P&TIT: A computer tool for predicting prototypical transcription promoter and terminator elements by conserved motifs
|
|
|
- Alexander Osborne
- 6 years ago
- Views:
Transcription
1 P&TIT: A computer tool for predicting prototypical transcription promoter and terminator elements by conserved motifs Marco Di Salvo a, Eva Pinatel b, Gianluca De Bellis b, Adelfia Talà a, Clelia Peano b, Pietro Alifano a a Department of Biological and Environmental Sciences and Technologies, University of Salento, Lecce, Italy b Institute of Biomedical Technologies National Research Council, Segrate, Milan, Italy. Abstract Over the last few decades, computational genomics has tremendously contributed to decipher biology from genome sequences and related data. Considerable effort has been devoted to prediction of transcription promoter and terminator sites that represent the essential punctuation marks for DNA transcription. In this study we have investigated the possibility to identify putative promoters and Rhodependent terminators in prokaryotes on the basis of evolutionarily conserved motifs. The final aim of this work is to develop computer software to predict location of bacterial promoters and terminators based on nucleic acid motifs. Introduction The evolutionary relatedness and/or functional analogies between transcription initiation and termination mechanisms in all three domains of life prompted us to explore the possibility to recognize promoter and terminator elements in a prokaryotic genome based on conserved structured motifs [1]. For promoter prediction we focused on possible G-quadruplex structures upstream of AT-rich elements. G-quadruplexes, non-canonical nucleic acid topologies with little established biological roles, are increasingly considered for conserved regulatory element discovery [2]. For Rho-dependent terminator prediction, based on current models for Rhodependent termination we used an algorithm based on C>G ratio, cytosine spacing, and hairpin structures (possible RNA polymerase pausing sites) downstream from a C>G content motif. Results of genome mining by were then validated by RNAseq data in the actinobacterium Streptomyces ambofaciens. Materials and methods Algorithms to identify the motifs of interest were generated by using the programming language Python v.3.5 taking as input data the annotated genome sequence of S. ambofaciens ATCC for promoter prediction, and RNAseq for validation. Methods to identify putative promoter and terminator elements are described afterwards. 102
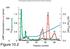
2 Results Method to identify putative promoters and results A three-step procedure was used to detect putative promoters (Figure 1): i.) The first step consisted in the identification of the putative promoter AT-rich element. To this purpose, within each intergenic sequence ( 50 nt) associated with transcribed genes, the 25-bp long region with the highest AT% (herein referred to as AT-max motif) was selected; ii.) The second step was the identification of putative G-quadruplex motifs extended up to 50 bp upstream from the 5 -end of the selected AT-max motif. Motif GxNyGxNyGxNyGx, with 2 x 4 and 1 y 10 is commonly used to predict the presence of G-quadruplexes [3]; iii.) The third step consisted in validation of promoter prediction by RNAseq data. We considered the 75-bp long elements (including the G- quadruplex and AT-max motifs) as possible promoters if there was an increase of read values by a factor of at least 1.5 between the read value at the AT-max motif 5 -end point and the read value 10 bp downstream from the AT-max motif 3 -end point, the latter with a read value 5. The results are shown in Table 1. In Figure 2 are represented some examples of putative promoter signals mapped by the RNAseq graphical user interface. Methods to identify putative Rhodependent terminators and results The procedure used to detect Rhodependent terminators was the following: i.) The first step consisted in the identification of putative RUT site. To this purpose, within each deduced intercistronic ( 78 nt) or intracistronic RNA sequence of transcribed genes, 78-nt long regions with C/G ratio 1.3 and regularly spaced cytosine residues (every nt) were selected (the so called RUT sites) [4]; ii.) The second step was the identification of hairpin structure (often serving as RNAP pausing sites) in a region extended up to 150 nt downstream from the 3 -end point of the C>G content motif; iii.) The third step consisted in validation of Rhodependent transcription terminator prediction by RNAseq data. We considered these regions as putative Rho-dependent terminators if there was a decrease of read value by a factor of at least 1.5 between the read value at the C>G content motif 5 -end point and the read value 150 nt downstream from the C>G content motif 3 -end point, the first with a read value 5. The results are shown in Tables 2 and 3. In Figure 3 are represented some examples of putative Rhodependent terminator signals mapped by the RNAseq graphical user interface. Methods identify putative intrinsic terminators and results In order to identify intrinsic (Rhoindependent) terminators, in all the intergenic sequences of the genome we looked for RNA hairpins followed by a run of U residues (min 3, max 8). We considered these regions as possible intrinsic terminators if there was a decrease of RNAseq read value by a factor of at least 1.5 between a point located 50 nt upstream from the U residues and a point located 5 nt downstream from the U residues, the first with the read value 5. The results are shown in Table 4. Conclusions In this study we investigated the possibility to identify promoter and terminator elements by detecting evolutionarily conserved motifs. The algorithm predicted putative promoters in about % of intergenic sequences associated with transcribed genes, and prediction was validated by RNAseq data in a percentage of cases ranging from 38.3% to 42.4% (Table 1). Interestingly, these high percentages very similar to those found in other evolutionarily distant promoters including yeast [5] and human gene 103
3 promoters more than 40% of which were shown to contain one or more G- quadruplexex [6]. Our results further support the idea that G-quadruplex is a prototypical motif involved in general promoter function/regulation. For Rho-dependent terminator prediction, the algorithm predicted putative Rhodependent terminators in about % of intercistronic sequences associated with transcribed genes, and prediction was validated by RNAseq data in a percentage of cases ranging from 31.1% to 36.4% (Table 2). In contrast, an algorithm predicted the presence of putative Rho-independent (intrinsic) terminators in a very small percentage of transcribed intercistronic sequences ranging from 6.3% to 6.7% with percentages of validation by RNAseq data ranging from 27.8% to 36.2% (Table 4). This very limited number of putative Rhoindependent terminators in S. ambofaciens suggests that Rho (or factor)-dependent termination mechanisms may be predominant in this microorganism. More interestingly, the presence of putative Rhodependent terminators was predicted in about % of transcribed intracistronic sequences, and results were validated in a percentage of cases ranging from 71.4% to 73.4% (Table 3). Future efforts will be dedicated to extending our analysis to other organisms with the final aim to better characterize what appear to be prototypical elements of the punctuation marks for DNA transcription. References [1] Burton SP, Burton ZF. The σ enigma: bacterial σ factors, archaeal TFB and eukaryotic TFIIB are homologs. Transcription 5:e (2014). [2] Rhodes D, Lipps HJ. G-quadruplexes and their regulatory roles in biology. Nucleic Acids Res 43: (2015). [3] Kikin O, D Antonio L and Bagga PS. (2006). QGRS Mapper: a web-based server for predicting G-quadruplexes in nucleotide sequenze. Nucleic Acids Res 34. [4] Alifano P, Rivellini F, Limauro D, Bruni CB, Carlomagno MS. A consensus motif common to all Rho-dependent prokaryotic transcription terminators. Cell 64: (1991). [5] Capra JA, Paeschke K, Singh M, Zakian VA. G- quadruplex DNA sequences are evolutionarily conserved and associated with distinct genomic features in Saccharomyces cerevisiae. PLoS Comput Biol 6:e (2010). [6] Huppert JL, Balasubramanian S. G- quadruplexes in promoters throughout the human genome. Nucleic Acids Res 35: (2007). 104
4 Table 1. Statistics of putative promoters that were identified by G-quadruplex and AT-max motif-based algorithm. Sampling time (h) Intergenic sequences with putative promoters (%) promoters (%) promoters with TANNNT consensus (%) , , , ,8 promoters with TTGAC consensus (%) Table 2. Statistics of putative extracistronic Rho-dependent terminators that were identified by C>G content RNA motif-based algorithm. Sampling time (h) Extracistronic sequences with Rho-dependent terminators (%) a extracistronic Rhodependent terminators (%) b a Percentages of extracistronic sequences that possess at least one possible RUT sequence. b Percentage of extracistronic sequences with at least one possible RUT sequence, which we consider as possible Rho-dependent terminators, according to of RNAseq data. Table 3. Statistics of putative intracistronic Rho-dependent terminators that were identified by C>G content RNA motif-based algorithm. Sampling time (h) Intracistronic sequences with Rho-dependent terminators (%) a intracistronic Rho-dependent terminators (%) b a Percentages of intracistronic sequences that possess at least one possible RUT sequence. b Percentage of intracistronic sequences with at least one possible RUT sequence, which we consider as possible Rho-dependent terminators, according to of RNAseq data. 105
5 Table 4. Statistics of putative Rho-independent terminators that were identified by the algorithm a Sampling time (h) Intercistronic sequences with Rho-independent terminators (%) a intercistronic Rho-independent terminators (%) a RNA hairpins followed by a run of U residues (min 3, max 8) Fig.1. Graphical description of the procedure used to detect putative promoters Fig. 2. Examples of putative promoter signals mapped by the RNAseq graphical user interface. 106
6 Fig. 3. Examples of putative Rho-dependent terminator signals mapped by the RNAseq graphical user interface 107
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Origin use and efficiency are similar among WT, rrm3, pif1-m2, and pif1-m2; rrm3 strains. A. Analysis of fork progression around confirmed and likely origins (from cerevisiae.oridb.org).
