Description of Changes and Corrections for PDB File Format Version 4.0. Provisional Document April 12, 2011
|
|
|
- Clifford Ramsey
- 6 years ago
- Views:
Transcription
1 Description of Changes and Corrections for PDB File Format Version 4.0 Provisional Document April 12, 2011 The wwpdb has reviewed the PDB archive and created a new set of corrected files that will be released in June This document describes the review and resulting changes and corrections. Version 4.0 PDB entries will follow the PDB Exchange Dictionary v.4.0 (
2 TABLE OF CONTENTS 1. Biological assemblies 2. Residual B factors 3. Peptide inhibitors/antibiotics 4. X-ray entries in nonstandard crystal frame 5. Entries with incomplete coordinate sets 6. Polymers containing nonstandard polymer linkages 7. Hybrid X-ray neutron diffraction structures 8. Partial occupancy 2
3 1. Biological assemblies 6126 entries were identified with missing or incomplete computational annotation of biological assemblies. Missing computational assembly data was obtained from curated PQS results as well as PISA where these were available. Coordinate sets for predicted biological assemblies were created for each corrected entry. Molecular images of the assemblies were reviewed for the predicted assemblies. Improbable assemblies were flagged from the visual inspection and removed from the entries. Biological assembly predictions were updated in 5837 entries entries were updated with both curated PQS and PISA results. 178 entries were updated with PISA results only and 78 entries were updated with PQS results only. Assemblies could not be predicted for 47 entries. 3
4 2. Residual B factors The ATOM records in PDB format files produced by the REFMAC program where TLS refinement was used in many cases contain residual rather than full isotropic B-values. The type of anisotropic displacement parameters (ADPs) deposited in entries obtained using TLS refinement with the REFMAC program was verified entries refined using REFMAC with TLS were evaluated. Each entry in this set was first assumed to contain only residual B-values for each atom site. Using these values, an attempt was made to back-calculate a new set of full B-values using the TLSANL program. A single round of refinement was performed using these back-calculated B-values. The refinement statistics (i.e. R-values) were compared with the depositor-reported statistics. Closer reproduction of reported results using back-calculated full B-values was interpreted to mean that the entry was likely to contain residual B-values. Detailed statistical analyses were also performed on the distribution of backcalculated refinement statistics and the differences in predicted and calculated values. These analyses did not reveal evidence of additional factors aside from TLS to account for the differences in reported refinement statistics, thus supporting the assignment procedure employed. In some cases, adjustments were made to residue ranges defining TLS groups to correct problems introduced by relabeling residues or polymer chains during PDB data processing and annotation. A factor contributing to the problems is the ambiguous manner in which REFMAC interprets overlapping residue ranges. This issued was verified with the program developer, which then allowed to properly represent TLS groups as sets of non-overlapping residue ranges. Entries have been flagged with a new mmcif data item and additional text in REMARK 3 identifying the deposited B-value content as LIKELY RESIDUAL. 4
5 Example: mmcif/pdbx format: _refine.pdbx_tls_residual_adp_flag LIKELY RESIDUAL PDB format: REMARK 3 B VALUES. REMARK 3 B VALUE TYPE : LIKELY RESIDUAL 7310 entries were found to contain residual B-values using the back-calculation procedure. Based on the back-calculation procedure and other information in the deposited entries, 212 entries were found to contain full B-values. No changes were made in these entries. The procedure could not be applied to 120 entries. These entries have been labeled as UNVERIFIED. In 1350 entries some correction was also made to the residue ranges defining TLS groups. Most of the entries were identified as likely residual after correction was made to the TLS ranges. A total of 7464 unique entries were corrected for TLS range and/or tagged as likely residual or as having unverified B-values. B value issueunverified: 120 TLS range issue: 1350 B value issuelikely residual: 7310 Total TLS range issue 1350 B value issue- likely residual 7310 B value issue- unverified 120 Total unique entries with TLS range 7464 and B value issues Full B values 212 Total unique entries reviewed 7676 Table 1: Entries involved in the B-value and TLS remediation. 5
6 3. Peptide inhibitors/antibiotics Small peptide inhibitors and antibiotic molecules have not been uniformly represented throughout the PDB archive. These molecules have been variously described as short polymers, as single molecules, and in some cases as a collection of molecules connected by LINK records. The detailed chemical annotation of these molecules has also been inconsistent. The representation of these molecules has been standardized according to the following scheme. Oligopeptides that are natural products or small synthetic oligopeptides containing predominately standard amino acids and standard peptide linkages have been represented as polymers. Other small oligopeptide molecules containing substantially modified amino acids or non-standard peptide linkages have been represented as single molecules. Sequences of natural products were matched to available sequence databases (UniProtKB for gene products and Norine for non-ribosomal products). Structures of evolutionarily related families of antibiotics were represented consistently. Consistent and informative keywords were added to the files to facilitate their identification. The sequence composition of the synthetic molecules was also checked and updated to match the literature. In some cases, the original authors had to be consulted to resolve discrepancies entries containing small polypeptide and antibiotic molecules have been reviewed. In 459 entries the small oligopeptide molecule is now represented as a single molecule, and in 570 entries the peptide-containing molecule has been represented as a polymer. Entries in which the molecular representation has been changed contain additional REMARK records documenting the nomenclature changes. The polymer sequence information for the consolidated single molecules has been preserved in the wwpdb Chemical Component Dictionary ( and in REMARK 630 of the PDB file format. A new reference file analogous to the Chemical Component Dictionary has been created to describe the chemical structure and biological functions of the oligopeptide molecules. This new reference file also contains the detailed documentation that has been used to standardize the representation and nomenclature of these molecules and to link their chemical description to other chemical and biological data resources. 6
7 4. X-ray entries in nonstandard crystal frame A small number of non-virus containing X-ray entries have been deposited in a non-standard coordinate orientation. For these entries, the transformation between Cartesian and fractional coordinates cannot be computed from the crystallographic cell constants alone. Non-virus X-ray entries with coordinates not oriented in the standard crystallographic frame have been flagged with a data item and REMARK as shown in the following example: mmcif/pdbx format: _pdbx_database_remark.id 285 _pdbx_database_remark.text ;THE ENTRY COORDINATES ARE NOT PRESENTED IN THE STANDARD CRYSTAL FRAME. ; PDB file format: REMARK 285 REMARK 285 THE ENTRY COORDINATES REMARK 285 ARE NOT PRESENTED IN THE STANDARD CRYSTAL FRAME. 148 non-virus X-ray entries were identified as deposited in non-standard orientation and these were flagged with the new PDB file format REMARK. 7
