Reporting Checklist for Nature Neuroscience
|
|
|
- Preston Harmon
- 5 years ago
- Views:
Transcription
1 Corresponding Author: Manuscript Number: Manuscript Type: Lennart Mucke NNRS49607AZ Resource Reporting Checklist for Nature Neuroscience # Main s: # lementary s: 4 # lementary Tables: 9 # lementary Videos: This checklist is used to ensure good reporting standards and to improve the reproducibility of published results. For more information, please read Reporting Life Sciences Research. Please note that in the event of publication, it is mandatory that authors include all relevant methodological and statistical information in the manuscript. Statistics reporting, by figure example example Please specify the following information for each panel reporting quantitative data, and where each item is reported (section, e.g., & paragraph number). Each figure should ideally contain an exact sample size (n) for each experimental group/condition, where n is an exact number and not a range, a clear definition of how n is defined (for example x cells from x slices from x animals from x litters, collected over x days), a description of the statistical used, the results of the s, any descriptive statistics and clearly defined error bars if applicable. For any experiments using custom statistics, please indicate the used and stats obtained for each experiment. Each figure should include a statement of how many times the experiment shown was replicated in the lab; the details of sample collection should be sufficiently clear so that the replicability of the experiment is obvious to the reader. For experiments reported in the text but not in the figures, please use the paragraph number instead of the figure number. Note: Mean and standard deviation are not appropriate on small samples, and plotting independent data points is usually more informative. When technical replicates are reported, error and significance measures reflect the experimental variability and not the variability of the biological process; it is misleading not to state this clearly. FIGURE NUMBER 1a results, TEST USED WHICH TEST? oneway ANOVA unpaired t Fig. EXACT VALUE 9, 9, 10, 15 n DEFINED? mice from at least 3 litters/group 15 slices from 10 mice Methods para 8 DESCRIPTIVE STATS (AVERAGE, VARIANCE) REPORTED? mean / SEM mean / SEM Fig. P VALUE EXACT VALUE p = p = Fig. DEGREES OF FREEDOM & F/t/z/R/ETC VALUE VALUE F(3, 36) =.97 t(8) =.808 Fig. 1
2 FIGURE NUMBER b c leme ntary 1a leme ntary 1b TEST USED WHICH TEST? paired t with Holm correction unpaired t with Holm correction, Welch's correction is used for dip and trip unpaired t (with Welch's correction if F was significant) with Benjamini Hochberg correction unpaired t (with Welch's correction if F was significant) with Benjamini Hochberg correction 1 1 EXACT VALUE n DEFINED? 8 mice 10 and 1 Cohort A: 3, Cohort B: 1 Cohort B: 10 and 1, Cohort D: 1, Cohort E: 10 mice per fractionation group mice per group mice per group & & 1 & 1 & DESCRIPTIVE STATS (AVERAGE, VARIANCE) REPORTED? mean / SEM mean / SEM mean / SEM mean / SEM 1 1 P VALUE EXACT VALUE PSD95: p=0.001, p=0.003; Tau: p=0.5, p=0.5; Alpha synuclein: p=0.0, p=0.04 dip386404: p<0.0001, p<0.0005; p19505: p=0.0, p=0.065; p400404: p>0.999, p>0.999; p175191: p=0.055, p=0.11; trip386404: p=0.0003, p=0.001 not significant for any comparison not significant for any comparison correcte d pvalue range reported on graph and figure correcte d pvalue range reported on graph and figure 1 1 DEGREES OF FREEDOM & F/t/z/R/ETC VALUE VALUE PSD95: t=5.45, df=7; Tau: t=1.46, df=7; Alpha synuclein: t=.996, df=7 dip386404: t=6.381, df=10.4; p19505: t=.49, df=0; p400404: t=1.983e007, df=0; p175191: t=.04, df=0; trip386404: t=4.556, df=16.3; not significant for any comparison not significant for any comparison no values to report no values to report 1 1
3 leme ntary b leme ntary 3b Unpaired t unpaired t Fisher's exact and Chi Square Pearson's Chisquare 3 Statisti cs Metho ds paragr aph 3 Cohort C: 4 Cohort F: 56 In MBRD: 1 R methylati on, 10 on, K methylati on, 11 ubiquitin ation, phosphor ylation; 3 R, 17 K, 16 S/T Not in MBRD: R methylati on, 4 on, K methylati on, 4 ubiquitin ation, 5 phosphor ylation; 13 R, 4 K, 60 S/T 41 K in tau, 11 sites modified by on and ubiquitin ation, 3 on sites without ubiquitin, 4 ubiquitin sites without on mice per group mice per group b & 3b & Modificat ions listed in Table 1 and Fig. 1, from tau sequence (not listed) Modificat ions listed in Table and Fig. 1, from tau sequence (not listed) mean / SEM mean / SEM b 3b p=0.57 p=0.958 Fisher's exact for R methylation p =0.53, p=1.0; Fisher's exact for Kmethylation p =0.57, p=1.0; Fisher's exact for phosphorylati on p =0.14, p=0.44; Chisquare for on p=0.053, p=0.1; Chisquare for ubiquitination p=0.035, p=0.17; p= b 3b Correcte d values in Statistics Methods paragrap h 3 t=1.51, df=6 t=0.054, df=9 b 3b 3
