PLANT-ASSOCIATED MICROBE GENOME INITIATIVE
|
|
|
- Merilyn York
- 5 years ago
- Views:
Transcription
1 PLANT-ASSOCIATED MICROBE GENOME INITIATIVE White Paper Developed by the Plant Microbiology Research Community In association with the APS Public Policy Board October 1, 2002 Drafting Committee: Jan E. Leach Plant Pathology, Kansas State University, 4024 Throckmorton, Manhattan, KS 66506, Scott Gold Plant Pathology, University of Georgia, Athens, GA 30602, Sue Tolin Plant Molecular Biology Lab, Room 102, 435 Old Glade Road, Blacksburg, VA, Kellye Eversole Eversole Associates, 3208 Park View Rd, Chevy Chase, MD 20815, This document and listings of the meeting participants can be found at:
2 Executive Summary Plant-associated microorganisms are critical to agricultural and food security and are key components in maintaining the balance of our ecosystems. Some of these diverse microbes, which include viruses, bacteria, oomycetes, fungi and nematodes, cause plant diseases, while others prevent diseases or enhance plant growth. Despite their importance, we know little about them on a genomic level. To intervene in disease and understand the basis of biological control or symbiotic relationships, a concerted and coordinated genomic analysis of these microbes is essential. Genome analysis, in this context, refers to the structural and functional analysis of the microbe DNA including the genes, the proteins encoded by those genes, as well as noncoding sequences involved in genome dynamics and function. The ultimate emphasis is on understanding genomic functions involved in plant-associations. Members of the American Phytopathological Society (APS) developed a prioritized list of plantassociated microbes for genome analysis. With this list as a foundation for discussions, a Workshop on Genomic Analysis of Plant-Associated Microorganisms was held in Washington D.C. on April 9-11, The workshop was organized by the Public Policy Board of APS, and was funded by the Department of Energy (DOE), the National Science Foundation (NSF), USDA-Agricultural Research Service (USDA- ARS) and USDA-National Research Initiatives (USDA-NRI). The workshop included academic, industrial, and governmental experts from the genomics and microbial research communities and observers from the federal funding agencies. After reviewing current and near-term technologies, workshop participants proposed a comprehensive, international initiative to obtain the genomic information needed to understand these important microbes and their interactions with host plants and the environment. Specifically, the recommendations call for a 5-year, $500 million international public effort for genome analysis of plant-associated microbes. The goals are to (1) obtain genome sequence information for several representative groups of microbes; (2) identify and determine function for the genes/proteins and other genomic elements involved in plant-microbe interactions; (3) develop and implement standardized bioinformatic tools and a database system that is applicable across all microbes; and (4) educate and train scientists with skills and knowledge of biological and computational sciences who will apply the information to the protection of our food sources and environment. Overview Plant-associated microbes play critical roles in agricultural and food safety and security and in the maintenance of ecosystem balance. Despite their importance, as of September, 2002, less than six percent of the microbes whose genomic sequence has been completed and made publicly available were plantassociated microbes ( To remedy this dearth of critical information, the microbe-plant interaction community, lead by the Public Policy Board of the American Phytopathological Society (APS), initiated a planning effort to develop a strategy to undertake the structural and functional genomic analysis of viruses, bacteria, oomycetes, fungi, and nematodes that interact with plants. This white paper, which is the product of a Workshop on Genome Analysis of Plant-Associated Microbes held in Washington D.C. (April 9-11, 2002), describes the rationale and recommendations for an international public initiative for genomic analysis of these plant-associated microbes. Background Microbes can be found on or within every higher organism, including plants. Plant-associated microbes play critical roles in plant health. Some microbes cause diseases whereas others prevent diseases or enhance plant growth. Microbes also interact with each other on or within plants with the result of more or less severe disease. Within each microbe group, the diversity of species that interacts with plants is tremendous. For example, more than 10,000 pathogenic species of fungi and viruses are estimated to cause diseases on plants, many of which are economically important. Pathogenic microbes continually erode our supply of food and fiber, causing losses to producers of over $200 billion annually. Recent terrorist activities have heightened concern that, in the wrong hands, these agents could be used to threaten our food, environment or economic security (Schaad et al. 1999, Countering Agricultural Bioterrorism, 2002,
3 National Research Council, The historic devastation wrought by the inadvertent introduction of exotic pathogens testifies to such potential. For example, the introduction of the chestnut blight pathogen, Cryphonectria parasitica, from Europe in 1904 effectively eliminated from North America the American chestnut, an important and beautiful timber tree. Similar potential damage is currently occurring with disease epidemics of forest trees including sudden oak death caused by Phytophthora ramorum and pitch canker of pine caused by Fusarium circinatum. Introductions of other pathogenic microbes have caused tremendous suffering to humans, including the famines that resulted from the Phytophthora late blight epidemic of potato in in Ireland, and the Cochliobolus brown spot epidemic of rice in Bengal in Despite their importance to our agricultural base and food security, we know little of the molecular genetics of these microbes. Our understanding of how these microbes cause disease or benefit plants is rudimentary, at best. With the recent advent of genomics, which includes the structural and functional analysis of the microbe genes and the proteins encoded by those genes, we see the opportunity to understand the nature of beneficial and pathogenic microbe-plant-associations at levels never before possible. The Promise of Genome Analysis of Plant-Associated Microbes Due to powerful automated sequence technologies and advances in bioinformatics, the task of sequencing entire genomes of organisms is now routine. The sequenced genomes of a relatively few model organisms are already enabling a wide variety of new discoveries, including new genes and metabolic pathways and insights into the mechanisms of microbial pathogenesis. The continuous development of ever improving bioinformatics tools is enhancing the discovery power of these sequences. With these exciting advances, the means to generate sequence and functional information and the tools to use that information to understand the basic biology of microorganisms that cause or prevent diseases on plants are now available. Genome analyses will enhance our knowledge base exponentially and thereby provide tools to abrogate the problems caused by plant pathogenic microbes through genetically-based approaches and allow development of improved beneficial microbes. Historically, the molecular basis for interactions between plants and microbes was studied using a gene-by-gene approach and host resistance was a major control approach. Now, coupled with the increasing availability of plant host genomic sequences, structural and functional genomic analysis of plant-associated microbes will increase the speed of identification of genes involved in host-pathogen interactions and will allow genome-wide approaches to understanding the role of a gene or pathway in interactions with plants. Some genes will potentially be useful as sources of pathogen-derived resistance as has already been demonstrated for many viral diseases. Other genes may be involved in recognition to trigger host defense responses to the pathogen. Comparisons of genomes of related strains or species will provide an understanding of the evolution of microbial genomes, particularly as they evolve in associations with plants. Researchers exploiting comparative genomics will be able to discern the molecular basis of how some microbes have evolved to form intimate biotrophic associations with plant cells, whereas others inhabit intercellular spaces and still others colonize only the vascular systems of their hosts. Comparisons of sequences within and between species will also provide information to develop accurate diagnostic tools. Such precision in diagnostics may be essential to track the source or origin of a pathogen (e.g., as for the anthrax strains recently released in the US, or plum pox virus recently introduced). This information may be essential for forensic purposes as well as to understand the genetic potential of the pathogen to become established or to cause an epidemic. Additionally, because of the evolutionary conservation between microbes of different genera, and because of the exchange of genetic information between some groups of microbes, genome analysis of plant-associated microbes will contribute to our understanding of how animal pathogens cause disease. For example bacterial genes involved in plant disease induction exhibit a great deal of similarity to bacterial genes involved in disease induction in animals and humans (Keen et al., 2000, Thus, identifying the full complement of genes involved in pathogenesis will help decipher the disease strategies of these organisms and will guide design of new control measures for use in both plants and animals. The knowledge of how microbes interact with plants is essential to the development of effective and environmentally-sound, chemically-based strategies for disease control. In the US, over $600 million