8 5. Entries with incomplete coordinate sets There are 3 entries in which the deposited coordinates are incomplete, containing only one residue or a few atoms, which have to be combined with another entry to generate the complete model. The missing coordinates corresponding to these entries have been located in other PDB entries. The coordinate data for these entries has been combined to fill in the missing portions of the incomplete entries. The coordinates in these 3 entries are now complete. These entries were marked with REVDAT indicating a change in ATOM and added a detailed description in REMARK 3. The three entries and their parent entries are: 1GRH - parent 1GRG 1TN1 - parent 1TN2 9LYZ - parent 6LYZ 8
9 6. Polymers containing nonstandard polymer linkages Previously, an effort was made to identify and merge duplicate residues (i.e., residues with identical chemistry and different 3-letter codes). In this process, a number of amino acids that appeared to be redundant but were in fact residues involved in a non-standard linkage, were accidentally merged with the corresponding standard residues. For example, the sequence and connectivity for the current PDB entry 1AT6 is shown below. In this entry, the highlighted residue ASP is linked through a sidechain peptide bond rather than through the main chain. This non-standard link shown in Figure 1 was not reflected in the entry. In the original, pre-remediation entry, this ASP residue was properly represented by the non-standard aminoacid residue, IAS. SEQRES LYS VAL PHE GLY ARG CYS GLU LEU ALA ALA ALA MET LYS SEQRES ARG HIS GLY LEU ASP ASN TYR ARG GLY TYR SER LEU GLY SEQRES ASN TRP VAL CYS ALA ALA LYS PHE GLU SER ASN PHE ASN SEQRES THR GLN ALA THR ASN ARG ASN THR ASP GLY SER THR ASP SEQRES TYR GLY ILE LEU GLN ILE ASN SER ARG TRP TRP CYS ASN SEQRES ASP GLY ARG THR PRO GLY SER ARG ASN LEU CYS ASN ILE SEQRES PRO CYS SER ALA LEU LEU SER SER ASP ILE THR ALA SER SEQRES VAL ASN CYS ALA LYS LYS ILE VAL SER ASP GLY ASN GLY SEQRES MET ASN ALA TRP VAL ALA TRP ARG ASN ARG CYS LYS GLY SEQRES THR ASP VAL GLN ALA TRP ILE ARG GLY CYS ARG LEU Figure 1. In entry 1AT6, the peptide bond between ASP and GLY involves the side-chain carboxylate group of the ASP instead of the main-chain carboxylate. This bond is circled in blue in the 3D diagram. The ASP residue is highlighted in yellow in the PDB SEQRES records above. There are 58 entries where such non-standard linkage information was removed. 9
10 Residue Change Entries with nonstandard linkages ASP was IAS 1at6 1c9p 1dlg 1DY5 1ejc 1ejd 1eyn 1jg3 1lsq 1q3g 1rtu 1ryw 1ybg 2ftm 2jv0 2ftl 2fi5 2fi4 2z2c 3ahs 3iss 3kqa 3kqj 3kr6 3lth DGL was FGA 148l 1ay3 1fjm 1p4n 1wco 2nym 2nyl 2ie3 2eax 2iae 2npp 3d2y 3d2z 3dw8 3gkr 3itb ILG was GGL 1aqv 1aqw 1aqx 1eva 1evb 1evc 1evd 1gac 1lcm 2gsr Restore the original definitions of the non-standard chemical components and add more detailed linkage information to their chemical component definitions in item chem_comp.type. The list of linking types includes: chem_comp.type L-gamma-peptide, C-delta linking D-gamma-peptide, C-delta linking L-beta-peptide, C-gamma linking D-beta-peptide, C-gamma linking Description Iso-peptide linking L-gamma peptide Iso-peptide linking D-gamma peptide Iso-peptide linking L-beta peptide Iso-peptide linking D-beta peptide The residues that have been added thus far are: IAS, FGA, GGL, and ACB. Others are being identified. The advantage of this approach is that the PDB sequence explicitly reflects nonstandard behavior, and this is reflected in the PDB SEQRES, SEQADV, and MODRES records. Collectively these records inform the PDB user (and software) that there is a special situation at this residue. Because this information is explicit in the sequence, it also informs users of PDB FASTA sequence data files. The sequences in the 58 PDB entries have been updated with the appropriate restored residues containing the detailed linking feature. SEQADV, MODRES records have been similarly updated. 10
11 7. Hybrid X-ray/neutron diffraction structures There are 54 released entries that were solved using more than one experimental method. Although the experimental method and data-collection details have been captured for both methods, the relationship between the method and the data collection details is not properly represented in the PDBx dictionary. The PDBx exchange dictionary has been extended to include the additional data items to handle hybrid X-ray/neutron diffraction methods. The key identifier for the diffraction dataset (i.e., diffrn_id and pdbx_diffrn_id) has been added to categories where data collection statistics and refinement statistics are stored (i.e. REFLNS and REFINE). The PDBx mmcif files in the PDB archive have been updated for these categories. 11
12 8. Partial occupancy In the 2009 remediation, occupancies were corrected in 490 X-ray and neutron entries. A mistake was made in 104 of these entries: for atoms with alternate conformer labels and with summed total occupancy less than 1.0, the occupancies were re-scaled as 1.0/n, where n is the number of conformers. The originally deposited occupancies of the affected atoms were restored and the remediation was then carried out properly, via: Atoms with multiple conformations but identical coordinates and B-values were merged and their occupancies were summed. Atoms which now have (total) occupancies <= 1.0 were left as deposited. Atoms with (total) occupancies > 1 were rescaled proportionally to a sum of 1.0 The occupancies have been corrected in these entries. 12
In silico measurements of twist and bend. moduli for beta solenoid protein self-
 In silico measurements of twist and bend moduli for beta solenoid protein self- assembly units Leonard P. Heinz, Krishnakumar M. Ravikumar, and Daniel L. Cox Department of Physics and Institute for Complex
In silico measurements of twist and bend moduli for beta solenoid protein self- assembly units Leonard P. Heinz, Krishnakumar M. Ravikumar, and Daniel L. Cox Department of Physics and Institute for Complex
Dr. R. Sankar, BSE 631 (2018)
 Pauling, Corey and Branson Diffraction of DNA http://www.nature.com/scitable/topicpage/dna-is-a-structure-that-encodes-biological-6493050 In short, stereochemistry is important in determining which helices
Pauling, Corey and Branson Diffraction of DNA http://www.nature.com/scitable/topicpage/dna-is-a-structure-that-encodes-biological-6493050 In short, stereochemistry is important in determining which helices
Basic concepts of molecular biology
 Basic concepts of molecular biology Gabriella Trucco Email: gabriella.trucco@unimi.it Life The main actors in the chemistry of life are molecules called proteins nucleic acids Proteins: many different
Basic concepts of molecular biology Gabriella Trucco Email: gabriella.trucco@unimi.it Life The main actors in the chemistry of life are molecules called proteins nucleic acids Proteins: many different
03-511/711 Computational Genomics and Molecular Biology, Fall
 03-511/711 Computational Genomics and Molecular Biology, Fall 2011 1 Problem Set 0 Due Tuesday, September 6th This homework is intended to be a self-administered placement quiz, to help you (and me) determine
03-511/711 Computational Genomics and Molecular Biology, Fall 2011 1 Problem Set 0 Due Tuesday, September 6th This homework is intended to be a self-administered placement quiz, to help you (and me) determine
11 questions for a total of 120 points
 Your Name: BYS 201, Final Exam, May 3, 2010 11 questions for a total of 120 points 1. 25 points Take a close look at these tables of amino acids. Some of them are hydrophilic, some hydrophobic, some positive
Your Name: BYS 201, Final Exam, May 3, 2010 11 questions for a total of 120 points 1. 25 points Take a close look at these tables of amino acids. Some of them are hydrophilic, some hydrophobic, some positive
Virtual bond representation
 Today s subjects: Virtual bond representation Coordination number Contact maps Sidechain packing: is it an instrumental way of selecting and consolidating a fold? ASA of proteins Interatomic distances
Today s subjects: Virtual bond representation Coordination number Contact maps Sidechain packing: is it an instrumental way of selecting and consolidating a fold? ASA of proteins Interatomic distances
First&year&tutorial&in&Chemical&Biology&(amino&acids,&peptide&and&proteins)&! 1.&!
 First&year&tutorial&in&Chemical&Biology&(amino&acids,&peptide&and&proteins& 1.& a. b. c. d. e. 2.& a. b. c. d. e. f. & UsingtheCahn Ingold Prelogsystem,assignstereochemicaldescriptorstothe threeaminoacidsshownbelow.