4 Likelihood Ratio paragr aph 3 41 K in tau, 11 modified by ubiquitin ation and on; methylati ons on those 11 sites and 1 not Representative figures Modificat ions listed in Table and Fig. 1, from tau sequence (not listed) 1. Are any representative images shown (including Western blots and immunohistochemistry/staining) in the paper? If so, what figure(s)?. For each representative image, is there a clear statement of how many times this experiment was successfully repeated and a discussion of any limitations in repeatability? If so, where is this reported (section, paragraph #)? Statistics and general methods 1. Is there a justification of the sample size? If so, how was it justified? Even if no sample size calculation was performed, authors should report why the sample size is adequate to measure their effect size.. Are statistical s justified as appropriate for every figure? a. If there is a section summarizing the statistical methods in the methods, is the statistical for each experiment clearly defined? b. Do the data meet the assumptions of the specific statistical you chose (e.g. normality for a parametric )? Where is this described (section, paragraph #)? c. Is there any estimate of variance within each group of data? Is the variance similar between groups that are being statistically compared? Where is this described (section, paragraph #)? p= paragrap h 3 Yes, a representative western blot is shown in figure a. The fullsized images are provided in lementary 4. The quantification of the experiment and the number of mice used per groups is shown adjacent to the blot in figure b and in the figure, respectively. No statistical methods were used to predetermine sample sizes but our sample sizes are similar to those reported in previous publications (PMID14773, PMID18886, PMID358556). Given that each modification has its own variance, we believe our claims are better justified by the repetition of each quantitative mass spectrometry experiment and 3 times within the manuscript as shown in lementary 1. The justification for statistical s is in the Statistics section of the methods. Yes, the statistical s used are listed adjacent to the data and in the statistical methods section. Data distribution was assumed to be normal based on visual inspection but this was not formally ed; for transparency, data are linked within the figure s. Unequal variance between groups was determined by an F with a pvalue less than Welch's correction was applied when comparing groups with unequal variance, as noted in the Statistics methods and figure s. 4
5 d. Are s specified as one or twosided? e. Are there adjustments for multiple comparisons? Yes, adjustments for multiple comparisons are listed in the statistical methods and figure s. 3. Are criteria for excluding data points reported? Was this criterion established prior to data collection? Where is this described (section, paragraph #)? 4. Define the method of randomization used to assign subjects (or samples) to the experimental groups and to collect and process data. If no randomization was used, state so. Where does this appear (section, paragraph #)? 5. Is a statement of the extent to which investigator knew the group allocation during the experiment and in assessing outcome included? If no blinding was done, state so. 6. For experiments in live vertebrates, is a statement of compliance with ethical guidelines/regulations included? 7. Is the species of the animals used reported? 8. Is the strain of the animals (including background strains of KO/ transgenic animals used) reported? 9. Is the sex of the animals/subjects used reported? 10. Is the age of the animals/subjects reported? 11. For animals housed in a vivarium, is the light/dark cycle reported? 