Borrelia burgdorferi Gene Expression In Vivo and Spirochete Pathogenicity
|
|
|
- Mervyn Woods
- 6 years ago
- Views:
Transcription
1 INFECTION AND IMMUNITY, Mar. 2000, p Vol. 68, No /00/$ Copyright 2000, American Society for Microbiology. All Rights Reserved. Borrelia burgdorferi Gene Expression In Vivo and Spirochete Pathogenicity JUAN ANGUITA, 1 SWAPNA SAMANTA, 1 BEATRIZ REVILLA, 1 KYOUNGHO SUK, 1 SUBRATA DAS, 1 STEPHEN W. BARTHOLD, 2 AND EROL FIKRIG 1 * Section of Rheumatology, Department of Internal Medicine, Yale University School of Medicine, New Haven, Connecticut 06520, 1 and Center for Comparative Medicine, Schools of Medicine and Veterinary Medicine, University of California at Davis, Davis, California Received 13 October 1999/Returned for modification 19 November 1999/Accepted 30 November 1999 Borrelia burgdorferi spirochetes that do not cause arthritis or carditis were developed and used to investigate Lyme disease pathogenesis. A clonal isolate of B. burgdorferi N40 (cn40), which induces disease in C3H/HeN (C3H) mice, was repeatedly passaged in vitro to generate nonpathogenic spirochetes. The passage 75 isolate (N40-75) was infectious for C3H mice but did not cause arthritis or carditis, and spirochetes were at low levels or absent in the joints or hearts, respectively. N40-75 could, however, cause disease in severe combined immunodeficient (SCID) mice, suggesting that the response in immunocompetent mice prevented effective spirochete dissemination and the subsequent development of arthritis and carditis. Administration of immune sera at 4 days after spirochete challenge aborted N40-75, but not cn40, infection in SCID mice. A B. burgdorferi genomic expression library was differentially probed with sera from cn40- and N40-75-infected mice, to identify genes that may not be effectively expressed by N40-75 in vivo. N40-75 was defective in the up-regulation of several genes that are preferentially expressed during mammalian infection, including dbpab, bba64, and genes that map to the cp32 family of plasmids. These data suggest that adaptation and gene expression may be required for B. burgdorferi to effectively colonize the host, evade humoral responses, and cause disease. Borrelia burgdorferi, the causative agent of Lyme disease, is the most common vector-borne pathogen in the United States (15). B. burgdorferi-infected mice develop arthritis and carditis, thereby partially mimicking the human illness (3, 5, 8, 10, 19, 43). The pathogenesis of murine Lyme borreliosis is multifactorial. The severity of disease has been associated with the number of B. burgdorferi in the affected organs (57), the virulence of specific spirochete isolates (12, 32, 40), and the host immune response to this organism (2, 11, 33, 35, 37, 53, 56). The severity of murine Lyme arthritis is genotype dependent. For example, C3H/HeN (C3H) mice develop more severe arthritis and have greater numbers of spirochetes in the joints than BALB/c mice (5, 36, 44). B. burgdorferi spirochetes adapt to different environments during their life cycle in Ixodes scapularis ticks and the reservoir host. Within engorging ticks, the B. burgdorferi protein expression dramatically changes, with the synthesis of OspC and down-regulation of OspA (21, 46). These changes are likely to occur in response to the incoming blood meal, temperature shifts, and alterations in spirochete density (21, 42, 49). In the mammalian host, B. burgdorferi must evade host antibodies that arise in response to infection, and selective gene expression and recombination events may facilitate spirochete survival (21, 58 60). Several genes, including eppa, ospe/f paralogues (erps), bbk32, bbk50, and the decorin-binding protein gene dbpa, are not expressed, or are expressed at low levels, by B. burgdorferi cultured in laboratory medium but are apparently up-regulated during infection (1, 16, 25, 51, 54). * Corresponding author. Mailing address: 608 Laboratory of Clinical Investigation, Section of Rheumatology, Department of Internal Medicine, Yale University School of Medicine, 333 Cedar St., New Haven, CT Phone: (203) Fax: (203) ef6@ .med.yale.edu. Present address: Samsung Biomedical Research Institute, Clinical Research Center, Seoul, South Korea The function of most of these conditionally expressed gene products is not known. OspA may be one of several plasminogen-binding proteins (28), DbpA adheres to decorin (30), and BBK32 mediates the attachment of B. burgdorferi to fibronectin (41). The examination of spirochetes continuously cultured in vitro has enhanced our understanding of B. burgdorferi pathogenesis. The sequential passage of uncloned B. burgdorferi in Barbour-Stoenner-Kelly (BSK) II medium results in selection of noninfectious subpopulations with variable plasmid and protein profiles (4, 34, 39, 45, 47). This suggests that specific genes are required for infection. A correlation between plasmid content, or specific B. burgdorferi genes, and infectivity has more recently been demonstrated using cloned spirochetes. Xu and colleagues have shown that specific plasmids are associated with the infectivity of B. burgdorferi B31 (55). Zhang and associates (58) used subtractive hybridization to identify a plasmid-encoded gene family, designated the vmp-like sequence locus (vls), which undergo antigenic variation by promiscuous recombination. This gene family is present in an infectious, clonal isolate of B. burgdorferi B31 but not in clonal in vitropassaged organisms, suggesting that it is necessary for spirochete infectivity (58). A clonal isolate of B. burgdorferi N40 (cn40) which causes severe arthritis and carditis in C3H mice has now been passaged in BSK II medium. Spirochetes passaged 75 times in vitro, designated N40-75, were investigated for infectivity and pathogenicity and compared to the parental isolate. The lack of arthritis and carditis was then correlated to a defect in differential N40-75 gene expression in vivo, resulting in an inability of N40-75 to rapidly adapt to the murine B. burgdorferi-specific antibody response. MATERIALS AND METHODS Mice. C3H, C57BL/6 (B6), C3H severe combined immunodeficient (C3Hscid), and C.B-17-scid mice were purchased from the Frederick Cancer Research 1222
2 VOL. 68, 2000 B. BURGDORFERI GENE EXPRESSION AND PATHOGENICITY 1223 Center, Frederick, Md. Mice were kept in filter-framed cages and euthanized with CO 2. B. burgdorferi and infections. cn40 is a clonal isolate of B. burgdorferi N40 that is infectious and pathogenic in mice (9). Spirochetes were grown in BSK II medium at 33 C. To obtain derivatives of cn40, B. burgdorferi spirochetes were serially passaged in BSK II medium every 3 to 5 days by inoculating 50 l of grown cultures into 7 ml of fresh medium. Clones of N40-75 were obtained by subsurface culture in solid BSK medium, as reported previously (58, 60). Every 5 to 10 passages, spirochetes were assessed for infectivity and the ability to induce carditis and arthritis in 3- to 6-week-old C3H mice. Individual mice in groups of five were inoculated with 10 4 or 10 7 spirochetes in the midline of the back by intradermal injection (26). The animals were necropsied at 21 and 60 days, time points that represent the peak of acute disease and the regression phase of inflammation (10), respectively. Mice were assessed for infection by culturing specimens of the blood, urinary bladder, and skin (at the inoculation site) in BSK II medium at 33 C (24). Cultures were read after 14 days, a time period sufficient to allow a single spirochete to grow to stationary phase. Quantitation of spirochetes in vitro. Quantitation of the spirochetes grown in vitro was performed as follows. Cultures were spun down and washed twice with phosphate-buffered saline (PBS). The spirochetes were resuspended in PBS, and the absorbance at 600 nm was determined. The absorbance at 600 nm was shown in parallel experiments to correlate with the number of spirochetes, as counted by dark-field microscopy, in a linear manner. One unit of absorbance at 600 nm was determined to equal spirochetes/ml. Histopathology of murine Lyme disease. Formalin-fixed, paraffin-embedded hearts and joints (both knees and tibiotarsi) were examined microscopically for evidence of disease (23). Arthritis and carditis were assessed as previously described (3, 5). All histopathologic assessments were made in a blinded fashion. In vivo serum treatments. Relative susceptibility of cn40 and N40-75 spirochetes to immune serum in the murine host was studied in C3H-scid mice. Mice were administered 50 l of immune sera at the time of spirochete inoculation and 4 and 8 days after infection (7, 20). Immune sera were obtained from immunocompetent C3H mice that had been infected with cn40 or N40-75 for 21 days. C3H-scid mice were necropsied 14 days after challenge and assessed for infection. DNA extraction and PCR. Cultured B. burgdorferi DNA was obtained by proteinase K digestion (100 g/ml, overnight at 55 C) followed by phenolchloroform-isoamyl alcohol (25:24:1) extraction and ethanol precipitation. PCR was performed on DNA samples using primers for flab and p39 (chromosomal targets), ospa (lp54 plasmid), erpt (lp28-1), bbk50 (lp36), ospd (lp38), ospc (cp26) and erps (cp32 family of plasmids [52]). Detection and quantitation of B. burgdorferi in infected tissues. DNA was extracted from ears, joints, and hearts of cn40- and N40-75-infected mice, using the QIAamp tissue kit (Qiagen Inc., Chatsworth, Calif.). Quantitation of B. burgdorferi was performed by PCR of twofold serial dilutions of the purified DNA. Primers corresponded to the flab gene. The number of spirochetes was calculated per milligram of DNA, using the following formula: reciprocal of the last dilution with a positive PCR 200/milligram of DNA used in the first dilution. The sensitivity of the assay was determined to be approximately 200 spirochetes in parallel experiments, and it did not depend on the source of the DNA (not shown). Immunoscreening of a genomic expression library. An N40 genomic expression library in lambda ZAP II (51) was differentially screened using immune sera from mice infected with cn40 or N40-75 for 21 days. Duplicate nitrocellulose filters were obtained for each plate, which contained approximately 10 4 plaques. Duplicate filters were screened with cn40- and N40-75-immune sera, and cn40- specific reactive plaques were rescued for further screening. Potential specific clones were subjected to three rounds of screening. Filters were blocked for 2 h using PBS with 3% bovine serum albumin (PBS-BSA) and incubated for 1 h with sera (1:100 dilution). Membrane filters were then washed three times using PBS with 0.05% Tween 20 and incubated for 30 min with goat anti-mouse immunoglobulin G in PBS-BSA (1:1,000 dilution), conjugated to alkaline phosphatase (Sigma Chemical Co., St. Louis, Mo.). After washing, the filters were incubated with tetramethylbenzidine substrate (Kirkegaard & Perry Laboratories, Inc., Gaithersburg, Md.). Positive clones were recovered in SM buffer with chloroform. After three rounds of screening, positive clones were further tested for possible reactivity with low titers of antibodies using N40-75-immune sera at a dilution of 1:50. Clones 2, 10, and 16 were then excised and sequenced using standard protocols. DNA homology comparisons were performed using BLAST search and the B. burgdorferi genome database (27). RNA purification and RT-PCR. RNA was extracted from the spleens of cn40- or N40-75-infected C3H mice by the guanidinium thiocyanate method (17), using a Micro RNA isolation kit (Stratagene, La Jolla, Calif.). To check for DNA contamination, RNA samples were subjected to PCR without further processing, using 100 ng of RNA and primers corresponding to flab, ospa, bba64 (27), dbpa, dbpb, bba65 (27), bba66 (27), p21, erpd, gene-1, and gene-2. Forty cycles were performed, with denaturation (94 C, 1 min), annealing (55 C, 1 min), and extension (72 C, 1 min) steps. If necessary, the samples were treated with DNase (Boehringer Mannheim, Indianapolis, Ind.) for 2 h at 37 C, extracted with phenol-chloroform-isoamyl alcohol (25:24:1), and ethanol precipitated. Aliquots of5to10 g were then used to obtain cdna with a reverse transcription Primer (reference) TABLE 1. Primers used Sequence flab...5 -AGA ATT ACT TCA AAG GCT-3 5 -TGT AGT TGT AAC ATT AAC-3 ospa...5 -GGT CAA ACC ACA CTT GAA GTT-3 5 -GTC AGT GTC ATT AAG TTC-3 erpt...5 -TTC ACA AGT AGC ACA ACA TGC TCC-3 5 -GCT ATC ACC ACC AAT TTC AAG TCC-3 bbk CAG CTT CTT CTA AAG AGG TAG AAG C-3 5 -TAA ATC GCA ACA AGC TTT AGC-3 ospd...5 -TAT CTT GTG TTC ATG ATA AAC-3 5 -ATC CAA TGT CTC TTT GAC TG-3 ospc...5 -GCC GTG AAA GAA GTT GAG ACC TTA-3 5 -TAA GAT TGT CCA GAC CAA GCA CTG-3 erps... E(13) (52)...5 -ATG TTT ATT ATT TGT GCT GTT T-3 E (310) (52)..5 -AAA CTA GTT TTA AAT GAT CC-3 dbpa...5 -GCT ACT GTA GTA GCT CGC A-3 5 -TTG AAT CGT CCT CTA AGG-3 dbpb...5 -CTG GAA ACA GTG GTC AAT TC-3 5 -TCT CTC CAC CAT TTT CAA T-3 bba TTC CTT CGT TGA AAT TTA AAG ATG-3 5 -TGA TTT CTT GCT CTC TGG AAG TC-3 bba AGC GGA TCT AAT TGA CAA GC-3 5 -TCC ATG TTG ATT GCT AAA TC-3 bba GAT TGT TTT TAT TTT CTT GCA CG-3 5 -TGA AAC GGC TTG AAT AGG TAT TC-3 p TTC TAA CTT CTA TTT TAA ATT-3 5 -AAT AAT GAG TTA AAA GTT AAG CAA-3 erpd...5 -GCG ATG ATC CTA ATG GCA-3 5 -CCA AGT TGT ACG TAT AGA GCT-3 gene AAC AAG CTC TAT TTA ACT AGT C-3 5 -GAT TCT TAC TGG TGA AAA G-3 gene GCA TTT AAG CAA TAT TCT TAG-3 5 -TCA CTT GAG CCC TTT GCT ATT-3 (RT)-PCR kit (Stratagene) utilizing random primers. PCR were then performed using the same primers and conditions and 2 to 5 l of the cdna samples. Primers. Sequences of the primers used are shown in Table 1. RESULTS B. burgdorferi N40 loses pathogenicity, but not infectivity, upon in vitro passage. A clonal isolate of B. burgdorferi N40 (cn40) that is highly infectious and pathogenic in C3H mice was continuously passaged in vitro to determine whether the spirochete would lose the ability to infect mice or cause disease. Despite being passaged 75 times in vitro, N40 retained its capacity to infect C3H mice, as demonstrated by recovery of the spirochete from selected organs at 2 to 3 weeks following inoculation with 10 4 organisms (Table 2). B. burgdorferi started to lose pathogenicity (the ability to cause arthritis and carditis) after 22 passages in vitro (Table 2). At passage 75, the spirochetes no longer elicited joint or cardiac inflammation in immunocompetent mice, regardless of their genetic (C3H or B6) background or dose (up to 10 7 ) of inoculum (Table 2). Arthritis and carditis were also not evident when N40-75-infected animals were examined for evidence of inflammation after 60 days (Table 2), indicating that the appearance of disease was not merely delayed. Characterization of N To understand why N40-75 was not pathogenic, we compared N40-75 and cn40 for antigenic and genetic differences. The protein profiles of cultured cn40 and N40-75 were similar except for a lower level of OspA/B and increased level of OspC expression by N40-75 (Fig. 1A).
