Unified nomenclature for the winged helix/forkhead transcription factors
|
|
|
- Asher Farmer
- 6 years ago
- Views:
Transcription
1 CORRESPONDENCE Unified nomenclature for the winged helix/forkhead transcription factors Klaus H. Kaestner, 1,4,5 Walter Knöchel, 2,4 and Daniel E. Martínez 3,4,5 1 Department of Genetics, University of Pennsylvania School of Medicine, Philadelphia, Pennsylvania USA; 2 Abteilung Biochemie, Universität Ulm, D Ulm, Germany; 3 Department of Biology, Pomona College, Claremont, California USA The winged helix/forkhead class of transcription factors is characterized by a 100-amino-acid, monomeric DNAbinding domain. The structure of the DNA-binding domain of one of the class members, hepatocyte nuclear factor 3 (HNF3 ), in a complex with a DNA target has been solved (Clark et al. 1993). The DNA-binding domain folds into a variant of the helix turn helix motif and is made up of three helices and two characteristic large loops, or wings. Therefore, the DNA-binding motif has been named the winged helix DNA-binding domain. Over the past 9 years since the identification of the first member of this class, the Drosophila melanogaster gene Fork head, >100 members of this gene family have been identified (for review, see Kaufmann and Knöchel 1996) in species ranging from yeast to human. The rapid accumulation of sequences by many different laboratories has led to the use of multiple names and classification systems, making it very difficult to follow the literature and to name newly characterized winged helix/ forkhead transcription factors. This problem was recognized and discussed at the first International Meeting on Forkhead/Winged Helix Proteins, held in La Jolla, California, in November At that time a proposal was developed to standardize the nomenclature for these proteins. Fox (Forkhead box) was adopted as the unified symbol for all chordate winged helix/forkhead transcription factors. A winged helix/forkhead nomenclature committee was elected to implement this proposal, in consultation with the community at large. This final proposal has been endorsed by >20 scientists 1 as well as the Human and Mouse Gene Nomenclature Committees. 4 Fox Nomenclature Committee. 5 Corresponding authors. kaestner@mail.med.upenn.edu; FAX (215) dmartinez@pomona.edu; FAX (909) The Fox subclasses All Fox proteins contain the characteristic 100-aminoacid winged helix domain, that defines this class of transcription factors. Other portions of the Fox proteins, which encode, for instance, transactivation or trans-repression domains, are highly divergent. We have utilized phylogenetic analysis to delineate 15 subclasses for all known chordate Fox proteins. The analysis included chordate sequences obtained from GenBank and sequences submitted directly to the nomenclature committee. The Fox domains of the proteins were aligned using Clustal W (Thompson et al. 1994), and a neighborjoining tree was generated using PAUP* 4.0 (Swofford 1999); (Fig. 1). This phylogenetic tree will be updated regularly as new sequences are discovered and may be downloaded from html. A complete phylogenetic analysis of all known forkhead proteins will be published elsewhere (D. Martínez and J.E. Signorovitch, pers. comm.). Numbering Fox proteins were assigned to individual subclasses based on the phylogenetic analysis described above. Subclasses were designated by a letter, and within each subclass proteins were given an Arabic numeral. Therefore, the actual name of any Fox protein is Fox, subclass N, member X, or for example, Foxd3. Abbreviations for the chordate Fox proteins will contain all uppercase letters for human (e.g., FOXD3); only the first letter capitalized for mouse (e.g., Foxd3); and the first and subclass letters capitalized for all other chordates (e.g., FoxD3). Current assignments for the chordate Fox proteins are listed in Table 1, together with previously used names. Whenever possible we have assigned the same name to ortholog proteins from different species. In a few cases where the phylogenetic affinities were not well resolved by the tree in Figure 1 (e.g., Foxd1 and Foxd2), a within-class phylo- 1 The following scientists have endorsed the use of this nomenclature system: Frederic G. Barr, William Biggs, Peter Carlsson, James E. Darnell, Sven Enerbäck, Peter Gruss, Brigid Hogan, Andrew D. Hollenbach, Robert Hromas, Tsutomu Kume, Trish Labosky, Eseng Lai, Suzanne C. Li, Naoyuki Miura, Sally A. Moody, Sharon Plon, Hiroshi Sasaki, Günther Schütz, Mathias Treier, Malcolm Whitman, Jeffrey Whitsett, Stella Zannini, and Ken Zaret. 142 GENES & DEVELOPMENT 14: by Cold Spring Harbor Laboratory Press ISSN /00 $5.00;
2 Fox nomenclature system Figure 1. (Continued on p. 144) Neighbor-joining phylogeny of chordate Fox proteins based on the amino acid sequence of the Fox domain. The distance measure used was mean character difference. The tree was rooted using Homo QRF1 (AF086040) as the outgroup. The numbers in the interior branches are bootstrap percentages. For each protein we indicate the organism (genus), the name, the accession number, and the proposed Fox name. GENES & DEVELOPMENT 143
3 Kaestner et al. 144 GENES & DEVELOPMENT
4 Fox nomenclature system Table 1. A proposed nomenclature for chordate winged helix proteins Accession nos. submitted to the Fox Nomenclature Committee by W. Knöchel (*) and S.A. Sullivan and S. Molly (**). GENES & DEVELOPMENT 145
5 Kaestner et al. genetic analysis was performed using the full sequence of the proteins (data not shown). We have used lowercase letters to distinguish between virtually identical proteins (e.g., Foxa4a and Foxa4b), presumably derived from duplicated genes, a case commonly found in polyploid species like Xenopus laevis. Please note that the phylogenetic tree includes several proteins that have not received a Fox designation because as yet, their phylogenetic relationships remain unclear due to limited sequence information. Naming new sequences A new Fox protein is defined as a fully sequenced gene, cdna, or protein that belongs to the Fox family of proteins based on sequence homology of its winged helix DNA-binding domain. We have established a Fox Nomenclature web site ( fox.html) that provides a form for submitting protein sequences to the Fox Nomenclature Committee. We encourage investigators who have discovered new Fox sequences to submit them to the committee for assignment of the proper Fox name. These sequences will be kept confidential until publication of the sequence by the investigators. We recommend that this new system of nomenclature be used in all future publications. References Clark, K.L., E.D. Halay, E. Lai, and S.K. Burley Co-crystal structure of the HNF-3/fork head DNA-recognition motif resembles histone H5. Nature 364: Kaufmann, E. and W. Knöchel Five years on the wings of fork head. Mech. Dev. 57: Swofford, D.L PAUP*: Phylogenetic analysis using parsimony and other methods. Sinauer, Sunderland, MA. Thompson, J.D., D.G. Higgins, and T.J. Gibson CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighing, position specific gap penalties and weight matrix choice. Nucleic Acid Res. 22: GENES & DEVELOPMENT
Human KIR sequences 2003
 Immunogenetics (2003) 55:227 239 DOI 10.1007/s00251-003-0572-y ORIGINAL PAPER C. A. Garcia J. Robinson L. A. Guethlein P. Parham J. A. Madrigal S. G. E. Marsh Human KIR sequences 2003 Received: 17 March
Immunogenetics (2003) 55:227 239 DOI 10.1007/s00251-003-0572-y ORIGINAL PAPER C. A. Garcia J. Robinson L. A. Guethlein P. Parham J. A. Madrigal S. G. E. Marsh Human KIR sequences 2003 Received: 17 March
