Identification of a Novel -Lactamase Produced by Xanthomonas campestris, a Phytopathogenic Bacterium
|
|
|
- Brent Cummings
- 6 years ago
- Views:
Transcription
1 ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, July 1999, p Vol. 43, No /99/$ Copyright 1999, American Society for Microbiology. All Rights Reserved. Identification of a Novel -Lactamase Produced by Xanthomonas campestris, a Phytopathogenic Bacterium SHU-FEN WENG, 1 * CHUN-YI CHEN, 1 YEONG-SHENG LEE, 1 JUEY-WEN LIN, 2 AND YI-HSIUNG TSENG 1 Institute of Molecular Biology 1 and Institute of Biochemistry, 2 National Chung Hsing University, Taichung 402, Taiwan, Republic of China Received 19 October 1998/Returned for modification 2 February 1999/Accepted 27 April 1999 The Xanthomonas campestris pv. campestris 11 chromosome encodes a periplasmic -lactamase of 30 kda. Gene replacement and complementation confirmed the presence of this enzyme. Its deduced amino acid sequence shows identity and conserved domains between it and Stenotrophomonas maltophilia L2 and other Ambler class A/Bush group 2 -lactamases. Southern hybridization detected a single homologous fragment in each of 12 other Xanthomonas strains, indicating that the presence of a -lactamase gene is common among xanthomonads. * Corresponding author. Mailing address: Institute of Molecular Biology, National Chung Hsing University, Taichung 402, Taiwan, Republic of China. Phone: Fax: E- mail: sfweng@dragon.nchu.edu.tw. Xanthomonas campestris pv. campestris is a gram-negative phytopathogenic bacterium causing black rot in crucifers (25). In a previous study it was found that among 86 strains of X. campestris pv. campestris isolated in Taiwan, 81 were resistant to ampicillin at a level of 50 g/ml (12a). Antibiotic resistance can be transferred by a bacterium through a variety of mechanisms, including transduction, conjugation, or transfection, and once a resistance gene is transferred, it may be incorporated into the genome or plasmids of the recipient bacterium (16). Since the genus Xanthomonas inhabits soil, infected plants, plant debris, and asymptomatic plants near acutely infected plants, the possibility of widespread ampicillin resistance in this genus raises environmental concerns. In this study, a periplasmic -lactamase was detected in X. campestris pv. campestris strain 11. The amino acid sequence deduced from the bla gene revealed a high degree of similarity between the -lactamase of strain 11 and the L2 serine -lactamase of Stenotrophomonas maltophilia (22). Bacterial strains, growth conditions, and DNA techniques. X. campestris pv. campestris strains 11, 11A, and 17 have been described (26). The other Xanthomonas strains were provided by S.-T. Hsu. Luria-Bertani (LB) medium and L agar (13) were used as general-purpose media with or without ampicillin (50 g/ml). X. campestris was cultivated at 28 C. The DNA techniques employed were those described by Sambrook et al. (18). DNA sequences were determined by the method of Sanger et al. (19). Detection of -lactamase in strain 11. To determine the cellular location of the -lactamase, strain 11 cells were fractionated by the method of Osborn and Munson (15), with minor modifications. Proteins from different fractions were analyzed by sodium dodecyl sulfate 12% polyacrylamide gel electrophoresis. Detection of -lactamase activity in situ was carried out with benzylpenicillin (100 g/ml) as the substrate, as described by Tai et al. (21). A single protein band in the periplasmic fraction was found to contain -lactamase activity and was not detected in cytoplasmic, membrane, and extracellular fractions (data not shown). This enzyme, with a molecular mass of 30 kda as determined by gel electrophoresis, has a size similar to those of other class A -lactamases detected in gram-negative organisms (3, 8, 20, 22), suggesting that the 30-kDa protein is indeed a -lactamase. Antibiotic sensitivity of strain 11. We carried out MIC determinations according to the microdilution method recommended by the National Committee for Clinical Laboratory Standards, using -lactams purchased from Sigma (St. Louis, Mo.). The MICs determined were as follows (in micrograms per milliliter): cephaloridine, 128; cephalothin, 256; penicillin, 64; ampicillin, 128; carbenicillin; 128; cefotaxime, 8; cefoxitin, 16; oxacillin, 256; and piperacillin, 128. Results indicated that strain 11 was highly resistant to the penicillins and narrowspectrum cephalosporins used in this study, was less resistant to cefoxitin (an expanded-spectrum cephalosporin), and was susceptible to cefotaxime (a broad-spectrum cephalosporin). Cloning and sequencing of the strain 11 bla gene. The strain 11 bla gene was cloned, by conferring ampicillin resistance in E. coli DH5, from a genomic library constructed with pbk- CMV. Deletion mapping located the bla gene in the 1.7-kb SstI-EcoRI fragment (Fig. 1). Sequence analysis of this 1.7-kb fragment revealed two open reading frames, orf295 and orf41, with opposite orientations (Fig. 2). orf295 (nucleotides [nt] 224 to 1111) potentially encoded a protein of 295 amino acids with an estimated molecular weight of 31,810. A Shine-Dalgarno sequence (AAGGA GG) was found 8 nt upstream of the orf295 initiation codon. Downstream from the orf295 termination codon (nt 1114 to 1167) were two inverted repeat sequences resembling a transcriptional termination signal (Fig. 2). orf41 (nt 1 to 123) is an incomplete open reading frame (Fig. 2). Its deduced amino acid sequence (41 residues) had 51% identity to the N-terminal regions of the AmpR proteins (required for the induction of -lactamases) from Citrobacter freundii (11), Enterobacter cloacae (14), Pseudomonas aeruginosa (12), and Proteus vulgaris (6). Based on the sequence identity, we predicted that orf41 was the 5 portion of the strain 11 ampr. The intergenic region between orf295 and orf41 (100 nt) did not contain the consensus sequences that are typical of the 35 and 10 regions of an Escherichia coli promoter (Fig. 2). The strain 11 bla gene had a G C content of 68.2%, and G Downloaded from on April 22, 2018 by guest 1792
2 VOL. 43, 1999 NOTES 1793 FIG. 1. Localization of the -lactamase gene of X. campestris pv. campestris strain 11. Plasmid pbk-1 contains a 4.5-kb Sau3A1 insert cloned into the multiple cloning sites of vector pbk-cmv. For deletion mapping, the 3.1-kb SstI-HindIII fragment, the 1.7-kb SstI-EcoRI fragment, the 0.8-kb SstI-XhoI fragment, and the 0.9-kb XhoI-EcoRI fragment from the pbk-1 insert were subcloned into the compatible sites of pbk-cmv to generate pbkh, pbke, pbkx, and pbkxe, respectively., transformants resistant to ampicillin;, transformants sensitive to ampicillin. Downloaded from and C were likely to appear in the third positions of the codons. The similarities between it and other X. campestris chromosomal genes (4) confirm that the strain 11 bla gene is indigenous. Computer analysis indicated that the strain 11 -lactamase has a signal peptide (17) of 22 residues (Fig. 2), consistent with the observation that the enzyme is a secretory protein destined for the periplasm and that, after processing, with the cleaved region being between Phe 22 and Ala 23, the mature protein would consist of 273 amino acids with a pi of 8.2 and a molecular weight (29,408) similar to that observed after sodium dodecyl sulfate-polyacrylamide gel electrophoresis (data not shown). Sequence comparison and classification of the strain 11 -lactamase. Several schemes have been proposed for the classification of bacterial -lactamases. Ambler (1) suggested a scheme containing four classes of -lactamases based on primary amino acid sequences. The most recent functional scheme by Bush et al. (5) considered the inhibition characteristics, substrate profiles, and molecular structures of the enzymes. Based on these parameters, they constructed a dendrogram showing relationships among 88 -lactamases. In this study, we constructed a dendrogram that clusters the strain 11 -lactamase with 20 other related sequences, including sequences which were not included in the dendrogram of Bush et al. (5). The highest degree of relatedness to the strain 11 -lactamase was found in the L2 serine -lactamase from S. maltophilia, an Ambler class A enzyme which falls in group 2e of the Bush scheme (Fig. 3). Consistent with this finding, the strain 11 -lactamase has the highest degree of identity (51.5%) with S. maltophilia L2 (22) and lower degrees with several other class A/group 2 enzymes: Yersinia enterocolitica BLA-A (45.0%), E. coli TOHO-1 (44.5%), and E. coli MEN-1 (44.5%) (2, 3, 8, 20) (Fig. 3). The conserved sequences present in the class A enzymes, including the consensus sequences STXK, SDN, and KTG and the highly conserved glutamic acid residue located 34 residues downstream from the SDN loop, are also found in the strain 11 -lactamase (Fig. 4). Northern blot analysis. The 317-bp XhoI-NotI fragment (bp 734 to 1050; Fig. 2) within the strain 11 bla gene was used as the probe for Northern hybridization with the RNA prepared from strain 11 cells grown in LB medium and treated with ampicillin 30 min prior to RNA extraction (24). A hybridization band of ca. 980 nt, with smearing in the lower area of the gel, was detected (data not shown). This transcript, having a size similar to that of the bla gene, appears to be monocistronic. Construction of the bla mutant strain 11bla::Tc and complementation test. Mutation in strain 11 bla was performed according to Lee et al. (10). A 2.0-kb tetracycline resistance (Tc r ) cartridge from mini-tn5tc (7) was inserted into the SmaI site of the pbke insert. The interrupted bla gene was electroporated into strain 11 (23), followed by replacement of the chromosomal wild-type version. One of the tetracycline-resistant (15 g/ml) and ampicillin-sensitive mutants (strain 11bla::Tc) thus obtained was verified by Southern hybridization to have undergone double crossover. The 1.7-kb SstI-EcoRI fragment containing the complete bla sequence was cloned into the broad-host-range vector prk415 on April 22, 2018 by guest
3 1794 NOTES ANTIMICROB. AGENTS CHEMOTHER. Downloaded from on April 22, 2018 by guest FIG. 2. Nucleotide sequence (1,190 bp) of the upstream region of the pbke insert. The deduced amino acids are represented by one-letter codes. Initiation and stop codons are boldfaced. The underlined regions (bp 1114 to 1139 and bp 1142 to 1167) are inverted repeats which potentially form a stem-loop structure resembling a transcription termination signal. Boldfaced letters in the repeats (16 out of 26 nucleotides in each region) represent the complementary bases. SD, ribosome-binding sequence.