First&year&tutorial&in&Chemical&Biology&(amino&acids,&peptide&and&proteins& 1.& a. b. c. d. e. 2.& a. b. c. d. e. f. & UsingtheCahn Ingold Prelogsystem,assignstereochemicaldescriptorstothe threeaminoacidsshownbelow.
Problem Set Unit The base ratios in the DNA and RNA for an onion (Allium cepa) are given below.
 Problem Set Unit 3 Name 1. Which molecule is found in both DNA and RNA? A. Ribose B. Uracil C. Phosphate D. Amino acid 2. Which molecules form the nucleotide marked in the diagram? A. phosphate, deoxyribose
Problem Set Unit 3 Name 1. Which molecule is found in both DNA and RNA? A. Ribose B. Uracil C. Phosphate D. Amino acid 2. Which molecules form the nucleotide marked in the diagram? A. phosphate, deoxyribose
CFSSP: Chou and Fasman Secondary Structure Prediction server
 Wide Spectrum, Vol. 1, No. 9, (2013) pp 15-19 CFSSP: Chou and Fasman Secondary Structure Prediction server T. Ashok Kumar Department of Bioinformatics, Noorul Islam College of Arts and Science, Kumaracoil
Wide Spectrum, Vol. 1, No. 9, (2013) pp 15-19 CFSSP: Chou and Fasman Secondary Structure Prediction server T. Ashok Kumar Department of Bioinformatics, Noorul Islam College of Arts and Science, Kumaracoil
466 Asn (N) to Ala (A) Generate beta dimer Interface
 Table S1: Amino acid changes to the HexA α-subunit to convert the dimer interface from α to β and to introduce the putative GM2A binding surface from β- onto the α- subunit Residue position (α-numbering)
Table S1: Amino acid changes to the HexA α-subunit to convert the dimer interface from α to β and to introduce the putative GM2A binding surface from β- onto the α- subunit Residue position (α-numbering)
Basic concepts of molecular biology
 Basic concepts of molecular biology Gabriella Trucco Email: gabriella.trucco@unimi.it What is life made of? 1665: Robert Hooke discovered that organisms are composed of individual compartments called cells
Basic concepts of molecular biology Gabriella Trucco Email: gabriella.trucco@unimi.it What is life made of? 1665: Robert Hooke discovered that organisms are composed of individual compartments called cells
Structural bioinformatics
 Structural bioinformatics Why structures? The representation of the molecules in 3D is more informative New properties of the molecules are revealed, which can not be detected by sequences Eran Eyal Plant
Structural bioinformatics Why structures? The representation of the molecules in 3D is more informative New properties of the molecules are revealed, which can not be detected by sequences Eran Eyal Plant
Supplementary Data for Monti, et al.
 Supplementary Data for Monti, et al. Supplementary Figure S1 Legend to Supplementary Figure S1 Tumor spectrum associated with germline p53 alleles (restricted to the 7 most frequent tissue targets). Structural
Supplementary Data for Monti, et al. Supplementary Figure S1 Legend to Supplementary Figure S1 Tumor spectrum associated with germline p53 alleles (restricted to the 7 most frequent tissue targets). Structural
Bioinformatics. ONE Introduction to Biology. Sami Khuri Department of Computer Science San José State University Biology/CS 123A Fall 2012
 Bioinformatics ONE Introduction to Biology Sami Khuri Department of Computer Science San José State University Biology/CS 123A Fall 2012 Biology Review DNA RNA Proteins Central Dogma Transcription Translation
Bioinformatics ONE Introduction to Biology Sami Khuri Department of Computer Science San José State University Biology/CS 123A Fall 2012 Biology Review DNA RNA Proteins Central Dogma Transcription Translation
Supplemental Table 1. Amino acid sequences of synthetic kisspeptins
 Supplemental Data Supplemental Table 1. Amino acid sequences of synthetic kisspeptins Kisspeptins Symbol Sequence Human kisspeptin-10 H-10 Tyr-Asn-Trp-Asn-Ser-Phe-Gly-Leu-Arg-Phe-NH 2 Rodent/Xenopus 1a
Supplemental Data Supplemental Table 1. Amino acid sequences of synthetic kisspeptins Kisspeptins Symbol Sequence Human kisspeptin-10 H-10 Tyr-Asn-Trp-Asn-Ser-Phe-Gly-Leu-Arg-Phe-NH 2 Rodent/Xenopus 1a
Amino Acid Sequences and Evolutionary Relationships. How do similarities in amino acid sequences of various species provide evidence for evolution?
 Amino Acid Sequences and Evolutionary Relationships Name: How do similarities in amino acid sequences of various species provide evidence for evolution? An important technique used in determining evolutionary
Amino Acid Sequences and Evolutionary Relationships Name: How do similarities in amino acid sequences of various species provide evidence for evolution? An important technique used in determining evolutionary
Amino Acid Sequences and Evolutionary Relationships
 Amino Acid Sequences and Evolutionary Relationships One technique used to determine evolutionary relationships is to study the biochemical similarity of organisms. Though molds, aardvarks, and humans appear
Amino Acid Sequences and Evolutionary Relationships One technique used to determine evolutionary relationships is to study the biochemical similarity of organisms. Though molds, aardvarks, and humans appear
From code to translation
 From code to translation What could be the role of the first peptides? Ádám Kun & Ádám Radványi Dpt. Plant Systematics, Ecology and Theoretical Biology, Eötvös University, Budapest, Hungary Parmenides
From code to translation What could be the role of the first peptides? Ádám Kun & Ádám Radványi Dpt. Plant Systematics, Ecology and Theoretical Biology, Eötvös University, Budapest, Hungary Parmenides
1) The penicillin family of antibiotics, discovered by Alexander Fleming in 1928, has the following general structure: O O
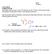 ame: TF ame: LS1a Fall 06 Problem Set #3 Due Friday 10/13 at noon in your TF s drop box on the 2 nd floor of the Science Center All questions including the (*extra*) ones should be turned in 1) The penicillin
ame: TF ame: LS1a Fall 06 Problem Set #3 Due Friday 10/13 at noon in your TF s drop box on the 2 nd floor of the Science Center All questions including the (*extra*) ones should be turned in 1) The penicillin
Case 7 A Storage Protein From Seeds of Brassica nigra is a Serine Protease Inhibitor
 Case 7 A Storage Protein From Seeds of Brassica nigra is a Serine Protease Inhibitor Focus concept Purification of a novel seed storage protein allows sequence analysis and determination of the protein
Case 7 A Storage Protein From Seeds of Brassica nigra is a Serine Protease Inhibitor Focus concept Purification of a novel seed storage protein allows sequence analysis and determination of the protein
1. DNA, RNA structure. 2. DNA replication. 3. Transcription, translation
 1. DNA, RNA structure 2. DNA replication 3. Transcription, translation DNA and RNA are polymers of nucleotides DNA is a nucleic acid, made of long chains of nucleotides Nucleotide Phosphate group Nitrogenous
