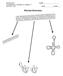
Supplementary Figure 1 Taxonomy of the family Camelidae. The graph shows the six. species of extant camelids distributed among three genera.
|
|
|
- Elvin Carpenter
- 5 years ago
- Views:
Transcription
1 Supplementary Figure 1 Taxonomy of the family Camelidae. The graph shows the six species of extant camelids distributed among three genera. 1
2 a b 2
3 c Supplementary Figure 2 17 bp-mer estimation of the genome size. (a) 17 bp-mer estimation of the genome size of Bactrian camel. (b) 17 bp-mer estimation of the genome size of dromedary. (c) 17 bp-mer estimation of the genome size of alpaca. The X-axis represents the sequencing depth. The Y-axis represents the proportion of a K-mer count in total K-mer counts at a given sequencing depth. The estimated genome size for Bactrian camel, dromedary and alpaca are 2.4Gb, 2.5Gb and 2.6Gb respectively. 3
4 Supplementary Figure 3 GC content distribution for the three camelids genomes. The x-axis shows the GC content and the y-axis represents the ratio of the bin number divided by the total windows. These three organisms have a similar GC content distribution. 4
5 A B Supplementary Figure 4 Error types of genome assemble. (A) type I error of genome assembly which only single reads from PE can map to the region. (B) type II error of genome assembly which PE reads cannot map to the region. 5
6 Supplementary Figure 5 Comparison of the distribution of several features in the final gene set for the four mammals. Window refers to the length of every point. No obvious unexpected differences exists among these four organisms, indicating the high quality of gene structure annotation. 6
7 Supplementary Figure 6 Protein identity among the gene sets of the three camelids. Bactrian camel and dromedary shared 94.93% average amino acid identity among 14,071 1:1 orthologs, Bactrian camel and alpaca had an identity of 91.55% for 16,145 orthologs, and dromedary and alpaca had 90.8% for 15,167 orthologs. The results showed a high similarity for the three organisms. 7
8 Supplementary Figure 7 Distributions of Ka/Ks among the gene sets of the three camelids. These single copy orthologs are 1:1 Bactrian camel-dromedary, 1:1 Bactrian camel-alpaca and 1:1 dromedary-alpaca. 8
9 Supplementary Figure 8 Orthologous protein composition inferred for ten genomes. The ten species are opossum (Monodelphis domestica), horse (Equus caballus), human (Homo sapiens), dog (Canis lupus familiaris), panda (Ailuropoda melanoleuca), Bactrian camel (Camelus bactrianus), cow (Bos taurus), dromedary (Camelus dromedarius), mouse (Mus musculus) and alpaca (Vicugna pacos). 9
10 Supplementary Figure 9 Phylogenetic tree based on four-fold degenerated sites in 7,398 single-copy orthologous genes by using the maximum likehood method. Single-copy orthologous genes were from the selected genomes, including opossum (Monodelphis domestica), human (Homo sapiens), mouse (Mus musculus), dromedary (Camelus dromedarius), Bactrian camel (Camelus bactrianus), alpaca (Vicugna pacos), cow (Bos taurus), horse (Equus caballus), dog (Canis lupus familiaris) and panda (Ailuropoda melanoleuca). Branch lengths represent the neutral divergence rates. Blue numbers on the branches represent a LRT values, which illustrate the reliability of branches calculated by PhyML. 10
11 Supplementary Figure 10 Estimation of divergence time. The numbers on the nodes represent the divergence times from present (million years ago, Mya). The red dots in four internal nodes indicate fossil calibration times for Homo sapiens-mus musculus divergence ( Mya) 1, Canis lupus familiaris-equus caballus divergence ( Mya) 1, Bos taurus-homo sapiens divergence ( Mya) 1, and Homo sapiens-monodelphis domestica divergence ( Mya) ( were used in the analysis. The graph showed the estimated divergence times with their 95% confidence intervals. 11
12 Supplementary Figure 11 Branch specific ω values by using the dataset of 7,398 single-copy orthologs. Different ω values showed different nature selected pressures among the evolutionary process. The ω values for Bactrian camel and dromedary are higher than other species, indicating an accelerated evolution. 12
13 Supplementary Figure 12 Heterozygous SNP density of genomes of the three camelids. The blue, red and green lines represent the genomic SNP densities of Bactrian camel, dromedary and alpaca, respectively. Heterozygous SNPs of each individual genome were annotated, then non-overlapping 50 kb windows were chosen and heterozygous density in each window was calculated. 13
14 Supplementary Figure 13 Hierarchical clustering of over-represented GO categories for expanded gene families in Bactrian camel genome. The over-represented GO categories ( molecular function group only) were clustered hierarchically using the method of Kosiol et al
15 Supplementary Figure 14 Hierarchical clustering of over-represented GO categories for expanded gene families in dromedary genome. The over-represented GO categories ( molecular function group only) were clustered hierarchically using the method of Kosiol et al
16 Supplementary Figure 15 Hierarchical clustering of over-represented GO categories for expanded gene families in alpaca genome. The over-represented GO categories ( molecular function group only) were clustered hierarchically using the method of Kosiol et al
17 Supplementary Figure 16 Hierarchical clustering of over-represented GO categories for unique amino acid residues changed genes in Bactrian camel genome. The over-represented GO categories ( molecular function group only) were clustered hierarchically using the method of Kosiol et al
18 Supplementary Figure 17 Hierarchical clustering of over-represented GO categories for unique amino acid residues changed genes in dromedary genome. The over-represented GO categories ( molecular function group only) were clustered hierarchically using the method of Kosiol et al
19 Supplementary Figure 18 Volcano plot of expression levels in renal cortex of Bactrian camel. The negative log 10 P-values (y-axis) are plotted against the log 2 fold changes (x-axis) in gene expression for comparison of control group with water-restricted conditions. Based on the normalized expression values, genes matching false discovery rate (FDR) 0.01 and log2(fold change) 1 were defined as differentially expressed genes (DEGs) in different conditions. The plot indicates the up-regulated genes (red), down-regulated genes (green), or not DEG (blue) in the renal cortex of Bactrian camels under WR. 19
20 Supplementary Figure 19 Volcano plot of expression levels in renal medulla of Bactrian camel. The negative log 10 P-values (y-axis) are plotted against the log 2 fold changes (x-axis) in gene expression for comparison of control group with water-restricted conditions. Based on the normalized expression values, genes matching false discovery rate (FDR) 0.01 and log2(fold change) 1 were defined as differentially expressed genes (DEGs) in different conditions. The plot indicates the up-regulated genes (red), down-regulated genes (green), or not DEG (blue) in the renal medulla of Bactrian camels under WR. 20
21 Supplementary Figure 20 Distribution of expression levels in renal cortex of Bactrian camel. The log 10 (RPKM of water-restricted condition) (y-axis) are plotted against the log 10 (normalized gene expression values of control group condition) (x-axis) in gene expression. Based on the normalized expression values, genes matching false discovery rate (FDR) 0.01 and log2(fold change) 1 were defined as differentially expressed genes (DEGs) in different conditions. The plot indicates the up-regulated genes (red), down-regulated genes (green), or not DEG (blue) in the renal cortex of Bactrian camels under WR. RPKM: reads per kilobase per million mapped reads. 21
22 Supplementary Figure 21 Distribution of expression levels in renal medulla of Bactrian camel. The log 10 (RPKM of water-restricted condition) (y-axis) are plotted against the log 10 (normalized gene expression values of control group condition) (x-axis) in gene expression. Based on the normalized expression values, genes matching false discovery rate (FDR) 0.01 and log2(fold change) 1 were defined as differentially expressed genes (DEGs) in different conditions. The plot indicates the up-regulated genes (red), down-regulated genes (green), or not DEG (blue) in the renal medulla of Bactrian camels during WR. RPKM: reads per kilobase per million mapped reads. 22
23 Supplementary Figure 22 GO classification of significantly up regulation of genes expression in renal cortex of Bactrian camel under water restriction condition. The enriched GO were shown mainly in three categories: biological process, cellular component and molecular function. 23
24 Supplementary Figure 23 GO classification of significantly down regulation of genes expression in renal cortex of Bactrian camel under water restriction condition. The enriched GO were shown mainly in three categories: biological process, cellular component and molecular function. 24
25 Supplementary Figure 24 GO classification of significantly up regulation of genes expression in renal medulla of Bactrian camel under water restriction condition. The enriched GO were shown mainly in three categories: biological process, cellular component and molecular function. 25
26 Supplementary Figure 25 GO classification of significantly down regulation of genes expression in renal medulla of Bactrian camel under water restriction condition. The enriched GO were shown mainly in three categories: biological process, cellular component and molecular function. 26
27 Supplementary Figure 26 A heat map of the expression patterns of AQP family. The heat map was generated based on hierarchical cluster analysis of expression of AQP family in renal cortex and renal medulla for control group and water restriction. 27
28 Supplementary Figure 27 Bactrian camels unique amino acid changes in the AQP4 gene. Red rectangle indicates Bactrian camel s unique amino acid change with R261C. 28
29 Supplementary Table 1 Statistics of raw data for the Bactrian camel genome*. Pair-end Libraries Solexa Reads Insert Size Average Reads Length (bp) Total Data (Gb) Sequence Coverage (X) Physical Coverage (X) 170 bp bp bp kb kb kb kb Total * The genome size is estimated to be 2.4 Gb for Bactrian camel genome (see Supplementary Table 7). 29
30 Supplementary Table 2 Statistics of raw data for the dromedary genome*. Pair-end Libraries Solexa Reads Insert Size Average Reads Length (bp) Total Data (Gb) Sequence Coverage (X) Physical Coverage (X) 170 bp bp bp kb kb kb Total * The genome size is estimated to be 2.3 Gb for dromedary genome (see Supplementary Table 7). 30