4 VOL. 43, 1999 NOTES 1795 Downloaded from on April 22, 2018 by guest FIG. 3. Dendrogram of -lactamases based on the sequence similarity between the strain 11 -lactamase and the 20 other sequences most closely related. The signal sequences were not included for comparison. Asterisks indicate the sequences not included in the dendrogram of Bush et al. (5). The Genetics Computer Group package was used for the construction.
5 1796 NOTES ANTIMICROB. AGENTS CHEMOTHER. Downloaded from FIG. 4. Alignment of the deduced amino acid sequences of the class A -lactamases from gram-negative bacteria S. maltophilia L2 (22), Y. enterocolitica BLA-A (20), E. coli TOHO-1 (8), and E. coli MEN-1 (2, 3). The consensus line displays the residues conserved in the sequences, with the boldfaced letters indicating the residues conserved in the strain 11 -lactamase. The consensus sequences STXK, SDN, KTG and the glutamic acid residue located 34 residues behind the SDN loop are underlined. The Genetics Computer Group package was used in these analyses. (9), resulting in prkse. Strain 11bla::Tc carrying prkse regained ampicillin resistance, indicating that the 1.7-kb SstI- EcoRI fragment indeed encodes a -lactamase. Distribution of bla gene among xanthomonads. Thirteen other strains of X. campestris, representing eight pathovars and one Xanthomonas oryzae pv. oryzae strain, were tested for resistance by being grown on LB agar containing ampicillin. All strains except X. campestris pv. mangiferaeindicae strain 38 were found to be ampicillin resistant. Southern hybridization revealed that for each of the EcoRI fragments of chromosomal DNA from these X. campestris strains, except strain 38, for which no signal was observed, a single band was detected. A 3.8-kb fragment was detected in X. campestris pathovars campestris (strains 2, 6, 11A, 17, and 85), begoniae (strain 59), dieffenbachiae (strain 65), and phaseoli (strain 73); a 10-kb fragment was detected in pathovars citri (strain 60), glycines (strain 69), and vesicatoria (strain 64); and a 5.5-kb fragment was detected in X. oryzae pv. oryzae (strain 21). Since only one fragment was detected, it is likely that only one copy of the bla gene is present in each of these Ap r strains. These results indicate that the bla genes are highly homologous and widespread among xanthomonads. Conclusion. In this study, we have identified a novel -lactamase from X. campestris, a plant-pathogenic bacterial species presently not involved in human or animal pathology. In addition, the presence of a -lactamase gene has been demonstrated to be widespread among xanthomonads. It is thought that unusual -lactamases produced by human pathogens are acquired from some other sources; the occurrence of this enzyme in the botanical world suggests one of the possible novel origins. Nucleotide sequence accession number. The nucleotide sequence determined here has been deposited in GenBank under accession no. AF We thank S.-T. Hsu for donating Xanthomonas strains and M.-T. Yang for providing the genomic bank of strain 11. We also thank S. T. Liu for a critical reading of the manuscript. This study was supported by grant no. NSC B B15 from the National Science Council of the Republic of China. on April 22, 2018 by guest
6 VOL. 43, 1999 NOTES 1797 REFERENCES 1. Ambler, R. P The structure of -lactamase. Philos. Trans. R. Soc. Lond. B 289: Barthelemy, M., J. Peduzzi, H. Bernard, C. Tancrede, and R. Labia Close amino acid sequence relationship between the new plasmid-mediated extended spectrum -lactamase MEN-1 and chromosomally encoded enzymes of Klebsiella oxytoca. Biochim. Biophys. Acta 1122: Bauernfeind, A., I. Stemplinger, R. Jungwirth, S. Ernst, and J. M. Casellas Sequences of -lactamase genes encoding CTX-M-1 (MEN-1) and CTX-M-2 and relationship of their amino acid sequences with those of other -lactamases. Antimicrob. Agents Chemother. 40: Bradbury, J. F Genus II. Xanthomonas Dowson 1939, 187. AL, p In N. R. Krieg and J. G. Holt (ed.), Bergey s manual of systematic bacteriology, vol. 1. Williams & Wilkins, Baltimore, Md. 5. Bush, K., G. A. Jacoby, and A. E. Medeiros A functional classification scheme for -lactamases and its correlation with molecular structure. Antimicrob. Agents Chemother. 39: Datz, M., B. Joris, E. A. M. Azab, M. Galleni, J. Van Beeumen, J.-M. Frere, and H. H. Martin A common system controls the induction of very different genes. The class-a -lactamase of Proteus vulgaris and the enterobacterial class-c -lactamase. Eur. J. Biochem. 226: de Lorenzo, V., M. Herrero, U. Jakubzik, and K. N. Timmis Mini-Tn5 transposon derivatives for insertion mutagenesis, promoter probing, and chromosomal insertion of cloned DNA in gram-negative eubacteria. J. Bacteriol. 172: Ishii, Y., A. Ohno, H. Taguchi, S. Imajo, M. Ishiguro, and H. Matsuzawa Cloning and sequence of the gene encoding a cefotaxime-hydrolyzing class A -lactamase isolated from Escherichia coli. Antimicrob. Agents Chemother. 39: Keen, N. T., S. Tamaki, D. Kobayashi, and D. Trollinger Improved broad-host-range plasmids for DNA cloning in Gram-negative bacteria. Gene 70: Lee, T.-C., N.-T. Lin, and Y.-H. Tseng Isolation and characterization of the reca gene of Xanthomonas campestris pv. campestris. Biochem. Biophys. Res. Commun. 221: Lindquist, S., F. Lindberg, and S. Normark Binding of the Citrobacter freundii AmpR regulator to a single DNA site provides both autoregulation and activation of the inducible ampc -lactamase gene. J. Bacteriol. 171: Lodge, J. M., S. D. Minchin, L. J. V. Piddock, and S. J. W. Busby Cloning, sequencing and analysis of the structural gene and regulatory region of the Pseudomonas aeruginosa chromosomal ampc -lactamase. Biochem. J. 272: a.Mao, J.-R., and Y.-H. Tseng. Unpublished data. 13. Miller, J. H Experiments in molecular genetics. Cold Spring Harbor Laboratory, Cold Spring Harbor, N.Y. 14. Naas, T., and P. Nordmann Analysis of a carbapenem-hydrolyzing class A -lactamase from Enterobacter cloacae and of its LysR-type regulatory protein. Proc. Natl. Acad. Sci. USA 91: Osborn, M. J., and R. Munson Separation of the inner (cytoplasmic) and outer membranes of Gram-negative bacteria. Methods Enzymol. 31: Pitout, J. D. D., C. C. Sanders, and W. E. Sanders, Jr Antimicrobial resistance with focus on -lactam resistance in gram-negative bacilli. Am. J. Med. 103: Pugsley, A. P The complete general secretory pathway in gram-negative bacteria. Microbiol. Rev. 57: Sambrook, J., E. F. Fritsch, and T. Maniatis Molecular cloning: a laboratory manual, 2nd ed. Cold Spring Harbor Laboratory, Cold Spring Harbor, N.Y. 19. Sanger, F., S. Nicklen, and A. R. Coulson DNA sequencing with chain-terminating inhibitors. Proc. Natl. Acad. Sci. USA 74: Seoane, A., and J. M. Garcia Lobo Nucleotide sequence of a new class A -lactamase gene from the chromosome of Yersinia enterocolitica: implications for the evolution of class A -lactamases. Mol. Gen. Genet. 