Proteomics and some of its Mass Spectrometric Applications
|
|
|
- Mark Bernard Conley
- 5 years ago
- Views:
Transcription
1 Proteomics and some of its Mass Spectrometric Applications What? Large scale screening of proteins, their expression, modifications and interactions by using high-throughput approaches 2 1
2 Why? The number of found genes (ORFs) in human genome is much less than the number of expressed proteins, and therefore, more information is needed about the working units of the cells From Garrett & Grisham, Biochemistry, 1999, Saunders College Publishing, USA 3 How? Proteomics branches : Proteomic analysis (analytical protein chemistry) Characterization of proteins and their post-translational modifications Expression proteomics (differential display proteomics) Profiling of expressed proteins using quantitative methods Cell-mapping proteomics (cataloging of protein-protein interactions) Identification of protein complexes Used approaches: Gel-based proteomics Mass spectrometry driven proteomics Protein arrays, Yeast two-hybrid arrays,... etc. 4 2
3 Proteomics approaches From Pevsner, 2003, p.248 Approaches to high-throughput protein analysis 5 Gel-based proteomics Older approach to screen the protein expression at the large scale The typical flow of gel-based proteomics (2DE & MS) Sample preparation First-dimension isoelectric focusing (IEF) Second-dimension SDS-PAGE Visualization & evaluation Expression analysis Protein identification by MS 6 3
4 Two-dimensional gel electrophoresis (2DE) ph MW st dimension separation based on the pi of proteins 2nd dimension separation based on the molecular weight of proteins Several visualization/detection possibilities 7 Quantitative proteomics - 2DE & MS From Simpson, 2002, p.4 General outflow of DIGE procedure (a control, b cancer) 8 4
5 An example of a quantitative expression analysis (2DE) Master gel, 14 controls, 14 cases, protein shown 6-fold upregulated t-test p=2.5* DE - Highlights and pitfalls Highlights Resolving capasity of hundreds of proteins at the same time (Sample fractionation needed for analyzing proteome more completely) Possibility to identify post-translational modifications (based on pi and MW) Pitfalls Laborous (time-consuming) and technically challenging Technical limitations e.g. staining method 10 5
6 Protein arrays Basic idea similar as in cdna arrays ; Substrate (protein, antibody etc) is bound on the surface of the array The sample is introduced to array binding Detection and analysis Used for several purposes Screening/profiling Protein - protein interactions Protein - small molecule interactions Kinase - substrate interactions etc. 11 Protein arrays cont. Both figures from Simpson, 2002 The use of protein arrays to identify enzyme - substrate interactions; a kinase modifies a substrate by transferring of a phosphate group to the substrate Antibody arrays: proteins are bound to antibodies and binding is detected 12 6
7 Protein arrays cont. New screening technique - not much knowledge yet Similarity to cdna arrays Technique exists Problems e.g. Specificity of substrates, also some proteins are more sticky than others Background variations Normalization among experiments 13 Yeast two-hybrid system Used to identify protein-protein interactions Systematic testing of binding candidates Used firstly with yeast (S. cerevisiae the whole genome know), but nowdays applied also to other organisms 14 7
8 Yeast two-hybrid cont. From Simpson 2002 Bait (i.e. a protein whose interacting proteins are searched) is bound to the DNA-binding domain of a reporter gene. Bait alone isn t capable to turn on the transcription of a reporter gene. Prey (candidate interacting protein) is bound to the activation region of reporter gene. Also prey alone isn t capable to turn on the transcription of a reporter gene. If bait and prey interact the transcription of a reporter gene is turned on measurement of reporter gene activity 15 Yeast two-hybrid cont. The method has been highly succesful in detecting many potential protein-protein interactions, but it has several limitations A bait capable to self-activation of the reporter gene is unsuitable If post-translational modifications (e.g. phosphorylation) are required for binding activity interaction is less likely to be detected Also if additional non-peptide factors (e.g. DNA sequence) are needed, the interaction is less likely to be detected Some interactions may lack biological content Discovery of novel interacting components not possible 16 8
9 MS driven techniques MS identification of proteins after quantitative analysis by 2DE Peptide mass fingerprinting (Maldi MS) Sequence based identification (MS/MS) Identification and quantitation using MS Labeling samples (e.g. ICAT, itraq) for quantitative analysis Identification of the post-translational modifications 17 Generation of peptides for MS analysis Proteins digestion Typically by trypsin; cleaves on the C-side of Arg and Lys i.e. generates peptides having R or K at the C-terminus Also other cleaving enzymes e.g. clostripain, endopeptidase Lys-C Peptide analysis Total mass of a peptide Peptide fragmentation (mass of single amino acids) 18 9
10 From Mann et al., 2001 Maldi-ToF MS Sample is cocrystallized with matrix and irradiated with laser, which leads to ionization of the peptides. The time-of-flight of the peptides is measured, which allows the determination of the peptide masses. 19 Peptide mass fingerprinting analysis 20 10
11 Peptide mass fingerprinting analysis cont. Database search for peptide masses Select: Database Taxonomy Enzyme Modifications Peptide tolerance Perform the search based on peptide masses 21 Peptide mass fingerprinting analysis cont
12 LC-MS analysis From Mann et al LC (Liquid Chromatogram) separation of complex mixtures of peptides A) Total ion current chromatogram B) Ion current for m/z = 591.5, selected ion current chromatogram C) Mass spectra at 117 min D) MS/MS spectrum at m/z and the interpreted amino acid sequence 23 Peptide fragmentation - ESI MS 24 12
13 m/z, amu m/ z, amu / Evaluation of the results - an example Human serum sample (w/o albumin depletion), 2DE and protein analysis using MS Proteins analyzed with Maldi and ESI Ion Trap α-antitrypsin No identification by Maldi Identification by ESI with high score (sample containing also human keratin...) Databases MW ~ 47kDa, pi ~ 5.4 Correct? 25 ICAT State 1 State 2 cicat labeling C 13 (0) C 13 (9) Combine, digest, fractionate by SCX HPLC µlc separation Fractionate to Sample plate MALDI sample plate, add matrix Intensity Intensity C 13 (0) Δm = 9.03 Da C 13 (9) m/z 1. MS scan to detect cicatabeled peptide pairs 2. Quantify and select differentially expressed proteins m/z Intensity 3. Select peptide for identification by MS/MS analysis m/z Courtesy of Timothy Griffin, Univ. Minnesota, USA 26 13
14 ICAT analysis Peptide identification by MS/MS or protein analysis by Maldi Quantitative analysis by ICAT peaks From Smolka et al Analysis of post-translational modifications From Mann et al Several strategies to analyze post-translational modifications Phosphorylated peak in MS; additional mass of Da 28 14
15 Proteomics summary There are several different approaches used in proteonome analysis; most of them are labourous at least at some point of analysis The methods are generally efficient for profiling the proteome and analyze expression differencies in case-control studies, but to obtain complete proteome usually one method is not enough References J Pevsner, Bioinformatics and Functional Genomics, John Wiley & Sons, NJ USA, 2003 RJ Simpson, Proteins and Proteomics, Cold Spring Harbor Laboratory Press, NY USA, 2002 Gygi SP, Rist B, Gerber SA, Turecek F, Gelb MH, Aebersold R, Quantitative analysis of complex protein mixtures using isotopecoded affinity tags, Nat Biotechnol 1999, 17: Mann M, Hendrickson RC, Pandey A, Analysis of Proteins and Proteomes by Mass Spectrometry, Annu. Rev. Biochem. 2001, 70: Smolka M, Zhou H, Aebersold R, Quantitative Protein Profiling Using Two-dimensional Gel Electrophoresis, Isotope-coded Affinity Tag Labeling, and Mass Spectrometry, Molecular and Cellular Proteomics 2002, 1.1:
Proteomics And Cancer Biomarker Discovery. Dr. Zahid Khan Institute of chemical Sciences (ICS) University of Peshawar. Overview. Cancer.