3 1224 ANGUITA ET AL. INFECT. IMMUN. Mouse Isolate TABLE 2. Loss of B. burgdorferi pathogenicity with in vitro passage in immunocompetent mice a Day of sacrifice Infection (no. of specimens positive/no. examined) Blood Inoculation site Bladder Arthritis (no. of mice with arthritis/no. examined) Carditis (no. of mice with carditis/no. examined) C3H cn /5 4/5 5/5 5/5 5/5 p /5 2/5 5/5 3/5 3/5 p /5 2/3 4/5 0/5 1/5 N /5 4/5 3/5 0/5 0/5 N /5 4/5 4/4 0/5 0/5 N40-75 (10 7 ) 21 0/5 3/4 2/3 0/5 0/5 B6 cn /5 5/5 5/5 0/5 5/5 N /5 3/5 4/5 0/5 0/5 a C3H and B6 mice were infected with cn40 or its derivatives (p22, p51, and N40-75) obtained by in vitro culturing. Mice were infected with 10 4 spirochetes, unless otherwise indicated, and necropsied at 21 or 60 days after infection. Arthritis and carditis were assessed in a blinded fashion. Nevertheless, the level of expression of OspA/B varied among different clonal N40-75 spirochetes (Fig. 1A). Moreover, the analysis of N40 and N40-75 extracts by Western blotting using rabbit cn40 hyperimmune sera showed no differences between the two passages (Fig. 1B). Western blot analysis of cn40 and N40-75 with cn40- and N40-75-infected mouse sera revealed some differences in the proteins recognized by both sera (Fig. 1C). The differences were more notable in the range of 19 to 25 kda, as well as 30 to 32 kda (Fig. 1C, arrows), indicating that infection with both passages induced differentiated antibody responses. DNA PCR targeted at genes expressed in the linear chromosome, as well as several plasmids, indicated the presence of these genes both in cn40 and N40-75 (Fig. 2), as well as several N40-75 clones (Fig. 2). The analysis of the amplification products obtained with cn40 and N40-75 DNA when using primers E(13) and E(310) (52) indicated the presence of ospe and p21, but no other erps, in both passages (not shown), in agreement with the fact that only these two genes have been reported to be sequenced from this isolate (N40) (52). The presence of erpt by PCR (Fig. 2) and the vls locus by Southern blotting (not shown) in both spirochetes indicated that plasmid lp28-1 had not been lost during in vitro passage. These data indicate that no readily apparent loss of DNA could be associated with the loss of pathogenicity resulting from in vitro passage. To determine whether pathogenicity was related to spirochete growth, cn40 and N40-75 multiplication rates in vitro and bacterial burden in vivo were examined. When equal numbers of cn40 and N40-75 ( ) were incubated in BSK II medium, N40-75 grew more rapidly than cn40 (Fig. 3). Differences were evident as early as 48 h (doubling times of 5.2 and 3.7 h for cn40 and N40-75, respectively) and continued throughout the culture period (t test, P 0.05). Once stationary-phase growth had been reached, the density of N40-75 was greater than that of cn40 (Fig. 3). In contrast, N40-75 was less abundant than cn40 during infection of immunocompetent mice. Two weeks after inoculation into mice, the number of N40-75 in the skin (ear tissues) was significantly (P 0.05) less than the number of cn40 (average standard error [SE] per milligram of DNA, 27,000 15,000 for N40-75 and 360, ,000 for cn40 [Fig. 4]). The number of N40-75 spirochetes in the joints of the infected mice was extremely low (6,000 6,000) compared with cn40 (340,000 88,000) (Fig. 4). N40-75 could not be detected by PCR in the heart tissues, whereas large numbers of cn40 (1,100, ,000) were apparent (Fig. 4). N40-75 causes arthritis and carditis in SCID mice. Pathogenicity and bacterial burden in five cn40- and five N infected C3H-scid mice were then examined to assess whether spirochete growth was influenced by the B. burgdorferi-specific response. All of the N40-75-infected SCID mice developed arthritis and carditis that was indistinguishable from cn40- induced disease. Moreover, the severity of the inflammation was comparable in cn40- and N40-75-infected SCID mice (not shown). Quantitative PCR revealed similar number of spirochetes in ear (N40-75, 200,000 10,000; cn40, 400, ,000), joint (N40-75, 50,000 10,000; cn40, 60,000 20,000), or heart (N40-75, 180, ,000; cn40, 70,000 60,000) tissues in N and cn40-infected SCID mice (Fig. 5). N40-75 does not evade immune clearance as rapidly as cn40. The SCID mouse studies suggest that the inability of N40-75 to efficiently colonize murine tissues may be related to the host immune response. Previous reports showed that B. burgdorferi-specific antibodies protect against infection (6, 7). Passive transfer experiments also demonstrated that B. burgdorferi spirochetes become resistant to immune clearance within 4 to 5 days following syringe inoculation (7, 20). We therefore determined whether B. burgdorferi-specific humoral responses were responsible for the decreased number of spirochetes in N40-75-infected immunocompetent mice. Antisera from B. burgdorferi-infected C3H mice were administered to cn40- or N40-75-infected SCID mice. As expected (20), administration of immune sera at the time of challenge with cn40 resulted in protection of the mice (Table 3). The protective effect of immune sera was not apparent at 4 days after cn40 inoculation (20), suggesting that syringe-administered cn40 spirochetes adapt to their new environment by this interval (Table 3). In contrast, when N40-75-infected SCID mice were administered immune sera from B. burgdorferi-infected immunocompetent mice, protection against spirochetal challenge was evident when the sera were administered at 4 days after infection (Table 3). Sera from either cn40- or N40-75-infected immunocompetent mice were equally protective (Table 3), suggesting that the protective antibody specificities in the two sera were similar. At day 8 of infection, immune sera were no longer protective (Table 3). These data suggest that N40-75 requires a longer period of time than cn40 to adapt and evade antibodies present in sera from B. burgdorferi-infected immunocompetent mice. N40-75 is deficient in in vivo gene expression. After entering the mammalian host, several B. burgdorferi genes are differentially expressed (21). We examined whether the delayed ability of N40-75 to host adapt was associated with alterations in the up- or down-regulation of in vivo-expressed genes or genes coding for antigens recognized by antibodies in immune sera.
4 VOL. 68, 2000 B. BURGDORFERI GENE EXPRESSION AND PATHOGENICITY 1225 Downloaded from FIG. 1. cn40 and N40-75 do not differ substantially when grown in vitro. cn40, N40-75, and several N40-75 clones were grown in vitro to stationary phase and analyzed by sodium dodecyl sulfate-polyacrylamide gel electrophoresis to compare protein contents (A). Western blot analysis with cn40 rabbit hyperimmune sera was also performed to test the presence of OspA, -B, and -C in both passages (B). The immune response to cn40 and N40-75 was also tested by Western blotting (C), by using 21-day-infected mouse sera with cn40 and N on July 1, 2018 by guest To identify antigens selectively recognized by sera from cn40- infected mice, a genomic expression library was differentially screened with immune sera from cn40- and N40-75-infected mice (51). Immune sera from B. burgdorferi-infected mice contain antibodies to antigens synthesized during infection. If the antigenic profiles of N40-75 and cn40 are different during infection, then the antibodies raised during infection may differ. Using this strategy, we isolated three clones that were recognized by cn40- but not N40-75-infected murine sera (Table 4). Clone 2 had an insert of 2,570 bp, containing the genes bba64 (a p35 homologue [29] in the published B31 genome sequence [27]), bba65, and bba66 (yet another p35 homologue) (Table 4). Clone 10 had an insert of 2,032 bp and included dbpa (incomplete) and dbpb (bba24 and bba25, respectively), which form an operon (22, 31). It also contained three small open reading frames (bba26, bba27, and bba28) (27) and the incomplete sequence for bba29. Clone 16 had an insert of 3,360 bp and contained sequences related to the ospef gene family: p21 (51), erpd (13, 50), and two unknown genes (Table 4). Both clones 2 and 10 mapped within the linear plasmid of 54 kb, while clone 16 contained sequences not present in the published B31 sequence (27). The presence of p21 and ErpD suggests that this clone may belong to the cp32 family of plasmids. N40-75 does not express bba64, bba65, bba66, and dbpab. To directly determine whether these specific genes, which were
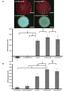
5 1226 ANGUITA ET AL. INFECT. IMMUN. FIG. 2. DNA contents of cn40 and N40-75, as well as of several N40-75 clones. DNA PCR for several genes was performed to detect any possible loss of DNA as a result of in vitro passage. The plasmids in which the amplified genes are present are indicated. identified by differential immunoscreening, were expressed by cn40 but not N40-75 during infection, we performed RT-PCR analysis of spleens from cn40- and N40-75-infected mice. dbpab, bba64, bba65, bba66, erpd, and gene-2 mrna could be readily detected in spleens of cn40-infected mice at 2 weeks of infection (Fig. 6). In contrast, N40-75-infected spleens lacked mrna for these genes. As expected, the flab gene (control) could be detected in cn40- or N40-75-infected animals, indicating that the spirochetes were present in these tissues. Neither cn40 or N40-75 expressed detectable levels of p21 mrna (Fig. 6), consistent with previous reports for this time period of FIG. 4. N40-75 does not effectively disseminate in the murine host. Ear, joint, and heart tissue was used to quantitate B. burgdorferi by DNA PCR using flab targets in mice infected for 3 weeks. Data represent the average SE of five individual mice., cn40;, N infection (18). Finally, gene-1 mrna was lacking in cn40 or N40-75 upon infection during 2 weeks (Fig. 6). DISCUSSION We characterized the pathogenicity of derivatives of a clonal population of B. burgdorferi cn40. Previous researchers have shown that B. burgdorferi passaged in vitro rapidly loses infectivity (40, 58). Our results differ in that cloned N40 spirochetes remained infectious in mice for up to 75 passages, suggesting that the infectivity of cn40 is relatively stable compared with other spirochetes passaged in vitro. Alternatively, infectivity could be maintained becaused short subculture intervals were used. It may be that prolonged growth under stationary-phase conditions leads to plasmid loss and more rapid reduction in infectivity. Indeed, both cn40 and N40-75 contained the plasmid expressing vls genes, supporting the contention that vls genes are important for infectivity and are not lost by cn40 spirochetes. Although N40-75 is infectious in C3H mice, these spirochetes do not cause arthritis or carditis in immunocompetent mice. The lack of disease in N40-75-infected mice correlated with a reduction, or lack, of spirochetal DNA in the joints and hearts. N40-75 could be readily cultured, or amplified by PCR, from the bladder, skin, and spleen, suggesting that the reduction in pathogenesis is due to an inability of N40-75 to effectively colonize specific tissues in sufficient numbers to cause disease. Lack of pathogenicity of N40-75 cannot be explained by assuming that the majority of cells in the uncloned N40-75 population are not infectious, since infection FIG. 3. N40-75 grows faster than cn40 in vitro. Equal amounts of cn40 and N40-75 were grown in BSK II medium and analyzed at different time points over 1 week. Data represent the average of five individual experiments SE. An asterisk indicates significant difference between cn40 and N40-75 at that particular time point (t test, P 0.05). FIG. 5. N40-75 disseminates efficiently in SCID mice. cn40 and N40-75 were quantitated in ears, joints, and hearts of C.B-17-scid mice after 2 weeks of infection as for Fig. 3., cn40;, N Data represent five individual mice.