228: Tai, P. C., N. Zyk, and N. Citri In situ detection of -lactamase activity in sodium dodecyl sulfate-polyacrylamide gels. Anal. Biochem. 144: Walsh, T. R., A. P. MacGowan, and P. M. Bennett Sequence analysis and enzyme kinetics of the L2 serine -lactamase from Stenotrophomonas maltophilia. Antimicrob. Agents Chemother. 41: Wang, T.-W., and Y.-H. Tseng Electrotransformation of Xanthomonas campestris by RF DNA of filamentous phage Lf. Lett. Appl. Microbiol. 14: Weng, S.-F., Y.-L. Liu, J.-W. Lin, and Y.-H. Tseng Transcriptional analysis of the threonine dehydrogenase gene of Xanthomonas campestris. Biochem. Biophys. Res. Commun. 240: Williams, P. H Black rot: a continuing threat to world crucifers. Plant Dis. 64: Yang, B.-Y., and Y.-H. Tseng Production of exopolysaccharide and levels of protease and pectinase activity in pathogenic and non-pathogenic strains of Xanthomonas campestris pv. campestris. Bot. Bull. Acad. Sin. 29: Downloaded from on April 22, 2018 by guest
Comparative Characterization of the Cephamycinase bla CMY-1 Gene and Its Relationship with Other -Lactamase Genes
 ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Aug. 1996, p. 1926 1930 Vol. 40, No. 8 0066-4804/96/$04.00 0 Copyright 1996, American Society for Microbiology Comparative Characterization of the Cephamycinase bla
ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Aug. 1996, p. 1926 1930 Vol. 40, No. 8 0066-4804/96/$04.00 0 Copyright 1996, American Society for Microbiology Comparative Characterization of the Cephamycinase bla
Cloning and Characterization of E. meningoseptica Beta Lactamase
 Cloning and Characterization of E. meningoseptica Beta Lactamase Authors: Lindsey Purcell, Jessica Matts, Patricia Canaan* Department of Biochemistry and Molecular Biology Abstract Elizabethkingia meningoseptica
Cloning and Characterization of E. meningoseptica Beta Lactamase Authors: Lindsey Purcell, Jessica Matts, Patricia Canaan* Department of Biochemistry and Molecular Biology Abstract Elizabethkingia meningoseptica
The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity
 Promega Notes Magazine Number 62, 1997, p. 02 The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity By Christine Andrews and Scott Lesley Promega
Promega Notes Magazine Number 62, 1997, p. 02 The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity By Christine Andrews and Scott Lesley Promega
Molecular Cell Biology - Problem Drill 11: Recombinant DNA
 Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
Prevalence and molecular characterization of clinical isolates of Escherichia coli expressing an AmpC phenotype
 J Antimicrob Chemother 2010; 65: 460 464 doi:10.1093/jac/dkp484 Advance publication 22 January 2010 Prevalence and molecular characterization of clinical isolates of Escherichia coli expressing an AmpC
J Antimicrob Chemother 2010; 65: 460 464 doi:10.1093/jac/dkp484 Advance publication 22 January 2010 Prevalence and molecular characterization of clinical isolates of Escherichia coli expressing an AmpC
The biomérieux solution. VITEK2 : A challenge with ESBL ESBL. Karen Bush
 International Newsletter n 4 December 2003 Through the IDENTIFYING RESISTANCE Newsletter, biomérieux s ambition is to contribute to the awareness and progress in the field of resistance to antibiotics.
International Newsletter n 4 December 2003 Through the IDENTIFYING RESISTANCE Newsletter, biomérieux s ambition is to contribute to the awareness and progress in the field of resistance to antibiotics.
Lecture Four. Molecular Approaches I: Nucleic Acids
 Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
2054, Chap. 14, page 1
 2054, Chap. 14, page 1 I. Recombinant DNA technology (Chapter 14) A. recombinant DNA technology = collection of methods used to perform genetic engineering 1. genetic engineering = deliberate modification
2054, Chap. 14, page 1 I. Recombinant DNA technology (Chapter 14) A. recombinant DNA technology = collection of methods used to perform genetic engineering 1. genetic engineering = deliberate modification
Recombinant DNA Technology. The Role of Recombinant DNA Technology in Biotechnology. yeast. Biotechnology. Recombinant DNA technology.
 PowerPoint Lecture Presentations prepared by Mindy Miller-Kittrell, North Carolina State University C H A P T E R 8 Recombinant DNA Technology The Role of Recombinant DNA Technology in Biotechnology Biotechnology?
PowerPoint Lecture Presentations prepared by Mindy Miller-Kittrell, North Carolina State University C H A P T E R 8 Recombinant DNA Technology The Role of Recombinant DNA Technology in Biotechnology Biotechnology?
Genetics Lecture 21 Recombinant DNA
 Genetics Lecture 21 Recombinant DNA Recombinant DNA In 1971, a paper published by Kathleen Danna and Daniel Nathans marked the beginning of the recombinant DNA era. The paper described the isolation of
Genetics Lecture 21 Recombinant DNA Recombinant DNA In 1971, a paper published by Kathleen Danna and Daniel Nathans marked the beginning of the recombinant DNA era. The paper described the isolation of
A Novel Class C -Lactamase (FOX-2) in Escherichia coli Conferring Resistance to Cephamycins
 ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Sept. 1997, p. 2041 2046 Vol. 41, No. 9 0066-4804/97/$04.00 0 Copyright 1997, American Society for Microbiology A Novel Class C -Lactamase (FOX-2) in Escherichia
ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Sept. 1997, p. 2041 2046 Vol. 41, No. 9 0066-4804/97/$04.00 0 Copyright 1997, American Society for Microbiology A Novel Class C -Lactamase (FOX-2) in Escherichia
AP Biology Gene Expression/Biotechnology REVIEW
 AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
The Prevalence of TEM-1 gene causing resistance to beta-lactam antibiotics in Klebsiella pneumoniae isolates from clinical samples and plasmid curing
 Available online at www.ijmrhs.com ISSN No: 2319-5886 International Journal of Medical Research & Health Sciences, 2016, 5, 11:557-561 The Prevalence of TEM-1 gene causing resistance to beta-lactam antibiotics
Available online at www.ijmrhs.com ISSN No: 2319-5886 International Journal of Medical Research & Health Sciences, 2016, 5, 11:557-561 The Prevalence of TEM-1 gene causing resistance to beta-lactam antibiotics
BCH 462 Competent Cells Formation and Transformation of Competent Cells with plasmid DNA.
 Lab#2 BCH 462 Competent Cells Formation and Transformation of Competent Cells with plasmid DNA. Outlines: 1-Insertion of foreign gene to the plasmid. 2-Competent cell. 3-Transformation of bacterial cell.
Lab#2 BCH 462 Competent Cells Formation and Transformation of Competent Cells with plasmid DNA. Outlines: 1-Insertion of foreign gene to the plasmid. 2-Competent cell. 3-Transformation of bacterial cell.