 Proteomics And Cancer Biomarker Discovery Dr. Zahid Khan Institute of chemical Sciences (ICS) University of Peshawar Overview Proteomics Cancer Aims Tools Data Base search Challenges Summary 1 Overview
Proteomics And Cancer Biomarker Discovery Dr. Zahid Khan Institute of chemical Sciences (ICS) University of Peshawar Overview Proteomics Cancer Aims Tools Data Base search Challenges Summary 1 Overview
Proteomics and Cancer
 Proteomics and Cancer Japan Society for the Promotion of Science (JSPS) Science Dialogue Program at Niitsu Senior High School Niitsu, Niigata September 4th 2006 Vladimir Valera, M.D, PhD JSPS Postdoctoral
Proteomics and Cancer Japan Society for the Promotion of Science (JSPS) Science Dialogue Program at Niitsu Senior High School Niitsu, Niigata September 4th 2006 Vladimir Valera, M.D, PhD JSPS Postdoctoral
Computing with large data sets
 Computing with large data sets Richard Bonneau, spring 2009 Lecture 14 (week 8): genomics 1 Central dogma Gene expression DNA RNA Protein v22.0480: computing with data, Richard Bonneau Lecture 14 places
Computing with large data sets Richard Bonneau, spring 2009 Lecture 14 (week 8): genomics 1 Central dogma Gene expression DNA RNA Protein v22.0480: computing with data, Richard Bonneau Lecture 14 places
Quantitative mass spec based proteomics
 Quantitative mass spec based proteomics Tuula Nyman Institute of Biotechnology tuula.nyman@helsinki.fi THE PROTEOME The complete protein complement expressed by a genome or by a cell or a tissue type (M.
Quantitative mass spec based proteomics Tuula Nyman Institute of Biotechnology tuula.nyman@helsinki.fi THE PROTEOME The complete protein complement expressed by a genome or by a cell or a tissue type (M.
Proteomics. Proteomics is the study of all proteins within organism. Challenges
 Proteomics Proteomics is the study of all proteins within organism. Challenges 1. The proteome is larger than the genome due to alternative splicing and protein modification. As we have said before we
Proteomics Proteomics is the study of all proteins within organism. Challenges 1. The proteome is larger than the genome due to alternative splicing and protein modification. As we have said before we
Advances in analytical biochemistry and systems biology: Proteomics
 Advances in analytical biochemistry and systems biology: Proteomics Brett Boghigian Department of Chemical & Biological Engineering Tufts University July 29, 2005 Proteomics The basics History Current
Advances in analytical biochemistry and systems biology: Proteomics Brett Boghigian Department of Chemical & Biological Engineering Tufts University July 29, 2005 Proteomics The basics History Current
MBios 478: Mass Spectrometry Applications [Dr. Wyrick] Slide #1. Lecture 25: Mass Spectrometry Applications
![MBios 478: Mass Spectrometry Applications [Dr. Wyrick] Slide #1. Lecture 25: Mass Spectrometry Applications MBios 478: Mass Spectrometry Applications [Dr. Wyrick] Slide #1. Lecture 25: Mass Spectrometry Applications](/thumbs/74/71368870.jpg) MBios 478: Mass Spectrometry Applications [Dr. Wyrick] Slide #1 Lecture 25: Mass Spectrometry Applications Measuring Protein Abundance o ICAT o DIGE Identifying Post-Translational Modifications Protein-protein
MBios 478: Mass Spectrometry Applications [Dr. Wyrick] Slide #1 Lecture 25: Mass Spectrometry Applications Measuring Protein Abundance o ICAT o DIGE Identifying Post-Translational Modifications Protein-protein
Proteomics: A Challenge for Technology and Information Science. What is proteomics?
 Proteomics: A Challenge for Technology and Information Science CBCB Seminar, November 21, 2005 Tim Griffin Dept. Biochemistry, Molecular Biology and Biophysics tgriffin@umn.edu What is proteomics? Proteomics
Proteomics: A Challenge for Technology and Information Science CBCB Seminar, November 21, 2005 Tim Griffin Dept. Biochemistry, Molecular Biology and Biophysics tgriffin@umn.edu What is proteomics? Proteomics
Strategies in proteomics
 Strategies in proteomics Systems biology - understand cellpathways, network, and complex interacting (includes Genomics, Proteomics, Metabolomics..) Biological processes - characterize protein complexes,
Strategies in proteomics Systems biology - understand cellpathways, network, and complex interacting (includes Genomics, Proteomics, Metabolomics..) Biological processes - characterize protein complexes,
Exam MOL3007 Functional Genomics
 Faculty of Medicine Department of Cancer Research and Molecular Medicine Exam MOL3007 Functional Genomics Thursday December 20 th 9.00-13.00 ECTS credits: 7.5 Number of pages (included front-page): 5 Supporting
Faculty of Medicine Department of Cancer Research and Molecular Medicine Exam MOL3007 Functional Genomics Thursday December 20 th 9.00-13.00 ECTS credits: 7.5 Number of pages (included front-page): 5 Supporting
Really high sensitivity mass spectrometry and Discovery and analysis of protein complexes
 Really high sensitivity mass spectrometry and Discovery and analysis of protein complexes The PRIME lab and AMS Importance of protein complexes in biology Methods for isolation of protein complexes In
Really high sensitivity mass spectrometry and Discovery and analysis of protein complexes The PRIME lab and AMS Importance of protein complexes in biology Methods for isolation of protein complexes In
Expression Proteomics: principles and examples of application
 Expression Proteomics: principles and examples of application Miguel Teixeira IBB Institute for Biotechnology and Bioengineering, Centro de Eng. Biológica e Química, IST Functional Genomics and Bioinformatics
Expression Proteomics: principles and examples of application Miguel Teixeira IBB Institute for Biotechnology and Bioengineering, Centro de Eng. Biológica e Química, IST Functional Genomics and Bioinformatics
Mass Spectrometry. Kyle Chau and Andrew Gioe
 Mass Spectrometry Kyle Chau and Andrew Gioe Computation of Molecular Mass - Mass Spectrum is a plot of intensity as a function of masscharge ratio, m/z. - Mass-charge ratio can be determined through accelerating
Mass Spectrometry Kyle Chau and Andrew Gioe Computation of Molecular Mass - Mass Spectrum is a plot of intensity as a function of masscharge ratio, m/z. - Mass-charge ratio can be determined through accelerating