31 Supplementary Table 3 Statistics of raw data for the alpaca genome*. Pair-end Libraries Insert Size Average Reads Length (bp) Total Data (Gb) Sequence Coverage (X) Physical Coverage (X) 170 bp bp Solexa 800 bp Reads 2 kb kb kb Total * The genome size is estimated to be 2.6 Gb for alpaca genome (see Supplementary Table 7). 31
32 Supplementary Table 4 Statistics after error correction for the Bactrian camel genome*. Pair-end Libraries Solexa Reads Insert Size Total Data (Gb) Average Reads Length (bp) Sequence Coverage (X) Physical Coverage (X) 170 bp bp bp kb kb kb kb Total * The genome size is estimated to be 2.4 Gb for Bactrian camel genome (see Supplementary Table 7). 32
33 Supplementary Table 5 Statistics after error correction for the dromedary genome*. Pair-end Libraries Solexa Reads Insert Size Average Reads Length (bp) Total Data (Gb) Sequence Coverage (X) Physical Coverage (X) 170 bp bp bp kb kb kb Total * The genome size is estimated to be 2.3 Gb for dromedary genome (see Supplementary Table 7). 33
34 Supplementary Table 6 Statistics after error correction for the alpaca genome*. Pair-end Libraries Solexa Reads Insert Size Average Reads Length (bp) Total Data (Gb) Sequence Coverage (X) Physical Coverage (X) 170 bp bp bp kb kb kb Total * The genome size is estimated to be 2.6 Gb for alpaca genome (see Supplementary Table 7). 34
35 Supplementary Table 7 17-mer statistics for the genomes of three species. Species K number K depth (X) Genome Size (bp) Used Base (bp) Used Read Physical coverage (X) C.bactrianus 51,384,018, ,446,858,035 62,606,245, ,389, C.dromedarius 45,493,835, ,274,691,798 59,913,535, ,231, V.pacos 92,057,544, ,630,215, ,039,540,270 1,123,874,
36 Supplementary Table 8 Statistics of the Bactrian camel genome assembly. Contig Scaffold Size (bp) Number Size (bp) Number N90 6,585 82,329 1,733, N80 11,069 59,395 3,388, N70 15,392 44,207 4,768, N60 19,894 32,837 6, N50 24,909 23,893 8,760, Longest 203, ,538, Total Size 1,992,031, ,005,902, Total Number (>100 bp) , ,480 Total Number (>2 kb) , ,067 36
37 Supplementary Table 9 Statistics of the dromedary genome assembly. Contig Scaffold Size (bp) Number Size (bp) Number N90 11,564 40, , N80 21,141 27,746 1,275, N70 30,944 19,994 2,122, N60 41,675 14,457 2,976, N50 54,135 10,265 4,123, Longest 539,544 23,736,781 Total Size 1,992,183,914 2,014,928,193 Total Number (>100 bp) 203, ,723 Total Number (>2 kb) 67,075 3,395 37
38 Supplementary Table 10 Statistics of the alpaca genome assembly. Contig Scaffold Size (bp) Number Size (bp) Number N90 13,519 33, , N80 26,079 22,965 1,948, N70 38,461 16,564 2,898, N60 51,492 11,988 3,985, N50 66,333 8,491 5,130, Longest 678, ,394, Total Size 2,036,384, ,049,118, Total Number ( 100 bp) , ,453 Total Number ( 2 kb) -- 53, ,314 38
39 Supplementary Table 11 Assessment of genome coverage by assembled transcripts of Bactrian camel. Dataset Number Total Length (bp) Covered by Assembly With >90% Sequence in one Scaffold Number With >50% Sequence in one Scaffold Percent(%) Number Percent(%) All >200bp >500bp >1000bp
40 Supplementary Table 12 Assessment of wild Bactrian camel genome* coverage by assembled transcripts of Bactrian camel. Dataset Number Total Length (bp) Covered by Assembly With >90% Sequence in one Scaffold Number With >50% Sequence in one Scaffold Percent(%) Number Percent(%) All >200bp >500bp >1000bp * The wild Bactrian camel genome was derived from the previous research 3. 40
41 Supplementary Table 13 Statistics of the aligned genomes. Genome # scaffolds/chr Genome size (Gb, with N) Genome Size (Gb, without N) Genome Size (Mb, masked) %masked V.pacos 319, C.bactrianus 140, C.dromedarius 117, H.sapiens , B.taurus 12, , Note: The genomes were masked with Ns in repeat sequence regions prior to LASTZ alignment. 41
42 Supplementary Table 14 Genome alignment statistics. Species vs Species Aligned Length(Gb) Target Genome Coverage Rate Query Genome Coverage Rate V.pacos vs H.sapiens % 88.85% V.pacos vs C.bactrianus % 88.81% V.pacos vs C.dromedarius % 83.64% C.bactrianus vs B.taurus % 95.44% C.dromedarius vs B.taurus % 86.55% C.dromedarius vs C.bactrianus % 90.63% 42
43 Supplementary Table 15 Segmental duplications in the three camelids genomes. Cutoff (kb) Number of Median size Genome SDs (bp) coverage (bp) >1 7,444 1,774 26,686,484 Bactrian camel > ,589 13,453,011 > ,133 7,525,705 > , ,532 >1 10,604 1,628 26,087,631 Dromedary > ,803 10,325,058 > ,554 4,356,878 > >1 24,295 2,124 36,422,148 Alpaca >5 3,823 7,152 19,304,290 > ,543 9,940,988 > , ,636 The segmental duplications in these three organisms were detected with different cutoff size (1 kb, 5 kb, 10 kb and 50 kb). 43
44 Supplementary Table 16 Statistics of the predicted protein coding genes for alpaca genome. De novo Homolog Gene set Number Average transcript length (bp) Average CDS length (bp) Average exon per gene Average exon length (bp) Average intron length (bp) AUGUSTUS 21,781 44,725 1, ,503 GENSCAN 45,949 30,335 1, ,201 H.sapiens 19,148 24,163 1, ,961 C.familiaris 20,254 21,856 1, ,752 S.scrofa 18,836 18,652 1, ,708 A.melanoleuca 19,703 22,209 1, ,840 C.bactrianus 22,356 23,511 1, B.taurus 19,737 22,012 1, ,761 GLEAN 20,864 24,788 1, ,147 Final set 20,864 24,788 1, ,147 44
45 Supplementary Table 17 Statistics of the predicted protein coding genes for Bactrian camel genome. Gene set Number Average transcript length (bp) Average CDS length (bp) Average exons per gene Average exon length (bp) Average intron length (bp) De novo AUGUSTUS 19,062 51,245 1, ,662 GENSCAN 39,581 34,865 1, ,357 B.taurus 30,670 24,738 1, ,915 Homolog C.familiaris 19,302 22,516 1, ,697 H.sapiens 31,506 16,391 1, ,580 S.scrofa 50,642 9, ,143 GLEAN 16,758 44,844 1, ,642 C.familiaris 17,323 26,634 1, ,901 Synteny H.sapiens 17,211 32,467 1, ,571 S.scrofa 15,107 21,527 1, ,779 Synteny-nr* 17,477 31,688 1, ,450 Final set 20,251 32,103 1, ,804 *Synteny-nr: Synteny non-redundancy. The synteny non-redundancy gene sets are predicted by LASTZ based on the C.familiaris, H.sapiens and S.scrofa. 45
46 Supplementary Table 18 Evaluation of completeness of the Bactrian camel genome assembly using core eukaryotic gene mapping approach. Parameter Number Percent (%) Total KOGs One KOG align one gene One KOG align several genes One KOG align no gene Note: KOG is mammal s orthologous gene found in the genome. 46
47 Supplementary Table 19 Evaluation of completeness of the dromedary genome assembly using core eukaryotic gene mapping approach. Parameter Number Percent (%) Total KOGs One KOG align one gene One KOG align several genes One KOG align no gene Note: KOG is mammal s orthologous gene found in the genome. 47
48 Supplementary Table 20 Evaluation of completeness of the alpaca genome assembly using core eukaryotic gene mapping approach. Parameter Number Percent (%) Total KOGs One KOG align one gene One KOG align several genes One KOG align no gene Note: KOG is mammal s orthologous gene found in the genome. 48
49 Supplementary Table 21 The number of genes with homology or functional classification using each annotation method for Bactrian camel genome. Number Percent (%) Total 20, % Annotated 19, SwissProt 18, Annotated TrEMBL 19, InterPro 16, KEGG 14, GO 13, Unannotated 1,
50 Supplementary Table 22 The number of genes with homology or functional classification using each annotation method for dromedary genome. Number Percent (%) Total 20, % Annotated InterPro 17, GO 14, KEGG 15, Swissprot 20, TrEMBL 18, Unannotated
51 Supplementary Table 23 The number of genes with homology or functional classification using each annotation method for alpaca genome. Number Percent (%) Total 20, % Annotated InterPro 16, GO 13, KEGG 14, Swissprot TrEMBL Unannotated 1,
52 Supplementary Table 24 Statistics of the repeats for Bactrian camel genome. RepBase TEs TE proteins De novo Combined TEs Length (bp) % in genome Length (bp) % in genome Length (bp) % in genome Length (bp) % in genome DNA 64,835, ,974, ,472, ,366, LINE 343,627, ,766, ,876, ,420, LTR 115,311, ,536, ,557, ,544, SINE 52,747, ,959, ,099, Other 9, , Unknown 1,003, ,648, ,649, Total 573,690, ,255, ,253, ,144, Repbase TEs: results of RepeatMasker analysis using Repbase; TE proteins: results of RepeatProteinMask analysis using Repbase; De novo: results of RepeatMasker analysis using the library predicted by the de novo method; Combined: the combined results for Repbase TEs, TE proteins, and the de novo method. 52
53 Supplementary Table 25 Statistics of the repeats for dromedary genome. RepBase TEs TE Proteins De novo Combined TEs Length (Mb) %in Genome Length (Mb) % in Genome Length (Mb) % in Genome Length (Mb) % in Genome DNA LINE LTR SINE Other Unknown Total Repbase TEs: results of RepeatMasker analysis using Repbase; TE proteins: results of RepeatProteinMask analysis using Repbase; De novo: results of RepeatMasker analysis using the library predicted by the de novo method; Combined: the combined results for Repbase TEs, TE proteins, and the de novo method. 53
54 Supplementary Table 26 Statistics of the repeats for alpaca genome. RepBase TEs TE Proteins De novo Combined TEs Length (bp) %in Genome Length (bp) % in Genome Length (bp) % in Genome Length (bp) % in Genome DNA 64,297, ,987, ,294, ,344, LINE 366,454, ,134, ,524, ,933, LTR 122,782, ,636, ,878, ,861, SINE 45,617, ,121, ,497, Other 6, , Unknown ,437, ,437, Total 595,690, ,738, ,070, ,532, Repbase TEs: results of RepeatMasker analysis using Repbase; TE proteins: results of RepeatProteinMask analysis using Repbase; De novo: results of RepeatMasker analysis using the library predicted by the de novo method; Combined: the combined results for Repbase TEs, TE proteins, and the de novo method. 54
55 Supplementary Table 27 Repeats comparison in five species genomes. Bactrian camel Alpaca Dromedary Cattle Human Length (Mb) % in genome Length (Mb) % in genome Length (Mb) % in genome Length (Mb) % in genome Length (Mb) % in genome DNA LINE LTR SINE Other Unknown Total 609,
56 Supplementary Table 28 Non-coding RNA genes in the Bactrian camel genome. Type Copy # Average length (bp) Total length (bp) % of genome mirna , trna , rrna snrna rrna , S , S , S S , snrna 1, , CD-box , HACA-box , splicing , Total ,
57 Supplementary Table 29 Non-coding RNA genes in the dromedary genome. Type Copy # Average length (bp) Total length (bp) % of genome trna , rrna , mirna , snrna CD-box , H-ACA box , scarna , Spliceosomal RNA , Total 2, ,
58 Supplementary Table 30 Non-coding RNA genes in the alpaca genome. Type Copy # Average length (bp) Total length (bp) % of genome mirna , trna , rrna snrna 18S ,, S , S S , CD-box , HACA-box , splicing , Total ,
