Comparative Analysis of Human Papillomavirus Detection by Polymerase Chain Reaction and Virapap/Viratype Kits
|
|
|
- Rosamund Gray
- 5 years ago
- Views:
Transcription
1 Comparative Analysis of Human Papillomavirus Detection by Polymerase Chain Reaction and Virapap/Viratype Kits GLENNA C. BURMER, M.D., PH.D., JAY D. PARKER, B.S., JEANNIE BATES, B.S., KATRINA EAST, B.S., AND BRUCE G. KULANDER, M.D. Cervical samples from 7 women referred by area physicians were analyzed for the presence of human papillomavirus (HPV) types,,, and 8 by the polymerase chain reaction (PCR) followed by Southern hybridization. Samples from S patients were concurrently analyzed by a commercial filter hybridization technique (Virapap and Viratype Kits, Life Technologies, Bethesda Research Labs, Gaithersburg, MD). The sensitivity of the Southern blot procedure combined with PCR was significantly higher than that of the Virapap and Viratype methods. HPV was detected in 7% of women who had positive results for dysplasia by PCR and in 7% by the Virapap method. HPV types /8 were found more commonly than types / in every diagnostic category. More than one HPV type was detected in % of HPV-positive patients. The prevalence of HPV in cytologically negative or indefinite patients as measured by PCR was % and %; in contrast, by the Virapap method, these values were 7% and %. These results demonstrate that PCR combined with Southern hybridization provides a higher level of sensitivity than methods that use hybridization without amplification of HPV DNA and also show that the prevalence of HPV is highest in cytologically positive smears. (Key words: Human papillomavirus; Polymerase catalyzed chain reaction; Virapap Kit; Cervical dysplasia) Am J Clin Pathol 99;9: 55-5 ALTHOUGH CERTAIN TYPES of human papillomaviruses (HPVs) have been strongly implicated in the genesis of cervical carcinoma, the exact prevalence of HPV and its role in the progressive development of neoplasia have not been clearly established. ' Prevalence data have depended on methods of detection such as in situ or filter hybridization with HPV-specific probes to detect the viruses. ' 9-5,8 Papillomaviruses have been detected in up to 9% of cervical carcinomas and between % and 8% of specimens exhibiting negative cytologic results. 8 ' 9 " 79 ' ' 7 ' ' Recent studies using the polymerase chain reaction (PCR) to amplify the virus have suggested that these prevalence rates are underestimates, and detection rates as high as 8% have been reported in cytologically negative patients. 5, ' Such high prevalence Received November 9, 989; received revised manuscript and accepted for publication March 9, 99. Supported in part by the NH NIDDK. grant DK97 and National Institute on Aging grant AG759 to Dr. Glenna Burmer. Address reprint requests to Dr. Burmer: SM-, Department of Pathology, School of Medicine, University of Washington, Seattle, Washington Department of Pathology, University of Washington School of Medicine and the Laboratory of Pathology of Seattle, Seattle, Washington rates in cytologically negative patients bring into question the clinical utility of screening for HPV. We have examined the prevalence of HPV,,, and 8 in 7 Seattle area patients referred to the Laboratory of Pathology of Seattle and the Swedish Hospital Medical Center, using the PCR combined with Southern hybridization to detect viral DNA sequences. In 5 cases, we have compared the sensitivity of detection by PCR and Southern hybridization with filter hybridization alone, using a commercially available assay, the Virapap and Viratype Kits (Life Technologies, Bethesda Research Labs, Gaithersburg, MD). Results obtained with the PCR and Virapap/Viratype assays were then correlated with cytologic classification and histopathologic diagnosis. Our data demonstrate the significantly increased sensitivity of detection when PCR is used to amplify the presence of the virus in patient samples. In contrast to recent reports, however, we detect significantly fewer positive samples by PCR in patients with negative or indefinite cytologic results. Our results therefore support the clinical utility of this assay for screening high-risk patients for papillomavirus. Study Design Experimental Design Two hundred seventy cervical samples were collected by Seattle area physicians between January and October 989, evaluated for the presence of HPV DNA at the University of Washington, and evaluated cytologically at the Laboratory of Pathology of Seattle. This referral-based patient population consisted of women attending clinics of area gynecologists and family practitioners. For cytologic evaluation, Papanicolaou (Pap) -stained cervical smears that had been obtained at the same time or within three months of HPV DNA analysis were evaluated by a cytopathologist (B.G.K.) without his knowledge of the results of the molecular studies. Smears were cat- 55 Downloaded from on 7 July 8
2 Vol. 9. No. 5 HUMAN PAPILLOMAVIRUS DETECTION BY PCR 555 egorized as negative, indefinite for dysplasia (I), cervical intraepithelial neoplasia (CIN) (mild dysplasia or condyloma), CIN (moderate dysplasia), or CIN (severe dysplasia). Patients with a negative diagnosis had a history of negative Pap smears and/or currently have negative findings for dysplasia; those with an indefinite for dysplasia diagnosis exhibited minimal cytologic alterations (e.g., inflammatory atypia) not diagnostic for condyloma or dysplasia. For HPV detection, cervical samples were collected by a Dacron swab using the Virapap Collection Kits according to the manufacturer's instructions (Life Technologies, Inc., Bethesda Research Labs, Gaithersburg, MD). Ten percent of the sample was immediately removed from the Virapap collection buffer, DNA was extracted, and ng of DNA was used as a template in PCR reactions using primers specific for HPV types,,, and 8 as detailed below. Ten percent of the PCR product was analyzed by Southern blotting using probes specific for each viral type. Controls for the PCR reactions consisted of primers amplified without template DNA or in the presence of human breast DNA (negative controls); and for positive controls, DNA extracted from HeLa cells (HPV 8), SiHa cells (HPV ), 9 and a patient with a condyloma known to be positive for HPV type /. Twenty-five percent of the remainder of the original sample was analyzed by the Virapap method, and viruspositive samples were further analyzed by the Viratype Kit according to the manufacturer's directions. Molecular analyses were all conducted without the examiner's prior knowledge of the corresponding cytologic results. DNA Extraction Genomic DNA was extracted from patient samples by standard methods. One hundred microliters from each Virapap Collection Kit was incubated overnight at 7 C in the presence of 5 fig/ml proteinase K and.5% (w/ v) sodium dodecyl sulfate in buffer containing mmol/ L TRIS-HC (ph 8.), 5 mmol/l NaCl, and mmol/ L EDTA, followed by serial phenol-chloroform (:) and chloroform extractions, and precipitated with. vol of mol/l sodium acetate (ph 5.) and.5 vol of ethanol at 7 C overnight. The DNA was resuspended in /il of water after centrifugation, and the concentration was determined spectrophotometrically. When less than ng of total DNA was present, 5 nl of the sample was used as a template in the PCR reaction; when larger amounts were recovered, ng was routinely used as a template for PCR. Polymerase Chain Reaction Genomic DNA was incubated in a total volume of nl in solution containing the following: 5 mmol/l KC; mmol/l TRIS-HC (ph 8.); 5 mmol/l MgCl ;.% (w/v) gelatin; Mmol/L each datp, dctp, dgtp, and dttp; ng/jul of oligonucleotide primers; and.5 units of Taq polymerase (Perkin-Elmer Cetus, Norwalk, CT). The mixture was overlaid with mineral oil and underwent cycles of PCR with denaturation for minute at 9 C, followed by -minute incubations at C (annealing) and minutes at 7 C (polymerization). 8 Samples were separately amplified for HPV types / and or 8 with the use of primers specific for each viral type (Table ). The oligonucleotide primers and probes were synthesized by our laboratory with the use of the beta-cyanoethyl phosphoramidite method on a Cruachem synthesizer (Cruachem Corporation, Herndon, VA). Southern and Dot Blot Hybridizations of PCR Products Ten microliters of the PCR product was electrophoresed on % (w/v) agarose, denatured for minutes by incubation in buffer containing.5 mol/l NaCl and.5 mol/ L NaOH, neutralized for minutes in mol/l NaCl with.5 mol/l TRIS-HC (ph 7.), and transferred to Nytran membranes (Schleicher and Schuell, Keene, NH). For dot blot hybridization, the DNA was denatured by addition of. vol of N NaOH and boiled for Table J. Oligonucleotide Primers and Probes Used in PCR Assay Primer Region Base Pair Reference Sequence /-U / l-d / l-p -U -D -P 8-U 8-D 8-P El El El , 5, 5, 5'-AAA GGT AAA GCG ACG GCT GTT-' 5'-TTT AAA TGG CCT AAT TAA ATC-' 5'-GGG AAC TAA CGG ACA GTG GAT ATG GCT ATT CTG AAG TGG AAG CT-' 5'-ATT AGT GAG TAT AGA CAT TA-' 5'-GGC TTT TGA CAG TTA ATA CA-' 5'-ATG GAA CAA CAT TAG AAC AGC AAT ACA ACA AAC CGT TCT G-' 5'-ACT ATG GCG CGC TTT GAG GAT CCA-' 5'-GGT TTC TGG CAC CGC AGG CA-' 5'-ATG GAG ACA CAT TGG AAA AAC TAA CTA ACA CTG GGT TAT-' U = upstream PCR primer: D = downstream PCR primer; P = probe. Downloaded from on 7 July 8