6 VOL. 68, 2000 B. BURGDORFERI GENE EXPRESSION AND PATHOGENICITY 1227 Spirochete challenge TABLE 3. Effect of immune sera on N or cn40-infected SCID mice a Immune serum Day of serum administration Infection (no. of mice infected/no. examined) Blood Inoculation site Bladder cn40 NMS 0 1/5 5/5 5/5 cn40 immune 0 0/5 0/5 0/5 4 2/5 4/5 5/5 8 1/4 4/4 4/4 N40-75 immune 0 0/5 0/5 0/5 4 0/5 3/5 3/4 N40-75 NMS 0 2/5 3/5 5/5 cn40 immune 0 0/5 0/5 0/5 4 0/5 0/5 0/5 8 1/4 4/4 3/4 N40-75 immune 0 0/5 0/5 0/5 4 0/5 0/5 0/5 a C3H-scid mice were inoculated with 10 4 spirochetes and then treated with immune sera from B. burgdorferi-infected mice. Immune sera were administered within 1 h of spirochete inoculation (day 0) or 4 or 8 days after inoculation. Immune sera from immunocompetent C3H mice that had been infected with cn40 or N40-75 for 21 days were used. NMS, normal mouse serum (control). of mice with 10 4 spirochetes rendered culture positive organs that were not significantly different between cn40- and N40-75-infected animals (Table 2). Moreover, infection with 1,000 times more N40-75 spirochetes (10 7 ) did not change the nonpathogenic phenotype of high-passaged N40, arguing against this explanation. This reasoning is supported by the obtention and characterization at the DNA level by PCR of several clones of N40-75, which showed no loss of different pieces of DNA tested. These clones were infectious in C3H mice, but they did not cause arthritis or carditis. The analysis of the proteins profiles of cn40 and N40-75 indicated differences in OspA/B and OspC. OspA/B appeared to be down-regulated in in vitro-grown N40-75, although the analysis of both passages by Western blotting using cn40- hyperimmune sera revealed that both contained the proteins. Moreover, the analysis of N40-75 clones revealed variability in the amount of OspA/B expressed. In contrast, OspC was upregulated in high-passage N40. The down-regulation in vivo of ospab and the up-regulation of ospc gene expression after syringe inoculation have been demonstrated (18, 38). Therefore, these differences are not likely to be related with the lack of N40-75 pathogenicity. In order to survive and efficiently colonize its hosts, B. burgdorferi must adapt to different environments during its life cycle. It is well known that when infected I. scapularis engorge, the spirochete protein expression alters before migrating to the tick salivary glands and entering the vertebrate host (4, 21, 45, 47, 55). These changes include the up-regulation of ospc and the down-regulation of ospa (21, 46). The adaptation to invade the mammalian host is associated with changes in several TABLE 4. cn40-specific clones selected by immunoscreening of an N40 genomic library with cn40- and N40-75-immune sera a Clone no. Size of insert (bp) Gene(s) present Location 2 2,570 bba64, a bba65, bba66 lp ,032 dbpab a lp ,360 p21, erpd, gene-1, b gene-2 c cp32 (?) a cn40-specific clones reacted with antibodies in immune sera from cn40- infected mice but not N40-75-infected mice. b Incomplete sequence; 3 sequence missing. c GenBank accession number AF GenBank accession number: U genes, including bbk32, bbk50, ospef homologues, and dbpa/b. There is a time period (around 4 days after syringe inoculation) in which cn40 is no longer vulnerable to the passive administration of infected mouse sera (7, 20). The process of adaptation and immune evasion probably involves changes in gene expression or recombination events in genes like vls (60). Zhang and collaborators have demonstrated that recombination of the vls gene occurs in vivo, after experimental infection of mice and as soon as 4 days after the initiation of the infection (60). The evasion of the B. burgdorferi-specific humoral responses is probably due to a combination of selective gene expression and recombination, although the relative contribution of each remains to be elucidated. These data provide evidence that long-term-cultured B. burgdorferi spirochetes are more susceptible to host immunity. Susceptibility of N40-75-infected SCID mice to the development of arthritis and carditis suggests that efficient colonization and dissemination in the host involves immune evasion. Passive transfer of immune sera to N40-75-infected SCID mice at 8 days following challenge did not influence the establishment of infection, indicating the N40-75 can adapt within this interval. The delayed adaptation period contribute to the lower number of N40-75 in immunocompetent mice. Spirochete genes selectively expressed during infection are likely to be important for the maintenance of B. burgdorferi in the mammalian host and, ultimately, the completion of the vector-host-vector cycle. Initial characterization of antibody responses arising against cn40 and N40-75 in vivo (Fig. 1C) indicated that cn40-immune sera reacted with a wider range of proteins. Our immunoscreening procedure resulted in the detection of three clones that represented genes expressed exclusively by cn40 and not N40-75 upon infection. These expression units included genes that are preferentially expressed in the vertebrate host, including dbpab (14) and ospef homologues (21). Our results also showed that these genes were not merely lost by N40-75, since we could readily amplify these sequences when using N40-75 DNA. Moreover, DNA subtraction techniques indicated that a loss of DNA contents had not occurred during in vitro passage (data not shown). Interestingly, other researchers have reported a low level of expression of both DbpA and -B in high ( 30)-passage B. burgdorferi (30, 31), consistent with our findings for N Clone 2 contained three genes that could have expressed an-
7 1228 ANGUITA ET AL. INFECT. IMMUN. FIG. 6. N40-75 does not up-regulate in vivo genes. cn40- and N40-75-infected mouse spleens were used to extract total RNA, and RT-PCR for B. burgdorferi dbpa, dbpb, bba64, bba65, bba66, p21, erpd, gene-1, and gene-2 was performed. As a control, flab was used to identify spirochetes in spleens of the infected animals. N40-75 DNA was used as a positive control, demonstrating the presence of the genes in high-passage N40. tigens that were only recognized by the immune sera from cn40-infected mice. The three genes, bba64, bba65, and bba66, were expressed exclusively by cn40 in infected mice. Clone 16 contained four genes: p21 (18), erpd, and two unknown genes. Of them, p21 was not detected in cn40- or N40-75-infected mice, consistent with a later time point expression during infection (18), erpd and gene-2 were specifically expressed by cn40 at 2 weeks of infection. The exact functions of most of the in vivo genes, as well as the signals that trigger the initiation of their expression once B. burgdorferi enters the mammalian host, remain unknown. Among the factors that could trigger changes in gene expression or recombination events, temperature has been the most extensively studied. Stevenson and collaborators (48, 49) have examined the effect of temperature shifting on the expression of ospc (49) and the erps (48). The temporal expression of p21, anospe homologue, cannot be explained solely by temperature alterations, since this gene is not expressed until 21 days after initiation of the infection (18). Moreover, Zhang and collaborators have recently excluded temperature as the factor initiating recombination at the vls locus, speculating about the necessity of other host factors triggering this phenomenon (60). A detailed study of the changes in gene expression occurring after infection should provide insight into host factors that influence those changes. Our results indicate that B. burgdorferi has to adapt to the mammalian host in order to be able to reach a certain number and induce inflammation. Our data also suggest that the adaptation process implies at least two phenomena: the evasion of immune responses arising during infection and the up-regulation of genes that are needed for new functions that need to be performed in the new environment. To comply with the first task, there are two possibilities that could explain our results: N40-75 has impaired the ability to induce recombination (i.e., at the vls locus), and/or the down-regulation of highly immunogenic genes that do not need to be expressed in the new environment is impaired in N The adaptation phenomenon would then be a complex process in which both immune evasion and up-regulation of certain genes need to be accomplished. This is true for immunocompetent mice, although our results with SCID mice suggest that immune evasion in the absence of an acquired immune response is not required for pathogenicity. However, until a good system is provided to study in vivo function of the genes obtained in the differential immunoscreening (i.e., a transformation system that works in infectious and pathogenic B. burgdorferi), no direct evidence will be obtained about the exact roles of these genes during infection. In conclusion, we have generated a derivative of cn40, designated N40-75, which do not cause arthritis or carditis in immunocompetent mice. The inability of N40-75 to adapt to the mammalian host correlates with its inability to induce disease. This defect in differential gene expression results in an increase and prolonged susceptibility of N40-75 to the antibodies in immune sera and an inability of these N40-75 spirochetes to effectively colonize and disseminate within the murine host due to that immune-based vulnerability. ACKNOWLEDGMENTS This work was supported by grants AI and AR from the National Institutes of Health, by the Arthritis Foundation, and by the American Heart Association. E.F. is a recipient of a Burroughs Wellcome Clinical Scientist Award in Translational Research. We thank Debbie Beck for technical assistance. REFERENCES 1. Akins, D. R., S. F. Porcella, T. G. Popova, D. Shevchenko, S. I. Baker, M. Li, M. V. Norgard, and J. D. Radolf Evidence for in vivo but not in vitro expression of a Borrelia burgdorferi outer surface protein F (OspF) homologue. Mol. Microbiol. 18: Anguita, J., D. H. Persing, M. Rincon, S. W. Barthold, and E. Fikrig Effect of anti-interleukin 12 treatment on murine lyme borreliosis. J. Clin. Investig. 97: Armstrong, A. L., S. W. Barthold, D. H. Persing, and D. S. Beck Carditis in Lyme disease susceptible and resistant strains of laboratory mice infected with Borrelia burgdorferi. Am. J. Trop. Med. Hyg. 47: Barbour, A. G Plasmid analysis of Borrelia burgdorferi, the Lyme