1. DNA, RNA structure 2. DNA replication 3. Transcription, translation DNA and RNA are polymers of nucleotides DNA is a nucleic acid, made of long chains of nucleotides Nucleotide Phosphate group Nitrogenous
Case 7 A Storage Protein From Seeds of Brassica nigra is a Serine Protease Inhibitor Last modified 29 September 2005
 Case 7 A Storage Protein From Seeds of Brassica nigra is a Serine Protease Inhibitor Last modified 9 September 005 Focus concept Purification of a novel seed storage protein allows sequence analysis and
Case 7 A Storage Protein From Seeds of Brassica nigra is a Serine Protease Inhibitor Last modified 9 September 005 Focus concept Purification of a novel seed storage protein allows sequence analysis and
Amino Acid Sequences and Evolutionary Relationships
 Amino Acid Sequences and Evolutionary Relationships Pre-Lab Discussion Homologous structures -- those structures believed to have a common origin but not necessarily a common function -- provide some of
Amino Acid Sequences and Evolutionary Relationships Pre-Lab Discussion Homologous structures -- those structures believed to have a common origin but not necessarily a common function -- provide some of
Thr Gly Tyr. Gly Lys Asn
 Your unique body characteristics (traits), such as hair color or blood type, are determined by the proteins your body produces. Proteins are the building blocks of life - in fact, about 45% of the human
Your unique body characteristics (traits), such as hair color or blood type, are determined by the proteins your body produces. Proteins are the building blocks of life - in fact, about 45% of the human
Pacific Symposium on Biocomputing 4: (1999)
 Applications of Knowledge Discovery to Molecular Biology: Identifying Structural Regularities in Proteins Shaobing Su, Diane J. Cook, and Lawrence B. Holder University of Texas at Arlington sandy su@sabre.com,
Applications of Knowledge Discovery to Molecular Biology: Identifying Structural Regularities in Proteins Shaobing Su, Diane J. Cook, and Lawrence B. Holder University of Texas at Arlington sandy su@sabre.com,
Cristian Micheletti SISSA (Trieste)
 Cristian Micheletti SISSA (Trieste) michelet@sissa.it Mar 2009 5pal - parvalbumin Calcium-binding protein HEADER CALCIUM-BINDING PROTEIN 25-SEP-91 5PAL 5PAL 2 COMPND PARVALBUMIN (ALPHA LINEAGE) 5PAL 3
Cristian Micheletti SISSA (Trieste) michelet@sissa.it Mar 2009 5pal - parvalbumin Calcium-binding protein HEADER CALCIUM-BINDING PROTEIN 25-SEP-91 5PAL 5PAL 2 COMPND PARVALBUMIN (ALPHA LINEAGE) 5PAL 3
6-Foot Mini Toober Activity
 Big Idea The interaction between the substrate and enzyme is highly specific. Even a slight change in shape of either the substrate or the enzyme may alter the efficient and selective ability of the enzyme
Big Idea The interaction between the substrate and enzyme is highly specific. Even a slight change in shape of either the substrate or the enzyme may alter the efficient and selective ability of the enzyme
Fundamentals of Protein Structure
 Outline Fundamentals of Protein Structure Yu (Julie) Chen and Thomas Funkhouser Princeton University CS597A, Fall 2005 Protein structure Primary Secondary Tertiary Quaternary Forces and factors Levels
Outline Fundamentals of Protein Structure Yu (Julie) Chen and Thomas Funkhouser Princeton University CS597A, Fall 2005 Protein structure Primary Secondary Tertiary Quaternary Forces and factors Levels
7.014 Solution Set 4
 7.014 Solution Set 4 Question 1 Shown below is a fragment of the sequence of a hypothetical bacterial gene. This gene encodes production of HWDWN, protein essential for metabolizing sugar yummose. The
7.014 Solution Set 4 Question 1 Shown below is a fragment of the sequence of a hypothetical bacterial gene. This gene encodes production of HWDWN, protein essential for metabolizing sugar yummose. The
Protein Structure Analysis
 BINF 731 Protein Structure Analysis http://binf.gmu.edu/vaisman/binf731/ Secondary Structure: Computational Problems Secondary structure characterization Secondary structure assignment Secondary structure
BINF 731 Protein Structure Analysis http://binf.gmu.edu/vaisman/binf731/ Secondary Structure: Computational Problems Secondary structure characterization Secondary structure assignment Secondary structure
Alpha-helices, beta-sheets and U-turns within a protein are stabilized by (hint: two words).
 1 Quiz1 Q1 2011 Alpha-helices, beta-sheets and U-turns within a protein are stabilized by (hint: two words) Value Correct Answer 1 noncovalent interactions 100% Equals hydrogen bonds (100%) Equals H-bonds
1 Quiz1 Q1 2011 Alpha-helices, beta-sheets and U-turns within a protein are stabilized by (hint: two words) Value Correct Answer 1 noncovalent interactions 100% Equals hydrogen bonds (100%) Equals H-bonds
蛋白質體學. Proteomics Amino acids, Peptides and Proteins 陳威戎 & 21
 蛋白質體學 Proteomics 2015 Amino acids, Peptides and Proteins 陳威戎 2015. 09. 14 & 21 Outline 1. Amino Acids 2. Peptides and Proteins 3. Covalent Structure of Proteins Amino Acids Proteins are polymers of amino
蛋白質體學 Proteomics 2015 Amino acids, Peptides and Proteins 陳威戎 2015. 09. 14 & 21 Outline 1. Amino Acids 2. Peptides and Proteins 3. Covalent Structure of Proteins Amino Acids Proteins are polymers of amino
Protein NMR II. Lecture 5
 Protein NMR II Lecture 5 Standard and NMR chemical shifts in proteins Residue N A A B O Ala 123.8 4.35 52.5 19.0 177.1 ys 118.8 4.65 58.8 28.6 174.8 Asp 120.4 4.76 54.1 40.8 177.2 Glu 120.2 4.29 56.7 29.7
Protein NMR II Lecture 5 Standard and NMR chemical shifts in proteins Residue N A A B O Ala 123.8 4.35 52.5 19.0 177.1 ys 118.8 4.65 58.8 28.6 174.8 Asp 120.4 4.76 54.1 40.8 177.2 Glu 120.2 4.29 56.7 29.7
Honors Research in Molecular Genetics Part 1
 Third Base Honors Research in Molecular enetics Part 1 ll bout ene Expression How do cells synthesize polypeptides and convert them to functional proteins? The message in your DN of who you are and how
Third Base Honors Research in Molecular enetics Part 1 ll bout ene Expression How do cells synthesize polypeptides and convert them to functional proteins? The message in your DN of who you are and how
X-ray structures of fructosyl peptide oxidases revealing residues responsible for gating oxygen access in the oxidative half reaction
 X-ray structures of fructosyl peptide oxidases revealing residues responsible for gating oxygen access in the oxidative half reaction Tomohisa Shimasaki 1, Hiromi Yoshida 2, Shigehiro Kamitori 2 & Koji
X-ray structures of fructosyl peptide oxidases revealing residues responsible for gating oxygen access in the oxidative half reaction Tomohisa Shimasaki 1, Hiromi Yoshida 2, Shigehiro Kamitori 2 & Koji
Station 1 DNA Evidence
 Station 1 DNA Evidence Cytochrome-c is a protein found in the mitochondria that is used in cellular respiration. This protein consists of a chain of 104 amino acids. The chart below shows the amino acid