3 55 BURMER ET AL. A.J.C.P. November 99 minutes at C, neutralized by addition of.5 vol of 5 mol/l ammonium acetate (ph 5.) at C, and spotted onto Nytran membranes with the use of a Vacudot-VS apparatus (American Bionetics, Inc., Hayward, CA). The membranes were baked at 8 C for one hour; prehybridized for four hours in 5X SSC, X Denhardt's solution, Mg/mL salmon sperm DNA, and.% (w/v) sodium dodecyl sulfate (SDS) and hybridized overnight at 8 C with [7- P]-end-labeled oligonucleotide probes specific for HPV types /,, or 8. The membranes were washed in 5X SSC/.5% (w/v) SDS at 7 C for 5 minutes, followed by two 5-minute washes in.5x SSC/.5% (w/v) SDS at 5 C, and exposed for two to three days to x-ray film at 7 C with two intensifying screens. Experimental Results SOUTHERN BLOT OF PCR AMPLIFIED HPV DNA 5789 A m PCR Analysis of Cervical Specimens Validation of the PCR Assay for Detection of HPV Types /,, and 8. Initial experiments were conducted with tissues and cell lines known to contain specific HPV types to establish the sensitivity and specificity of the PCR assay used in these studies. PCR primers and probes specific for types /,, and 8 were synthesized after a Genebank dot plot analysis of regions exhibiting the least homology between viral types and within the region for HPV types and 8, and El region for HPV types / (Table ). The choice of sequences with least homology ensured the greatest specificity for viral subtypes. The region was chosen to maximize the sensitivity of detection of integrated, low copy number viral sequences in dysplastic lesions, which may have lost viral sequences less necessary for maintenance of the transformed state.' Figure shows a representative Southern blot of /il of the amplification product after cycles of PCR of DNA from HeLa cells (B, lane ), which contain -5 integrated copies of HPV 8 per genome ; SiHa cells (C, lane ), which contain - copies of HPV per genome 9 ; a patient with a urethral condyloma known to be positive for HPV / (A, lane ); and seven patient samples. The patient samples in lanes 7 and 8 are positive for HPV /, and the patient in lane 5 had positive results for HPV 8 and faintly positive results for HPV. The hybridization is specific for each viral type, and crosshybridization was not observed between the type-specific probes and the products of these PCR reactions, except in cases in which dual infection was apparent (e.g., Fig., patient sample 5). Hybridization also was not seen when samples of cultures from herpes simplex types I and II were amplified with HPV-specific primers and hybridized with probes for HPV types /,, or 8 (unpublished data). B c m~ ~^K ^sk! FIG.. Autoradiogram of Southern blot of PCR products after amplification with primers specific for the El region of HPV / (Panel A and Table, /-U and /-D), the region of HPV 8 (Panel B and Table, 8-U and 8-D), and HPV (Panel C and Table, - U and -D). DNA from HeLa cells (Lane ), SiHa cells (Lane ), a condyloma containing HPV type / (Lane ), and seven patient samples was used as a template in a -cycle PCR reaction. Ten percent of the PCR product was electrophoresed on a % (weight/volume) ethidium bromide-stained agarose gel, transferred to Nytran membranes, and each blot separately hybridized with end-labeled probes specific for HPV types / (/ -P), 8 (8-P) or (-P). Thirty percent of the PCR product from the type / positive control patient, SiHa cells, and HeLa cells (Fig. I, lanes,, and, respectively) was then directly sequenced with the use of an internally nested, kinased primer. The sequencing autoradiograms verified the identity of the amplified fragment as specific for types / II,, and 8 (unpublished data). The sensitivity of the PCR method was then tested by performance of serial dilutions of HeLa DNA before PCR and performance of dot blots with nl (%) of the reaction mixture after cycles of PCR (Fig. ). Adjacent wells were loaded with diluted HeLa DNA without am-,-a Downloaded from on 7 July 8
4 Vol. 9 No. 5 HUMAN PAPILLOMAVIRUS DETECTION BY PCR 557 WITH PCR NO PCR DOT BLOT OF HELA DNA A B C D E F G FIG.. Dot blot of products of PCR reaction of serially diluted HeLa DNA compared with DNA without PCR. The HeLa genomic DNA template was diluted and underwent cycles of PCR with type 8 specific primers and blotted onto Nytran (WITH PCR), or blotted at each dilution without amplification (NO PCR). DNA dilutions are as follows: Mg (A), ng (B), pg (Q, pg (D), pg ( ),. pg (F), and primers only (G). plification. The results demonstrate our ability to detect HPV 8 in pg of HeLa DNA, which contains approximately HeLa cell equivalent of DNA or -5 copies of virus. In contrast, our dot blot procedure using a labeled oligonucleotide probe was incapable of detecting HPV in jug of HeLa DNA, which represents an increase in sensitivity of, using PCR in conjunction with the dot blot, over using the dot blot alone for HPV detection. Although these results are not directly comparable to those of the Virapap/Viratype assay, which is a modified dot blot procedure that uses radiolabeled RNA probes corresponding to the full-length HPV genome, a minimum of, HeLa cells (e.g., ng of HeLa DNA) has previously been reported to be the lower limit of sensitivity of detection of HPV DNA by dot blot hybridization procedures that use full-length nick-translated HPV DNA probes. 7 ' 5, Therefore, this represents an increase in sensitivity of approximately, using PCR compared with commonly used filter hybridization methods. PCR Analysis of Clinical Samples. Two hundred seventy patients were analyzed by PCR and Southern hybridization for the prevalence of HPV types /,, and 8. Fifty percent of cases ( of 7) were positive for the presence of at least one virus by PCR; 8% ( of ) of these cases contained type or, 7% (9 of ) were positive for type or 8, and % ( of ) contained evidence of infection by viruses from both the / and /8 groups (Table ). Cases were then analyzed according to cytopathologic diagnosis. Within the group of patients without previous history and without current cytologic evidence for atypia, % (5 of 7) had positive results for HPV (Table ). Within the group of patients indefinite for dysplasia, 7% ( of 59) had positive results for HPV. A higher percentage of patients with cytologic results positive for dysplasia contained detectable HPV (%, 7%, and 7% for CIN,, and, respectively, Table ). The increased prevalence of HPV is reflected by an increase in both types / and /8 within the CIN population; however, the percentage of cases positive for HPV types / decreased in patients with moderate or severe dysplasia. In all diagnostic categories, the frequency of infection with types /8 was at least.9-fold greater than for types /. This difference was statistically significant for the diagnostic categories analyzed as a whole (P <. by a conditional binomial test). Analysis by Virapap/Viratype Kits One hundred fifty-four cases were concurrently analyzed for the presence of HPV by dot blot hybridization with the use of the Virapap and Viratype Kits. Overall, 9% (5 of 5) of cases had detectable HPV by the Virapap method (Table ). Of cases negative or indefinite for dysplasia, 8% ( of 7) contained evidence of HPV by the Virapap method. The percentage of Virapap-positive samples increased in cytologically positive cases (5% for CIN and 5% for CIN and ). Within the subset of 5 PCR amplified cases that were analyzed concurrently by the Virapap method, the percentages of positive cases in each diagnostic category did not differ significantly from the pool of total PCR cases. Cases were then analyzed according to specific viral types as distinguished by the Viratype Kit (/, /8, and //5, Table ). Because of the reduced sensitivity of the commercial Viratype Kit relative to Virapap screening, only of the 5 Virapap-positive cases could Table. PCR Positive and Cases in Each Diagnostic Class Diagnosis n / /8 Both Positive Percent Positive Indefinite CIN CIN CIN ± 5 7 ± ± ± 5+ Downloaded from on 7 July 8
5 558 BURMER ET AL. A.J.C.P. November 99 Table. PCR or Virapap Positive Cases in Each Diagnostic Category Table 5. Comparison of Virapap and PCR/Southern Hybridization Diagnosis n PCR Positive Virapap Positive % Indefinite CIN CIN CIN ± 8 ± ± + 7 ± Standard errors are shown for the percentage of PCR and Virapap positive cases. 7± 5 ± 5 5 ± 7 5 ± 5 ± 9 ± Virapap Positive Positive 7 PCR/Southern 9 8 Shown are the number of cases exhibiting concordance and discordance. 5 be definitively subtyped. The prevalences of types /8 and //5 were equal ( of = 8%) and.7 times more prevalent than type / ( of = 8%); cases ( of = 5%) were positive for more than one subgroup of HPV. The increased frequency of /8 and //5 over the / types existed in all diagnostic categories; this difference was statistically significant as assessed by a conditional binomial test (P <.5). Comparison of Virapap/Viratype Methods with PCR/Southern Blot Complete concordance between PCR/Southern and Virapap assays (both assays positive or negative for a given case) was obtained in % of cases ( of 5, Table 5). Of the 5 discordant cases, ( of 5 = 77%) consisted of those detected as positive by PCR and were negative by the Virapap method. Five cases were positive by the Virapap method and negative by both the PCR and Viratype assays; it is uncertain whether these cases represent false positive results for the Virapap assay or false negative results for both the PCR and Viratype tests. Seven cases were positive by both the Virapap and Viratype assays, yet negative by PCR. Of these seven cases, five were positive only for HPV type //5, which was not examined by our PCR-based assay. Twelve cases were identified by the Viratype assay as positive for HPV //5, alone or in combination with types / or /8 (Table ). Although our PCR assay was not designed to detect HPV //5, seven of the Viratype-positive assays were detected as positive for HPV by PCR. In addition to the two cases that were Viratype positive for / or /8, our PCR assay detected the presence of these subtypes in the remaining five cases that were classified by the Viratype method as only positive for //5. These cases may represent instances of crosshybridization between the Viratype probes for //5 and / or /8 DNA. Alternatively, these may be cases of dual infection in which the Viratype Kit was only able to detect //5 DNA because the second viral type was present at a concentration below the threshold of detection by this method. All cases positive by the Viratype method for / or /8 were also found positive for the same virus by the PCR assay; in addition, of the cases were positive by PCR for both / and or 8. When cases were analyzed according to cytologic classification, the PCR assay was found to be more sensitive than the Virapap method in detecting the presence of HPV in every diagnostic category (Table ). This difference is highly statistically significant when comparing the PCR group as a whole with the Virapap screen (P <. by McNemar's test, Table 5). Discussion Estimates of the prevalence of HPV infection rely on the sensitivity of the technique used to detect virus in patient samples. We have compared the sensitivity and specificity of a commercially available, established filter hybridization method (Virapap and Viratype) with an assay using the PCR in conjunction with Southern hybrid- Table. Viratype Positive Cases: Classification by Diagnosis and HPV Type Diagnosis Positive / /8 //5 Two or More Indefinite CIN CIN CIN (8 ± 5) 5 (8 ± 9) (8 ±9) (5±7) Shown in parentheses is the percentage of the total contributed by each viral group, plus or minus the standard error. Downloaded from on 7 July 8