8 VOL. 68, 2000 B. BURGDORFERI GENE EXPRESSION AND PATHOGENICITY 1229 disease agent. J. Clin. Microbiol. 26: Barthold, S. W., D. S. Beck, G. M. Hansen, G. A. Terwilliger, and K. D. Moody Lyme borreliosis in selected strains and ages of laboratory mice. J. Infect. Dis. 162: Barthold, S. W., and L. K. Bockenstedt Passive immunizing activity of sera from mice infected with Borrelia burgdorferi. Infect. Immun. 61: Barthold, S. W., M. de Souza, and S. Feng Serum-mediated resolution of Lyme arthritis in mice. Lab. Investig. 74: Barthold, S. W., M. S. de Souza, J. L. Janotka, A. L. Smith, and D. H. Persing Chronic Lyme borreliosis in the laboratory mouse. Am. J. Pathol. 143: Barthold, S. W., E. Fikrig, L. K. Bockenstedt, and D. H. Persing Circumvention of outer surface protein A immunity by host-adapted Borrelia burgdorferi. Infect. Immun. 63: Barthold, S. W., D. H. Persing, A. L. Armstrong, and R. A. Peeples Kinetics of Borrelia burgdorferi dissemination and evolution of disease after intradermal inoculation of mice. Am. J. Pathol. 139: Brunet, L. R., C. Sellitto, A. Spielman, and S. R. I. Telford Antibody response of the mouse reservoir of Borrelia burgdorferi in nature. Infect. Immun. 63: Carroll, J. A., and F. C. Gherardini Membrane protein variations associated with in vitro passage of Borrelia burgdorferi. Infect. Immun. 64: Casjens, S., R. van Vugt, K. Tilly, P. A. Rosa, and B. Stevenson Homology throughout the multiple 32-kilobase circular plasmids present in Lyme disease spirochetes. J. Bacteriol. 179: Cassat, D. R., N. K. Patel, N. D. Ulbrandt, and M. S. Hanson DbpA, but not OspA, is expressed by Borrelia burgdorferi during spirochetemia and is a target for protective antibodies. Infect. Immun. 66: Centers for Disease Control and Prevention Lyme disease United States, Morbid. Mortal. Weekly Rep. 46: Champion, C. I., D. R. Blanco, J. T. Skare, D. A. Haake, M. Giladi, D. Foley, J. N. Miller, and M. A. Lovett A 9.0-kilobase-pair circular plasmid of Borrelia burgdorferi encodes an exported protein: evidence for expression only during infection. Infect. Immun. 62: Chomczynski, P., and N. Sacchi Single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction. Anal. Biochem. 162: Das, S., S. W. Barthold, S. S. Giles, R. R. Montgomery, S. R. I. Telford, and E. Fikrig Temporal pattern of Borrelia burgdorferi p21 expression in ticks and the mammalian host. J. Clin. Investig. 99: Defosse, D. L., P. H. Duray, and R. C. Johnson The NIH-3 immunodeficient mouse is a model for Lyme borreliosis myositis and carditis. Am. J. Pathol. 141: de Silva, A., E. Fikrig, E. Hodzic, F. S. Kantor, S. R. I. Telford, and S. W. Barthold Immune evasion by tickborne and host-adapted Borrelia burgdorferi. J. Infect. Dis. 177: de Silva, A. M., and E. Fikrig Arthropod- and host-specific gene expression by Borrelia burgdorferi. J. Clin. Investig. 99: Feng, S., E. Hodzic, B. Stevenson, and S. W. Barthold Humoral immunity to Borrelia burgdorferi N40 decorin binding proteins during infection of laboratory mice. Infect. Immun. 66: Fikrig, E., S. W. Barthold, and R. A. Flavell OspA vaccination of mice with established Borrelia burgdorferi infection alters disease but not infection. Infect. Immun. 61: Fikrig, E., S. W. Barthold, F. S. Kantor, and R. A. Flavell Long-term protection of mice from Lyme disease by vaccination with OspA. Infect. Immun. 60: Fikrig, E., S. W. Barthold, W. Sun, W. Feng, S. R. I. Telford, and R. A. Flavell Borrelia burgdorferi P35 and P37 proteins, expressed in vivo, elicit protective immunity. Immunity 6: Fikrig, E., H. Tao, F. S. Kantor, S. W. Barthold, and R. A. Flavell Evasion of protective immunity by Borrelia burgdorferi by truncation of outer surface protein B. Proc. Natl. Acad. Sci. USA 90: Fraser, C. M., S. Casjens, W. M. Huang, G. G. Sutton, R. Clayton, R. Lathigra, O. White, K. A. Ketchum, R. Dodson, E. K. Hickey, M. Gwinn, B. Dougherty, J. F. Tomb, R. D. Fleischmann, D. Richardson, J. Peterson, A. R. Kerlavage, J. Quackenbush, S. Salzberg, M. Hanson, R. van Vugt, N. Palmer, M. D. Adams, J. Gocayne, J. C. Venter, et al Genomic sequence of a Lyme disease spirochaete, Borrelia burgdorferi. Nature 390: Fuchs, H., R. Wallich, M. M. Simon, and M. D. Kramer The outer surface protein A of the spirochete Borrelia burgdorferi is a plasmin(ogen) receptor. Proc. Natl. Acad. Sci. USA 91: Gilmore, R. D., Jr., K. J. Kappel, and B. J. Johnson Molecular characterization of a 35-kilodalton protein of Borrelia burgdorferi, an antigen of diagnostic importance in early Lyme disease. J. Clin. Microbiol. 35: Guo, B. P., S. J. Norris, L. C. Rosenberg, and M. Hook Adherence of Borrelia burgdorferi to the proteoglycan decorin. Infect. Immun. 63: Hagman, K. E., P. Lahdenne, T. G. Popova, S. F. Porcella, D. R. Akins, J. D. Radolf, and M. V. Norgard Decorin-binding protein of Borrelia burgdorferi is encoded within a two-gene operon and is protective in the murine model of Lyme borreliosis. Infect. Immun. 66: Hughes, C. A., S. M. Engstrom, L. A. Coleman, C. B. Kodner, and R. C. Johnson Protective immunity is induced by a Borrelia burgdorferi mutant that lacks OspA and OspB. Infect. Immun. 61: Keane-Myers, A., and S. P. Nickell Role of IL-4 and IFN-gamma in modulation of immunity to Borrelia burgdorferi in mice. J. Immunol. 155: Kenefick, K. B., L. C. Lim, J. D. Alder, J. L. Schmitz, C. J. Czuprynski, and R. F. Schell Induction of interleukin-1 release by high- and lowpassage isolates of Borrelia burgdorferi. J. Infect. Dis. 167: Lengl-Janssen, B., A. F. Strauss, A. C. Steere, and T. Kamradt The T helper cell response in Lyme arthritis: differential recognition of Borrelia burgdorferi outer surface protein A in patients with treatment-resistant or treatment-responsive Lyme arthritis. J. Exp. Med. 180: Ma, Y., K. P. Seiler, E. J. Eichwald, J. H. Weis, C. Teuscher, and J. J. Weis Distinct characteristics of resistance to Borrelia burgdorferi-induced arthritis in C57BL/6N mice. Infect. Immun. 66: Matyniak, J. E., and S. L. Reiner T helper phenotype and genetic susceptibility in experimental Lyme disease. J. Exp. Med. 181: Montgomery, R. R., S. E. Malawista, K. J. Feen, and L. K. Bockenstedt Direct demonstration of antigenic substitution of Borrelia burgdorferi ex vivo: exploration of the paradox of the early immune response to outer surface proteins A and C in Lyme disease. J. Exp. Med. 183: Moody, K. D., S. W. Barthold, and G. A. Terwilliger Lyme borreliosis in laboratory animals: effect of host species and in vitro passage of Borrelia burgdorferi. Am. J. Trop. Med. Hyg. 43: Norris, S. J., J. K. Howell, S. A. Garza, M. S. Ferdows, and A. G. Barbour High- and low-infectivity phenotypes of clonal populations of in vitrocultured Borrelia burgdorferi. Infect. Immun. 63: Probert, W. S., and B. J. B. Johnson Identification of a 47 kda fibronectin-binding protein expressed by Borrelia burgdorferi B31. Mol. Microbiol. 30: Ramamoorthy, R., and M. T. Phillip Differential expression of Borrelia burgdorferi proteins during growth in vitro. Infect. Immun. 66: Schaible, U. E., S. Gay, C. Museteanu, M. D. Kramer, G. Zimmer, K. Eichmann, U. Museteanu, and M. M. Simon Lyme borreliosis in the severe combined immunodeficiency (scid) mouse manifests predominantly in the joints, heart, and liver. Am. J. Pathol. 137: Schaible, U. E., M. D. Kramer, R. Wallich, T. Tran, and M. M. Simon Experimental Borrelia burgdorferi infection in inbred mouse strains: antibody response and association of H-2 genes with resistance and susceptibility to development of arthritis. Eur. J. Immunol. 21: Schwan, T. G., W. Burgdorfer, and C. F. Garon Changes in infectivity and plasmid profile of the Lyme disease spirochete, Borrelia burgdorferi, as a result of in vitro cultivation. Infect. Immun. 56: Schwan, T. G., J. Piesman, W. T. Golde, M. C. Dolan, and P. A. Rosa Induction of an outer surface protein on Borrelia burgdorferi during tick feeding. Proc. Natl. Acad. Sci. USA 92: Simpson, W. J., C. F. Garon, and T. G. Schwan Analysis of supercoiled circular plasmids in infectious and non-infectious Borrelia burgdorferi. Microb. Pathog. 8: Stevenson, B., J. L. Bono, T. G. Schwan, and P. Rosa Borrelia burgdorferi Erp proteins are immunogenic in mammals infected by tick bite, and their synthesis is inducible in cultured bacteria. Infect. Immun. 66: Stevenson, B., T. G. Schwan, and P. A. Rosa Temperature-related differential expression of antigens in the Lyme disease spirochete, Borrelia burgdorferi. Infect. Immun. 63: Stevenson, B., K. Tilly, and P. A. Rosa A family of genes located on four separate 32-kilobase circular plasmids in Borrelia burgdorferi B31. J. Bacteriol. 178: Suk, K., S. Das, W. Sun, B. Jwang, S. W. Barthold, R. A. Flavell, and E. Fikrig Borrelia burgdorferi genes selectively expressed in the infected host. Proc. Natl. Acad. Sci. USA 92: Sung, S. Y., C. P. Lavoie, J. A. Carlyon, and R. T. Marconi Genetic divergence and evolutionary instability in ospe-related members of the upstream homology box gene family in Borrelia burgdorferi sensu lato complex isolates. Infect. Immun. 66: Szczepanski, A., and J. L. Benach Lyme borreliosis: host responses to Borrelia burgdorferi. Microbiol. Rev. 55: Wallich, R., C. Brenner, M. D. Kramer, and M. M. Simon Molecular cloning and immunological characterization of a novel linear-plasmid-encoded gene, pg, of Borrelia burgdorferi expressed only in vivo. Infect. Immun. 63: Xu, Y., C. Kodner, L. Coleman, and R. C. Johnson Correlation of plasmids with infectivity of Borrelia burgdorferi sensu stricto type strain B31. Infect. Immun. 64: Yang, L., Y. Ma, R. Schoenfeld, M. Griffiths, E. Eichwald, B. Araneo, and J. J. Weis Evidence for B-lymphocyte mitogen activity in Borrelia burgdorferi-infected mice. Infect. Immun. 60:
9 1230 ANGUITA ET AL. INFECT. IMMUN. 57. Yang, L., J. H. Weis, E. Eichwald, C. P. Kolbert, D. H. Persing, and J. J. Weis Heritable susceptibility to severe Borrelia burgdorferi-induced arthritis is dominant and is associated with persistence of large numbers of spirochetes in tissues. Infect. Immun. 62: Zhang, J. R., J. M. Hardham, A. G. Barbour, and S. J. Norris Antigenic variation in Lyme disease borreliae by promiscuous recombination of VMP-like sequence cassettes. Cell 89: Zhang, J. R., and S. J. Norris Genetic variation of the Borrelia burgdorferi gene vlse involves cassette-specific, segmental gene conversion. Infect. Immun. 66: Zhang, J. R., and S. J. Norris Kinetics and in vivo induction of genetic variation of vlse in Borrelia burgdorferi. Infect. Immun. 66: Editor: D. L. Burns
Supplementary. Table 1: Oligonucleotides and Plasmids. complementary to positions from 77 of the SRα '- GCT CTA GAG AAC TTG AAG TAC AGA CTG C
 Supplementary Table 1: Oligonucleotides and Plasmids 913954 5'- GCT CTA GAG AAC TTG AAG TAC AGA CTG C 913955 5'- CCC AAG CTT ACA GTG TGG CCA TTC TGC TG 223396 5'- CGA CGC GTA CAG TGT GGC CAT TCT GCT G
Supplementary Table 1: Oligonucleotides and Plasmids 913954 5'- GCT CTA GAG AAC TTG AAG TAC AGA CTG C 913955 5'- CCC AAG CTT ACA GTG TGG CCA TTC TGC TG 223396 5'- CGA CGC GTA CAG TGT GGC CAT TCT GCT G
Influence of Cultivation Media on Genetic Regulatory Patterns in Borrelia burgdorferi
 INFECTION AND IMMUNITY, June 2001, p. 4159 4163 Vol. 69, No. 6 0019-9567/01/$04.00 0 DOI: 10.1128/IAI.69.6.4159 4163.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved. Influence
INFECTION AND IMMUNITY, June 2001, p. 4159 4163 Vol. 69, No. 6 0019-9567/01/$04.00 0 DOI: 10.1128/IAI.69.6.4159 4163.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved. Influence
Supplemental Data Supplemental Figure 1.