4 are spent annually on agricultural fungicides alone. Similar to what is being observed with the heavy use of antibiotics targeted to human or animal pathogens, changes in pathogen biology more and more frequently render some of these chemicals ineffective. Furthermore, increased regulatory policies are restricting the use of existing agrochemicals for pathogen control. With genome information, multiple tactics for control of pathogens could be developed. For example, the identification of more precise targets in the pathogen may allow for design of more specific and effective chemicals that are environmentally benign. In sum, functional, structural and comparative genome analyses will provide insights into how microbes infect and establish either disease or beneficial interactions with host plants, how they reproduce and spread in the plants and how they persist and are disseminated in the environment. This information will be critical for the development of more effective, environmentally benign control measures and for accurate diagnostic and tracking methods. Which plant-associated microbes should be sequenced? Even with the costs of genome sequencing decreasing, it is clearly not feasible to sequence genomes of all plant-associated species at this time. Through a series of task forces with broad input, the APS established a prioritized list of species for genome sequencing. This list, published in a White Paper entitled Microbial Genomic Sequencing: Perspectives of the American Phytopathological Society ( was developed using the following criteria: Economic importance and relevance to U.S. and world-wide agriculture and forestry; Unique biological or environmental features; Broad interest to a significant community of scientists or agriculturists; Genetic tractability (i.e., the ease with which genetic studies, such as crosses or genome modifications, can be performed); and Availability of tools and other biological resources (e.g., gene libraries, genetic maps, and the genetic tractability of the host so that the system could be addressed from both sides of the hostpathogen interaction, etc.). The intention at its publication was that the APS list, compiled in , would be subject to change. A forum was held at the APS meetings in July, 2002 to discuss the list criteria and content, and define a means to periodically update the list. As a result of these discussions, and to encourage even wider input and a more accurate up-to-date list, a mechanism was established whereby the research community can revisit the list criteria and content, and make recommendations through the chairs of each pertinent APS subject matter committee ( The subject matter committees will annually evaluate the list. In addition, input is invited on the need for specialized lists, such as the list of organisms that are potential introduced threats to U.S. agriculture ( Community discussions preceding this white paper A workshop was held in Washington, D.C. April 9-11, 2002 to develop a strategy to expedite genome analysis of plant-associated microbes. The workshop was organized by the APS Public Policy Board, and was funded by the Department of Energy (DOE), the National Science Foundation (NSF), USDA-Agricultural Research Service (USDA-ARS) and USDA-National Research Initiative (USDA-NRI). Participants included academic, industrial, and governmental experts from the genomics and microbial research communities and observers from the federal funding agencies. The participants agreed on several key points: The lack of publicly available information on the genomes of plant-associated microbes is a major impediment to research, and, as a result, to crop biosecurity; The APS list of microbes for sequencing is a good starting point, but requires periodic review and updating; Current technological and biological resources would allow for sequence analysis of a significant number of microbes from this very diverse group, given appropriate funding resources;
5 The availability of genome sequence would enable and enhance the development of tools for functional and comparative genomics; and There is an urgent need for development and employment of flexible and broadly applicable bioinformatic tools and the establishment and continual updating of databases and genetic resource centers. An advisory committee was appointed to oversee the development of a Plant-Associated Microbe Genome Initiative. Discussions toward this initiative were broadened by two means. Firstly, a draft of this position paper was published in several relevant society newsletters and through lists, and the research community was invited to send ideas or comments by (microbegenomics@scisoc.org). These comments were posted at the APS PPB website ( Secondly, a public open forum was held at the Annual APS meetings in Milwaukee, WI (July 27, 2002). At this forum, the Initiative recommendations were summarized and feedback on them solicited. All comments and suggestions from these various sources was used to develop this final position paper. This final position paper, referred to as the Plant-Associated Microbe Genome Initiative, will be distributed to appropriate federal research funding agencies to garner support and guide program development for promoting genome analysis of these important microbes. Recommendations for Genome Analysis of Plant-Associated Microbes To remedy the paucity of genomic information available for plant-associated microbes, the workshop participants developed the Plant-Associated Microbe Genome Initiative and proposed a five-year concentrated effort. The recommendations for the initiative reflect the diversity of the microbes that interact with plants and the tremendous differences in the current status of the genome analysis of those organisms. Due to the diverse group of organisms, the recommendations are meant to provide a broad outline of how an international comprehensive effort might be approached and are not meant to provide specific technical detail of how the analyses should be accomplished. To be effective, the effort must be collaborative, well-coordinated, and international. Industrial and public partnerships should be encouraged with the stipulation that all information must be made rapidly available to the public. Finally, the effort must involve strong education and training programs. Four areas are described below that should be targeted in this five-year initiative. The participants recommend a $500 million investment to be distributed equally among these areas over the five years, except that in the first few years the proportion of resources necessarily will be greater to sequence analysis than to functional analyses. 1. Genome sequence analysis Genome sequence information is the key to gene discovery and the first step towards learning the function of each gene. There are vast differences in the amounts of sequence information currently available for different plant-associated microbes. For example, due to their smaller genome size, many different plant viruses and a few plant bacteria have already been sequenced; whereas, no genomes of plant-associated oomycetes, fungi or nematodes have been sequenced to completion. These differences in available information, which vary widely within and between microbe groups, impact the entry level for analysis of each group. They also impact the cost of sequencing the genomes of different microbe groups. No consensus was reached on the depth of sequence needed (i.e., draft versus complete genome sequences). One component of workshop participants suggested that to obtain useful SNP and genotype information more than one strain of an organism should be sequenced. As an example, this component recommended that the genomes of three strains of a given bacterial species should be sequenced completely followed by sequencing to 8X coverage of an additional strains. Another component suggested that if a complete genome sequence were available from one type strain, then lower coverage draft sequence of additional strains would provide information sufficient for most purposes. Although there were other permutations of this discussion, the emerging conclusions are that the depth of sequencing will depend on the end user needs (the questions to be addressed), subject to the complexity of the microbial genome of interest and available resources.
6 Focus initial resources largely on sequence analysis to remedy the paucity in sequence information available for plant-associated microbes; Target 2,000 Mb of sequence total over all microbes over five years; and Determine the depth of sequence or coverage for each microbe on a case-by-case basis, based on the specific needs or questions, to be evaluated by the research community. 2. Functional analysis Sophisticated technologies, including array-based technologies, are available for gene expression and proteomic and metabolomic analyses and will enable the dissection of the functions of genes and proteins involved in interactions of microbes with plants and the environment. Additionally, highthroughput genome-wide gene deletion and tagging methods are being developed. Although limited, there is publicly-available genome sequence information currently for a few plant-associated microbes that is sufficient to launch functional genomic studies. These microbes can serve as model systems for functional genomic efforts to identify genes/proteins and biochemical pathways that are involved in interactions with plants. Focus initial resources on those few microbes with sufficient structural genomic and genetic resources available; and Invest in improvement of technologies for functional analyses. 3. Standardized bioinformatic tools and database system The underpinning for successful structural and functional genome analysis is the ability to integrate, manipulate, analyze, compare, and store all of data generated by these technologies. It was clear from the intensity of discussions surrounding the bioinformatic tools and database systems that this is an area of major deficiency and concern. It was also clear that this is a problem not only for analysis and storage of data for the genomes of plant-associated microbes, but for all databases. Participants discussed the merits of fixing the existing systems as opposed to developing new, state-of-the-art systems, or using one major database as opposed to many smaller, interconnected databases. Critical features highlighted were that these tools must be (1) flexible (e.g., usable for comparisons among different genomes, including microbes and plants), (2) scalable, (3) inter-operable, (4) easily implemented, (5) readily updateable, and (6) stably funded and maintained. Guidelines should be developed for standardization of public databases, and these should include common (1) means for electronic submission of data, (2) terminology, and (3) criteria for quality of data submitted. Increase resources allocated to the standardization of databases and bioinformatic tools, as these are essential to the total science of genomics and the application of that science; Develop mechanisms for discussions, planning, and research at an international level; and Develop a framework to finance and manage the maintenance and continual updating of databases and information resources. 4. Train the next generation of plant microbiologists To capitalize on the information provided from the analysis of the genomes of plant-associated microbes, a key goal of the Plant-Associated Microbe Genome Initiative must be to train new and update current plant microbiologists to ensure a research community skilled in microbial and plant biology and in computational sciences. Additionally, these scholars will need the resources to interact and collaborate in an international research arena. International collaborations are particularly important for analysis of pathogens exotic to the US. Train plant microbiologists in the skills of genomics and computational sciences (e.g., this training should involve scientists at all levels of their careers to ensure an up-to-date, skilled pool of plant microbiologists); and Provide funding and opportunities for domestic and international research collaborations.