6 Vol. 9. No. 5 HUMAN PAPILLOMAVIRUS DETECTION BY PCR 559 ization. Our results demonstrate the increased sensitivity of using PCR to detect HPV in patients with or without cytologic evidence of cervical intraepithelial neoplasia. The validity of our comparisons between the PCRbased assay and the dot blot hybridization assay has been demonstrated in several ways. The prevalence of HPV within our cytologically negative population as detected by the Virapap method is consistent with previous reports of the prevalence of HPV infection in Seattle area patients as assessed by Southern or dot blot hybridization methods. 9 Moreover, the increased prevalence of HPV types and 8 over types / within our patient population as detected by both the Viratype method and our PCR assay is consistent with a previous report. 9 The decreased prevalence of types / in higher categories of cytologic dysplasia compared with types /8 has been well established by traditional hybridization studies.' 5 ' ' 9 ' ' In addition, the frequency of dual infections within HPVpositive cases, as detected by both our Viratype and PCR assays, is similar to that reported previously. 5 The major difference between data obtained by our PCR assay and those obtained by Southern or dot blot hybridization alone resides in the increased sensitivity of detection of HPV in every diagnostic category. This difference is made particularly evident by the fact that, in many cases, only % of the patient's sample was used as a template in the PCR assay (% of original sample was used for DNA extraction, of which approximately % was used as a template in the PCR reaction), compared with the volumes used for analysis in the Virapap and Viratype Kits. Moreover, our assay only detected types /,, and 8, and our estimates of prevalence still exceed those detected by the Virapap Kit, which also includes probes for detecting HPV types //5. A practical disadvantage of the PCR procedure is that, because of its ability to amplify minute quantities of template DNA, there is an increased possibility of contamination, which may produce false positive results. We have guarded against this possibility by including negative controls (PCR reaction buffer, Taq polymerase, primers, and no template DNA) within each group of patient samples and by technical procedures that include cryopreservation of pretested, aliquoted PCR reaction mixes containing all components except for the template DNA. These procedures limit the number of manipulations to a minimum, decreasing the likelihood of contamination of patient samples. Despite the increased sensitivity of the PCR assay, the prevalence of HPV within cytologically negative patients (%) remained far short of recent estimates from other countries that report prevalence rates of 8%. 5 ' - The probability that this results from major differences in sensitivity between the PCR assays is unlikely, as demonstrated by our ability to detect HPV DNA in pg of genomic DNA. The differences between these data may reflect regional differences in prevalence and distribution of HPV types, differences in the definition of a cytologically negative or indefinite population, differences in sampling technique, or a combination of these factors. The increased prevalence of HPV types and 8 within cytologically positive cases as detected by PCR further supports the role of these HPV subtypes in the progression of cervical dysplasia. The relative ease, specificity, and sensitivity of the PCR assay will provide a powerful tool for the clinical identification of patients who may be at increased risk for the development of cervical carcinoma. References. Bedell MA, Jones KH, Laimins LA. The -E7 region of human papillomavirus type 8 is sufficient for transformation of NIH T and Rat- cells. J Virol 987;:5-.. Burmer GC, Rabinovitch PS, Loeb LA. Analysis of c-ki-ras mutations in human colon carcinoma by cell sorting, polymerase chain reaction, and DNA sequencing. Cancer Res 989;9:-.. Cole ST, Danos O. Nucleotide sequence and comparative analysis of the human papillomavirus type 8 genome. J Mol Biol 987:9: Crum CP, Mitao M, Levine RU, Silverstein S. Cervical papillomaviruses segregate within morphologically distinct precancerous lesions. J Virol 985;5: Dartmann K, Schwarz E, Gissmann L, Zur Hausen H. The nucleotide sequence and genome organization of human papilloma virus type. Virology 985;5:-.. Davis LG, Dibner MD, Battey JF. Preparation of DNA from eucaryotic cells. In: Basic methods in molecular biology. New York: Elsevier, 98:-. 7. Demeter T, Kulski JK, Rakoczy P, Sterrett GF, Pixley EC. Detection of human papillomavirus DNA in cell scrapes and formalin-fixed, paraffin-embedded tissue of the uterine cervix by filter in situ hybridization. J Med Virol 988;: De Villiers EM, Wagner D, Schneider A, et al. Human papillomavirus infections in women with and without abnormal cervical cytology. Lancet 987;: Kiviat NB, Koutsky LA, Paavonen JA, et al. Prevalence of genital papillomavirus infection among women attending a college student health clinic or a sexually transmitted disease clinic. J Infect Dis 989;59:9-.. Koss LG. Diagnostic cytopathology, vol. rd ed. Philadelphia: JB Lippincott, 979: Lorincz AT, Temple GF, Patterson JA, Jenson AB, Kurman RJ, Lancaster WD. Correlation of cellular atypia and human papillomavirus deoxyribonucleic acid sequences in exfoliated cells of the uterine cervix. Obstet Gynecol 98;8: Matlashewski G, Schneider J, Banks L, Jones N, Murray A, Crawford L. Human papillomavirus type DNA cooperates with activated ras in transforming primary cells. EMBO J 987;:7-7.. McCance DJ, Clarkson PK, Dyson JL, Walker PG, Singer A. Human papillomavirus types and in multifocal intraepithelial neoplasia of the female lower genital tract. Br J Obstet Gynaecol 985;9:9-.. McCance DJ, Campion MJ, Singer A. Non-invasive detection of cervical papillomavirus DNA. Lancet 98;: McNicol PJ, Guijon FB, Paraskevas M, Brunham RC. Comparison of filter in situ deoxyribonucleic acid hybridization with cytologic, colposcopic, and histopathologic examination for detection of human papillomavirus infection in women with cervical intraepithelial neoplasia. Am J Obstet Gynecol 989;:5-7. Downloaded from on 7 July 8
7 5 BURMER ET AL. A.J.C.P. November 99. Pfister H. Human papillomaviruses and genital cancer. Adv Cancer Res 987;8: Pratili MA, Le Doussal V, Harvey P, et al. Recherche de papillomavirus humains dans des cellules epitheliales du col uterin: frequence des types et 8. Resultats preliminaires d'une etude clinique, cytologique et virologique. J Gynecol Obstet Biol Reprod (Paris) 98;5: Saiki RK, Scharf S, Faloona F, et al. Enzymatic amplification of beta-globin sequences and restriction site analysis for diagnosis of sickle cell anemia. Science 985;: Schneider A, Kraus H, Schuhmann R, Gissmann L. Papillomavirus infection of the lower genital tract: detection of viral DNA in gynecological swabs. Int J Cancer 985;5:-8.. Schneider A, Sawada E, Gissmann L, Shah K. Human papillomaviruses in women with a history of abnormal Papanicolaou smears and in their male partners. Obstet Gynecol 987;9: Schwarz E, Durst M, Demankowski C, et al. DNA sequence and genome organization of genital human papillomavirus type B. EMBOJ 98;:-8.. Schwarz E, Freese UK, Gissmann L, et al. Structure and transcription of human papillomavirus sequences in cervical carcinoma cells. Nature 985;:-.. Seedorf K, KrammerG, Durst M, Suhai S, Rowekamp WG. Human papillomavirus type DNA sequence. Virology 985; 5: Shibata DK, Arnheim N, Martin WJ. Detection of human papillomavirus in paraffin-embedded tissue using the polymerase chain reaction. J Exp Med 988;: Tidy JA, Parry GCN, Ward P, et al. High rate of human papillomavirus infection in cytologically normal cervices. Lancet 989; :.. Tidy JA, Vousden KH, Farrell PJ. Relation between infection with a subtype of HPV and cervical neoplasia. Lancet 989; : Toon PG, Arrand JR, Wilson LP, Sharp DS. Human papillomavirus infection of the uterine cervix of women without cytological signs of neoplasia. Br Med J 98;9:-. 8. Wagner D, Ikenberg H, Boehm N, Gissmann L. Identification of human papillomavirus in cervical swabs by deoxyribonucleic acid in situ hybridization. Obstet Gynecol 98;: Yee C, Krishnan-Hewlett I, Baker CC, Schlegel R, Howley PM. Presence and expression of human papillomavirus sequences in human cervical carcinoma cell lines. Am J Pathol 985; 9:-.. Young LS, Bevan IS, Johnson MA, et al. The polymerase chain reaction: a new epidemiological tool for investigating cervical human papillomavirus infection. Br Med J 989;98:-8.. Zur Hausen H, Gissmann L, Schlehofer JR. Viruses in the etiology of human genital cancer. Prog Med Virol 98;:7-8.. Zur Hausen H, Schneider A. The role of papillomaviruses in human anogenital cancer. In: Salzman NP, Howley PM, eds. The Papovaviridae. The papillomaviruses. New York: Plenum, 987: 5-. Downloaded from on 7 July 8
Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH).
 Bisulfite Treatment of DNA Dilute DNA sample to 2µg DNA in 50µl ddh 2 O. Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH). Incubate in a 37ºC water bath for 30 minutes. To 55µl samples
Bisulfite Treatment of DNA Dilute DNA sample to 2µg DNA in 50µl ddh 2 O. Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH). Incubate in a 37ºC water bath for 30 minutes. To 55µl samples
Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR
 Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR 1 The problem We wish to clone a yet unknown gene from a known
Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR 1 The problem We wish to clone a yet unknown gene from a known
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
Supplementary Figure 1A A404 Cells +/- Retinoic Acid
 Supplementary Figure 1A A44 Cells +/- Retinoic Acid 1 1 H3 Lys4 di-methylation SM-actin VEC cfos (-) RA (+) RA 14 1 1 8 6 4 H3 Lys79 di-methylation SM-actin VEC cfos (-) RA (+) RA Supplementary Figure
Supplementary Figure 1A A44 Cells +/- Retinoic Acid 1 1 H3 Lys4 di-methylation SM-actin VEC cfos (-) RA (+) RA 14 1 1 8 6 4 H3 Lys79 di-methylation SM-actin VEC cfos (-) RA (+) RA Supplementary Figure
Cat. # Product Size DS130 DynaExpress TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1
 Product Name: Kit Component TA PCR Cloning Kit (ptakn-2) Cat. # Product Size DS130 TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1 2 Ligation Buffer
Product Name: Kit Component TA PCR Cloning Kit (ptakn-2) Cat. # Product Size DS130 TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1 2 Ligation Buffer
Supplemental Data Supplemental Figure 1.
