Mice and Poly-I:C treatment PCR and immunoblotting
|
|
|
- Andra Crawford
- 5 years ago
- Views:
Transcription
1 Mice and Poly-I:C treatment All experimental mice were housed in Sanford/Burnham Medical Research Institute in pathogen-free environment with controlled temperature and humidity. Shp2 flox/flox :Mx1-Cre - mice were bred with Shp2 flox/flox :Mx1-Cre + mice to produce control and knockout littermates. Pten flox/flox mice were purchased from Jackson Laboratory and those mice were original deposited by Dr. Hong Wu 1. Pten flox/flox mice were bred with Shp2 flox/flox :Mx1-Cre + to produce Shp2 flox/flox :Pten flox/flox :Mx1-Cre + double knockout animals. CD mice were kindly donated by Dr. Robert Rickert. Poly-I:C was reconstituted according to manufacturer s (GE Amersham) instructions. Five doses of Poly-I:C (10 mg/kg) were given to control and experimental groups by intraperitoneal injection every other day. PCR and immunoblotting DNA from peripheral blood was extracted using DNeasy Blood & Tissue Kit (Qiagen). Primers for genotyping are 5 AAGAGGAAATAGGAAGCATTGAGGA3 and 5 TAGGGAATGTGACAAGAAAGCAGTC3. PCR products for floxed and knockout alleles are 1000 bp and 400 bp, respectively. For immunoblotting, cells were lysed in RIPA buffer and protein extract was separated by SDS-PAGE and then transferred to PVDF membrane. Home-made anti-shp2 and anti-erk antibodies were used to detect Shp2 and Erk expression. Antibody against GFP (Santa Cruz), Flag (cell signaling), Gata2 (Santa Cruz) were used to detect respective proteins. For SCF stimulation, EML cells were starved 6 hours in UltraCULTURE serum free medium (Lonza) supplemented with 2 mm glutamine and then stimulated with 100 ng/ml recombinant mouse SCF (R&D) for indicated time periods. For detection of phosphorylated protein, anti-phospho AKT(Ser 473), anti-phospho Erk (Thr202/Tyr204), anti-phospho Stat3(Tyr727), anti-phospho JNK (Thr183/Tyr185), anti-phospho p38 (Thr180/Tyr182) (all from Cell Signaling) were used. Antibody against Tubulin (Sigma), AKT, Stat3, p38 (Cell Signaling) and JNK (Santa Cruz) were used to detect the specific proteins.
2 CBC count Peripheral blood was collected trough intra orbital bleeding. Complete blood cell (CBC) counting was performed with the VetScan HMII Hematology System (Abaxis). In vitro CFU and in vivo CFU-S assays For in vitro colony forming unit (CFU) assay, 50,000 bone marrow cells were seeded in MethoCult GF M3434 media (Stemcell Technologies) according to the manufacturer s recommendations. Colonies were visualized and counted microscopically at day 14. The experiment was performed in duplicate for each sample. To assess the CFU of LSK cells, 250 LSK cells were sorted out and seeded into MethoCult GF M3434 media (Stemcell Technologies). For in vivo CFU-S assay, 10 5 bone marrow cells were transplanted to lethally irradiated recipient mice. Spleens were harvested from recipients 12 days post transplantation and fixed in Bouin s solution for colony numeration Flow cytometry Lineage staining was performed using mouse hematopoietic lineage flow panel from ebioscience (biotin-conjugated antibodies against CD3 (145-2C11), CD45R/B220 (RA3-6B2), CD11b (M1/70), TER-119, Ly-6G (RB6-8C5)) supplemented with biotin-conjugated antibodies against mouse CD4 (L3T4), mouse CD8 (CD8b, Ly-3) (ebioscience). Streptavidin conjugated with APC-CY7 was used for secondary staining. For further staining of HSCs, antibody against CD117 (ckit)-pe, FITC, PE-CY5(2B8); Sca1-FITC, APC (D7); Flk2-PE-CY5(A2F10); CD150-APC(mShad150); CD48-PE (HM48-1 ) (ebioscience); CD34-APC, PE-CY5 (MEC14.7) (Biolegend) were used. In the competitive reconstitution experiment, we used anti-mouse CD45.1-FITC (A20) and antimouse CD45.2-FITC (104) antibodies (ebioscience). Lineage - cells were enriched by lineage depletion kit (Miltenyi) according to manufacturer s instruction. For myeloid and lymphoid progenitor cell staining, additional antibodies against mouse IL-7Ra-PE-CY5 (A7R34), anti-mouse CD16/32-PE(93) (ebioscience) were used. CMPs were gated as Lin - Sca-1 - c-kit + FcgR lo CD34 +. CLPs were gated as Lin - IL-7R - Sca-1 lo c-kit lo. GMPs were
3 gated as Lin - Sca-1 - c-kit + FcgR hi CD34 +. MEPs were gated as Lin - Sca-1 - c-kit + FcgR lo CD34 -. Correspondent isotype controls (ebioscience or Santa Cruz) were used for proper gating. Cell cycle analysis Pyronin Y/Hoechest33342 co-staining was used to distinguish the G0/G1 phase of LSK cells. Around 50,000 LSK cells were sorted into ice-cold 100% ethanol and incubated for at least 16 hours. We developed a new method of carrier cells to avoid the loss of LSK cells during the sequential staining process. EML cell line is a bone marrow progenitor cell line. EML cells were stained with 1 μm DDAO-SE (Invitrogen) and then fixed in ice-cold 100% ethanol for at least 16 hours. EML cells were mixed with LSK cells and stained in 0.5 μg/ml Pyronin Y and 2 μg/ml Hoechest33342, 45 mins before FACS analysis. We gated on DDAO-SE - cells for additional G 0 /G 1 gating of LSK cells. For the 5-FU challenge assay, one week after the final injection of poly-i:c, fl/fl:cre + or fl/fl:cre mice were injected intraperitoneally with three sequential doses of 5-FU (150 mg/kg, Fluka) every 7 days. Mice were monitored carefully every morning for consecutive 28 days. Homing, migration assay For the in vivo homing assay, lineage-depleted bone marrow cells were stained with 5 μm CFDA-SE (invitrogen) for 5 mins and injected through tail vein into lethally irradiated recipient mice. Bone marrow from recipients was harvested and analyzed 16 hours later. CDFA-SE + cells were considered as homed cells. To assess the homed progenitor cells, we adopted the method as previously described 2. Five million bone marrow nucleated cells from donors were transplanted into lethally irradiated recipient mice. Recipients were euthanized 24 hours later to harvest the femur. Nucleated cells (0.9x10 5 to 2.5x10 5 ) were plated in MethoCult 3434 medium (stem cell technologies). Colonies were numerated on day 8 and normalized against the seeding cell number. Bone marrow cells from lethally irradiated mice without transplantation were also plated as negative control and no colonies were detectable. For the in vitro migration assay, 10 6 lineage-depleted bone marrow cells were seeded to the top chamber of 5 μm pore size 24-well transwell plate (Corning) and migrated against 5 nm SCF. Migrated cells were numerated after 3-hour incubation at 37 C, 5%CO 2.