Supplementary Figure 1 Origin use and efficiency are similar among WT, rrm3, pif1-m2, and pif1-m2; rrm3 strains. A. Analysis of fork progression around confirmed and likely origins (from cerevisiae.oridb.org).
Expression of the genome. Books: 1. Molecular biology of the gene: Watson et al 2. Genetics: Peter J. Russell
 Expression of the genome Books: 1. Molecular biology of the gene: Watson et al 2. Genetics: Peter J. Russell 1 Transcription 1. Francis Crick (1956) named the flow of information from DNA RNA protein the
Expression of the genome Books: 1. Molecular biology of the gene: Watson et al 2. Genetics: Peter J. Russell 1 Transcription 1. Francis Crick (1956) named the flow of information from DNA RNA protein the
Positional Preference of Rho-Independent Transcriptional Terminators in E. Coli
 Positional Preference of Rho-Independent Transcriptional Terminators in E. Coli Annie Vo Introduction Gene expression can be regulated at the transcriptional level through the activities of terminators.
Positional Preference of Rho-Independent Transcriptional Terminators in E. Coli Annie Vo Introduction Gene expression can be regulated at the transcriptional level through the activities of terminators.
Identifying Regulatory Regions using Multiple Sequence Alignments
 Identifying Regulatory Regions using Multiple Sequence Alignments Prerequisites: BLAST Exercise: Detecting and Interpreting Genetic Homology. Resources: ClustalW is available at http://www.ebi.ac.uk/tools/clustalw2/index.html
Identifying Regulatory Regions using Multiple Sequence Alignments Prerequisites: BLAST Exercise: Detecting and Interpreting Genetic Homology. Resources: ClustalW is available at http://www.ebi.ac.uk/tools/clustalw2/index.html
Prokaryotic Physiology. March 3, 2017
 1. (10 pts) Explain the replication of both strands of DNA in prokaryotes. At a minimum explain the direction of synthesis, synthesis of the leading and lagging strand, separation of the strands and the
1. (10 pts) Explain the replication of both strands of DNA in prokaryotes. At a minimum explain the direction of synthesis, synthesis of the leading and lagging strand, separation of the strands and the
DNA Transcription. Dr Aliwaini
 DNA Transcription 1 DNA Transcription-Introduction The synthesis of an RNA molecule from DNA is called Transcription. All eukaryotic cells have five major classes of RNA: ribosomal RNA (rrna), messenger
DNA Transcription 1 DNA Transcription-Introduction The synthesis of an RNA molecule from DNA is called Transcription. All eukaryotic cells have five major classes of RNA: ribosomal RNA (rrna), messenger
Biochemistry 111. Carl Parker x A Braun
 Biochemistry 111 Carl Parker x6368 101A Braun csp@caltech.edu Central Dogma of Molecular Biology DNA-Dependent RNA Polymerase Requires a DNA Template Synthesizes RNA in a 5 to 3 direction Requires ribonucleoside
Biochemistry 111 Carl Parker x6368 101A Braun csp@caltech.edu Central Dogma of Molecular Biology DNA-Dependent RNA Polymerase Requires a DNA Template Synthesizes RNA in a 5 to 3 direction Requires ribonucleoside
RNA Expression of the information in a gene generally involves production of an RNA molecule transcribed from a DNA template. RNA differs from DNA
 RNA Expression of the information in a gene generally involves production of an RNA molecule transcribed from a DNA template. RNA differs from DNA that it has a hydroxyl group at the 2 position of the
RNA Expression of the information in a gene generally involves production of an RNA molecule transcribed from a DNA template. RNA differs from DNA that it has a hydroxyl group at the 2 position of the
BIOCHEMISTRY REVIEW. Overview of Biomolecules. Chapter 12 Transcription
 BIOCHEMISTRY REVIEW Overview of Biomolecules Chapter 12 Transcription 2 3 4 5 Are You Getting It?? Which are general characteristics of transcription? (multiple answers) a) An entire DNA molecule is transcribed
BIOCHEMISTRY REVIEW Overview of Biomolecules Chapter 12 Transcription 2 3 4 5 Are You Getting It?? Which are general characteristics of transcription? (multiple answers) a) An entire DNA molecule is transcribed
GENETICS - CLUTCH CH.10 TRANSCRIPTION.
 !! www.clutchprep.com CONCEPT: OVERVIEW OF TRANSCRIPTION Transcription is the process of using DNA as a template to RNA RNA polymerase is the enzyme that transcribes DNA - There are many different types
!! www.clutchprep.com CONCEPT: OVERVIEW OF TRANSCRIPTION Transcription is the process of using DNA as a template to RNA RNA polymerase is the enzyme that transcribes DNA - There are many different types
DNA RNA. Protein. Enzymes in the central dogma. Cellular enzymes (Mostly) RNA virus enzymes. DNA polymerase. Reverse transcriptase.
 Enzymes in the central dogma Cellular enzymes (Mostly) RNA virus enzymes DNA DNA polymerase Reverse transcriptase RNA polymerase RNA-dependent RNA polymerase RNA Ribosome Protein The process of transcription
Enzymes in the central dogma Cellular enzymes (Mostly) RNA virus enzymes DNA DNA polymerase Reverse transcriptase RNA polymerase RNA-dependent RNA polymerase RNA Ribosome Protein The process of transcription
M1 - Biochemistry. Nucleic Acid Structure II/Transcription I
 M1 - Biochemistry Nucleic Acid Structure II/Transcription I PH Ratz, PhD (Resources: Lehninger et al., 5th ed., Chapters 8, 24 & 26) 1 Nucleic Acid Structure II/Transcription I Learning Objectives: 1.
M1 - Biochemistry Nucleic Acid Structure II/Transcription I PH Ratz, PhD (Resources: Lehninger et al., 5th ed., Chapters 8, 24 & 26) 1 Nucleic Acid Structure II/Transcription I Learning Objectives: 1.