 Supplemental Data Supplemental Figure 1. Silique arrangement in the wild-type, jhs, and complemented lines. Wild-type (WT) (A), the jhs1 mutant (B,C), and the jhs1 mutant complemented with JHS1 (Com) (D)
Supplemental Data Supplemental Figure 1. Silique arrangement in the wild-type, jhs, and complemented lines. Wild-type (WT) (A), the jhs1 mutant (B,C), and the jhs1 mutant complemented with JHS1 (Com) (D)
Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC
 Supplementary Appendixes Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC ACG TAG CTC CGG CTG GA-3 for vimentin, /5AmMC6/TCC CTC GCG CGT GGC TTC CGC
Supplementary Appendixes Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC ACG TAG CTC CGG CTG GA-3 for vimentin, /5AmMC6/TCC CTC GCG CGT GGC TTC CGC
PCR analysis was performed to show the presence and the integrity of the var1csa and var-
 Supplementary information: Methods: Table S1: Primer Name Nucleotide sequence (5-3 ) DBL3-F tcc ccg cgg agt gaa aca tca tgt gac tg DBL3-R gac tag ttt ctt tca ata aat cac tcg c DBL5-F cgc cct agg tgc ttc
Supplementary information: Methods: Table S1: Primer Name Nucleotide sequence (5-3 ) DBL3-F tcc ccg cgg agt gaa aca tca tgt gac tg DBL3-R gac tag ttt ctt tca ata aat cac tcg c DBL5-F cgc cct agg tgc ttc
Supplementary. Table 1: Oligonucleotides and Plasmids. complementary to positions from 77 of the SRα '- GCT CTA GAG AAC TTG AAG TAC AGA CTG C
 Supplementary Table 1: Oligonucleotides and Plasmids 913954 5'- GCT CTA GAG AAC TTG AAG TAC AGA CTG C 913955 5'- CCC AAG CTT ACA GTG TGG CCA TTC TGC TG 223396 5'- CGA CGC GTA CAG TGT GGC CAT TCT GCT G
Supplementary Table 1: Oligonucleotides and Plasmids 913954 5'- GCT CTA GAG AAC TTG AAG TAC AGA CTG C 913955 5'- CCC AAG CTT ACA GTG TGG CCA TTC TGC TG 223396 5'- CGA CGC GTA CAG TGT GGC CAT TCT GCT G
Legends for supplementary figures 1-3
 High throughput resistance profiling of Plasmodium falciparum infections based on custom dual indexing and Illumina next generation sequencing-technology Sidsel Nag 1,2 *, Marlene D. Dalgaard 3, Poul-Erik
High throughput resistance profiling of Plasmodium falciparum infections based on custom dual indexing and Illumina next generation sequencing-technology Sidsel Nag 1,2 *, Marlene D. Dalgaard 3, Poul-Erik
Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis
 1 2 3 4 5 6 7 8 9 10 11 12 Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis Information Research). Exons
1 2 3 4 5 6 7 8 9 10 11 12 Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis Information Research). Exons
Hes6. PPARα. PPARγ HNF4 CD36
 SUPPLEMENTARY INFORMATION Supplementary Table Positions and Sequences of ChIP primers -63 AGGTCACTGCCA -79 AGGTCTGCTGTG Hes6-0067 GGGCAaAGTTCA ACOT -395 GGGGCAgAGTTCA PPARα -309 GGCTCAaAGTTCAaGTTCA CPTa
SUPPLEMENTARY INFORMATION Supplementary Table Positions and Sequences of ChIP primers -63 AGGTCACTGCCA -79 AGGTCTGCTGTG Hes6-0067 GGGCAaAGTTCA ACOT -395 GGGGCAgAGTTCA PPARα -309 GGCTCAaAGTTCAaGTTCA CPTa
Supplementary Information
 Supplementary Information A general solution for opening double-stranded DNA for isothermal amplification Gangyi Chen, Juan Dong, Yi Yuan, Na Li, Xin Huang, Xin Cui* and Zhuo Tang* Supplementary Materials
Supplementary Information A general solution for opening double-stranded DNA for isothermal amplification Gangyi Chen, Juan Dong, Yi Yuan, Na Li, Xin Huang, Xin Cui* and Zhuo Tang* Supplementary Materials
Supplemental material
 Supplemental material Diversity of O-antigen repeat-unit structures can account for the substantial sequence variation of Wzx translocases Yaoqin Hong and Peter R. Reeves School of Molecular Bioscience,
Supplemental material Diversity of O-antigen repeat-unit structures can account for the substantial sequence variation of Wzx translocases Yaoqin Hong and Peter R. Reeves School of Molecular Bioscience,
Supporting information for Biochemistry, 1995, 34(34), , DOI: /bi00034a013
 Supporting information for Biochemistry, 1995, 34(34), 10807 10815, DOI: 10.1021/bi00034a013 LESNIK 10807-1081 Terms & Conditions Electronic Supporting Information files are available without a subscription
Supporting information for Biochemistry, 1995, 34(34), 10807 10815, DOI: 10.1021/bi00034a013 LESNIK 10807-1081 Terms & Conditions Electronic Supporting Information files are available without a subscription
Supplemental Information. Human Senataxin Resolves RNA/DNA Hybrids. Formed at Transcriptional Pause Sites. to Promote Xrn2-Dependent Termination
 Supplemental Information Molecular Cell, Volume 42 Human Senataxin Resolves RNA/DNA Hybrids Formed at Transcriptional Pause Sites to Promote Xrn2-Dependent Termination Konstantina Skourti-Stathaki, Nicholas
Supplemental Information Molecular Cell, Volume 42 Human Senataxin Resolves RNA/DNA Hybrids Formed at Transcriptional Pause Sites to Promote Xrn2-Dependent Termination Konstantina Skourti-Stathaki, Nicholas
Supporting Information
 Supporting Information Transfection of DNA Cages into Mammalian Cells Email: a.turberfield@physics.ox.ac.uk Table of Contents Supporting Figure 1 DNA tetrahedra used in transfection experiments 2 Supporting
Supporting Information Transfection of DNA Cages into Mammalian Cells Email: a.turberfield@physics.ox.ac.uk Table of Contents Supporting Figure 1 DNA tetrahedra used in transfection experiments 2 Supporting
PCR-based Markers and Cut Flower Longevity in Carnation
 PCRbased Markers and Cut Flower Longevity in Carnation Laura De Benedetti, Luca Braglia, Simona Bruna, Gianluca Burchi *, Antonio Mercuri and Tito Schiva Istituto Sperimentale per la Floricoltura, Corso
PCRbased Markers and Cut Flower Longevity in Carnation Laura De Benedetti, Luca Braglia, Simona Bruna, Gianluca Burchi *, Antonio Mercuri and Tito Schiva Istituto Sperimentale per la Floricoltura, Corso
Y-chromosomal haplogroup typing Using SBE reaction
 Schematic of multiplex PCR followed by SBE reaction Multiplex PCR Exo SAP purification SBE reaction 5 A 3 ddatp ddgtp 3 T 5 A G 3 T 5 3 5 G C 5 3 3 C 5 ddttp ddctp 5 T 3 T C 3 A 5 3 A 5 5 C 3 3 G 5 3 G
Schematic of multiplex PCR followed by SBE reaction Multiplex PCR Exo SAP purification SBE reaction 5 A 3 ddatp ddgtp 3 T 5 A G 3 T 5 3 5 G C 5 3 3 C 5 ddttp ddctp 5 T 3 T C 3 A 5 3 A 5 5 C 3 3 G 5 3 G
MacBlunt PCR Cloning Kit Manual
 MacBlunt PCR Cloning Kit Manual Shipping and Storage MacBlunt PCR Cloning Kits are shipped on dry ice. Each kit contains a box with cloning reagents and an attached bag with Eco-Blue Competent Cells (optional).
MacBlunt PCR Cloning Kit Manual Shipping and Storage MacBlunt PCR Cloning Kits are shipped on dry ice. Each kit contains a box with cloning reagents and an attached bag with Eco-Blue Competent Cells (optional).