4 CHIP assay The CHIP assay was conducted as described before 3. Antibody against mouse Gata2 (sc-8334 Santa Cruz) was used. The primers used for Gata2 binding to kit promoter were: Forward 5' GTG GTT GTC CTT TAT TGT CTA GGG 3' ; Reverse5' GCT GTG CCC TCT AAG ACC AGG AG 3'. The primers used for Gata2 binding to c-kit -114kb and +5kb region were adopted from a previous publication 4. Q-PCR primers and gene expression analysis SYBR green qpcr master mix (stratagene) was used for quantitative-pcr assay. Target gene expression was calculated by ΔCT method. Most q-pcr primers in this study were adopted from PrimerBank ( Others were designed using invitrogen online tool OligoPerfect Designer. ( shp2 forward 5' T C C A T G G T C A C T T G T C T G G A 3' shp2 reverse 5' G A C G T G G G T C A C T T T G G A C T 3' c-kit forward 5' GCC ACG TCT CAG CCA TCT G 3' c-kit reverse 5' GTC GCC AGC TTC AAC TAT TAA CT 3' gata1 forward 5' TGG GGA CCT CAG AAC CCT TG 3' gata1 reverse 5' GGC TGC ATT TGG GGA AGT G 3' gata 2 forward 5' CAC CCC GCC GTA TTG AAT G 3' gata 2 reverse 5' CCT GCG AGT CGA GAT GGT TG 3' sp1 forward 5' GCC GCC TTT TCT CAG ACT C 3' sp11 reverse 5' TTG GGT GAC TCA ATT CTG CTG 3' scl forward 5' CAC TAG GCA GTG GGT TCT TTG 3' scl reverse 5' GGT GTG AGG ACC ATC AGA AAT CT 3' Retroviral infection of bone marrow cells
5 Retroviral construct MSCV-Cre-IRES-hCD2 that co-expresses Cre recombinase and human CD2 (hcd2) marker and control construct MSCV-hCD2 was a gift from Robert Rickert. Retrovirus was prepared using packaging cell line Plat-E following manufacture s protocol (Cell Biolabs). 2-3 months old +/+ or fl/fl mice were injected with 500 μl (10 mg/ml) 5-FU and bone marrow was harvested 5 days after the injection. Bone marrow cells were pre-stimulated with DMEM medium containing 15% FBS and 10 ng/μl recombinant mouse IL-6, 6 ng/μl recombinant mouse IL-3 (PropTech), 100 ng/μl recombinant mouse SCF (R&D) and 10% WEHI conditional medium (BD bioscience) for 24 hrs. Retroviral infection was performed by spin inoculation at 3000 rpm for 90 mins. Bone marrow cells were stained with PE-anti-hCD2 antibody (Biolegend) 48 hrs later to detect positively infected cells. REFERENCES 1. Groszer M, Erickson R, Scripture-Adams DD, et al. Negative regulation of neural stem/progenitor cell proliferation by the Pten tumor suppressor gene in vivo. Science. 2001;294: Katayama Y, Hidalgo A, Furie BC, Vestweber D, Furie B, Frenette PS. PSGL-1 participates in E-selectin-mediated progenitor homing to bone marrow: evidence for cooperation between E-selectin ligands and alpha4 integrin. Blood. 2003;102: Zhang SS, Hao E, Yu J, et al. Coordinated regulation by Shp2 tyrosine phosphatase of signaling events controlling insulin biosynthesis in pancreatic beta-cells. Proc Natl Acad Sci U S A. 2009;106: Jing H, Vakoc CR, Ying L, et al. Exchange of GATA factors mediates transitions in looped chromatin organization at a developmentally regulated gene locus. Mol Cell. 2008;29:
6 Figure S1. Related to Fig. 1 (A, B, C) Comparable bone marrow cellularity (A), bone marrow LSK frequency (B) and CBC counts among different control genotypes as shown (C). Figure S2. Related to Fig. 3 (A, B) competitive reconstitution assay with donor to competitor ratio at 4:1 in the injection first/transplantation later setting (A) or transplantation first/injection later setting (B) (n = 4-9). (C) Reciprocal bone marrow transplantation. Poly-I:C treated fl/fl:cre - or fl/fl:cre + mice were lethally irradiated and used as recipients. Wild-type whole bone marrow or sorted LSK cells from CD45.1 mice were used as donor cells. Hematopoiesis in recipient mice was monitored at different time points post transplantation. (D) CBC counts in the recipients of whole bone marrow reciprocal transplantation. No difference was detected between wild-type and knockout groups (n = 5). (E) CBC counts in the recipients of LSK cell reciprocal transplantation. No difference was detected between wild-type and knockout groups (n = 5). Figure S3. Related to Fig. 5 (A) Representative FACS plots illustrate Pyronin Y/Hoechest33342 staining in LSK cells. (B) Quantification of G 0, G 1 and S-G 2 -M phase cell percentage in LSK subpopulation. More knockout LSK cells entered G 1 phase, with reduced number in the quiescent G 0 phase. Results were obtained from 3 independent experiments. (C) Increased mortality rate to 5-FU myeloablation in Shp2 knockout animals. Percentage of survived animals was shown. Data were analyzed by log-rank (Mantel-Cox) test (n=11). (D) Representative FACS plots illustrate the Annexin V staining of different bone marrow compartments.
7 Figure S4. Related to Fig. 6 (A) Kit-staining profile of lineage - fl/fl bone marrow infected with control retrovirus or Cre expressing retrovirus. Cre expression led to loss of Kit + cells in lineage - gated population. (B) Control +/+ or fl/fl bone marrow was infected with Cre-IRES-hCD2 retrovirus and transplanted into lethally irradiated mice. As hcd2 transcription is under control by the same promoter for Cre, hcd2 detection on cell surface was used as marker for Cre-expressing cells. The right panel is a representative FACS plot for hcd2 on day 0 of transplantation and left panel is representative FACS plot for hcd2 on day 30 of transplantation. On day 0, around 50% of bone marrow cells either from +/+ or fl/fl mice were hcd2-positive. On day 30, hcd2-positive fl/fl bone marrow cells were barely detectable, while the chimerism of +/+ cells remained higher than 50%. Figure S5. Related to Fig. 7 (A) qrt-pcr of Gata1, Gata2, Scl and Sp1 mrna in sorted 34 - LSK and 34 + LSK cells. β-actin was used as reference gene for normalization. (B) The mean fluorescent intensity (MFI) of Kit staining of single or combinatorial chemical inhibitors (LY294002, U0126 and AG490) treated EML cells. (C) CHIP assay demonstrated the binding of Gata2 to Kit promoter was significantly reduced in single (25 μm LY294002, 10 μm U0126 and 25 μm AG490) or combined (500nm) chemical inhibitors treated EML cells.
8
9
10
11
12
Supplemental Information Inventory
 Cell Stem Cell, Volume 6 Supplemental Information Distinct Hematopoietic Stem Cell Subtypes Are Differentially Regulated by TGF-β1 Grant A. Challen, Nathan C. Boles, Stuart M. Chambers, and Margaret A.
Cell Stem Cell, Volume 6 Supplemental Information Distinct Hematopoietic Stem Cell Subtypes Are Differentially Regulated by TGF-β1 Grant A. Challen, Nathan C. Boles, Stuart M. Chambers, and Margaret A.
Supplementary Figure 1A A404 Cells +/- Retinoic Acid
 Supplementary Figure 1A A44 Cells +/- Retinoic Acid 1 1 H3 Lys4 di-methylation SM-actin VEC cfos (-) RA (+) RA 14 1 1 8 6 4 H3 Lys79 di-methylation SM-actin VEC cfos (-) RA (+) RA Supplementary Figure
Supplementary Figure 1A A44 Cells +/- Retinoic Acid 1 1 H3 Lys4 di-methylation SM-actin VEC cfos (-) RA (+) RA 14 1 1 8 6 4 H3 Lys79 di-methylation SM-actin VEC cfos (-) RA (+) RA Supplementary Figure
Supplemental Data Supplemental Figure 1.