 Supplemental Data Supplemental Figure 1. Silique arrangement in the wild-type, jhs, and complemented lines. Wild-type (WT) (A), the jhs1 mutant (B,C), and the jhs1 mutant complemented with JHS1 (Com) (D)
Supplemental Data Supplemental Figure 1. Silique arrangement in the wild-type, jhs, and complemented lines. Wild-type (WT) (A), the jhs1 mutant (B,C), and the jhs1 mutant complemented with JHS1 (Com) (D)
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
PCR analysis was performed to show the presence and the integrity of the var1csa and var-
 Supplementary information: Methods: Table S1: Primer Name Nucleotide sequence (5-3 ) DBL3-F tcc ccg cgg agt gaa aca tca tgt gac tg DBL3-R gac tag ttt ctt tca ata aat cac tcg c DBL5-F cgc cct agg tgc ttc
Supplementary information: Methods: Table S1: Primer Name Nucleotide sequence (5-3 ) DBL3-F tcc ccg cgg agt gaa aca tca tgt gac tg DBL3-R gac tag ttt ctt tca ata aat cac tcg c DBL5-F cgc cct agg tgc ttc
Hes6. PPARα. PPARγ HNF4 CD36
 SUPPLEMENTARY INFORMATION Supplementary Table Positions and Sequences of ChIP primers -63 AGGTCACTGCCA -79 AGGTCTGCTGTG Hes6-0067 GGGCAaAGTTCA ACOT -395 GGGGCAgAGTTCA PPARα -309 GGCTCAaAGTTCAaGTTCA CPTa
SUPPLEMENTARY INFORMATION Supplementary Table Positions and Sequences of ChIP primers -63 AGGTCACTGCCA -79 AGGTCTGCTGTG Hes6-0067 GGGCAaAGTTCA ACOT -395 GGGGCAgAGTTCA PPARα -309 GGCTCAaAGTTCAaGTTCA CPTa
Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR
 Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR 1 The problem We wish to clone a yet unknown gene from a known
Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR 1 The problem We wish to clone a yet unknown gene from a known
Supplementary Figure 1A A404 Cells +/- Retinoic Acid
 Supplementary Figure 1A A44 Cells +/- Retinoic Acid 1 1 H3 Lys4 di-methylation SM-actin VEC cfos (-) RA (+) RA 14 1 1 8 6 4 H3 Lys79 di-methylation SM-actin VEC cfos (-) RA (+) RA Supplementary Figure
Supplementary Figure 1A A44 Cells +/- Retinoic Acid 1 1 H3 Lys4 di-methylation SM-actin VEC cfos (-) RA (+) RA 14 1 1 8 6 4 H3 Lys79 di-methylation SM-actin VEC cfos (-) RA (+) RA Supplementary Figure
Supplemental material
 Supplemental material Diversity of O-antigen repeat-unit structures can account for the substantial sequence variation of Wzx translocases Yaoqin Hong and Peter R. Reeves School of Molecular Bioscience,
Supplemental material Diversity of O-antigen repeat-unit structures can account for the substantial sequence variation of Wzx translocases Yaoqin Hong and Peter R. Reeves School of Molecular Bioscience,
Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH).
 Bisulfite Treatment of DNA Dilute DNA sample to 2µg DNA in 50µl ddh 2 O. Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH). Incubate in a 37ºC water bath for 30 minutes. To 55µl samples
Bisulfite Treatment of DNA Dilute DNA sample to 2µg DNA in 50µl ddh 2 O. Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH). Incubate in a 37ºC water bath for 30 minutes. To 55µl samples
Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis
 1 2 3 4 5 6 7 8 9 10 11 12 Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis Information Research). Exons
1 2 3 4 5 6 7 8 9 10 11 12 Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis Information Research). Exons
Arabidopsis actin depolymerizing factor AtADF4 mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB
 Arabidopsis actin depolymerizing factor mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB Files in this Data Supplement: Supplemental Table S1 Supplemental Table
Arabidopsis actin depolymerizing factor mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB Files in this Data Supplement: Supplemental Table S1 Supplemental Table
Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system
 Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system Dr. Tim Welsink Molecular Biology Transient Gene Expression OUTLINE Short
Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system Dr. Tim Welsink Molecular Biology Transient Gene Expression OUTLINE Short
Project 07/111 Final Report October 31, Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines
 Project 07/111 Final Report October 31, 2007. Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines Project Leader: Dr Douglas C. Hodgins (519-824-4120 Ex 54758, fax 519-824-5930)
Project 07/111 Final Report October 31, 2007. Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines Project Leader: Dr Douglas C. Hodgins (519-824-4120 Ex 54758, fax 519-824-5930)
PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells
 Supplementary Information for: PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells Ju Hye Jang 1, Hyun Kim 2, Mi Jung Jang 2, Ju Hyun Cho 1,2,* 1 Research Institute
Supplementary Information for: PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells Ju Hye Jang 1, Hyun Kim 2, Mi Jung Jang 2, Ju Hyun Cho 1,2,* 1 Research Institute
Supplemental Table 1. Mutant ADAMTS3 alleles detected in HEK293T clone 4C2. WT CCTGTCACTTTGGTTGATAGC MVLLSLWLIAAALVEVR
 Supplemental Dataset Supplemental Table 1. Mutant ADAMTS3 alleles detected in HEK293T clone 4C2. DNA sequence Amino acid sequence WT CCTGTCACTTTGGTTGATAGC MVLLSLWLIAAALVEVR Allele 1 CCTGTC------------------GATAGC
Supplemental Dataset Supplemental Table 1. Mutant ADAMTS3 alleles detected in HEK293T clone 4C2. DNA sequence Amino acid sequence WT CCTGTCACTTTGGTTGATAGC MVLLSLWLIAAALVEVR Allele 1 CCTGTC------------------GATAGC
Disease and selection in the human genome 3
 Disease and selection in the human genome 3 Ka/Ks revisited Please sit in row K or forward RBFD: human populations, adaptation and immunity Neandertal Museum, Mettman Germany Sequence genome Measure expression
Disease and selection in the human genome 3 Ka/Ks revisited Please sit in row K or forward RBFD: human populations, adaptation and immunity Neandertal Museum, Mettman Germany Sequence genome Measure expression
ΔPDD1 x ΔPDD1. ΔPDD1 x wild type. 70 kd Pdd1. Pdd3
 Supplemental Fig. S1 ΔPDD1 x wild type ΔPDD1 x ΔPDD1 70 kd Pdd1 50 kd 37 kd Pdd3 Supplemental Fig. S1. ΔPDD1 strains express no detectable Pdd1 protein. Western blot analysis of whole-protein extracts
Supplemental Fig. S1 ΔPDD1 x wild type ΔPDD1 x ΔPDD1 70 kd Pdd1 50 kd 37 kd Pdd3 Supplemental Fig. S1. ΔPDD1 strains express no detectable Pdd1 protein. Western blot analysis of whole-protein extracts
Supporting information for Biochemistry, 1995, 34(34), , DOI: /bi00034a013
 Supporting information for Biochemistry, 1995, 34(34), 10807 10815, DOI: 10.1021/bi00034a013 LESNIK 10807-1081 Terms & Conditions Electronic Supporting Information files are available without a subscription
Supporting information for Biochemistry, 1995, 34(34), 10807 10815, DOI: 10.1021/bi00034a013 LESNIK 10807-1081 Terms & Conditions Electronic Supporting Information files are available without a subscription
Supplemental Data. mir156-regulated SPL Transcription. Factors Define an Endogenous Flowering. Pathway in Arabidopsis thaliana
 Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
ORFs and genes. Please sit in row K or forward
 ORFs and genes Please sit in row K or forward https://www.flickr.com/photos/teseum/3231682806/in/photostream/ Question: why do some strains of Vibrio cause cholera and others don t? Methods Mechanisms
ORFs and genes Please sit in row K or forward https://www.flickr.com/photos/teseum/3231682806/in/photostream/ Question: why do some strains of Vibrio cause cholera and others don t? Methods Mechanisms
strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular
![strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular](/thumbs/72/66361212.jpg) Additional file 2 Identification of AOX1 in P. pastoris GS115 with a Mut s phenotype Results and Discussion The HBsAg producing strain was originally identified as a Mut s (methanol utilization slow) strain
Additional file 2 Identification of AOX1 in P. pastoris GS115 with a Mut s phenotype Results and Discussion The HBsAg producing strain was originally identified as a Mut s (methanol utilization slow) strain
Dierks Supplementary Fig. S1
 Dierks Supplementary Fig. S1 ITK SYK PH TH K42R wt K42R (kinase deficient) R29C E42K Y323F R29C E42K Y323F (reduced phospholipid binding) (enhanced phospholipid binding) (reduced Cbl binding) E42K Y323F
Dierks Supplementary Fig. S1 ITK SYK PH TH K42R wt K42R (kinase deficient) R29C E42K Y323F R29C E42K Y323F (reduced phospholipid binding) (enhanced phospholipid binding) (reduced Cbl binding) E42K Y323F
Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC
 Supplementary Appendixes Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC ACG TAG CTC CGG CTG GA-3 for vimentin, /5AmMC6/TCC CTC GCG CGT GGC TTC CGC
Supplementary Appendixes Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC ACG TAG CTC CGG CTG GA-3 for vimentin, /5AmMC6/TCC CTC GCG CGT GGC TTC CGC
Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were
 1 Supplemental methods 2 3 4 5 6 7 8 9 1 11 12 13 14 15 16 17 18 19 21 22 23 Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were monitored by quantitative reverse-transcription
1 Supplemental methods 2 3 4 5 6 7 8 9 1 11 12 13 14 15 16 17 18 19 21 22 23 Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were monitored by quantitative reverse-transcription
Cat. # Product Size DS130 DynaExpress TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1
 Product Name: Kit Component TA PCR Cloning Kit (ptakn-2) Cat. # Product Size DS130 TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1 2 Ligation Buffer
Product Name: Kit Component TA PCR Cloning Kit (ptakn-2) Cat. # Product Size DS130 TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1 2 Ligation Buffer
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/10/494/eaan6284/dc1 Supplementary Materials for Activation of master virulence regulator PhoP in acidic ph requires the Salmonella-specific protein UgtL Jeongjoon
www.sciencesignaling.org/cgi/content/full/10/494/eaan6284/dc1 Supplementary Materials for Activation of master virulence regulator PhoP in acidic ph requires the Salmonella-specific protein UgtL Jeongjoon
Legends for supplementary figures 1-3
 High throughput resistance profiling of Plasmodium falciparum infections based on custom dual indexing and Illumina next generation sequencing-technology Sidsel Nag 1,2 *, Marlene D. Dalgaard 3, Poul-Erik
High throughput resistance profiling of Plasmodium falciparum infections based on custom dual indexing and Illumina next generation sequencing-technology Sidsel Nag 1,2 *, Marlene D. Dalgaard 3, Poul-Erik
Y-chromosomal haplogroup typing Using SBE reaction
 Schematic of multiplex PCR followed by SBE reaction Multiplex PCR Exo SAP purification SBE reaction 5 A 3 ddatp ddgtp 3 T 5 A G 3 T 5 3 5 G C 5 3 3 C 5 ddttp ddctp 5 T 3 T C 3 A 5 3 A 5 5 C 3 3 G 5 3 G
Schematic of multiplex PCR followed by SBE reaction Multiplex PCR Exo SAP purification SBE reaction 5 A 3 ddatp ddgtp 3 T 5 A G 3 T 5 3 5 G C 5 3 3 C 5 ddttp ddctp 5 T 3 T C 3 A 5 3 A 5 5 C 3 3 G 5 3 G
Table S1. Bacterial strains (Related to Results and Experimental Procedures)
 Table S1. Bacterial strains (Related to Results and Experimental Procedures) Strain number Relevant genotype Source or reference 1045 AB1157 Graham Walker (Donnelly and Walker, 1989) 2458 3084 (MG1655)
Table S1. Bacterial strains (Related to Results and Experimental Procedures) Strain number Relevant genotype Source or reference 1045 AB1157 Graham Walker (Donnelly and Walker, 1989) 2458 3084 (MG1655)
Materials Protein synthesis kit. This kit consists of 24 amino acids, 24 transfer RNAs, four messenger RNAs and one ribosome (see below).