Supporting Information
 Supporting Information Table S1. Oligonucleotide sequences used in this work Oligo DNA A B C D CpG-A CpG-B CpG-C CpG-D Sequence 5 ACA TTC CTA AGT CTG AAA CAT TAC AGC TTG CTA CAC GAG AAG AGC CGC CAT AGT
Supporting Information Table S1. Oligonucleotide sequences used in this work Oligo DNA A B C D CpG-A CpG-B CpG-C CpG-D Sequence 5 ACA TTC CTA AGT CTG AAA CAT TAC AGC TTG CTA CAC GAG AAG AGC CGC CAT AGT
Arabidopsis actin depolymerizing factor AtADF4 mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB
 Arabidopsis actin depolymerizing factor mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB Files in this Data Supplement: Supplemental Table S1 Supplemental Table
Arabidopsis actin depolymerizing factor mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB Files in this Data Supplement: Supplemental Table S1 Supplemental Table
Amplification of Human Papillomavirus DNA Sequences by Using Conserved Primers
 JOURNAL OF CLINICAL MICROBIOLOGY, Dec. 1989, p. 2660-2665 0095-1137/89/122660-06$02.00/0 Copyright 1989, American Society for Microbiology Vol. 27, No. 12 Amplification of Human Papillomavirus DNA Sequences
JOURNAL OF CLINICAL MICROBIOLOGY, Dec. 1989, p. 2660-2665 0095-1137/89/122660-06$02.00/0 Copyright 1989, American Society for Microbiology Vol. 27, No. 12 Amplification of Human Papillomavirus DNA Sequences
An engineered tryptophan zipper-type peptide as a molecular recognition scaffold
 SUPPLEMENTARY MATERIAL An engineered tryptophan zipper-type peptide as a molecular recognition scaffold Zihao Cheng and Robert E. Campbell* Supplementary Methods Library construction for FRET-based screening
SUPPLEMENTARY MATERIAL An engineered tryptophan zipper-type peptide as a molecular recognition scaffold Zihao Cheng and Robert E. Campbell* Supplementary Methods Library construction for FRET-based screening
Supporting Information. Trifluoroacetophenone-Linked Nucleotides and DNA for Studying of DNA-protein Interactions by 19 F NMR Spectroscopy
 Supporting Information Trifluoroacetophenone-Linked Nucleotides and DNA for Studying of DNA-protein Interactions by 19 F NMR Spectroscopy Agata Olszewska, Radek Pohl and Michal Hocek # * Institute of Organic
Supporting Information Trifluoroacetophenone-Linked Nucleotides and DNA for Studying of DNA-protein Interactions by 19 F NMR Spectroscopy Agata Olszewska, Radek Pohl and Michal Hocek # * Institute of Organic
Dierks Supplementary Fig. S1
 Dierks Supplementary Fig. S1 ITK SYK PH TH K42R wt K42R (kinase deficient) R29C E42K Y323F R29C E42K Y323F (reduced phospholipid binding) (enhanced phospholipid binding) (reduced Cbl binding) E42K Y323F
Dierks Supplementary Fig. S1 ITK SYK PH TH K42R wt K42R (kinase deficient) R29C E42K Y323F R29C E42K Y323F (reduced phospholipid binding) (enhanced phospholipid binding) (reduced Cbl binding) E42K Y323F
Supporting Online Information
 Supporting Online Information Isolation of Human Genomic DNA Sequences with Expanded Nucleobase Selectivity Preeti Rathi, Sara Maurer, Grzegorz Kubik and Daniel Summerer* Department of Chemistry and Chemical
Supporting Online Information Isolation of Human Genomic DNA Sequences with Expanded Nucleobase Selectivity Preeti Rathi, Sara Maurer, Grzegorz Kubik and Daniel Summerer* Department of Chemistry and Chemical
ORFs and genes. Please sit in row K or forward
 ORFs and genes Please sit in row K or forward https://www.flickr.com/photos/teseum/3231682806/in/photostream/ Question: why do some strains of Vibrio cause cholera and others don t? Methods Mechanisms
ORFs and genes Please sit in row K or forward https://www.flickr.com/photos/teseum/3231682806/in/photostream/ Question: why do some strains of Vibrio cause cholera and others don t? Methods Mechanisms
PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells
 Supplementary Information for: PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells Ju Hye Jang 1, Hyun Kim 2, Mi Jung Jang 2, Ju Hyun Cho 1,2,* 1 Research Institute
Supplementary Information for: PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells Ju Hye Jang 1, Hyun Kim 2, Mi Jung Jang 2, Ju Hyun Cho 1,2,* 1 Research Institute
Supporting Information. Copyright Wiley-VCH Verlag GmbH & Co. KGaA, Weinheim, 2006
 Supporting Information Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Supporting Information for Expanding the Genetic
Supporting Information Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Supporting Information for Expanding the Genetic
Disease and selection in the human genome 3
 Disease and selection in the human genome 3 Ka/Ks revisited Please sit in row K or forward RBFD: human populations, adaptation and immunity Neandertal Museum, Mettman Germany Sequence genome Measure expression
Disease and selection in the human genome 3 Ka/Ks revisited Please sit in row K or forward RBFD: human populations, adaptation and immunity Neandertal Museum, Mettman Germany Sequence genome Measure expression
strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular
![strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular](/thumbs/72/66361212.jpg) Additional file 2 Identification of AOX1 in P. pastoris GS115 with a Mut s phenotype Results and Discussion The HBsAg producing strain was originally identified as a Mut s (methanol utilization slow) strain
Additional file 2 Identification of AOX1 in P. pastoris GS115 with a Mut s phenotype Results and Discussion The HBsAg producing strain was originally identified as a Mut s (methanol utilization slow) strain
hcd1tg/hj1tg/ ApoE-/- hcd1tg/hj1tg/ ApoE+/+
 ApoE+/+ ApoE-/- ApoE-/- H&E (1x) Supplementary Figure 1. No obvious pathology is observed in the colon of diseased ApoE-/me. Colon samples were fixed in 1% formalin and laid out in Swiss rolls for paraffin
ApoE+/+ ApoE-/- ApoE-/- H&E (1x) Supplementary Figure 1. No obvious pathology is observed in the colon of diseased ApoE-/me. Colon samples were fixed in 1% formalin and laid out in Swiss rolls for paraffin
Supplemental Data. mir156-regulated SPL Transcription. Factors Define an Endogenous Flowering. Pathway in Arabidopsis thaliana
 Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
Engineering D66N mutant using quick change site directed mutagenesis. Harkewal Singh 09/01/2010
 Engineering D66N mutant using quick change site directed mutagenesis Harkewal Singh 09/01/2010 1 1- What is quick change site directed mutagenesis? 2- An overview of the kit contents. 3- A brief information
Engineering D66N mutant using quick change site directed mutagenesis Harkewal Singh 09/01/2010 1 1- What is quick change site directed mutagenesis? 2- An overview of the kit contents. 3- A brief information
Supplementary Figures
 Supplementary Figures Supplementary Fig. 1 Characterization of GSCs. a. Immunostaining of primary GSC spheres from GSC lines. Nestin (neural progenitor marker, red), TLX (green). Merged images of nestin,
Supplementary Figures Supplementary Fig. 1 Characterization of GSCs. a. Immunostaining of primary GSC spheres from GSC lines. Nestin (neural progenitor marker, red), TLX (green). Merged images of nestin,
Supplemental Table 1. Mutant ADAMTS3 alleles detected in HEK293T clone 4C2. WT CCTGTCACTTTGGTTGATAGC MVLLSLWLIAAALVEVR
 Supplemental Dataset Supplemental Table 1. Mutant ADAMTS3 alleles detected in HEK293T clone 4C2. DNA sequence Amino acid sequence WT CCTGTCACTTTGGTTGATAGC MVLLSLWLIAAALVEVR Allele 1 CCTGTC------------------GATAGC
Supplemental Dataset Supplemental Table 1. Mutant ADAMTS3 alleles detected in HEK293T clone 4C2. DNA sequence Amino acid sequence WT CCTGTCACTTTGGTTGATAGC MVLLSLWLIAAALVEVR Allele 1 CCTGTC------------------GATAGC
Gene synthesis by circular assembly amplification
 Gene synthesis by circular assembly amplification Duhee Bang & George M Church Supplementary figures and text: Supplementary Figure 1. Dpo4 gene (1.05kb) construction by various methods. Supplementary
Gene synthesis by circular assembly amplification Duhee Bang & George M Church Supplementary figures and text: Supplementary Figure 1. Dpo4 gene (1.05kb) construction by various methods. Supplementary
Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were
 1 Supplemental methods 2 3 4 5 6 7 8 9 1 11 12 13 14 15 16 17 18 19 21 22 23 Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were monitored by quantitative reverse-transcription
1 Supplemental methods 2 3 4 5 6 7 8 9 1 11 12 13 14 15 16 17 18 19 21 22 23 Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were monitored by quantitative reverse-transcription
RPA-AB RPA-C Supplemental Figure S1: SDS-PAGE stained with Coomassie Blue after protein purification.
 RPA-AB RPA-C (a) (b) (c) (d) (e) (f) Supplemental Figure S: SDS-PAGE stained with Coomassie Blue after protein purification. (a) RPA; (b) RPA-AB; (c) RPA-CDE; (d) RPA-CDE core; (e) RPA-DE; and (f) RPA-C
RPA-AB RPA-C (a) (b) (c) (d) (e) (f) Supplemental Figure S: SDS-PAGE stained with Coomassie Blue after protein purification. (a) RPA; (b) RPA-AB; (c) RPA-CDE; (d) RPA-CDE core; (e) RPA-DE; and (f) RPA-C
Sequencing of DNA lesions facilitated by site-specific excision via base. excision repair DNA glycosylases yielding ligatable gaps
 Supporting information Sequencing of DNA lesions facilitated by site-specific excision via base excision repair DNA glycosylases yielding ligatable gaps Jan Riedl, Aaron M. Fleming, and Cynthia J. Burrows*
Supporting information Sequencing of DNA lesions facilitated by site-specific excision via base excision repair DNA glycosylases yielding ligatable gaps Jan Riedl, Aaron M. Fleming, and Cynthia J. Burrows*
Materials Protein synthesis kit. This kit consists of 24 amino acids, 24 transfer RNAs, four messenger RNAs and one ribosome (see below).