LECTURE-3. Protein Chemistry to proteomics HANDOUT. Proteins are the most dynamic and versatile macromolecules in a living cell, which
 LECTURE-3 Protein Chemistry to proteomics HANDOUT PREAMBLE Proteins are the most dynamic and versatile macromolecules in a living cell, which regulates essential activities of the cell. The classical protein
LECTURE-3 Protein Chemistry to proteomics HANDOUT PREAMBLE Proteins are the most dynamic and versatile macromolecules in a living cell, which regulates essential activities of the cell. The classical protein
Využití cílené proteomiky pro kontrolu falšování potravin: identifikace peptidových markerů v mase pomocí LC- Q Exactive MS/MS
 Využití cílené proteomiky pro kontrolu falšování potravin: identifikace peptidových markerů v mase pomocí LC- Q Exactive MS/MS Michal Godula Ph.D. Thermo Fisher Scientific The world leader in serving science
Využití cílené proteomiky pro kontrolu falšování potravin: identifikace peptidových markerů v mase pomocí LC- Q Exactive MS/MS Michal Godula Ph.D. Thermo Fisher Scientific The world leader in serving science
SGN-6106 Computational Systems Biology I
 SGN-6106 Computational Systems Biology I A View of Modern Measurement Systems in Cell Biology Kaisa-Leena Taattola 21.2.2007 1 The cell a complex system (Source: Lehninger Principles of Biochemistry 4th
SGN-6106 Computational Systems Biology I A View of Modern Measurement Systems in Cell Biology Kaisa-Leena Taattola 21.2.2007 1 The cell a complex system (Source: Lehninger Principles of Biochemistry 4th
11/22/13. Proteomics, functional genomics, and systems biology. Biosciences 741: Genomics Fall, 2013 Week 11
 Proteomics, functional genomics, and systems biology Biosciences 741: Genomics Fall, 2013 Week 11 1 Figure 6.1 The future of genomics Functional Genomics The field of functional genomics represents the
Proteomics, functional genomics, and systems biology Biosciences 741: Genomics Fall, 2013 Week 11 1 Figure 6.1 The future of genomics Functional Genomics The field of functional genomics represents the
Basic protein and peptide science for proteomics. Henrik Johansson
 Basic protein and peptide science for proteomics Henrik Johansson Proteins are the main actors in the cell Membranes Transport and storage Chemical factories DNA Building proteins Structure Proteins mediate
Basic protein and peptide science for proteomics Henrik Johansson Proteins are the main actors in the cell Membranes Transport and storage Chemical factories DNA Building proteins Structure Proteins mediate
Lecture 5: 8/31. CHAPTER 5 Techniques in Protein Biochemistry
 Lecture 5: 8/31 CHAPTER 5 Techniques in Protein Biochemistry Chapter 5 Outline The proteome is the entire set of proteins expressed and modified by a cell under a particular set of biochemical conditions.
Lecture 5: 8/31 CHAPTER 5 Techniques in Protein Biochemistry Chapter 5 Outline The proteome is the entire set of proteins expressed and modified by a cell under a particular set of biochemical conditions.
Lecture 8: Affinity Chromatography-III
 Lecture 8: Affinity Chromatography-III Key words: Chromatography; Affinity chromatography; Protein Purification During this lecture, we shall be studying few more examples of affinity chromatography. The
Lecture 8: Affinity Chromatography-III Key words: Chromatography; Affinity chromatography; Protein Purification During this lecture, we shall be studying few more examples of affinity chromatography. The
2D separations and analysis of proteins in biological samples
 January 18, 2005 2D separations and analysis of proteins in biological samples Helen Kim 934-3880 helenkim@uab.edu McCallum Building, room 460 Jan 18, 2005 HelenKim/UAB/PharmTox 1 Learning objectives 2-D
January 18, 2005 2D separations and analysis of proteins in biological samples Helen Kim 934-3880 helenkim@uab.edu McCallum Building, room 460 Jan 18, 2005 HelenKim/UAB/PharmTox 1 Learning objectives 2-D
Improving Productivity with Applied Biosystems GPS Explorer
 Product Bulletin TOF MS Improving Productivity with Applied Biosystems GPS Explorer Software Purpose GPS Explorer Software is the application layer software for the Applied Biosystems 4700 Proteomics Discovery
Product Bulletin TOF MS Improving Productivity with Applied Biosystems GPS Explorer Software Purpose GPS Explorer Software is the application layer software for the Applied Biosystems 4700 Proteomics Discovery
27041, Week 02. Review of Week 01
 27041, Week 02 Review of Week 01 The human genome sequencing project (HGP) 2 CBS, Department of Systems Biology Systems Biology and emergent properties 3 CBS, Department of Systems Biology Different model
27041, Week 02 Review of Week 01 The human genome sequencing project (HGP) 2 CBS, Department of Systems Biology Systems Biology and emergent properties 3 CBS, Department of Systems Biology Different model
NPTEL
 NPTEL Syllabus Proteomics: Principles and Techniques - Video course COURSE OUTLINE An introduction to proteomics: Basics of protein structure and function, An overview of systems biology, Evolution from
NPTEL Syllabus Proteomics: Principles and Techniques - Video course COURSE OUTLINE An introduction to proteomics: Basics of protein structure and function, An overview of systems biology, Evolution from
Genomics, Transcriptomics and Proteomics
 Genomics, Transcriptomics and Proteomics GENES ARNm PROTEINS Genomics Transcriptomics Proteomics (10 6 protein forms?) Maturation PTMs Partners Localisation EDyP Lab : «Exploring the Dynamics of Proteomes»
Genomics, Transcriptomics and Proteomics GENES ARNm PROTEINS Genomics Transcriptomics Proteomics (10 6 protein forms?) Maturation PTMs Partners Localisation EDyP Lab : «Exploring the Dynamics of Proteomes»
Peptide biomarkers as a way to determine meat authenticity
 Peptide biomarkers as a way to determine meat authenticity Miguel A. Sentandreu Laboratory of Meat Science IATA (CSIC), SPAIN The problem of meat authentication Clear and reliable information about food
Peptide biomarkers as a way to determine meat authenticity Miguel A. Sentandreu Laboratory of Meat Science IATA (CSIC), SPAIN The problem of meat authentication Clear and reliable information about food
What are proteomics? And what can they tell us about seed maturation and germination?