Reservoir of Bacterial Exotoxin Genes in the Environment
 Reservoir of Bacterial Exotoxin Genes in the Environment Veronica Casas, Joseph Magbanua, Gerico Sobrepeña, and Stanley Maloy * Center for Microbial Sciences San Diego State University 5500 Campanile Drive
Reservoir of Bacterial Exotoxin Genes in the Environment Veronica Casas, Joseph Magbanua, Gerico Sobrepeña, and Stanley Maloy * Center for Microbial Sciences San Diego State University 5500 Campanile Drive
Representation in Supervised Machine Learning Application to Biological Problems
 Representation in Supervised Machine Learning Application to Biological Problems Frank Lab Howard Hughes Medical Institute & Columbia University 2010 Robert Howard Langlois Hughes Medical Institute What
Representation in Supervised Machine Learning Application to Biological Problems Frank Lab Howard Hughes Medical Institute & Columbia University 2010 Robert Howard Langlois Hughes Medical Institute What
Identifying Regulatory Regions using Multiple Sequence Alignments
 Identifying Regulatory Regions using Multiple Sequence Alignments Prerequisites: BLAST Exercise: Detecting and Interpreting Genetic Homology. Resources: ClustalW is available at http://www.ebi.ac.uk/tools/clustalw2/index.html
Identifying Regulatory Regions using Multiple Sequence Alignments Prerequisites: BLAST Exercise: Detecting and Interpreting Genetic Homology. Resources: ClustalW is available at http://www.ebi.ac.uk/tools/clustalw2/index.html
Project Report. Genetic Assessment of the Sunda Pangolin (Manis javanica) Using Degraded and Fresh Samples
 Genetic Assessment of the Sunda Pangolin (Manis javanica) Using Degraded and Fresh Samples Project Report Đỗ Văn Thao 1, Nguyễn Văn Thành 1, Dương Thúy Hà 1, và Lê Đức Minh 1,2 1 Hanoi University of Science
Genetic Assessment of the Sunda Pangolin (Manis javanica) Using Degraded and Fresh Samples Project Report Đỗ Văn Thao 1, Nguyễn Văn Thành 1, Dương Thúy Hà 1, và Lê Đức Minh 1,2 1 Hanoi University of Science
Textbook Reading Guidelines
 Understanding Bioinformatics by Marketa Zvelebil and Jeremy Baum Last updated: May 1, 2009 Textbook Reading Guidelines Preface: Read the whole preface, and especially: For the students with Life Science
Understanding Bioinformatics by Marketa Zvelebil and Jeremy Baum Last updated: May 1, 2009 Textbook Reading Guidelines Preface: Read the whole preface, and especially: For the students with Life Science
NGS Approaches to Epigenomics
 I519 Introduction to Bioinformatics, 2013 NGS Approaches to Epigenomics Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Background: chromatin structure & DNA methylation Epigenomic
I519 Introduction to Bioinformatics, 2013 NGS Approaches to Epigenomics Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Background: chromatin structure & DNA methylation Epigenomic
Supplemental Data. Osakabe et al. (2013). Plant Cell /tpc
 Supplemental Figure 1. Phylogenetic analysis of KUP in various species. The amino acid sequences of KUPs from green algae and land plants were identified with a BLAST search and aligned using ClustalW
Supplemental Figure 1. Phylogenetic analysis of KUP in various species. The amino acid sequences of KUPs from green algae and land plants were identified with a BLAST search and aligned using ClustalW
Figure S1 Correlation in size of analogous introns in mouse and teleost Piccolo genes. Mouse intron size was plotted against teleost intron size for t
 Figure S1 Correlation in size of analogous introns in mouse and teleost Piccolo genes. Mouse intron size was plotted against teleost intron size for the pcloa genes of zebrafish, green spotted puffer (listed
Figure S1 Correlation in size of analogous introns in mouse and teleost Piccolo genes. Mouse intron size was plotted against teleost intron size for the pcloa genes of zebrafish, green spotted puffer (listed
Taxonomic status of the brown teal (Anas chlorotis) in Fiordland
 Taxonomic status of the brown teal (Anas chlorotis) in Fiordland Neil J Gemmell and Heather J Flint Department of Zoology University of Canterbury Private Bag 4800 Christchurch Published by Department
Taxonomic status of the brown teal (Anas chlorotis) in Fiordland Neil J Gemmell and Heather J Flint Department of Zoology University of Canterbury Private Bag 4800 Christchurch Published by Department
CS313 Exercise 1 Cover Page Fall 2017
 CS313 Exercise 1 Cover Page Fall 2017 Due by the start of class on Monday, September 18, 2017. Name(s): In the TIME column, please estimate the time you spent on the parts of this exercise. Please try
CS313 Exercise 1 Cover Page Fall 2017 Due by the start of class on Monday, September 18, 2017. Name(s): In the TIME column, please estimate the time you spent on the parts of this exercise. Please try
Following text taken from Suresh Kumar. Bioinformatics Web - Comprehensive educational resource on Bioinformatics. 6th May.2005
 Bioinformatics is the recording, annotation, storage, analysis, and searching/retrieval of nucleic acid sequence (genes and RNAs), protein sequence and structural information. This includes databases of
Bioinformatics is the recording, annotation, storage, analysis, and searching/retrieval of nucleic acid sequence (genes and RNAs), protein sequence and structural information. This includes databases of
In 1996, the genome of Saccharomyces cerevisiae was completed due to the work of
 Summary: Kellis, M. et al. Nature 423,241-253. Background In 1996, the genome of Saccharomyces cerevisiae was completed due to the work of approximately 600 scientists world-wide. This group of researchers
Summary: Kellis, M. et al. Nature 423,241-253. Background In 1996, the genome of Saccharomyces cerevisiae was completed due to the work of approximately 600 scientists world-wide. This group of researchers
RNA does not adopt the classic B-DNA helix conformation when it forms a self-complementary double helix
 Reason: RNA has ribose sugar ring, with a hydroxyl group (OH) If RNA in B-from conformation there would be unfavorable steric contact between the hydroxyl group, base, and phosphate backbone. RNA structure
Reason: RNA has ribose sugar ring, with a hydroxyl group (OH) If RNA in B-from conformation there would be unfavorable steric contact between the hydroxyl group, base, and phosphate backbone. RNA structure
Scoring Alignments. Genome 373 Genomic Informatics Elhanan Borenstein
 Scoring Alignments Genome 373 Genomic Informatics Elhanan Borenstein A quick review Course logistics Genomes (so many genomes) The computational bottleneck Python: Programs, input and output Number and
Scoring Alignments Genome 373 Genomic Informatics Elhanan Borenstein A quick review Course logistics Genomes (so many genomes) The computational bottleneck Python: Programs, input and output Number and
Genome-Scale Predictions of the Transcription Factor Binding Sites of Cys 2 His 2 Zinc Finger Proteins in Yeast June 17 th, 2005
 Genome-Scale Predictions of the Transcription Factor Binding Sites of Cys 2 His 2 Zinc Finger Proteins in Yeast June 17 th, 2005 John Brothers II 1,3 and Panayiotis V. Benos 1,2 1 Bioengineering and Bioinformatics
Genome-Scale Predictions of the Transcription Factor Binding Sites of Cys 2 His 2 Zinc Finger Proteins in Yeast June 17 th, 2005 John Brothers II 1,3 and Panayiotis V. Benos 1,2 1 Bioengineering and Bioinformatics
This place covers: Methods or systems for genetic or protein-related data processing in computational molecular biology.