 Supplemental Data Supplemental Figure 1. Silique arrangement in the wild-type, jhs, and complemented lines. Wild-type (WT) (A), the jhs1 mutant (B,C), and the jhs1 mutant complemented with JHS1 (Com) (D)
Supplemental Data Supplemental Figure 1. Silique arrangement in the wild-type, jhs, and complemented lines. Wild-type (WT) (A), the jhs1 mutant (B,C), and the jhs1 mutant complemented with JHS1 (Com) (D)
Dierks Supplementary Fig. S1
 Dierks Supplementary Fig. S1 ITK SYK PH TH K42R wt K42R (kinase deficient) R29C E42K Y323F R29C E42K Y323F (reduced phospholipid binding) (enhanced phospholipid binding) (reduced Cbl binding) E42K Y323F
Dierks Supplementary Fig. S1 ITK SYK PH TH K42R wt K42R (kinase deficient) R29C E42K Y323F R29C E42K Y323F (reduced phospholipid binding) (enhanced phospholipid binding) (reduced Cbl binding) E42K Y323F
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
Supplementary Figures
 Supplementary Figures Supplementary Fig. 1 Characterization of GSCs. a. Immunostaining of primary GSC spheres from GSC lines. Nestin (neural progenitor marker, red), TLX (green). Merged images of nestin,
Supplementary Figures Supplementary Fig. 1 Characterization of GSCs. a. Immunostaining of primary GSC spheres from GSC lines. Nestin (neural progenitor marker, red), TLX (green). Merged images of nestin,
Hes6. PPARα. PPARγ HNF4 CD36
 SUPPLEMENTARY INFORMATION Supplementary Table Positions and Sequences of ChIP primers -63 AGGTCACTGCCA -79 AGGTCTGCTGTG Hes6-0067 GGGCAaAGTTCA ACOT -395 GGGGCAgAGTTCA PPARα -309 GGCTCAaAGTTCAaGTTCA CPTa
SUPPLEMENTARY INFORMATION Supplementary Table Positions and Sequences of ChIP primers -63 AGGTCACTGCCA -79 AGGTCTGCTGTG Hes6-0067 GGGCAaAGTTCA ACOT -395 GGGGCAgAGTTCA PPARα -309 GGCTCAaAGTTCAaGTTCA CPTa
Supplementary. Table 1: Oligonucleotides and Plasmids. complementary to positions from 77 of the SRα '- GCT CTA GAG AAC TTG AAG TAC AGA CTG C
 Supplementary Table 1: Oligonucleotides and Plasmids 913954 5'- GCT CTA GAG AAC TTG AAG TAC AGA CTG C 913955 5'- CCC AAG CTT ACA GTG TGG CCA TTC TGC TG 223396 5'- CGA CGC GTA CAG TGT GGC CAT TCT GCT G
Supplementary Table 1: Oligonucleotides and Plasmids 913954 5'- GCT CTA GAG AAC TTG AAG TAC AGA CTG C 913955 5'- CCC AAG CTT ACA GTG TGG CCA TTC TGC TG 223396 5'- CGA CGC GTA CAG TGT GGC CAT TCT GCT G
Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC
 Supplementary Appendixes Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC ACG TAG CTC CGG CTG GA-3 for vimentin, /5AmMC6/TCC CTC GCG CGT GGC TTC CGC
Supplementary Appendixes Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC ACG TAG CTC CGG CTG GA-3 for vimentin, /5AmMC6/TCC CTC GCG CGT GGC TTC CGC
Anti-Pim-1 (Cat#3247), anti-met (Cat#3127), anti-ron (Cat#2654), Anti-EGFR
 Supplementary Methods Antibodies Anti-Pim-1 (Cat#3247), anti-met (Cat#3127), anti-ron (Cat#2654), Anti-EGFR (Cat#2646), anti-igf1r (Cat#3018), anti-insr (Cat#3020), anti-akt (pan, Cat#4691), anti-phospho-akt
Supplementary Methods Antibodies Anti-Pim-1 (Cat#3247), anti-met (Cat#3127), anti-ron (Cat#2654), Anti-EGFR (Cat#2646), anti-igf1r (Cat#3018), anti-insr (Cat#3020), anti-akt (pan, Cat#4691), anti-phospho-akt
Supplemental Data. mir156-regulated SPL Transcription. Factors Define an Endogenous Flowering. Pathway in Arabidopsis thaliana
 Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
Supplemental Information. Human Senataxin Resolves RNA/DNA Hybrids. Formed at Transcriptional Pause Sites. to Promote Xrn2-Dependent Termination
 Supplemental Information Molecular Cell, Volume 42 Human Senataxin Resolves RNA/DNA Hybrids Formed at Transcriptional Pause Sites to Promote Xrn2-Dependent Termination Konstantina Skourti-Stathaki, Nicholas
Supplemental Information Molecular Cell, Volume 42 Human Senataxin Resolves RNA/DNA Hybrids Formed at Transcriptional Pause Sites to Promote Xrn2-Dependent Termination Konstantina Skourti-Stathaki, Nicholas
Supplementary methods. RNA isolation, cdna synthesis, and quantitative real-time PCR. Total RNAs were
 Supplementary methods RNA isolation, cdna synthesis, and quantitative real-time PCR. Total RNAs were extracted from cells by using an RNeasy Mini Kit (QIAGEN) and then reverse-transcribed using the SuperScript
Supplementary methods RNA isolation, cdna synthesis, and quantitative real-time PCR. Total RNAs were extracted from cells by using an RNeasy Mini Kit (QIAGEN) and then reverse-transcribed using the SuperScript
Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis
 1 2 3 4 5 6 7 8 9 10 11 12 Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis Information Research). Exons
1 2 3 4 5 6 7 8 9 10 11 12 Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis Information Research). Exons
Overexpression Normal expression Overexpression Normal expression. 26 (21.1%) N (%) P-value a N (%)
 SUPPLEMENTARY TABLES Table S1. Alteration of ZNF322A protein expression levels in relation to clinicopathological parameters in 123 Asian and 74 Caucasian lung cancer patients. Asian patients Caucasian
SUPPLEMENTARY TABLES Table S1. Alteration of ZNF322A protein expression levels in relation to clinicopathological parameters in 123 Asian and 74 Caucasian lung cancer patients. Asian patients Caucasian
Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of
 Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of the cell line) were immunostained for HA, acetylated
Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of the cell line) were immunostained for HA, acetylated
SUPPLEMENTARY INFORMATION
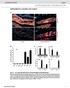 doi: 10.1038/nature07182 SUPPLEMENTAL FIGURES AND TABLES Fig. S1. myf5-expressing cells give rise to brown fat depots and skeletal muscle (a) Perirenal BAT from control (cre negative) and myf5-cre:r26r3-yfp
doi: 10.1038/nature07182 SUPPLEMENTAL FIGURES AND TABLES Fig. S1. myf5-expressing cells give rise to brown fat depots and skeletal muscle (a) Perirenal BAT from control (cre negative) and myf5-cre:r26r3-yfp
PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells
 Supplementary Information for: PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells Ju Hye Jang 1, Hyun Kim 2, Mi Jung Jang 2, Ju Hyun Cho 1,2,* 1 Research Institute
Supplementary Information for: PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells Ju Hye Jang 1, Hyun Kim 2, Mi Jung Jang 2, Ju Hyun Cho 1,2,* 1 Research Institute
hcd1tg/hj1tg/ ApoE-/- hcd1tg/hj1tg/ ApoE+/+
 ApoE+/+ ApoE-/- ApoE-/- H&E (1x) Supplementary Figure 1. No obvious pathology is observed in the colon of diseased ApoE-/me. Colon samples were fixed in 1% formalin and laid out in Swiss rolls for paraffin
ApoE+/+ ApoE-/- ApoE-/- H&E (1x) Supplementary Figure 1. No obvious pathology is observed in the colon of diseased ApoE-/me. Colon samples were fixed in 1% formalin and laid out in Swiss rolls for paraffin
Nongenetic Reprogramming of the Ligand Specificity. of Growth Factor Receptors by Bispecific DNA Aptamers
 Supporting Information For Nongenetic Reprogramming of the Ligand Specificity of Growth Factor Receptors by Bispecific DNA Aptamers Ryosuke Ueki,* Saki Atsuta, Ayaka Ueki and Shinsuke Sando* Department
Supporting Information For Nongenetic Reprogramming of the Ligand Specificity of Growth Factor Receptors by Bispecific DNA Aptamers Ryosuke Ueki,* Saki Atsuta, Ayaka Ueki and Shinsuke Sando* Department
Arabidopsis actin depolymerizing factor AtADF4 mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB
 Arabidopsis actin depolymerizing factor mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB Files in this Data Supplement: Supplemental Table S1 Supplemental Table
Arabidopsis actin depolymerizing factor mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB Files in this Data Supplement: Supplemental Table S1 Supplemental Table
MicroRNAs Modulate Hematopoietic Lineage Differentiation
 Chen et al., page 1 MicroRNAs Modulate Hematopoietic Lineage Differentiation Chang-Zheng Chen, Ling Li, Harvey F. Lodish, David. Bartel Supplemental Online Material Methods Cell isolation Murine bone marrow
Chen et al., page 1 MicroRNAs Modulate Hematopoietic Lineage Differentiation Chang-Zheng Chen, Ling Li, Harvey F. Lodish, David. Bartel Supplemental Online Material Methods Cell isolation Murine bone marrow
Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR
 Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR 1 The problem We wish to clone a yet unknown gene from a known
Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR 1 The problem We wish to clone a yet unknown gene from a known
Supplemental Information. Andrianne, Assabban et al.