 Protein Synthesis Instructions The purpose of today s lab is to: Understand how a cell manufactures proteins from amino acids, using information stored in the genetic code. Assemble models of four very
Protein Synthesis Instructions The purpose of today s lab is to: Understand how a cell manufactures proteins from amino acids, using information stored in the genetic code. Assemble models of four very
ΔPDD1 x ΔPDD1. ΔPDD1 x wild type. 70 kd Pdd1. Pdd3
 Supplemental Fig. S1 ΔPDD1 x wild type ΔPDD1 x ΔPDD1 70 kd Pdd1 50 kd 37 kd Pdd3 Supplemental Fig. S1. ΔPDD1 strains express no detectable Pdd1 protein. Western blot analysis of whole-protein extracts
Supplemental Fig. S1 ΔPDD1 x wild type ΔPDD1 x ΔPDD1 70 kd Pdd1 50 kd 37 kd Pdd3 Supplemental Fig. S1. ΔPDD1 strains express no detectable Pdd1 protein. Western blot analysis of whole-protein extracts
II 0.95 DM2 (RPP1) DM3 (At3g61540) b
 Table S2. F 2 Segregation Ratios at 16 C, Related to Figure 2 Cross n c Phenotype Model e 2 Locus A Locus B Normal F 1 -like Enhanced d Uk-1/Uk-3 149 64 36 49 DM2 (RPP1) DM1 (SSI4) a Bla-1/Hh-0 F 3 111
Table S2. F 2 Segregation Ratios at 16 C, Related to Figure 2 Cross n c Phenotype Model e 2 Locus A Locus B Normal F 1 -like Enhanced d Uk-1/Uk-3 149 64 36 49 DM2 (RPP1) DM1 (SSI4) a Bla-1/Hh-0 F 3 111
Overexpression Normal expression Overexpression Normal expression. 26 (21.1%) N (%) P-value a N (%)
 SUPPLEMENTARY TABLES Table S1. Alteration of ZNF322A protein expression levels in relation to clinicopathological parameters in 123 Asian and 74 Caucasian lung cancer patients. Asian patients Caucasian
SUPPLEMENTARY TABLES Table S1. Alteration of ZNF322A protein expression levels in relation to clinicopathological parameters in 123 Asian and 74 Caucasian lung cancer patients. Asian patients Caucasian
Multiplexing Genome-scale Engineering
 Multiplexing Genome-scale Engineering Harris Wang, Ph.D. Department of Systems Biology Department of Pathology & Cell Biology http://wanglab.c2b2.columbia.edu Rise of Genomics An Expanding Toolbox Esvelt
Multiplexing Genome-scale Engineering Harris Wang, Ph.D. Department of Systems Biology Department of Pathology & Cell Biology http://wanglab.c2b2.columbia.edu Rise of Genomics An Expanding Toolbox Esvelt
Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system
 Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system Dr. Tim Welsink Molecular Biology Transient Gene Expression OUTLINE Short
Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system Dr. Tim Welsink Molecular Biology Transient Gene Expression OUTLINE Short
Project 07/111 Final Report October 31, Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines
 Project 07/111 Final Report October 31, 2007. Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines Project Leader: Dr Douglas C. Hodgins (519-824-4120 Ex 54758, fax 519-824-5930)
Project 07/111 Final Report October 31, 2007. Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines Project Leader: Dr Douglas C. Hodgins (519-824-4120 Ex 54758, fax 519-824-5930)
I. Things to keep in mind before you start this protocol
 Inverse PCR and Sequencing Protocol on 5 Fly Preps For recovery of sequences flanking piggybac elements This protocol is an adaptation of "Inverse PCR and Cycle Sequencing Protocols" by E. Jay Rehm Berkeley
Inverse PCR and Sequencing Protocol on 5 Fly Preps For recovery of sequences flanking piggybac elements This protocol is an adaptation of "Inverse PCR and Cycle Sequencing Protocols" by E. Jay Rehm Berkeley
Table S1. Bacterial strains (Related to Results and Experimental Procedures)
 Table S1. Bacterial strains (Related to Results and Experimental Procedures) Strain number Relevant genotype Source or reference 1045 AB1157 Graham Walker (Donnelly and Walker, 1989) 2458 3084 (MG1655)
Table S1. Bacterial strains (Related to Results and Experimental Procedures) Strain number Relevant genotype Source or reference 1045 AB1157 Graham Walker (Donnelly and Walker, 1989) 2458 3084 (MG1655)
SUPPLEMENTARY MATERIALS AND METHODS. E. coli strains, plasmids, and growth conditions. Escherichia coli strain P90C (1)
 SUPPLEMENTARY MATERIALS AND METHODS E. coli strains, plasmids, and growth conditions. Escherichia coli strain P90C (1) dinb::kan (lab stock) derivative was used as wild-type. MG1655 alka tag dinb (2) is
SUPPLEMENTARY MATERIALS AND METHODS E. coli strains, plasmids, and growth conditions. Escherichia coli strain P90C (1) dinb::kan (lab stock) derivative was used as wild-type. MG1655 alka tag dinb (2) is
BioInformatics and Computational Molecular Biology. Course Website
 BioInformatics and Computational Molecular Biology Course Website http://bioinformatics.uchc.edu What is Bioinformatics Bioinformatics upgrades the information content of biological measurements. Discovery
BioInformatics and Computational Molecular Biology Course Website http://bioinformatics.uchc.edu What is Bioinformatics Bioinformatics upgrades the information content of biological measurements. Discovery
Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of
 Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of the cell line) were immunostained for HA, acetylated
Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of the cell line) were immunostained for HA, acetylated
2ml of 1M stock 10x TBE (1 Litre) Tris Base 107.8g 55g (harmful, wear mask) EDTA 7.4g
 Phytoplasma Detection Protocol Buffers: Hybridisation buffer 100ml hybridisation buffer 2.92g Sodium chloride 4g Blocking reagent (add slowly while stirring) Mix at room temperature for 2 hours Can be
Phytoplasma Detection Protocol Buffers: Hybridisation buffer 100ml hybridisation buffer 2.92g Sodium chloride 4g Blocking reagent (add slowly while stirring) Mix at room temperature for 2 hours Can be
Sexing Bovine Preimplantation Embryos by PCR
 Sexing Bovine Preimplantation Embryos by PCR Katherine E.M. Hendricks 1, Leydson F. Martins 2, Justin M. Fear 1 and Peter J. Hansen 1 1 Dept. of Animal Sciences, University of Florida and Department of
Sexing Bovine Preimplantation Embryos by PCR Katherine E.M. Hendricks 1, Leydson F. Martins 2, Justin M. Fear 1 and Peter J. Hansen 1 1 Dept. of Animal Sciences, University of Florida and Department of
A netlike rolling circle nucleic acid amplification technique
 Electronic Supplementary Material (ESI) for Analyst. This journal is The Royal Society of Chemistry 2014 Supplementary Information A netlike rolling circle nucleic acid amplification technique Xiaoli Zhu,
Electronic Supplementary Material (ESI) for Analyst. This journal is The Royal Society of Chemistry 2014 Supplementary Information A netlike rolling circle nucleic acid amplification technique Xiaoli Zhu,
SUPPORTING INFORMATION
 SUPPORTING INFORMATION Investigation of the Biosynthesis of the Lasso Peptide Chaxapeptin Using an E. coli-based Production System Helena Martin-Gómez, Uwe Linne, Fernando Albericio, Judit Tulla-Puche,*
SUPPORTING INFORMATION Investigation of the Biosynthesis of the Lasso Peptide Chaxapeptin Using an E. coli-based Production System Helena Martin-Gómez, Uwe Linne, Fernando Albericio, Judit Tulla-Puche,*
NESTED Sequence-based Typing (SBT) protocol for epidemiological typing of Legionella pneumophila directly from clinical samples
 NESTED Sequence-based Typing (SBT) protocol for epidemiological typing of Legionella pneumophila directly from clinical samples VERSION 2.0 SUMMARY This procedure describes the use of nested Sequence-Based
NESTED Sequence-based Typing (SBT) protocol for epidemiological typing of Legionella pneumophila directly from clinical samples VERSION 2.0 SUMMARY This procedure describes the use of nested Sequence-Based
NAME:... MODEL ANSWER... STUDENT NUMBER:... Maximum marks: 50. Internal Examiner: Hugh Murrell, Computer Science, UKZN
 COMP710, Bioinformatics with Julia, Test One, Thursday the 20 th of April, 2017, 09h30-11h30 1 NAME:...... MODEL ANSWER... STUDENT NUMBER:...... Maximum marks: 50 Internal Examiner: Hugh Murrell, Computer
COMP710, Bioinformatics with Julia, Test One, Thursday the 20 th of April, 2017, 09h30-11h30 1 NAME:...... MODEL ANSWER... STUDENT NUMBER:...... Maximum marks: 50 Internal Examiner: Hugh Murrell, Computer
COMPARISON OF DETECTION THRESHOLD OF DIFFERENT PASTEURELLA MULTOCIDA SPECIFIC PCRs
 Indian J. Anim. Res., 46 (1) : 28-33, 2012 AGRICULTURAL RESEARCH COMMUNICATION CENTRE www.arccjournals.com / indianjournals.com COMPARISON OF DETECTION THRESHOLD OF DIFFERENT PASTEURELLA MULTOCIDA SPECIFIC
Indian J. Anim. Res., 46 (1) : 28-33, 2012 AGRICULTURAL RESEARCH COMMUNICATION CENTRE www.arccjournals.com / indianjournals.com COMPARISON OF DETECTION THRESHOLD OF DIFFERENT PASTEURELLA MULTOCIDA SPECIFIC
Supporting Information
 Supporting Information Barderas et al. 10.1073/pnas.0801221105 SI Text: Docking of gastrin to Constructed scfv Models Interactive predocking of the 4-WL-5 motif into the central pocket observed in the
Supporting Information Barderas et al. 10.1073/pnas.0801221105 SI Text: Docking of gastrin to Constructed scfv Models Interactive predocking of the 4-WL-5 motif into the central pocket observed in the