Station 1 DNA Evidence Cytochrome-c is a protein found in the mitochondria that is used in cellular respiration. This protein consists of a chain of 104 amino acids. The chart below shows the amino acid
DNA.notebook March 08, DNA Overview
 DNA Overview Deoxyribonucleic Acid, or DNA, must be able to do 2 things: 1) give instructions for building and maintaining cells. 2) be copied each time a cell divides. DNA is made of subunits called nucleotides
DNA Overview Deoxyribonucleic Acid, or DNA, must be able to do 2 things: 1) give instructions for building and maintaining cells. 2) be copied each time a cell divides. DNA is made of subunits called nucleotides
Daily Agenda. Warm Up: Review. Translation Notes Protein Synthesis Practice. Redos
 Daily Agenda Warm Up: Review Translation Notes Protein Synthesis Practice Redos 1. What is DNA Replication? 2. Where does DNA Replication take place? 3. Replicate this strand of DNA into complimentary
Daily Agenda Warm Up: Review Translation Notes Protein Synthesis Practice Redos 1. What is DNA Replication? 2. Where does DNA Replication take place? 3. Replicate this strand of DNA into complimentary
Amino Acids and Proteins
 Various Functions of Proteins SB203 Amino Acids and Proteins Jirundon Yuvaniyama, Ph.D. Department of Biochemistry Faculty of Science Mahidol University Enzymes Transport proteins utrient and storage proteins
Various Functions of Proteins SB203 Amino Acids and Proteins Jirundon Yuvaniyama, Ph.D. Department of Biochemistry Faculty of Science Mahidol University Enzymes Transport proteins utrient and storage proteins
Granby Transcription and Translation Services plc
 ompany Resources ranby Transcription and Translation Services plc has invested heavily in the Protein Synthesis business. mongst the resources available to new recruits are: the latest cellphones which
ompany Resources ranby Transcription and Translation Services plc has invested heavily in the Protein Synthesis business. mongst the resources available to new recruits are: the latest cellphones which
NORWEGIAN UNIVERSITY OF SCIENCE AND TECHNOLOGY DEPARTMENT OF BIOTECHNOLOGY Professor Bjørn E. Christensen, Department of Biotechnology
 Page 1 NRWEGIAN UNIVERSITY F SCIENCE AND TECNLGY DEPARTMENT F BITECNLGY Professor Bjørn E. Christensen, Department of Biotechnology Contact during the exam: phone: 73593327, 92634016 EXAM TBT4135 BIPLYMERS
Page 1 NRWEGIAN UNIVERSITY F SCIENCE AND TECNLGY DEPARTMENT F BITECNLGY Professor Bjørn E. Christensen, Department of Biotechnology Contact during the exam: phone: 73593327, 92634016 EXAM TBT4135 BIPLYMERS
The Stringent Response
 The Stringent Response When amino acids are limiting a response is triggered to shut down a wide range of biosynthetic processes This process is called the Stringent Response It results in the synthesis
The Stringent Response When amino acids are limiting a response is triggered to shut down a wide range of biosynthetic processes This process is called the Stringent Response It results in the synthesis
a) Give the sequence of the mrna transcribed from this gene and indicate the 5 and 3 ends of the mrna.
 ame: Section : 7.014 Problem Set 4 nswers to this problem set are to be turned in at the box outside 68120 by 11:45 am, Wednesday, March 9. Problem sets will not be accepted late. Solutions will be posted
ame: Section : 7.014 Problem Set 4 nswers to this problem set are to be turned in at the box outside 68120 by 11:45 am, Wednesday, March 9. Problem sets will not be accepted late. Solutions will be posted
Homework. A bit about the nature of the atoms of interest. Project. The role of electronega<vity
 Homework Why cited articles are especially useful. citeulike science citation index When cutting and pasting less is more. Project Your protein: I will mail these out this weekend If you haven t gotten
Homework Why cited articles are especially useful. citeulike science citation index When cutting and pasting less is more. Project Your protein: I will mail these out this weekend If you haven t gotten
Supporting Information Contents
 Supporting Information Choy Theng Loh, Kiyoshi Ozawa, Kellie L. Tuck, Nicholas Barlow, Thomas Huber, Gottfried Otting, and Bim Graham Lanthanide tags for site-specific ligation to an unnatural amino acid
Supporting Information Choy Theng Loh, Kiyoshi Ozawa, Kellie L. Tuck, Nicholas Barlow, Thomas Huber, Gottfried Otting, and Bim Graham Lanthanide tags for site-specific ligation to an unnatural amino acid
7.014 Quiz II Handout
 7.014 Quiz II Handout Quiz II: Wednesday, March 17 12:05-12:55 54-100 **This will be a closed book exam** Quiz Review Session: Friday, March 12 7:00-9:00 pm room 54-100 Open Tutoring Session: Tuesday,
7.014 Quiz II Handout Quiz II: Wednesday, March 17 12:05-12:55 54-100 **This will be a closed book exam** Quiz Review Session: Friday, March 12 7:00-9:00 pm room 54-100 Open Tutoring Session: Tuesday,
Solutions to 7.02 Quiz II 10/27/05
 Solutions to 7.02 Quiz II 10/27/05 Class Average = 83 Standard Deviation = 9 Range Grade % 87-100 A 43 74-86 B 39 55-73 C 17 > 54 D 1 Question 1 (56 points) While studying deep sea bacteria, you discover
Solutions to 7.02 Quiz II 10/27/05 Class Average = 83 Standard Deviation = 9 Range Grade % 87-100 A 43 74-86 B 39 55-73 C 17 > 54 D 1 Question 1 (56 points) While studying deep sea bacteria, you discover
Algorithms in Bioinformatics ONE Transcription Translation
 Algorithms in Bioinformatics ONE Transcription Translation Sami Khuri Department of Computer Science San José State University sami.khuri@sjsu.edu Biology Review DNA RNA Proteins Central Dogma Transcription
Algorithms in Bioinformatics ONE Transcription Translation Sami Khuri Department of Computer Science San José State University sami.khuri@sjsu.edu Biology Review DNA RNA Proteins Central Dogma Transcription
Programme Good morning and summary of last week Levels of Protein Structure - I Levels of Protein Structure - II
 Programme 8.00-8.10 Good morning and summary of last week 8.10-8.30 Levels of Protein Structure - I 8.30-9.00 Levels of Protein Structure - II 9.00-9.15 Break 9.15-11.15 Exercise: Building a protein model
Programme 8.00-8.10 Good morning and summary of last week 8.10-8.30 Levels of Protein Structure - I 8.30-9.00 Levels of Protein Structure - II 9.00-9.15 Break 9.15-11.15 Exercise: Building a protein model
Computational Methods for Protein Structure Prediction
 Computational Methods for Protein Structure Prediction Ying Xu 2017/12/6 1 Outline introduction to protein structures the problem of protein structure prediction why it is possible to predict protein structures
Computational Methods for Protein Structure Prediction Ying Xu 2017/12/6 1 Outline introduction to protein structures the problem of protein structure prediction why it is possible to predict protein structures
Endomorphin-1 and Endomorphin-2. These items may be or may be not in stock anymore. Please contact us, if you are interestet in purchasing.