 Danseed Symposium Kobæk Strand, Skelskør 14 March, 2017 What are proteomics? And what can they tell us about seed maturation and germination? Ian Max Møller Department of Molecular Biology and Genetics
Danseed Symposium Kobæk Strand, Skelskør 14 March, 2017 What are proteomics? And what can they tell us about seed maturation and germination? Ian Max Møller Department of Molecular Biology and Genetics
A New Strategy for Quantitative Proteomics Using Isotope-Coded Protein Labels
 ICPL TM -Isotope Coded Label A New Strategy for Quantitative Proteomics Using Isotope-Coded Labels Alexander Schmidt Department for Max-Planck-Institute of Biochemistry Spotfire User Conference 2004, Cologne
ICPL TM -Isotope Coded Label A New Strategy for Quantitative Proteomics Using Isotope-Coded Labels Alexander Schmidt Department for Max-Planck-Institute of Biochemistry Spotfire User Conference 2004, Cologne
Overview. Tools for Protein Sample Preparation, 2-D Electrophoresis, and Imaging and Analysis
 Expression Proteomics // Tools for Protein Separation and Analysis www.expressionproteomics.com 1 2 3 4 Overview Tools for Protein Sample Preparation, 2-D Electrophoresis, and Imaging and Analysis overview
Expression Proteomics // Tools for Protein Separation and Analysis www.expressionproteomics.com 1 2 3 4 Overview Tools for Protein Sample Preparation, 2-D Electrophoresis, and Imaging and Analysis overview
基于质谱的蛋白质药物定性定量分析技术及应用
 基于质谱的蛋白质药物定性定量分析技术及应用 蛋白质组学质谱数据分析 单抗全蛋白测序及鉴定 Bioinformatics Solutions Inc. Waterloo, ON, Canada 上海 中国 2017 1 3 PEAKS Studio GUI Introduction Overview of PEAKS Studio GUI Project View Tasks, running info,
基于质谱的蛋白质药物定性定量分析技术及应用 蛋白质组学质谱数据分析 单抗全蛋白测序及鉴定 Bioinformatics Solutions Inc. Waterloo, ON, Canada 上海 中国 2017 1 3 PEAKS Studio GUI Introduction Overview of PEAKS Studio GUI Project View Tasks, running info,
Strategies for Quantitative Proteomics. Atelier "Protéomique Quantitative" La Grande Motte, France - June 26, 2007
 Strategies for Quantitative Proteomics Atelier "Protéomique Quantitative", France - June 26, 2007 Bruno Domon, Ph.D. Institut of Molecular Systems Biology ETH Zurich Zürich, Switzerland OUTLINE Introduction
Strategies for Quantitative Proteomics Atelier "Protéomique Quantitative", France - June 26, 2007 Bruno Domon, Ph.D. Institut of Molecular Systems Biology ETH Zurich Zürich, Switzerland OUTLINE Introduction
Automation of MALDI-TOF Analysis for Proteomics
 Application Note #MT-50 BIFLEX III / REFLEX III Automation of MALDI-TOF Analysis for Proteomics D. Suckau, C. Köster, P.Hufnagel, K.-O. Kräuter and U. Rapp, Bruker Daltonik GmbH, Bremen, Germany Introduction
Application Note #MT-50 BIFLEX III / REFLEX III Automation of MALDI-TOF Analysis for Proteomics D. Suckau, C. Köster, P.Hufnagel, K.-O. Kräuter and U. Rapp, Bruker Daltonik GmbH, Bremen, Germany Introduction
Proteomics. Manickam Sugumaran. Department of Biology University of Massachusetts Boston, MA 02125
 Proteomics Manickam Sugumaran Department of Biology University of Massachusetts Boston, MA 02125 Genomic studies produced more than 75,000 potential gene sequence targets. (The number may be even higher
Proteomics Manickam Sugumaran Department of Biology University of Massachusetts Boston, MA 02125 Genomic studies produced more than 75,000 potential gene sequence targets. (The number may be even higher
Key questions of proteomics. Bioinformatics 2. Proteomics. Foundation of proteomics. What proteins are there? Protein digestion
 s s Key questions of proteomics What proteins are there? Bioinformatics 2 Lecture X roteomics How much is there of each of the proteins? - Absolute quantitation - Stoichiometry What (modification/splice)
s s Key questions of proteomics What proteins are there? Bioinformatics 2 Lecture X roteomics How much is there of each of the proteins? - Absolute quantitation - Stoichiometry What (modification/splice)
Exam MOL3007 Functional Genomics
 Faculty of Medicine Department of Cancer Research and Molecular Medicine Exam MOL3007 Functional Genomics Tuesday May 29 th 9.00-13.00 ECTS credits: 7.5 Number of pages (included front-page): 5 Supporting
Faculty of Medicine Department of Cancer Research and Molecular Medicine Exam MOL3007 Functional Genomics Tuesday May 29 th 9.00-13.00 ECTS credits: 7.5 Number of pages (included front-page): 5 Supporting
Mass Spectrometry-based Methods of Proteome Analysis
 1 Mass Spectrometry-based Methods of Proteome Analysis Boris L. Zybailov and Michael P. Washburn Stowers Institute for Medical Research, Kansas City, MO, USA 1 Principles and Instrumentation 3 1.1 Proteome
1 Mass Spectrometry-based Methods of Proteome Analysis Boris L. Zybailov and Michael P. Washburn Stowers Institute for Medical Research, Kansas City, MO, USA 1 Principles and Instrumentation 3 1.1 Proteome
Mass Spectrometry and Proteomics - Lecture 6 - Matthias Trost Newcastle University
 Mass Spectrometry and Proteomics - Lecture 6 - Matthias Trost Newcastle University matthias.trost@ncl.ac.uk Previously Quantitation techniques Label-free TMT SILAC Peptide identification and FDR 194 Lecture
Mass Spectrometry and Proteomics - Lecture 6 - Matthias Trost Newcastle University matthias.trost@ncl.ac.uk Previously Quantitation techniques Label-free TMT SILAC Peptide identification and FDR 194 Lecture
Proteomics Background and clinical utility
 Proteomics Background and clinical utility H.H. Helgason MD Antoni van Leeuwenhoek Hospital The Netherlands Cancer Institute Amsterdam Introduction Background Definitions Protein biomarkers Technical aspects
Proteomics Background and clinical utility H.H. Helgason MD Antoni van Leeuwenhoek Hospital The Netherlands Cancer Institute Amsterdam Introduction Background Definitions Protein biomarkers Technical aspects
Center for Mass Spectrometry and Proteomics Phone (612) (612)
 Outline Database search types Peptide Mass Fingerprint (PMF) Precursor mass-based Sequence tag Results comparison across programs Manual inspection of results Terminology Mass tolerance MS/MS search FASTA
Outline Database search types Peptide Mass Fingerprint (PMF) Precursor mass-based Sequence tag Results comparison across programs Manual inspection of results Terminology Mass tolerance MS/MS search FASTA
FACTORS THAT AFFECT PROTEIN IDENTIFICATION BY MASS SPECTROMETRY HAOFEI TIFFANY WANG. (Under the Direction of Ron Orlando) ABSTRACT
 FACTORS THAT AFFECT PROTEIN IDENTIFICATION BY MASS SPECTROMETRY by HAOFEI TIFFANY WANG (Under the Direction of Ron Orlando) ABSTRACT Mass spectrometry combined with database search utilities is a valuable
FACTORS THAT AFFECT PROTEIN IDENTIFICATION BY MASS SPECTROMETRY by HAOFEI TIFFANY WANG (Under the Direction of Ron Orlando) ABSTRACT Mass spectrometry combined with database search utilities is a valuable
Innovations for Protein Research. Protein Research. Powerful workflows built on solid science
 Innovations for Protein Research Protein Research Powerful workflows built on solid science Your partner in discovering innovative solutions for quantitative protein and biomarker research Applied Biosystems/MDS
Innovations for Protein Research Protein Research Powerful workflows built on solid science Your partner in discovering innovative solutions for quantitative protein and biomarker research Applied Biosystems/MDS
What is a proteome and how is the link between an organisms genome and a proteome.