59 Supplementary Table 31 Statistics for the orthologous gene families of these ten species genomes. Species Genes number Unclustered genes Family number Unique families Average genes per family Bactrian camel 20, , Dromedary 20, , Alpaca 20,864 1,867 15, Cattle 21, , Horse 20, , Dog 19, , Panda 19, , Human 21,642 1,195 15, Mouse 22,843 1,851 15, Opossum 19,448 1,195 15, Note: Unclustered genes refer to those specific to the current species; unique families refer to those specific to the current species. 59
60 Supplementary Table 32 Branch specific ω values of ten species. Species Mean ω 95% confidence interval of ω Low Opossum Human Mouse Dromedary Bactrian camel Alpaca Cattle Horse Dog Panda Up 60
61 Supplementary Table 33 Statistics of identified SNPs for Bactrian camel genome. Type Analyzed regions Heterozygote Heterozygote rate ( 10-3 ) Genome 1,976,686,993 2,289, Autosomes 1,883,590,687 2,237, Chr.X* 93,096,306 52, CDS 30,798,739 21, Autosomes 29,713,812 20, Chr.X* 1,084, *The X-chromosome-derived scaffolds were identified by LASTZ alignment to the cattle X chromosome. 61
62 Supplementary Table 34 Statistics of identified SNPs for dromedary genome. Type Analyzed regions Heterozygote Heterozygote rate ( 10-3 ) Genome 1,976,042,088 1,462, Autosomes 1,884,026,699 1,443, Chr.X* 92,015,389 18, CDS 30,874,565 13, Autosomes 29,788,584 13, Chr.X* 1,085, *The X-chromosome-derived scaffolds were identified by LASTZ alignment to the cattle X chromosome. 62
63 Supplementary Table 35 Statistics of identified SNPs for alpaca genome. Type Analyzed regions Heterozygote Heterozygote rate ( 10-3 ) Genome 2,006,396,020 5,346, Autosomes 1,907,527,860 5,237, Chr.X* 98,868, , CDS 42,584,726 81, Autosomes 40,525,019 79, Chr.X* 2,059,707 1, *The X-chromosome-derived scaffolds were identified by LASTZ alignment to the cattle X chromosome. 63
64 Supplementary Table 36 GO enrichment analysis for expanded gene families in Bactrian camel genome. GO ID Description Taxonomy P-value Number of genes GO: olfactory receptor activity MF 1.45e GO: G-protein coupled receptor activity MF 1.29e GO: G-protein coupled receptor protein BP 1.04e signaling pathway GO: cellular component organization or BP 1.4e biogenesis at cellular level GO: receptor activity MF 4.4e GO: cellular macromolecular complex BP 1.3e assembly GO: cellular component biogenesis BP 1.5e GO: intracellular non-membrane-bounded CC 6.0e organelle GO: integral to membrane CC 9.6e GO: signal transduction BP 6.2e GO: nucleosome CC 2.3e GO: nucleosome assembly BP 6.7e GO: intracellular organelle part CC 1.8e GO: GTP binding MF 4.6e GO: macromolecular complex CC 6.1e GO: GTPase activity MF 1.4e GO: microtubule CC 4.6e GO: protein polymerization BP 1.3e GO: biological regulation BP 1.5e GO: regulation of cellular process BP 4.3e GO: structural molecule activity MF 8.1e GO: cell part CC 2.3e GO: odorant binding MF 2.1e-11 6 GO: cellular process BP 2.8e GO: microtubule-based movement BP 4.2e GO: membrane CC 4.9e GO: intracellular organelle CC 2.8e GO: nucleus CC 2.1e GO: pheromone binding MF 4.6e-06 3 GO: cysteine-type endopeptidase inhibitor MF 9.0e-06 5 activity GO: ferric iron binding MF 3.3e-05 5 GO: endopeptidase inhibitor activity MF 4.5e-05 9 GO: cellular iron ion homeostasis BP 5.1e-05 5 GO: DNA binding MF GO: sulfotransferase activity MF GO: viral reproduction BP GO: structural constituent of ribosome MF GO: ribosome CC GO: ribosome biogenesis BP GO: protein catabolic process BP GO: serine-type endopeptidase inhibitor MF activity GO: metalloendopeptidase activity MF P-values were adjusted by Benjimini-Hochberg false discovery rate method 4. 64
65 Supplementary Table 37 GO enrichment analysis for expanded gene families in dromedary genome. GO ID Description Taxonomy P-value Number of genes GO: RNA-directed DNA polymerase MF activity GO: RNA-dependent DNA replication BP GO: nucleotidyltransferase activity MF GO: RNA binding MF 1.5e GO: structural constituent of ribosome MF 2.6e GO: ribosome CC 2.6e GO: cellular macromolecule biosynthetic BP 9.7e process GO: translation BP 6.1e GO: cellular biosynthetic process BP 5.7e GO: structural molecule activity MF 2.7e GO: intracellular CC 2.1e non-membrane-bounded organelle GO: cytoplasmic part CC 1.87e GO: transferase activity, transferring MF 4.63e phosphorus-containing groups GO: cellular macromolecule metabolic BP 5.7e process GO: cytoplasm CC 7.2e GO: macromolecule metabolic process BP 6.8e GO: macromolecular complex CC 7.7e GO: cellular metabolic process BP 1.5e GO: nucleic acid binding MF 4.1e GO: primary metabolic process BP 4.5e GO: transferase activity MF 2.5e GO: nucleic acid metabolic process BP 7.2e GO: cellular process BP 1.0e GO: pheromone receptor activity MF 2.3e GO: nucleobase, nucleoside, nucleotide BP 3.2e and nucleic acid metabolic process GO: metabolic process BP 1.8e GO: cellular protein metabolic process BP 3.5e GO: intracellular organelle CC 2.2e GO: gene expression BP 2.77e GO: intracellular part CC 4.37e GO: protein metabolic process BP 3.68e GO: iron ion binding MF 2.09e GO: G-protein coupled receptor activity MF 5.8e GO: ferric iron binding MF 1.6e GO: iron ion transport BP 3.7e GO: cellular iron ion homeostasis BP 3.7e GO: transmembrane receptor activity MF 1.24e GO: receptor activity MF 1.4e GO: large ribosomal subunit CC 3.6e GO: olfactory receptor activity MF 1.0e GO: cytochrome-c oxidase activity MF 5.8e GO: rrna binding MF 7.14e GO: G-protein coupled receptor protein signaling pathway BP 4.56e
66 GO: intracellular CC 1.68e GO: chromosomal part CC 2.13e GO: keratin filament CC 4.84e GO: cell surface receptor linked BP 5.57e signaling pathway GO: heme binding MF 6.58e GO: hydrogen ion transmembrane MF 3.63e transporter activity GO: chromosome, centromeric region CC 5.45e-06 9 GO: monooxygenase activity MF 1.32e GO: mitochondrion CC 2.3e GO: dutp metabolic process BP 7.8e-05 5 GO: protein folding BP 9.3e GO: chromatin CC GO: intracellular organelle part CC GO: small ribosomal subunit CC GO: translational elongation BP GO: catalytic activity MF GO: aspartic-type endopeptidase activity MF GO: MHC class I protein complex CC GO: cell part CC GO: MHC protein complex CC GO: nucleosome CC GO: antigen processing and presentation BP GO: nucleosome assembly BP GO: prefoldin complex CC GO: homophilic cell adhesion BP P-values were adjusted by Benjimini-Hochberg false discovery rate method 4. 66
67 Supplementary Table 38 GO enrichment analysis for expanded gene families in alpaca genome. GO ID Description Taxonomy P-value Number of genes GO: structural constituent of ribosome MF 3.98e GO: ribosome CC 3.98e GO: translation BP 7.89e GO: structural molecule activity MF 5.898e GO: gene expression BP 2.22e GO: cellular macromolecule BP 5.096e biosynthetic process GO: intracellular CC 5.18e non-membrane-bounded organelle GO: macromolecule metabolic process BP 7.45e GO: cellular macromolecule metabolic BP 3.2e process GO: macromolecular complex CC 2.3e GO: regulation of transcription, BP 4.3e DNA-dependent GO: transition metal ion binding MF 5.5e GO: intracellular CC 1.68e GO: ferric iron binding MF 6.3e GO: primary metabolic process BP 1.1e GO: iron ion transport BP 1.5e GO: metal ion binding MF 2.3e GO: cellular iron ion homeostasis BP 3.45e GO: protein metabolic process BP 6.49e GO: small ribosomal subunit CC 6.98e GO: cellular protein metabolic process BP 8.19e GO: zinc ion binding MF 2.56e GO: cellular process BP 4.32e GO: cell part CC 8.7e GO: intracellular organelle CC 5.55e GO: biological regulation BP 3.9e GO: gamete generation BP 5.1e-10 7 GO: homophilic cell adhesion BP 2.9e GO: nucleic acid binding MF 2.01e GO: regulation of cellular process BP 5.79e GO: cell adhesion BP 6.42e GO: ephrin receptor activity MF 3.58e-05 5 GO: olfactory receptor activity MF GO: defense response BP GO: plasma membrane CC GO: microtubule CC GO: protein polymerization BP GO: transmembrane receptor protein BP tyrosine kinase signaling pathway GO: protein catabolic process BP GO: microtubule-based movement BP GO: metalloendopeptidase activity MF P-values were adjusted by Benjimini-Hochberg false discovery rate method 4. 67
68 Supplementary Table 39 GO enrichment of unique amino acid residues changed gene in Bactrian camel genome. GO ID Description Taxonomy P-value Number of genes GO: protein phosphorylation BP 3.6e GO: protein kinase activity MF 1.18e GO: catalytic activity MF 1.62e GO: transferase activity, transferring MF 2.7e phosphorus-containing groups GO: transferase activity MF 3.15e GO: phosphotransferase activity, MF 4.8e alcohol group as acceptor GO: protein serine/threonine kinase MF 5.33e activity GO: kinase activity MF 6.3e GO: ATP binding MF GO: phosphate-containing compound BP metabolic process GO: macromolecule modification BP GO: protein modification process BP GO: vacuolar proton-transporting CC V-type ATPase, V1 domain GO: small molecule binding MF P-values were adjusted by Benjimini-Hochberg false discovery rate method 4. 68
69 Supplementary Table 40 GO enrichment of unique amino acid residues changed gene in dromedary genome. GO ID Description Taxonomy P-value Number of genes GO: peptidyl-histidine modification BP GO: protein kinase activity MF GO: protein phosphorylation BP GO: ATP binding MF GO: catalytic activity MF GO: protein serine/threonine kinase MF activity GO: phosphate-containing compound BP metabolic process GO: small molecule binding MF GO: small molecule metabolic BP process GO: phosphotransferase activity, MF alcohol group as acceptor GO: protein tyrosine kinase activity MF GO: peptidyl-amino acid modification BP GO: purine ribonucleoside MF triphosphate binding GO: nucleotide binding MF P-values were adjusted by Benjimini-Hochberg false discovery rate method 4. 69
70 Supplementary Table 41 The top 10 GO terms of the gained genes in the Bactrian camel genome. GO term GO description P-value GO: sensory perception of smell 0.00 GO: olfactory receptor activity 0.00 GO: immune response 0.00 GO: chemokine activity 2.69E-13 GO: cytokine receptor binding 4.40E-13 GO: G-protein coupled receptor protein signaling pathway 2.80E-12 GO: negative regulation of megakaryocyte differentiation 4.84E-12 GO: zinc ion binding 2.35E-11 GO: antigen processing and presentation 3.02E-11 GO: sensory perception of taste 1.07E-06 P-values were adjusted by Benjimini-Hochberg false discovery rate method 4. 70