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/10/494/eaan6284/dc1 Supplementary Materials for Activation of master virulence regulator PhoP in acidic ph requires the Salmonella-specific protein UgtL Jeongjoon
www.sciencesignaling.org/cgi/content/full/10/494/eaan6284/dc1 Supplementary Materials for Activation of master virulence regulator PhoP in acidic ph requires the Salmonella-specific protein UgtL Jeongjoon
evaluated with UAS CLB eliciting UAS CIT -N Libraries increase in the
 Supplementary Figures Supplementary Figure 1: Promoter scaffold library assemblies. Many ensembless of libraries were evaluated in this work. As a legend, the box outline color in top half of the figure
Supplementary Figures Supplementary Figure 1: Promoter scaffold library assemblies. Many ensembless of libraries were evaluated in this work. As a legend, the box outline color in top half of the figure
Lecture 11: Gene Prediction
 Lecture 11: Gene Prediction Study Chapter 6.11-6.14 1 Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and end positions of genes in a genome Where are
Lecture 11: Gene Prediction Study Chapter 6.11-6.14 1 Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and end positions of genes in a genome Where are
Supplementary Figure 1. Analysis of replication rates by Pol. Nature Structural & Molecular Biology: doi: /nsmb.3207
 Supplementary Figure 1 Analysis of replication rates by Pol (a) Electrophoretic Mobility Shift Assay (EMSA) of Pol -DV binding to template-primer DNA (Fried, M. & Crothers, D.M. Equilibria and kinetics
Supplementary Figure 1 Analysis of replication rates by Pol (a) Electrophoretic Mobility Shift Assay (EMSA) of Pol -DV binding to template-primer DNA (Fried, M. & Crothers, D.M. Equilibria and kinetics
11th Meeting of the Science Working Group. Lima, Peru, October 2012 SWG-11-JM-11
 11th Meeting of the Science Working Group Lima, Peru, 15-19 October 2012 Russian population genetics studies of jack mackerel in the South Pacific P.K.Afanasiev M.A.Rabchun A.I.Glubokov Introduction. In
11th Meeting of the Science Working Group Lima, Peru, 15-19 October 2012 Russian population genetics studies of jack mackerel in the South Pacific P.K.Afanasiev M.A.Rabchun A.I.Glubokov Introduction. In
Genomics and Gene Recognition Genes and Blue Genes
 Genomics and Gene Recognition Genes and Blue Genes November 1, 2004 Prokaryotic Gene Structure prokaryotes are simplest free-living organisms studying prokaryotes can give us a sense what is the minimum
Genomics and Gene Recognition Genes and Blue Genes November 1, 2004 Prokaryotic Gene Structure prokaryotes are simplest free-living organisms studying prokaryotes can give us a sense what is the minimum
SAY IT WITH DNA: Protein Synthesis Activity by Larry Flammer
 TEACHER S GUIDE SAY IT WITH DNA: Protein Synthesis Activity by Larry Flammer SYNOPSIS This activity uses the metaphor of decoding a secret message for the Protein Synthesis process. Students teach themselves
TEACHER S GUIDE SAY IT WITH DNA: Protein Synthesis Activity by Larry Flammer SYNOPSIS This activity uses the metaphor of decoding a secret message for the Protein Synthesis process. Students teach themselves
Anti-Pim-1 (Cat#3247), anti-met (Cat#3127), anti-ron (Cat#2654), Anti-EGFR
 Supplementary Methods Antibodies Anti-Pim-1 (Cat#3247), anti-met (Cat#3127), anti-ron (Cat#2654), Anti-EGFR (Cat#2646), anti-igf1r (Cat#3018), anti-insr (Cat#3020), anti-akt (pan, Cat#4691), anti-phospho-akt
Supplementary Methods Antibodies Anti-Pim-1 (Cat#3247), anti-met (Cat#3127), anti-ron (Cat#2654), Anti-EGFR (Cat#2646), anti-igf1r (Cat#3018), anti-insr (Cat#3020), anti-akt (pan, Cat#4691), anti-phospho-akt
Primer Design Workshop. École d'été en géné-que des champignons 2012 Dr. Will Hintz University of Victoria
 Primer Design Workshop École d'été en géné-que des champignons 2012 Dr. Will Hintz University of Victoria Scenario You have discovered the presence of a novel endophy5c organism living inside the cells
Primer Design Workshop École d'été en géné-que des champignons 2012 Dr. Will Hintz University of Victoria Scenario You have discovered the presence of a novel endophy5c organism living inside the cells
G+C content. 1 Introduction. 2 Chromosomes Topology & Counts. 3 Genome size. 4 Replichores and gene orientation. 5 Chirochores.
 1 Introduction 2 Chromosomes Topology & Counts 3 Genome size 4 Replichores and gene orientation 5 Chirochores 6 7 Codon usage 121 marc.bailly-bechet@univ-lyon1.fr Bacterial genome structures Introduction
1 Introduction 2 Chromosomes Topology & Counts 3 Genome size 4 Replichores and gene orientation 5 Chirochores 6 7 Codon usage 121 marc.bailly-bechet@univ-lyon1.fr Bacterial genome structures Introduction
Supplementary Methods Quantitative RT-PCR. For mrna, total RNA was prepared using TRIzol reagent (Invitrogen) and genomic DNA was eliminated with TURB
 Supplementary Methods Quantitative RT-PCR. For mrna, total RNA was prepared using TRIzol reagent (Invitrogen) and genomic DNA was eliminated with TURBO DNA-free Kit (Ambion). One µg of total RNA was reverse
Supplementary Methods Quantitative RT-PCR. For mrna, total RNA was prepared using TRIzol reagent (Invitrogen) and genomic DNA was eliminated with TURBO DNA-free Kit (Ambion). One µg of total RNA was reverse
Supplementary Information. Construction of Lasso Peptide Fusion Proteins
 Supplementary Information Construction of Lasso Peptide Fusion Proteins Chuhan Zong 1, Mikhail O. Maksimov 2, A. James Link 2,3 * Departments of 1 Chemistry, 2 Chemical and Biological Engineering, and
Supplementary Information Construction of Lasso Peptide Fusion Proteins Chuhan Zong 1, Mikhail O. Maksimov 2, A. James Link 2,3 * Departments of 1 Chemistry, 2 Chemical and Biological Engineering, and
Det matematisk-naturvitenskapelige fakultet
 UNIVERSITETET I OSLO Det matematisk-naturvitenskapelige fakultet Exam in: MBV4010 Arbeidsmetoder i molekylærbiologi og biokjemi I MBV4010 Methods in molecular biology and biochemistry I Day of exam: Friday
UNIVERSITETET I OSLO Det matematisk-naturvitenskapelige fakultet Exam in: MBV4010 Arbeidsmetoder i molekylærbiologi og biokjemi I MBV4010 Methods in molecular biology and biochemistry I Day of exam: Friday
Supporting Information
 Supporting Information Wiley-VCH 2007 69451 Weinheim, Germany Site-Specific Control of Distances between Gold Nanoparticles using Phosphorothioate Anchors on DNA and a Short Bifunctional Molecular Fastener
Supporting Information Wiley-VCH 2007 69451 Weinheim, Germany Site-Specific Control of Distances between Gold Nanoparticles using Phosphorothioate Anchors on DNA and a Short Bifunctional Molecular Fastener
Supplemental Data. Bennett et al. (2010). Plant Cell /tpc
 BRN1 ---------MSSSNGGVPPGFRFHPTDEELLHYYLKKKISYEKFEMEVIKEVDLNKIEPWDLQDRCKIGSTPQNEWYFFSHKDRKYPTGS 81 BRN2 --------MGSSSNGGVPPGFRFHPTDEELLHYYLKKKISYQKFEMEVIREVDLNKLEPWDLQERCKIGSTPQNEWYFFSHKDRKYPTGS 82 SMB
BRN1 ---------MSSSNGGVPPGFRFHPTDEELLHYYLKKKISYEKFEMEVIKEVDLNKIEPWDLQDRCKIGSTPQNEWYFFSHKDRKYPTGS 81 BRN2 --------MGSSSNGGVPPGFRFHPTDEELLHYYLKKKISYQKFEMEVIREVDLNKLEPWDLQERCKIGSTPQNEWYFFSHKDRKYPTGS 82 SMB
2
 1 2 3 4 5 6 7 Supplemental Table 1. Magnaporthe oryzae strains generated in this study. Strain background Genotype Strain name Description Guy-11 H1:RFP H1:RFP Strain expressing Histone H1- encoding gene
1 2 3 4 5 6 7 Supplemental Table 1. Magnaporthe oryzae strains generated in this study. Strain background Genotype Strain name Description Guy-11 H1:RFP H1:RFP Strain expressing Histone H1- encoding gene
SUPPLEMENTAL MATERIAL GENOTYPING WITH MULTIPLEXING TARGETED RESEQUENCING
 SUPPLEMENTAL MATERIAL GENOTYPING WITH MULTIPLEXING TARGETED RESEQUENCING All of the patients and control subjects were sequenced and genotyped in the same way. Shotgun libraries of approximately 250 bp
SUPPLEMENTAL MATERIAL GENOTYPING WITH MULTIPLEXING TARGETED RESEQUENCING All of the patients and control subjects were sequenced and genotyped in the same way. Shotgun libraries of approximately 250 bp
SUPPORTING INFORMATION FILE
 Intrinsic and extrinsic connections of Tet3 dioxygenase with CXXC zinc finger modules Nan Liu, Mengxi Wang, Wen Deng, Christine S. Schmidt, Weihua Qin, Heinrich Leonhardt and Fabio Spada Department of
Intrinsic and extrinsic connections of Tet3 dioxygenase with CXXC zinc finger modules Nan Liu, Mengxi Wang, Wen Deng, Christine S. Schmidt, Weihua Qin, Heinrich Leonhardt and Fabio Spada Department of
CHAPTER 3 MATERIALS AND METHODS
 CHAPTER 3 MATERIALS AND METHODS CHAPTER THREE MATERIALS AND METHODS 3.1 MATERIALS 3.1.1 CHEMICALS* Chemical Name Absolute ethanol Acetic Acid Agarose Ammonium acetate Betaine Bromophenol Blue Bovine Serum
CHAPTER 3 MATERIALS AND METHODS CHAPTER THREE MATERIALS AND METHODS 3.1 MATERIALS 3.1.1 CHEMICALS* Chemical Name Absolute ethanol Acetic Acid Agarose Ammonium acetate Betaine Bromophenol Blue Bovine Serum
SUPPLEMENTARY INFORMATION
 doi:1.138/nature11496 Cl. 8 Cl. E93 Rag1 -/- 3H9 + BM Rag1 -/- BM CD CD c-kit c-kit c-kit wt Spleen c-kit B22 B22 IgM IgM IgM Supplementary Figure 1. FACS analysis of single-cell-derived pre-b cell clones.
doi:1.138/nature11496 Cl. 8 Cl. E93 Rag1 -/- 3H9 + BM Rag1 -/- BM CD CD c-kit c-kit c-kit wt Spleen c-kit B22 B22 IgM IgM IgM Supplementary Figure 1. FACS analysis of single-cell-derived pre-b cell clones.