 Answer to exam questions Bi2014 autumn 2013. Question 1a. What is a proteome and how is the link between an organisms genome and a proteome. The answer should discuss the following topics: What is a genome,
Answer to exam questions Bi2014 autumn 2013. Question 1a. What is a proteome and how is the link between an organisms genome and a proteome. The answer should discuss the following topics: What is a genome,
Enhancers mutations that make the original mutant phenotype more extreme. Suppressors mutations that make the original mutant phenotype less extreme
 Interactomics and Proteomics 1. Interactomics The field of interactomics is concerned with interactions between genes or proteins. They can be genetic interactions, in which two genes are involved in the
Interactomics and Proteomics 1. Interactomics The field of interactomics is concerned with interactions between genes or proteins. They can be genetic interactions, in which two genes are involved in the
Spectrum Mill MS Proteomics Workbench. Comprehensive tools for MS proteomics
 Spectrum Mill MS Proteomics Workbench Comprehensive tools for MS proteomics Meeting the challenge of proteomics data analysis Mass spectrometry is a core technology for proteomics research, but large-scale
Spectrum Mill MS Proteomics Workbench Comprehensive tools for MS proteomics Meeting the challenge of proteomics data analysis Mass spectrometry is a core technology for proteomics research, but large-scale
Combination of Isobaric Tagging Reagents and Cysteinyl Peptide Enrichment for In-Depth Quantification
 Combination of Isobaric Tagging Reagents and Cysteinyl Peptide Enrichment for In-Depth Quantification Protein Expression Analysis using the TripleTOF 5600 System and itraq Reagents Vojtech Tambor 1, Christie
Combination of Isobaric Tagging Reagents and Cysteinyl Peptide Enrichment for In-Depth Quantification Protein Expression Analysis using the TripleTOF 5600 System and itraq Reagents Vojtech Tambor 1, Christie
The best proteomics: combination of upfront reduction of sample complexity, reproducible resolution, and effective follow up
 The best proteomics: combination of upfront reduction of sample complexity, reproducible resolution, and effective follow up Helen Kim, Ph.D. Dept of Pharmacology & Toxicology University of Alabama at
The best proteomics: combination of upfront reduction of sample complexity, reproducible resolution, and effective follow up Helen Kim, Ph.D. Dept of Pharmacology & Toxicology University of Alabama at
Medicinal Chemistry of Modern Antibiotics
 Chemistry 259 Medicinal Chemistry of Modern Antibiotics Spring 2012 Lecture 5: Modern Target Discovery & MOA Thomas Hermann Department of Chemistry & Biochemistry University of California, San Diego Drug
Chemistry 259 Medicinal Chemistry of Modern Antibiotics Spring 2012 Lecture 5: Modern Target Discovery & MOA Thomas Hermann Department of Chemistry & Biochemistry University of California, San Diego Drug
Medicinal Chemistry of Modern Antibiotics
 Chemistry 259 Medicinal Chemistry of Modern Antibiotics Spring 2008 Lecture 5: Modern Target Discovery & MOA Thomas Hermann Department of Chemistry & Biochemistry University of California, San Diego Drug
Chemistry 259 Medicinal Chemistry of Modern Antibiotics Spring 2008 Lecture 5: Modern Target Discovery & MOA Thomas Hermann Department of Chemistry & Biochemistry University of California, San Diego Drug
Protein microarray and proteomics
 17 2 2005 4 Chinese Bulletin of Life Sciences Vol. 17, No. 2 Apr., 2005 1004-0374(2005)02-0132-05 510515 ; Q503;Q51 A Protein microarray and proteomics FEI Jia, MA Wen-Li*, ZHENG Wen-Ling (Institute of
17 2 2005 4 Chinese Bulletin of Life Sciences Vol. 17, No. 2 Apr., 2005 1004-0374(2005)02-0132-05 510515 ; Q503;Q51 A Protein microarray and proteomics FEI Jia, MA Wen-Li*, ZHENG Wen-Ling (Institute of
Chapter 9 Proteomics. From genomics to proteomics
 Chapter 9 Proteomics From genomics to proteomics From structural genomics, functional genomics to proteomics http://www.nature.com/scitable/topicpage/the-proteome-discovering-the-structure-and-function-613
Chapter 9 Proteomics From genomics to proteomics From structural genomics, functional genomics to proteomics http://www.nature.com/scitable/topicpage/the-proteome-discovering-the-structure-and-function-613
Topics. Protein Bioinformatics ( ) Progression of Protein Analysis by Mass Spectrometry. Protein identification by mass spectrometry
 Protein Bioinformatics (26.655) Lecture 15: Quantitative Proteomics Tuesday, May 5, 28 Robert N. Cole, Ph.D. Mass Spectrometry/Proteomics Facility at Johns Hopkins School of Medicine 371 Broadway Research
Protein Bioinformatics (26.655) Lecture 15: Quantitative Proteomics Tuesday, May 5, 28 Robert N. Cole, Ph.D. Mass Spectrometry/Proteomics Facility at Johns Hopkins School of Medicine 371 Broadway Research
Among the many unprecedented and fundamental. Proteomics Approaches. in Drug Discovery. Understanding the. human genome is. important, but most
 Proteomics Approaches in Drug Discovery Understanding the human genome is important, but most pharmaceuticals target proteins. Among the many unprecedented and fundamental changes that occurred in our
Proteomics Approaches in Drug Discovery Understanding the human genome is important, but most pharmaceuticals target proteins. Among the many unprecedented and fundamental changes that occurred in our
Introduction to Proteomics
 Introduction to Proteomics Tasso Miliotis, PhD AstraZeneca R&D Gothenburg Translational Sciences tasso.miliotis@astrazeneca.com 1 Site Management Mölndal May 2008 Outline Drug Discovery AZ R&D Sample Preparation
Introduction to Proteomics Tasso Miliotis, PhD AstraZeneca R&D Gothenburg Translational Sciences tasso.miliotis@astrazeneca.com 1 Site Management Mölndal May 2008 Outline Drug Discovery AZ R&D Sample Preparation
Protein analysis. Dr. Mamoun Ahram Summer semester, Resources This lecture Campbell and Farrell s Biochemistry, Chapters 5
 Protein analysis Dr. Mamoun Ahram Summer semester, 2015-2016 Resources This lecture Campbell and Farrell s Biochemistry, Chapters 5 Bases of protein separation Proteins can be purified on the basis Solubility
Protein analysis Dr. Mamoun Ahram Summer semester, 2015-2016 Resources This lecture Campbell and Farrell s Biochemistry, Chapters 5 Bases of protein separation Proteins can be purified on the basis Solubility
Outline and learning objectives. From Proteomics to Systems Biology. Integration of omics - information
 From to Systems Biology Outline and learning objectives Omics science provides global analysis tools to study entire systems How to obtain omics - What can we learn Limitations Integration of omics - In-class
From to Systems Biology Outline and learning objectives Omics science provides global analysis tools to study entire systems How to obtain omics - What can we learn Limitations Integration of omics - In-class
PROTEOMICS. Timothy Palzkill Baylor College of Medicine. KLUWER ACADEMIC PUBLISHERS New York / Boston / Dordrecht / London / Moscow