71 Supplementary Table 42 The top 10 GO terms of the gained genes in the dromedary genome. GO term GO description P-value GO: sensory perception of smell 0.00 GO: olfactory receptor activity 0.00 GO: Sequestering of actin monomers 1.51e-05 GO: Response to pheromone 3.32e-05 GO: Pheromone receptor activity 3.32e-05 GO: Homophilic cell adhesion 3.49e-05 GO: Cytochrome-c oxidase activity 1.14E-04 GO: Antigen binding 3.00E-04 GO: Cellular response to reactive oxygen species 3.16E-04 GO: NEDD8 ligase activity 3.16E-04 P-values were adjusted by Benjimini-Hochberg false discovery rate method 4. 71
72 Supplementary Table 43 Rapidly evolving GO categories associated with fat and energy in the Bactrian camel and dromedary genomes compared with alpaca genome. GO ID Bactrian camel dn/ds Bactrian camel versus alpaca Alpaca dn/ds P-value Dromedary dn/ds Dromedary versus alpaca Alpaca dn/ds P-value GO description GO: ATP catabolic process GO: E-05 ATPase activity ATPase activity coupled to GO: E E-16 transmembrane movement of substances GO: E E-07 mitochondrion GO: E-07 mitochondrial matrix GO: E lipid transport GO: E-05 response to insulin stimulus 72
73 Supplementary Table 44 Rapidly evolving GO categories associated with fat and energy in the Bactrian camel genome compared with dromedary genome. GO ID Bactrian camel dn/ds Dromedary dn/ds GO description P-value GO: fatty acid metabolic process GO: cellular lipid metabolic process GO: triglyceride biosynthetic process GO: lipid catabolic process GO: cholesterol homeostasis 2.31E-12 GO: cholesterol metabolic process GO: glycerophospholipid biosynthetic process GO: fat cell differentiation 2.54E-09 GO: carbohydrate metabolic process 8.06E-05 GO: generation of precursor metabolites and energy 1.61E-06 GO: cellular response to insulin stimulus 4.44E-10 GO: insulin receptor signaling pathway 7.54E-05 73
74 Supplementary Table 45 Transcriptome sequenced data statistics for control group and water restriction. Total reads (M) Total base (Gb) Map reads (M) Map ratio (%) Renal cortex (CG) Renal medulla (CG) Renal cortex (WR) Renal medulla (WR)
75 Supplementary Table 46 Transcriptomic representation of genes of the aldosterone-regulated sodium reabsorption in renal cortex for control group and water restriction. Gene ID Gene Name RPKM of CG RPKM of WR Log2 Ratio (WR/CG) Up-Down-Regulation (WR/CG) P-value (WR/CG) FDR (WR/CG) Ala_bactrian_19666 ATP1A Up 0 0 Ala_bactrian_09473 ATP1A Up 2.41E E-35 Ala_bactrian_02829 ATP1A Up 8.57E E-01 Ala_bactrian_15986 ATP1B Up 0 0 Ala_bactrian_03560 ATP1B Up 1.81E E-04 Ala_bactrian_17974 ATP1B Up 4.49E E-18 Ala_bactrian_06438 ATP1B Up Ala_bactrian_10566 FXYD Down 0 0 Ala_bactrian_14226 HSD11B Up 6.75E E-01 Ala_bactrian_10747 IGF Up Ala_bactrian_04915 INSR Up 0 0 Ala_bactrian_08579 IRS Up 8.59E E-105 Ala_bactrian_15617 IRS Up 5.27E E-01 Ala_bactrian_19183 KCNJ Up 8.70E E-80 Ala_bactrian_11246 KRAS Down 4.42E E-07 Ala_bactrian_03613 MAPK Up 1.94E E-170 Ala_bactrian_14091 MAPK Up 1.31E E-28 Ala_bactrian_12234 NEDD4L Up 5.05E E-100 Ala_bactrian_02710 NR3C Up 1.52E E-25 Ala_bactrian_09741 NR3C Up 4.83E E-37 Ala_bactrian_10426 PDPK Up 7.60E E-38 Ala_bactrian_03414 PIK3CA Up 8.09E E-01 75
76 Ala_bactrian_10548 PIK3CB Up 9.41E E-68 Ala_bactrian_14406 PIK3CD Up 5.21E E-60 Ala_bactrian_00530 PIK3CG Up 3.28E E-02 Ala_bactrian_02487 PIK3R Up 1.73E E-26 Ala_bactrian_08669 PIK3R Up 9.32E E-70 Ala_bactrian_18999 PIK3R Up 6.84E E-18 Ala_bactrian_00529 PIK3R Up 1.75E E-11 Ala_bactrian_11097 PRKCA Up 2.08E E-134 Ala_bactrian_01079 PRKCB Down 8.31E E-01 Ala_bactrian_02418 PRKCG Up 7.24E E-01 Ala_bactrian_09164 SCNN1A Up 0 0 Ala_bactrian_16445 SCNN1B Up 2.12E E-34 Ala_bactrian_00164 SCNN1G Up 1.07E E-109 Ala_bactrian_15032 SFN Down 6.89E E-16 Ala_bactrian_10043 SGK Up 6.75E E-216 Ala_bactrian_13495 SLC9A3R Up 5.68E E-05 The genes showed in this table belong to the KEGG pathway (map04960). 76
77 Supplementary Table 47 Transcriptomic representation of genes of the aldosterone-regulated sodium reabsorption in renal medulla for control group and water restriction. Gene ID Gene Name RPKM of CG RPKM of WR Log2 Ratio (WR/CG) Up-Down-Regulation (WR/CG) P-value (WR/CG) FDR (WR/CG) Ala_bactrian_19666 ATP1A Up 0 0 Ala_bactrian_09473 ATP1A Down 1.87E E-11 Ala_bactrian_02829 ATP1A Up 2.66E E-01 Ala_bactrian_18032 ATP1A Up 3.53E E-02 Ala_bactrian_15986 ATP1B Up 8.26E E-85 Ala_bactrian_03560 ATP1B Down 5.16E E-01 Ala_bactrian_17974 ATP1B Up 1.81E E-06 Ala_bactrian_06438 ATP1B Down 4.80E E-01 Ala_bactrian_10566 FXYD Up 1.77E E-210 Ala_bactrian_14226 HSD11B Up 0 0 Ala_bactrian_10747 IGF Down 1.41E E-01 Ala_bactrian_04915 INSR Down 1.54E E-72 Ala_bactrian_08579 IRS Down 7.44E E-01 Ala_bactrian_15617 IRS Down 9.99E E-73 Ala_bactrian_19183 KCNJ Up 2.16E E-10 Ala_bactrian_11246 KRAS Down 3.17E E-24 Ala_bactrian_03613 MAPK Down 9.59E E-17 Ala_bactrian_14091 MAPK Up 7.05E E-56 Ala_bactrian_12234 NEDD4L Down 4.65E E-84 Ala_bactrian_02710 NR3C Down 9.71E E-14 Ala_bactrian_09741 NR3C Down 8.92E E-40 Ala_bactrian_10426 PDPK Down 1.24E E-51 77
78 Ala_bactrian_03414 PIK3CA Down 4.74E E-92 Ala_bactrian_10548 PIK3CB Down 2.30E E-76 Ala_bactrian_14406 PIK3CD Down 2.31E E-18 Ala_bactrian_00530 PIK3CG Down 5.68E E-10 Ala_bactrian_02487 PIK3R Down 9.66E E-01 Ala_bactrian_08669 PIK3R Up 6.54E E-23 Ala_bactrian_18999 PIK3R Down 1.91E E-15 Ala_bactrian_00529 PIK3R Down 9.12E E-06 Ala_bactrian_11097 PRKCA Down 2.46E E-98 Ala_bactrian_01079 PRKCB Down 1.09E E-09 Ala_bactrian_02418 PRKCG Down 3.09E E-01 Ala_bactrian_09164 SCNN1A Up 1.03E E-99 Ala_bactrian_16445 SCNN1B Up 1.71E E-113 Ala_bactrian_00164 SCNN1G Up 2.29E E-72 Ala_bactrian_15032 SFN Up 5.10E E-04 Ala_bactrian_10043 SGK Down 8.68E E-03 Ala_bactrian_13495 SLC9A3R Up 1.55E E-90 The genes showed in this table belong to the KEGG pathway (map04960). 78
79 Supplementary Table 48 Transcriptomic representation of genes of the aquaporins family in renal cortex for control group and water restriction. Gene ID Gene Name RPKM of CG RPKM of WR Log2 Ratio (WR/CG) Up-Down-Regulation (WR/CG) P-value (WR/CG) FDR (WR/CG) Ala_bactrian_10657 AQP Up 0 0 Ala_bactrian_17951 AQP Up 0 0 Ala_bactrian_00286 AQP Up E E-251 Ala_bactrian_10592 AQP Down E-02 Ala_bactrian_13653 AQP Up E E-67 Ala_bactrian_10970 AQP Down Ala_bactrian_01284 AQP Up
80 Supplementary Table 49 Transcriptomic representation of genes of the aquaporins family in renal medulla for control group and water restriction. Gene ID Gene Name RPKM of CG RPKM of WR Log2 Ratio (WR/CG) Up-Down-Regulation (WR/CG) P-value (WR/CG) FDR (WR/CG) Ala_bactrian_10657 AQP Up 0 0 Ala_bactrian_17951 AQP Up 0 0 Ala_bactrian_00286 AQP Up 0 0 Ala_bactrian_10592 AQP Up 1.53E E-04 Ala_bactrian_13653 AQP Up 2.37E E-20 Ala_bactrian_10970 AQP Down Ala_bactrian_17742 AQP Up Ala_bactrian_01284 AQP Up
Molecular Cell Biology - Problem Drill 01: Introduction to Molecular Cell Biology
 Molecular Cell Biology - Problem Drill 01: Introduction to Molecular Cell Biology Question No. 1 of 10 1. Which statement describes how an organism is organized from most simple to most complex? Question
Molecular Cell Biology - Problem Drill 01: Introduction to Molecular Cell Biology Question No. 1 of 10 1. Which statement describes how an organism is organized from most simple to most complex? Question
Supplementary Table 1. Summary of whole genome shotgun sequence used for genome assembly
 Supplementary Tables Supplementary Table 1. Summary of whole genome shotgun sequence used for genome assembly Library Read length Raw data Filtered data insert size (bp) * Total Sequence depth Total Sequence
Supplementary Tables Supplementary Table 1. Summary of whole genome shotgun sequence used for genome assembly Library Read length Raw data Filtered data insert size (bp) * Total Sequence depth Total Sequence
Supplemental Material for: A high-throughput RNA-seq approach to profile transcriptional responses
 Supplemental Material for: A high-throughput RNA-seq approach to profile transcriptional responses G. A. Moyerbrailean 1, G. O. Davis 1, C. T. Harvey 1, D. Watza 1, X. Wen 2, R. Pique-Regi 1,3,, F. Luca
Supplemental Material for: A high-throughput RNA-seq approach to profile transcriptional responses G. A. Moyerbrailean 1, G. O. Davis 1, C. T. Harvey 1, D. Watza 1, X. Wen 2, R. Pique-Regi 1,3,, F. Luca
Supplement to: The Genomic Sequence of the Chinese Hamster Ovary (CHO)-K1 cell line
 Supplement to: The Genomic Sequence of the Chinese Hamster Ovary (CHO)-K1 cell line Table of Contents SUPPLEMENTARY TEXT:... 2 FILTERING OF RAW READS PRIOR TO ASSEMBLY:... 2 COMPARATIVE ANALYSIS... 2 IMMUNOGENIC
Supplement to: The Genomic Sequence of the Chinese Hamster Ovary (CHO)-K1 cell line Table of Contents SUPPLEMENTARY TEXT:... 2 FILTERING OF RAW READS PRIOR TO ASSEMBLY:... 2 COMPARATIVE ANALYSIS... 2 IMMUNOGENIC
2012 GENERAL [5 points]
![2012 GENERAL [5 points] 2012 GENERAL [5 points]](/thumbs/75/72452386.jpg) GENERAL [5 points] 2012 Mark all processes that are part of the 'standard dogma of molecular' [ ] DNA replication [ ] transcription [ ] translation [ ] reverse transposition [ ] DNA restriction [ ] DNA
GENERAL [5 points] 2012 Mark all processes that are part of the 'standard dogma of molecular' [ ] DNA replication [ ] transcription [ ] translation [ ] reverse transposition [ ] DNA restriction [ ] DNA
Sequencing and assembly of the sheep genome reference sequence
 Sequencing and assembly of the sheep genome reference sequence Yu Jiang Kunming Institute of Zoology, CAS, China the International Sheep Genomics Consortium (ISGC) ISGC Presentations Yu Jiang, Kunming
Sequencing and assembly of the sheep genome reference sequence Yu Jiang Kunming Institute of Zoology, CAS, China the International Sheep Genomics Consortium (ISGC) ISGC Presentations Yu Jiang, Kunming
Bayesian Decomposition
 Bayesian Decomposition Michael Ochs Making Proteins A Closer Look at Translation Post-Translational Modification RNA Splicing mirna Identifying Pathways A 1 2 3 B C D A B C D Bioinformatics www.promega.com
Bayesian Decomposition Michael Ochs Making Proteins A Closer Look at Translation Post-Translational Modification RNA Splicing mirna Identifying Pathways A 1 2 3 B C D A B C D Bioinformatics www.promega.com
Chapter 6: Transcription and RNA Processing in Eukaryotes
 3. Basic Genetics Plant Molecular Biology Chapter 6: Transcription and RNA Processing in Eukaryotes - Genetic organization in eukaryote - Transcription in eukaryote - - RNA processing in eukaryote - Translation
3. Basic Genetics Plant Molecular Biology Chapter 6: Transcription and RNA Processing in Eukaryotes - Genetic organization in eukaryote - Transcription in eukaryote - - RNA processing in eukaryote - Translation
Key Area 1.3: Gene Expression
 Key Area 1.3: Gene Expression RNA There is a second type of nucleic acid in the cell, called RNA. RNA plays a vital role in the production of protein from the code in the DNA. What is gene expression?