Table of Contents. Description Kit Components Reagents not supplied in the kit Equipment required Storage...
 Table of Contents Description... 2 Kit Components... 2 Reagents not supplied in the kit... 2 Equipment required... 2 Storage... 2 References... 2 Principle... 3 Protocol... 4 Identification of HPV types...
Table of Contents Description... 2 Kit Components... 2 Reagents not supplied in the kit... 2 Equipment required... 2 Storage... 2 References... 2 Principle... 3 Protocol... 4 Identification of HPV types...
S4B fluorescence (AU)
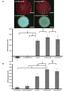 A S4B fluorescence (AU) S4B fluorescence (AU) dsbb csgba csgd dsbb csgba bcsa 5000 * NS NS 4000 * 3000 2000 1000 0 ΔcsgBAΔbcsA ΔcsgDΔdsbBΔbcsA ΔcsgBA ΔdsbBΔcsgBA ΔcsgDΔdsbB B -1000 4000 * * NS 3500 * 3000
A S4B fluorescence (AU) S4B fluorescence (AU) dsbb csgba csgd dsbb csgba bcsa 5000 * NS NS 4000 * 3000 2000 1000 0 ΔcsgBAΔbcsA ΔcsgDΔdsbBΔbcsA ΔcsgBA ΔdsbBΔcsgBA ΔcsgDΔdsbB B -1000 4000 * * NS 3500 * 3000
Search for and Analysis of Single Nucleotide Polymorphisms (SNPs) in Rice (Oryza sativa, Oryza rufipogon) and Establishment of SNP Markers
 DNA Research 9, 163 171 (2002) Search for and Analysis of Single Nucleotide Polymorphisms (SNPs) in Rice (Oryza sativa, Oryza rufipogon) and Establishment of SNP Markers Shinobu Nasu, Junko Suzuki, Rieko
DNA Research 9, 163 171 (2002) Search for and Analysis of Single Nucleotide Polymorphisms (SNPs) in Rice (Oryza sativa, Oryza rufipogon) and Establishment of SNP Markers Shinobu Nasu, Junko Suzuki, Rieko
TILLING Resources for Japonica and Indica Rice
 TILLING Resources for Japonica and Indica Rice Luca Comai, Steve Henikoff, Tom Tai, Hei Leung TILLING Resources for Japonica and Indica Rice Outline Who: Seattle TILLING Project (Luca Comai, Steve Henikoff)
TILLING Resources for Japonica and Indica Rice Luca Comai, Steve Henikoff, Tom Tai, Hei Leung TILLING Resources for Japonica and Indica Rice Outline Who: Seattle TILLING Project (Luca Comai, Steve Henikoff)
Lecture 19A. DNA computing
 Lecture 19A. DNA computing What exactly is DNA (deoxyribonucleic acid)? DNA is the material that contains codes for the many physical characteristics of every living creature. Your cells use different
Lecture 19A. DNA computing What exactly is DNA (deoxyribonucleic acid)? DNA is the material that contains codes for the many physical characteristics of every living creature. Your cells use different
Lewis x/cd15 expression in human myeloid cell differentiation. is regulated by sialidase activity
 Lewis x/cd15 expression in human myeloid cell differentiation is regulated by sialidase activity Samah Zeineb Gadhoum 1, 2 and Robert Sackstein* 1, 2, 3, 4 From the Departments of Dermatology 1 and Medicine
Lewis x/cd15 expression in human myeloid cell differentiation is regulated by sialidase activity Samah Zeineb Gadhoum 1, 2 and Robert Sackstein* 1, 2, 3, 4 From the Departments of Dermatology 1 and Medicine
Homework. A bit about the nature of the atoms of interest. Project. The role of electronega<vity
 Homework Why cited articles are especially useful. citeulike science citation index When cutting and pasting less is more. Project Your protein: I will mail these out this weekend If you haven t gotten
Homework Why cited articles are especially useful. citeulike science citation index When cutting and pasting less is more. Project Your protein: I will mail these out this weekend If you haven t gotten
Nongenetic Reprogramming of the Ligand Specificity. of Growth Factor Receptors by Bispecific DNA Aptamers
 Supporting Information For Nongenetic Reprogramming of the Ligand Specificity of Growth Factor Receptors by Bispecific DNA Aptamers Ryosuke Ueki,* Saki Atsuta, Ayaka Ueki and Shinsuke Sando* Department
Supporting Information For Nongenetic Reprogramming of the Ligand Specificity of Growth Factor Receptors by Bispecific DNA Aptamers Ryosuke Ueki,* Saki Atsuta, Ayaka Ueki and Shinsuke Sando* Department
NOTES Cloning and Partial DNA Sequencing of Two New Human Papillomavirus Types Associated with Condylomas and Low-Grade
 JOURNAL OF VROLOGY. June 1989. p. 2829-2834 002'-538X/89/062829-06$02.00/0 Copyright C 1989. American Society for Microbiology Vol. 63, No. 6 NOTES Cloning and Partial DNA Sequencing of Two New Human Papillomavirus
JOURNAL OF VROLOGY. June 1989. p. 2829-2834 002'-538X/89/062829-06$02.00/0 Copyright C 1989. American Society for Microbiology Vol. 63, No. 6 NOTES Cloning and Partial DNA Sequencing of Two New Human Papillomavirus
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Blanken MO, Rovers MM, Molenaar JM, et al. Respiratory syncytial
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Blanken MO, Rovers MM, Molenaar JM, et al. Respiratory syncytial
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2014 Electronic Supplementary Information Multiplexed Detection of Lung Cancer Cells at the Single-Molecule
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2014 Electronic Supplementary Information Multiplexed Detection of Lung Cancer Cells at the Single-Molecule
A green method of staining DNA in polyacrylamide gel electrophoresis based on fluorescent copper nanoclusters synthesized in situ
 Electronic Supplementary Material A green method of staining DNA in polyacrylamide gel electrophoresis based on fluorescent copper nanoclusters synthesized in situ Xiaoli Zhu 1, Hai Shi 1, Yalan Shen 1,
Electronic Supplementary Material A green method of staining DNA in polyacrylamide gel electrophoresis based on fluorescent copper nanoclusters synthesized in situ Xiaoli Zhu 1, Hai Shi 1, Yalan Shen 1,
Event-specific Method for the Quantification of Soybean Event DP Using Real-time PCR. Validated Method
 Event-specific Method for the Quantification of Soybean Event DP-356043-5 Using Real-time PCR Validated Method 29 March 2010 Joint Research Centre Institute for Health and Consumer Protection Molecular
Event-specific Method for the Quantification of Soybean Event DP-356043-5 Using Real-time PCR Validated Method 29 March 2010 Joint Research Centre Institute for Health and Consumer Protection Molecular
Electronic Supplementary Information
 Electronic Supplementary Information Ultrasensitive and Selective DNA Detection Based on Nicking Endonuclease Assisted Signal Amplification and Its Application in Cancer Cells Detection Sai Bi, Jilei Zhang,
Electronic Supplementary Information Ultrasensitive and Selective DNA Detection Based on Nicking Endonuclease Assisted Signal Amplification and Its Application in Cancer Cells Detection Sai Bi, Jilei Zhang,
SUPPLEMENTAL TABLE S1. Additional descriptions of plasmid constructions and the oligonucleotides used Plasmid or Oligonucleotide
 SUPPLEMENTAL TABLE S1. Additional descriptions of plasmid constructions and the oligonucleotides used Plasmid or Oligonucleotide former/ working Description a designation Plasmids pes213a b pes213-tn5
SUPPLEMENTAL TABLE S1. Additional descriptions of plasmid constructions and the oligonucleotides used Plasmid or Oligonucleotide former/ working Description a designation Plasmids pes213a b pes213-tn5
Analysis of human papillomavirus type 16 E6-E7 transcription in cervical carcinomas and normal cervical epithelium using the polymerase chain reaction
 Journal of General Virology (0),,. Printed in Great Britain Analysis of human papillomavirus type E-E transcription in cervical carcinomas and normal cervical epithelium using the polymerase chain reaction
Journal of General Virology (0),,. Printed in Great Britain Analysis of human papillomavirus type E-E transcription in cervical carcinomas and normal cervical epithelium using the polymerase chain reaction
SUPPLEMENTARY INFORMATION
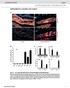 doi: 10.1038/nature07182 SUPPLEMENTAL FIGURES AND TABLES Fig. S1. myf5-expressing cells give rise to brown fat depots and skeletal muscle (a) Perirenal BAT from control (cre negative) and myf5-cre:r26r3-yfp
doi: 10.1038/nature07182 SUPPLEMENTAL FIGURES AND TABLES Fig. S1. myf5-expressing cells give rise to brown fat depots and skeletal muscle (a) Perirenal BAT from control (cre negative) and myf5-cre:r26r3-yfp
High-throughput cloning and expression in recalcitrant bacteria
 High-throughput cloning and expression in recalcitrant bacteria Eric R Geertsma & Bert Poolman Supplementary text and figures: Supplementary Figure 1 Frequency of SfiI sites yielding identical 3 extensions
High-throughput cloning and expression in recalcitrant bacteria Eric R Geertsma & Bert Poolman Supplementary text and figures: Supplementary Figure 1 Frequency of SfiI sites yielding identical 3 extensions
Polymerase Chain Reaction (PCR)
 Laboratory for Environmental Pathogens Research Department of Environmental Sciences University of Toledo Polymerase Chain Reaction (PCR) Background information The polymerase chain reaction (PCR) is an
Laboratory for Environmental Pathogens Research Department of Environmental Sciences University of Toledo Polymerase Chain Reaction (PCR) Background information The polymerase chain reaction (PCR) is an