 G16B BIOINFORMATICS, i.e. INFORMATION AND COMMUNICATION TECHNOLOGY [ICT] SPECIALLY ADAPTED FOR GENETIC OR PROTEIN-RELATED DATA PROCESSING IN COMPUTATIONAL MOLECULAR BIOLOGY Methods or systems for genetic
G16B BIOINFORMATICS, i.e. INFORMATION AND COMMUNICATION TECHNOLOGY [ICT] SPECIALLY ADAPTED FOR GENETIC OR PROTEIN-RELATED DATA PROCESSING IN COMPUTATIONAL MOLECULAR BIOLOGY Methods or systems for genetic
TIGR THE INSTITUTE FOR GENOMIC RESEARCH
 Introduction to Genome Annotation: Overview of What You Will Learn This Week C. Robin Buell May 21, 2007 Types of Annotation Structural Annotation: Defining genes, boundaries, sequence motifs e.g. ORF,
Introduction to Genome Annotation: Overview of What You Will Learn This Week C. Robin Buell May 21, 2007 Types of Annotation Structural Annotation: Defining genes, boundaries, sequence motifs e.g. ORF,
Enhancers. Activators and repressors of transcription
 Enhancers Can be >50 kb away from the gene they regulate. Can be upstream from a promoter, downstream from a promoter, within an intron, or even downstream of the final exon of a gene. Are often cell type
Enhancers Can be >50 kb away from the gene they regulate. Can be upstream from a promoter, downstream from a promoter, within an intron, or even downstream of the final exon of a gene. Are often cell type
RNA was extracted from midguts of adult females 24 hr after blood feeding on a healthy
 Supplemental Data Cell Host & Microbe, Volume 5 The STAT Pathway Mediates Late-Phase Immunity against Plasmodium in the Mosquito Anopheles gambiae Lalita Gupta, Alvaro Molina-Cruz, Sanjeev Kumar, Janneth
Supplemental Data Cell Host & Microbe, Volume 5 The STAT Pathway Mediates Late-Phase Immunity against Plasmodium in the Mosquito Anopheles gambiae Lalita Gupta, Alvaro Molina-Cruz, Sanjeev Kumar, Janneth
BIOINFORMATICS Introduction
 BIOINFORMATICS Introduction Mark Gerstein, Yale University bioinfo.mbb.yale.edu/mbb452a 1 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu What is Bioinformatics? (Molecular) Bio -informatics One idea
BIOINFORMATICS Introduction Mark Gerstein, Yale University bioinfo.mbb.yale.edu/mbb452a 1 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu What is Bioinformatics? (Molecular) Bio -informatics One idea
Genetic variation of captive green peafowl Pavo muticus in Thailand based on D-loop sequences
 Short Communication Genetic variation of captive green peafowl Pavo muticus in Thailand based on D-loop sequences AMPORN WIWEGWEAW* AND WINA MECKVICHAI Department of Biology, Faculty of Science, Chulalongkorn
Short Communication Genetic variation of captive green peafowl Pavo muticus in Thailand based on D-loop sequences AMPORN WIWEGWEAW* AND WINA MECKVICHAI Department of Biology, Faculty of Science, Chulalongkorn
Expression of the genome. Books: 1. Molecular biology of the gene: Watson et al 2. Genetics: Peter J. Russell
 Expression of the genome Books: 1. Molecular biology of the gene: Watson et al 2. Genetics: Peter J. Russell 1 Transcription 1. Francis Crick (1956) named the flow of information from DNA RNA protein the
Expression of the genome Books: 1. Molecular biology of the gene: Watson et al 2. Genetics: Peter J. Russell 1 Transcription 1. Francis Crick (1956) named the flow of information from DNA RNA protein the
VALLIAMMAI ENGINEERING COLLEGE
 VALLIAMMAI ENGINEERING COLLEGE SRM Nagar, Kattankulathur 603 203 DEPARTMENT OF COMPUTER SCIENCE AND ENGINEERING QUESTION BANK VII SEMESTER BM6005 BIO INFORMATICS Regulation 2013 Academic Year 2018-19 Prepared
VALLIAMMAI ENGINEERING COLLEGE SRM Nagar, Kattankulathur 603 203 DEPARTMENT OF COMPUTER SCIENCE AND ENGINEERING QUESTION BANK VII SEMESTER BM6005 BIO INFORMATICS Regulation 2013 Academic Year 2018-19 Prepared
Chapter 25: Regulating Eukaryotic Transcription The Ligand Responsive Activators
 Chapter 25: Regulating Eukaryotic Transcription The Ligand Responsive Activators At least 5 potential gene expression control points Superfamily of Gene Regulators Activation of gene structure Initiation
Chapter 25: Regulating Eukaryotic Transcription The Ligand Responsive Activators At least 5 potential gene expression control points Superfamily of Gene Regulators Activation of gene structure Initiation
Heme utilization in the Caenorhabditis elegans hypodermal cells is facilitated by hemeresponsive
 Supplemental Data Heme utilization in the Caenorhabditis elegans hypodermal cells is facilitated by hemeresponsive gene-2 Caiyong Chen 1, Tamika K. Samuel 1, Michael Krause 2, Harry A. Dailey 3, and Iqbal
Supplemental Data Heme utilization in the Caenorhabditis elegans hypodermal cells is facilitated by hemeresponsive gene-2 Caiyong Chen 1, Tamika K. Samuel 1, Michael Krause 2, Harry A. Dailey 3, and Iqbal
Introduction to Bioinformatics
 Introduction to Bioinformatics Changhui (Charles) Yan Old Main 401 F http://www.cs.usu.edu www.cs.usu.edu/~cyan 1 How Old Is The Discipline? "The term bioinformatics is a relatively recent invention, not
Introduction to Bioinformatics Changhui (Charles) Yan Old Main 401 F http://www.cs.usu.edu www.cs.usu.edu/~cyan 1 How Old Is The Discipline? "The term bioinformatics is a relatively recent invention, not
Chapter 14 Regulation of Transcription
 Chapter 14 Regulation of Transcription Cis-acting sequences Distance-independent cis-acting elements Dissecting regulatory elements Transcription factors Overview transcriptional regulation Transcription
Chapter 14 Regulation of Transcription Cis-acting sequences Distance-independent cis-acting elements Dissecting regulatory elements Transcription factors Overview transcriptional regulation Transcription
GREG GIBSON SPENCER V. MUSE
 A Primer of Genome Science ience THIRD EDITION TAGCACCTAGAATCATGGAGAGATAATTCGGTGAGAATTAAATGGAGAGTTGCATAGAGAACTGCGAACTG GREG GIBSON SPENCER V. MUSE North Carolina State University Sinauer Associates, Inc.