 Supplemental Information. Andrianne, Assabban et al. Figure S1. Related to Figure 1. Sensitivity of Zfp36 -/- mice to Imiquimod treatment following different experimental protocols. Zfp36-deficient mice
Supplemental Information. Andrianne, Assabban et al. Figure S1. Related to Figure 1. Sensitivity of Zfp36 -/- mice to Imiquimod treatment following different experimental protocols. Zfp36-deficient mice
SUPPLEMENTARY INFORMATION
 doi:1.138/nature11496 Cl. 8 Cl. E93 Rag1 -/- 3H9 + BM Rag1 -/- BM CD CD c-kit c-kit c-kit wt Spleen c-kit B22 B22 IgM IgM IgM Supplementary Figure 1. FACS analysis of single-cell-derived pre-b cell clones.
doi:1.138/nature11496 Cl. 8 Cl. E93 Rag1 -/- 3H9 + BM Rag1 -/- BM CD CD c-kit c-kit c-kit wt Spleen c-kit B22 B22 IgM IgM IgM Supplementary Figure 1. FACS analysis of single-cell-derived pre-b cell clones.
Materials Protein synthesis kit. This kit consists of 24 amino acids, 24 transfer RNAs, four messenger RNAs and one ribosome (see below).
 Protein Synthesis Instructions The purpose of today s lab is to: Understand how a cell manufactures proteins from amino acids, using information stored in the genetic code. Assemble models of four very
Protein Synthesis Instructions The purpose of today s lab is to: Understand how a cell manufactures proteins from amino acids, using information stored in the genetic code. Assemble models of four very
Project 07/111 Final Report October 31, Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines
 Project 07/111 Final Report October 31, 2007. Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines Project Leader: Dr Douglas C. Hodgins (519-824-4120 Ex 54758, fax 519-824-5930)
Project 07/111 Final Report October 31, 2007. Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines Project Leader: Dr Douglas C. Hodgins (519-824-4120 Ex 54758, fax 519-824-5930)
Supplemental Table 1. Mutant ADAMTS3 alleles detected in HEK293T clone 4C2. WT CCTGTCACTTTGGTTGATAGC MVLLSLWLIAAALVEVR
 Supplemental Dataset Supplemental Table 1. Mutant ADAMTS3 alleles detected in HEK293T clone 4C2. DNA sequence Amino acid sequence WT CCTGTCACTTTGGTTGATAGC MVLLSLWLIAAALVEVR Allele 1 CCTGTC------------------GATAGC
Supplemental Dataset Supplemental Table 1. Mutant ADAMTS3 alleles detected in HEK293T clone 4C2. DNA sequence Amino acid sequence WT CCTGTCACTTTGGTTGATAGC MVLLSLWLIAAALVEVR Allele 1 CCTGTC------------------GATAGC
Y-chromosomal haplogroup typing Using SBE reaction
 Schematic of multiplex PCR followed by SBE reaction Multiplex PCR Exo SAP purification SBE reaction 5 A 3 ddatp ddgtp 3 T 5 A G 3 T 5 3 5 G C 5 3 3 C 5 ddttp ddctp 5 T 3 T C 3 A 5 3 A 5 5 C 3 3 G 5 3 G
Schematic of multiplex PCR followed by SBE reaction Multiplex PCR Exo SAP purification SBE reaction 5 A 3 ddatp ddgtp 3 T 5 A G 3 T 5 3 5 G C 5 3 3 C 5 ddttp ddctp 5 T 3 T C 3 A 5 3 A 5 5 C 3 3 G 5 3 G
Supplemental Table 1. Primers used for PCR.
 Supplemental Table 1. Primers used for PCR. Gene Type Primer Sequence Genotyping and semi-quantitative RT-PCR F 5 -TTG CCC GAT CAC CAT CTG TA-3 rwa1-1 R 5 -TGT AGC GAT CAA GGC CTG ATC TAA-3 LB 5 -TAG CAT
Supplemental Table 1. Primers used for PCR. Gene Type Primer Sequence Genotyping and semi-quantitative RT-PCR F 5 -TTG CCC GAT CAC CAT CTG TA-3 rwa1-1 R 5 -TGT AGC GAT CAA GGC CTG ATC TAA-3 LB 5 -TAG CAT
Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH).
 Bisulfite Treatment of DNA Dilute DNA sample to 2µg DNA in 50µl ddh 2 O. Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH). Incubate in a 37ºC water bath for 30 minutes. To 55µl samples
Bisulfite Treatment of DNA Dilute DNA sample to 2µg DNA in 50µl ddh 2 O. Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH). Incubate in a 37ºC water bath for 30 minutes. To 55µl samples
Cat. # Product Size DS130 DynaExpress TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1
 Product Name: Kit Component TA PCR Cloning Kit (ptakn-2) Cat. # Product Size DS130 TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1 2 Ligation Buffer
Product Name: Kit Component TA PCR Cloning Kit (ptakn-2) Cat. # Product Size DS130 TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1 2 Ligation Buffer
Disease and selection in the human genome 3
 Disease and selection in the human genome 3 Ka/Ks revisited Please sit in row K or forward RBFD: human populations, adaptation and immunity Neandertal Museum, Mettman Germany Sequence genome Measure expression
Disease and selection in the human genome 3 Ka/Ks revisited Please sit in row K or forward RBFD: human populations, adaptation and immunity Neandertal Museum, Mettman Germany Sequence genome Measure expression
ΔPDD1 x ΔPDD1. ΔPDD1 x wild type. 70 kd Pdd1. Pdd3
 Supplemental Fig. S1 ΔPDD1 x wild type ΔPDD1 x ΔPDD1 70 kd Pdd1 50 kd 37 kd Pdd3 Supplemental Fig. S1. ΔPDD1 strains express no detectable Pdd1 protein. Western blot analysis of whole-protein extracts
Supplemental Fig. S1 ΔPDD1 x wild type ΔPDD1 x ΔPDD1 70 kd Pdd1 50 kd 37 kd Pdd3 Supplemental Fig. S1. ΔPDD1 strains express no detectable Pdd1 protein. Western blot analysis of whole-protein extracts
Supporting information for Biochemistry, 1995, 34(34), , DOI: /bi00034a013
 Supporting information for Biochemistry, 1995, 34(34), 10807 10815, DOI: 10.1021/bi00034a013 LESNIK 10807-1081 Terms & Conditions Electronic Supporting Information files are available without a subscription
Supporting information for Biochemistry, 1995, 34(34), 10807 10815, DOI: 10.1021/bi00034a013 LESNIK 10807-1081 Terms & Conditions Electronic Supporting Information files are available without a subscription
Supplemental material
 Supplemental material Diversity of O-antigen repeat-unit structures can account for the substantial sequence variation of Wzx translocases Yaoqin Hong and Peter R. Reeves School of Molecular Bioscience,
Supplemental material Diversity of O-antigen repeat-unit structures can account for the substantial sequence variation of Wzx translocases Yaoqin Hong and Peter R. Reeves School of Molecular Bioscience,
Flow cytometry Stained cells were analyzed and sorted by SORP FACS Aria (BD Biosciences).