NATIONAL SCIENCE AND TECHNOLOGY COUNCIL. National Research and Development Strategy for Microbial Forensics
 NATIONAL SCIENCE AND TECHNOLOGY COUNCIL National Research and Development Strategy for Microbial Forensics National Strategy to Support Research in Microbial Forensics Attribution Investigations and National
NATIONAL SCIENCE AND TECHNOLOGY COUNCIL National Research and Development Strategy for Microbial Forensics National Strategy to Support Research in Microbial Forensics Attribution Investigations and National
Crop Biosecurity: Are We Prepared?
 Crop Biosecurity: Are We Prepared? White Paper Developed by the Public Policy Board of the American Phytopathological Society (APS) May 2003 Drafting Committee: Dr. John L. Sherwood Chair APS Public Policy
Crop Biosecurity: Are We Prepared? White Paper Developed by the Public Policy Board of the American Phytopathological Society (APS) May 2003 Drafting Committee: Dr. John L. Sherwood Chair APS Public Policy
Course Descriptions. BIOL: Biology. MICB: Microbiology. [1]
![Course Descriptions. BIOL: Biology. MICB: Microbiology. [1] Course Descriptions. BIOL: Biology. MICB: Microbiology. [1]](/thumbs/72/66855241.jpg) Course Descriptions BIOL: Biology http://www.calendar.ubc.ca/vancouver/courses.cfm?code=biol [1] BIOL 112 (3) Biology of the Cell The principles of cellular and molecular biology using bacterial and eukaryotic
Course Descriptions BIOL: Biology http://www.calendar.ubc.ca/vancouver/courses.cfm?code=biol [1] BIOL 112 (3) Biology of the Cell The principles of cellular and molecular biology using bacterial and eukaryotic
5th National Plant Disease Recovery System Workshop Apr 14-16, 2013, Falls Church, VA
 Refining the Generic Model for Disease Recovery Plans for the National Plant Disease Recovery System (or) Are We Finally Getting Closer to Generic Plans? 5th National Plant Disease Recovery System Workshop
Refining the Generic Model for Disease Recovery Plans for the National Plant Disease Recovery System (or) Are We Finally Getting Closer to Generic Plans? 5th National Plant Disease Recovery System Workshop
Ribotyping Easily Fills in for Whole Genome Sequencing to Characterize Food-borne Pathogens David Sistanich
 Ribotyping Easily Fills in for Whole Genome Sequencing to Characterize Food-borne Pathogens David Sistanich Technical Support Specialist Hygiena, LLC Whole genome sequencing (WGS) has become the ultimate
Ribotyping Easily Fills in for Whole Genome Sequencing to Characterize Food-borne Pathogens David Sistanich Technical Support Specialist Hygiena, LLC Whole genome sequencing (WGS) has become the ultimate
Following text taken from Suresh Kumar. Bioinformatics Web - Comprehensive educational resource on Bioinformatics. 6th May.2005
 Bioinformatics is the recording, annotation, storage, analysis, and searching/retrieval of nucleic acid sequence (genes and RNAs), protein sequence and structural information. This includes databases of
Bioinformatics is the recording, annotation, storage, analysis, and searching/retrieval of nucleic acid sequence (genes and RNAs), protein sequence and structural information. This includes databases of
Agricultural biosecurity: threats to crop production. Michael Jeger Division of Biology, Imperial College London Beijing workshop, 31 Oct 3 Nov 2010
 Agricultural biosecurity: threats to crop production Michael Jeger Division of Biology, Imperial College London Beijing workshop, 31 Oct 3 Nov 2010 Outline of presentation Crop production, origins and
Agricultural biosecurity: threats to crop production Michael Jeger Division of Biology, Imperial College London Beijing workshop, 31 Oct 3 Nov 2010 Outline of presentation Crop production, origins and
Abstract... i. Committee Membership... iii. Foreword... vii. 1 Scope Introduction Standard Precautions References...
 Vol. 28 No. 12 Replaces MM18-P Vol. 27 No. 22 Interpretive Criteria for Identification of Bacteria and Fungi by DNA Target Sequencing; Approved Guideline Sequencing DNA targets of cultured isolates provides
Vol. 28 No. 12 Replaces MM18-P Vol. 27 No. 22 Interpretive Criteria for Identification of Bacteria and Fungi by DNA Target Sequencing; Approved Guideline Sequencing DNA targets of cultured isolates provides
White pine blister rust on pine
 White pine blister rust on pine History of white pine blister rust First recorded in Europe from the Baltic region in 1854 By 1900 it widespread across Europe - rapid spread by forest tree nurseries Introduced
White pine blister rust on pine History of white pine blister rust First recorded in Europe from the Baltic region in 1854 By 1900 it widespread across Europe - rapid spread by forest tree nurseries Introduced
Infectious Diseases Institute Faculty Hires See below for research interests, areas of expertise and contact information
 Infectious Diseases Institute Faculty Hires See below for research interests, areas of expertise and contact information Matt Anderson, Ph.D. Anderson.3196@osu.edu Microbial Communities, Host Defense and
Infectious Diseases Institute Faculty Hires See below for research interests, areas of expertise and contact information Matt Anderson, Ph.D. Anderson.3196@osu.edu Microbial Communities, Host Defense and
The Microbiology of Food
 The Microbiology of Food Bio 8 - Fall 2018 Tu 6:30PM - 9:00PM Anderson, Room 309 Instructor: Benjamin Wolfe (Assistant Professor, Department of Biology) e-mail: benjamin.wolfe@tufts.edu Office Hours: 3:30-:30pm
The Microbiology of Food Bio 8 - Fall 2018 Tu 6:30PM - 9:00PM Anderson, Room 309 Instructor: Benjamin Wolfe (Assistant Professor, Department of Biology) e-mail: benjamin.wolfe@tufts.edu Office Hours: 3:30-:30pm
2008 Workshop Statistical Workshop for Microarray Data Analysis
 Historical APS Special Sessions, 2010 2003 YEAR SECTION TITLE 2010 of Pathogens 10th I. E. Melhus Graduate Student Symposium: Seed Epidemiology, Management, and Phytosanitary Concerns 2010 of Pathogens
Historical APS Special Sessions, 2010 2003 YEAR SECTION TITLE 2010 of Pathogens 10th I. E. Melhus Graduate Student Symposium: Seed Epidemiology, Management, and Phytosanitary Concerns 2010 of Pathogens
Building a Foundation to Combine Site-Specific Biological and Physical Data for Next-Generation Precision Agriculture
 Building a Foundation to Combine Site-Specific Biological and Physical Data for Next-Generation Precision Agriculture Kellye Eversole 7 December 2017 Microbe-Assisted Crop Production Symposium Vienna,
Building a Foundation to Combine Site-Specific Biological and Physical Data for Next-Generation Precision Agriculture Kellye Eversole 7 December 2017 Microbe-Assisted Crop Production Symposium Vienna,
Just the Facts: A Basic Introduction to the Science Underlying NCBI Resources
 National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools News About NCBI Site Map
National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools News About NCBI Site Map
DOCTOR OF PHILOSOPHY PROGRAM IN ORAL BIOLOGY
 Doctor of Philosophy Program in Oral Biology FACULTY OF DENTISTRY Naresuan University 183 Doctor of Philosophy Program in Oral Biology Oral biology at Naresuan University covers the study of the structure
Doctor of Philosophy Program in Oral Biology FACULTY OF DENTISTRY Naresuan University 183 Doctor of Philosophy Program in Oral Biology Oral biology at Naresuan University covers the study of the structure
Review of lecture 1: Significance of Plant Disease. Lecture 2: Disease Concept
 Review of lecture 1: Significance of Plant Disease 10% of all food production is lost to disease (30% to all pests) The introduction of exotic plant pathogens has caused great losses: e.g., American chestnut
Review of lecture 1: Significance of Plant Disease 10% of all food production is lost to disease (30% to all pests) The introduction of exotic plant pathogens has caused great losses: e.g., American chestnut
Regulatory Requirements for Use of Transgenic Plants in the Greenhouse
 Regulatory Requirements for Use of Transgenic Plants in the Greenhouse Agenda Introduction Biosafety Levels Containment Frank A. Cantone, Ph.D., CBSP Biological Safety Officer Environmental Health & Safety
Regulatory Requirements for Use of Transgenic Plants in the Greenhouse Agenda Introduction Biosafety Levels Containment Frank A. Cantone, Ph.D., CBSP Biological Safety Officer Environmental Health & Safety