 PROTEOMICS PROTEOMICS by Timothy Palzkill Baylor College of Medicine KLUWER ACADEMIC PUBLISHERS New York / Boston / Dordrecht / London / Moscow ebook ISBN: 0-306-46895-6 Print ISBN: 0-792-37565-3 2002
PROTEOMICS PROTEOMICS by Timothy Palzkill Baylor College of Medicine KLUWER ACADEMIC PUBLISHERS New York / Boston / Dordrecht / London / Moscow ebook ISBN: 0-306-46895-6 Print ISBN: 0-792-37565-3 2002
Protein purification prior to proteomics analysis
 Protein purification prior to proteomics analysis Stephen Barnes, PhD Department of Pharmacology & Toxicology and Mass Spectrometry Shared Facility, UAB The dynamic range of protein abundances Proteins
Protein purification prior to proteomics analysis Stephen Barnes, PhD Department of Pharmacology & Toxicology and Mass Spectrometry Shared Facility, UAB The dynamic range of protein abundances Proteins
Experimental Techniques 2
 Experimental Techniques 2 High-throughput interaction detection Yeast two-hybrid - pairwise organisms as machines to learn about organisms yeast, worm, fly, human,... low intersection between repeated
Experimental Techniques 2 High-throughput interaction detection Yeast two-hybrid - pairwise organisms as machines to learn about organisms yeast, worm, fly, human,... low intersection between repeated
Zakopane, April, 2011
 Clinical Proteomics: A Technology to shape the future? Zakopane, April, 2011 Protein Chemistry/Proteomics/Peptide Synthesis and Array Unit Institute of Biomedicine/Anatomy Biomedicum Helsinki, 00014 University
Clinical Proteomics: A Technology to shape the future? Zakopane, April, 2011 Protein Chemistry/Proteomics/Peptide Synthesis and Array Unit Institute of Biomedicine/Anatomy Biomedicum Helsinki, 00014 University
Faster, easier, flexible proteomics solutions
 Agilent HPLC-Chip LC/MS Faster, easier, flexible proteomics solutions Our measure is your success. products applications soft ware services Phospho- Anaysis Intact Glycan & Glycoprotein Agilent s HPLC-Chip
Agilent HPLC-Chip LC/MS Faster, easier, flexible proteomics solutions Our measure is your success. products applications soft ware services Phospho- Anaysis Intact Glycan & Glycoprotein Agilent s HPLC-Chip
A Highly Accurate Mass Profiling Approach to Protein Biomarker Discovery Using HPLC-Chip/ MS-Enabled ESI-TOF MS
 Application Note PROTEOMICS METABOLOMICS GENOMICS INFORMATICS GLYILEVALCYSGLUGLNALASERLEUASPARG CYSVALLYSPROLYSPHETYRTHRLEUHISLYS A Highly Accurate Mass Profiling Approach to Protein Biomarker Discovery
Application Note PROTEOMICS METABOLOMICS GENOMICS INFORMATICS GLYILEVALCYSGLUGLNALASERLEUASPARG CYSVALLYSPROLYSPHETYRTHRLEUHISLYS A Highly Accurate Mass Profiling Approach to Protein Biomarker Discovery
Bioinformatics (Lec 19) Picture Copyright: the National Museum of Health
 3/29/05 1 Picture Copyright: AccessExcellence @ the National Museum of Health PCR 3/29/05 2 Schematic outline of a typical PCR cycle Target DNA Primers dntps DNA polymerase 3/29/05 3 Gel Electrophoresis
3/29/05 1 Picture Copyright: AccessExcellence @ the National Museum of Health PCR 3/29/05 2 Schematic outline of a typical PCR cycle Target DNA Primers dntps DNA polymerase 3/29/05 3 Gel Electrophoresis
PROTEIN AND PROTEOME ANALYTICS IDENTIFICATION, CHARACTERIZATION AND EXPRESSION ANALYSIS OF PROTEINS
 FRAUNHOFER INSTITUTE FOR INTERFACIAL ENGINEERING AND BIOTECHNOLOGY PROTEIN AND PROTEOME ANALYTICS IDENTIFICATION, CHARACTERIZATION AND EXPRESSION ANALYSIS OF PROTEINS 1 1 2 2 PROTEINS PROTEOMICS Proteins,
FRAUNHOFER INSTITUTE FOR INTERFACIAL ENGINEERING AND BIOTECHNOLOGY PROTEIN AND PROTEOME ANALYTICS IDENTIFICATION, CHARACTERIZATION AND EXPRESSION ANALYSIS OF PROTEINS 1 1 2 2 PROTEINS PROTEOMICS Proteins,
Combining Techniques to Answer Molecular Questions
 Combining Techniques to Answer Molecular Questions UNIT FM02 How to cite this article: Curr. Protoc. Essential Lab. Tech. 9:FM02.1-FM02.5. doi: 10.1002/9780470089941.etfm02s9 INTRODUCTION This manual is
Combining Techniques to Answer Molecular Questions UNIT FM02 How to cite this article: Curr. Protoc. Essential Lab. Tech. 9:FM02.1-FM02.5. doi: 10.1002/9780470089941.etfm02s9 INTRODUCTION This manual is
BIOLOGY 429 PROTEOMICS LABORATORY Fall 2004
 Dr. Karin Sauer Dr. Anna Tan-Wilson BIOLOGY 429 PROTEOMICS LABORATORY Fall 2004 COURSE BULLETIN DESCRIPTION: Proteomics techniques for the analysis of global changes in protein patterns. Methods include
Dr. Karin Sauer Dr. Anna Tan-Wilson BIOLOGY 429 PROTEOMICS LABORATORY Fall 2004 COURSE BULLETIN DESCRIPTION: Proteomics techniques for the analysis of global changes in protein patterns. Methods include
Appendix. Table of contents
 Appendix Table of contents 1. Appendix figures 2. Legends of Appendix figures 3. References Appendix Figure S1 -Tub STIL (41) DVFFYQADDEHYIPR (55) (64) AVLLDLEPR (72) 1 451aa (1071) YLNENQLSQLSVTR (1084)
Appendix Table of contents 1. Appendix figures 2. Legends of Appendix figures 3. References Appendix Figure S1 -Tub STIL (41) DVFFYQADDEHYIPR (55) (64) AVLLDLEPR (72) 1 451aa (1071) YLNENQLSQLSVTR (1084)
Genomes, Proteomes, and Bioinformatics
 URL: http://www.iupac.org/symposia/proceedings/phuket97/oxender.html 1999 IUPAC Genomes, Proteomes, and Bioinformatics Dale L. Oxender, Brian Moldover, and James D. Cavalcoli Parke-Davie Pharmaceutical
URL: http://www.iupac.org/symposia/proceedings/phuket97/oxender.html 1999 IUPAC Genomes, Proteomes, and Bioinformatics Dale L. Oxender, Brian Moldover, and James D. Cavalcoli Parke-Davie Pharmaceutical
Biology: the basis for smart proteomic approaches to protein analysis
 QuickTime and a TIFF (Uncompressed) decompressor are needed to see this picture. Biology: the basis for smart proteomic approaches to protein analysis Helen Kim, PhD Department of Pharmacology & Toxicology
QuickTime and a TIFF (Uncompressed) decompressor are needed to see this picture. Biology: the basis for smart proteomic approaches to protein analysis Helen Kim, PhD Department of Pharmacology & Toxicology
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/1144622/dc1 Supporting Online Material for Genome Transplantation in Bacteria: Changing One Species to Another Carole Lartigue, John I. Glass, * Nina Alperovich, Rembert
www.sciencemag.org/cgi/content/full/1144622/dc1 Supporting Online Material for Genome Transplantation in Bacteria: Changing One Species to Another Carole Lartigue, John I. Glass, * Nina Alperovich, Rembert
Proteomics. Areas of Application for Proteomics. Most Commonly Used Proteomics Techniques: Limitations: Examples
 Proteomics Areas of Application for Proteomics Most Commonly Used Proteomics Techniques: Antibody arrays Protein activity arrays 2-D gels ICAT technology SELDI Limitations: protein sources surfaces and
Proteomics Areas of Application for Proteomics Most Commonly Used Proteomics Techniques: Antibody arrays Protein activity arrays 2-D gels ICAT technology SELDI Limitations: protein sources surfaces and
Supplementary Fig. 1 Identification of Nedd4 as an IRS-2-associated protein in camp-treated FRTL-5 cells.