Answers to Module 1. An obligate aerobe is an organism that has an absolute requirement of oxygen for growth.
 Answers to Module 1 Short Answers 1) What is an obligate aerobe? An obligate aerobe is an organism that has an absolute requirement of oxygen for growth. What about facultative anaerobe? 2) Distinguish
Answers to Module 1 Short Answers 1) What is an obligate aerobe? An obligate aerobe is an organism that has an absolute requirement of oxygen for growth. What about facultative anaerobe? 2) Distinguish
Genetic Engineering & Recombinant DNA
 Genetic Engineering & Recombinant DNA Chapter 10 Copyright The McGraw-Hill Companies, Inc) Permission required for reproduction or display. Applications of Genetic Engineering Basic science vs. Applied
Genetic Engineering & Recombinant DNA Chapter 10 Copyright The McGraw-Hill Companies, Inc) Permission required for reproduction or display. Applications of Genetic Engineering Basic science vs. Applied
BIOLOGY LTF DIAGNOSTIC TEST DNA to PROTEIN & BIOTECHNOLOGY
 Biology Multiple Choice 016074 BIOLOGY LTF DIAGNOSTIC TEST DNA to PROTEIN & BIOTECHNOLOGY Test Code: 016074 Directions: Each of the questions or incomplete statements below is followed by five suggested
Biology Multiple Choice 016074 BIOLOGY LTF DIAGNOSTIC TEST DNA to PROTEIN & BIOTECHNOLOGY Test Code: 016074 Directions: Each of the questions or incomplete statements below is followed by five suggested
Computational Biology I LSM5191
 Computational Biology I LSM5191 Lecture 5 Notes: Genetic manipulation & Molecular Biology techniques Broad Overview of: Enzymatic tools in Molecular Biology Gel electrophoresis Restriction mapping DNA
Computational Biology I LSM5191 Lecture 5 Notes: Genetic manipulation & Molecular Biology techniques Broad Overview of: Enzymatic tools in Molecular Biology Gel electrophoresis Restriction mapping DNA
Lecture 25 (11/15/17)
 Lecture 25 (11/15/17) Reading: Ch9; 328-332 Ch25; 990-995, 1005-1012 Problems: Ch9 (study-guide: applying); 1,2 Ch9 (study-guide: facts); 7,8 Ch25 (text); 1-3,5-7,9,10,13-15 Ch25 (study-guide: applying);
Lecture 25 (11/15/17) Reading: Ch9; 328-332 Ch25; 990-995, 1005-1012 Problems: Ch9 (study-guide: applying); 1,2 Ch9 (study-guide: facts); 7,8 Ch25 (text); 1-3,5-7,9,10,13-15 Ch25 (study-guide: applying);
Efficient Multi-site-directed Mutagenesis directly from Genomic Template.
 Efficient Multi-site-directed Mutagenesis directly from Genomic Template. Fengtao Luo 1, Xiaolan Du 1, Tujun Weng 1, Xuan Wen 1, Junlan Huang 1, Lin Chen 1 Running title: Multi-site-directed Mutagenesis
Efficient Multi-site-directed Mutagenesis directly from Genomic Template. Fengtao Luo 1, Xiaolan Du 1, Tujun Weng 1, Xuan Wen 1, Junlan Huang 1, Lin Chen 1 Running title: Multi-site-directed Mutagenesis
Cloning and Sequencing of the Gene Encoding Curvaticin FS47, an Anti- Listerial Bacteriocin Produced by Lactobacillus curvatus FS47
 Cloning and Sequencing of the Gene Encoding Curvaticin FS47, an Anti- Listerial Bacteriocin Produced by Lactobacillus curvatus FS47 S. Macwana, L. Ma, M.A. Cousin, and P.M. Muriana Story in Brief Curvaticin
Cloning and Sequencing of the Gene Encoding Curvaticin FS47, an Anti- Listerial Bacteriocin Produced by Lactobacillus curvatus FS47 S. Macwana, L. Ma, M.A. Cousin, and P.M. Muriana Story in Brief Curvaticin
Supporting Information-Tables
 Supporting Information-Tables Table S1. Bacterial strains and plasmids used in this work Bacterial strains Description Source of reference Streptococcus pneumoniae 1 Cp1015 non-capsulated and βl susceptible
Supporting Information-Tables Table S1. Bacterial strains and plasmids used in this work Bacterial strains Description Source of reference Streptococcus pneumoniae 1 Cp1015 non-capsulated and βl susceptible
CHAPTER 20 DNA TECHNOLOGY AND GENOMICS. Section A: DNA Cloning
 Section A: DNA Cloning 1. DNA technology makes it possible to clone genes for basic research and commercial applications: an overview 2. Restriction enzymes are used to make recombinant DNA 3. Genes can
Section A: DNA Cloning 1. DNA technology makes it possible to clone genes for basic research and commercial applications: an overview 2. Restriction enzymes are used to make recombinant DNA 3. Genes can
M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION
 M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION Chapter 7 Microbial Genetics Lecture prepared by Mindy Miller-Kittrell, University of Tennessee, Knoxville The Structure and Replication
M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION Chapter 7 Microbial Genetics Lecture prepared by Mindy Miller-Kittrell, University of Tennessee, Knoxville The Structure and Replication
Year III Pharm.D Dr. V. Chitra
 Year III Pharm.D Dr. V. Chitra 1 Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Only one strand of DNA serves
Year III Pharm.D Dr. V. Chitra 1 Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Only one strand of DNA serves
DNA Technology. Asilomar Singer, Zinder, Brenner, Berg
 DNA Technology Asilomar 1973. Singer, Zinder, Brenner, Berg DNA Technology The following are some of the most important molecular methods we will be using in this course. They will be used, among other
DNA Technology Asilomar 1973. Singer, Zinder, Brenner, Berg DNA Technology The following are some of the most important molecular methods we will be using in this course. They will be used, among other
Antisense RNA Insert Design for Plasmid Construction to Knockdown Target Gene Expression
 Vol. 1:7-15 Antisense RNA Insert Design for Plasmid Construction to Knockdown Target Gene Expression Ji, Tom, Lu, Aneka, Wu, Kaylee Department of Microbiology and Immunology, University of British Columbia
Vol. 1:7-15 Antisense RNA Insert Design for Plasmid Construction to Knockdown Target Gene Expression Ji, Tom, Lu, Aneka, Wu, Kaylee Department of Microbiology and Immunology, University of British Columbia
2. In Figure 10-4, why is edna made only from mrna and not also from trnas and ribosomal RNAs?
 2. In Figure 10-4, why is edna made only from mrna and not also from trnas and ribosomal RNAs? Answer: edna is made from mrna and not from trnas or rrnas because polyt primers are used to prime the first
2. In Figure 10-4, why is edna made only from mrna and not also from trnas and ribosomal RNAs? Answer: edna is made from mrna and not from trnas or rrnas because polyt primers are used to prime the first
Genome Sequence Assembly
 Genome Sequence Assembly Learning Goals: Introduce the field of bioinformatics Familiarize the student with performing sequence alignments Understand the assembly process in genome sequencing Introduction:
Genome Sequence Assembly Learning Goals: Introduce the field of bioinformatics Familiarize the student with performing sequence alignments Understand the assembly process in genome sequencing Introduction:
7 Gene Isolation and Analysis of Multiple
 Genetic Techniques for Biological Research Corinne A. Michels Copyright q 2002 John Wiley & Sons, Ltd ISBNs: 0-471-89921-6 (Hardback); 0-470-84662-3 (Electronic) 7 Gene Isolation and Analysis of Multiple
Genetic Techniques for Biological Research Corinne A. Michels Copyright q 2002 John Wiley & Sons, Ltd ISBNs: 0-471-89921-6 (Hardback); 0-470-84662-3 (Electronic) 7 Gene Isolation and Analysis of Multiple
Synthetic Biology. Sustainable Energy. Therapeutics Industrial Enzymes. Agriculture. Accelerating Discoveries, Expanding Possibilities. Design.