Embracing Complexity to Achieve a New Vision for Agriculture
 Embracing Complexity to Achieve a New Vision for Agriculture Kellye Eversole 16 February 2018 Symposium: Phytobiome Research to Improve Agricultural Productivity The Challenge 32 Growing Seasons Agricultural
Embracing Complexity to Achieve a New Vision for Agriculture Kellye Eversole 16 February 2018 Symposium: Phytobiome Research to Improve Agricultural Productivity The Challenge 32 Growing Seasons Agricultural
European Technology Platform for Global Animal Health. Action Plan
 European Technology Platform for Global Animal Health Action Plan European Technology T Platform for Global Animal Health Action Plan 2 Table of contents Executive Summary 7 Chapter 1: The Action Plan
European Technology Platform for Global Animal Health Action Plan European Technology T Platform for Global Animal Health Action Plan 2 Table of contents Executive Summary 7 Chapter 1: The Action Plan
BIOLOGICAL CONTROL MAKING IT WORK
 BIOLOGICAL CONTROL MAKING IT WORK PART 1: SUMMARY & RECOMMENDATIONS On April 4 and 5,1991,52 individuals from academia, the private sector BRIAN F. CHABOT Director of Research Cornell Cooperative Extension
BIOLOGICAL CONTROL MAKING IT WORK PART 1: SUMMARY & RECOMMENDATIONS On April 4 and 5,1991,52 individuals from academia, the private sector BRIAN F. CHABOT Director of Research Cornell Cooperative Extension
CHAPTERS 16 & 17: DNA Technology
 CHAPTERS 16 & 17: DNA Technology 1. What is the function of restriction enzymes in bacteria? 2. How do bacteria protect their DNA from the effects of the restriction enzymes? 3. How do biologists make
CHAPTERS 16 & 17: DNA Technology 1. What is the function of restriction enzymes in bacteria? 2. How do bacteria protect their DNA from the effects of the restriction enzymes? 3. How do biologists make
Unit 2: Metabolism and Survival Sub-Topic (2.7) Genetic Control of Metabolism (2.8) Ethical considerations in the use of microorganisms
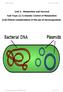 Unit 2: Metabolism and Survival Sub-Topic (2.7) Genetic Control of Metabolism (2.8) Ethical considerations in the use of microorganisms Duncanrig Secondary JHM&MHC 2015 Page 1 of 18 On completion of this
Unit 2: Metabolism and Survival Sub-Topic (2.7) Genetic Control of Metabolism (2.8) Ethical considerations in the use of microorganisms Duncanrig Secondary JHM&MHC 2015 Page 1 of 18 On completion of this
Plant biosecurity challenges & needs at Australia s border. Mark Whattam AQIS
 Plant biosecurity challenges & needs at Australia s border Mark Whattam AQIS Presentation outline 1. At the front line: AQIS Operational Science Program 2. An interconnected world - the threats are real!
Plant biosecurity challenges & needs at Australia s border Mark Whattam AQIS Presentation outline 1. At the front line: AQIS Operational Science Program 2. An interconnected world - the threats are real!
MICROBIOLOGY (MICR) Microbiology (MICR) 1
 Microbiology (MICR) 1 MICROBIOLOGY (MICR) MICR 1513 Inquiry-Based Biology Description: Directed inquiry and hands-on study of biological principles. Restricted to elementary education majors or related
Microbiology (MICR) 1 MICROBIOLOGY (MICR) MICR 1513 Inquiry-Based Biology Description: Directed inquiry and hands-on study of biological principles. Restricted to elementary education majors or related
Genetics Lecture 21 Recombinant DNA
 Genetics Lecture 21 Recombinant DNA Recombinant DNA In 1971, a paper published by Kathleen Danna and Daniel Nathans marked the beginning of the recombinant DNA era. The paper described the isolation of
Genetics Lecture 21 Recombinant DNA Recombinant DNA In 1971, a paper published by Kathleen Danna and Daniel Nathans marked the beginning of the recombinant DNA era. The paper described the isolation of
Refresher on gene expression - DNA: The stuff of life
 Plant Pathology 602 Plant-Microbe Interactions Lecture 2 Molecular methods for studying hostpathogen interactions I Sophien Kamoun kamoun.1@osu.edu The Ohio State University Ohio Agricultural Research
Plant Pathology 602 Plant-Microbe Interactions Lecture 2 Molecular methods for studying hostpathogen interactions I Sophien Kamoun kamoun.1@osu.edu The Ohio State University Ohio Agricultural Research
Microbial Metabolism Systems Microbiology
 1 Microbial Metabolism Systems Microbiology Ching-Tsan Huang ( 黃慶璨 ) Office: Agronomy Hall, Room 111 Tel: (02) 33664454 E-mail: cthuang@ntu.edu.tw MIT OCW Systems Microbiology aims to integrate basic biological
1 Microbial Metabolism Systems Microbiology Ching-Tsan Huang ( 黃慶璨 ) Office: Agronomy Hall, Room 111 Tel: (02) 33664454 E-mail: cthuang@ntu.edu.tw MIT OCW Systems Microbiology aims to integrate basic biological
Viral Genomes. Genomes may consist of: 1. Double Stranded DNA 2. Double Stranded RNA 3. Single-stranded RNA 4. Single-stranded DNA
 Chapter 19 Viral Genomes Genomes may consist of: 1. Double Stranded DNA 2. Double Stranded RNA 3. Single-stranded RNA 4. Single-stranded DNA Genome is usually organized as a single linear or circular molecule
Chapter 19 Viral Genomes Genomes may consist of: 1. Double Stranded DNA 2. Double Stranded RNA 3. Single-stranded RNA 4. Single-stranded DNA Genome is usually organized as a single linear or circular molecule
Microbial Biotechnology agustin krisna wardani
 Microbial Biotechnology agustin krisna wardani 1. The Structure of Microbes Microbes (microorganisms) are tiny organisms that are too small to be seen individually by the naked eye and must be viewed with
Microbial Biotechnology agustin krisna wardani 1. The Structure of Microbes Microbes (microorganisms) are tiny organisms that are too small to be seen individually by the naked eye and must be viewed with
SAMPLE. Interpretive Criteria for Identification of Bacteria and Fungi by Targeted DNA Sequencing
 MM18 Interpretive Criteria for Identification of Bacteria and Fungi by Targeted DNA Sequencing This guideline includes information on sequencing DNA targets of cultured isolates, provides a quantitative
MM18 Interpretive Criteria for Identification of Bacteria and Fungi by Targeted DNA Sequencing This guideline includes information on sequencing DNA targets of cultured isolates, provides a quantitative
Science and Regulation: A Canadian Approach to Invasive Alien Species
 Science and Regulation: A Canadian Approach to Invasive Alien Species Eric Allen Natural Resources Canada Lesley Cree Canadian Food Inspection Agency ! Canada s National Plant Protection Organization Canadian
Science and Regulation: A Canadian Approach to Invasive Alien Species Eric Allen Natural Resources Canada Lesley Cree Canadian Food Inspection Agency ! Canada s National Plant Protection Organization Canadian
MicroSEQ TM ID Rapid Microbial Identification System:
 MicroSEQ TM ID Rapid Microbial Identification System: the complete solution for reliable genotypic microbial identification 1 The world leader in serving science Rapid molecular methods for pharmaceutical
MicroSEQ TM ID Rapid Microbial Identification System: the complete solution for reliable genotypic microbial identification 1 The world leader in serving science Rapid molecular methods for pharmaceutical
From Simple to Complex Phytobiomes and the 2050 Vision for Agriculture
 From Simple to Complex Phytobiomes and the 2050 Vision for Agriculture Kellye Eversole Executive Director, Alliance Exploring Phytobiomes Workshop Plant and Animal Genome XXV San Diego, California, USA
From Simple to Complex Phytobiomes and the 2050 Vision for Agriculture Kellye Eversole Executive Director, Alliance Exploring Phytobiomes Workshop Plant and Animal Genome XXV San Diego, California, USA
MICROBIO, IMMUN, PATHOLOGY-MIP (MIP)
 Microbio, Immun, Pathology-MIP (MIP) 1 MICROBIO, IMMUN, PATHOLOGY-MIP (MIP) Courses MIP 101 Introduction to Human Disease (GT-SC2) Credits: 3 (3-0-0) Survey of human systems and diseases. Additional Information:
Microbio, Immun, Pathology-MIP (MIP) 1 MICROBIO, IMMUN, PATHOLOGY-MIP (MIP) Courses MIP 101 Introduction to Human Disease (GT-SC2) Credits: 3 (3-0-0) Survey of human systems and diseases. Additional Information:
Enabling the adoption of new diagnostics within the UK healthcare system:
 Enabling the adoption of new diagnostics within the UK healthcare system: The key role of diagnostics in the AMR challenge Fiona Carragher FRCPath @DepCSOFiona Deputy Chief Scientific Officer for England