1. For animals housed in a vivarium, is the housing group (i.e. number of animals per cage) reported? Tau modifications with final value of zero, reflecting that this modification was not found in this sample, were excluded from the quantitative comparison, as stated in Quantitative Comparison of Tau Modifications in the methods. The frequency of zerovalues found for each tau modification was assessed separately. Mice were assigned to groups randomly based on mouse number and genotype, which is stated in the Mice and Hypothermia Induction section of the methods. Yes, the biochemical and mass spectrometry analysis was performed blinded to genotype and fractionation, as listed in the methods sections Mice and Hypothermia Induction. Yes, this is stated in the Mice and Hypothermia Induction section of the methods. Yes, the species of animal used is reported throughout the manuscript with strain listed in the Mice and Hypothermia Induction section of the methods. Yes, the strain is listed in the Mice and Hypothermia Induction section of the methods. Male and female animals were used in this study and the sex of all animals used is listed in detail in lementary Tables 1 and 3. The age of the animals used in the study is listed in figure, supplementary figure 1 and lementary Tables 1 and 3. Yes, the light/dark cycle is reported in the Mice and Hypothermia Induction section of the methods. Yes, mice were group housed as listed in the Mice and Hypothermia Induction section of the methods. 5
6 13. For behavioral experiments, is the time of day reported (e.g. light or dark cycle)? 14. Is the previous history of the animals/subjects (e.g. prior drug administration, surgery, behavioral ing) reported? a. If multiple behavioral s were conducted in the same group of animals, is this reported? 15. If any animals/subjects were excluded from analysis, is this reported? Reagents a. How were the criteria for exclusion defined? Where is this described (section, paragraph #)? b. Specify reasons for any discrepancy between the number of animals at the beginning and end of the study. Where is this described (section, paragraph #)? 1. Have antibodies been validated for use in the system under study (assay and species)? a. Is antibody catalog number given? Where does this appear (section, paragraph #)? b. Where were the validation data reported (citation, supplementary information, Antibodypedia)? Where does this appear (section, paragraph #)?. Cell line identity a. Are any cell lines used in this paper listed in the database of commonly misidentified cell lines maintained by ICLAC and NCBI Biosample? Yes, it is reported in the Mice and Hypothermia Induction section of the methods. All mice used in this study were naive, as reported in the Mice and Hypothermia Induction section of the methods. No animals were excluded from this study. The antibodies used have been validated for western blot by the manufacturer when purchased from a commercial source. The O GlcNAc antibody was validated in the literature in mouse and rat brain, and was confirmed to be specific to modified tau by our lab using mouse lacking endogenous tau and recombinant tau as negative controls. Yes, the catalog number is given in the Western Blotting section of the methods. The manufacturer provided validation of these antibodies for western blotting. 6
7 b. If yes, include in the Methods section a scientific justification of their useindicate here in which section and paragraph the justification can be found. c. For each cell line, include in the Methods section a statement that specifies: the source of the cell lines have the cell lines been authenticated? If so, by which method? have the cell lines been ed for mycoplasma contamination? Data deposition Data deposition in a public repository is mandatory for: a. Protein, DNA and RNA sequences b. Macromolecular structures c. Crystallographic data for small molecules d. Microarray data Deposition is strongly recommended for many other datasets for which structured public repositories exist; more details on our data policy are available here. We encourage the provision of other source data in supplementary information or in unstructured repositories such as Figshare and Dryad. We encourage publication of Data Descriptors (see Scientific Data) to maximize data reuse. 1. Are accession codes for deposit dates provided? Computer code/software Raw mass spectrometry is freely available in MS viewer and the access codes are listed in paragraph 8 of the Peptide Identification and LabelFree Quantitation by Mass Spectrometry section of the online methods. Any custom algorithm/software that is central to the methods must be supplied by the authors in a usable and readable form for readers at the time of publication. However, referees may ask for this information at any time during the review process. 1. Identify all custom software or scripts that were required to conduct the study and where in the procedures each was used.. If computer code was used to generate results that are central to the paper's conclusions, include a statement in the Methods section under "Code availability" to indicate whether and how the code can be accessed. Include version information as necessary and any restrictions on availability. Human subjects 7