 Synthetic Biology Accelerating Discoveries, Expanding Possibilities Sustainable Energy Therapeutics Industrial Enzymes Agriculture Design Build Generate Solutions to Advance Synthetic Biology Research
Synthetic Biology Accelerating Discoveries, Expanding Possibilities Sustainable Energy Therapeutics Industrial Enzymes Agriculture Design Build Generate Solutions to Advance Synthetic Biology Research
BS 50 Genetics and Genomics Week of Nov 29
 BS 50 Genetics and Genomics Week of Nov 29 Additional Practice Problems for Section Problem 1. A linear piece of DNA is digested with restriction enzymes EcoRI and HinDIII, and the products are separated
BS 50 Genetics and Genomics Week of Nov 29 Additional Practice Problems for Section Problem 1. A linear piece of DNA is digested with restriction enzymes EcoRI and HinDIII, and the products are separated
MIT Department of Biology 7.013: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr.
 MIT Department of Biology 7.01: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel iv) Would Xba I be useful for cloning? Why or why not?
MIT Department of Biology 7.01: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel iv) Would Xba I be useful for cloning? Why or why not?
3 Designing Primers for Site-Directed Mutagenesis
 3 Designing Primers for Site-Directed Mutagenesis 3.1 Learning Objectives During the next two labs you will learn the basics of site-directed mutagenesis: you will design primers for the mutants you designed
3 Designing Primers for Site-Directed Mutagenesis 3.1 Learning Objectives During the next two labs you will learn the basics of site-directed mutagenesis: you will design primers for the mutants you designed
Learning Objectives :
 Learning Objectives : Understand the basic differences between genomic and cdna libraries Understand how genomic libraries are constructed Understand the purpose for having overlapping DNA fragments in
Learning Objectives : Understand the basic differences between genomic and cdna libraries Understand how genomic libraries are constructed Understand the purpose for having overlapping DNA fragments in
Supplementary Material. Increased heterocyst frequency by patn disruption in Anabaena leads to enhanced photobiological
 Supplementary Material Increased heterocyst frequency by patn disruption in Anabaena leads to enhanced photobiological hydrogen production at high light intensity and high cell density Applied Microbiology
Supplementary Material Increased heterocyst frequency by patn disruption in Anabaena leads to enhanced photobiological hydrogen production at high light intensity and high cell density Applied Microbiology
Introducing new DNA into the genome requires cloning the donor sequence, delivery of the cloned DNA into the cell, and integration into the genome.
 Key Terms Chapter 32: Genetic Engineering Cloning describes propagation of a DNA sequence by incorporating it into a hybrid construct that can be replicated in a host cell. A cloning vector is a plasmid
Key Terms Chapter 32: Genetic Engineering Cloning describes propagation of a DNA sequence by incorporating it into a hybrid construct that can be replicated in a host cell. A cloning vector is a plasmid
MOLECULAR GENETICS: TRANSFORMATION AND CLONING adapted by Dr. D. L. Vogelien
 Introduction MOLECULAR GENETICS: TRANSFORMATION AND CLONING adapted by Dr. D. L. Vogelien The field of molecular genetics has resulted in a number of practical applications that have been of tremendous
Introduction MOLECULAR GENETICS: TRANSFORMATION AND CLONING adapted by Dr. D. L. Vogelien The field of molecular genetics has resulted in a number of practical applications that have been of tremendous
Isolation of Candidatus Liberibacter Genes by RAPD and New PCR Detection Technique
 Fourteenth IOCV Conference, 2000 Short Communications Isolation of Candidatus Liberibacter Genes by RAPD and New PCR Detection Technique A. Hocquellet, J. M. Bové, and M. Garnier ABSTRACT. The RAPD technique
Fourteenth IOCV Conference, 2000 Short Communications Isolation of Candidatus Liberibacter Genes by RAPD and New PCR Detection Technique A. Hocquellet, J. M. Bové, and M. Garnier ABSTRACT. The RAPD technique
Purification and Characterization of a DNA Plasmid Part A CHEM 4581: Biochemistry Laboratory I Version: January 18, 2008
 Purification and Characterization of a DNA Plasmid Part A CHEM 4581: Biochemistry Laboratory I Version: January 18, 2008 INTRODUCTION DNA Plasmids. A plasmid is a small double-stranded, circular DNA molecule
Purification and Characterization of a DNA Plasmid Part A CHEM 4581: Biochemistry Laboratory I Version: January 18, 2008 INTRODUCTION DNA Plasmids. A plasmid is a small double-stranded, circular DNA molecule
DNA Transcription. Visualizing Transcription. The Transcription Process
 DNA Transcription By: Suzanne Clancy, Ph.D. 2008 Nature Education Citation: Clancy, S. (2008) DNA transcription. Nature Education 1(1) If DNA is a book, then how is it read? Learn more about the DNA transcription
DNA Transcription By: Suzanne Clancy, Ph.D. 2008 Nature Education Citation: Clancy, S. (2008) DNA transcription. Nature Education 1(1) If DNA is a book, then how is it read? Learn more about the DNA transcription
ONTARIO SCIENCE CENTRE. Teacher Guide. Way to Glow Program
 ONTARIO SCIENCE CENTRE Teacher Guide Way to Glow Program Table of Contents Bacterial transformation background information 3 Experimental procedure 5 Expected results 7 Post-program activity sheet 8 Post-program
ONTARIO SCIENCE CENTRE Teacher Guide Way to Glow Program Table of Contents Bacterial transformation background information 3 Experimental procedure 5 Expected results 7 Post-program activity sheet 8 Post-program
Lecture 3 (FW) January 28, 2009 Cloning of DNA; PCR amplification Reading assignment: Cloning, ; ; 330 PCR, ; 329.
 Lecture 3 (FW) January 28, 2009 Cloning of DNA; PCR amplification Reading assignment: Cloning, 240-245; 286-87; 330 PCR, 270-274; 329. Take Home Lesson(s) from Lecture 2: 1. DNA is a double helix of complementary
Lecture 3 (FW) January 28, 2009 Cloning of DNA; PCR amplification Reading assignment: Cloning, 240-245; 286-87; 330 PCR, 270-274; 329. Take Home Lesson(s) from Lecture 2: 1. DNA is a double helix of complementary
Bac 32, a Novel Bacteriocin Widely Disseminated among Clinical Isolates of Enterococcus faecium
 ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Apr. 2006, p. 1202 1212 Vol. 50, No. 4 0066-4804/06/$08.00 0 doi:10.1128/aac.50.4.1202 1212.2006 Copyright 2006, American Society for Microbiology. All Rights Reserved.
ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Apr. 2006, p. 1202 1212 Vol. 50, No. 4 0066-4804/06/$08.00 0 doi:10.1128/aac.50.4.1202 1212.2006 Copyright 2006, American Society for Microbiology. All Rights Reserved.
The Mosaic Nature of Genomes
 The Mosaic Nature of Genomes n DNA sequence is not static Mutations of single bases Large deletions Large insertions of sequence n Transferred from other species n New functions useful in particular situations
The Mosaic Nature of Genomes n DNA sequence is not static Mutations of single bases Large deletions Large insertions of sequence n Transferred from other species n New functions useful in particular situations
Genetics - Problem Drill 19: Dissection of Gene Function: Mutational Analysis of Model Organisms
 Genetics - Problem Drill 19: Dissection of Gene Function: Mutational Analysis of Model Organisms No. 1 of 10 1. The mouse gene knockout is based on. (A) Homologous recombination (B) Site-specific recombination
Genetics - Problem Drill 19: Dissection of Gene Function: Mutational Analysis of Model Organisms No. 1 of 10 1. The mouse gene knockout is based on. (A) Homologous recombination (B) Site-specific recombination
Plasmids-Mediated Antibiotic Resistance in E. coli
 Plasmids-Mediated Antibiotic Resistance in E. coli (Received: 08.12.1998) Nariman A. H. Aly*; Amany Tharwat** and Wesam F. El-Baz** *Microbial Genetics Dept., Genetic Engineering and Biotechnology Div.,
Plasmids-Mediated Antibiotic Resistance in E. coli (Received: 08.12.1998) Nariman A. H. Aly*; Amany Tharwat** and Wesam F. El-Baz** *Microbial Genetics Dept., Genetic Engineering and Biotechnology Div.,
BIOTECHNOLOGY. Course Syllabus. Section A: Engineering Mathematics. Subject Code: BT. Course Structure. Engineering Mathematics. General Biotechnology
 BIOTECHNOLOGY Subject Code: BT Course Structure Sections/Units Section A Section B Unit 1 Unit 2 Unit 3 Unit 4 Unit 5 Unit 6 Unit 7 Section C Section D Section E Topics Engineering Mathematics General
BIOTECHNOLOGY Subject Code: BT Course Structure Sections/Units Section A Section B Unit 1 Unit 2 Unit 3 Unit 4 Unit 5 Unit 6 Unit 7 Section C Section D Section E Topics Engineering Mathematics General
1a. What is the ratio of feathered to unfeathered shanks in the offspring of the above cross?