 Protein Synthesis Instructions The purpose of today s lab is to: Understand how a cell manufactures proteins from amino acids, using information stored in the genetic code. Assemble models of four very
Protein Synthesis Instructions The purpose of today s lab is to: Understand how a cell manufactures proteins from amino acids, using information stored in the genetic code. Assemble models of four very
Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of
 Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of the cell line) were immunostained for HA, acetylated
Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of the cell line) were immunostained for HA, acetylated
Multiplexing Genome-scale Engineering
 Multiplexing Genome-scale Engineering Harris Wang, Ph.D. Department of Systems Biology Department of Pathology & Cell Biology http://wanglab.c2b2.columbia.edu Rise of Genomics An Expanding Toolbox Esvelt
Multiplexing Genome-scale Engineering Harris Wang, Ph.D. Department of Systems Biology Department of Pathology & Cell Biology http://wanglab.c2b2.columbia.edu Rise of Genomics An Expanding Toolbox Esvelt
Supporting Information. Copyright Wiley-VCH Verlag GmbH & Co. KGaA, Weinheim, 2006
 Supporting Information Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Supporting Information for Expanding the Genetic
Supporting Information Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Supporting Information for Expanding the Genetic
Supplementary Figures
 Supplementary Figures Supplementary Fig. 1 Characterization of GSCs. a. Immunostaining of primary GSC spheres from GSC lines. Nestin (neural progenitor marker, red), TLX (green). Merged images of nestin,
Supplementary Figures Supplementary Fig. 1 Characterization of GSCs. a. Immunostaining of primary GSC spheres from GSC lines. Nestin (neural progenitor marker, red), TLX (green). Merged images of nestin,
Lecture 11: Gene Prediction
 Lecture 11: Gene Prediction Study Chapter 6.11-6.14 1 Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and end positions of genes in a genome Where are
Lecture 11: Gene Prediction Study Chapter 6.11-6.14 1 Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and end positions of genes in a genome Where are
RPA-AB RPA-C Supplemental Figure S1: SDS-PAGE stained with Coomassie Blue after protein purification.
 RPA-AB RPA-C (a) (b) (c) (d) (e) (f) Supplemental Figure S: SDS-PAGE stained with Coomassie Blue after protein purification. (a) RPA; (b) RPA-AB; (c) RPA-CDE; (d) RPA-CDE core; (e) RPA-DE; and (f) RPA-C
RPA-AB RPA-C (a) (b) (c) (d) (e) (f) Supplemental Figure S: SDS-PAGE stained with Coomassie Blue after protein purification. (a) RPA; (b) RPA-AB; (c) RPA-CDE; (d) RPA-CDE core; (e) RPA-DE; and (f) RPA-C
Supplemental Table 1. Primers used for PCR.
 Supplemental Table 1. Primers used for PCR. Gene Type Primer Sequence Genotyping and semi-quantitative RT-PCR F 5 -TTG CCC GAT CAC CAT CTG TA-3 rwa1-1 R 5 -TGT AGC GAT CAA GGC CTG ATC TAA-3 LB 5 -TAG CAT
Supplemental Table 1. Primers used for PCR. Gene Type Primer Sequence Genotyping and semi-quantitative RT-PCR F 5 -TTG CCC GAT CAC CAT CTG TA-3 rwa1-1 R 5 -TGT AGC GAT CAA GGC CTG ATC TAA-3 LB 5 -TAG CAT
Supplementary Information
 Supplementary Information A general solution for opening double-stranded DNA for isothermal amplification Gangyi Chen, Juan Dong, Yi Yuan, Na Li, Xin Huang, Xin Cui* and Zhuo Tang* Supplementary Materials
Supplementary Information A general solution for opening double-stranded DNA for isothermal amplification Gangyi Chen, Juan Dong, Yi Yuan, Na Li, Xin Huang, Xin Cui* and Zhuo Tang* Supplementary Materials
Specificity of Infection-Induced Immunity among Borrelia burgdorferi Sensu Lato Species
 INFECTION AND IMMUNITY, Jan. 1999, p. 36 42 Vol. 67, No. 1 0019-9567/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. Specificity of Infection-Induced Immunity among
INFECTION AND IMMUNITY, Jan. 1999, p. 36 42 Vol. 67, No. 1 0019-9567/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. Specificity of Infection-Induced Immunity among
hcd1tg/hj1tg/ ApoE-/- hcd1tg/hj1tg/ ApoE+/+
 ApoE+/+ ApoE-/- ApoE-/- H&E (1x) Supplementary Figure 1. No obvious pathology is observed in the colon of diseased ApoE-/me. Colon samples were fixed in 1% formalin and laid out in Swiss rolls for paraffin
ApoE+/+ ApoE-/- ApoE-/- H&E (1x) Supplementary Figure 1. No obvious pathology is observed in the colon of diseased ApoE-/me. Colon samples were fixed in 1% formalin and laid out in Swiss rolls for paraffin
Overexpression Normal expression Overexpression Normal expression. 26 (21.1%) N (%) P-value a N (%)
 SUPPLEMENTARY TABLES Table S1. Alteration of ZNF322A protein expression levels in relation to clinicopathological parameters in 123 Asian and 74 Caucasian lung cancer patients. Asian patients Caucasian
SUPPLEMENTARY TABLES Table S1. Alteration of ZNF322A protein expression levels in relation to clinicopathological parameters in 123 Asian and 74 Caucasian lung cancer patients. Asian patients Caucasian
Supporting Information
 Supporting Information Barderas et al. 10.1073/pnas.0801221105 SI Text: Docking of gastrin to Constructed scfv Models Interactive predocking of the 4-WL-5 motif into the central pocket observed in the
Supporting Information Barderas et al. 10.1073/pnas.0801221105 SI Text: Docking of gastrin to Constructed scfv Models Interactive predocking of the 4-WL-5 motif into the central pocket observed in the
Supplemental Information. Human Senataxin Resolves RNA/DNA Hybrids. Formed at Transcriptional Pause Sites. to Promote Xrn2-Dependent Termination
 Supplemental Information Molecular Cell, Volume 42 Human Senataxin Resolves RNA/DNA Hybrids Formed at Transcriptional Pause Sites to Promote Xrn2-Dependent Termination Konstantina Skourti-Stathaki, Nicholas
Supplemental Information Molecular Cell, Volume 42 Human Senataxin Resolves RNA/DNA Hybrids Formed at Transcriptional Pause Sites to Promote Xrn2-Dependent Termination Konstantina Skourti-Stathaki, Nicholas
Supporting Information
 Supporting Information Table S1. Oligonucleotide sequences used in this work Oligo DNA A B C D CpG-A CpG-B CpG-C CpG-D Sequence 5 ACA TTC CTA AGT CTG AAA CAT TAC AGC TTG CTA CAC GAG AAG AGC CGC CAT AGT
Supporting Information Table S1. Oligonucleotide sequences used in this work Oligo DNA A B C D CpG-A CpG-B CpG-C CpG-D Sequence 5 ACA TTC CTA AGT CTG AAA CAT TAC AGC TTG CTA CAC GAG AAG AGC CGC CAT AGT
Gene synthesis by circular assembly amplification
 Gene synthesis by circular assembly amplification Duhee Bang & George M Church Supplementary figures and text: Supplementary Figure 1. Dpo4 gene (1.05kb) construction by various methods. Supplementary
Gene synthesis by circular assembly amplification Duhee Bang & George M Church Supplementary figures and text: Supplementary Figure 1. Dpo4 gene (1.05kb) construction by various methods. Supplementary
Anti-Pim-1 (Cat#3247), anti-met (Cat#3127), anti-ron (Cat#2654), Anti-EGFR
 Supplementary Methods Antibodies Anti-Pim-1 (Cat#3247), anti-met (Cat#3127), anti-ron (Cat#2654), Anti-EGFR (Cat#2646), anti-igf1r (Cat#3018), anti-insr (Cat#3020), anti-akt (pan, Cat#4691), anti-phospho-akt
Supplementary Methods Antibodies Anti-Pim-1 (Cat#3247), anti-met (Cat#3127), anti-ron (Cat#2654), Anti-EGFR (Cat#2646), anti-igf1r (Cat#3018), anti-insr (Cat#3020), anti-akt (pan, Cat#4691), anti-phospho-akt
An engineered tryptophan zipper-type peptide as a molecular recognition scaffold
 SUPPLEMENTARY MATERIAL An engineered tryptophan zipper-type peptide as a molecular recognition scaffold Zihao Cheng and Robert E. Campbell* Supplementary Methods Library construction for FRET-based screening
SUPPLEMENTARY MATERIAL An engineered tryptophan zipper-type peptide as a molecular recognition scaffold Zihao Cheng and Robert E. Campbell* Supplementary Methods Library construction for FRET-based screening
II 0.95 DM2 (RPP1) DM3 (At3g61540) b
 Table S2. F 2 Segregation Ratios at 16 C, Related to Figure 2 Cross n c Phenotype Model e 2 Locus A Locus B Normal F 1 -like Enhanced d Uk-1/Uk-3 149 64 36 49 DM2 (RPP1) DM1 (SSI4) a Bla-1/Hh-0 F 3 111
Table S2. F 2 Segregation Ratios at 16 C, Related to Figure 2 Cross n c Phenotype Model e 2 Locus A Locus B Normal F 1 -like Enhanced d Uk-1/Uk-3 149 64 36 49 DM2 (RPP1) DM1 (SSI4) a Bla-1/Hh-0 F 3 111
SUPPLEMENTARY MATERIALS AND METHODS. E. coli strains, plasmids, and growth conditions. Escherichia coli strain P90C (1)
 SUPPLEMENTARY MATERIALS AND METHODS E. coli strains, plasmids, and growth conditions. Escherichia coli strain P90C (1) dinb::kan (lab stock) derivative was used as wild-type. MG1655 alka tag dinb (2) is
SUPPLEMENTARY MATERIALS AND METHODS E. coli strains, plasmids, and growth conditions. Escherichia coli strain P90C (1) dinb::kan (lab stock) derivative was used as wild-type. MG1655 alka tag dinb (2) is
G+C content. 1 Introduction. 2 Chromosomes Topology & Counts. 3 Genome size. 4 Replichores and gene orientation. 5 Chirochores.
 1 Introduction 2 Chromosomes Topology & Counts 3 Genome size 4 Replichores and gene orientation 5 Chirochores 6 7 Codon usage 121 marc.bailly-bechet@univ-lyon1.fr Bacterial genome structures Introduction
1 Introduction 2 Chromosomes Topology & Counts 3 Genome size 4 Replichores and gene orientation 5 Chirochores 6 7 Codon usage 121 marc.bailly-bechet@univ-lyon1.fr Bacterial genome structures Introduction
MacBlunt PCR Cloning Kit Manual
 MacBlunt PCR Cloning Kit Manual Shipping and Storage MacBlunt PCR Cloning Kits are shipped on dry ice. Each kit contains a box with cloning reagents and an attached bag with Eco-Blue Competent Cells (optional).
MacBlunt PCR Cloning Kit Manual Shipping and Storage MacBlunt PCR Cloning Kits are shipped on dry ice. Each kit contains a box with cloning reagents and an attached bag with Eco-Blue Competent Cells (optional).
