the Quantification of Endogenous Cellular Kinase Activity using HTRF Technology Brad Larson and Peter Banks, Applications Department,