Transcription. By : Lucia Dhiantika Witasari M.Biotech., Apt
 Transcription By : Lucia Dhiantika Witasari M.Biotech., Apt REGULATION OF GENE EXPRESSION 11/26/2010 2 RNA Messenger RNAs (mrnas) encode the amino acid sequence of one or more polypeptides specified by
Transcription By : Lucia Dhiantika Witasari M.Biotech., Apt REGULATION OF GENE EXPRESSION 11/26/2010 2 RNA Messenger RNAs (mrnas) encode the amino acid sequence of one or more polypeptides specified by
Mechanisms of Transcription. School of Life Science Shandong University
 Mechanisms of Transcription School of Life Science Shandong University Ch 12: Mechanisms of Transcription 1. RNA polymerase and the transcription cycle 2. The transcription cycle in bacteria 3. Transcription
Mechanisms of Transcription School of Life Science Shandong University Ch 12: Mechanisms of Transcription 1. RNA polymerase and the transcription cycle 2. The transcription cycle in bacteria 3. Transcription
Lecture 11. Initiation of RNA Pol II transcription. Transcription Initiation Complex
 Lecture 11 *Eukaryotic Transcription Gene Organization RNA Processing 5 cap 3 polyadenylation splicing Translation Initiation of RNA Pol II transcription Consensus sequence of promoter TATA Transcription
Lecture 11 *Eukaryotic Transcription Gene Organization RNA Processing 5 cap 3 polyadenylation splicing Translation Initiation of RNA Pol II transcription Consensus sequence of promoter TATA Transcription
Chapter 11 Part A: Metabolism: The synthesis of nucleic acids and proteins
 Chapter 11 Part A: Metabolism: The synthesis of nucleic acids and proteins I. Synthesis of DNA = REPLICATION A. Components of DNA (Fig. 11-1) 1. Composed of 4 different nucleotides that are joined by the
Chapter 11 Part A: Metabolism: The synthesis of nucleic acids and proteins I. Synthesis of DNA = REPLICATION A. Components of DNA (Fig. 11-1) 1. Composed of 4 different nucleotides that are joined by the
Lecture 21 Regulation of transcription
 Lecture 21 Regulation of transcription Key learning goals: Understand what a promoter sequence is Understand the role of sigma factors in prokaryotic RNAP initiation Understand what an open complex is,
Lecture 21 Regulation of transcription Key learning goals: Understand what a promoter sequence is Understand the role of sigma factors in prokaryotic RNAP initiation Understand what an open complex is,
Profile HMMs. 2/10/05 CAP5510/CGS5166 (Lec 10) 1 START STATE 1 STATE 2 STATE 3 STATE 4 STATE 5 STATE 6 END
 Profile HMMs START STATE 1 STATE 2 STATE 3 STATE 4 STATE 5 STATE 6 END 2/10/05 CAP5510/CGS5166 (Lec 10) 1 Profile HMMs with InDels Insertions Deletions Insertions & Deletions DELETE 1 DELETE 2 DELETE 3
Profile HMMs START STATE 1 STATE 2 STATE 3 STATE 4 STATE 5 STATE 6 END 2/10/05 CAP5510/CGS5166 (Lec 10) 1 Profile HMMs with InDels Insertions Deletions Insertions & Deletions DELETE 1 DELETE 2 DELETE 3
Transcription Regulation And Gene Expression in Eukaryotes FS 2016 Graduate Course G2 P Matthias and RG Clerc Pharmazentrum Hörsaal 2 16h15-18h00
 Transcription Regulation And Gene Expression in Eukaryotes FS 2016 Graduate Course G2 P Matthias and RG Clerc Pharmazentrum Hörsaal 2 16h15-18h00 The general problem RG Clerc March 2, 2016 RNA Transcription
Transcription Regulation And Gene Expression in Eukaryotes FS 2016 Graduate Course G2 P Matthias and RG Clerc Pharmazentrum Hörsaal 2 16h15-18h00 The general problem RG Clerc March 2, 2016 RNA Transcription
Chapter 3. DNA, RNA, and Protein Synthesis
 Chapter 3. DNA, RNA, and Protein Synthesis 4. Transcription Gene Expression Regulatory region (promoter) 5 flanking region Upstream region Coding region 3 flanking region Downstream region Transcription
Chapter 3. DNA, RNA, and Protein Synthesis 4. Transcription Gene Expression Regulatory region (promoter) 5 flanking region Upstream region Coding region 3 flanking region Downstream region Transcription
FUNCTIONAL BIOINFORMATICS
 Molecular Biology-2018 1 FUNCTIONAL BIOINFORMATICS PREDICTING THE FUNCTION OF AN UNKNOWN PROTEIN Suppose you have found the amino acid sequence of an unknown protein and wish to find its potential function.
Molecular Biology-2018 1 FUNCTIONAL BIOINFORMATICS PREDICTING THE FUNCTION OF AN UNKNOWN PROTEIN Suppose you have found the amino acid sequence of an unknown protein and wish to find its potential function.
Annotating the Genome (H)
 Annotating the Genome (H) Annotation principles (H1) What is annotation? In general: annotation = explanatory note* What could be useful as an annotation of a DNA sequence? an amino acid sequence? What
Annotating the Genome (H) Annotation principles (H1) What is annotation? In general: annotation = explanatory note* What could be useful as an annotation of a DNA sequence? an amino acid sequence? What
Transcription. Manzur Ali PP, DBT,M.E.S College,Marampally
 Transcription Manzur Ali PP, DBT,M.E.S College,Marampally manzursir@gmail.com RNA transcription is actively regulated Not all DNA is transcribed in a given cell (less than 50% even in prokaryotes) For
Transcription Manzur Ali PP, DBT,M.E.S College,Marampally manzursir@gmail.com RNA transcription is actively regulated Not all DNA is transcribed in a given cell (less than 50% even in prokaryotes) For
Transcription in Prokaryotes. Jörg Bungert, PhD Phone:
 Transcription in Prokaryotes Jörg Bungert, PhD Phone: 352-273-8098 Email: jbungert@ufl.edu Objectives Understand the basic mechanism of transcription. Know the function of promoter elements and associating
Transcription in Prokaryotes Jörg Bungert, PhD Phone: 352-273-8098 Email: jbungert@ufl.edu Objectives Understand the basic mechanism of transcription. Know the function of promoter elements and associating
CLASS 3.5: 03/29/07 EUKARYOTIC TRANSCRIPTION I: PROMOTERS AND ENHANCERS
 CLASS 3.5: 03/29/07 EUKARYOTIC TRANSCRIPTION I: PROMOTERS AND ENHANCERS A. Promoters and Polymerases (RNA pols): 1. General characteristics - Initiation of transcription requires a. Transcription factors
CLASS 3.5: 03/29/07 EUKARYOTIC TRANSCRIPTION I: PROMOTERS AND ENHANCERS A. Promoters and Polymerases (RNA pols): 1. General characteristics - Initiation of transcription requires a. Transcription factors
Eukaryotic & Prokaryotic Transcription. RNA polymerases
 Eukaryotic & Prokaryotic Transcription RNA polymerases RNA Polymerases A. E. coli RNA polymerase 1. core enzyme = ββ'(α)2 has catalytic activity but cannot recognize start site of transcription ~500,000
Eukaryotic & Prokaryotic Transcription RNA polymerases RNA Polymerases A. E. coli RNA polymerase 1. core enzyme = ββ'(α)2 has catalytic activity but cannot recognize start site of transcription ~500,000
Chapter 17 Lecture. Concepts of Genetics. Tenth Edition. Regulation of Gene Expression in Eukaryotes
 Chapter 17 Lecture Concepts of Genetics Tenth Edition Regulation of Gene Expression in Eukaryotes Chapter Contents 17.1 Eukaryotic Gene Regulation Can Occur at Any of the Steps Leading from DNA to Protein
Chapter 17 Lecture Concepts of Genetics Tenth Edition Regulation of Gene Expression in Eukaryotes Chapter Contents 17.1 Eukaryotic Gene Regulation Can Occur at Any of the Steps Leading from DNA to Protein
SIBC504: TRANSCRIPTION & RNA PROCESSING Assistant Professor Dr. Chatchawan Srisawat
 SIBC504: TRANSCRIPTION & RNA PROCESSING Assistant Professor Dr. Chatchawan Srisawat TRANSCRIPTION: AN OVERVIEW Transcription: the synthesis of a single-stranded RNA from a doublestranded DNA template.