Supporting Information
 Supporting Information Transfection of DNA Cages into Mammalian Cells Email: a.turberfield@physics.ox.ac.uk Table of Contents Supporting Figure 1 DNA tetrahedra used in transfection experiments 2 Supporting
Supporting Information Transfection of DNA Cages into Mammalian Cells Email: a.turberfield@physics.ox.ac.uk Table of Contents Supporting Figure 1 DNA tetrahedra used in transfection experiments 2 Supporting
NAME:... MODEL ANSWER... STUDENT NUMBER:... Maximum marks: 50. Internal Examiner: Hugh Murrell, Computer Science, UKZN
 COMP710, Bioinformatics with Julia, Test One, Thursday the 20 th of April, 2017, 09h30-11h30 1 NAME:...... MODEL ANSWER... STUDENT NUMBER:...... Maximum marks: 50 Internal Examiner: Hugh Murrell, Computer
COMP710, Bioinformatics with Julia, Test One, Thursday the 20 th of April, 2017, 09h30-11h30 1 NAME:...... MODEL ANSWER... STUDENT NUMBER:...... Maximum marks: 50 Internal Examiner: Hugh Murrell, Computer
Supplementary Information. Construction of Lasso Peptide Fusion Proteins
 Supplementary Information Construction of Lasso Peptide Fusion Proteins Chuhan Zong 1, Mikhail O. Maksimov 2, A. James Link 2,3 * Departments of 1 Chemistry, 2 Chemical and Biological Engineering, and
Supplementary Information Construction of Lasso Peptide Fusion Proteins Chuhan Zong 1, Mikhail O. Maksimov 2, A. James Link 2,3 * Departments of 1 Chemistry, 2 Chemical and Biological Engineering, and
A Second Allele of eppa in Borrelia burgdorferi Strain B31 Is Located on the Previously Undetected Circular Plasmid cp9-2
 JOURNAL OF BACTERIOLOGY, Nov. 2000, p. 6254 6258 Vol. 182, No. 21 0021-9193/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. A Second Allele of eppa in Borrelia burgdorferi
JOURNAL OF BACTERIOLOGY, Nov. 2000, p. 6254 6258 Vol. 182, No. 21 0021-9193/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. A Second Allele of eppa in Borrelia burgdorferi
evaluated with UAS CLB eliciting UAS CIT -N Libraries increase in the
 Supplementary Figures Supplementary Figure 1: Promoter scaffold library assemblies. Many ensembless of libraries were evaluated in this work. As a legend, the box outline color in top half of the figure
Supplementary Figures Supplementary Figure 1: Promoter scaffold library assemblies. Many ensembless of libraries were evaluated in this work. As a legend, the box outline color in top half of the figure
Genomics and Gene Recognition Genes and Blue Genes
 Genomics and Gene Recognition Genes and Blue Genes November 1, 2004 Prokaryotic Gene Structure prokaryotes are simplest free-living organisms studying prokaryotes can give us a sense what is the minimum
Genomics and Gene Recognition Genes and Blue Genes November 1, 2004 Prokaryotic Gene Structure prokaryotes are simplest free-living organisms studying prokaryotes can give us a sense what is the minimum
Supplementary Fig. 1. Isolation and in vitro expansion of EpCAM + cholangiocytes. For collagenase perfusion, enzyme solution was injected from the
 Supplementary Fig. 1. Isolation and in vitro expansion of EpCAM + cholangiocytes. For collagenase perfusion, enzyme solution was injected from the portal vein for digesting adult livers, whereas it was
Supplementary Fig. 1. Isolation and in vitro expansion of EpCAM + cholangiocytes. For collagenase perfusion, enzyme solution was injected from the portal vein for digesting adult livers, whereas it was
Supplemental Data. Bennett et al. (2010). Plant Cell /tpc
 BRN1 ---------MSSSNGGVPPGFRFHPTDEELLHYYLKKKISYEKFEMEVIKEVDLNKIEPWDLQDRCKIGSTPQNEWYFFSHKDRKYPTGS 81 BRN2 --------MGSSSNGGVPPGFRFHPTDEELLHYYLKKKISYQKFEMEVIREVDLNKLEPWDLQERCKIGSTPQNEWYFFSHKDRKYPTGS 82 SMB
BRN1 ---------MSSSNGGVPPGFRFHPTDEELLHYYLKKKISYEKFEMEVIKEVDLNKIEPWDLQDRCKIGSTPQNEWYFFSHKDRKYPTGS 81 BRN2 --------MGSSSNGGVPPGFRFHPTDEELLHYYLKKKISYQKFEMEVIREVDLNKLEPWDLQERCKIGSTPQNEWYFFSHKDRKYPTGS 82 SMB
S4B fluorescence (AU)
 A S4B fluorescence (AU) S4B fluorescence (AU) dsbb csgba csgd dsbb csgba bcsa 5000 * NS NS 4000 * 3000 2000 1000 0 ΔcsgBAΔbcsA ΔcsgDΔdsbBΔbcsA ΔcsgBA ΔdsbBΔcsgBA ΔcsgDΔdsbB B -1000 4000 * * NS 3500 * 3000
A S4B fluorescence (AU) S4B fluorescence (AU) dsbb csgba csgd dsbb csgba bcsa 5000 * NS NS 4000 * 3000 2000 1000 0 ΔcsgBAΔbcsA ΔcsgDΔdsbBΔbcsA ΔcsgBA ΔdsbBΔcsgBA ΔcsgDΔdsbB B -1000 4000 * * NS 3500 * 3000
SUPPORTING INFORMATION FILE
 Intrinsic and extrinsic connections of Tet3 dioxygenase with CXXC zinc finger modules Nan Liu, Mengxi Wang, Wen Deng, Christine S. Schmidt, Weihua Qin, Heinrich Leonhardt and Fabio Spada Department of
Intrinsic and extrinsic connections of Tet3 dioxygenase with CXXC zinc finger modules Nan Liu, Mengxi Wang, Wen Deng, Christine S. Schmidt, Weihua Qin, Heinrich Leonhardt and Fabio Spada Department of
Supporting Online Information
 Supporting Online Information Isolation of Human Genomic DNA Sequences with Expanded Nucleobase Selectivity Preeti Rathi, Sara Maurer, Grzegorz Kubik and Daniel Summerer* Department of Chemistry and Chemical
Supporting Online Information Isolation of Human Genomic DNA Sequences with Expanded Nucleobase Selectivity Preeti Rathi, Sara Maurer, Grzegorz Kubik and Daniel Summerer* Department of Chemistry and Chemical
Supplemental Information. Target-Mediated Protection of Endogenous. MicroRNAs in C. elegans. Inventory of Supplementary Information
 Developmental Cell, Volume 20 Supplemental Information Target-Mediated Protection of Endogenous MicroRNAs in C. elegans Saibal Chatterjee, Monika Fasler, Ingo Büssing, and Helge Großhans Inventory of Supplementary
Developmental Cell, Volume 20 Supplemental Information Target-Mediated Protection of Endogenous MicroRNAs in C. elegans Saibal Chatterjee, Monika Fasler, Ingo Büssing, and Helge Großhans Inventory of Supplementary
2
 1 2 3 4 5 6 7 Supplemental Table 1. Magnaporthe oryzae strains generated in this study. Strain background Genotype Strain name Description Guy-11 H1:RFP H1:RFP Strain expressing Histone H1- encoding gene
1 2 3 4 5 6 7 Supplemental Table 1. Magnaporthe oryzae strains generated in this study. Strain background Genotype Strain name Description Guy-11 H1:RFP H1:RFP Strain expressing Histone H1- encoding gene
SAY IT WITH DNA: Protein Synthesis Activity by Larry Flammer
 TEACHER S GUIDE SAY IT WITH DNA: Protein Synthesis Activity by Larry Flammer SYNOPSIS This activity uses the metaphor of decoding a secret message for the Protein Synthesis process. Students teach themselves
TEACHER S GUIDE SAY IT WITH DNA: Protein Synthesis Activity by Larry Flammer SYNOPSIS This activity uses the metaphor of decoding a secret message for the Protein Synthesis process. Students teach themselves
SUPPLEMENTARY INFORMATION
 doi:1.138/nature11496 Cl. 8 Cl. E93 Rag1 -/- 3H9 + BM Rag1 -/- BM CD CD c-kit c-kit c-kit wt Spleen c-kit B22 B22 IgM IgM IgM Supplementary Figure 1. FACS analysis of single-cell-derived pre-b cell clones.
doi:1.138/nature11496 Cl. 8 Cl. E93 Rag1 -/- 3H9 + BM Rag1 -/- BM CD CD c-kit c-kit c-kit wt Spleen c-kit B22 B22 IgM IgM IgM Supplementary Figure 1. FACS analysis of single-cell-derived pre-b cell clones.
Variations in Barbour-Stoenner-Kelly Culture Medium Modulate Infectivity and Pathogenicity of Borrelia burgdorferi Clinical Isolates
 INFECTION AND IMMUNITY, Nov. 2004, p. 6702 6706 Vol. 72, No. 11 0019-9567/04/$08.00 0 DOI: 10.1128/IAI.72.11.6702 6706.2004 Copyright 2004, American Society for Microbiology. All Rights Reserved. Variations
INFECTION AND IMMUNITY, Nov. 2004, p. 6702 6706 Vol. 72, No. 11 0019-9567/04/$08.00 0 DOI: 10.1128/IAI.72.11.6702 6706.2004 Copyright 2004, American Society for Microbiology. All Rights Reserved. Variations
SUPPLEMENTARY INFORMATION
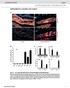 doi: 10.1038/nature07182 SUPPLEMENTAL FIGURES AND TABLES Fig. S1. myf5-expressing cells give rise to brown fat depots and skeletal muscle (a) Perirenal BAT from control (cre negative) and myf5-cre:r26r3-yfp
doi: 10.1038/nature07182 SUPPLEMENTAL FIGURES AND TABLES Fig. S1. myf5-expressing cells give rise to brown fat depots and skeletal muscle (a) Perirenal BAT from control (cre negative) and myf5-cre:r26r3-yfp
SUPPORTING INFORMATION
 SUPPORTING INFORMATION Investigation of the Biosynthesis of the Lasso Peptide Chaxapeptin Using an E. coli-based Production System Helena Martin-Gómez, Uwe Linne, Fernando Albericio, Judit Tulla-Puche,*
SUPPORTING INFORMATION Investigation of the Biosynthesis of the Lasso Peptide Chaxapeptin Using an E. coli-based Production System Helena Martin-Gómez, Uwe Linne, Fernando Albericio, Judit Tulla-Puche,*
Expression of Recombinant Proteins
 Expression of Recombinant Proteins Uses of Cloned Genes sequencing reagents (eg, probes) protein production insufficient natural quantities modify/mutagenesis library screening Expression Vector Features
Expression of Recombinant Proteins Uses of Cloned Genes sequencing reagents (eg, probes) protein production insufficient natural quantities modify/mutagenesis library screening Expression Vector Features
11th Meeting of the Science Working Group. Lima, Peru, October 2012 SWG-11-JM-11
 11th Meeting of the Science Working Group Lima, Peru, 15-19 October 2012 Russian population genetics studies of jack mackerel in the South Pacific P.K.Afanasiev M.A.Rabchun A.I.Glubokov Introduction. In
11th Meeting of the Science Working Group Lima, Peru, 15-19 October 2012 Russian population genetics studies of jack mackerel in the South Pacific P.K.Afanasiev M.A.Rabchun A.I.Glubokov Introduction. In
BioInformatics and Computational Molecular Biology. Course Website
 BioInformatics and Computational Molecular Biology Course Website http://bioinformatics.uchc.edu What is Bioinformatics Bioinformatics upgrades the information content of biological measurements. Discovery
BioInformatics and Computational Molecular Biology Course Website http://bioinformatics.uchc.edu What is Bioinformatics Bioinformatics upgrades the information content of biological measurements. Discovery
SUPPLEMENTAL MATERIAL GENOTYPING WITH MULTIPLEXING TARGETED RESEQUENCING
 SUPPLEMENTAL MATERIAL GENOTYPING WITH MULTIPLEXING TARGETED RESEQUENCING All of the patients and control subjects were sequenced and genotyped in the same way. Shotgun libraries of approximately 250 bp
SUPPLEMENTAL MATERIAL GENOTYPING WITH MULTIPLEXING TARGETED RESEQUENCING All of the patients and control subjects were sequenced and genotyped in the same way. Shotgun libraries of approximately 250 bp
A Genetically Encoded Toolbox for Glycocalyx Engineering: Tunable Control of Cell Adhesion,
 TITLE A Genetically Encoded Toolbox for Glycocalyx Engineering: Tunable Control of Cell Adhesion, Survival, and Cancer Cell Behaviors AUTHORS Carolyn R. Shurer *, Marshall J. Colville *, Vivek K. Gupta,
TITLE A Genetically Encoded Toolbox for Glycocalyx Engineering: Tunable Control of Cell Adhesion, Survival, and Cancer Cell Behaviors AUTHORS Carolyn R. Shurer *, Marshall J. Colville *, Vivek K. Gupta,
MCB421 FALL2005 EXAM#3 ANSWERS Page 1 of 12. ANSWER: Both transposon types form small duplications of adjacent host DNA sequences.