A Primer of Genome Science ience THIRD EDITION TAGCACCTAGAATCATGGAGAGATAATTCGGTGAGAATTAAATGGAGAGTTGCATAGAGAACTGCGAACTG GREG GIBSON SPENCER V. MUSE North Carolina State University Sinauer Associates, Inc.
Nature Genetics: doi: /ng.3556 INTEGRATED SUPPLEMENTARY FIGURE TEMPLATE. Supplementary Figure 1
 INTEGRATED SUPPLEMENTARY FIGURE TEMPLATE Supplementary Figure 1 REF6 expression in transgenic lines. (a,b) Expression of REF6 in REF6-HA ref6 and REF6ΔZnF-HA ref6 plants detected by RT qpcr (a) and immunoblot
INTEGRATED SUPPLEMENTARY FIGURE TEMPLATE Supplementary Figure 1 REF6 expression in transgenic lines. (a,b) Expression of REF6 in REF6-HA ref6 and REF6ΔZnF-HA ref6 plants detected by RT qpcr (a) and immunoblot
Supplemental Data. Bai et al. Plant Cell. (2012) /tpc A
 A B Unconserved Conserved AIF4 AIF2 AIF3 AIF1 UPB1 IBH1 Supplemental Figure 1. IBH1, UPB1 and AIFs belong to the same HLH family. (A), Part of a phylogenetic tree constructed using conserved domains shows
A B Unconserved Conserved AIF4 AIF2 AIF3 AIF1 UPB1 IBH1 Supplemental Figure 1. IBH1, UPB1 and AIFs belong to the same HLH family. (A), Part of a phylogenetic tree constructed using conserved domains shows
translation The building blocks of proteins are? amino acids nitrogen containing bases like A, G, T, C, and U Complementary base pairing links
 The actual process of assembling the proteins on the ribosome is called? translation The building blocks of proteins are? Complementary base pairing links Define and name the Purines amino acids nitrogen
The actual process of assembling the proteins on the ribosome is called? translation The building blocks of proteins are? Complementary base pairing links Define and name the Purines amino acids nitrogen
Update of human and mouse forkhead box (FOX) gene families
 Update of human and mouse forkhead box (FOX) gene families Brian C. Jackson, 1 Christopher Carpenter, 1 Daniel W. Nebert 2* and Vasilis Vasiliou 1* 1 Molecular Toxicology and Environmental Health Sciences
Update of human and mouse forkhead box (FOX) gene families Brian C. Jackson, 1 Christopher Carpenter, 1 Daniel W. Nebert 2* and Vasilis Vasiliou 1* 1 Molecular Toxicology and Environmental Health Sciences
BIOLOGY 111. CHAPTER 6: DNA: The Molecule of Life
 BIOLOGY 111 CHAPTER 6: DNA: The Molecule of Life Chromosomes and Inheritance Learning Outcomes 6.1 Describe the structure of the DNA molecule and how this structure allows for the storage of information,
BIOLOGY 111 CHAPTER 6: DNA: The Molecule of Life Chromosomes and Inheritance Learning Outcomes 6.1 Describe the structure of the DNA molecule and how this structure allows for the storage of information,
What Are the Chemical Structures and Functions of Nucleic Acids?
 THE NUCLEIC ACIDS What Are the Chemical Structures and Functions of Nucleic Acids? Nucleic acids are polymers specialized for the storage, transmission, and use of genetic information. DNA = deoxyribonucleic
THE NUCLEIC ACIDS What Are the Chemical Structures and Functions of Nucleic Acids? Nucleic acids are polymers specialized for the storage, transmission, and use of genetic information. DNA = deoxyribonucleic
To investigate the heredity of the WFP gene, we selected plants that were homozygous
 Supplementary information Supplementary Note ST-12 WFP allele is semi-dominant To investigate the heredity of the WFP gene, we selected plants that were homozygous for chromosome 1 of Nipponbare and heterozygous
Supplementary information Supplementary Note ST-12 WFP allele is semi-dominant To investigate the heredity of the WFP gene, we selected plants that were homozygous for chromosome 1 of Nipponbare and heterozygous
Reproductive Biology and Endocrinology BioMed Central Review Smad signalling in the ovary Noora Kaivo-oja, Luke A Jeffery, Olli Ritvos and David G Mot
 Reproductive Biology and Endocrinology BioMed Central Review Smad signalling in the ovary Noora Kaivo-oja, Luke A Jeffery, Olli Ritvos and David G Mottershead* Open Access Address: Programme for Developmental
Reproductive Biology and Endocrinology BioMed Central Review Smad signalling in the ovary Noora Kaivo-oja, Luke A Jeffery, Olli Ritvos and David G Mottershead* Open Access Address: Programme for Developmental
Genome 373: Genomic Informatics. Elhanan Borenstein
 Genome 373: Genomic Informatics Elhanan Borenstein Genome 373 This course is intended to introduce students to the breadth of problems and methods in computational analysis of genomes and biological systems,
Genome 373: Genomic Informatics Elhanan Borenstein Genome 373 This course is intended to introduce students to the breadth of problems and methods in computational analysis of genomes and biological systems,
Biology 644: Bioinformatics
 Processes Activation Repression Initiation Elongation.... Processes Splicing Editing Degradation Translation.... Transcription Translation DNA Regulators DNA-Binding Transcription Factors Chromatin Remodelers....
Processes Activation Repression Initiation Elongation.... Processes Splicing Editing Degradation Translation.... Transcription Translation DNA Regulators DNA-Binding Transcription Factors Chromatin Remodelers....
CHAPTER 18 LECTURE NOTES: CONTROL OF GENE EXPRESSION PART B: CONTROL IN EUKARYOTES
 CHAPTER 18 LECTURE NOTES: CONTROL OF GENE EXPRESSION PART B: CONTROL IN EUKARYOTES I. Introduction A. No operon structures in eukaryotes B. Regulation of gene expression is frequently tissue specific.
CHAPTER 18 LECTURE NOTES: CONTROL OF GENE EXPRESSION PART B: CONTROL IN EUKARYOTES I. Introduction A. No operon structures in eukaryotes B. Regulation of gene expression is frequently tissue specific.
GENETICS - CLUTCH CH.15 GENOMES AND GENOMICS.