 Mice C57BL/6-Ly5.1 or -Ly5.2 congenic mice were used for LSK transduction and competitive repopulation assays. Animal care was in accordance with the guidelines of Keio University for animal and recombinant
Mice C57BL/6-Ly5.1 or -Ly5.2 congenic mice were used for LSK transduction and competitive repopulation assays. Animal care was in accordance with the guidelines of Keio University for animal and recombinant
Supporting Information
 Supporting Information Table S1. Oligonucleotide sequences used in this work Oligo DNA A B C D CpG-A CpG-B CpG-C CpG-D Sequence 5 ACA TTC CTA AGT CTG AAA CAT TAC AGC TTG CTA CAC GAG AAG AGC CGC CAT AGT
Supporting Information Table S1. Oligonucleotide sequences used in this work Oligo DNA A B C D CpG-A CpG-B CpG-C CpG-D Sequence 5 ACA TTC CTA AGT CTG AAA CAT TAC AGC TTG CTA CAC GAG AAG AGC CGC CAT AGT
PCR analysis was performed to show the presence and the integrity of the var1csa and var-
 Supplementary information: Methods: Table S1: Primer Name Nucleotide sequence (5-3 ) DBL3-F tcc ccg cgg agt gaa aca tca tgt gac tg DBL3-R gac tag ttt ctt tca ata aat cac tcg c DBL5-F cgc cct agg tgc ttc
Supplementary information: Methods: Table S1: Primer Name Nucleotide sequence (5-3 ) DBL3-F tcc ccg cgg agt gaa aca tca tgt gac tg DBL3-R gac tag ttt ctt tca ata aat cac tcg c DBL5-F cgc cct agg tgc ttc
Legends for supplementary figures 1-3
 High throughput resistance profiling of Plasmodium falciparum infections based on custom dual indexing and Illumina next generation sequencing-technology Sidsel Nag 1,2 *, Marlene D. Dalgaard 3, Poul-Erik
High throughput resistance profiling of Plasmodium falciparum infections based on custom dual indexing and Illumina next generation sequencing-technology Sidsel Nag 1,2 *, Marlene D. Dalgaard 3, Poul-Erik
II 0.95 DM2 (RPP1) DM3 (At3g61540) b
 Table S2. F 2 Segregation Ratios at 16 C, Related to Figure 2 Cross n c Phenotype Model e 2 Locus A Locus B Normal F 1 -like Enhanced d Uk-1/Uk-3 149 64 36 49 DM2 (RPP1) DM1 (SSI4) a Bla-1/Hh-0 F 3 111
Table S2. F 2 Segregation Ratios at 16 C, Related to Figure 2 Cross n c Phenotype Model e 2 Locus A Locus B Normal F 1 -like Enhanced d Uk-1/Uk-3 149 64 36 49 DM2 (RPP1) DM1 (SSI4) a Bla-1/Hh-0 F 3 111
Supporting Information
 Supporting Information Transfection of DNA Cages into Mammalian Cells Email: a.turberfield@physics.ox.ac.uk Table of Contents Supporting Figure 1 DNA tetrahedra used in transfection experiments 2 Supporting
Supporting Information Transfection of DNA Cages into Mammalian Cells Email: a.turberfield@physics.ox.ac.uk Table of Contents Supporting Figure 1 DNA tetrahedra used in transfection experiments 2 Supporting
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/10/494/eaan6284/dc1 Supplementary Materials for Activation of master virulence regulator PhoP in acidic ph requires the Salmonella-specific protein UgtL Jeongjoon
www.sciencesignaling.org/cgi/content/full/10/494/eaan6284/dc1 Supplementary Materials for Activation of master virulence regulator PhoP in acidic ph requires the Salmonella-specific protein UgtL Jeongjoon
Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were
 1 Supplemental methods 2 3 4 5 6 7 8 9 1 11 12 13 14 15 16 17 18 19 21 22 23 Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were monitored by quantitative reverse-transcription
1 Supplemental methods 2 3 4 5 6 7 8 9 1 11 12 13 14 15 16 17 18 19 21 22 23 Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were monitored by quantitative reverse-transcription
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3240 Supplementary Figure 1 GBM cell lines display similar levels of p100 to p52 processing but respond differentially to TWEAK-induced TERT expression according to TERT promoter mutation
DOI: 10.1038/ncb3240 Supplementary Figure 1 GBM cell lines display similar levels of p100 to p52 processing but respond differentially to TWEAK-induced TERT expression according to TERT promoter mutation
Luo et al. Supplemental Figures and Materials and Methods
 Luo et al. Supplemental Figures and Materials and Methods The supplemental figures demonstrate that nuclear NFAT is situated at PODs, overexpressed PML does not increase NFAT nuclear localization, and
Luo et al. Supplemental Figures and Materials and Methods The supplemental figures demonstrate that nuclear NFAT is situated at PODs, overexpressed PML does not increase NFAT nuclear localization, and
FACS. Chromatin immunoprecipitation (ChIP) and ChIP-chip arrays 3 4
 FACS Stem progenitor sorting for progenitor analysis Adult bone marrow cells were depleted with Miltenyi lineage depletion column (Miltenyi). For identification of CMPs, GMPs, and MEPs the cells were incubated
FACS Stem progenitor sorting for progenitor analysis Adult bone marrow cells were depleted with Miltenyi lineage depletion column (Miltenyi). For identification of CMPs, GMPs, and MEPs the cells were incubated
Lecture 11: Gene Prediction
 Lecture 11: Gene Prediction Study Chapter 6.11-6.14 1 Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and end positions of genes in a genome Where are
Lecture 11: Gene Prediction Study Chapter 6.11-6.14 1 Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and end positions of genes in a genome Where are
Supplemental Figure 1
 mrna level (fold induction) Supplemental Figure 1 a-flag pmx-rank WT pmx-rank Mt 0 1 3 6 0 1 3 6 (h) c-fos b-actin 5 4 3 2 1 0 0 1 3 6 0 1 3 6 (h) pmx-rank WT pmx-rank Mt TRAP+ Mononuclear pre-ocs/well
mrna level (fold induction) Supplemental Figure 1 a-flag pmx-rank WT pmx-rank Mt 0 1 3 6 0 1 3 6 (h) c-fos b-actin 5 4 3 2 1 0 0 1 3 6 0 1 3 6 (h) pmx-rank WT pmx-rank Mt TRAP+ Mononuclear pre-ocs/well
Supplementary Methods Quantitative RT-PCR. For mrna, total RNA was prepared using TRIzol reagent (Invitrogen) and genomic DNA was eliminated with TURB
 Supplementary Methods Quantitative RT-PCR. For mrna, total RNA was prepared using TRIzol reagent (Invitrogen) and genomic DNA was eliminated with TURBO DNA-free Kit (Ambion). One µg of total RNA was reverse
Supplementary Methods Quantitative RT-PCR. For mrna, total RNA was prepared using TRIzol reagent (Invitrogen) and genomic DNA was eliminated with TURBO DNA-free Kit (Ambion). One µg of total RNA was reverse
ORFs and genes. Please sit in row K or forward
 ORFs and genes Please sit in row K or forward https://www.flickr.com/photos/teseum/3231682806/in/photostream/ Question: why do some strains of Vibrio cause cholera and others don t? Methods Mechanisms
ORFs and genes Please sit in row K or forward https://www.flickr.com/photos/teseum/3231682806/in/photostream/ Question: why do some strains of Vibrio cause cholera and others don t? Methods Mechanisms
Supplementary Fig. 1: Characterization of Asxl2 -/- mouse model. (a) HSCs and their
 Supplementary Fig. 1: Characterization of Asxl2 -/- mouse model. (a) HSCs and their differentiated cell populations were sorted from the BM of WT mice using respective surface markers, and Asxl2 mrna expression
Supplementary Fig. 1: Characterization of Asxl2 -/- mouse model. (a) HSCs and their differentiated cell populations were sorted from the BM of WT mice using respective surface markers, and Asxl2 mrna expression
Multiplexing Genome-scale Engineering
 Multiplexing Genome-scale Engineering Harris Wang, Ph.D. Department of Systems Biology Department of Pathology & Cell Biology http://wanglab.c2b2.columbia.edu Rise of Genomics An Expanding Toolbox Esvelt
Multiplexing Genome-scale Engineering Harris Wang, Ph.D. Department of Systems Biology Department of Pathology & Cell Biology http://wanglab.c2b2.columbia.edu Rise of Genomics An Expanding Toolbox Esvelt
RPA-AB RPA-C Supplemental Figure S1: SDS-PAGE stained with Coomassie Blue after protein purification.