 Page 1 of 12 (10pts) 1. There are two mechanisms for transposition used by bacterial transposable elements: replicative (Tn3) and non-replicative (Tn5 and Tn10). Compare and contrast the two mechanisms
Page 1 of 12 (10pts) 1. There are two mechanisms for transposition used by bacterial transposable elements: replicative (Tn3) and non-replicative (Tn5 and Tn10). Compare and contrast the two mechanisms
PCR-based Markers and Cut Flower Longevity in Carnation
 PCRbased Markers and Cut Flower Longevity in Carnation Laura De Benedetti, Luca Braglia, Simona Bruna, Gianluca Burchi *, Antonio Mercuri and Tito Schiva Istituto Sperimentale per la Floricoltura, Corso
PCRbased Markers and Cut Flower Longevity in Carnation Laura De Benedetti, Luca Braglia, Simona Bruna, Gianluca Burchi *, Antonio Mercuri and Tito Schiva Istituto Sperimentale per la Floricoltura, Corso
Primer Design Workshop. École d'été en géné-que des champignons 2012 Dr. Will Hintz University of Victoria
 Primer Design Workshop École d'été en géné-que des champignons 2012 Dr. Will Hintz University of Victoria Scenario You have discovered the presence of a novel endophy5c organism living inside the cells
Primer Design Workshop École d'été en géné-que des champignons 2012 Dr. Will Hintz University of Victoria Scenario You have discovered the presence of a novel endophy5c organism living inside the cells
SUPPLEMENTARY MATERIALS
 SUPPLEMENTARY MATERIALS Supplementary Table S1: Water sampling sites in Brisbane River and their characteristics Sampling sites GPS coordinates Site characteristics Suspected source of fecal pollution
SUPPLEMENTARY MATERIALS Supplementary Table S1: Water sampling sites in Brisbane River and their characteristics Sampling sites GPS coordinates Site characteristics Suspected source of fecal pollution
Search for and Analysis of Single Nucleotide Polymorphisms (SNPs) in Rice (Oryza sativa, Oryza rufipogon) and Establishment of SNP Markers
 DNA Research 9, 163 171 (2002) Search for and Analysis of Single Nucleotide Polymorphisms (SNPs) in Rice (Oryza sativa, Oryza rufipogon) and Establishment of SNP Markers Shinobu Nasu, Junko Suzuki, Rieko
DNA Research 9, 163 171 (2002) Search for and Analysis of Single Nucleotide Polymorphisms (SNPs) in Rice (Oryza sativa, Oryza rufipogon) and Establishment of SNP Markers Shinobu Nasu, Junko Suzuki, Rieko
Supplementary Methods Quantitative RT-PCR. For mrna, total RNA was prepared using TRIzol reagent (Invitrogen) and genomic DNA was eliminated with TURB
 Supplementary Methods Quantitative RT-PCR. For mrna, total RNA was prepared using TRIzol reagent (Invitrogen) and genomic DNA was eliminated with TURBO DNA-free Kit (Ambion). One µg of total RNA was reverse
Supplementary Methods Quantitative RT-PCR. For mrna, total RNA was prepared using TRIzol reagent (Invitrogen) and genomic DNA was eliminated with TURBO DNA-free Kit (Ambion). One µg of total RNA was reverse
MOLECULAR CLONING AND SEQUENCING OF FISH MPR 46
 MOLECULAR CLONING AND SEQUENCING OF FISH MPR 46 INTRODUCTION The mammalian MPR 46 (human, mouse, bovine) sequences show extensive homologies. Among the non-mammalian vertebrates, only a partial sequence
MOLECULAR CLONING AND SEQUENCING OF FISH MPR 46 INTRODUCTION The mammalian MPR 46 (human, mouse, bovine) sequences show extensive homologies. Among the non-mammalian vertebrates, only a partial sequence
-15 diopter negative lenses in wild-type and homozygous CHRM2-deleted mice, and
 Supplementary Materials Supplementary Figure 1: Myopia induction was performed using uniocular -10 and -15 diopter negative lenses in wild-type and homozygous CHRM2-deleted mice, and results at 2, 4 and
Supplementary Materials Supplementary Figure 1: Myopia induction was performed using uniocular -10 and -15 diopter negative lenses in wild-type and homozygous CHRM2-deleted mice, and results at 2, 4 and
Lecture 19A. DNA computing
 Lecture 19A. DNA computing What exactly is DNA (deoxyribonucleic acid)? DNA is the material that contains codes for the many physical characteristics of every living creature. Your cells use different
Lecture 19A. DNA computing What exactly is DNA (deoxyribonucleic acid)? DNA is the material that contains codes for the many physical characteristics of every living creature. Your cells use different
Engineering D66N mutant using quick change site directed mutagenesis. Harkewal Singh 09/01/2010
 Engineering D66N mutant using quick change site directed mutagenesis Harkewal Singh 09/01/2010 1 1- What is quick change site directed mutagenesis? 2- An overview of the kit contents. 3- A brief information
Engineering D66N mutant using quick change site directed mutagenesis Harkewal Singh 09/01/2010 1 1- What is quick change site directed mutagenesis? 2- An overview of the kit contents. 3- A brief information
Complexity of the Ruminococcus flavefaciens FD-1 cellulosome reflects an expansion of family-related protein-protein interactions
 Complexity of the Ruminococcus flavefaciens FD-1 cellulosome reflects an expansion of family-related protein-protein interactions Vered Israeli-Ruimy 1,*, Pedro Bule 2,*, Sadanari Jindou 3, Bareket Dassa
Complexity of the Ruminococcus flavefaciens FD-1 cellulosome reflects an expansion of family-related protein-protein interactions Vered Israeli-Ruimy 1,*, Pedro Bule 2,*, Sadanari Jindou 3, Bareket Dassa
Efficient genome replication of hepatitis B virus using adenovirus vector: a compact pregenomic RNA-expression unit
 Efficient genome replication of hepatitis B virus using adenovirus vector: a compact pregenomic RNA-expression unit Mariko Suzuki 1, Saki Kondo 1, Manabu Yamasaki 2, Norie Matsuda 2, Akio Nomoto 2, Tetsuro
Efficient genome replication of hepatitis B virus using adenovirus vector: a compact pregenomic RNA-expression unit Mariko Suzuki 1, Saki Kondo 1, Manabu Yamasaki 2, Norie Matsuda 2, Akio Nomoto 2, Tetsuro
Appendix 1a. Microsatellite analysis of P1-hyg, P2-neo and their. Amplified cdna (base pairs) P1-hyg P2-neo All progeny A A
 Supplementary information Appendix 1a. Microsatellite analysis of P1-hyg, P2-neo and their double drug resistant progeny (ABI 377). Primer set a,b Amplified cdna (base pairs) P1-hyg P2-neo All progeny
Supplementary information Appendix 1a. Microsatellite analysis of P1-hyg, P2-neo and their double drug resistant progeny (ABI 377). Primer set a,b Amplified cdna (base pairs) P1-hyg P2-neo All progeny
Supplementary Figure 1. The level of pri-mir-8 gradually decreases while those of BR-C and E74 increase during 3rd instar larval development.
 qrt-pcr RT-PCR Relative pri-mir-8 level 1.2 1.0 0.8 0.6 0.4 0.2 0.0 Early 3rd (72h) Mid 3rd (96h) Late 3rd (119h) BR-C E74 mtl rrna Early 3rd (72h) Mid 3rd (96h) Late 3rd (119h) Supplementary Figure 1.
qrt-pcr RT-PCR Relative pri-mir-8 level 1.2 1.0 0.8 0.6 0.4 0.2 0.0 Early 3rd (72h) Mid 3rd (96h) Late 3rd (119h) BR-C E74 mtl rrna Early 3rd (72h) Mid 3rd (96h) Late 3rd (119h) Supplementary Figure 1.
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Coding and noncoding transcription at the vg locus. (a,b) qpcr analysis of developmental and tissue-specific vg mrna (a) and PRE/TRE (b) transcription, shown as percentage of TBP
Supplementary Figure 1 Coding and noncoding transcription at the vg locus. (a,b) qpcr analysis of developmental and tissue-specific vg mrna (a) and PRE/TRE (b) transcription, shown as percentage of TBP
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Blanken MO, Rovers MM, Molenaar JM, et al. Respiratory syncytial
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Blanken MO, Rovers MM, Molenaar JM, et al. Respiratory syncytial
SUPPLEMENTAL TABLE S1. Additional descriptions of plasmid constructions and the oligonucleotides used Plasmid or Oligonucleotide
 SUPPLEMENTAL TABLE S1. Additional descriptions of plasmid constructions and the oligonucleotides used Plasmid or Oligonucleotide former/ working Description a designation Plasmids pes213a b pes213-tn5
SUPPLEMENTAL TABLE S1. Additional descriptions of plasmid constructions and the oligonucleotides used Plasmid or Oligonucleotide former/ working Description a designation Plasmids pes213a b pes213-tn5
Chapter 13 Chromatin Structure and its Effects on Transcription
 Chapter 13 Chromatin Structure and its Effects on Transcription Students must be positive that they understand standard PCR. There is a resource on the web for this purpose. Warn them before this class.
Chapter 13 Chromatin Structure and its Effects on Transcription Students must be positive that they understand standard PCR. There is a resource on the web for this purpose. Warn them before this class.
Supplementary Information
 Supplementary Information Microbead-based biomimetic synthetic neighbors enhance survival and function of rat pancreatic β-cells Wei Li, a Samuel Lee, b Minglin Ma, a, f Soo Min Kim, b Patrick Guye, c
Supplementary Information Microbead-based biomimetic synthetic neighbors enhance survival and function of rat pancreatic β-cells Wei Li, a Samuel Lee, b Minglin Ma, a, f Soo Min Kim, b Patrick Guye, c
High-throughput cloning and expression in recalcitrant bacteria
 High-throughput cloning and expression in recalcitrant bacteria Eric R Geertsma & Bert Poolman Supplementary text and figures: Supplementary Figure 1 Frequency of SfiI sites yielding identical 3 extensions
High-throughput cloning and expression in recalcitrant bacteria Eric R Geertsma & Bert Poolman Supplementary text and figures: Supplementary Figure 1 Frequency of SfiI sites yielding identical 3 extensions
Infection of Mice with Lyme Disease Spirochetes Constitutively Producing Outer Surface Proteins A and B
 INFECTION AND IMMUNITY, June 2007, p. 2786 2794 Vol. 75, No. 6 0019-9567/07/$08.00 0 doi:10.1128/iai.01307-06 Copyright 2007, American Society for Microbiology. All Rights Reserved. Infection of Mice with
INFECTION AND IMMUNITY, June 2007, p. 2786 2794 Vol. 75, No. 6 0019-9567/07/$08.00 0 doi:10.1128/iai.01307-06 Copyright 2007, American Society for Microbiology. All Rights Reserved. Infection of Mice with
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2014 Electronic Supplementary Information Multiplexed Detection of Lung Cancer Cells at the Single-Molecule
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2014 Electronic Supplementary Information Multiplexed Detection of Lung Cancer Cells at the Single-Molecule
SUPPLEMENTARY INFORMATION
 1. RNA/DNA sequences used in this study 2. Height and stiffness measurements on hybridized molecules 3. Stiffness maps at varying concentrations of target DNA 4. Stiffness measurements on RNA/DNA hybrids.
1. RNA/DNA sequences used in this study 2. Height and stiffness measurements on hybridized molecules 3. Stiffness maps at varying concentrations of target DNA 4. Stiffness measurements on RNA/DNA hybrids.
Supporting Information
 Supporting Information Wiley-VCH 2007 69451 Weinheim, Germany Site-Specific Control of Distances between Gold Nanoparticles using Phosphorothioate Anchors on DNA and a Short Bifunctional Molecular Fastener
Supporting Information Wiley-VCH 2007 69451 Weinheim, Germany Site-Specific Control of Distances between Gold Nanoparticles using Phosphorothioate Anchors on DNA and a Short Bifunctional Molecular Fastener
Nongenetic Reprogramming of the Ligand Specificity. of Growth Factor Receptors by Bispecific DNA Aptamers
 Supporting Information For Nongenetic Reprogramming of the Ligand Specificity of Growth Factor Receptors by Bispecific DNA Aptamers Ryosuke Ueki,* Saki Atsuta, Ayaka Ueki and Shinsuke Sando* Department
Supporting Information For Nongenetic Reprogramming of the Ligand Specificity of Growth Factor Receptors by Bispecific DNA Aptamers Ryosuke Ueki,* Saki Atsuta, Ayaka Ueki and Shinsuke Sando* Department