 !! www.clutchprep.com CONCEPT: OVERVIEW OF GENOMICS Genomics is the study of genomes in their entirety Bioinformatics is the analysis of the information content of genomes - Genes, regulatory sequences,
!! www.clutchprep.com CONCEPT: OVERVIEW OF GENOMICS Genomics is the study of genomes in their entirety Bioinformatics is the analysis of the information content of genomes - Genes, regulatory sequences,
Bioinformatics Tools. Stuart M. Brown, Ph.D Dept of Cell Biology NYU School of Medicine
 Bioinformatics Tools Stuart M. Brown, Ph.D Dept of Cell Biology NYU School of Medicine Bioinformatics Tools Stuart M. Brown, Ph.D Dept of Cell Biology NYU School of Medicine Overview This lecture will
Bioinformatics Tools Stuart M. Brown, Ph.D Dept of Cell Biology NYU School of Medicine Bioinformatics Tools Stuart M. Brown, Ph.D Dept of Cell Biology NYU School of Medicine Overview This lecture will
DNA: Structure and Function
 DNA: Structure and Function Biology's biggest moment in the 20th century, as heralded in six paragraphs in The New York Times, May 16, 1953. 2 Research of DNA Structure Chargaff s Rule of Ratios Amount
DNA: Structure and Function Biology's biggest moment in the 20th century, as heralded in six paragraphs in The New York Times, May 16, 1953. 2 Research of DNA Structure Chargaff s Rule of Ratios Amount
Bioinformatics for Biologists. Comparative Protein Analysis
 Bioinformatics for Biologists Comparative Protein nalysis: Part I. Phylogenetic Trees and Multiple Sequence lignments Robert Latek, PhD Sr. Bioinformatics Scientist Whitehead Institute for Biomedical Research
Bioinformatics for Biologists Comparative Protein nalysis: Part I. Phylogenetic Trees and Multiple Sequence lignments Robert Latek, PhD Sr. Bioinformatics Scientist Whitehead Institute for Biomedical Research
Just the Facts: A Basic Introduction to the Science Underlying NCBI Resources
 National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools News About NCBI Site Map
National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools News About NCBI Site Map
Chimp Sequence Annotation: Region 2_3
 Chimp Sequence Annotation: Region 2_3 Jeff Howenstein March 30, 2007 BIO434W Genomics 1 Introduction We received region 2_3 of the ChimpChunk sequence, and the first step we performed was to run RepeatMasker
Chimp Sequence Annotation: Region 2_3 Jeff Howenstein March 30, 2007 BIO434W Genomics 1 Introduction We received region 2_3 of the ChimpChunk sequence, and the first step we performed was to run RepeatMasker
Transcription factors
 Atlas of Genetics and Cytogenetics in Oncology and Haematology Transcription factors I Introduction * II Initiation of transcription III Transcription factors family pdf version I Introduction III.1 Helix-Turn-Helix
Atlas of Genetics and Cytogenetics in Oncology and Haematology Transcription factors I Introduction * II Initiation of transcription III Transcription factors family pdf version I Introduction III.1 Helix-Turn-Helix
Predicting Protein Structure and Examining Similarities of Protein Structure by Spectral Analysis Techniques
 Predicting Protein Structure and Examining Similarities of Protein Structure by Spectral Analysis Techniques, Melanie Abeysundera, Krista Collins, Chris Field Department of Mathematics and Statistics Dalhousie
Predicting Protein Structure and Examining Similarities of Protein Structure by Spectral Analysis Techniques, Melanie Abeysundera, Krista Collins, Chris Field Department of Mathematics and Statistics Dalhousie
The Genetic Code and Transcription. Chapter 12 Honors Genetics Ms. Susan Chabot
 The Genetic Code and Transcription Chapter 12 Honors Genetics Ms. Susan Chabot TRANSCRIPTION Copy SAME language DNA to RNA Nucleic Acid to Nucleic Acid TRANSLATION Copy DIFFERENT language RNA to Amino
The Genetic Code and Transcription Chapter 12 Honors Genetics Ms. Susan Chabot TRANSCRIPTION Copy SAME language DNA to RNA Nucleic Acid to Nucleic Acid TRANSLATION Copy DIFFERENT language RNA to Amino
Sequence Databases and database scanning
 Sequence Databases and database scanning Marjolein Thunnissen Lund, 2012 Types of databases: Primary sequence databases (proteins and nucleic acids). Composite protein sequence databases. Secondary databases.
Sequence Databases and database scanning Marjolein Thunnissen Lund, 2012 Types of databases: Primary sequence databases (proteins and nucleic acids). Composite protein sequence databases. Secondary databases.
The Reliability and Stability of an Inferred Phylogenetic Tree from Empirical Data
 The Reliability and Stability of an Inferred Phylogenetic Tree from Empirical Data Yukako Katsura, 1,2 Craig E. Stanley Jr, 1 Sudhir Kumar, 1 and Masatoshi Nei*,1,2 1 Department of Biology and Institute
The Reliability and Stability of an Inferred Phylogenetic Tree from Empirical Data Yukako Katsura, 1,2 Craig E. Stanley Jr, 1 Sudhir Kumar, 1 and Masatoshi Nei*,1,2 1 Department of Biology and Institute
1. Mitosis = growth, repair, asexual reproduc4on
 Places Muta4ons get passed on: Cell Reproduc4on: 2 types of cell reproduc4on: 1. Mitosis = growth, repair, asexual reproduc4on Photocopy machine Growth/Repair Passed on in the same body 2. Meiosis = sexual
Places Muta4ons get passed on: Cell Reproduc4on: 2 types of cell reproduc4on: 1. Mitosis = growth, repair, asexual reproduc4on Photocopy machine Growth/Repair Passed on in the same body 2. Meiosis = sexual
Comparative Bioinformatics. BSCI348S Fall 2003 Midterm 1
 BSCI348S Fall 2003 Midterm 1 Multiple Choice: select the single best answer to the question or completion of the phrase. (5 points each) 1. The field of bioinformatics a. uses biomimetic algorithms to
BSCI348S Fall 2003 Midterm 1 Multiple Choice: select the single best answer to the question or completion of the phrase. (5 points each) 1. The field of bioinformatics a. uses biomimetic algorithms to
Introduction to Bioinformatics Online Course: IBT
 Introduction to Bioinformatics Online Course: IBT Multiple Sequence Alignment Building Multiple Sequence Alignment Lec5: Interpreting your MSA Using Logos Using Logos - Logos are a terrific way to generate
Introduction to Bioinformatics Online Course: IBT Multiple Sequence Alignment Building Multiple Sequence Alignment Lec5: Interpreting your MSA Using Logos Using Logos - Logos are a terrific way to generate
Concepts of Genetics, 10e (Klug/Cummings/Spencer/Palladino) Chapter 1 Introduction to Genetics
 1 Concepts of Genetics, 10e (Klug/Cummings/Spencer/Palladino) Chapter 1 Introduction to Genetics 1) What is the name of the company or institution that has access to the health, genealogical, and genetic
1 Concepts of Genetics, 10e (Klug/Cummings/Spencer/Palladino) Chapter 1 Introduction to Genetics 1) What is the name of the company or institution that has access to the health, genealogical, and genetic
CSE : Computational Issues in Molecular Biology. Lecture 19. Spring 2004
 CSE 397-497: Computational Issues in Molecular Biology Lecture 19 Spring 2004-1- Protein structure Primary structure of protein is determined by number and order of amino acids within polypeptide chain.
CSE 397-497: Computational Issues in Molecular Biology Lecture 19 Spring 2004-1- Protein structure Primary structure of protein is determined by number and order of amino acids within polypeptide chain.
Gene expression DNA RNA. Protein DNA. Replication. Initiation Elongation Processing Export. DNA RNA Protein. Transcription. Degradation.