8 1. Which IRB approved the protocol? Where is this stated (section, paragraph #)?. Is demographic information on all subjects provided? 3. Is the number of human subjects, their age and sex clearly defined? 4. Are the inclusion and exclusion criteria (if any) clearly specified? 5. How well were the groups matched? Where is this information described (section, paragraph #)? 6. Is a statement included confirming that informed consent was obtained from all subjects? 7. For publication of patient photos, is a statement included confirming that consent to publish was obtained? fmri studies For papers reporting functional imaging (fmri) results please ensure that these minimal reporting guidelines are met and that all this information is clearly provided in the methods: 1. Were any subjects scanned but then rejected for the analysis after the data was collected? a. If yes, is the number rejected and reasons for rejection described?. Is the number of blocks, trials or experimental units per session and/ or subjects specified? 3. Is the length of each trial and interval between trials specified? 4. Is a blocked, eventrelated, or mixed design being used? If applicable, please specify the block length or how the eventrelated or mixed design was optimized. 8
9 5. Is the task design clearly described? 6. How was behavioral performance measured? 7. Is an ANOVA or factorial design being used? 8. For data acquisition, is a whole brain scan used? If not, state area of acquisition. a. How was this region determined? 9. Is the field strength (in Tesla) of the MRI system stated? a. Is the pulse sequence type (gradient/spin echo, EPI/spiral) stated? b. Are the fieldofview, matrix size, slice thickness, and TE/TR/ flip angle clearly stated? 10. Are the software and specific parameters (model/functions, smoothing kernel size if applicable, etc.) used for data processing and preprocessing clearly stated? 11. Is the coordinate space for the anatomical/functional imaging data clearly defined as subject/native space or standardized stereotaxic space, e.g., original Talairach, MNI305, ICBM15, etc? Where (section, paragraph #)? 1. If there was data normalization/standardization to a specific space template, are the type of transformation (linear vs. nonlinear) used and image types being transformed clearly described? Where (section, paragraph #)? 13. How were anatomical locations determined, e.g., via an automated labeling algorithm (AAL), standardized coordinate database (Talairach daemon), probabilistic atlases, etc.? 14. Were any additional regressors (behavioral covariates, motion etc) used? 15. Is the contrast construction clearly defined? 16. Is a mixed/random effects or fixed inference used? a. If fixed effects inference used, is this justified? 17. Were repeated measures used (multiple measurements per subject)? 9
10 a. If so, are the method to account for within subject correlation and the assumptions made about variance clearly stated? 18. If the threshold used for inference and visualization in figures varies, is this clearly stated? 19. Are statistical inferences for multiple comparisons? a. If not, is this labeled as? 0. Are the results based on an ROI (region of interest) analysis? a. If so, is the rationale clearly described? b. How were the ROI s defined (functional vs anatomical localization)? 1. Is there correction for multiple comparisons within each voxel?. For clusterwise significance, is the clusterdefining threshold and the significance level defined? Additional comments Additional Comments 10
Reporting Checklist for Nature Neuroscience
 Corresponding Author: Manuscript Number: Manuscript Type: Leonard Petrucelli NNA52530C Article Reporting Checklist for Nature Neuroscience # Main s: 8 # Supplementary s: 10 # Supplementary Tables: 3 #
Corresponding Author: Manuscript Number: Manuscript Type: Leonard Petrucelli NNA52530C Article Reporting Checklist for Nature Neuroscience # Main s: 8 # Supplementary s: 10 # Supplementary Tables: 3 #
Reporting Checklist for Nature Neuroscience
 Corresponding Author: Manuscript Number: Manuscript Type: Michael Cole NNA51 Article Reporting Checklist for Nature Neuroscience # Main Figures: 5 # lementary Figures: 4 # lementary Tables: 0 # lementary
Corresponding Author: Manuscript Number: Manuscript Type: Michael Cole NNA51 Article Reporting Checklist for Nature Neuroscience # Main Figures: 5 # lementary Figures: 4 # lementary Tables: 0 # lementary
Reporting Checklist for Nature Neuroscience