 Problem Set 5 answers 1. Whether or not the shanks of chickens contains feathers is due to two independently assorting genes. Individuals have unfeathered shanks when they are homozygous for recessive
Problem Set 5 answers 1. Whether or not the shanks of chickens contains feathers is due to two independently assorting genes. Individuals have unfeathered shanks when they are homozygous for recessive
Spostiamo ora la nostra attenzione sui batteri, e batteriofagi
 Spostiamo ora la nostra attenzione sui batteri, e batteriofagi Bacteria Mutate Spontaneously and Grow at an Exponential Rate. Useful for genetics studies, development of genetic engineering Teoria dell'adattamento
Spostiamo ora la nostra attenzione sui batteri, e batteriofagi Bacteria Mutate Spontaneously and Grow at an Exponential Rate. Useful for genetics studies, development of genetic engineering Teoria dell'adattamento
Construction of plant complementation vector and generation of transgenic plants
 MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
SI Results Peculiarities in the consumption of ammonium and glucose by AmtB- strains SI Materials and Methods Strain Construction
 SI Results Peculiarities in the consumption of ammonium and glucose by AmtB - strains. The AmtB - strain failed to consume all the ammonium in the medium under NH 3 -limiting conditions (.5 mm total ammonium
SI Results Peculiarities in the consumption of ammonium and glucose by AmtB - strains. The AmtB - strain failed to consume all the ammonium in the medium under NH 3 -limiting conditions (.5 mm total ammonium
Certificate of Analysis
 Certificate of Analysis pet6xhn Expression Vector Set Contents Product Information... 1 pet6xhn-n, pet6xhn-c, and pet6xhn-gfpuv Vector Information... 2 Location of Features... 4 Additional Information...
Certificate of Analysis pet6xhn Expression Vector Set Contents Product Information... 1 pet6xhn-n, pet6xhn-c, and pet6xhn-gfpuv Vector Information... 2 Location of Features... 4 Additional Information...
Bacterial defense mechanisms against bacteriophages. B.G.E.T Jayashantha B.Sc (ug) Microbiology (Special), University of Kelaniya, Sri Lanka
 Bacterial defense mechanisms against bacteriophages B.G.E.T Jayashantha B.Sc (ug) Microbiology (Special), University of Kelaniya, Sri Lanka Objective To study the defense mechanisms associated with bacteria
Bacterial defense mechanisms against bacteriophages B.G.E.T Jayashantha B.Sc (ug) Microbiology (Special), University of Kelaniya, Sri Lanka Objective To study the defense mechanisms associated with bacteria
Identification of Beta-Lactamase Enzymes and Prediction of Successful Beta-Lactam Therapy
 JOURNAL OF CLINICAL MICROBIOLOGY, May 1983, p. 791-798 95-1137/83/5791-8$2./ Copyright C 1983, American Society for Microbiology Vol. 17, No. 5 Relative Substrate Affinity Index Values: a Method for Identification
JOURNAL OF CLINICAL MICROBIOLOGY, May 1983, p. 791-798 95-1137/83/5791-8$2./ Copyright C 1983, American Society for Microbiology Vol. 17, No. 5 Relative Substrate Affinity Index Values: a Method for Identification
Chapter 8 Lecture Outline. Transcription, Translation, and Bioinformatics
 Chapter 8 Lecture Outline Transcription, Translation, and Bioinformatics Replication, Transcription, Translation n Repetitive processes Build polymers of nucleotides or amino acids n All have 3 major steps
Chapter 8 Lecture Outline Transcription, Translation, and Bioinformatics Replication, Transcription, Translation n Repetitive processes Build polymers of nucleotides or amino acids n All have 3 major steps
The Regulation of Bacterial Gene Expression
 The Regulation of Bacterial Gene Expression Constitutive genes are expressed at a fixed rate Other genes are expressed only as needed Inducible genes Repressible genes Catabolite repression Pre-transcriptional
The Regulation of Bacterial Gene Expression Constitutive genes are expressed at a fixed rate Other genes are expressed only as needed Inducible genes Repressible genes Catabolite repression Pre-transcriptional
The dialkylglycine decarboxylase repressor DgdR. Functional aspects and relation to other LysR proteins
 11/29/2011 1 The dialkylglycine decarboxylase repressor DgdR. Functional aspects and relation to other LysR proteins 1. Dialkylglycine amino acids 2. In vivo and in vitro studies on the DgdR repressor
11/29/2011 1 The dialkylglycine decarboxylase repressor DgdR. Functional aspects and relation to other LysR proteins 1. Dialkylglycine amino acids 2. In vivo and in vitro studies on the DgdR repressor
Multiple choice questions (numbers in brackets indicate the number of correct answers)
 1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
Concepts: What are RFLPs and how do they act like genetic marker loci?
 Restriction Fragment Length Polymorphisms (RFLPs) -1 Readings: Griffiths et al: 7th Edition: Ch. 12 pp. 384-386; Ch.13 pp404-407 8th Edition: pp. 364-366 Assigned Problems: 8th Ch. 11: 32, 34, 38-39 7th
Restriction Fragment Length Polymorphisms (RFLPs) -1 Readings: Griffiths et al: 7th Edition: Ch. 12 pp. 384-386; Ch.13 pp404-407 8th Edition: pp. 364-366 Assigned Problems: 8th Ch. 11: 32, 34, 38-39 7th
Molecular Biology: DNA sequencing
 Molecular Biology: DNA sequencing Author: Prof Marinda Oosthuizen Licensed under a Creative Commons Attribution license. SEQUENCING OF LARGE TEMPLATES As we have seen, we can obtain up to 800 nucleotides
Molecular Biology: DNA sequencing Author: Prof Marinda Oosthuizen Licensed under a Creative Commons Attribution license. SEQUENCING OF LARGE TEMPLATES As we have seen, we can obtain up to 800 nucleotides
How antimicrobial agents work
 Physical and Chemical Control of Microbes Physical Agents heat or radiation Chemical Agents disinfectants or antiseptics Important Terms 1. Sterilization process of killing all viable microbes 2. Bactericide
Physical and Chemical Control of Microbes Physical Agents heat or radiation Chemical Agents disinfectants or antiseptics Important Terms 1. Sterilization process of killing all viable microbes 2. Bactericide
The UspA1 Protein and a Second Type of UspA2 Protein Mediate Adherence of Moraxella catarrhalis to Human Epithelial Cells In Vitro
 JOURNAL OF BACTERIOLOGY, Mar. 2000, p. 1364 1373 Vol. 182, No. 5 0021-9193/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. The UspA1 Protein and a Second Type of UspA2
JOURNAL OF BACTERIOLOGY, Mar. 2000, p. 1364 1373 Vol. 182, No. 5 0021-9193/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. The UspA1 Protein and a Second Type of UspA2
Contents... vii. List of Figures... xii. List of Tables... xiv. Abbreviatons... xv. Summary... xvii. 1. Introduction In vitro evolution...
 vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
2054, Chap. 13, page 1
 2054, Chap. 13, page 1 I. Microbial Recombination and Plasmids (Chapter 13) A. recombination = process of combining genetic material from 2 organisms to produce a genotype different from either parent
2054, Chap. 13, page 1 I. Microbial Recombination and Plasmids (Chapter 13) A. recombination = process of combining genetic material from 2 organisms to produce a genotype different from either parent
A Novel CTX-M -Lactamase (CTX-M-8) in Cefotaxime-Resistant Enterobacteriaceae Isolated in Brazil
 ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, July 2000, p. 1936 1942 Vol. 44, No. 7 0066-4804/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. A Novel CTX-M -Lactamase (CTX-M-8)
ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, July 2000, p. 1936 1942 Vol. 44, No. 7 0066-4804/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. A Novel CTX-M -Lactamase (CTX-M-8)
Themes: RNA and RNA Processing. Messenger RNA (mrna) What is a gene? RNA is very versatile! RNA-RNA interactions are very important!