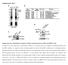 Supplementary Fig. 1 Supplementary Fig. 1 Identification of Nedd4 as an IRS-2-associated protein in camp-treated FRTL-5 cells. (a) FRTL-5 cells were treated with 1 mm dibutyryl camp for 24 h, and the lysates
Supplementary Fig. 1 Supplementary Fig. 1 Identification of Nedd4 as an IRS-2-associated protein in camp-treated FRTL-5 cells. (a) FRTL-5 cells were treated with 1 mm dibutyryl camp for 24 h, and the lysates
Kinetics Review. Tonight at 7 PM Phys 204 We will do two problems on the board (additional ones than in the problem sets)
 Quiz 1 Kinetics Review Tonight at 7 PM Phys 204 We will do two problems on the board (additional ones than in the problem sets) I will post the problems with solutions on Toolkit for those that can t make
Quiz 1 Kinetics Review Tonight at 7 PM Phys 204 We will do two problems on the board (additional ones than in the problem sets) I will post the problems with solutions on Toolkit for those that can t make
Genome Biology and Biotechnology
 Genome Biology and Biotechnology 10. The proteome Prof. M. Zabeau Department of Plant Systems Biology Flanders Interuniversity Institute for Biotechnology (VIB) University of Gent International course
Genome Biology and Biotechnology 10. The proteome Prof. M. Zabeau Department of Plant Systems Biology Flanders Interuniversity Institute for Biotechnology (VIB) University of Gent International course
Motivation From Protein to Gene
 MOLECULAR BIOLOGY 2003-4 Topic B Recombinant DNA -principles and tools Construct a library - what for, how Major techniques +principles Bioinformatics - in brief Chapter 7 (MCB) 1 Motivation From Protein
MOLECULAR BIOLOGY 2003-4 Topic B Recombinant DNA -principles and tools Construct a library - what for, how Major techniques +principles Bioinformatics - in brief Chapter 7 (MCB) 1 Motivation From Protein
Proteomics software at MSI. Pratik Jagtap Minnesota Supercomputing institute
 Proteomics software at MSI. Pratik Jagtap Minnesota Supercomputing institute http://www.mass.msi.umn.edu/ Proteomics software at MSI. proteomics : emerging technology proteomics workflow search algorithms
Proteomics software at MSI. Pratik Jagtap Minnesota Supercomputing institute http://www.mass.msi.umn.edu/ Proteomics software at MSI. proteomics : emerging technology proteomics workflow search algorithms
ENCODE DCC Antibody Validation Document
 ENCODE DCC Antibody Validation Document Date of Submission 09/12/12 Name: Trupti Kawli Email: trupti@stanford.edu Lab Snyder Antibody Name: SREBP1 (sc-8984) Target: SREBP1 Company/ Source: Santa Cruz Biotechnology
ENCODE DCC Antibody Validation Document Date of Submission 09/12/12 Name: Trupti Kawli Email: trupti@stanford.edu Lab Snyder Antibody Name: SREBP1 (sc-8984) Target: SREBP1 Company/ Source: Santa Cruz Biotechnology
From Proteomics to Systems Biology. Integration of omics - information
 From Proteomics to Systems Biology Integration of omics - information Outline and learning objectives Omics science provides global analysis tools to study entire systems How to obtain omics - data What
From Proteomics to Systems Biology Integration of omics - information Outline and learning objectives Omics science provides global analysis tools to study entire systems How to obtain omics - data What
CHROMATOGRAPHY OF DIFFICULT PROTEINS. Andrew J. Alpert, Ph.D. PolyLC Inc. Columbia, MD U.S.A.
 CHROMATOGRAPHY OF DIFFICULT PROTEINS Andrew J. Alpert, Ph.D. PolyLC Inc. Columbia, MD U.S.A. PROTEOMICS ANALYSES A butterfly and a caterpillar have the same genome but different proteomes. Techniques for
CHROMATOGRAPHY OF DIFFICULT PROTEINS Andrew J. Alpert, Ph.D. PolyLC Inc. Columbia, MD U.S.A. PROTEOMICS ANALYSES A butterfly and a caterpillar have the same genome but different proteomes. Techniques for
Proteomics. Areas of Application for Proteomics Most Commonly Used Proteomics Techniques: Limitations: Examples
 Proteomics Areas of Application for Proteomics Most Commonly Used Proteomics Techniques: Antibody arrays Protein activity arrays 2-D gels ICAT technology SELDI Limitations: protein sources surfaces and
Proteomics Areas of Application for Proteomics Most Commonly Used Proteomics Techniques: Antibody arrays Protein activity arrays 2-D gels ICAT technology SELDI Limitations: protein sources surfaces and
Areas of Application for Proteomics Most Commonly Used Proteomics Techniques:
 Proteomics Areas of Application for Proteomics Most Commonly Used Proteomics Techniques: Antibody arrays Protein activity arrays 2-D gels ICAT technology SELDI Limitations: Examples protein sources surfaces
Proteomics Areas of Application for Proteomics Most Commonly Used Proteomics Techniques: Antibody arrays Protein activity arrays 2-D gels ICAT technology SELDI Limitations: Examples protein sources surfaces
Towards unbiased biomarker discovery
 Towards unbiased biomarker discovery High-throughput molecular profiling technologies are routinely applied for biomarker discovery to make the drug discovery process more efficient and enable personalised
Towards unbiased biomarker discovery High-throughput molecular profiling technologies are routinely applied for biomarker discovery to make the drug discovery process more efficient and enable personalised
Topic 2: Proteins. 2-1 specific proteins can be purified from cell extracts. Molecular Biology and Public Health ( 分子生物学与公共卫生 )
 HAPTER20: Techniques of Molecular Biology Molecular Biology and Public Health ( 分子生物学与公共卫生 ) -------Protein manipulation ( 蛋白操作 ) Topic 2: Proteins 1. Protein purification ( 蛋白质纯化 ) 2. Affinity chromatography
HAPTER20: Techniques of Molecular Biology Molecular Biology and Public Health ( 分子生物学与公共卫生 ) -------Protein manipulation ( 蛋白操作 ) Topic 2: Proteins 1. Protein purification ( 蛋白质纯化 ) 2. Affinity chromatography
New workflows for protein expression analysis ICAT. Reagent Technology
 New workflows for protein expression analysis ICAT Reagent Technology Breakthrough technology for protein expression analysis accelerate In protein-expression analysis studies, ICAT reagent technology
New workflows for protein expression analysis ICAT Reagent Technology Breakthrough technology for protein expression analysis accelerate In protein-expression analysis studies, ICAT reagent technology
Chapter 5: Proteins: Primary Structure
 Instant download and all chapters Test Bank Fundamentals of Biochemistry Life at the Molecular Level 4th Edition Donald Voet https://testbanklab.com/download/test-bank-fundamentals-biochemistry-life-molecular-level-
Instant download and all chapters Test Bank Fundamentals of Biochemistry Life at the Molecular Level 4th Edition Donald Voet https://testbanklab.com/download/test-bank-fundamentals-biochemistry-life-molecular-level-
Biomarker Discovery using Surface Plasmon Resonance Imaging
 F e a t u r e A r t i c l e Feature Article Biomarker Discovery using Surface Plasmon Resonance Imaging Elodie LY-MORIN, Sophie BELLON, Géraldine MÉLIZZI, Chiraz FRYDMAN Surface Plasmon Resonance (SPR)
F e a t u r e A r t i c l e Feature Article Biomarker Discovery using Surface Plasmon Resonance Imaging Elodie LY-MORIN, Sophie BELLON, Géraldine MÉLIZZI, Chiraz FRYDMAN Surface Plasmon Resonance (SPR)
ENCODE DCC Antibody Validation Document
 ENCODE DCC Antibody Validation Document Date of Submission 06/15/2012 Name: Trupti Kawli Email: trupti@stanford.edu Lab Snyder Antibody Name: CDP (sc6327) Target: CDP Company/ Source: Santa Cruz Catalog
ENCODE DCC Antibody Validation Document Date of Submission 06/15/2012 Name: Trupti Kawli Email: trupti@stanford.edu Lab Snyder Antibody Name: CDP (sc6327) Target: CDP Company/ Source: Santa Cruz Catalog
ProMass HR Applications!