 Corresponding Author: Manuscript Number: Manuscript Type: McColl NNRS50586AZ Resource Reporting Checklist for Nature Neuroscience # Main Figures: 8 # lementary Figures: 7 # lementary Tables: 13 # lementary
Corresponding Author: Manuscript Number: Manuscript Type: McColl NNRS50586AZ Resource Reporting Checklist for Nature Neuroscience # Main Figures: 8 # lementary Figures: 7 # lementary Tables: 13 # lementary
Reporting Checklist for Nature Neuroscience
 Corresponding Author: Manuscript Number: Manuscript Type: Feltri NNA54268B Article Reporting Checklist for Nature Neuroscience # Main Figures: 7 # Supplementary Figures: 5 # Supplementary Tables: 1 # Supplementary
Corresponding Author: Manuscript Number: Manuscript Type: Feltri NNA54268B Article Reporting Checklist for Nature Neuroscience # Main Figures: 7 # Supplementary Figures: 5 # Supplementary Tables: 1 # Supplementary
Reporting Checklist for Nature Neuroscience
 Corresponding Author: Manuscript Number: Manuscript Type: Yang Dan NNA56598 Article Reporting Checklist for Nature Neuroscience # Main Figures: 8 # Supplementary Figures: 12 # Supplementary Tables: 0 #
Corresponding Author: Manuscript Number: Manuscript Type: Yang Dan NNA56598 Article Reporting Checklist for Nature Neuroscience # Main Figures: 8 # Supplementary Figures: 12 # Supplementary Tables: 0 #
Reporting Checklist for Nature Neuroscience
 Corresponding Author: Manuscript Number: Manuscript Type: xiangdong fu, yuanchao xue NNA52804A Article Reporting Checklist Nature Neuroscience # Main Figures: 5 # Supplementary Figures: 7 # Supplementary
Corresponding Author: Manuscript Number: Manuscript Type: xiangdong fu, yuanchao xue NNA52804A Article Reporting Checklist Nature Neuroscience # Main Figures: 5 # Supplementary Figures: 7 # Supplementary
Reporting Checklist for Nature Neuroscience
 Corresponding Author: Manuscript Number: Manuscript Type: Liu NNA52890T Article Reporting Checklist for Nature Neuroscience # Main Figures: 4 # Supplementary Figures: 3 # Supplementary Tables: 1 # Supplementary
Corresponding Author: Manuscript Number: Manuscript Type: Liu NNA52890T Article Reporting Checklist for Nature Neuroscience # Main Figures: 4 # Supplementary Figures: 3 # Supplementary Tables: 1 # Supplementary
Reporting Checklist for Nature Neuroscience
 Corresponding Author: Manuscript Number: Manuscript Type: Robert Froemke NNA7412T Article Reporting Checklist for Nature Neuroscience # Main s: 7 # Supplementary s: 14 # Supplementary Tables: 0 # Supplementary
Corresponding Author: Manuscript Number: Manuscript Type: Robert Froemke NNA7412T Article Reporting Checklist for Nature Neuroscience # Main s: 7 # Supplementary s: 14 # Supplementary Tables: 0 # Supplementary
Reporting Checklist for Nature Neuroscience
 Corresponding Author: Manuscript Number: Manuscript Type: Chay T. Kuo NNA47008 Article Reporting Checklist for Nature Neuroscience # Main Figures: 7 # Supplementary Figures: 11 # Supplementary Tables:
Corresponding Author: Manuscript Number: Manuscript Type: Chay T. Kuo NNA47008 Article Reporting Checklist for Nature Neuroscience # Main Figures: 7 # Supplementary Figures: 11 # Supplementary Tables:
Reporting Checklist for Nature Neuroscience
 Corresponding Author: Manuscript Number: Manuscript Type: David Dupret NNBC5329A Brief Communication Reporting Checklist for Nature Neuroscience # s: 3 # s: 11 # Tables: 0 # Videos: 0 This checklist is
Corresponding Author: Manuscript Number: Manuscript Type: David Dupret NNBC5329A Brief Communication Reporting Checklist for Nature Neuroscience # s: 3 # s: 11 # Tables: 0 # Videos: 0 This checklist is
Reporting Checklist for Nature Neuroscience
 Corresponding Author: Manuscript Number: Manuscript Type: RuRong Ji NNA59063A Article Reporting Checklist for Nature Neuroscience # Main ures: 8 # Supplementary ures: 15 # Supplementary Tables: 0 # Supplementary
Corresponding Author: Manuscript Number: Manuscript Type: RuRong Ji NNA59063A Article Reporting Checklist for Nature Neuroscience # Main ures: 8 # Supplementary ures: 15 # Supplementary Tables: 0 # Supplementary
Reporting Checklist for Nature Neuroscience
 Corresponding Author: Manuscript Number: Manuscript Type: Rachel Wilson NNA48246B Article Reporting Checklist for Nature Neuroscience # Main Figures: 8 # Supplementary Figures: 7 # Supplementary Tables:
Corresponding Author: Manuscript Number: Manuscript Type: Rachel Wilson NNA48246B Article Reporting Checklist for Nature Neuroscience # Main Figures: 8 # Supplementary Figures: 7 # Supplementary Tables:
Reporting Checklist for Nature Neuroscience