 These items may be or may be not in stock anymore. Please contact us, if you are interestet in purchasing. Cyclic Nucleotide Antisera Antiserum against Cyclic AMP ( rabbit ) lyophilized RIA-Titer: 1 :
These items may be or may be not in stock anymore. Please contact us, if you are interestet in purchasing. Cyclic Nucleotide Antisera Antiserum against Cyclic AMP ( rabbit ) lyophilized RIA-Titer: 1 :
 Problem: The GC base pairs are more stable than AT base pairs. Why? 5. Triple-stranded DNA was first observed in 1957. Scientists later discovered that the formation of triplestranded DNA involves a type
Problem: The GC base pairs are more stable than AT base pairs. Why? 5. Triple-stranded DNA was first observed in 1957. Scientists later discovered that the formation of triplestranded DNA involves a type
Dynamic Programming Algorithms
 Dynamic Programming Algorithms Sequence alignments, scores, and significance Lucy Skrabanek ICB, WMC February 7, 212 Sequence alignment Compare two (or more) sequences to: Find regions of conservation
Dynamic Programming Algorithms Sequence alignments, scores, and significance Lucy Skrabanek ICB, WMC February 7, 212 Sequence alignment Compare two (or more) sequences to: Find regions of conservation
NRPS Code Project Summary
 NRPS Code Project Summary Nick ill. ata formatting/trimming he data used in this project was obtained from a paper which detailed a machine-learning approach to the prediction of amino-acids encoded by
NRPS Code Project Summary Nick ill. ata formatting/trimming he data used in this project was obtained from a paper which detailed a machine-learning approach to the prediction of amino-acids encoded by
NAME:... MODEL ANSWER... STUDENT NUMBER:... Maximum marks: 50. Internal Examiner: Hugh Murrell, Computer Science, UKZN
 COMP710, Bioinformatics with Julia, Test One, Thursday the 20 th of April, 2017, 09h30-11h30 1 NAME:...... MODEL ANSWER... STUDENT NUMBER:...... Maximum marks: 50 Internal Examiner: Hugh Murrell, Computer
COMP710, Bioinformatics with Julia, Test One, Thursday the 20 th of April, 2017, 09h30-11h30 1 NAME:...... MODEL ANSWER... STUDENT NUMBER:...... Maximum marks: 50 Internal Examiner: Hugh Murrell, Computer
Materials Protein synthesis kit. This kit consists of 24 amino acids, 24 transfer RNAs, four messenger RNAs and one ribosome (see below).
 Protein Synthesis Instructions The purpose of today s lab is to: Understand how a cell manufactures proteins from amino acids, using information stored in the genetic code. Assemble models of four very
Protein Synthesis Instructions The purpose of today s lab is to: Understand how a cell manufactures proteins from amino acids, using information stored in the genetic code. Assemble models of four very
Ali Yaghi. Tamara Wahbeh. Mamoun Ahram
 28 Ali Yaghi Tamara Wahbeh Mamoun Ahram This sheet is a continuation of protein purification methods. Isoelectric focusing Separation of proteins based on Isoelectric points(charge),and it is a horizontal
28 Ali Yaghi Tamara Wahbeh Mamoun Ahram This sheet is a continuation of protein purification methods. Isoelectric focusing Separation of proteins based on Isoelectric points(charge),and it is a horizontal
ENZYMES AND METABOLIC PATHWAYS
 ENZYMES AND METABOLIC PATHWAYS This document is licensed under the Attribution-NonCommercial-ShareAlike 2.5 Italy license, available at http://creativecommons.org/licenses/by-nc-sa/2.5/it/ 1. Enzymes build
ENZYMES AND METABOLIC PATHWAYS This document is licensed under the Attribution-NonCommercial-ShareAlike 2.5 Italy license, available at http://creativecommons.org/licenses/by-nc-sa/2.5/it/ 1. Enzymes build
Supplementary Table 1: List of CH3 domain interface residues in the first chain (A) and
 Supplementary Tables Supplementary Table 1: List of CH3 domain interface residues in the first chain (A) and their side chain contacting residues in the second chain (B) a Interface Res. in Contacting
Supplementary Tables Supplementary Table 1: List of CH3 domain interface residues in the first chain (A) and their side chain contacting residues in the second chain (B) a Interface Res. in Contacting
1/4/18 NUCLEIC ACIDS. Nucleic Acids. Nucleic Acids. ECS129 Instructor: Patrice Koehl
 NUCLEIC ACIDS ECS129 Instructor: Patrice Koehl Nucleic Acids Nucleotides DNA Structure RNA Synthesis Function Secondary structure Tertiary interactions Wobble hypothesis DNA RNA Replication Transcription
NUCLEIC ACIDS ECS129 Instructor: Patrice Koehl Nucleic Acids Nucleotides DNA Structure RNA Synthesis Function Secondary structure Tertiary interactions Wobble hypothesis DNA RNA Replication Transcription
NUCLEIC ACIDS. ECS129 Instructor: Patrice Koehl
 NUCLEIC ACIDS ECS129 Instructor: Patrice Koehl Nucleic Acids Nucleotides DNA Structure RNA Synthesis Function Secondary structure Tertiary interactions Wobble hypothesis DNA RNA Replication Transcription
NUCLEIC ACIDS ECS129 Instructor: Patrice Koehl Nucleic Acids Nucleotides DNA Structure RNA Synthesis Function Secondary structure Tertiary interactions Wobble hypothesis DNA RNA Replication Transcription
De novo sequencing in the identification of mass data. Wang Quanhui Liu Siqi Beijing Institute of Genomics, CAS
 De novo sequencing in the identification of mass data Wang Quanhui Liu Siqi Beijing Institute of Genomics, CAS The difficulties in mass data analysis Although the techniques of genomic sequencing are being
De novo sequencing in the identification of mass data Wang Quanhui Liu Siqi Beijing Institute of Genomics, CAS The difficulties in mass data analysis Although the techniques of genomic sequencing are being
Solutions to Problem Set 1
 MIT Department of Biology 7.014 Introductory Biology, Spring 004 Question 1 Solutions to 7.014 Problem Set 1 a) Describe the conditions of the atmosphere on prebiotic earth and how these conditions differ
MIT Department of Biology 7.014 Introductory Biology, Spring 004 Question 1 Solutions to 7.014 Problem Set 1 a) Describe the conditions of the atmosphere on prebiotic earth and how these conditions differ
Gene Expression Translation U C A G A G
 Why? ene Expression Translation How do cells synthesize polypeptides and convert them to functional proteins? The message in your DN of who you are and how your body works is carried out by cells through
Why? ene Expression Translation How do cells synthesize polypeptides and convert them to functional proteins? The message in your DN of who you are and how your body works is carried out by cells through
CS 4491/CS 7990 SPECIAL TOPICS IN BIOINFORMATICS
 1 CS 4491/CS 7990 SPECIAL TOPICS IN BIOINFORMATICS * Some contents are adapted from Dr. Jean Gao at UT Arlington Mingon Kang, PhD Computer Science, Kennesaw State University 2 Genetics The discovery of
1 CS 4491/CS 7990 SPECIAL TOPICS IN BIOINFORMATICS * Some contents are adapted from Dr. Jean Gao at UT Arlington Mingon Kang, PhD Computer Science, Kennesaw State University 2 Genetics The discovery of
7.013 Spring 2005 Problem Set 1
 MIT Department of Biology 7.013: Introductory Biology Spring 005 Instructors: rofessor azel Sive, rofessor Tyler Jacks, Dr. laudette Gardel AME TA Section # 7.013 Spring 005 roblem Set 1 FRIDAY February
MIT Department of Biology 7.013: Introductory Biology Spring 005 Instructors: rofessor azel Sive, rofessor Tyler Jacks, Dr. laudette Gardel AME TA Section # 7.013 Spring 005 roblem Set 1 FRIDAY February
Comprehensive analysis of proteolysis in long-ripened hard cooked Old Saare cheese
 Comprehensive analysis of proteolysis in long-ripened hard cooked Old Saare cheese Minna Varikmaa, Tiina Kriščiunaite, Natalja Kabanova, Irina Stulova, Viktoria Põžjanova, Raivo Vilu Competence Centre
Comprehensive analysis of proteolysis in long-ripened hard cooked Old Saare cheese Minna Varikmaa, Tiina Kriščiunaite, Natalja Kabanova, Irina Stulova, Viktoria Põžjanova, Raivo Vilu Competence Centre
Chapter Twelve Protein Synthesis: Translation of the Genetic Message
 Mary K. Campbell Shawn O. Farrell international.cengage.com/ Chapter Twelve Protein Synthesis: Translation of the Genetic Message Paul D. Adams University of Arkansas 1 Translating the Genetic Message
Mary K. Campbell Shawn O. Farrell international.cengage.com/ Chapter Twelve Protein Synthesis: Translation of the Genetic Message Paul D. Adams University of Arkansas 1 Translating the Genetic Message
7.013 Problem Set 3 FRIDAY October 8th, 2004
 MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert. Weinberg, Dr. laudette ardel Name: T: 7.013 Problem Set 3 FRIDY October 8th, 2004 Problem
MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert. Weinberg, Dr. laudette ardel Name: T: 7.013 Problem Set 3 FRIDY October 8th, 2004 Problem
Residue Contact Prediction for Protein Structure using 2-Norm Distances
 Residue Contact Prediction for Protein Structure using 2-Norm Distances Nikita V Mahajan Department of Computer Science &Engg GH Raisoni College of Engineering, Nagpur LGMalik Department of Computer Science
Residue Contact Prediction for Protein Structure using 2-Norm Distances Nikita V Mahajan Department of Computer Science &Engg GH Raisoni College of Engineering, Nagpur LGMalik Department of Computer Science
(a) Which enzyme(s) make 5' - 3' phosphodiester bonds? (c) Which enzyme(s) make single-strand breaks in DNA backbones?