SIBC504: TRANSCRIPTION & RNA PROCESSING Assistant Professor Dr. Chatchawan Srisawat TRANSCRIPTION: AN OVERVIEW Transcription: the synthesis of a single-stranded RNA from a doublestranded DNA template.
TIGR THE INSTITUTE FOR GENOMIC RESEARCH
 Introduction to Genome Annotation: Overview of What You Will Learn This Week C. Robin Buell May 21, 2007 Types of Annotation Structural Annotation: Defining genes, boundaries, sequence motifs e.g. ORF,
Introduction to Genome Annotation: Overview of What You Will Learn This Week C. Robin Buell May 21, 2007 Types of Annotation Structural Annotation: Defining genes, boundaries, sequence motifs e.g. ORF,
We can now identify three major pathways of information flow in the cell (in replication, information passes from one DNA molecule to other DNA
 1 We can now identify three major pathways of information flow in the cell (in replication, information passes from one DNA molecule to other DNA molecules; in transcription, information passes from DNA
1 We can now identify three major pathways of information flow in the cell (in replication, information passes from one DNA molecule to other DNA molecules; in transcription, information passes from DNA
RNA POLYMERASE FUNCTIONS E-BOOK
 08 March, 2018 RNA POLYMERASE 1 2 3 FUNCTIONS E-BOOK Document Filetype: PDF 431.06 KB 0 RNA POLYMERASE 1 2 3 FUNCTIONS E-BOOK It catalyzes the transcription of DNA to synthesize precursors of mrna and
08 March, 2018 RNA POLYMERASE 1 2 3 FUNCTIONS E-BOOK Document Filetype: PDF 431.06 KB 0 RNA POLYMERASE 1 2 3 FUNCTIONS E-BOOK It catalyzes the transcription of DNA to synthesize precursors of mrna and
DNA Transcription. Visualizing Transcription. The Transcription Process
 DNA Transcription By: Suzanne Clancy, Ph.D. 2008 Nature Education Citation: Clancy, S. (2008) DNA transcription. Nature Education 1(1) If DNA is a book, then how is it read? Learn more about the DNA transcription
DNA Transcription By: Suzanne Clancy, Ph.D. 2008 Nature Education Citation: Clancy, S. (2008) DNA transcription. Nature Education 1(1) If DNA is a book, then how is it read? Learn more about the DNA transcription
Year III Pharm.D Dr. V. Chitra
 Year III Pharm.D Dr. V. Chitra 1 Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Only one strand of DNA serves
Year III Pharm.D Dr. V. Chitra 1 Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Only one strand of DNA serves
RNA synthesis/transcription I Biochemistry 302. February 6, 2004 Bob Kelm
 RNA synthesis/transcription I Biochemistry 302 February 6, 2004 Bob Kelm Overview of RNA classes Messenger RNA (mrna) Encodes protein Relatively short half-life ( 3 min in E. coli, 30 min in eukaryotic
RNA synthesis/transcription I Biochemistry 302 February 6, 2004 Bob Kelm Overview of RNA classes Messenger RNA (mrna) Encodes protein Relatively short half-life ( 3 min in E. coli, 30 min in eukaryotic
Bioinformatics. Lecturer: Antinisca Di Marco Tutor: Francesco Gallo
 Bioinformatics Lecturer: Antinisca Di Marco Tutor: Francesco Gallo E mails: nome.cognome@univaq.it For appointment Di Marco: Tuersday 3.30 4:30 p.m. Friday 10:00-11:00 a.m. please ask appointment via email
Bioinformatics Lecturer: Antinisca Di Marco Tutor: Francesco Gallo E mails: nome.cognome@univaq.it For appointment Di Marco: Tuersday 3.30 4:30 p.m. Friday 10:00-11:00 a.m. please ask appointment via email
Genetic Parts in Bacterial Gene Expression
 Genetic Parts in Bacterial Gene Expression The Focus Promoters Operators Transcription Factors Transcriptional Terminators Ribosome Binding Sites An Abstract Annotation We ll Pay Particular Attention To:
Genetic Parts in Bacterial Gene Expression The Focus Promoters Operators Transcription Factors Transcriptional Terminators Ribosome Binding Sites An Abstract Annotation We ll Pay Particular Attention To:
30 Gene expression: Transcription
 30 Gene expression: Transcription Gene structure. o Exons coding region of DNA. o Introns non-coding region of DNA. o Introns are interspersed between exons of a single gene. o Promoter region helps enzymes
30 Gene expression: Transcription Gene structure. o Exons coding region of DNA. o Introns non-coding region of DNA. o Introns are interspersed between exons of a single gene. o Promoter region helps enzymes
There are four major types of introns. Group I introns, found in some rrna genes, are self-splicing: they can catalyze their own removal.