 RPA-AB RPA-C (a) (b) (c) (d) (e) (f) Supplemental Figure S: SDS-PAGE stained with Coomassie Blue after protein purification. (a) RPA; (b) RPA-AB; (c) RPA-CDE; (d) RPA-CDE core; (e) RPA-DE; and (f) RPA-C
RPA-AB RPA-C (a) (b) (c) (d) (e) (f) Supplemental Figure S: SDS-PAGE stained with Coomassie Blue after protein purification. (a) RPA; (b) RPA-AB; (c) RPA-CDE; (d) RPA-CDE core; (e) RPA-DE; and (f) RPA-C
DNA aptamer to c-met inhibits cancer cell migration
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2014 Electronic Supplementary Information For DNA aptamer to c-met inhibits cancer cell migration Ryosuke
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2014 Electronic Supplementary Information For DNA aptamer to c-met inhibits cancer cell migration Ryosuke
S4B fluorescence (AU)
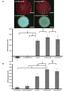 A S4B fluorescence (AU) S4B fluorescence (AU) dsbb csgba csgd dsbb csgba bcsa 5000 * NS NS 4000 * 3000 2000 1000 0 ΔcsgBAΔbcsA ΔcsgDΔdsbBΔbcsA ΔcsgBA ΔdsbBΔcsgBA ΔcsgDΔdsbB B -1000 4000 * * NS 3500 * 3000
A S4B fluorescence (AU) S4B fluorescence (AU) dsbb csgba csgd dsbb csgba bcsa 5000 * NS NS 4000 * 3000 2000 1000 0 ΔcsgBAΔbcsA ΔcsgDΔdsbBΔbcsA ΔcsgBA ΔdsbBΔcsgBA ΔcsgDΔdsbB B -1000 4000 * * NS 3500 * 3000
Supplementary Figure 1. The level of pri-mir-8 gradually decreases while those of BR-C and E74 increase during 3rd instar larval development.
 qrt-pcr RT-PCR Relative pri-mir-8 level 1.2 1.0 0.8 0.6 0.4 0.2 0.0 Early 3rd (72h) Mid 3rd (96h) Late 3rd (119h) BR-C E74 mtl rrna Early 3rd (72h) Mid 3rd (96h) Late 3rd (119h) Supplementary Figure 1.
qrt-pcr RT-PCR Relative pri-mir-8 level 1.2 1.0 0.8 0.6 0.4 0.2 0.0 Early 3rd (72h) Mid 3rd (96h) Late 3rd (119h) BR-C E74 mtl rrna Early 3rd (72h) Mid 3rd (96h) Late 3rd (119h) Supplementary Figure 1.
The GOODELL laboratory
 The GOODELL laboratory Author Nathan Boles Feb.5, 2009 Title Hematopoietic Progenitor Staining Introduction This protocol describes the analysis or isolation of the early hematopoietic progenitors. HBSS+
The GOODELL laboratory Author Nathan Boles Feb.5, 2009 Title Hematopoietic Progenitor Staining Introduction This protocol describes the analysis or isolation of the early hematopoietic progenitors. HBSS+
MacBlunt PCR Cloning Kit Manual
 MacBlunt PCR Cloning Kit Manual Shipping and Storage MacBlunt PCR Cloning Kits are shipped on dry ice. Each kit contains a box with cloning reagents and an attached bag with Eco-Blue Competent Cells (optional).
MacBlunt PCR Cloning Kit Manual Shipping and Storage MacBlunt PCR Cloning Kits are shipped on dry ice. Each kit contains a box with cloning reagents and an attached bag with Eco-Blue Competent Cells (optional).
Table S1. Bacterial strains (Related to Results and Experimental Procedures)
 Table S1. Bacterial strains (Related to Results and Experimental Procedures) Strain number Relevant genotype Source or reference 1045 AB1157 Graham Walker (Donnelly and Walker, 1989) 2458 3084 (MG1655)
Table S1. Bacterial strains (Related to Results and Experimental Procedures) Strain number Relevant genotype Source or reference 1045 AB1157 Graham Walker (Donnelly and Walker, 1989) 2458 3084 (MG1655)
Supplementary Figure 1 Tmod3 expression and phosphorylation. (a) Expression of Tmod3 in 3T3-L1 preadipocytes and differentiated adipocytes.
 1 2 3 1 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 Supplementary Figure 1 Tmod3 expression and phosphorylation. (a) Expression of Tmod3 in 3T3-L1 preadipocytes and differentiated
1 2 3 1 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 Supplementary Figure 1 Tmod3 expression and phosphorylation. (a) Expression of Tmod3 in 3T3-L1 preadipocytes and differentiated
Fig. S1: Nkx2.2 is specifically deleted in the intestinal epithelium.
 Fig. S1: Nkx2.2 is specifically deleted in the intestinal epithelium. PCR of representative tissues with oligonucleotides to identify recombination of the Nkx2.2flox allele. Nkx2.2 is specifically deleted
Fig. S1: Nkx2.2 is specifically deleted in the intestinal epithelium. PCR of representative tissues with oligonucleotides to identify recombination of the Nkx2.2flox allele. Nkx2.2 is specifically deleted
Supplementary Information. Construction of Lasso Peptide Fusion Proteins
 Supplementary Information Construction of Lasso Peptide Fusion Proteins Chuhan Zong 1, Mikhail O. Maksimov 2, A. James Link 2,3 * Departments of 1 Chemistry, 2 Chemical and Biological Engineering, and
Supplementary Information Construction of Lasso Peptide Fusion Proteins Chuhan Zong 1, Mikhail O. Maksimov 2, A. James Link 2,3 * Departments of 1 Chemistry, 2 Chemical and Biological Engineering, and
show PB hemoglobin concentration (Hgb) and platelet counts (Plt) in Kras ex3op/ex3op mice
 Supplemental Data Supplemental Figure Legends Supplemental Figure 1. Hematologic profile of Kras / mice. (A) Scatter plots show PB hemoglobin concentration (Hgb) and platelet counts (Plt) in Kras / mice
Supplemental Data Supplemental Figure Legends Supplemental Figure 1. Hematologic profile of Kras / mice. (A) Scatter plots show PB hemoglobin concentration (Hgb) and platelet counts (Plt) in Kras / mice
Supplementary Fig. 1. Isolation and in vitro expansion of EpCAM + cholangiocytes. For collagenase perfusion, enzyme solution was injected from the
 Supplementary Fig. 1. Isolation and in vitro expansion of EpCAM + cholangiocytes. For collagenase perfusion, enzyme solution was injected from the portal vein for digesting adult livers, whereas it was
Supplementary Fig. 1. Isolation and in vitro expansion of EpCAM + cholangiocytes. For collagenase perfusion, enzyme solution was injected from the portal vein for digesting adult livers, whereas it was
Supporting Information. Copyright Wiley-VCH Verlag GmbH & Co. KGaA, Weinheim, 2006
 Supporting Information Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Supporting Information for Expanding the Genetic
Supporting Information Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Supporting Information for Expanding the Genetic
We designed the targeting vector in such a way that the first exon was flanked by two LoxP sites and
 Mice We designed the targeting vector in such a way that the first exon was flanked by two LoxP sites and an Frt flanked neo cassette (Fig. S1A). Targeted BA1 (C57BL/6 129/SvEv) hybrid embryonic stem cells
Mice We designed the targeting vector in such a way that the first exon was flanked by two LoxP sites and an Frt flanked neo cassette (Fig. S1A). Targeted BA1 (C57BL/6 129/SvEv) hybrid embryonic stem cells
Supplemental methods Supplemental figure and legend...7. Supplemental table.. 8
 Supplemental Digital Content (SDC) Contents Supplemental methods..2-6 Supplemental figure and legend...7 Supplemental table.. 8 1 SDC, Supplemental Methods Flow cytometric analysis of intracellular phosphorylated
Supplemental Digital Content (SDC) Contents Supplemental methods..2-6 Supplemental figure and legend...7 Supplemental table.. 8 1 SDC, Supplemental Methods Flow cytometric analysis of intracellular phosphorylated
An evolutionarily conserved negative feedback mechanism in the hippo pathway reflects functional difference between LATS1 and LATS2
 /, Supplementary Advance Publications Materials 2015 2016 An evolutionarily conserved negative feedback mechanism in the hippo pathway reflects functional difference between LATS1 and LATS2 Supplementary
/, Supplementary Advance Publications Materials 2015 2016 An evolutionarily conserved negative feedback mechanism in the hippo pathway reflects functional difference between LATS1 and LATS2 Supplementary
88.7 <GFP-A>: EGFP GFP. Null. % of Cells! <APC-A> B220!