Key Area 1.3: Gene Expression RNA There is a second type of nucleic acid in the cell, called RNA. RNA plays a vital role in the production of protein from the code in the DNA. What is gene expression?
Big Idea 3C Basic Review
 Big Idea 3C Basic Review 1. A gene is a. A sequence of DNA that codes for a protein. b. A sequence of amino acids that codes for a protein. c. A sequence of codons that code for nucleic acids. d. The end
Big Idea 3C Basic Review 1. A gene is a. A sequence of DNA that codes for a protein. b. A sequence of amino acids that codes for a protein. c. A sequence of codons that code for nucleic acids. d. The end
Identification of Cassava MicroRNAs under Abiotic Stress
 Identification of Cassava MicroRNAs under Abiotic Stress Carolina Ballén-Taborda 1, Germán lata 2, Sarah Ayling 3, Fausto Rodríguez-Zapata 1, Luis Augusto Becerra Lopez-Lavalle 1, Jorge Duitama 1 & Joe
Identification of Cassava MicroRNAs under Abiotic Stress Carolina Ballén-Taborda 1, Germán lata 2, Sarah Ayling 3, Fausto Rodríguez-Zapata 1, Luis Augusto Becerra Lopez-Lavalle 1, Jorge Duitama 1 & Joe
Videos. Lesson Overview. Fermentation
 Lesson Overview Fermentation Videos Bozeman Transcription and Translation: https://youtu.be/h3b9arupxzg Drawing transcription and translation: https://youtu.be/6yqplgnjr4q Objectives 29a) I can contrast
Lesson Overview Fermentation Videos Bozeman Transcription and Translation: https://youtu.be/h3b9arupxzg Drawing transcription and translation: https://youtu.be/6yqplgnjr4q Objectives 29a) I can contrast
CHAPTER 21 LECTURE SLIDES
 CHAPTER 21 LECTURE SLIDES Prepared by Brenda Leady University of Toledo To run the animations you must be in Slideshow View. Use the buttons on the animation to play, pause, and turn audio/text on or off.
CHAPTER 21 LECTURE SLIDES Prepared by Brenda Leady University of Toledo To run the animations you must be in Slideshow View. Use the buttons on the animation to play, pause, and turn audio/text on or off.
What is necessary for life?
 Life What is necessary for life? Most life familiar to us: Eukaryotes FREE LIVING Or Parasites First appeared ~ 1.5-2 10 9 years ago Requirements: DNA, proteins, lipids, carbohydrates, complex structure,
Life What is necessary for life? Most life familiar to us: Eukaryotes FREE LIVING Or Parasites First appeared ~ 1.5-2 10 9 years ago Requirements: DNA, proteins, lipids, carbohydrates, complex structure,
Chapter 8 Lecture Outline. Transcription, Translation, and Bioinformatics
 Chapter 8 Lecture Outline Transcription, Translation, and Bioinformatics Replication, Transcription, Translation n Repetitive processes Build polymers of nucleotides or amino acids n All have 3 major steps
Chapter 8 Lecture Outline Transcription, Translation, and Bioinformatics Replication, Transcription, Translation n Repetitive processes Build polymers of nucleotides or amino acids n All have 3 major steps
COMPUTATIONAL PREDICTION AND CHARACTERIZATION OF A TRANSCRIPTOME USING CASSAVA (MANIHOT ESCULENTA) RNA-SEQ DATA
 COMPUTATIONAL PREDICTION AND CHARACTERIZATION OF A TRANSCRIPTOME USING CASSAVA (MANIHOT ESCULENTA) RNA-SEQ DATA AOBAKWE MATSHIDISO, SCOTT HAZELHURST, CHRISSIE REY Wits Bioinformatics, University of the
COMPUTATIONAL PREDICTION AND CHARACTERIZATION OF A TRANSCRIPTOME USING CASSAVA (MANIHOT ESCULENTA) RNA-SEQ DATA AOBAKWE MATSHIDISO, SCOTT HAZELHURST, CHRISSIE REY Wits Bioinformatics, University of the
The Central Dogma. DNA makes RNA makes Proteins
 The Central Dogma DNA makes RNA makes Proteins TRANSCRIPTION DNA RNA transcript RNA polymerase RNA PROCESSING Exon RNA transcript (pre-) Intron Aminoacyl-tRNA synthetase NUCLEUS CYTOPLASM FORMATION OF
The Central Dogma DNA makes RNA makes Proteins TRANSCRIPTION DNA RNA transcript RNA polymerase RNA PROCESSING Exon RNA transcript (pre-) Intron Aminoacyl-tRNA synthetase NUCLEUS CYTOPLASM FORMATION OF
I. Gene Expression Figure 1: Central Dogma of Molecular Biology
 I. Gene Expression Figure 1: Central Dogma of Molecular Biology Central Dogma: Gene Expression: RNA Structure RNA nucleotides contain the pentose sugar Ribose instead of deoxyribose. Contain the bases
I. Gene Expression Figure 1: Central Dogma of Molecular Biology Central Dogma: Gene Expression: RNA Structure RNA nucleotides contain the pentose sugar Ribose instead of deoxyribose. Contain the bases
Molecular Cell Biology - Problem Drill 08: Transcription, Translation and the Genetic Code
 Molecular Cell Biology - Problem Drill 08: Transcription, Translation and the Genetic Code Question No. 1 of 10 1. Which of the following statements about how genes function is correct? Question #1 (A)
Molecular Cell Biology - Problem Drill 08: Transcription, Translation and the Genetic Code Question No. 1 of 10 1. Which of the following statements about how genes function is correct? Question #1 (A)
DNA makes RNA makes Proteins. The Central Dogma
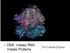 DNA makes RNA makes Proteins The Central Dogma TRANSCRIPTION DNA RNA transcript RNA polymerase RNA PROCESSING Exon RNA transcript (pre-mrna) Intron Aminoacyl-tRNA synthetase NUCLEUS CYTOPLASM FORMATION
DNA makes RNA makes Proteins The Central Dogma TRANSCRIPTION DNA RNA transcript RNA polymerase RNA PROCESSING Exon RNA transcript (pre-mrna) Intron Aminoacyl-tRNA synthetase NUCLEUS CYTOPLASM FORMATION
9/3/2009. DNA RNA Proteins. DNA Genetic program RNAs Ensure synthesis of proteins Proteins Ensure all cellular functions Carbohydrates (sugars) Energy
 Structure Properties Functions of the cell Chemical organization of the cell Based on molecular substrate : DNA contains information RNA ensures protein synthesis Proteins ensure vitality Relations between
Structure Properties Functions of the cell Chemical organization of the cell Based on molecular substrate : DNA contains information RNA ensures protein synthesis Proteins ensure vitality Relations between
Molecular characterization of Pst isolates from Western Canada
 #bgri2014 Molecular characterization of Pst isolates from Western Canada André Laroche, Michele Frick, Yong Xu, Byron Puchalski, Brent Puchalski, Harpinder Randhawa, Robert Graf and Denis Gaudet Agriculture
#bgri2014 Molecular characterization of Pst isolates from Western Canada André Laroche, Michele Frick, Yong Xu, Byron Puchalski, Brent Puchalski, Harpinder Randhawa, Robert Graf and Denis Gaudet Agriculture
RNA Expression of the information in a gene generally involves production of an RNA molecule transcribed from a DNA template. RNA differs from DNA
 RNA Expression of the information in a gene generally involves production of an RNA molecule transcribed from a DNA template. RNA differs from DNA that it has a hydroxyl group at the 2 position of the
RNA Expression of the information in a gene generally involves production of an RNA molecule transcribed from a DNA template. RNA differs from DNA that it has a hydroxyl group at the 2 position of the
8/21/2014. From Gene to Protein
 From Gene to Protein Chapter 17 Objectives Describe the contributions made by Garrod, Beadle, and Tatum to our understanding of the relationship between genes and enzymes Briefly explain how information
From Gene to Protein Chapter 17 Objectives Describe the contributions made by Garrod, Beadle, and Tatum to our understanding of the relationship between genes and enzymes Briefly explain how information
RNA ID missing Word ID missing Word DNA ID missing Word
 Table #1 Vocab Term RNA ID missing Word ID missing Word DNA ID missing Word Definition Define Base pairing rules of A=T and C=G are used for this process DNA duplicates, or makes a copy of, itself. Synthesis
Table #1 Vocab Term RNA ID missing Word ID missing Word DNA ID missing Word Definition Define Base pairing rules of A=T and C=G are used for this process DNA duplicates, or makes a copy of, itself. Synthesis
What is necessary for life?
 Life What is necessary for life? Most life familiar to us: Eukaryotes FREE LIVING Or Parasites First appeared ~ 1.5-2 10 9 years ago Requirements: DNA, proteins, lipids, carbohydrates, complex structure,
Life What is necessary for life? Most life familiar to us: Eukaryotes FREE LIVING Or Parasites First appeared ~ 1.5-2 10 9 years ago Requirements: DNA, proteins, lipids, carbohydrates, complex structure,
M1 - Biochemistry. Nucleic Acid Structure II/Transcription I
 M1 - Biochemistry Nucleic Acid Structure II/Transcription I PH Ratz, PhD (Resources: Lehninger et al., 5th ed., Chapters 8, 24 & 26) 1 Nucleic Acid Structure II/Transcription I Learning Objectives: 1.
M1 - Biochemistry Nucleic Acid Structure II/Transcription I PH Ratz, PhD (Resources: Lehninger et al., 5th ed., Chapters 8, 24 & 26) 1 Nucleic Acid Structure II/Transcription I Learning Objectives: 1.