 EXAMPLE QUESTIONS AND ANSWERS 1. Topoisomerase does which one of the following? (a) Makes new DNA strands. (b) Unties knots in DNA molecules. (c) Joins the ends of double-stranded DNA molecules. (d) Is
EXAMPLE QUESTIONS AND ANSWERS 1. Topoisomerase does which one of the following? (a) Makes new DNA strands. (b) Unties knots in DNA molecules. (c) Joins the ends of double-stranded DNA molecules. (d) Is
Important points from last time
 Important points from last time Subst. rates differ site by site Fit a Γ dist. to variation in rates Γ generally has two parameters but in biology we fix one to ensure a mean equal to 1 and the other parameter
Important points from last time Subst. rates differ site by site Fit a Γ dist. to variation in rates Γ generally has two parameters but in biology we fix one to ensure a mean equal to 1 and the other parameter
Degenerate Code. Translation. trna. The Code is Degenerate trna / Proofreading Ribosomes Translation Mechanism
 Translation The Code is Degenerate trna / Proofreading Ribosomes Translation Mechanism Degenerate Code There are 64 possible codon triplets There are 20 naturally-encoding amino acids Several codons specify
Translation The Code is Degenerate trna / Proofreading Ribosomes Translation Mechanism Degenerate Code There are 64 possible codon triplets There are 20 naturally-encoding amino acids Several codons specify
Abbreviations. Inhibitor synthesis
 Supplementary Material I for: Bailey et al. manuscript An analysis of subdomain orientation, conformational change and disorder in relation to crystal packing of aspartic proteinases. Abbreviations Boc
Supplementary Material I for: Bailey et al. manuscript An analysis of subdomain orientation, conformational change and disorder in relation to crystal packing of aspartic proteinases. Abbreviations Boc
Structure and Function of the First Full-Length Murein Peptide Ligase (Mpl) Cell Wall Recycling Protein
 Paper Presentation PLoS ONE 2011 Structure and Function of the First Full-Length Murein Peptide Ligase (Mpl) Cell Wall Recycling Protein Debanu Das, Mireille Herve, Julie Feuerhelm, etc. and Dominique
Paper Presentation PLoS ONE 2011 Structure and Function of the First Full-Length Murein Peptide Ligase (Mpl) Cell Wall Recycling Protein Debanu Das, Mireille Herve, Julie Feuerhelm, etc. and Dominique
 www.lessonplansinc.com Topic: Gene Mutations WS Summary: Students will learn about frame shift mutations and base substitution mutations. Goals & Objectives: Students will be able to demonstrate how mutations
www.lessonplansinc.com Topic: Gene Mutations WS Summary: Students will learn about frame shift mutations and base substitution mutations. Goals & Objectives: Students will be able to demonstrate how mutations
Structure formation and association of biomolecules. Prof. Dr. Martin Zacharias Lehrstuhl für Molekulardynamik (T38) Technische Universität München
 Structure formation and association of biomolecules Prof. Dr. Martin Zacharias Lehrstuhl für Molekulardynamik (T38) Technische Universität München Motivation Many biomolecules are chemically synthesized
Structure formation and association of biomolecules Prof. Dr. Martin Zacharias Lehrstuhl für Molekulardynamik (T38) Technische Universität München Motivation Many biomolecules are chemically synthesized
Proteins: Wide range of func2ons. Polypep2des. Amino Acid Monomers
 Proteins: Wide range of func2ons Proteins coded in DNA account for more than 50% of the dry mass of most cells Protein func9ons structural support storage transport cellular communica9ons movement defense
Proteins: Wide range of func2ons Proteins coded in DNA account for more than 50% of the dry mass of most cells Protein func9ons structural support storage transport cellular communica9ons movement defense
Custom [ 125 I] Radioiodination
![Custom [ 125 I] Radioiodination Custom [ 125 I] Radioiodination](/thumbs/80/82436545.jpg) Page 1 Custom [ 125 I] Radioiodination Radioiodination of proteins and peptides may be performed by different methods, namely by targeting an iodine-accepting group on the protein or by the conjugation
Page 1 Custom [ 125 I] Radioiodination Radioiodination of proteins and peptides may be performed by different methods, namely by targeting an iodine-accepting group on the protein or by the conjugation
Level 2 Biology, 2017
 91159 911590 2SUPERVISOR S Level 2 Biology, 2017 91159 Demonstrate understanding of gene expression 2.00 p.m. Wednesday 22 November 2017 Credits: Four Achievement Achievement with Merit Achievement with
91159 911590 2SUPERVISOR S Level 2 Biology, 2017 91159 Demonstrate understanding of gene expression 2.00 p.m. Wednesday 22 November 2017 Credits: Four Achievement Achievement with Merit Achievement with
Nucleic acid and protein Flow of genetic information
 Nucleic acid and protein Flow of genetic information References: Glick, BR and JJ Pasternak, 2003, Molecular Biotechnology: Principles and Applications of Recombinant DNA, ASM Press, Washington DC, pages.
Nucleic acid and protein Flow of genetic information References: Glick, BR and JJ Pasternak, 2003, Molecular Biotechnology: Principles and Applications of Recombinant DNA, ASM Press, Washington DC, pages.
This is the knowledge that you should understand upon completing this section:
 DN 11 Syllabus hecklist This is the knowledge that you should understand upon completing this section: 11.1 DN DN occurs bound to proteins in chromosomes in the nucs and as unbound DN in the mitochondria.
DN 11 Syllabus hecklist This is the knowledge that you should understand upon completing this section: 11.1 DN DN occurs bound to proteins in chromosomes in the nucs and as unbound DN in the mitochondria.