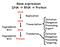 Gene expression DNA RNA Protein DNA DNA Degradation RNA Degradation Protein Replication Transcription Translation Initiation Elongation Processing Export Initiation Elongation Processing Targeting Chapter
Gene expression DNA RNA Protein DNA DNA Degradation RNA Degradation Protein Replication Transcription Translation Initiation Elongation Processing Export Initiation Elongation Processing Targeting Chapter
Introduction to Bioinformatics
 Introduction to Bioinformatics Dr. Taysir Hassan Abdel Hamid Lecturer, Information Systems Department Faculty of Computer and Information Assiut University taysirhs@aun.edu.eg taysir_soliman@hotmail.com
Introduction to Bioinformatics Dr. Taysir Hassan Abdel Hamid Lecturer, Information Systems Department Faculty of Computer and Information Assiut University taysirhs@aun.edu.eg taysir_soliman@hotmail.com
Why Use BLAST? David Form - August 15,
 Wolbachia Workshop 2017 Bioinformatics BLAST Basic Local Alignment Search Tool Finding Model Organisms for Study of Disease Can yeast be used as a model organism to study cystic fibrosis? BLAST Why Use
Wolbachia Workshop 2017 Bioinformatics BLAST Basic Local Alignment Search Tool Finding Model Organisms for Study of Disease Can yeast be used as a model organism to study cystic fibrosis? BLAST Why Use
DNA and the Production of Proteins Course Notes. Cell Biology. Sub-Topic 1.3 DNA and the Production of Proteins
 Cell Biology Sub-Topic 1.3 DNA and the Production of Proteins On completion of this subtopic I will be able to state that: Chromosomes contain genetic information that gives rise to an organism s characteristics.
Cell Biology Sub-Topic 1.3 DNA and the Production of Proteins On completion of this subtopic I will be able to state that: Chromosomes contain genetic information that gives rise to an organism s characteristics.
Lecture Overview. Overview of the Genetic Information. Chapter 3 DNA & RNA Lecture 6
 Visual Anatomy & Physiology First Edition Martini & Ober Chapter 3 DNA & RNA Lecture 6 Lecture Overview What is the cell s genetic information? How/where is the genetic information stored in eukaryotic
Visual Anatomy & Physiology First Edition Martini & Ober Chapter 3 DNA & RNA Lecture 6 Lecture Overview What is the cell s genetic information? How/where is the genetic information stored in eukaryotic
Article A Teaching Approach From the Exhaustive Search Method to the Needleman Wunsch Algorithm
 Article A Teaching Approach From the Exhaustive Search Method to the Needleman Wunsch Algorithm Zhongneng Xu * Yayun Yang Beibei Huang, From the Department of Ecology, Jinan University, Guangzhou 510632,
Article A Teaching Approach From the Exhaustive Search Method to the Needleman Wunsch Algorithm Zhongneng Xu * Yayun Yang Beibei Huang, From the Department of Ecology, Jinan University, Guangzhou 510632,
COMPARATIVE ANALYSIS AND INSILICO CHARACTERIZATION OF LEA, DREB AND OsDHODH1 GENES INVOLVED IN DROUGHT RESISTANCE OF ORYZA SATIVA
 International Journal of Bio-Technology and Research (IJBTR) ISSN (P):2249-6858; ISSN (E):2249-796X Vol. 7, Issue 3, Jun 2017, 47-56 TJPRC Pvt. Ltd. COMPARATIVE ANALYSIS AND INSILICO CHARACTERIZATION OF
International Journal of Bio-Technology and Research (IJBTR) ISSN (P):2249-6858; ISSN (E):2249-796X Vol. 7, Issue 3, Jun 2017, 47-56 TJPRC Pvt. Ltd. COMPARATIVE ANALYSIS AND INSILICO CHARACTERIZATION OF
Lecture 7 Motif Databases and Gene Finding
 Introduction to Bioinformatics for Medical Research Gideon Greenspan gdg@cs.technion.ac.il Lecture 7 Motif Databases and Gene Finding Motif Databases & Gene Finding Motifs Recap Motif Databases TRANSFAC
Introduction to Bioinformatics for Medical Research Gideon Greenspan gdg@cs.technion.ac.il Lecture 7 Motif Databases and Gene Finding Motif Databases & Gene Finding Motifs Recap Motif Databases TRANSFAC
CHAPTER 11 DNA NOTES PT. 4: PROTEIN SYNTHESIS TRANSCRIPTION & TRANSLATION
 CHAPTER 11 DNA NOTES PT. 4: PROTEIN SYNTHESIS TRANSCRIPTION & TRANSLATION DNA and the Language of Life RECAP Synthesis= Making something Protein Synthesis= Making Proteins Three steps in Protein Synthesis
CHAPTER 11 DNA NOTES PT. 4: PROTEIN SYNTHESIS TRANSCRIPTION & TRANSLATION DNA and the Language of Life RECAP Synthesis= Making something Protein Synthesis= Making Proteins Three steps in Protein Synthesis
2/10/17. Contents. Applications of HMMs in Epigenomics
 2/10/17 I529: Machine Learning in Bioinformatics (Spring 2017) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2017 Background:
2/10/17 I529: Machine Learning in Bioinformatics (Spring 2017) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2017 Background:
All organisms respond to cellular stress by shutting off the regular protein synthesis and
 1. INTRODUCTION All organisms respond to cellular stress by shutting off the regular protein synthesis and starting the synthesis of a new set of proteins called heat shock proteins or more generally,
1. INTRODUCTION All organisms respond to cellular stress by shutting off the regular protein synthesis and starting the synthesis of a new set of proteins called heat shock proteins or more generally,
Molecular Evolution. Lectures Papers Lab. Dr. Walter Salzburger. Structure of the course: Structure i
 Dr. Walter Salzburger Molecular Evolution Herbstsemester 2008 Freitag 13:15-15 Uhr 2 Kreditpunkte Structure i Structure of the course: The Nature of Molecular Evolution Molecules as Documents of Evolutionary
Dr. Walter Salzburger Molecular Evolution Herbstsemester 2008 Freitag 13:15-15 Uhr 2 Kreditpunkte Structure i Structure of the course: The Nature of Molecular Evolution Molecules as Documents of Evolutionary
Lecture Overview. Overview of the Genetic Information. Marieb s Human Anatomy and Physiology. Chapter 3 DNA & RNA Protein Synthesis Lecture 6
 Marieb s Human Anatomy and Physiology Marieb Hoehn Chapter 3 DNA & RNA Protein Synthesis Lecture 6 Lecture Overview The Genetic Information Structure of DNA/RNA DNA Replication Overview of protein synthesis
Marieb s Human Anatomy and Physiology Marieb Hoehn Chapter 3 DNA & RNA Protein Synthesis Lecture 6 Lecture Overview The Genetic Information Structure of DNA/RNA DNA Replication Overview of protein synthesis
Name: Period: Date: BIOLOGY HONORS DNA REVIEW GUIDE (extremely in detail) by Trung Pham. 5. What two bases are classified as purines? pyrimidine?