Enabling the adoption of new diagnostics within the UK healthcare system: The key role of diagnostics in the AMR challenge Fiona Carragher FRCPath @DepCSOFiona Deputy Chief Scientific Officer for England
Promises and Problems Associated with Agricultural Biotechnology
 Workshop A Promises and Problems Associated with Agricultural Biotechnology DONALD P. WEEKS University of Nebraska Lincoln Lincoln, NE Participants in Workshop A addressed the following questions: What
Workshop A Promises and Problems Associated with Agricultural Biotechnology DONALD P. WEEKS University of Nebraska Lincoln Lincoln, NE Participants in Workshop A addressed the following questions: What
May 9, Meeting Summary. Facilitating Antibacterial Drug Development
 May 9, 2012 Meeting Summary Facilitating Antibacterial Drug Development Origins of the Current Public Health Crisis of Antibacterial Resistance Antibacterial drugs play a critical role in the ability to
May 9, 2012 Meeting Summary Facilitating Antibacterial Drug Development Origins of the Current Public Health Crisis of Antibacterial Resistance Antibacterial drugs play a critical role in the ability to
2018 Media Kit. Table of Contents Click the links above to jump to the desired page
 2018 Media Kit The American Society for Microbiology is the oldest and largest single life science membership organization in the world. Membership has grown from 59 scientists in 1899 to more than 39,000
2018 Media Kit The American Society for Microbiology is the oldest and largest single life science membership organization in the world. Membership has grown from 59 scientists in 1899 to more than 39,000
2019 Media Kit. Table of Contents Click the links above to jump to the desired page
 2019 Media Kit The American Society for Microbiology is the oldest and largest single life science membership organization in the world. Membership has grown from 59 scientists in 1899 to more than 39,000
2019 Media Kit The American Society for Microbiology is the oldest and largest single life science membership organization in the world. Membership has grown from 59 scientists in 1899 to more than 39,000
Biodiversity Conservation for Sustainable Agroecosystems Workshop
 Biodiversity Conservation for Sustainable Agroecosystems Workshop CO-CHAIRS MARIE BOEHM Center for Studies in Agriculture, Law and Environment, University of Saskatchewan BOB MORGAN Saskatchewan Wheat
Biodiversity Conservation for Sustainable Agroecosystems Workshop CO-CHAIRS MARIE BOEHM Center for Studies in Agriculture, Law and Environment, University of Saskatchewan BOB MORGAN Saskatchewan Wheat
Mini-Symposium MICROBIOTA. Free. Meet the speaker. 14. November :30 19:00 Bohnenkamp Haus. Everyone welcome. Sponsored by:
 Mini-Symposium MICROBIOTA Free Meet the speaker Everyone welcome Sponsored by: 14. November 2017 13:30 19:00 Bohnenkamp Haus PROGRAM 13:30 14:30 Julia Vorholt The leaf microbiota: disassembling and rebuilding
Mini-Symposium MICROBIOTA Free Meet the speaker Everyone welcome Sponsored by: 14. November 2017 13:30 19:00 Bohnenkamp Haus PROGRAM 13:30 14:30 Julia Vorholt The leaf microbiota: disassembling and rebuilding
Introduction to Bioinformatics
 Introduction to Bioinformatics Alla L Lapidus, Ph.D. SPbSU St. Petersburg Term Bioinformatics Term Bioinformatics was invented by Paulien Hogeweg (Полина Хогевег) and Ben Hesper in 1970 as "the study of
Introduction to Bioinformatics Alla L Lapidus, Ph.D. SPbSU St. Petersburg Term Bioinformatics Term Bioinformatics was invented by Paulien Hogeweg (Полина Хогевег) and Ben Hesper in 1970 as "the study of
Lawrence Livermore National Laboratory 2
 August 28, 2012 LLNL-PRES-573112 This work was performed under the auspices of the U.S. Department of Energy by Lawrence Livermore National Laboratory under contract DE-AC52-07NA27344. Lawrence Livermore
August 28, 2012 LLNL-PRES-573112 This work was performed under the auspices of the U.S. Department of Energy by Lawrence Livermore National Laboratory under contract DE-AC52-07NA27344. Lawrence Livermore
Generated by Foxit PDF Creator Foxit Software For evaluation only. Biotechnology in Plant Pathology
 Biotechnology in Plant Pathology Plant Biotechnology Definition: The use of tissue culture & genetic engineering techniques to produce genetically modified plants that show improved desirable characteristics.
Biotechnology in Plant Pathology Plant Biotechnology Definition: The use of tissue culture & genetic engineering techniques to produce genetically modified plants that show improved desirable characteristics.
Introduction, by Tortora, Funke and Case, 11th Ed. TENTATIVE LECTURE OUTLINE DATE TOPIC CHAPTER
 MICROBIOLOGY 220 Spring 2014 TTh Section Professor: Scott Rose Text: Microbiology; An Introduction, by Tortora, Funke and Case, 11th Ed. TENTATIVE LECTURE OUTLINE DATE TOPIC CHAPTER Jan. 23 Introduction
MICROBIOLOGY 220 Spring 2014 TTh Section Professor: Scott Rose Text: Microbiology; An Introduction, by Tortora, Funke and Case, 11th Ed. TENTATIVE LECTURE OUTLINE DATE TOPIC CHAPTER Jan. 23 Introduction
Course outlines for senior Microbiology and Virology units of study within the Bachelor of Medical Science Last updated 18 November 2006
 Course outlines for senior Microbiology and Virology units of study within the Bachelor of Medical Science Last updated 18 November 2006 These course outlines are a guide only. They are provided for the
Course outlines for senior Microbiology and Virology units of study within the Bachelor of Medical Science Last updated 18 November 2006 These course outlines are a guide only. They are provided for the
Regulation of enzyme synthesis
 Regulation of enzyme synthesis The lac operon is an example of an inducible operon - it is normally off, but when a molecule called an inducer is present, the operon turns on. The trp operon is an example
Regulation of enzyme synthesis The lac operon is an example of an inducible operon - it is normally off, but when a molecule called an inducer is present, the operon turns on. The trp operon is an example
The Swedish Foundation for Strategic Research (SSF) announces a. call for proposals in the areas of
 1(8) 2008-09-16 Announcement The Swedish Foundation for Strategic Research (SSF) announces a call for proposals in the areas of Design, development and validation of new predictive models and new biomarkers
1(8) 2008-09-16 Announcement The Swedish Foundation for Strategic Research (SSF) announces a call for proposals in the areas of Design, development and validation of new predictive models and new biomarkers
USDA Agricultural Biotechnology Research
 USDA Agricultural Biotechnology Research Peter Burfening, Ph.D. National Program Leader, Animal Genome Program National Research Initiative CSREES, USDA Agricultural Biotechnology A range of tools, including
USDA Agricultural Biotechnology Research Peter Burfening, Ph.D. National Program Leader, Animal Genome Program National Research Initiative CSREES, USDA Agricultural Biotechnology A range of tools, including
Science to Support Plant Protection for Horticulture AAFC s Science & Technology Branch
 Science to Support Plant Protection for Horticulture AAFC s Science & Technology Branch Crop, Plant Protection and the Environment Committee March 15, 2018 Dr. Della Johnston Outline Strategic direction
Science to Support Plant Protection for Horticulture AAFC s Science & Technology Branch Crop, Plant Protection and the Environment Committee March 15, 2018 Dr. Della Johnston Outline Strategic direction
NATIONAL INSTITUTE FOR MICROBIAL FORENSICS & FOOD AND AGRICULTURAL BIOSECURITY
 NATIONAL INSTITUTE FOR MICROBIAL FORENSICS & FOOD AND AGRICULTURAL BIOSECURITY NIMFFAB OKLAHOMA STATE UNIVERSITY Jacque Fletcher - NPDRS Workshop March 2011 National Academy of Sciences 2002 A strong national
NATIONAL INSTITUTE FOR MICROBIAL FORENSICS & FOOD AND AGRICULTURAL BIOSECURITY NIMFFAB OKLAHOMA STATE UNIVERSITY Jacque Fletcher - NPDRS Workshop March 2011 National Academy of Sciences 2002 A strong national
ISA Tree academy October 1, 2014 Waco, TX
 ISA Tree academy October 1, 2014 Waco, TX Sheila McBride Diagnostician at the Texas Plant Disease Diagnostic Lab Texas A&M AgriLife Extension Service College Station, TX 77843 What is a PLANT DISEASE?