 Themes: RNA is very versatile! RNA and RNA Processing Chapter 14 RNA-RNA interactions are very important! Prokaryotes and Eukaryotes have many important differences. Messenger RNA (mrna) Carries genetic
Themes: RNA is very versatile! RNA and RNA Processing Chapter 14 RNA-RNA interactions are very important! Prokaryotes and Eukaryotes have many important differences. Messenger RNA (mrna) Carries genetic
Gene Transfer 11/4/13. Fredrick Griffith in the 1920s did an experiment. Not until 1944 was DNA shown to be the moveable element
 Gene Transfer Fredrick Griffith in the 1920s did an experiment. Not until 19 was DN shown to be the moveable element Dead pathogen cells able to make a capsule were able to pass this ability to the live
Gene Transfer Fredrick Griffith in the 1920s did an experiment. Not until 19 was DN shown to be the moveable element Dead pathogen cells able to make a capsule were able to pass this ability to the live
AP Biology. Chapter 20. Biotechnology: DNA Technology & Genomics. Biotechnology. The BIG Questions. Evolution & breeding of food plants
 What do you notice about these phrases? radar racecar Madam I m Adam Able was I ere I saw Elba a man, a plan, a canal, Panama Was it a bar or a bat I saw? Chapter 20. Biotechnology: DNA Technology & enomics
What do you notice about these phrases? radar racecar Madam I m Adam Able was I ere I saw Elba a man, a plan, a canal, Panama Was it a bar or a bat I saw? Chapter 20. Biotechnology: DNA Technology & enomics
Title: Description of the First Escherichia coli Clinical Isolate Harboring Colistin-
 AAC Accepted Manuscript Posted Online 24 October 2016 Antimicrob. Agents Chemother. doi:10.1128/aac.01945-16 Copyright 2016, American Society for Microbiology. All Rights Reserved. 1 2 Title: Description
AAC Accepted Manuscript Posted Online 24 October 2016 Antimicrob. Agents Chemother. doi:10.1128/aac.01945-16 Copyright 2016, American Society for Microbiology. All Rights Reserved. 1 2 Title: Description
Microbial Biotechnology agustin krisna wardani
 Microbial Biotechnology agustin krisna wardani 1. The Structure of Microbes Microbes (microorganisms) are tiny organisms that are too small to be seen individually by the naked eye and must be viewed with
Microbial Biotechnology agustin krisna wardani 1. The Structure of Microbes Microbes (microorganisms) are tiny organisms that are too small to be seen individually by the naked eye and must be viewed with
Sample Question Paper Session Class XII Biotechnology (045) Marking Scheme
 Sample Question Paper Session 2015-16 Class XII Biotechnology (045) Marking Scheme SECTION-A 1. Methylation Student will add a methyl group to one or two bases within the sequence recognized by enzyme.
Sample Question Paper Session 2015-16 Class XII Biotechnology (045) Marking Scheme SECTION-A 1. Methylation Student will add a methyl group to one or two bases within the sequence recognized by enzyme.
Vectors for Gene Cloning: Plasmids and Bacteriophages
 Vectors for Gene Cloning: Plasmids and Bacteriophages DNA molecule must be able to replicate within the host cell to be able to act as a vector for gene cloning, so that numerous copies of the recombinant
Vectors for Gene Cloning: Plasmids and Bacteriophages DNA molecule must be able to replicate within the host cell to be able to act as a vector for gene cloning, so that numerous copies of the recombinant
7.013 Practice Quiz
 MIT Department of Biology 7.013: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel 7.013 Practice Quiz 2 2004 1 Question 1 A. The primer
MIT Department of Biology 7.013: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel 7.013 Practice Quiz 2 2004 1 Question 1 A. The primer
MOK. Media Optimization Kit
 MOK Media Optimization Kit The Media Optimization Kit determines the best medium formulation for maximizing accumulation of recombinant proteins expressed in E. coli, utilizing a series of Athena s superior
MOK Media Optimization Kit The Media Optimization Kit determines the best medium formulation for maximizing accumulation of recombinant proteins expressed in E. coli, utilizing a series of Athena s superior
Microbiology 微生物学 Spring-Summer
 Microbiology 微生物学 2017 Spring-Summer Relevant Information and Resources Course slides can be found at http://mypage.zju.edu.cn/haichun 教学工作 Course-related questions will be answered through emails. Textbook:
Microbiology 微生物学 2017 Spring-Summer Relevant Information and Resources Course slides can be found at http://mypage.zju.edu.cn/haichun 教学工作 Course-related questions will be answered through emails. Textbook:
Your Gene GGT Term_T7 ATG. Protease Cleavage Site. Name Affinity Tag Protease Cleavage Site Qty Storage. pd454-fh8 FH8 PCS_TEV 10Rx -20
 IP-Free E. coli Inducible Expression Vectors E. coli expression vectors are available with the following promoters: T5 or T7 (IPTG-inducible), rhabad (rhamnose-inducible), ara (arabinose and IPTG-inducible)
IP-Free E. coli Inducible Expression Vectors E. coli expression vectors are available with the following promoters: T5 or T7 (IPTG-inducible), rhabad (rhamnose-inducible), ara (arabinose and IPTG-inducible)
Practice Test #3. Multiple Choice Identify the choice that best completes the statement or answers the question.
 Practice Test #3 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. An application of using DNA technology to help environmental scientists would be _. a.
Practice Test #3 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. An application of using DNA technology to help environmental scientists would be _. a.
RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by. 5 AGACACAAACACCAUUGUCACACUCCACAGC; Rand-2 OMe,
 Materials and methods Oligonucleotides and DNA constructs RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by Dharmacon Inc. (Lafayette, CO). The sequences were: 122-2 OMe, 5
Materials and methods Oligonucleotides and DNA constructs RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by Dharmacon Inc. (Lafayette, CO). The sequences were: 122-2 OMe, 5
NEW OUTCOMES IN MUTATION RATES ANALYSIS
 NEW OUTCOMES IN MUTATION RATES ANALYSIS Valentina Agoni valentina.agoni@unipv.it The GC-content is very variable in different genome regions and species but although many hypothesis we still do not know
NEW OUTCOMES IN MUTATION RATES ANALYSIS Valentina Agoni valentina.agoni@unipv.it The GC-content is very variable in different genome regions and species but although many hypothesis we still do not know
Cloning of a Gene Responsible for the Biosynthesis of Glutathione in Escherichia coli B
 APPLIED AND ENVIRONMENTAL MICROBIOLOGY, Dec. 1982, P. 1444-1448 Vol. 44, No. 6 0099-2240/82/121444-05$02.00/0 Copyright C 1982, American Society for Microbiology Cloning of a Gene Responsible for the Biosynthesis
APPLIED AND ENVIRONMENTAL MICROBIOLOGY, Dec. 1982, P. 1444-1448 Vol. 44, No. 6 0099-2240/82/121444-05$02.00/0 Copyright C 1982, American Society for Microbiology Cloning of a Gene Responsible for the Biosynthesis
SME-Type Carbapenem-Hydrolyzing Class A -Lactamases from Geographically Diverse Serratia marcescens Strains
 ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Nov. 2000, p. 3035 3039 Vol. 44, No. 11 0066-4804/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. SME-Type Carbapenem-Hydrolyzing
ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Nov. 2000, p. 3035 3039 Vol. 44, No. 11 0066-4804/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. SME-Type Carbapenem-Hydrolyzing
Chapter 13 - Concept Mapping
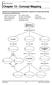 Chapter 13 - Concept Mapping Using the terms and phrases provided below, complete the concept map showing the discovery of DNA structure. amount of base pairs five-carbon sugar purine DNA polymerases Franklin
Chapter 13 - Concept Mapping Using the terms and phrases provided below, complete the concept map showing the discovery of DNA structure. amount of base pairs five-carbon sugar purine DNA polymerases Franklin
NAME TA SEC Problem Set 3 FRIDAY March 5, Problem sets will NOT be accepted late.