Lesson Overview. Fermentation 13.1 RNA
 13.1 RNA The Role of RNA Genes contain coded DNA instructions that tell cells how to build proteins. The first step in decoding these genetic instructions is to copy part of the base sequence from DNA
13.1 RNA The Role of RNA Genes contain coded DNA instructions that tell cells how to build proteins. The first step in decoding these genetic instructions is to copy part of the base sequence from DNA
Videos. Bozeman Transcription and Translation: Drawing transcription and translation:
 Videos Bozeman Transcription and Translation: https://youtu.be/h3b9arupxzg Drawing transcription and translation: https://youtu.be/6yqplgnjr4q Objectives 29a) I can contrast RNA and DNA. 29b) I can explain
Videos Bozeman Transcription and Translation: https://youtu.be/h3b9arupxzg Drawing transcription and translation: https://youtu.be/6yqplgnjr4q Objectives 29a) I can contrast RNA and DNA. 29b) I can explain
Find this material useful? You can help our team to keep this site up and bring you even more content consider donating via the link on our site.
 Find this material useful? You can help our team to keep this site up and bring you even more content consider donating via the link on our site. Still having trouble understanding the material? Check
Find this material useful? You can help our team to keep this site up and bring you even more content consider donating via the link on our site. Still having trouble understanding the material? Check
Bioinformatics: Sequence Analysis. COMP 571 Luay Nakhleh, Rice University
 Bioinformatics: Sequence Analysis COMP 571 Luay Nakhleh, Rice University Course Information Instructor: Luay Nakhleh (nakhleh@rice.edu); office hours by appointment (office: DH 3119) TA: Leo Elworth (DH
Bioinformatics: Sequence Analysis COMP 571 Luay Nakhleh, Rice University Course Information Instructor: Luay Nakhleh (nakhleh@rice.edu); office hours by appointment (office: DH 3119) TA: Leo Elworth (DH
Introduction to Cellular Biology and Bioinformatics. Farzaneh Salari
 Introduction to Cellular Biology and Bioinformatics Farzaneh Salari Outline Bioinformatics Cellular Biology A Bioinformatics Problem What is bioinformatics? Computer Science Statistics Bioinformatics Mathematics...
Introduction to Cellular Biology and Bioinformatics Farzaneh Salari Outline Bioinformatics Cellular Biology A Bioinformatics Problem What is bioinformatics? Computer Science Statistics Bioinformatics Mathematics...
Genes and How They Work. Chapter 15
 Genes and How They Work Chapter 15 The Nature of Genes They proposed the one gene one enzyme hypothesis. Today we know this as the one gene one polypeptide hypothesis. 2 The Nature of Genes The central
Genes and How They Work Chapter 15 The Nature of Genes They proposed the one gene one enzyme hypothesis. Today we know this as the one gene one polypeptide hypothesis. 2 The Nature of Genes The central
Molecular Genetics. The flow of genetic information from DNA. DNA Replication. Two kinds of nucleic acids in cells: DNA and RNA.
 Molecular Genetics DNA Replication Two kinds of nucleic acids in cells: DNA and RNA. DNA function 1: DNA transmits genetic information from parents to offspring. DNA function 2: DNA controls the functions
Molecular Genetics DNA Replication Two kinds of nucleic acids in cells: DNA and RNA. DNA function 1: DNA transmits genetic information from parents to offspring. DNA function 2: DNA controls the functions
Gene Expression: Transcription, Translation, RNAs and the Genetic Code
 Lecture 28-29 Gene Expression: Transcription, Translation, RNAs and the Genetic Code Central dogma of molecular biology During transcription, the information in a DNA sequence (a gene) is copied into a
Lecture 28-29 Gene Expression: Transcription, Translation, RNAs and the Genetic Code Central dogma of molecular biology During transcription, the information in a DNA sequence (a gene) is copied into a
BIO 311C Spring Lecture 36 Wednesday 28 Apr.
 BIO 311C Spring 2010 1 Lecture 36 Wednesday 28 Apr. Synthesis of a Polypeptide Chain 5 direction of ribosome movement along the mrna 3 ribosome mrna NH 2 polypeptide chain direction of mrna movement through
BIO 311C Spring 2010 1 Lecture 36 Wednesday 28 Apr. Synthesis of a Polypeptide Chain 5 direction of ribosome movement along the mrna 3 ribosome mrna NH 2 polypeptide chain direction of mrna movement through
MOLECULAR GENETICS PROTEIN SYNTHESIS. Molecular Genetics Activity #2 page 1
 AP BIOLOGY MOLECULAR GENETICS ACTIVITY #2 NAME DATE HOUR PROTEIN SYNTHESIS Molecular Genetics Activity #2 page 1 GENETIC CODE PROTEIN SYNTHESIS OVERVIEW Molecular Genetics Activity #2 page 2 PROTEIN SYNTHESIS
AP BIOLOGY MOLECULAR GENETICS ACTIVITY #2 NAME DATE HOUR PROTEIN SYNTHESIS Molecular Genetics Activity #2 page 1 GENETIC CODE PROTEIN SYNTHESIS OVERVIEW Molecular Genetics Activity #2 page 2 PROTEIN SYNTHESIS
Review of Protein (one or more polypeptide) A polypeptide is a long chain of..
 Gene expression Review of Protein (one or more polypeptide) A polypeptide is a long chain of.. In a protein, the sequence of amino acid determines its which determines the protein s A protein with an enzymatic
Gene expression Review of Protein (one or more polypeptide) A polypeptide is a long chain of.. In a protein, the sequence of amino acid determines its which determines the protein s A protein with an enzymatic
The Major Function Of Rna Is To Carry Out The Genetic Instructions For Protein Synthesis
 The Major Function Of Rna Is To Carry Out The Genetic Instructions For Protein Synthesis For example, protein synthesis in human mitochondria relies on a genetic code that Leder and Nirenberg were able
The Major Function Of Rna Is To Carry Out The Genetic Instructions For Protein Synthesis For example, protein synthesis in human mitochondria relies on a genetic code that Leder and Nirenberg were able
Unit 1 Human cells. 1. Division and differentiation in human cells
 Unit 1 Human cells 1. Division and differentiation in human cells Stem cells Describe the process of differentiation. Explain how differentiation is brought about with reference to genes. Name the two
Unit 1 Human cells 1. Division and differentiation in human cells Stem cells Describe the process of differentiation. Explain how differentiation is brought about with reference to genes. Name the two
Protein Synthesis & Gene Expression
 DNA provides the instructions for how to build proteins Each gene dictates how to build a single protein in prokaryotes The sequence of nucleotides (AGCT) in DNA dictates the order of amino acids that
DNA provides the instructions for how to build proteins Each gene dictates how to build a single protein in prokaryotes The sequence of nucleotides (AGCT) in DNA dictates the order of amino acids that
Evidence for convergent evolution of ALU repeats in human and mouse
 Evidence for convergent evolution of ALU repeats in human and mouse Aristotelis Tsirigos - IBM Computational Genomics Group May 2 I. ALU repeats: a class of mobile elements II. Evolution of ALUs III. Genomic
Evidence for convergent evolution of ALU repeats in human and mouse Aristotelis Tsirigos - IBM Computational Genomics Group May 2 I. ALU repeats: a class of mobile elements II. Evolution of ALUs III. Genomic
Unit II Problem 3 Genetics: Summary of Basic Concepts in Molecular Biology
 Unit II Problem 3 Genetics: Summary of Basic Concepts in Molecular Biology - The central dogma (principle) of molecular biology: Information from DNA are transcribed to mrna which will be further translated
Unit II Problem 3 Genetics: Summary of Basic Concepts in Molecular Biology - The central dogma (principle) of molecular biology: Information from DNA are transcribed to mrna which will be further translated
Unit 1: DNA and the Genome. Sub-Topic (1.3) Gene Expression
 Unit 1: DNA and the Genome Sub-Topic (1.3) Gene Expression Unit 1: DNA and the Genome Sub-Topic (1.3) Gene Expression On completion of this subtopic I will be able to State the meanings of the terms genotype,
Unit 1: DNA and the Genome Sub-Topic (1.3) Gene Expression Unit 1: DNA and the Genome Sub-Topic (1.3) Gene Expression On completion of this subtopic I will be able to State the meanings of the terms genotype,
Dissecting the Human Protein-Protein Interaction Network via Phylogenetic Decomposition
 Dissecting the Human Protein-Protein Interaction Network via Phylogenetic Decomposition Cho-Yi Chen 1, Andy Ho 2, Hsin-Yuan Huang 3, Hsueh-Fen Juan 1,2,*, Hsuan-Cheng Huang 4,* 1 Genome and Systems Biology
Dissecting the Human Protein-Protein Interaction Network via Phylogenetic Decomposition Cho-Yi Chen 1, Andy Ho 2, Hsin-Yuan Huang 3, Hsueh-Fen Juan 1,2,*, Hsuan-Cheng Huang 4,* 1 Genome and Systems Biology
Nucleic acids deoxyribonucleic acid (DNA) ribonucleic acid (RNA) nucleotide
 Nucleic Acids Nucleic acids are molecules that store information for cellular growth and reproduction There are two types of nucleic acids: - deoxyribonucleic acid (DNA) and ribonucleic acid (RNA) These
Nucleic Acids Nucleic acids are molecules that store information for cellular growth and reproduction There are two types of nucleic acids: - deoxyribonucleic acid (DNA) and ribonucleic acid (RNA) These
Chapter 8: DNA and RNA
 Chapter 8: DNA and RNA Lecture Outline Enger, E. D., Ross, F. C., & Bailey, D. B. (2012). Concepts in biology (14th ed.). New York: McGraw- Hill. 1 8-1 DNA and the Importance of Proteins Proteins play
Chapter 8: DNA and RNA Lecture Outline Enger, E. D., Ross, F. C., & Bailey, D. B. (2012). Concepts in biology (14th ed.). New York: McGraw- Hill. 1 8-1 DNA and the Importance of Proteins Proteins play
Pre-Lab Questions. 1. Use the following data to construct a cladogram of the major plant groups.