 ProMass HR Applications! ProMass HR Features Ø ProMass HR includes features for high resolution data processing. Ø ProMass HR includes the standard ProMass deconvolution algorithm as well as the full Positive
ProMass HR Applications! ProMass HR Features Ø ProMass HR includes features for high resolution data processing. Ø ProMass HR includes the standard ProMass deconvolution algorithm as well as the full Positive
Laboratory Water Quality Affects Protein Separation by 2D Gel Electrophoresis
 Laboratory Water Quality Affects Protein Separation by 2D Gel Electrophoresis 2D gel electrophoresis remains a dominant technique in proteomics. Obtaining high quality gels requires careful and tedious
Laboratory Water Quality Affects Protein Separation by 2D Gel Electrophoresis 2D gel electrophoresis remains a dominant technique in proteomics. Obtaining high quality gels requires careful and tedious
Application Note # ET-20 BioPharma Compass: A fully Automated Solution for Characterization and QC of Intact and Digested Proteins
 Application Note # ET-20 BioPharma Compass: A fully Automated Solution for Characterization and QC of Intact and Digested Proteins BioPharma Compass TM is a fully automated solution for the rapid characterization
Application Note # ET-20 BioPharma Compass: A fully Automated Solution for Characterization and QC of Intact and Digested Proteins BioPharma Compass TM is a fully automated solution for the rapid characterization
Comparability Analysis of Protein Therapeutics by Bottom-Up LC-MS with Stable Isotope-Tagged Reference Standards
 Comparability Analysis of Protein Therapeutics by Bottom-Up LC-MS with Stable Isotope-Tagged Reference Standards 16 September 2011 Abbott Bioresearch Center, Worcester, MA USA Manuilov, A. V., C. H. Radziejewski
Comparability Analysis of Protein Therapeutics by Bottom-Up LC-MS with Stable Isotope-Tagged Reference Standards 16 September 2011 Abbott Bioresearch Center, Worcester, MA USA Manuilov, A. V., C. H. Radziejewski
Spectral Counting Approaches and PEAKS
 Spectral Counting Approaches and PEAKS INBRE Proteomics Workshop, April 5, 2017 Boris Zybailov Department of Biochemistry and Molecular Biology University of Arkansas for Medical Sciences 1. Introduction
Spectral Counting Approaches and PEAKS INBRE Proteomics Workshop, April 5, 2017 Boris Zybailov Department of Biochemistry and Molecular Biology University of Arkansas for Medical Sciences 1. Introduction
Proteomics and Mass Spectrometry
 Molecular Cell Biology Lecture. Oct. 18, 2018 Proteomics and Mass Spectrometry Ron Bose, MD PhD Biochemistry and Molecular Cell Biology Programs Lab: Couch Research Building (4515 McKinley), 3 rd floor
Molecular Cell Biology Lecture. Oct. 18, 2018 Proteomics and Mass Spectrometry Ron Bose, MD PhD Biochemistry and Molecular Cell Biology Programs Lab: Couch Research Building (4515 McKinley), 3 rd floor
Unit title: Protein Structure and Function (SCQF level 8)
 Higher National Unit specification General information Unit code: H92J 35 Superclass: RH Publication date: May 2015 Source: Scottish Qualifications Authority Version: 01 Unit purpose This Unit is designed
Higher National Unit specification General information Unit code: H92J 35 Superclass: RH Publication date: May 2015 Source: Scottish Qualifications Authority Version: 01 Unit purpose This Unit is designed
Application Note TOF/MS
 Application Note TOF/MS New Level of Confidence for Protein Identification: Results Dependent Analysis and Peptide Mass Fingerprinting Using the 4700 Proteomics Discovery System Purpose The Applied Biosystems
Application Note TOF/MS New Level of Confidence for Protein Identification: Results Dependent Analysis and Peptide Mass Fingerprinting Using the 4700 Proteomics Discovery System Purpose The Applied Biosystems
Detecting Challenging Post Translational Modifications (PTMs) using CESI-MS
 Detecting Challenging Post Translational Modifications (PTMs) using CESI-MS Sensitive workflow for the detection of PTMs in biological samples from small sample volumes Klaus Faserl, 1 Bettina Sarg, 1
Detecting Challenging Post Translational Modifications (PTMs) using CESI-MS Sensitive workflow for the detection of PTMs in biological samples from small sample volumes Klaus Faserl, 1 Bettina Sarg, 1
Protein Microarrays?
 Protein Microarrays? Protein Chemistry/Proteomics Unit and the Neuroscience Program,Biomedicum Helsinki E-Mail: marc.baumann@helsinki.fi (http://research.med.helsinki.fi/corefacilities/proteinchem) What
Protein Microarrays? Protein Chemistry/Proteomics Unit and the Neuroscience Program,Biomedicum Helsinki E-Mail: marc.baumann@helsinki.fi (http://research.med.helsinki.fi/corefacilities/proteinchem) What
NPTEL VIDEO COURSE PROTEOMICS PROF. SANJEEVA SRIVASTAVA
 LECTURE-24 Quantitative proteomics: Stable isotope labeling by amino acids in cell culture (SILAC) TRANSCRIPT Welcome to the proteomics course. In today s lecture we will talk about Quantitative proteomics:
LECTURE-24 Quantitative proteomics: Stable isotope labeling by amino acids in cell culture (SILAC) TRANSCRIPT Welcome to the proteomics course. In today s lecture we will talk about Quantitative proteomics:
This is the author's accepted version of the manuscript.
 This is the author's accepted version of the manuscript. The definitive version is published in Nature Communications Online Edition: 2015/4/16 (Japan time), doi:10.1038/ncomms7780. The final version published
This is the author's accepted version of the manuscript. The definitive version is published in Nature Communications Online Edition: 2015/4/16 (Japan time), doi:10.1038/ncomms7780. The final version published
Investigation of a Mammalian Cellular Model for Differential Protein Expression Analysis Using 1D PAGE and Cleavable ICAT Reagents
 pplication Note 1D PGE and Cleavable ICT Reagents Investigation of a Mammalian Cellular Model for Differential Protein Expression nalysis Using 1D PGE and Cleavable ICT Reagents Overview The use of two-dimensional
pplication Note 1D PGE and Cleavable ICT Reagents Investigation of a Mammalian Cellular Model for Differential Protein Expression nalysis Using 1D PGE and Cleavable ICT Reagents Overview The use of two-dimensional
Assessment of Active Biopharmaceutical Ingredients Prior To and Following Removal of Interfering Excipients
 Assessment of Active Biopharmaceutical Ingredients Prior To and Following Removal of Interfering Excipients Andrew J Reason and Howard R Morris BioPharmaSpec Ltd, and BioPharmaSpec Inc, The EMA guideline
Assessment of Active Biopharmaceutical Ingredients Prior To and Following Removal of Interfering Excipients Andrew J Reason and Howard R Morris BioPharmaSpec Ltd, and BioPharmaSpec Inc, The EMA guideline