 MIT Department of Biology 7.013: Introductory Biology - Spring 2004 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. laudette ardel NME T SE 7.013 Problem Set 3 FRIDY March 5, 2004 Problem
MIT Department of Biology 7.013: Introductory Biology - Spring 2004 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. laudette ardel NME T SE 7.013 Problem Set 3 FRIDY March 5, 2004 Problem
H. Wu, B.-G. Liu, J.-H. Liu, Y.-S. Pan, L. Yuan and G.-Z. Hu
 Phenotypic and molecular characterization of CTX-M-14 extended-spectrum β-lactamase and plasmid-mediated ACT-like AmpC β-lactamase produced by Klebsiella pneumoniae isolates from chickens in Henan Province,
Phenotypic and molecular characterization of CTX-M-14 extended-spectrum β-lactamase and plasmid-mediated ACT-like AmpC β-lactamase produced by Klebsiella pneumoniae isolates from chickens in Henan Province,
Lectures of Dr.Mohammad Alfaham. The Bacterial Genetics
 Lectures of Dr.Mohammad Alfaham The Bacterial Genetics is the total collection of genes carried by a bacterium both on its chromosome and on its extrachromosomal genetic elements (plasmids) A Gene A gene
Lectures of Dr.Mohammad Alfaham The Bacterial Genetics is the total collection of genes carried by a bacterium both on its chromosome and on its extrachromosomal genetic elements (plasmids) A Gene A gene
Relationship between nucleotide sequence and 3D protein structure of six genes in Escherichia coli, by analysis of DNA sequence using a Markov model
 Relationship between nucleotide sequence and 3D protein structure of six genes in Escherichia coli, by analysis of DNA sequence using a Markov model Yuko Ohfuku 1,, 3*, Hideo Tanaka and Masami Uebayasi
Relationship between nucleotide sequence and 3D protein structure of six genes in Escherichia coli, by analysis of DNA sequence using a Markov model Yuko Ohfuku 1,, 3*, Hideo Tanaka and Masami Uebayasi
Chapter 15 Gene Technologies and Human Applications
 Chapter Outline Chapter 15 Gene Technologies and Human Applications Section 1: The Human Genome KEY IDEAS > Why is the Human Genome Project so important? > How do genomics and gene technologies affect
Chapter Outline Chapter 15 Gene Technologies and Human Applications Section 1: The Human Genome KEY IDEAS > Why is the Human Genome Project so important? > How do genomics and gene technologies affect
HiPer Transformation Teaching Kit
 HiPer Transformation Teaching Kit Product Code: HTBM017 Number of experiments that can be performed: 10 Duration of Experiment: 4 days Day 1- Preparation of media and revival of E. coli Host Day 2- Inoculation
HiPer Transformation Teaching Kit Product Code: HTBM017 Number of experiments that can be performed: 10 Duration of Experiment: 4 days Day 1- Preparation of media and revival of E. coli Host Day 2- Inoculation
pfb and pfb-neo Retroviral Vectors
 pfb and pfb-neo Retroviral Vectors INSTRUCTION MANUAL Catalog #217563 (pfb Retroviral Vector) and #217561 (pfb-neo Retroviral Vector) Revision A For In Vitro Use Only 217561-12 LIMITED PRODUCT WARRANTY
pfb and pfb-neo Retroviral Vectors INSTRUCTION MANUAL Catalog #217563 (pfb Retroviral Vector) and #217561 (pfb-neo Retroviral Vector) Revision A For In Vitro Use Only 217561-12 LIMITED PRODUCT WARRANTY
Patentability/Literature Research
 Patentability/Literature Research The probiotic-based solution to combat cholera as presented here comprises of two major innovative components: (1) the metabolite-dependent pathogen inhibition by a probiotic
Patentability/Literature Research The probiotic-based solution to combat cholera as presented here comprises of two major innovative components: (1) the metabolite-dependent pathogen inhibition by a probiotic
An extended-spectrum AmpC-type L-lactamase obtained by in vitro antibiotic selection
 FEMS Microbiology Letters 165 (1998) 85^90 An extended-spectrum AmpC-type L-lactamase obtained by in vitro antibiotic selection Mar èa-isabel Morosini a, Mar èa-cristina Negri a, Brian Shoichet b, Mar
FEMS Microbiology Letters 165 (1998) 85^90 An extended-spectrum AmpC-type L-lactamase obtained by in vitro antibiotic selection Mar èa-isabel Morosini a, Mar èa-cristina Negri a, Brian Shoichet b, Mar
Biotechnology and Genomics in Public Health. Sharon S. Krag, PhD Johns Hopkins University
 This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike License. Your use of this material constitutes acceptance of that license and the conditions of use of materials on this
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike License. Your use of this material constitutes acceptance of that license and the conditions of use of materials on this
7.1 The lac Operon 7-1
 7.1 The lac Operon The lac operon was the first operon discovered It contains 3 genes coding for E. coli proteins that permit the bacteria to use the sugar lactose Galactoside permease (lacy) which transports
7.1 The lac Operon The lac operon was the first operon discovered It contains 3 genes coding for E. coli proteins that permit the bacteria to use the sugar lactose Galactoside permease (lacy) which transports
that no inhibitory secondary structures that might block the From these findings we concluded that all or part of the
 Proc. Natl. Acad. Sci. USA Vol. 89, pp. 2605-2609, April 1992 Biochemistry A uridine-rich sequence required for translation of prokaryotic mrna (Shine-Dalgarno sequence/ribosomal protein Sl/protein synthesis)
Proc. Natl. Acad. Sci. USA Vol. 89, pp. 2605-2609, April 1992 Biochemistry A uridine-rich sequence required for translation of prokaryotic mrna (Shine-Dalgarno sequence/ribosomal protein Sl/protein synthesis)
Received 3 August 2000/Accepted 26 October 2000
 JOURNAL OF BACTERIOLOGY, Feb. 2001, p. 1096 1100 Vol. 183, No. 3 0021-9193/01/$04.00 0 DOI: 10.1128/JB.183.3.1096 1100.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved. Transcript
JOURNAL OF BACTERIOLOGY, Feb. 2001, p. 1096 1100 Vol. 183, No. 3 0021-9193/01/$04.00 0 DOI: 10.1128/JB.183.3.1096 1100.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved. Transcript
Use of Drosophila Melanogaster as a Model System in the Study of Human Sodium- Dependent Multivitamin Transporter. Michael Brinton BIOL 230W.
 Use of Drosophila Melanogaster as a Model System in the Study of Human Sodium- Dependent Multivitamin Transporter Michael Brinton BIOL 230W.001 28 October 2013 TA: Sashi Gollapudi Introduction Many human
Use of Drosophila Melanogaster as a Model System in the Study of Human Sodium- Dependent Multivitamin Transporter Michael Brinton BIOL 230W.001 28 October 2013 TA: Sashi Gollapudi Introduction Many human
HOUR EXAM II BIOLOGY 422 FALL, In the spirit of the honor code, I pledge that I have neither given nor received help on this exam.
 Name First Last (Please Print) PID Number - HOUR EXAM II BIOLOGY 422 FALL, 2010 In the spirit of the honor code, I pledge that I have neither given nor received help on this exam. 1 Signature 2 3 4 5 6
Name First Last (Please Print) PID Number - HOUR EXAM II BIOLOGY 422 FALL, 2010 In the spirit of the honor code, I pledge that I have neither given nor received help on this exam. 1 Signature 2 3 4 5 6
THE BENEFITS AND USES OF MICROBES
 MODULE 4 MICROBES AND MICROBIAL BIOTECHNOLOGY U N I T 2 THE BENEFITS AND USES OF MICROBES A. MICROBIAL BIOTECHNOLOGY 1 Read What is biotechnology? and decide which of the words below can be used instead
MODULE 4 MICROBES AND MICROBIAL BIOTECHNOLOGY U N I T 2 THE BENEFITS AND USES OF MICROBES A. MICROBIAL BIOTECHNOLOGY 1 Read What is biotechnology? and decide which of the words below can be used instead
Genetic support of Extended- Spectrum ß-Lactamases
 Genetic support of Extended- Spectrum ß-Lactamases Laurent Poirel Dept of Microbiology (Pr Nordmann) Bicêtre Hospital. South-Paris Medical School. France ESCMID Conference on ESBL 29-31 May 2006 TEM-like
Genetic support of Extended- Spectrum ß-Lactamases Laurent Poirel Dept of Microbiology (Pr Nordmann) Bicêtre Hospital. South-Paris Medical School. France ESCMID Conference on ESBL 29-31 May 2006 TEM-like