 B. C. 1 Cell Number (X18) A. % of Cells % of Max 8 Con 6 4 88.7 1 13 14 : EGFP 4 6 4 1 WT 8 % of% ofcells Max 6 6 4 G. 1 13 B Hdac3-/- 14 15 1 13 14 15 Ter119 Ly6C Wild type WT 4
B. C. 1 Cell Number (X18) A. % of Cells % of Max 8 Con 6 4 88.7 1 13 14 : EGFP 4 6 4 1 WT 8 % of% ofcells Max 6 6 4 G. 1 13 B Hdac3-/- 14 15 1 13 14 15 Ter119 Ly6C Wild type WT 4
An engineered tryptophan zipper-type peptide as a molecular recognition scaffold
 SUPPLEMENTARY MATERIAL An engineered tryptophan zipper-type peptide as a molecular recognition scaffold Zihao Cheng and Robert E. Campbell* Supplementary Methods Library construction for FRET-based screening
SUPPLEMENTARY MATERIAL An engineered tryptophan zipper-type peptide as a molecular recognition scaffold Zihao Cheng and Robert E. Campbell* Supplementary Methods Library construction for FRET-based screening
SUPPLEMENTAL DATA SUPPLEMENTAL FIGURE LEGENDS
 SUPPLEMENTAL DATA SUPPLEMENTAL FIGURE LEGENDS SUPPLEMENTAL FIGURE S1. Identification of BmCREC. (A) Amino acid sequences of BmCREC show the peptides identified in LC-MS/MS analysis (marked by red letters
SUPPLEMENTAL DATA SUPPLEMENTAL FIGURE LEGENDS SUPPLEMENTAL FIGURE S1. Identification of BmCREC. (A) Amino acid sequences of BmCREC show the peptides identified in LC-MS/MS analysis (marked by red letters
7-amino actinomycin D (7ADD) was added to all samples 10 minutes prior to analysis on the flow cytometer in order to gate 7AAD viable cells.
 Antibody staining for Ho uptake analyses For HSC staining, 10 7 BM cells from Ho perfused mice were stained with biotinylated lineage antibodies (CD3, CD5, B220, CD11b, Gr-1, CD41, Ter119), anti Sca-1-PECY7,
Antibody staining for Ho uptake analyses For HSC staining, 10 7 BM cells from Ho perfused mice were stained with biotinylated lineage antibodies (CD3, CD5, B220, CD11b, Gr-1, CD41, Ter119), anti Sca-1-PECY7,
Gene synthesis by circular assembly amplification
 Gene synthesis by circular assembly amplification Duhee Bang & George M Church Supplementary figures and text: Supplementary Figure 1. Dpo4 gene (1.05kb) construction by various methods. Supplementary
Gene synthesis by circular assembly amplification Duhee Bang & George M Church Supplementary figures and text: Supplementary Figure 1. Dpo4 gene (1.05kb) construction by various methods. Supplementary
Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system
 Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system Dr. Tim Welsink Molecular Biology Transient Gene Expression OUTLINE Short
Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system Dr. Tim Welsink Molecular Biology Transient Gene Expression OUTLINE Short
strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular
![strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular](/thumbs/72/66361212.jpg) Additional file 2 Identification of AOX1 in P. pastoris GS115 with a Mut s phenotype Results and Discussion The HBsAg producing strain was originally identified as a Mut s (methanol utilization slow) strain
Additional file 2 Identification of AOX1 in P. pastoris GS115 with a Mut s phenotype Results and Discussion The HBsAg producing strain was originally identified as a Mut s (methanol utilization slow) strain
evaluated with UAS CLB eliciting UAS CIT -N Libraries increase in the
 Supplementary Figures Supplementary Figure 1: Promoter scaffold library assemblies. Many ensembless of libraries were evaluated in this work. As a legend, the box outline color in top half of the figure
Supplementary Figures Supplementary Figure 1: Promoter scaffold library assemblies. Many ensembless of libraries were evaluated in this work. As a legend, the box outline color in top half of the figure
Supplementary Figure 1: Analysis of monocyte subsets and lineage relationships. (a) Gating strategy for definition of MDP and cmop populations in BM
 Supplementary Figure 1: Analysis of monocyte subsets and lineage relationships. (a) Gating strategy for definition of MDP and cmop populations in BM of Cx3cr1 GFP/+ mice related to Fig. 1a. MDP was defined
Supplementary Figure 1: Analysis of monocyte subsets and lineage relationships. (a) Gating strategy for definition of MDP and cmop populations in BM of Cx3cr1 GFP/+ mice related to Fig. 1a. MDP was defined
Homework. A bit about the nature of the atoms of interest. Project. The role of electronega<vity
 Homework Why cited articles are especially useful. citeulike science citation index When cutting and pasting less is more. Project Your protein: I will mail these out this weekend If you haven t gotten
Homework Why cited articles are especially useful. citeulike science citation index When cutting and pasting less is more. Project Your protein: I will mail these out this weekend If you haven t gotten
Supporting Online Information
 Supporting Online Information Isolation of Human Genomic DNA Sequences with Expanded Nucleobase Selectivity Preeti Rathi, Sara Maurer, Grzegorz Kubik and Daniel Summerer* Department of Chemistry and Chemical
Supporting Online Information Isolation of Human Genomic DNA Sequences with Expanded Nucleobase Selectivity Preeti Rathi, Sara Maurer, Grzegorz Kubik and Daniel Summerer* Department of Chemistry and Chemical
SUPPORTING INFORMATION FILE
 Intrinsic and extrinsic connections of Tet3 dioxygenase with CXXC zinc finger modules Nan Liu, Mengxi Wang, Wen Deng, Christine S. Schmidt, Weihua Qin, Heinrich Leonhardt and Fabio Spada Department of
Intrinsic and extrinsic connections of Tet3 dioxygenase with CXXC zinc finger modules Nan Liu, Mengxi Wang, Wen Deng, Christine S. Schmidt, Weihua Qin, Heinrich Leonhardt and Fabio Spada Department of
gacgacgaggagaccaccgctttg aggcacattgaaggtctcaaacatg
 Supplementary information Supplementary table 1: primers for cloning and sequencing cloning for E- Ras ggg aat tcc ctt gag ctg ctg ggg aat ggc ttt gcc ggt cta gag tat aaa gga agc ttt gaa tcc Tpbp Oct3/4
Supplementary information Supplementary table 1: primers for cloning and sequencing cloning for E- Ras ggg aat tcc ctt gag ctg ctg ggg aat ggc ttt gcc ggt cta gag tat aaa gga agc ttt gaa tcc Tpbp Oct3/4
NAME:... MODEL ANSWER... STUDENT NUMBER:... Maximum marks: 50. Internal Examiner: Hugh Murrell, Computer Science, UKZN
 COMP710, Bioinformatics with Julia, Test One, Thursday the 20 th of April, 2017, 09h30-11h30 1 NAME:...... MODEL ANSWER... STUDENT NUMBER:...... Maximum marks: 50 Internal Examiner: Hugh Murrell, Computer
COMP710, Bioinformatics with Julia, Test One, Thursday the 20 th of April, 2017, 09h30-11h30 1 NAME:...... MODEL ANSWER... STUDENT NUMBER:...... Maximum marks: 50 Internal Examiner: Hugh Murrell, Computer
* Correspondence: Ralf Dressel: 1 Supplementary Figures and Tables. - Supplementary Tables 1 to 3. - Supplementary Figures 1 to 6
 s Efficient killing of murine pluripotent stem cells by natural killer (NK) cells requires activation by cytokines and partly depends on the activating NK receptor NKG2D Carina Gröschel, Daniela Hübscher,
s Efficient killing of murine pluripotent stem cells by natural killer (NK) cells requires activation by cytokines and partly depends on the activating NK receptor NKG2D Carina Gröschel, Daniela Hübscher,
Supplemental Data. Bennett et al. (2010). Plant Cell /tpc
 BRN1 ---------MSSSNGGVPPGFRFHPTDEELLHYYLKKKISYEKFEMEVIKEVDLNKIEPWDLQDRCKIGSTPQNEWYFFSHKDRKYPTGS 81 BRN2 --------MGSSSNGGVPPGFRFHPTDEELLHYYLKKKISYQKFEMEVIREVDLNKLEPWDLQERCKIGSTPQNEWYFFSHKDRKYPTGS 82 SMB
BRN1 ---------MSSSNGGVPPGFRFHPTDEELLHYYLKKKISYEKFEMEVIKEVDLNKIEPWDLQDRCKIGSTPQNEWYFFSHKDRKYPTGS 81 BRN2 --------MGSSSNGGVPPGFRFHPTDEELLHYYLKKKISYQKFEMEVIREVDLNKLEPWDLQERCKIGSTPQNEWYFFSHKDRKYPTGS 82 SMB
In vivo BrdU Incorporation Assay for Murine Hematopioetic Stem Cells Ningfei An, Yubin Kang *
 In vivo BrdU Incorporation Assay for Murine Hematopioetic Stem Cells Ningfei An, Yubin Kang * Division of Hematology-Oncology, Department of Medicine, Medical University of South Carolina, Charleston,
In vivo BrdU Incorporation Assay for Murine Hematopioetic Stem Cells Ningfei An, Yubin Kang * Division of Hematology-Oncology, Department of Medicine, Medical University of South Carolina, Charleston,
Antibody used for FC Figure S1. Multimodal characterization of NIR dyes in vitro Figure S2. Ex vivo analysis of HL60 cells homing
 Antibody used for FC The following antibodies were used following manufacturer s instructions: anti-human CD4 (clone HI3, IgG1, k - Becton Dickinson), anti-human CD33 (clone WM3, IgG1, k- Becton Dickinson),
Antibody used for FC The following antibodies were used following manufacturer s instructions: anti-human CD4 (clone HI3, IgG1, k - Becton Dickinson), anti-human CD33 (clone WM3, IgG1, k- Becton Dickinson),
G+C content. 1 Introduction. 2 Chromosomes Topology & Counts. 3 Genome size. 4 Replichores and gene orientation. 5 Chirochores.