 BIOLOGY HONORS DNA REVIEW GUIDE (extremely in detail) by Trung Pham 1. What is the base pair rule for DNA? RNA? 2. What is the sugar found in RNA called? 3. is replaced by the base uracil in RNA? 4. What
BIOLOGY HONORS DNA REVIEW GUIDE (extremely in detail) by Trung Pham 1. What is the base pair rule for DNA? RNA? 2. What is the sugar found in RNA called? 3. is replaced by the base uracil in RNA? 4. What
Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid
 Supplementary Figures Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid sequence of Drosophila RCC1. Same colors are for Figure 1 with sequence of β-wedge that interacts with Ran in
Supplementary Figures Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid sequence of Drosophila RCC1. Same colors are for Figure 1 with sequence of β-wedge that interacts with Ran in
Introduction and History of Genome Modification. Adam Clore, PhD Director, Synthetic Biology Design
 Introduction and History of Genome Modification Adam Clore, PhD Director, Synthetic Biology Design Early Non-site Directed Genome Modification Homologous recombination in yeast TARGET GENE 5 Arm URA3 3
Introduction and History of Genome Modification Adam Clore, PhD Director, Synthetic Biology Design Early Non-site Directed Genome Modification Homologous recombination in yeast TARGET GENE 5 Arm URA3 3
BETA STRAND Prof. Alejandro Hochkoeppler Department of Pharmaceutical Sciences and Biotechnology University of Bologna
 Prof. Alejandro Hochkoeppler Department of Pharmaceutical Sciences and Biotechnology University of Bologna E-mail: a.hochkoeppler@unibo.it C-ter NH and CO groups: right, left, right (plane of the slide)
Prof. Alejandro Hochkoeppler Department of Pharmaceutical Sciences and Biotechnology University of Bologna E-mail: a.hochkoeppler@unibo.it C-ter NH and CO groups: right, left, right (plane of the slide)
Chapter 13 - Regulation of Gene Expression
 Chapter 13 - Regulation of Gene Expression 1. Describe the typical components of an operon in an E. coli (prokaryotic) cell. (p. 238-239) a. regulator gene - b. promoter - c. operator - d. structural gene
Chapter 13 - Regulation of Gene Expression 1. Describe the typical components of an operon in an E. coli (prokaryotic) cell. (p. 238-239) a. regulator gene - b. promoter - c. operator - d. structural gene
Differential Gene Expression
 Biology 4361 Developmental Biology Differential Gene Expression September 28, 2006 Chromatin Structure ~140 bp ~60 bp Transcriptional Regulation: 1. Packing prevents access CH 3 2. Acetylation ( C O )
Biology 4361 Developmental Biology Differential Gene Expression September 28, 2006 Chromatin Structure ~140 bp ~60 bp Transcriptional Regulation: 1. Packing prevents access CH 3 2. Acetylation ( C O )
 Watson BM Gene Capitolo 11 Watson et al., BIOLOGIA MOLECOLARE DEL GENE, Zanichelli editore S.p.A. ? Le proteine della trasposizione Watson et al., BIOLOGIA MOLECOLARE DEL GENE, Zanichelli editore S.p.A.
Watson BM Gene Capitolo 11 Watson et al., BIOLOGIA MOLECOLARE DEL GENE, Zanichelli editore S.p.A. ? Le proteine della trasposizione Watson et al., BIOLOGIA MOLECOLARE DEL GENE, Zanichelli editore S.p.A.
File S1. Program overview and features
 File S1 Program overview and features Query list filtering. Further filtering may be applied through user selected query lists (Figure. 2B, Table S3) that restrict the results and/or report specifically
File S1 Program overview and features Query list filtering. Further filtering may be applied through user selected query lists (Figure. 2B, Table S3) that restrict the results and/or report specifically
Structural Analysis of the EGR Family of Transcription Factors: Templates for Predicting Protein DNA Interactions
 Introduction Structural Analysis of the EGR Family of Transcription Factors: Templates for Predicting Protein DNA Interactions Jamie Duke, Rochester Institute of Technology Mentor: Carlos Camacho, University
Introduction Structural Analysis of the EGR Family of Transcription Factors: Templates for Predicting Protein DNA Interactions Jamie Duke, Rochester Institute of Technology Mentor: Carlos Camacho, University
Grundlagen der Bioinformatik Summer Lecturer: Prof. Daniel Huson
 Grundlagen der Bioinformatik, SoSe 11, D. Huson, April 11, 2011 1 1 Introduction Grundlagen der Bioinformatik Summer 2011 Lecturer: Prof. Daniel Huson Office hours: Thursdays 17-18h (Sand 14, C310a) 1.1
Grundlagen der Bioinformatik, SoSe 11, D. Huson, April 11, 2011 1 1 Introduction Grundlagen der Bioinformatik Summer 2011 Lecturer: Prof. Daniel Huson Office hours: Thursdays 17-18h (Sand 14, C310a) 1.1
Prediction of Transcription Factors that Regulate Common Binding Motifs Dana Wyman and Emily Alsentzer CS 229, Fall 2014
 Prediction of Transcription Factors that Regulate Common Binding Motifs Dana Wyman and Emily Alsentzer CS 229, Fall 2014 Introduction A. Background Proper regulation of mrna levels is essential to nearly
Prediction of Transcription Factors that Regulate Common Binding Motifs Dana Wyman and Emily Alsentzer CS 229, Fall 2014 Introduction A. Background Proper regulation of mrna levels is essential to nearly
BIOCHEMISTRY Nucleic Acids
 BIOCHEMISTRY Nucleic Acids BIOB111 CHEMISTRY & BIOCHEMISTRY Session 17 Session Plan Types of Nucleic Acids Nucleosides Nucleotides Primary Structure of Nucleic Acids DNA Double Helix DNA Replication Types
BIOCHEMISTRY Nucleic Acids BIOB111 CHEMISTRY & BIOCHEMISTRY Session 17 Session Plan Types of Nucleic Acids Nucleosides Nucleotides Primary Structure of Nucleic Acids DNA Double Helix DNA Replication Types
Supplementary Online Material. the flowchart of Supplemental Figure 1, with the fraction of known human loci retained
 SOM, page 1 Supplementary Online Material Materials and Methods Identification of vertebrate mirna gene candidates The computational procedure used to identify vertebrate mirna genes is summarized in the
SOM, page 1 Supplementary Online Material Materials and Methods Identification of vertebrate mirna gene candidates The computational procedure used to identify vertebrate mirna genes is summarized in the
2/19/13. Contents. Applications of HMMs in Epigenomics
 2/19/13 I529: Machine Learning in Bioinformatics (Spring 2013) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Background:
2/19/13 I529: Machine Learning in Bioinformatics (Spring 2013) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Background:
Molecular Genetics. Before You Read. Read to Learn
 12 Molecular Genetics section 3 DNA,, and Protein DNA codes for, which guides protein synthesis. What You ll Learn the different types of involved in transcription and translation the role of polymerase
12 Molecular Genetics section 3 DNA,, and Protein DNA codes for, which guides protein synthesis. What You ll Learn the different types of involved in transcription and translation the role of polymerase
DNA Transcription. Visualizing Transcription. The Transcription Process
 DNA Transcription By: Suzanne Clancy, Ph.D. 2008 Nature Education Citation: Clancy, S. (2008) DNA transcription. Nature Education 1(1) If DNA is a book, then how is it read? Learn more about the DNA transcription