ISA Tree academy October 1, 2014 Waco, TX Sheila McBride Diagnostician at the Texas Plant Disease Diagnostic Lab Texas A&M AgriLife Extension Service College Station, TX 77843 What is a PLANT DISEASE?
Growing Together: How Viruses Have Shaped Human Evolution. Shirlee Wohl Katherine Wu
 Growing Together: How Viruses Have Shaped Human Evolution Shirlee Wohl Katherine Wu What comes to your mind when you hear the word virus? flu public health cold vaccine infection disease cough biology
Growing Together: How Viruses Have Shaped Human Evolution Shirlee Wohl Katherine Wu What comes to your mind when you hear the word virus? flu public health cold vaccine infection disease cough biology
Rein Drenkhan, PhD. Associate professor of forest pathology
 Some experiences and results from the Norwegian-Estonian Research Cooperation Programme project: DNA-based early detection and diagnostics of alien invasive forest pathogens and tracing of their introduction
Some experiences and results from the Norwegian-Estonian Research Cooperation Programme project: DNA-based early detection and diagnostics of alien invasive forest pathogens and tracing of their introduction
West Africa Centre for Crop Improvement. New Four Year Ph D Programme In Plant Breeding f or West Africa Centre For Crop Improvement
 West Africa Centre for Crop Improvement New Four Year Ph D Programme In Plant Breeding f or West Africa Centre For Crop Improvement Introduction The West Africa Centre for Crop Improvement (WACCI), a partnership
West Africa Centre for Crop Improvement New Four Year Ph D Programme In Plant Breeding f or West Africa Centre For Crop Improvement Introduction The West Africa Centre for Crop Improvement (WACCI), a partnership
State of Texas Assessments of Academic Readiness (STAAR ) Performance Level Descriptors Biology
 State of Texas Assessments of Academic Readiness (STAAR ) Biology Scientific process skills are not assessed in isolation but are incorporated into questions that assess the biology content. These process
State of Texas Assessments of Academic Readiness (STAAR ) Biology Scientific process skills are not assessed in isolation but are incorporated into questions that assess the biology content. These process
2018 Media Kit. Table of Contents Click the links above to jump to the desired page
 2018 Media Kit The American Society for Microbiology is the oldest and largest single life science membership organization in the world. Membership has grown from 59 scientists in 1899 to more than 39,000
2018 Media Kit The American Society for Microbiology is the oldest and largest single life science membership organization in the world. Membership has grown from 59 scientists in 1899 to more than 39,000
ATIP Avenir Program 2018 Young group leader
 ATIP Avenir Program 2018 Young group leader Important dates - October 17th (4 pm) 2017 : opening of the registrations online - November 23 th 2017: deadline for the online submission, the mailing of the
ATIP Avenir Program 2018 Young group leader Important dates - October 17th (4 pm) 2017 : opening of the registrations online - November 23 th 2017: deadline for the online submission, the mailing of the
Business and Industry Advisory Committee to the OECD - Comité Consultatif Economique et Industriel Auprès de l OCDE
 Business and Industry Advisory Committee to the OECD - Comité Consultatif Economique et Industriel Auprès de l OCDE In Committee BIOTECHNOLOGY: A KEY CONTRIBUTOR TO SUSTAINABLE ECONOMIC GROWTH A Vision
Business and Industry Advisory Committee to the OECD - Comité Consultatif Economique et Industriel Auprès de l OCDE In Committee BIOTECHNOLOGY: A KEY CONTRIBUTOR TO SUSTAINABLE ECONOMIC GROWTH A Vision
The Institute of Microbiology of the Czech Academy of Sciences is looking for. four POSTDOCTORAL FELLOW positions in the fields of:
 4 Postdoctoral Fellow positions in microbiology, molecular biology and ecology The Institute of Microbiology of the Czech Academy of Sciences, Prague, Czech Republic The Institute of Microbiology of the
4 Postdoctoral Fellow positions in microbiology, molecular biology and ecology The Institute of Microbiology of the Czech Academy of Sciences, Prague, Czech Republic The Institute of Microbiology of the
Chapter 10 Microbial Genetics: New Genes for Old Germs
 Chapter 10 Microbial Genetics: New Genes for Old Germs Objectives: After reading Chapter Ten, you should understand The structure and complexity of the bacterial chromosome and the significance of plasmids.
Chapter 10 Microbial Genetics: New Genes for Old Germs Objectives: After reading Chapter Ten, you should understand The structure and complexity of the bacterial chromosome and the significance of plasmids.
Drosophila White Paper 2003 August 13, 2003
 Drosophila White Paper 2003 August 13, 2003 Explanatory Note: The first Drosophila White Paper was written in 1999. Revisions to this document were made in 2000 and the final version was published as the
Drosophila White Paper 2003 August 13, 2003 Explanatory Note: The first Drosophila White Paper was written in 1999. Revisions to this document were made in 2000 and the final version was published as the
Bruker Introduces Bologna Work ow for Rapid and Cost-E ective Clinical Microbiology Diagnosis of Bloodstream Infections with Broad Species Coverage
 NEWS RELEASE Bruker Introduces Bologna Work ow for Rapid and Cost-E ective Clinical Microbiology Diagnosis of Bloodstream Infections with Broad Species Coverage 4/20/2018 60-90 Minute Microbial ID of 2,700
NEWS RELEASE Bruker Introduces Bologna Work ow for Rapid and Cost-E ective Clinical Microbiology Diagnosis of Bloodstream Infections with Broad Species Coverage 4/20/2018 60-90 Minute Microbial ID of 2,700
ngs metagenomics target variation amplicon bioinformatics diagnostics dna trio indel high-throughput gene structural variation ChIP-seq mendelian
 Metagenomics T TM storage genetics assembly ncrna custom genotyping RNA-seq de novo mendelian ChIP-seq exome genomics indel ngs trio prediction metagenomics SNP resequencing bioinformatics diagnostics
Metagenomics T TM storage genetics assembly ncrna custom genotyping RNA-seq de novo mendelian ChIP-seq exome genomics indel ngs trio prediction metagenomics SNP resequencing bioinformatics diagnostics
Implementation Plan A collaborative national research, development and technology transfer center for tropical hardwood stewardship.
 Tropical Hardwood Tree Improvement and Regeneration Center (Tropical HTIRC) Implementation Plan 2011-2016 A collaborative national research, development and technology transfer center for tropical hardwood
Tropical Hardwood Tree Improvement and Regeneration Center (Tropical HTIRC) Implementation Plan 2011-2016 A collaborative national research, development and technology transfer center for tropical hardwood
EVOLUTION OF A MEETING: 20+ YEARS OF RESEARCH COMMUNICATION AND COORDINATION
 EVOLUTION OF A MEETING: 20+ YEARS OF RESEARCH COMMUNICATION AND COORDINATION Michael L. McManus 1 (retired) and Kurt W. Gottschalk 2 1 USDA Forest Service, Northern Research Station, Hamden CT 06514 2
EVOLUTION OF A MEETING: 20+ YEARS OF RESEARCH COMMUNICATION AND COORDINATION Michael L. McManus 1 (retired) and Kurt W. Gottschalk 2 1 USDA Forest Service, Northern Research Station, Hamden CT 06514 2
DNA/RNA MICROARRAYS NOTE: USE THIS KIT WITHIN 6 MONTHS OF RECEIPT.