 1 2 Continuous genes - Intron: Many eukaryotic genes contain coding regions called exons and noncoding regions called intervening sequences or introns. The average human gene contains from eight to nine
1 2 Continuous genes - Intron: Many eukaryotic genes contain coding regions called exons and noncoding regions called intervening sequences or introns. The average human gene contains from eight to nine
The discovery of the role of RNA RNA structure, synthesis and function
 Central Dogma The discovery of the role of RNA RNA structure, synthesis and function! Fundamental observations in genetics!! Genes are located in nuclei (in eukaryotes)!! Polypeptides are synthesised in
Central Dogma The discovery of the role of RNA RNA structure, synthesis and function! Fundamental observations in genetics!! Genes are located in nuclei (in eukaryotes)!! Polypeptides are synthesised in
SCBC203 Gene Expression. Assoc. Prof. Rutaiwan Tohtong Department of Biochemistry Faculty of Science PR318
 SCBC203 Gene Expression Assoc. Prof. Rutaiwan Tohtong Department of Biochemistry Faculty of Science PR318 Rutaiwan.toh@mahidol.ac.th 1 Gene Expression Gene expression is a process where by the genetic
SCBC203 Gene Expression Assoc. Prof. Rutaiwan Tohtong Department of Biochemistry Faculty of Science PR318 Rutaiwan.toh@mahidol.ac.th 1 Gene Expression Gene expression is a process where by the genetic
DNA Topoisomerases relieve the supercoiling stress ahead of the fork
 DNA Topoisomerases relieve the supercoiling stress ahead of the fork Tw 1) T w : # of turns around the central axis 2) W r : # of times the double helix crosses itself 3) Linking Number: L k = T w + W
DNA Topoisomerases relieve the supercoiling stress ahead of the fork Tw 1) T w : # of turns around the central axis 2) W r : # of times the double helix crosses itself 3) Linking Number: L k = T w + W
Multiple choice questions (numbers in brackets indicate the number of correct answers)
 1 February 15, 2013 Multiple choice questions (numbers in brackets indicate the number of correct answers) 1. Which of the following statements are not true Transcriptomes consist of mrnas Proteomes consist
1 February 15, 2013 Multiple choice questions (numbers in brackets indicate the number of correct answers) 1. Which of the following statements are not true Transcriptomes consist of mrnas Proteomes consist
EECS730: Introduction to Bioinformatics
 EECS730: Introduction to Bioinformatics Lecture 08: Gene finding aatgcatgcggctatgctaatgcatgcggctatgctaagctgggatccgatgacaatgcatgcggctatgctaatgcatgcggc tatgcaagctgggatccgatgactatgctaagctgggatccgatgacaatgcatgcggctatgctaatgaatggtcttgggatt
EECS730: Introduction to Bioinformatics Lecture 08: Gene finding aatgcatgcggctatgctaatgcatgcggctatgctaagctgggatccgatgacaatgcatgcggctatgctaatgcatgcggc tatgcaagctgggatccgatgactatgctaagctgggatccgatgacaatgcatgcggctatgctaatgaatggtcttgggatt
CSE/Beng/BIMM 182: Biological Data Analysis. Instructor: Vineet Bafna TA: Nitin Udpa
 CSE/Beng/BIMM 182: Biological Data Analysis Instructor: Vineet Bafna TA: Nitin Udpa Today We will explore the syllabus through a series of questions? Please ASK All logistical information will be given
CSE/Beng/BIMM 182: Biological Data Analysis Instructor: Vineet Bafna TA: Nitin Udpa Today We will explore the syllabus through a series of questions? Please ASK All logistical information will be given
Transcription. DNA to RNA
 Transcription from DNA to RNA The Central Dogma of Molecular Biology replication DNA RNA Protein transcription translation Why call it transcription and translation? transcription is such a direct copy
Transcription from DNA to RNA The Central Dogma of Molecular Biology replication DNA RNA Protein transcription translation Why call it transcription and translation? transcription is such a direct copy
Computational gene finding
 Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) Lec 1 Lec 2 Lec 3 The biological context Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) Lec 1 Lec 2 Lec 3 The biological context Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
Gene Identification in silico
 Gene Identification in silico Nita Parekh, IIIT Hyderabad Presented at National Seminar on Bioinformatics and Functional Genomics, at Bioinformatics centre, Pondicherry University, Feb 15 17, 2006. Introduction
Gene Identification in silico Nita Parekh, IIIT Hyderabad Presented at National Seminar on Bioinformatics and Functional Genomics, at Bioinformatics centre, Pondicherry University, Feb 15 17, 2006. Introduction
Answers to Module 1. An obligate aerobe is an organism that has an absolute requirement of oxygen for growth.
 Answers to Module 1 Short Answers 1) What is an obligate aerobe? An obligate aerobe is an organism that has an absolute requirement of oxygen for growth. What about facultative anaerobe? 2) Distinguish
Answers to Module 1 Short Answers 1) What is an obligate aerobe? An obligate aerobe is an organism that has an absolute requirement of oxygen for growth. What about facultative anaerobe? 2) Distinguish
RNA Metabolism Chap 26, part I
 RNA Metabolism Chap 26, part I mrna (selective and regulated) trna rrna other (specialized) RNAs (eukaryotes!!!) processing transcriptome (Surprisingly, much of your genome is transcribed!) RNA is the
RNA Metabolism Chap 26, part I mrna (selective and regulated) trna rrna other (specialized) RNAs (eukaryotes!!!) processing transcriptome (Surprisingly, much of your genome is transcribed!) RNA is the
Identification of individual motifs on the genome scale. Some slides are from Mayukh Bhaowal
 Identification of individual motifs on the genome scale Some slides are from Mayukh Bhaowal Two papers Nature 423, 241-254 (15 May 2003) Sequencing and comparison of yeast species to identify genes and
Identification of individual motifs on the genome scale Some slides are from Mayukh Bhaowal Two papers Nature 423, 241-254 (15 May 2003) Sequencing and comparison of yeast species to identify genes and
Transcription Eukaryotic Cells
 Transcription Eukaryotic Cells Packet #20 1 Introduction Transcription is the process in which genetic information, stored in a strand of DNA (gene), is copied into a strand of RNA. Protein-encoding genes
Transcription Eukaryotic Cells Packet #20 1 Introduction Transcription is the process in which genetic information, stored in a strand of DNA (gene), is copied into a strand of RNA. Protein-encoding genes
Chapter 8. Microbial Genetics. Lectures prepared by Christine L. Case. Copyright 2010 Pearson Education, Inc.
 Chapter 8 Microbial Genetics Lectures prepared by Christine L. Case Structure and Function of Genetic Material Learning Objectives 8-1 Define genetics, genome, chromosome, gene, genetic code, genotype,
Chapter 8 Microbial Genetics Lectures prepared by Christine L. Case Structure and Function of Genetic Material Learning Objectives 8-1 Define genetics, genome, chromosome, gene, genetic code, genotype,
This place covers: Methods or systems for genetic or protein-related data processing in computational molecular biology.
 G16B BIOINFORMATICS, i.e. INFORMATION AND COMMUNICATION TECHNOLOGY [ICT] SPECIALLY ADAPTED FOR GENETIC OR PROTEIN-RELATED DATA PROCESSING IN COMPUTATIONAL MOLECULAR BIOLOGY Methods or systems for genetic
G16B BIOINFORMATICS, i.e. INFORMATION AND COMMUNICATION TECHNOLOGY [ICT] SPECIALLY ADAPTED FOR GENETIC OR PROTEIN-RELATED DATA PROCESSING IN COMPUTATIONAL MOLECULAR BIOLOGY Methods or systems for genetic
Genome-Wide Survey of MicroRNA - Transcription Factor Feed-Forward Regulatory Circuits in Human. Supporting Information
 Genome-Wide Survey of MicroRNA - Transcription Factor Feed-Forward Regulatory Circuits in Human Angela Re #, Davide Corá #, Daniela Taverna and Michele Caselle # equal contribution * corresponding author,
Genome-Wide Survey of MicroRNA - Transcription Factor Feed-Forward Regulatory Circuits in Human Angela Re #, Davide Corá #, Daniela Taverna and Michele Caselle # equal contribution * corresponding author,
CS313 Exercise 1 Cover Page Fall 2017
 CS313 Exercise 1 Cover Page Fall 2017 Due by the start of class on Monday, September 18, 2017. Name(s): In the TIME column, please estimate the time you spent on the parts of this exercise. Please try
CS313 Exercise 1 Cover Page Fall 2017 Due by the start of class on Monday, September 18, 2017. Name(s): In the TIME column, please estimate the time you spent on the parts of this exercise. Please try
Transcription is the first step of gene expression, in which a particular segment of DNA is copied into RNA by the enzyme, RNA polymerase.
 Transcription in Bacteria Transcription in Bacteria Transcription in Bacteria Transcription is the first step of gene expression, in which a particular segment of DNA is copied into RNA by the enzyme,
Transcription in Bacteria Transcription in Bacteria Transcription in Bacteria Transcription is the first step of gene expression, in which a particular segment of DNA is copied into RNA by the enzyme,
In 1996, the genome of Saccharomyces cerevisiae was completed due to the work of
 Summary: Kellis, M. et al. Nature 423,241-253. Background In 1996, the genome of Saccharomyces cerevisiae was completed due to the work of approximately 600 scientists world-wide. This group of researchers
Summary: Kellis, M. et al. Nature 423,241-253. Background In 1996, the genome of Saccharomyces cerevisiae was completed due to the work of approximately 600 scientists world-wide. This group of researchers
Computational Methods for Protein Structure Prediction and Fold Recognition... 1 I. Cymerman, M. Feder, M. PawŁowski, M.A. Kurowski, J.M.