DNA Transcription By: Suzanne Clancy, Ph.D. 2008 Nature Education Citation: Clancy, S. (2008) DNA transcription. Nature Education 1(1) If DNA is a book, then how is it read? Learn more about the DNA transcription
Gene expression analysis. Biosciences 741: Genomics Fall, 2013 Week 5. Gene expression analysis
 Gene expression analysis Biosciences 741: Genomics Fall, 2013 Week 5 Gene expression analysis From EST clusters to spotted cdna microarrays Long vs. short oligonucleotide microarrays vs. RT-PCR Methods
Gene expression analysis Biosciences 741: Genomics Fall, 2013 Week 5 Gene expression analysis From EST clusters to spotted cdna microarrays Long vs. short oligonucleotide microarrays vs. RT-PCR Methods
Applications of HMMs in Epigenomics
 I529: Machine Learning in Bioinformatics (Spring 2013) Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Contents Background:
I529: Machine Learning in Bioinformatics (Spring 2013) Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Contents Background:
O C. 5 th C. 3 rd C. the national health museum
 Elements of Molecular Biology Cells Cells is a basic unit of all living organisms. It stores all information to replicate itself Nucleus, chromosomes, genes, All living things are made of cells Prokaryote,
Elements of Molecular Biology Cells Cells is a basic unit of all living organisms. It stores all information to replicate itself Nucleus, chromosomes, genes, All living things are made of cells Prokaryote,
Gene Identification in silico
 Gene Identification in silico Nita Parekh, IIIT Hyderabad Presented at National Seminar on Bioinformatics and Functional Genomics, at Bioinformatics centre, Pondicherry University, Feb 15 17, 2006. Introduction
Gene Identification in silico Nita Parekh, IIIT Hyderabad Presented at National Seminar on Bioinformatics and Functional Genomics, at Bioinformatics centre, Pondicherry University, Feb 15 17, 2006. Introduction
Chapter 13 - Concept Mapping
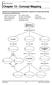 Chapter 13 - Concept Mapping Using the terms and phrases provided below, complete the concept map showing the discovery of DNA structure. amount of base pairs five-carbon sugar purine DNA polymerases Franklin
Chapter 13 - Concept Mapping Using the terms and phrases provided below, complete the concept map showing the discovery of DNA structure. amount of base pairs five-carbon sugar purine DNA polymerases Franklin
Lecture 2: Central Dogma of Molecular Biology & Intro to Programming
 Lecture 2: Central Dogma of Molecular Biology & Intro to Programming Central Dogma of Molecular Biology Proteins: workhorse molecules of biological systems Proteins are synthesized from the genetic blueprints
Lecture 2: Central Dogma of Molecular Biology & Intro to Programming Central Dogma of Molecular Biology Proteins: workhorse molecules of biological systems Proteins are synthesized from the genetic blueprints
Homology Modeling of Mouse orphan G-protein coupled receptors Q99MX9 and G2A
 Quality in Primary Care (2016) 24 (2): 49-57 2016 Insight Medical Publishing Group Research Article Homology Modeling of Mouse orphan G-protein Research Article Open Access coupled receptors Q99MX9 and
Quality in Primary Care (2016) 24 (2): 49-57 2016 Insight Medical Publishing Group Research Article Homology Modeling of Mouse orphan G-protein Research Article Open Access coupled receptors Q99MX9 and
a-dB. Code assigned:
 This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
Chem 465 Biochemistry II
 Chem 465 Biochemistry II Name: 2 points Multiple choice (4 points apiece): 1. Which of the following is not true of trna molecules? A) The 3'-terminal sequence is -CCA. B) Their anticodons are complementary
Chem 465 Biochemistry II Name: 2 points Multiple choice (4 points apiece): 1. Which of the following is not true of trna molecules? A) The 3'-terminal sequence is -CCA. B) Their anticodons are complementary
Outline. Computational Genomics and Molecular Biology. Genes Encode Proteins. DNA forms a double stranded helix. DNA replication T C A G
 Computational Genomics and Molecular Biology Dannie Durand Fall 2004 Lecture 1 Genes Encode Proteins DNA forms a double stranded helix GTGCACCTGACTCCTGAG A gene is a DNA sequence V H L T P E A protein
Computational Genomics and Molecular Biology Dannie Durand Fall 2004 Lecture 1 Genes Encode Proteins DNA forms a double stranded helix GTGCACCTGACTCCTGAG A gene is a DNA sequence V H L T P E A protein
GNET 744 Biological Sequence Analysis, Protein Structure and Genome-Wide Data
 GNET 744 Biological Sequence Analysis, Protein Structure and Genome-Wide Data This course provides an introduction to basic protein structure/function analyses, sequencing informatics and macromolecular
GNET 744 Biological Sequence Analysis, Protein Structure and Genome-Wide Data This course provides an introduction to basic protein structure/function analyses, sequencing informatics and macromolecular
An Evolutionary Optimization for Multiple Sequence Alignment
 195 An Evolutionary Optimization for Multiple Sequence Alignment 1 K. Lohitha Lakshmi, 2 P. Rajesh 1 M tech Scholar Department of Computer Science, VVIT Nambur, Guntur,A.P. 2 Assistant Prof Department
195 An Evolutionary Optimization for Multiple Sequence Alignment 1 K. Lohitha Lakshmi, 2 P. Rajesh 1 M tech Scholar Department of Computer Science, VVIT Nambur, Guntur,A.P. 2 Assistant Prof Department
ALGORITHMS IN BIO INFORMATICS. Chapman & Hall/CRC Mathematical and Computational Biology Series A PRACTICAL INTRODUCTION. CRC Press WING-KIN SUNG
 Chapman & Hall/CRC Mathematical and Computational Biology Series ALGORITHMS IN BIO INFORMATICS A PRACTICAL INTRODUCTION WING-KIN SUNG CRC Press Taylor & Francis Group Boca Raton London New York CRC Press
Chapman & Hall/CRC Mathematical and Computational Biology Series ALGORITHMS IN BIO INFORMATICS A PRACTICAL INTRODUCTION WING-KIN SUNG CRC Press Taylor & Francis Group Boca Raton London New York CRC Press
DNA. translation. base pairing rules for DNA Replication. thymine. cytosine. amino acids. The building blocks of proteins are?
 2 strands, has the 5-carbon sugar deoxyribose, and has the nitrogen base Thymine. The actual process of assembling the proteins on the ribosome is called? DNA translation Adenine pairs with Thymine, Thymine
2 strands, has the 5-carbon sugar deoxyribose, and has the nitrogen base Thymine. The actual process of assembling the proteins on the ribosome is called? DNA translation Adenine pairs with Thymine, Thymine
Introduction to Bioinformatics CPSC 265. What is bioinformatics? Textbooks
 Introduction to Bioinformatics CPSC 265 Thanks to Jonathan Pevsner, Ph.D. Textbooks Johnathan Pevsner, who I stole most of these slides from (thanks!) has written a textbook, Bioinformatics and Functional
Introduction to Bioinformatics CPSC 265 Thanks to Jonathan Pevsner, Ph.D. Textbooks Johnathan Pevsner, who I stole most of these slides from (thanks!) has written a textbook, Bioinformatics and Functional