 Pre-Lab Questions Name: 1. Use the following data to construct a cladogram of the major plant groups. Table 1: Characteristics of Major Plant Groups Organism Vascular Flowers Seeds Tissue Mosses 0 0 0
Pre-Lab Questions Name: 1. Use the following data to construct a cladogram of the major plant groups. Table 1: Characteristics of Major Plant Groups Organism Vascular Flowers Seeds Tissue Mosses 0 0 0
The Nature of Genes. The Nature of Genes. The Nature of Genes. The Nature of Genes. The Nature of Genes. The Genetic Code. Genes and How They Work
 Genes and How They Work Chapter 15 Early ideas to explain how genes work came from studying human diseases. Archibald Garrod studied alkaptonuria, 1902 Garrod recognized that the disease is inherited via
Genes and How They Work Chapter 15 Early ideas to explain how genes work came from studying human diseases. Archibald Garrod studied alkaptonuria, 1902 Garrod recognized that the disease is inherited via
Section C: The Control of Gene Expression
 Section C: The Control of Gene Expression 1. Each cell of a multicellular eukaryote expresses only a small fraction of its genes 2. The control of gene expression can occur at any step in the pathway from
Section C: The Control of Gene Expression 1. Each cell of a multicellular eukaryote expresses only a small fraction of its genes 2. The control of gene expression can occur at any step in the pathway from
Fig Ch 17: From Gene to Protein
 Fig. 17-1 Ch 17: From Gene to Protein Basic Principles of Transcription and Translation RNA is the intermediate between genes and the proteins for which they code Transcription is the synthesis of RNA
Fig. 17-1 Ch 17: From Gene to Protein Basic Principles of Transcription and Translation RNA is the intermediate between genes and the proteins for which they code Transcription is the synthesis of RNA
Chapter 11. Transcription. The biochemistry and molecular biology department of CMU
 Chapter 11 Transcription The biochemistry and molecular biology department of CMU Transcription The synthesis of RNA molecules using DNA strands as the templates so that the genetic information can be
Chapter 11 Transcription The biochemistry and molecular biology department of CMU Transcription The synthesis of RNA molecules using DNA strands as the templates so that the genetic information can be
Nucleic Acids. OpenStax College. 1 DNA and RNA
 OpenStax-CNX module: m44403 1 Nucleic Acids OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 4.0 By the end of this section, you will be
OpenStax-CNX module: m44403 1 Nucleic Acids OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 4.0 By the end of this section, you will be
Relationship of Gene s Types and Introns
 Chi To BME 230 Final Project Relationship of Gene s Types and Introns Abstract: The relationship in gene ontology classification and the modification of the length of introns through out the evolution
Chi To BME 230 Final Project Relationship of Gene s Types and Introns Abstract: The relationship in gene ontology classification and the modification of the length of introns through out the evolution
Transcription Eukaryotic Cells
 Transcription Eukaryotic Cells Packet #20 1 Introduction Transcription is the process in which genetic information, stored in a strand of DNA (gene), is copied into a strand of RNA. Protein-encoding genes
Transcription Eukaryotic Cells Packet #20 1 Introduction Transcription is the process in which genetic information, stored in a strand of DNA (gene), is copied into a strand of RNA. Protein-encoding genes
NUCLEIC ACID METABOLISM. Omidiwura, B.R.O
 NUCLEIC ACID METABOLISM Omidiwura, B.R.O Nucleic Acids Nucleic acids are molecules that store information for cellular growth and reproduction There are two types of nucleic acids: - deoxyribonucleic acid
NUCLEIC ACID METABOLISM Omidiwura, B.R.O Nucleic Acids Nucleic acids are molecules that store information for cellular growth and reproduction There are two types of nucleic acids: - deoxyribonucleic acid
Overview of Human Genetics
 Overview of Human Genetics 1 Structure and function of nucleic acids. 2 Structure and composition of the human genome. 3 Mendelian genetics. Lander et al. (Nature, 2001) MAT 394 (ASU) Human Genetics Spring
Overview of Human Genetics 1 Structure and function of nucleic acids. 2 Structure and composition of the human genome. 3 Mendelian genetics. Lander et al. (Nature, 2001) MAT 394 (ASU) Human Genetics Spring
Sequencing the genomes of Nicotiana sylvestris and Nicotiana tomentosiformis Nicolas Sierro
 Sequencing the genomes of Nicotiana sylvestris and Nicotiana tomentosiformis Nicolas Sierro Philip Morris International R&D, Philip Morris Products S.A., Neuchatel, Switzerland Introduction Nicotiana sylvestris
Sequencing the genomes of Nicotiana sylvestris and Nicotiana tomentosiformis Nicolas Sierro Philip Morris International R&D, Philip Morris Products S.A., Neuchatel, Switzerland Introduction Nicotiana sylvestris
Chapter 17. From Gene to Protein. Slide 1. Slide 2. Slide 3. Gene Expression. Which of the following is the best example of gene expression? Why?
 Slide 1 Chapter 17 From Gene to Protein PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
Slide 1 Chapter 17 From Gene to Protein PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
How does the human genome stack up? Genomic Size. Genome Size. Number of Genes. Eukaryotic genomes are generally larger.
 How does the human genome stack up? Organism Human (Homo sapiens) Laboratory mouse (M. musculus) Mustard weed (A. thaliana) Roundworm (C. elegans) Fruit fly (D. melanogaster) Yeast (S. cerevisiae) Bacterium
How does the human genome stack up? Organism Human (Homo sapiens) Laboratory mouse (M. musculus) Mustard weed (A. thaliana) Roundworm (C. elegans) Fruit fly (D. melanogaster) Yeast (S. cerevisiae) Bacterium
Chapter 2. An Introduction to Genes and Genomes
 PowerPoint Lectures for Introduction to Biotechnology, Second Edition William J.Thieman and Michael A.Palladino Chapter 2 An Introduction to Genes and Genomes Lectures by Lara Dowland Chapter Contents
PowerPoint Lectures for Introduction to Biotechnology, Second Edition William J.Thieman and Michael A.Palladino Chapter 2 An Introduction to Genes and Genomes Lectures by Lara Dowland Chapter Contents
Multiple choice questions (numbers in brackets indicate the number of correct answers)
 1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
Human Biology 175 Lecture Notes: Cells. Section 1 Introduction
 Human Biology 175 Lecture Notes: Cells Section 1 Introduction A) 1) All organisms are composed of at least one cell. a) Unicellular example: b) Multicellular example(s): 2) All cells are similar in their
Human Biology 175 Lecture Notes: Cells Section 1 Introduction A) 1) All organisms are composed of at least one cell. a) Unicellular example: b) Multicellular example(s): 2) All cells are similar in their
Fermentation. Lesson Overview. Lesson Overview 13.1 RNA
 13.1 RNA THINK ABOUT IT DNA is the genetic material of cells. The sequence of nucleotide bases in the strands of DNA carries some sort of code. In order for that code to work, the cell must be able to
13.1 RNA THINK ABOUT IT DNA is the genetic material of cells. The sequence of nucleotide bases in the strands of DNA carries some sort of code. In order for that code to work, the cell must be able to
Text Reference, Campbell v.8, chapter 17 PROTEIN SYNTHESIS
 AP BIOLOGY Text Reference, Campbell v.8, chapter 17 ACTIVITY 1.22 NAME DATE HOUR PROTEIN SYNTHESIS GENETIC CODE PROTEIN SYNTHESIS OVERVIEW PROTEIN SYNTHESIS TRANSCRIPTION PROTEIN SYNTHESIS TRANSLATION
AP BIOLOGY Text Reference, Campbell v.8, chapter 17 ACTIVITY 1.22 NAME DATE HOUR PROTEIN SYNTHESIS GENETIC CODE PROTEIN SYNTHESIS OVERVIEW PROTEIN SYNTHESIS TRANSCRIPTION PROTEIN SYNTHESIS TRANSLATION
Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions. Supplementary Material
 Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions Joshua N. Burton 1, Andrew Adey 1, Rupali P. Patwardhan 1, Ruolan Qiu 1, Jacob O. Kitzman 1, Jay Shendure 1 1 Department
Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions Joshua N. Burton 1, Andrew Adey 1, Rupali P. Patwardhan 1, Ruolan Qiu 1, Jacob O. Kitzman 1, Jay Shendure 1 1 Department
Chapter 12: Molecular Biology of the Gene
 Biology Textbook Notes Chapter 12: Molecular Biology of the Gene p. 214-219 The Genetic Material (12.1) - Genetic Material must: 1. Be able to store information that pertains to the development, structure,
Biology Textbook Notes Chapter 12: Molecular Biology of the Gene p. 214-219 The Genetic Material (12.1) - Genetic Material must: 1. Be able to store information that pertains to the development, structure,
Supplementary Figure 1. The tree of Chinese jujube and its growing environment. The jujube has a very long lifecycle, even more than 1000 productive
 Supplementary Figure 1. The tree of Chinese jujube and its growing environment. The jujube has a very long lifecycle, even more than 1000 productive years. It is well adapted to drought and salinity. Supplementary
Supplementary Figure 1. The tree of Chinese jujube and its growing environment. The jujube has a very long lifecycle, even more than 1000 productive years. It is well adapted to drought and salinity. Supplementary
Self-test Quiz for Chapter 12 (From DNA to Protein: Genotype to Phenotype)
 Self-test Quiz for Chapter 12 (From DNA to Protein: Genotype to Phenotype) Question#1: One-Gene, One-Polypeptide The figure below shows the results of feeding trials with one auxotroph strain of Neurospora
Self-test Quiz for Chapter 12 (From DNA to Protein: Genotype to Phenotype) Question#1: One-Gene, One-Polypeptide The figure below shows the results of feeding trials with one auxotroph strain of Neurospora
FUNCTIONAL BIOINFORMATICS
 Molecular Biology-2018 1 FUNCTIONAL BIOINFORMATICS PREDICTING THE FUNCTION OF AN UNKNOWN PROTEIN Suppose you have found the amino acid sequence of an unknown protein and wish to find its potential function.
Molecular Biology-2018 1 FUNCTIONAL BIOINFORMATICS PREDICTING THE FUNCTION OF AN UNKNOWN PROTEIN Suppose you have found the amino acid sequence of an unknown protein and wish to find its potential function.
The Nature of Genes. The Nature of Genes. Genes and How They Work. Chapter 15/16
 Genes and How They Work Chapter 15/16 The Nature of Genes Beadle and Tatum proposed the one gene one enzyme hypothesis. Today we know this as the one gene one polypeptide hypothesis. 2 The Nature of Genes
Genes and How They Work Chapter 15/16 The Nature of Genes Beadle and Tatum proposed the one gene one enzyme hypothesis. Today we know this as the one gene one polypeptide hypothesis. 2 The Nature of Genes
Genetics Biology 331 Exam 3B Spring 2015
 Genetics Biology 331 Exam 3B Spring 2015 MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1) DNA methylation may be a significant mode of genetic regulation
Genetics Biology 331 Exam 3B Spring 2015 MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1) DNA methylation may be a significant mode of genetic regulation
From DNA to Protein: Genotype to Phenotype
 12 From DNA to Protein: Genotype to Phenotype 12.1 What Is the Evidence that Genes Code for Proteins? The gene-enzyme relationship is one-gene, one-polypeptide relationship. Example: In hemoglobin, each
12 From DNA to Protein: Genotype to Phenotype 12.1 What Is the Evidence that Genes Code for Proteins? The gene-enzyme relationship is one-gene, one-polypeptide relationship. Example: In hemoglobin, each
Quick Review of Protein Synthesis
 Collin College BIOL. 2401 Quick Review of Protein Synthesis. Proteins and Protein Synthesis Proteins are the molecular units that do most of the work in a cell. They function as molecular catalysts, help
Collin College BIOL. 2401 Quick Review of Protein Synthesis. Proteins and Protein Synthesis Proteins are the molecular units that do most of the work in a cell. They function as molecular catalysts, help
RNA-SEQUENCING ANALYSIS
 RNA-SEQUENCING ANALYSIS Joseph Powell SISG- 2018 CONTENTS Introduction to RNA sequencing Data structure Analyses Transcript counting Alternative splicing Allele specific expression Discovery APPLICATIONS
RNA-SEQUENCING ANALYSIS Joseph Powell SISG- 2018 CONTENTS Introduction to RNA sequencing Data structure Analyses Transcript counting Alternative splicing Allele specific expression Discovery APPLICATIONS
Feedback D. Incorrect! No, although this is a correct characteristic of RNA, this is not the best response to the questions.
 Biochemistry - Problem Drill 23: RNA No. 1 of 10 1. Which of the following statements best describes the structural highlights of RNA? (A) RNA can be single or double stranded. (B) G-C pairs have 3 hydrogen
Biochemistry - Problem Drill 23: RNA No. 1 of 10 1. Which of the following statements best describes the structural highlights of RNA? (A) RNA can be single or double stranded. (B) G-C pairs have 3 hydrogen
Section 10.3 Outline 10.3 How Is the Base Sequence of a Messenger RNA Molecule Translated into Protein?