 Contents Computational Methods for Protein Structure Prediction and Fold Recognition........................... 1 I. Cymerman, M. Feder, M. PawŁowski, M.A. Kurowski, J.M. Bujnicki 1 Primary Structure Analysis...................
Contents Computational Methods for Protein Structure Prediction and Fold Recognition........................... 1 I. Cymerman, M. Feder, M. PawŁowski, M.A. Kurowski, J.M. Bujnicki 1 Primary Structure Analysis...................
Outline. Gene Finding Questions. Recap: Prokaryotic gene finding Eukaryotic gene finding The human gene complement Regulation
 Tues, Nov 29: Gene Finding 1 Online FCE s: Thru Dec 12 Thurs, Dec 1: Gene Finding 2 Tues, Dec 6: PS5 due Project presentations 1 (see course web site for schedule) Thurs, Dec 8 Final papers due Project
Tues, Nov 29: Gene Finding 1 Online FCE s: Thru Dec 12 Thurs, Dec 1: Gene Finding 2 Tues, Dec 6: PS5 due Project presentations 1 (see course web site for schedule) Thurs, Dec 8 Final papers due Project
Synthetic cells: do bacteria need all its genes? No.
 NO NEED TO REFER TO THE SLIDES. بسم هللا الرحمن الرحيم Do we need all the non coding regions of the DNA? Two weeks ago, they discovered that the genome of a plant is very small (recall that plant genome
NO NEED TO REFER TO THE SLIDES. بسم هللا الرحمن الرحيم Do we need all the non coding regions of the DNA? Two weeks ago, they discovered that the genome of a plant is very small (recall that plant genome
Introduction and History of Genome Modification. Adam Clore, PhD Director, Synthetic Biology Design
 Introduction and History of Genome Modification Adam Clore, PhD Director, Synthetic Biology Design Early Non-site Directed Genome Modification Homologous recombination in yeast TARGET GENE 5 Arm URA3 3
Introduction and History of Genome Modification Adam Clore, PhD Director, Synthetic Biology Design Early Non-site Directed Genome Modification Homologous recombination in yeast TARGET GENE 5 Arm URA3 3
Big Idea 3C Basic Review
 Big Idea 3C Basic Review 1. A gene is a. A sequence of DNA that codes for a protein. b. A sequence of amino acids that codes for a protein. c. A sequence of codons that code for nucleic acids. d. The end
Big Idea 3C Basic Review 1. A gene is a. A sequence of DNA that codes for a protein. b. A sequence of amino acids that codes for a protein. c. A sequence of codons that code for nucleic acids. d. The end
Biology. Biology. Slide 1 of 39. End Show. Copyright Pearson Prentice Hall
 Biology Biology 1 of 39 12-3 RNA and Protein Synthesis 2 of 39 Essential Question What is transcription and translation and how do they take place? 3 of 39 12 3 RNA and Protein Synthesis Genes are coded
Biology Biology 1 of 39 12-3 RNA and Protein Synthesis 2 of 39 Essential Question What is transcription and translation and how do they take place? 3 of 39 12 3 RNA and Protein Synthesis Genes are coded
Biology. Biology. Slide 1 of 39. End Show. Copyright Pearson Prentice Hall
 Biology Biology 1 of 39 12-3 RNA and Protein Synthesis 2 of 39 12 3 RNA and Protein Synthesis Genes are coded DNA instructions that control the production of proteins. Genetic messages can be decoded by
Biology Biology 1 of 39 12-3 RNA and Protein Synthesis 2 of 39 12 3 RNA and Protein Synthesis Genes are coded DNA instructions that control the production of proteins. Genetic messages can be decoded by
Bis2A 12.2 Eukaryotic Transcription
 OpenStax-CNX module: m56061 1 Bis2A 12.2 Eukaryotic Transcription Mitch Singer Based on Eukaryotic Transcription by OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative
OpenStax-CNX module: m56061 1 Bis2A 12.2 Eukaryotic Transcription Mitch Singer Based on Eukaryotic Transcription by OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative
1. The diagram below shows an error in the transcription of a DNA template to messenger RNA (mrna).
 1. The diagram below shows an error in the transcription of a DNA template to messenger RNA (mrna). Which statement best describes the error shown in the diagram? (A) The mrna strand contains the uracil
1. The diagram below shows an error in the transcription of a DNA template to messenger RNA (mrna). Which statement best describes the error shown in the diagram? (A) The mrna strand contains the uracil
CHAPTERS , 17: Eukaryotic Genetics
 CHAPTERS 14.1 14.6, 17: Eukaryotic Genetics 1. Review the levels of DNA packing within the eukaryote nucleus. Label each level. (A similar diagram is on pg 188 of your textbook.) 2. How do the coding regions
CHAPTERS 14.1 14.6, 17: Eukaryotic Genetics 1. Review the levels of DNA packing within the eukaryote nucleus. Label each level. (A similar diagram is on pg 188 of your textbook.) 2. How do the coding regions
and Promoter Sequence Data
 : Combining Gene Expression and Promoter Sequence Data Outline 1. Motivation Functionally related genes cluster together genes sharing cis-elements cluster together transcriptional regulation is modular
: Combining Gene Expression and Promoter Sequence Data Outline 1. Motivation Functionally related genes cluster together genes sharing cis-elements cluster together transcriptional regulation is modular
Chromatographic Separation of the three forms of RNA Polymerase II.
 Chromatographic Separation of the three forms of RNA Polymerase II. α-amanitin α-amanitin bound to Pol II Function of the three enzymes. Yeast Pol II. RNA Polymerase Subunit Structures 10-7 Subunit structure.
Chromatographic Separation of the three forms of RNA Polymerase II. α-amanitin α-amanitin bound to Pol II Function of the three enzymes. Yeast Pol II. RNA Polymerase Subunit Structures 10-7 Subunit structure.