 DNA/RNA MICROARRAYS This protocol is based on the EDVOTEK protocol DNA/RNA Microarrays. 10 groups of students NOTE: USE THIS KIT WITHIN 6 MONTHS OF RECEIPT. 1. EXPERIMENT OBJECTIVE The objective of this
DNA/RNA MICROARRAYS This protocol is based on the EDVOTEK protocol DNA/RNA Microarrays. 10 groups of students NOTE: USE THIS KIT WITHIN 6 MONTHS OF RECEIPT. 1. EXPERIMENT OBJECTIVE The objective of this
10. BIOTECHNOLOGY (Code No. 045)
 10. BIOTECHNOLOGY (Code No. 045) An unprecedented growth of human knowledge in the field of Biological Sciences coupled with equally significant developments in the field of technology have brought significant
10. BIOTECHNOLOGY (Code No. 045) An unprecedented growth of human knowledge in the field of Biological Sciences coupled with equally significant developments in the field of technology have brought significant
Could modern methods of genetics improve the disease resistance in farm animals?
 Could modern methods of genetics improve the disease resistance in farm animals? (introductory lecture) A. Kuznetsov Content Innate immune response Adaptive immune response Breeding programs Transgenesis
Could modern methods of genetics improve the disease resistance in farm animals? (introductory lecture) A. Kuznetsov Content Innate immune response Adaptive immune response Breeding programs Transgenesis
Research Summary 2011 Potato Pathology & Genomics University of Minnesota Professor Jim Bradeen
 Bradeen Research Summary 2011 Research Summary 2011 Potato Pathology & Genomics University of Minnesota Professor Jim Bradeen jbradeen@umn.edu 1. Understanding the Molecular Basis of Tuber Disease Resistance
Bradeen Research Summary 2011 Research Summary 2011 Potato Pathology & Genomics University of Minnesota Professor Jim Bradeen jbradeen@umn.edu 1. Understanding the Molecular Basis of Tuber Disease Resistance
UNIT ONE Performance Objective Critical Attributes Benchmarks/Assessment
 Curriculum Standard: The student will demonstrate an understanding of the ways biology affects their lives and the industry of agriculture. The student will use the scientific method and research techniques
Curriculum Standard: The student will demonstrate an understanding of the ways biology affects their lives and the industry of agriculture. The student will use the scientific method and research techniques
DNA Banking for the 21 st Century
 DNA Banking for the 21 st Century A White Paper of Recommendations from the U.S. Workshop on DNA Banking The importance of DNA data to modern studies of systematics and evolution cannot be overstated.
DNA Banking for the 21 st Century A White Paper of Recommendations from the U.S. Workshop on DNA Banking The importance of DNA data to modern studies of systematics and evolution cannot be overstated.
Forecast diagnostics for antimicrobial resistance (AMR)
 Forecast diagnostics for antimicrobial resistance (AMR) Authors: Ann Van den Bruel, Philip Turner NIHR Diagnostic Evidence Cooperative Oxford, University of Oxford Context When asked to make forecasts
Forecast diagnostics for antimicrobial resistance (AMR) Authors: Ann Van den Bruel, Philip Turner NIHR Diagnostic Evidence Cooperative Oxford, University of Oxford Context When asked to make forecasts
USGCRP/CCSP Strategic Planning Building Block Global Carbon Cycle 1
 USGCRP/CCSP Strategic Planning Building Block Global Carbon Cycle 1 Summary The global carbon cycle is changing rapidly as a result of human actions, altering Earth s climate. Many research priorities
USGCRP/CCSP Strategic Planning Building Block Global Carbon Cycle 1 Summary The global carbon cycle is changing rapidly as a result of human actions, altering Earth s climate. Many research priorities
Microbiology An Introduction Tortora Funke Case Eleventh Edition
 Microbiology An Introduction Tortora Funke Case Eleventh Edition Pearson Education Limited Edinburgh Gate Harlow Essex CM20 2JE England and Associated Companies throughout the world Visit us on the World
Microbiology An Introduction Tortora Funke Case Eleventh Edition Pearson Education Limited Edinburgh Gate Harlow Essex CM20 2JE England and Associated Companies throughout the world Visit us on the World
Agenda Item 3 CX/ASIA 12/18/3 August 2012
 E Agenda Item 3 CX/ASIA 12/18/3 August 2012 JOINT FAO/WHO FOOD STANDARDS PROGRAMME FAO/WHO COORDINATING COMMITTEE FOR ASIA 18 th Session Tokyo, Japan, 5 9 November 2012 DRAFT STRATEGIC PLAN OF THE CODEX
E Agenda Item 3 CX/ASIA 12/18/3 August 2012 JOINT FAO/WHO FOOD STANDARDS PROGRAMME FAO/WHO COORDINATING COMMITTEE FOR ASIA 18 th Session Tokyo, Japan, 5 9 November 2012 DRAFT STRATEGIC PLAN OF THE CODEX
Testimony of the. Federation of American Societies for Experimental Biology. Prepared for the. House Committee on Appropriations
 Contact: Benjamin H. Krinsky, PhD Senior Legislative Affairs Officer Federation of American Societies for Experimental Biology (FASEB) bkrinsky@faseb.org Testimony of the Federation of American Societies
Contact: Benjamin H. Krinsky, PhD Senior Legislative Affairs Officer Federation of American Societies for Experimental Biology (FASEB) bkrinsky@faseb.org Testimony of the Federation of American Societies
NIAID Strategic Plan for Biodefense Research February 2002
 NIAID Strategic Plan for Biodefense Research February 2002 U. S. DEPARTMENT OF HEALTH AND HUMAN SERVICES National Institutes of Health National Institute of Allergy and Infectious Diseases NIAID Strategic
NIAID Strategic Plan for Biodefense Research February 2002 U. S. DEPARTMENT OF HEALTH AND HUMAN SERVICES National Institutes of Health National Institute of Allergy and Infectious Diseases NIAID Strategic
Questionnaire on the use of High Throughput Sequencing, Bioinformatics and Computational Genomics (HTS-BCG) in the OIE Reference Centre network
 Questionnaire on the use of High Throughput Sequencing, Bioinformatics and Computational Genomics (HTS-BCG) in the OIE Reference Centre network Massimo Palmarini MRC-University of Glasgow Centre for Virus
Questionnaire on the use of High Throughput Sequencing, Bioinformatics and Computational Genomics (HTS-BCG) in the OIE Reference Centre network Massimo Palmarini MRC-University of Glasgow Centre for Virus
Overview of CIDT Challenges and Opportunities
 Overview of CIDT Challenges and Opportunities Peter Gerner-Smidt, MD, DSc Enteric Diseases Laboratory Branch InFORM II Phoenix, AZ, 19 November 2015 National Center for Emerging and Zoonotic Infectious
Overview of CIDT Challenges and Opportunities Peter Gerner-Smidt, MD, DSc Enteric Diseases Laboratory Branch InFORM II Phoenix, AZ, 19 November 2015 National Center for Emerging and Zoonotic Infectious
AUTHENTIC RESEARCH IN THE CLASSROOM: THE METAGENOMICS MODEL Brian Gibbens, Ph.D. 10/9/2012
 AUTHENTIC RESEARCH IN THE CLASSROOM: THE METAGENOMICS MODEL Brian Gibbens, Ph.D. 10/9/2012 1 Outline What is Authentic Research? Why is it important to study microbes? What is Metagenomics? Why Teach Metagenomics?
AUTHENTIC RESEARCH IN THE CLASSROOM: THE METAGENOMICS MODEL Brian Gibbens, Ph.D. 10/9/2012 1 Outline What is Authentic Research? Why is it important to study microbes? What is Metagenomics? Why Teach Metagenomics?
Lecture Series 10 The Genetics of Viruses and Prokaryotes
 Lecture Series 10 The Genetics of Viruses and Prokaryotes The Genetics of Viruses and Prokaryotes A. Using Prokaryotes and Viruses for Genetic Experiments B. Viruses: Reproduction and Recombination C.