 Section 10.3 Outline 10.3 How Is the Base Sequence of a Messenger RNA Molecule Translated into Protein? Messenger RNA Carries Information for Protein Synthesis from the DNA to Ribosomes Ribosomes Consist
Section 10.3 Outline 10.3 How Is the Base Sequence of a Messenger RNA Molecule Translated into Protein? Messenger RNA Carries Information for Protein Synthesis from the DNA to Ribosomes Ribosomes Consist
From DNA to Protein: Genotype to Phenotype
 12 From DNA to Protein: Genotype to Phenotype 12.1 What Is the Evidence that Genes Code for Proteins? The gene-enzyme relationship is one-gene, one-polypeptide relationship. Example: In hemoglobin, each
12 From DNA to Protein: Genotype to Phenotype 12.1 What Is the Evidence that Genes Code for Proteins? The gene-enzyme relationship is one-gene, one-polypeptide relationship. Example: In hemoglobin, each
Prokaryotic Transcription
 Prokaryotic Transcription Transcription Basics DNA is the genetic material Nucleic acid Capable of self-replication and synthesis of RNA RNA is the middle man Nucleic acid Structure and base sequence are
Prokaryotic Transcription Transcription Basics DNA is the genetic material Nucleic acid Capable of self-replication and synthesis of RNA RNA is the middle man Nucleic acid Structure and base sequence are
1. ADHERE AND DEFEND: Our bacterium has entered the host. Now it needs to adhere and get past the normal microbiota.
 North Seattle College Stage 02 Colonization and Infection This explanatory model will tell the story of how one bacterium adheres to a host and, through binary fission, ends up making two daughter cells.
North Seattle College Stage 02 Colonization and Infection This explanatory model will tell the story of how one bacterium adheres to a host and, through binary fission, ends up making two daughter cells.
GENETICS - CLUTCH CH.15 GENOMES AND GENOMICS.
 !! www.clutchprep.com CONCEPT: OVERVIEW OF GENOMICS Genomics is the study of genomes in their entirety Bioinformatics is the analysis of the information content of genomes - Genes, regulatory sequences,
!! www.clutchprep.com CONCEPT: OVERVIEW OF GENOMICS Genomics is the study of genomes in their entirety Bioinformatics is the analysis of the information content of genomes - Genes, regulatory sequences,
Molecular Genetics Student Objectives
 Molecular Genetics Student Objectives Exam 1: Enduring understanding 3.A: Heritable information provides for continuity of life. Essential knowledge 3.A.1: DNA, and in some cases RNA, is the primary source
Molecular Genetics Student Objectives Exam 1: Enduring understanding 3.A: Heritable information provides for continuity of life. Essential knowledge 3.A.1: DNA, and in some cases RNA, is the primary source
Ch Molecular Biology of the Gene
 Ch. 12 - Molecular Biology of the Gene AP BIOLOGY CHAPTER GUIDE 1. In the middle of the unraveling the mysteries of DNA, researchers knew that genetic material must be able to. It must be stable so it
Ch. 12 - Molecular Biology of the Gene AP BIOLOGY CHAPTER GUIDE 1. In the middle of the unraveling the mysteries of DNA, researchers knew that genetic material must be able to. It must be stable so it
Chimp Sequence Annotation: Region 2_3
 Chimp Sequence Annotation: Region 2_3 Jeff Howenstein March 30, 2007 BIO434W Genomics 1 Introduction We received region 2_3 of the ChimpChunk sequence, and the first step we performed was to run RepeatMasker
Chimp Sequence Annotation: Region 2_3 Jeff Howenstein March 30, 2007 BIO434W Genomics 1 Introduction We received region 2_3 of the ChimpChunk sequence, and the first step we performed was to run RepeatMasker
RNA : functional role
 RNA : functional role Hamad Yaseen, PhD MLS Department, FAHS Hamad.ali@hsc.edu.kw RNA mrna rrna trna 1 From DNA to Protein -Outline- From DNA to RNA From RNA to Protein From DNA to RNA Transcription: Copying
RNA : functional role Hamad Yaseen, PhD MLS Department, FAHS Hamad.ali@hsc.edu.kw RNA mrna rrna trna 1 From DNA to Protein -Outline- From DNA to RNA From RNA to Protein From DNA to RNA Transcription: Copying
Translation BIT 220 Chapter 13
 Translation BIT 220 Chapter 13 Making protein from mrna Most genes encode for proteins -some make RNA as end product Proteins -Monomer Amino Acid 20 amino acids -peptides -polypeptides -Structure of Amino
Translation BIT 220 Chapter 13 Making protein from mrna Most genes encode for proteins -some make RNA as end product Proteins -Monomer Amino Acid 20 amino acids -peptides -polypeptides -Structure of Amino
36. The double bonds in naturally-occuring fatty acids are usually isomers. A. cis B. trans C. both cis and trans D. D- E. L-
 36. The double bonds in naturally-occuring fatty acids are usually isomers. A. cis B. trans C. both cis and trans D. D- E. L- 37. The essential fatty acids are A. palmitic acid B. linoleic acid C. linolenic
36. The double bonds in naturally-occuring fatty acids are usually isomers. A. cis B. trans C. both cis and trans D. D- E. L- 37. The essential fatty acids are A. palmitic acid B. linoleic acid C. linolenic
1/24/2012. Cell. Plasma Membrane
 Chapter 3 Outline Plasma Membrane Cytoplasm and Its Organelles Cell and Gene Expression Protein Synthesis and Secretion DNA Synthesis and Cell Division Cell Basic unit of structure and function in body
Chapter 3 Outline Plasma Membrane Cytoplasm and Its Organelles Cell and Gene Expression Protein Synthesis and Secretion DNA Synthesis and Cell Division Cell Basic unit of structure and function in body
Higher Human Biology Unit 1: Human Cells Pupils Learning Outcomes
 Higher Human Biology Unit 1: Human Cells Pupils Learning Outcomes 1.1 Division and Differentiation in Human Cells I can state that cellular differentiation is the process by which a cell develops more
Higher Human Biology Unit 1: Human Cells Pupils Learning Outcomes 1.1 Division and Differentiation in Human Cells I can state that cellular differentiation is the process by which a cell develops more
DNA REPLICATION. DNA structure. Semiconservative replication. DNA structure. Origin of replication. Replication bubbles and forks.
 DNA REPLICATION 5 4 Phosphate 3 DNA structure Nitrogenous base 1 Deoxyribose 2 Nucleotide DNA strand = DNA polynucleotide 2004 Biology Olympiad Preparation Program 2 2004 Biology Olympiad Preparation Program
DNA REPLICATION 5 4 Phosphate 3 DNA structure Nitrogenous base 1 Deoxyribose 2 Nucleotide DNA strand = DNA polynucleotide 2004 Biology Olympiad Preparation Program 2 2004 Biology Olympiad Preparation Program
9/19/13. cdna libraries, EST clusters, gene prediction and functional annotation. Biosciences 741: Genomics Fall, 2013 Week 3
 cdna libraries, EST clusters, gene prediction and functional annotation Biosciences 741: Genomics Fall, 2013 Week 3 1 2 3 4 5 6 Figure 2.14 Relationship between gene structure, cdna, and EST sequences
cdna libraries, EST clusters, gene prediction and functional annotation Biosciences 741: Genomics Fall, 2013 Week 3 1 2 3 4 5 6 Figure 2.14 Relationship between gene structure, cdna, and EST sequences
DNA. Essential Question: How does the structure of the DNA molecule allow it to carry information?
 DNA Essential Question: How does the structure of the DNA molecule allow it to carry information? Fun Website to Explore! http://learn.genetics.utah.edu/content/molecules/ DNA History Griffith Experimented
DNA Essential Question: How does the structure of the DNA molecule allow it to carry information? Fun Website to Explore! http://learn.genetics.utah.edu/content/molecules/ DNA History Griffith Experimented
From Gene to Protein transcription, messenger RNA (mrna) translation, RNA processing triplet code, template strand, codons,
 From Gene to Protein I. Transcription and translation are the two main processes linking gene to protein. A. RNA is chemically similar to DNA, except that it contains ribose as its sugar and substitutes
From Gene to Protein I. Transcription and translation are the two main processes linking gene to protein. A. RNA is chemically similar to DNA, except that it contains ribose as its sugar and substitutes
Chapter 17. From Gene to Protein
 Chapter 17 From Gene to Protein Overview: The Flow of Genetic Information The information content of DNA is in the form of specific sequences of nucleotides The DNA inherited by an organism leads to specific
Chapter 17 From Gene to Protein Overview: The Flow of Genetic Information The information content of DNA is in the form of specific sequences of nucleotides The DNA inherited by an organism leads to specific
BIOCHEMISTRY REVIEW. Overview of Biomolecules. Chapter 12 Transcription
 BIOCHEMISTRY REVIEW Overview of Biomolecules Chapter 12 Transcription 2 3 4 5 Are You Getting It?? Which are general characteristics of transcription? (multiple answers) a) An entire DNA molecule is transcribed
BIOCHEMISTRY REVIEW Overview of Biomolecules Chapter 12 Transcription 2 3 4 5 Are You Getting It?? Which are general characteristics of transcription? (multiple answers) a) An entire DNA molecule is transcribed
BEADLE & TATUM EXPERIMENT
 FROM DNA TO PROTEINS: gene expression Chapter 14 LECTURE OBJECTIVES What Is the Evidence that Genes Code for Proteins? How Does Information Flow from Genes to Proteins? How Is the Information Content in
FROM DNA TO PROTEINS: gene expression Chapter 14 LECTURE OBJECTIVES What Is the Evidence that Genes Code for Proteins? How Does Information Flow from Genes to Proteins? How Is the Information Content in
1.5 Nucleic Acids and Their Functions Page 1 S. Preston 1
 AS Unit 1: Basic Biochemistry and Cell Organisation Name: Date: Topic 1.5 Nucleic Acids and their functions Page 1 From the syllabus: 1.5 Nucleic Acids and Their Functions Page 1 S. Preston 1 l. Nucleic
AS Unit 1: Basic Biochemistry and Cell Organisation Name: Date: Topic 1.5 Nucleic Acids and their functions Page 1 From the syllabus: 1.5 Nucleic Acids and Their Functions Page 1 S. Preston 1 l. Nucleic
CH 17 :From Gene to Protein
 CH 17 :From Gene to Protein Defining a gene gene gene Defining a gene is problematic because one gene can code for several protein products, some genes code only for RNA, two genes can overlap, and there
CH 17 :From Gene to Protein Defining a gene gene gene Defining a gene is problematic because one gene can code for several protein products, some genes code only for RNA, two genes can overlap, and there
Figure A summary of spontaneous alterations likely to require DNA repair.
 DNA Damage Figure 5-46. A summary of spontaneous alterations likely to require DNA repair. The sites on each nucleotide that are known to be modified by spontaneous oxidative damage (red arrows), hydrolytic
DNA Damage Figure 5-46. A summary of spontaneous alterations likely to require DNA repair. The sites on each nucleotide that are known to be modified by spontaneous oxidative damage (red arrows), hydrolytic
Cufflinks Scripture Both. GENCODE/ UCSC/ RefSeq 0.18% 0.92% Scripture
 Cabili_SuppFig_1 a 1.9.8.7.6.5.4.3.2.1 Partially compatible % Partially recovered RefSeq coding Fully compatible % Fully recovered RefSeq coding Cufflinks Scripture Both b 18.5% GENCODE/ UCSC/ RefSeq Cufflinks
Cabili_SuppFig_1 a 1.9.8.7.6.5.4.3.2.1 Partially compatible % Partially recovered RefSeq coding Fully compatible % Fully recovered RefSeq coding Cufflinks Scripture Both b 18.5% GENCODE/ UCSC/ RefSeq Cufflinks