Genomics and Gene Recognition Genes and Blue Genes
 Genomics and Gene Recognition Genes and Blue Genes November 1, 2004 Prokaryotic Gene Structure prokaryotes are simplest free-living organisms studying prokaryotes can give us a sense what is the minimum
Genomics and Gene Recognition Genes and Blue Genes November 1, 2004 Prokaryotic Gene Structure prokaryotes are simplest free-living organisms studying prokaryotes can give us a sense what is the minimum
I. Gene Expression Figure 1: Central Dogma of Molecular Biology
 I. Gene Expression Figure 1: Central Dogma of Molecular Biology Central Dogma: Gene Expression: RNA Structure RNA nucleotides contain the pentose sugar Ribose instead of deoxyribose. Contain the bases
I. Gene Expression Figure 1: Central Dogma of Molecular Biology Central Dogma: Gene Expression: RNA Structure RNA nucleotides contain the pentose sugar Ribose instead of deoxyribose. Contain the bases
Transcription is the first stage of gene expression
 Transcription is the first stage of gene expression RNA synthesis is catalyzed by RNA polymerase, which pries the DNA strands apart and hooks together the RNA nucleotides The RNA is complementary to the
Transcription is the first stage of gene expression RNA synthesis is catalyzed by RNA polymerase, which pries the DNA strands apart and hooks together the RNA nucleotides The RNA is complementary to the
Chapter 12. DNA TRANSCRIPTION and TRANSLATION
 Chapter 12 DNA TRANSCRIPTION and TRANSLATION 12-3 RNA and Protein Synthesis WARM UP What are proteins? Where do they come from? From DNA to RNA to Protein DNA in our cells carry the instructions for making
Chapter 12 DNA TRANSCRIPTION and TRANSLATION 12-3 RNA and Protein Synthesis WARM UP What are proteins? Where do they come from? From DNA to RNA to Protein DNA in our cells carry the instructions for making
Chapter 24: Promoters and Enhancers
 Chapter 24: Promoters and Enhancers A typical gene transcribed by RNA polymerase II has a promoter that usually extends upstream from the site where transcription is initiated the (#1) of transcription
Chapter 24: Promoters and Enhancers A typical gene transcribed by RNA polymerase II has a promoter that usually extends upstream from the site where transcription is initiated the (#1) of transcription
Computational Biology I LSM5191 (2003/4)
 Computational Biology I LSM5191 (2003/4) Aylwin Ng, D.Phil Lecture Notes: Transcriptome: Molecular Biology of Gene Expression I Flow of information: DNA to polypeptide DNA Start Exon1 Intron Exon2 Termination
Computational Biology I LSM5191 (2003/4) Aylwin Ng, D.Phil Lecture Notes: Transcriptome: Molecular Biology of Gene Expression I Flow of information: DNA to polypeptide DNA Start Exon1 Intron Exon2 Termination
NCBI & Other Genome Databases. BME 110/BIOL 181 CompBio Tools
 NCBI & Other Genome Databases BME 110/BIOL 181 CompBio Tools Todd Lowe March 31, 2011 Admin Reading Dummies Ch 3 Assigned Review: "The impact of next-generation sequencing technology on genetics" by E.
NCBI & Other Genome Databases BME 110/BIOL 181 CompBio Tools Todd Lowe March 31, 2011 Admin Reading Dummies Ch 3 Assigned Review: "The impact of next-generation sequencing technology on genetics" by E.
Overview of Transcription
 Overview of Transcription The general process is similar in prokaryotes and eukaryotes. Three phases: - Initiation The word gene was coined in 1909 (W. Johannsen). The central dogma (1950s). - Elongation
Overview of Transcription The general process is similar in prokaryotes and eukaryotes. Three phases: - Initiation The word gene was coined in 1909 (W. Johannsen). The central dogma (1950s). - Elongation
Basi s c i Fea e tu t re r s s of f R NA N Sy S nth t esi s s i s
 Transcription Dr.H.B.Mahesha, Yuvaraja s College, University of Mysore, Mysuru. It is the process of transcribing or making a copy of Genetic information stored in a DNA strand into a Complementary strand
Transcription Dr.H.B.Mahesha, Yuvaraja s College, University of Mysore, Mysuru. It is the process of transcribing or making a copy of Genetic information stored in a DNA strand into a Complementary strand
Independent Study Guide The Blueprint of Life, from DNA to Protein (Chapter 7)
 Independent Study Guide The Blueprint of Life, from DNA to Protein (Chapter 7) I. General Principles (Chapter 7 introduction) a. Morse code distinct series of dots and dashes encode the 26 letters of the
Independent Study Guide The Blueprint of Life, from DNA to Protein (Chapter 7) I. General Principles (Chapter 7 introduction) a. Morse code distinct series of dots and dashes encode the 26 letters of the
Genome annotation. Erwin Datema (2011) Sandra Smit (2012, 2013)
 Genome annotation Erwin Datema (2011) Sandra Smit (2012, 2013) Genome annotation AGACAAAGATCCGCTAAATTAAATCTGGACTTCACATATTGAAGTGATATCACACGTTTCTCTAAT AATCTCCTCACAATATTATGTTTGGGATGAACTTGTCGTGATTTGCCATTGTAGCAATCACTTGAA
Genome annotation Erwin Datema (2011) Sandra Smit (2012, 2013) Genome annotation AGACAAAGATCCGCTAAATTAAATCTGGACTTCACATATTGAAGTGATATCACACGTTTCTCTAAT AATCTCCTCACAATATTATGTTTGGGATGAACTTGTCGTGATTTGCCATTGTAGCAATCACTTGAA
Chapter 6: Transcription and RNA Processing in Eukaryotes
 3. Basic Genetics Plant Molecular Biology Chapter 6: Transcription and RNA Processing in Eukaryotes - Genetic organization in eukaryote - Transcription in eukaryote - - RNA processing in eukaryote - Translation
3. Basic Genetics Plant Molecular Biology Chapter 6: Transcription and RNA Processing in Eukaryotes - Genetic organization in eukaryote - Transcription in eukaryote - - RNA processing in eukaryote - Translation
The Genetic Code and Transcription. Chapter 12 Honors Genetics Ms. Susan Chabot
 The Genetic Code and Transcription Chapter 12 Honors Genetics Ms. Susan Chabot TRANSCRIPTION Copy SAME language DNA to RNA Nucleic Acid to Nucleic Acid TRANSLATION Copy DIFFERENT language RNA to Amino
The Genetic Code and Transcription Chapter 12 Honors Genetics Ms. Susan Chabot TRANSCRIPTION Copy SAME language DNA to RNA Nucleic Acid to Nucleic Acid TRANSLATION Copy DIFFERENT language RNA to Amino
Yesterday s Picture UNIT 3B
 Warm-Up Plasmids are circular pieces of DNA which bacterial cells are able to take up from the environment, then replicate and transcribe. Eukaryotic cells, by contrast, contain large, linear (non-circular)
Warm-Up Plasmids are circular pieces of DNA which bacterial cells are able to take up from the environment, then replicate and transcribe. Eukaryotic cells, by contrast, contain large, linear (non-circular)
MODULE TSS2: SEQUENCE ALIGNMENTS (ADVANCED)
 MODULE TSS2: SEQUENCE ALIGNMENTS (ADVANCED) Lesson Plan: Title MEG LAAKSO AND JAMIE SIDERS Identifying the TSS for a gene in D. eugracilis using sequence alignment with the D. melanogaster ortholog Objectives
MODULE TSS2: SEQUENCE ALIGNMENTS (ADVANCED) Lesson Plan: Title MEG LAAKSO AND JAMIE SIDERS Identifying the TSS for a gene in D. eugracilis using sequence alignment with the D. melanogaster ortholog Objectives
7.03, 2005, Lecture 20 EUKARYOTIC GENES AND GENOMES I
 7.03, 2005, Lecture 20 EUKARYOTIC GENES AND GENOMES I For the last several lectures we have been looking at how one can manipulate prokaryotic genomes and how prokaryotic genes are regulated. In the next
7.03, 2005, Lecture 20 EUKARYOTIC GENES AND GENOMES I For the last several lectures we have been looking at how one can manipulate prokaryotic genomes and how prokaryotic genes are regulated. In the next
3.1 Storing Information. Polypeptides. Protein Structure. Helical Secondary Structure. Types of Protein Structure. Molecular Biology Fourth Edition
 Lecture PowerPoint to accompany Molecular Biology Fourth Edition Robert F. Weaver Chapter 3 An Introduction to Gene Function 3.1 Storing Information Producing a protein from DNA information involves both
Lecture PowerPoint to accompany Molecular Biology Fourth Edition Robert F. Weaver Chapter 3 An Introduction to Gene Function 3.1 Storing Information Producing a protein from DNA information involves both
XII International PhD Workshop OWD 2010, October 2010
 XII International PhD Workshop OWD 2010, 23 26 October 2010 Data Mining in Molecular Biology: A Journey From Raw Datasets to Biologically Meaningful Conclusions Roman Jaksik, Silesian University of Technology
XII International PhD Workshop OWD 2010, 23 26 October 2010 Data Mining in Molecular Biology: A Journey From Raw Datasets to Biologically Meaningful Conclusions Roman Jaksik, Silesian University of Technology
BS 50 Genetics and Genomics Week of Oct 24
 BS 50 Genetics and Genomics Week of Oct 24 Additional Practice Problems for Section Question 1: The following table contains a list of statements that apply to replication, transcription, both, or neither.