Lecture Series 10 The Genetics of Viruses and Prokaryotes The Genetics of Viruses and Prokaryotes A. Using Prokaryotes and Viruses for Genetic Experiments B. Viruses: Reproduction and Recombination C.
INTRODUCTION TO GENETIC EPIDEMIOLOGY (EPID0754) Prof. Dr. Dr. K. Van Steen
 INTRODUCTION TO GENETIC EPIDEMIOLOGY (EPID0754) Prof. Dr. Dr. K. Van Steen CHAPTER 1: SETTING THE PACE 1 Course Responsible Contact details 2 Administrative Issues Course details and examination methods
INTRODUCTION TO GENETIC EPIDEMIOLOGY (EPID0754) Prof. Dr. Dr. K. Van Steen CHAPTER 1: SETTING THE PACE 1 Course Responsible Contact details 2 Administrative Issues Course details and examination methods
National Animal Genome Research Program (NSRP-8) 2018 Project proposal renewal request. Reviewer Comments and Response to Reviewers (in blue)
 National Animal Genome Research Program (NSRP-8) 2018 Project proposal renewal request Reviewer Comments and Response to Reviewers (in blue) Reviewer number 1 It is very encouraging to perceive the progress
National Animal Genome Research Program (NSRP-8) 2018 Project proposal renewal request Reviewer Comments and Response to Reviewers (in blue) Reviewer number 1 It is very encouraging to perceive the progress
New Life, Old Bottles Regulating First-Generation Products of Synthetic Biology. Michael Rodemeyer University of Virginia
 New Life, Old Bottles Regulating First-Generation Products of Synthetic Biology Michael Rodemeyer University of Virginia What is synthetic biology? An emerging set of tools and techniques driven by rapid
New Life, Old Bottles Regulating First-Generation Products of Synthetic Biology Michael Rodemeyer University of Virginia What is synthetic biology? An emerging set of tools and techniques driven by rapid
Burton's Microbiology for the Health Sciences
 Burton's Microbiology for the Health Sciences Section I. Introduction to Microbiology Burton's Microbiology for the Health Sciences Chapter 1. Microbiology - The Science 1 Chapter 1 Outline Introduction
Burton's Microbiology for the Health Sciences Section I. Introduction to Microbiology Burton's Microbiology for the Health Sciences Chapter 1. Microbiology - The Science 1 Chapter 1 Outline Introduction
Unit 8: Genomics Guided Reading Questions (150 pts total)
 Name: AP Biology Biology, Campbell and Reece, 7th Edition Adapted from chapter reading guides originally created by Lynn Miriello Chapter 18 The Genetics of Viruses and Bacteria Unit 8: Genomics Guided
Name: AP Biology Biology, Campbell and Reece, 7th Edition Adapted from chapter reading guides originally created by Lynn Miriello Chapter 18 The Genetics of Viruses and Bacteria Unit 8: Genomics Guided
Biotechnology: Unlocking Nature s Secrets
 Course Syllabus Biotechnology: Unlocking Nature s Secrets Course Description Can we bring back extinct species? Will the cures for cancer, malaria, and other diseases come from the combination of natural
Course Syllabus Biotechnology: Unlocking Nature s Secrets Course Description Can we bring back extinct species? Will the cures for cancer, malaria, and other diseases come from the combination of natural
Microbiology, Molecular Biology and Biochemistry
 Microbiology, Molecular Biology and Biochemistry Bruce L. Miller, Interim Dept. Head, Dept. of Microbiology, Molecular Biology and Biochemistry (142 Life Sc. Bldg. 83844-3052; phone 208/885-7966; mmbb@uidaho.edu;
Microbiology, Molecular Biology and Biochemistry Bruce L. Miller, Interim Dept. Head, Dept. of Microbiology, Molecular Biology and Biochemistry (142 Life Sc. Bldg. 83844-3052; phone 208/885-7966; mmbb@uidaho.edu;
Lecture Four. Molecular Approaches I: Nucleic Acids
 Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
DEPARTMENT: MICROBIOLOGY PROGRAMME: B SC. Statements of Programme Specific Outcomes (PSOs)
 DEPARTMENT: MICROBIOLOGY PROGRAMME: B SC Statements of Programme Specific Outcomes (PSOs) 1. Understand the contributions of various scientist in microbiology and scope of various branches of it 2. Understand
DEPARTMENT: MICROBIOLOGY PROGRAMME: B SC Statements of Programme Specific Outcomes (PSOs) 1. Understand the contributions of various scientist in microbiology and scope of various branches of it 2. Understand
Warm-Up. Describe how the Hershey-Chase experiment proved that DNA is the heritable molecule of genes.
 Warm-Up Describe how the Hershey-Chase experiment proved that DNA is the heritable molecule of genes. Yesterday s Pictures 3D χ " = $ (o e)" e DNA protein protein protein protein protein Host Cell DNA
Warm-Up Describe how the Hershey-Chase experiment proved that DNA is the heritable molecule of genes. Yesterday s Pictures 3D χ " = $ (o e)" e DNA protein protein protein protein protein Host Cell DNA
Course outcomes. M.Sc Microbiology (CBCS) Code Course title Course Type HPW Credits
 Course outcomes M.Sc Microbiology (CBCS) Semester-I General MB 101 Microbiology General Microbiology This focuses on general principles of microbiology, microbial cell structure and function and their
Course outcomes M.Sc Microbiology (CBCS) Semester-I General MB 101 Microbiology General Microbiology This focuses on general principles of microbiology, microbial cell structure and function and their
National Institute of Biology - NIB
 National Institute of Biology - NIB NIB was established by the Republic of Slovenia as a public, non-profit research institute in 1960 and is today the leading research institution in biological sciences.
National Institute of Biology - NIB NIB was established by the Republic of Slovenia as a public, non-profit research institute in 1960 and is today the leading research institution in biological sciences.
A Study on the Validity of Perceived Risks versus Benefits of Genetically Engineered Foods. among US Consumers. Gaganjot Kaur
 A Study on the Validity of Perceived Risks versus Benefits of Genetically Engineered Foods among US Consumers Gaganjot Kaur Assistant Professor, Apex Institute of Technology, University Business School,
A Study on the Validity of Perceived Risks versus Benefits of Genetically Engineered Foods among US Consumers Gaganjot Kaur Assistant Professor, Apex Institute of Technology, University Business School,
SPO GRANTS (2001) PHASES
 2002 SPO GRANTS (2001) PHASES MAIN PHASES Legal organisation of the Institute of Biotechnology Ankara University Biotechnology Labs Organisation of Central Lab of the Institute of Biotechnology/ Purchase
2002 SPO GRANTS (2001) PHASES MAIN PHASES Legal organisation of the Institute of Biotechnology Ankara University Biotechnology Labs Organisation of Central Lab of the Institute of Biotechnology/ Purchase
Crop Biosecurity: Are We Prepared?
 Crop Biosecurity: Are We Prepared? White Paper Developed by the Public Policy Board of the American Phytopathological Society (APS) May 2003 Drafting Committee: Dr. John L. Sherwood Chair APS Public Policy
Crop Biosecurity: Are We Prepared? White Paper Developed by the Public Policy Board of the American Phytopathological Society (APS) May 2003 Drafting Committee: Dr. John L. Sherwood Chair APS Public Policy
RISK ASSESSMENT OF GENETICALLY MODIFIED MICRO-ORAGNISMS: A FORMAT THAT OFFERS ONE POSSIBLE WAY OF ACHIEVING GOOD PRACTICE
 RISK ASSESSMENT OF GENETICALLY MODIFIED MICRO-ORAGNISMS: A FORMAT THAT OFFERS ONE POSSIBLE WAY OF ACHIEVING GOOD PRACTICE 1 Regulation 6 of the Genetically Modified Organisms (Contained Use) Regulations
RISK ASSESSMENT OF GENETICALLY MODIFIED MICRO-ORAGNISMS: A FORMAT THAT OFFERS ONE POSSIBLE WAY OF ACHIEVING GOOD PRACTICE 1 Regulation 6 of the Genetically Modified Organisms (Contained Use) Regulations