Cambridge International Examinations Cambridge International Advanced Subsidiary and Advanced Level
 ambridge International Examinations ambridge International Advanced Subsidiary and Advanced Level *8744875516* BIOLOGY 9700/22 Paper 2 AS Level Structured Questions October/November 2016 1 hour 15 minutes
ambridge International Examinations ambridge International Advanced Subsidiary and Advanced Level *8744875516* BIOLOGY 9700/22 Paper 2 AS Level Structured Questions October/November 2016 1 hour 15 minutes
iclicker Question #28B - after lecture Shown below is a diagram of a typical eukaryotic gene which encodes a protein: start codon stop codon 2 3
 Bio 111 Handout for Molecular Biology 4 This handout contains: Today s iclicker Questions Information on Exam 3 Solutions Fall 2008 Exam 3 iclicker Question #28A - before lecture Which of the following
Bio 111 Handout for Molecular Biology 4 This handout contains: Today s iclicker Questions Information on Exam 3 Solutions Fall 2008 Exam 3 iclicker Question #28A - before lecture Which of the following
1. DNA replication. (a) Why is DNA replication an essential process?
 ame Section 7.014 Problem Set 3 Please print out this problem set and record your answers on the printed copy. Answers to this problem set are to be turned in to the box outside 68120 by 5:00pm on Friday
ame Section 7.014 Problem Set 3 Please print out this problem set and record your answers on the printed copy. Answers to this problem set are to be turned in to the box outside 68120 by 5:00pm on Friday
MOLEBIO LAB #3: Electrophoretic Separation of Proteins
 MOLEBIO LAB #3: Electrophoretic Separation of Proteins Introduction: Proteins occupy a central position in the structure and function of all living organisms. Some proteins serve as structural components
MOLEBIO LAB #3: Electrophoretic Separation of Proteins Introduction: Proteins occupy a central position in the structure and function of all living organisms. Some proteins serve as structural components
Cambridge International Examinations Cambridge International Advanced Subsidiary and Advanced Level
 ambridge International Examinations ambridge International Advanced Subsidiary and Advanced Level *8744875516* BIOLOGY 9700/22 Paper 2 AS Level Structured Questions October/November 2016 1 hour 15 minutes
ambridge International Examinations ambridge International Advanced Subsidiary and Advanced Level *8744875516* BIOLOGY 9700/22 Paper 2 AS Level Structured Questions October/November 2016 1 hour 15 minutes
Unit 1. DNA and the Genome
 Unit 1 DNA and the Genome Gene Expression Key Area 3 Vocabulary 1: Transcription Translation Phenotype RNA (mrna, trna, rrna) Codon Anticodon Ribosome RNA polymerase RNA splicing Introns Extrons Gene Expression
Unit 1 DNA and the Genome Gene Expression Key Area 3 Vocabulary 1: Transcription Translation Phenotype RNA (mrna, trna, rrna) Codon Anticodon Ribosome RNA polymerase RNA splicing Introns Extrons Gene Expression
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Frequencies of DQ2.5, DQ2.2, DQ8 and DQ7.5, the most common haplotypes in celiac disease cases. The map illustrates the differences among European countries and shows a clear gradient
Supplementary Figure 1 Frequencies of DQ2.5, DQ2.2, DQ8 and DQ7.5, the most common haplotypes in celiac disease cases. The map illustrates the differences among European countries and shows a clear gradient
Using DNA sequence, distinguish species in the same genus from one another.
 Species Identification: Penguins 7. It s Not All Black and White! Name: Objective Using DNA sequence, distinguish species in the same genus from one another. Background In this activity, we will observe
Species Identification: Penguins 7. It s Not All Black and White! Name: Objective Using DNA sequence, distinguish species in the same genus from one another. Background In this activity, we will observe
ORFs and genes. Please sit in row K or forward
 ORFs and genes Please sit in row K or forward https://www.flickr.com/photos/teseum/3231682806/in/photostream/ Question: why do some strains of Vibrio cause cholera and others don t? Methods Mechanisms
ORFs and genes Please sit in row K or forward https://www.flickr.com/photos/teseum/3231682806/in/photostream/ Question: why do some strains of Vibrio cause cholera and others don t? Methods Mechanisms
Supporting Information
 Supporting Information Supplementary Figure-1 Results of Phage Display Screening Against TNT The sequences seen in Supplementary Figure-1 represent the phages sequenced after the third and fourth rounds
Supporting Information Supplementary Figure-1 Results of Phage Display Screening Against TNT The sequences seen in Supplementary Figure-1 represent the phages sequenced after the third and fourth rounds
Station 1: DNA Structure Use the figure above to answer each of the following questions. 1.This is the subunit that DNA is composed of. 2.
 1. Station 1: DNA Structure Use the figure above to answer each of the following questions. 1.This is the subunit that DNA is composed of. 2.This subunit is composed of what 3 parts? 3.What molecules make
1. Station 1: DNA Structure Use the figure above to answer each of the following questions. 1.This is the subunit that DNA is composed of. 2.This subunit is composed of what 3 parts? 3.What molecules make
Aipotu II: Biochemistry
 Aipotu II: Biochemistry Introduction: The Biological Phenomenon Under Study In this lab, you will continue to explore the biological mechanisms behind the expression of flower color in a hypothetical plant.
Aipotu II: Biochemistry Introduction: The Biological Phenomenon Under Study In this lab, you will continue to explore the biological mechanisms behind the expression of flower color in a hypothetical plant.
Packing of Secondary Structures
 7.88 Lecture Notes - 5 7.24/7.88J/5.48J The Protein Folding and Human Disease Packing of Secondary Structures Packing of Helices against sheets Packing of sheets against sheets Parallel Orthogonal Table:
7.88 Lecture Notes - 5 7.24/7.88J/5.48J The Protein Folding and Human Disease Packing of Secondary Structures Packing of Helices against sheets Packing of sheets against sheets Parallel Orthogonal Table:
Ch Biophysical Chemistry
 Ch 247.53. Biophysical Chemistry Nina Rosario L. Rojas 2012-2013 sem 1 Review of Protein Structure Why structure? Primary, secondary, tertiary structure Disulfide bonds scheme 2 STRUCTURE- REGULAR STRUCTURE
Ch 247.53. Biophysical Chemistry Nina Rosario L. Rojas 2012-2013 sem 1 Review of Protein Structure Why structure? Primary, secondary, tertiary structure Disulfide bonds scheme 2 STRUCTURE- REGULAR STRUCTURE
Zool 3200: Cell Biology Exam 3 3/6/15
 Name: Trask Zool 3200: Cell Biology Exam 3 3/6/15 Answer each of the following questions in the space provided; circle the correct answer or answers for each multiple choice question and circle either
Name: Trask Zool 3200: Cell Biology Exam 3 3/6/15 Answer each of the following questions in the space provided; circle the correct answer or answers for each multiple choice question and circle either
Outline. Pseudogenes. Pseudo-genes. The genetic code (DNA version) What is a gene? What is a gene? Dead genes Vitamin C Urate oxidase. Alan R.
 Pseudogenes Alan R. Rogers January 15, 2016 Dead genes Vitamin C Urate oxidase ψmyh16 GBA Globins 1 / 35 2 / 35 Pseudo-genes Genes are DNA sequences that code for protein. Some genes are broken and cannot
Pseudogenes Alan R. Rogers January 15, 2016 Dead genes Vitamin C Urate oxidase ψmyh16 GBA Globins 1 / 35 2 / 35 Pseudo-genes Genes are DNA sequences that code for protein. Some genes are broken and cannot
2018 Protein Modeling Exam Key
 2018 Protein Modeling Exam Key Multiple Choice: 1. Which of the following amino acids has a negative charge at ph 7? a. Gln b. Glu c. Ser d. Cys 2. Which of the following is an example of secondary structure?
2018 Protein Modeling Exam Key Multiple Choice: 1. Which of the following amino acids has a negative charge at ph 7? a. Gln b. Glu c. Ser d. Cys 2. Which of the following is an example of secondary structure?