BS 50 Genetics and Genomics Week of Oct 24 Additional Practice Problems for Section Question 1: The following table contains a list of statements that apply to replication, transcription, both, or neither.
The Central Dogma of Molecular Biology
 The Central Dogma of Molecular Biology In the Central Dogma of Molecular Biology, this process occurs when mrna is made from DNA? A. TranscripBon B. TranslaBon C. ReplicaBon 1 DNA: The ultimate instruction
The Central Dogma of Molecular Biology In the Central Dogma of Molecular Biology, this process occurs when mrna is made from DNA? A. TranscripBon B. TranslaBon C. ReplicaBon 1 DNA: The ultimate instruction
DIFFERENT ASPECTS OF GENE REGULATION
 TARTU UNIVERSITY FACULTY OF BILOGY AND GEOGRAPHY DIFFERENT ASPECTS OF GENE REGULATION TARTU 2005 Sten Ilmjärv 2 TABLE OF CONTENT TABLE OF CONTENT...2 GLOSSARY...3 INTRODUCTION...4 1. THE GENE...5 2. GENE
TARTU UNIVERSITY FACULTY OF BILOGY AND GEOGRAPHY DIFFERENT ASPECTS OF GENE REGULATION TARTU 2005 Sten Ilmjärv 2 TABLE OF CONTENT TABLE OF CONTENT...2 GLOSSARY...3 INTRODUCTION...4 1. THE GENE...5 2. GENE
RNA Structure and the Versatility of RNA. Mitesh Shrestha
 RNA Structure and the Versatility of RNA Mitesh Shrestha Ribonucleic Acid (RNA) Nitrogenous Bases (Adenine, Uracil, Guanine, Cytosine) Ribose Sugar Ribonucleic Acid (RNA) Phosphate Group RNA world Hypothesis
RNA Structure and the Versatility of RNA Mitesh Shrestha Ribonucleic Acid (RNA) Nitrogenous Bases (Adenine, Uracil, Guanine, Cytosine) Ribose Sugar Ribonucleic Acid (RNA) Phosphate Group RNA world Hypothesis
The Nature of Genes. The Nature of Genes. Genes and How They Work. Chapter 15/16
 Genes and How They Work Chapter 15/16 The Nature of Genes Beadle and Tatum proposed the one gene one enzyme hypothesis. Today we know this as the one gene one polypeptide hypothesis. 2 The Nature of Genes
Genes and How They Work Chapter 15/16 The Nature of Genes Beadle and Tatum proposed the one gene one enzyme hypothesis. Today we know this as the one gene one polypeptide hypothesis. 2 The Nature of Genes
Class XII Chapter 6 Molecular Basis of Inheritance Biology
 Question 1: Group the following as nitrogenous bases and nucleosides: Adenine, Cytidine, Thymine, Guanosine, Uracil and Cytosine. Nitrogenous bases present in the list are adenine, thymine, uracil, and
Question 1: Group the following as nitrogenous bases and nucleosides: Adenine, Cytidine, Thymine, Guanosine, Uracil and Cytosine. Nitrogenous bases present in the list are adenine, thymine, uracil, and
DNA SEQUENCING BY SANGER METHOD
 DNA SEQUENCING BY SANGER METHOD First method described by Sanger and Coulson,1975 for DNA sequencing was called plus and minus. This method used E.coli DNA polymerase I and DNA ploymerase from bacteriophage
DNA SEQUENCING BY SANGER METHOD First method described by Sanger and Coulson,1975 for DNA sequencing was called plus and minus. This method used E.coli DNA polymerase I and DNA ploymerase from bacteriophage
Lecture 2: Central Dogma of Molecular Biology & Intro to Programming
 Lecture 2: Central Dogma of Molecular Biology & Intro to Programming Central Dogma of Molecular Biology Proteins: workhorse molecules of biological systems Proteins are synthesized from the genetic blueprints
Lecture 2: Central Dogma of Molecular Biology & Intro to Programming Central Dogma of Molecular Biology Proteins: workhorse molecules of biological systems Proteins are synthesized from the genetic blueprints
ECS 234: Genomic Data Integration ECS 234
 : Genomic Data Integration Heterogeneous Data Integration DNA Sequence Microarray Proteomics >gi 12004594 gb AF217406.1 Saccharomyces cerevisiae uridine nucleosidase (URH1) gene, complete cds ATGGAATCTGCTGATTTTTTTACCTCACGAAACTTATTAAAACAGATAATTTCCCTCATCTGCAAGGTTG
: Genomic Data Integration Heterogeneous Data Integration DNA Sequence Microarray Proteomics >gi 12004594 gb AF217406.1 Saccharomyces cerevisiae uridine nucleosidase (URH1) gene, complete cds ATGGAATCTGCTGATTTTTTTACCTCACGAAACTTATTAAAACAGATAATTTCCCTCATCTGCAAGGTTG
BIOINFORMATICS AND SYSTEM BIOLOGY (INTERNATIONAL PROGRAM)
 BIOINFORMATICS AND SYSTEM BIOLOGY (INTERNATIONAL PROGRAM) PROGRAM TITLE DEGREE TITLE Master of Science Program in Bioinformatics and System Biology (International Program) Master of Science (Bioinformatics
BIOINFORMATICS AND SYSTEM BIOLOGY (INTERNATIONAL PROGRAM) PROGRAM TITLE DEGREE TITLE Master of Science Program in Bioinformatics and System Biology (International Program) Master of Science (Bioinformatics
G-Quadruplex DNA Sequences Are Evolutionarily Conserved and Associated with Distinct Genomic Features in Saccharomyces cerevisiae
 G-Quadruplex DNA Sequences Are Evolutionarily Conserved and Associated with Distinct Genomic Features in Saccharomyces cerevisiae John A. Capra 1", Katrin Paeschke 2", Mona Singh 1 *, Virginia A. Zakian
G-Quadruplex DNA Sequences Are Evolutionarily Conserved and Associated with Distinct Genomic Features in Saccharomyces cerevisiae John A. Capra 1", Katrin Paeschke 2", Mona Singh 1 *, Virginia A. Zakian
Transcription and Post Transcript Modification
 Transcription and Post Transcript Modification You Should Be Able To 1. Describe transcription. 2. Compare and contrast eukaryotic + prokaryotic transcription. 3. Explain mrna processing in eukaryotes.
Transcription and Post Transcript Modification You Should Be Able To 1. Describe transcription. 2. Compare and contrast eukaryotic + prokaryotic transcription. 3. Explain mrna processing in eukaryotes.
Resources. How to Use This Presentation. Chapter 10. Objectives. Table of Contents. Griffith s Discovery of Transformation. Griffith s Experiments
 How to Use This Presentation To View the presentation as a slideshow with effects select View on the menu bar and click on Slide Show. To advance through the presentation, click the right-arrow key or
How to Use This Presentation To View the presentation as a slideshow with effects select View on the menu bar and click on Slide Show. To advance through the presentation, click the right-arrow key or