 1 Introduction 2 Chromosomes Topology & Counts 3 Genome size 4 Replichores and gene orientation 5 Chirochores 6 7 Codon usage 121 marc.bailly-bechet@univ-lyon1.fr Bacterial genome structures Introduction
1 Introduction 2 Chromosomes Topology & Counts 3 Genome size 4 Replichores and gene orientation 5 Chirochores 6 7 Codon usage 121 marc.bailly-bechet@univ-lyon1.fr Bacterial genome structures Introduction
A Genetically Encoded Toolbox for Glycocalyx Engineering: Tunable Control of Cell Adhesion,
 TITLE A Genetically Encoded Toolbox for Glycocalyx Engineering: Tunable Control of Cell Adhesion, Survival, and Cancer Cell Behaviors AUTHORS Carolyn R. Shurer *, Marshall J. Colville *, Vivek K. Gupta,
TITLE A Genetically Encoded Toolbox for Glycocalyx Engineering: Tunable Control of Cell Adhesion, Survival, and Cancer Cell Behaviors AUTHORS Carolyn R. Shurer *, Marshall J. Colville *, Vivek K. Gupta,
Supplementary Information
 Supplementary Information Microbead-based biomimetic synthetic neighbors enhance survival and function of rat pancreatic β-cells Wei Li, a Samuel Lee, b Minglin Ma, a, f Soo Min Kim, b Patrick Guye, c
Supplementary Information Microbead-based biomimetic synthetic neighbors enhance survival and function of rat pancreatic β-cells Wei Li, a Samuel Lee, b Minglin Ma, a, f Soo Min Kim, b Patrick Guye, c
Dynamic enhancer-gene body contacts during transcription elongation
 Dynamic enhancer-gene body contacts during transcription elongation Kiwon Lee, Chris C.-S. Hsiung,, Peng Huang, Arjun Raj, *, and Gerd A. Blobel Division of Hematology, The Children s Hospital of Philadelphia,
Dynamic enhancer-gene body contacts during transcription elongation Kiwon Lee, Chris C.-S. Hsiung,, Peng Huang, Arjun Raj, *, and Gerd A. Blobel Division of Hematology, The Children s Hospital of Philadelphia,
Lewis x/cd15 expression in human myeloid cell differentiation. is regulated by sialidase activity
 Lewis x/cd15 expression in human myeloid cell differentiation is regulated by sialidase activity Samah Zeineb Gadhoum 1, 2 and Robert Sackstein* 1, 2, 3, 4 From the Departments of Dermatology 1 and Medicine
Lewis x/cd15 expression in human myeloid cell differentiation is regulated by sialidase activity Samah Zeineb Gadhoum 1, 2 and Robert Sackstein* 1, 2, 3, 4 From the Departments of Dermatology 1 and Medicine
Supplemental Information. Target-Mediated Protection of Endogenous. MicroRNAs in C. elegans. Inventory of Supplementary Information
 Developmental Cell, Volume 20 Supplemental Information Target-Mediated Protection of Endogenous MicroRNAs in C. elegans Saibal Chatterjee, Monika Fasler, Ingo Büssing, and Helge Großhans Inventory of Supplementary
Developmental Cell, Volume 20 Supplemental Information Target-Mediated Protection of Endogenous MicroRNAs in C. elegans Saibal Chatterjee, Monika Fasler, Ingo Büssing, and Helge Großhans Inventory of Supplementary
Supporting Information
 Supporting Information Sen et al. 10.1073/pnas.1114232109 SI Materials and Methods Cell Culture. Human embryonic lung fibroblasts (HELF) were maintained in Eagle s minimum essential medium (Mediatech)
Supporting Information Sen et al. 10.1073/pnas.1114232109 SI Materials and Methods Cell Culture. Human embryonic lung fibroblasts (HELF) were maintained in Eagle s minimum essential medium (Mediatech)
SUPPLEMENTAL METHODS. Chromatin immunoprecipitation (ChIP) assay. Cell culture. Non-ablated mouse transplantation. Hematology and flow cytometry
 SUPPLEMENTAL METHODS Cell culture EML C1 cells were cultured in IMDM 20% horse serum, 1% antibiotic-antimycotic (Gibco by Life Technologies) 15% SCF conditioned medium from BHK/ MKL cells or 50 100 ng/ml
SUPPLEMENTAL METHODS Cell culture EML C1 cells were cultured in IMDM 20% horse serum, 1% antibiotic-antimycotic (Gibco by Life Technologies) 15% SCF conditioned medium from BHK/ MKL cells or 50 100 ng/ml
SUPPLEMENTARY MATERIALS AND METHODS. E. coli strains, plasmids, and growth conditions. Escherichia coli strain P90C (1)
 SUPPLEMENTARY MATERIALS AND METHODS E. coli strains, plasmids, and growth conditions. Escherichia coli strain P90C (1) dinb::kan (lab stock) derivative was used as wild-type. MG1655 alka tag dinb (2) is
SUPPLEMENTARY MATERIALS AND METHODS E. coli strains, plasmids, and growth conditions. Escherichia coli strain P90C (1) dinb::kan (lab stock) derivative was used as wild-type. MG1655 alka tag dinb (2) is
Hematopoietic stem cell transplantation without irradiation
 nature methods Hematopoietic stem cell transplantation without irradiation Claudia Waskow, Vikas Madan, Susanne Bartels, Céline Costa, Rosel Blasig & Hans-Reimer Rodewald Supplementary figures and text:
nature methods Hematopoietic stem cell transplantation without irradiation Claudia Waskow, Vikas Madan, Susanne Bartels, Céline Costa, Rosel Blasig & Hans-Reimer Rodewald Supplementary figures and text:
Supplementary Figure 1. Brwd1 transcripts in Brwd1 mut B cells. Nature Immunology: doi: /ni.3249
 Supplementary Figure 1 Brwd1 transcripts in Brwd1 mut B cells. (a) Analysis of Brwd1 transcripts in isolated small pre-b (Lin B220 + CD43 IgM FSC lo ) and splenic B (B220 + ) cells from WT and Brwd1 mut
Supplementary Figure 1 Brwd1 transcripts in Brwd1 mut B cells. (a) Analysis of Brwd1 transcripts in isolated small pre-b (Lin B220 + CD43 IgM FSC lo ) and splenic B (B220 + ) cells from WT and Brwd1 mut
SUPPLEMENTARY INFORMATION. Material and methods
 SUPPLEMENTARY INFORMATION Material and methods Cell culture Human hepatocellular carcinoma (HepG) cells and human embryonic kidney (HEK)93 cells were grown in Dulbecco s modified Eagle s medium (Invitrogen
SUPPLEMENTARY INFORMATION Material and methods Cell culture Human hepatocellular carcinoma (HepG) cells and human embryonic kidney (HEK)93 cells were grown in Dulbecco s modified Eagle s medium (Invitrogen
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES Supplementary Figure 1 Histogram displaying the distribution of DNA barcode copy numbers for each virus library. 100,000 BB88 cells were infected by lentivirus that carried the corresponding
SUPPLEMENTARY FIGURES Supplementary Figure 1 Histogram displaying the distribution of DNA barcode copy numbers for each virus library. 100,000 BB88 cells were infected by lentivirus that carried the corresponding