 Corponding Author: Manuscript Number: Manuscript Type: Gregory C. DeAngelis NNA49009A Article Reporting Checklist for Nature Neuroscience # Main Figu: 7 # lementary Figu: # lementary Tables: 0 # lementary
Corponding Author: Manuscript Number: Manuscript Type: Gregory C. DeAngelis NNA49009A Article Reporting Checklist for Nature Neuroscience # Main Figu: 7 # lementary Figu: # lementary Tables: 0 # lementary
Reporting Checklist for Nature Neuroscience
 Corresponding Author: Manuscript Number: Manuscript Type: Hillel Adesnik NNA58169T Article Reporting Checklist for Nature Neuroscience # Main ures: 5 # lementary ures: 6 # lementary Tables: 1 # lementary
Corresponding Author: Manuscript Number: Manuscript Type: Hillel Adesnik NNA58169T Article Reporting Checklist for Nature Neuroscience # Main ures: 5 # lementary ures: 6 # lementary Tables: 1 # lementary
Reporting Checklist for Nature Neuroscience
 Corresponding Author: Marcelo Coba Manuscript Number: NNRS3314D Manuscript Type: Resource Reporting Checklist for Nature Neuroscience # Main Figures: 7 # lementary Figures: 4 # lementary Tables: 11 # lementary
Corresponding Author: Marcelo Coba Manuscript Number: NNRS3314D Manuscript Type: Resource Reporting Checklist for Nature Neuroscience # Main Figures: 7 # lementary Figures: 4 # lementary Tables: 11 # lementary
Reporting Checklist for Nature Neuroscience
 Corresponding Author: Manuscript Number: Manuscript Type: Ofer Yizhar, PhD NNA54616C Article Reporting Checklist for Nature Neuroscience # Main s: 5 # lementary s: 10 # lementary Tables: 0 # lementary
Corresponding Author: Manuscript Number: Manuscript Type: Ofer Yizhar, PhD NNA54616C Article Reporting Checklist for Nature Neuroscience # Main s: 5 # lementary s: 10 # lementary Tables: 0 # lementary
Reporting Checklist for Nature Neuroscience
 Corresponding Author: Manuscript Number: Manuscript Type: Atsushi Miyawaki NNT51493A Technical Report Reporting Checklist Nature Neuroscience # Main Figures: 7 # Supplementary Figures: 12 # Supplementary
Corresponding Author: Manuscript Number: Manuscript Type: Atsushi Miyawaki NNT51493A Technical Report Reporting Checklist Nature Neuroscience # Main Figures: 7 # Supplementary Figures: 12 # Supplementary
Reporting Checklist for Nature Neuroscience
 Corresponding Author: Manuscript Number: Manuscript Type: Benjamin R. Rost NNT51693 Technical Report Reporting Checklt for Nature Neuroscience # Main Figures: 6 # Supplementary Figures: 9 # Supplementary
Corresponding Author: Manuscript Number: Manuscript Type: Benjamin R. Rost NNT51693 Technical Report Reporting Checklt for Nature Neuroscience # Main Figures: 6 # Supplementary Figures: 9 # Supplementary
Reporting Checklist for Nature Neuroscience
 Correspondg Author: Manuscript Number: Manuscript Type: Spencer Smith NNA48000A Article Reportg Checklist for Nature Neuroscience # Ma Figures: 6 # Supplementary Figures: 8 # Supplementary Tables: 1 #
Correspondg Author: Manuscript Number: Manuscript Type: Spencer Smith NNA48000A Article Reportg Checklist for Nature Neuroscience # Ma Figures: 6 # Supplementary Figures: 8 # Supplementary Tables: 1 #
Life Sciences Reporting Summary
 Life Sciences Reporting Summary Adams Waldorf, Rajagopal, Gale Initial submission Revised version Final submission Nature Research wishes to improve the reproducibility of the work that we publish. This
Life Sciences Reporting Summary Adams Waldorf, Rajagopal, Gale Initial submission Revised version Final submission Nature Research wishes to improve the reproducibility of the work that we publish. This
Life Sciences Reporting Summary
 Life Sciences Reporting Summary Corresponding author(s): Rich, Jeremy N. Initial submission Revised version Final submission Nature Research wishes to improve the reproducibility of the work that we publish.
Life Sciences Reporting Summary Corresponding author(s): Rich, Jeremy N. Initial submission Revised version Final submission Nature Research wishes to improve the reproducibility of the work that we publish.
Our web collection on statistics for biologists may be useful.
 Reporting Summary Corresponding author(s): FELIPE CAVA Nature Research wishes to improve the reproducibility of the work that we publish. This form provides structure for consistency and transparency in
Reporting Summary Corresponding author(s): FELIPE CAVA Nature Research wishes to improve the reproducibility of the work that we publish. This form provides structure for consistency and transparency in
Jeffrey W. Elias, PhD UCD SOM Office of Research Grants Facilitation.
 NIH IS CHANGING/ENHANCING THE REVIEW CRITERIA AND FOCUSING ON MISSING OPPORTUNITIES TO EXAMINE SEX DIFFERENCES JOURNALS ARE CHANGING/ENHANCING THE REVIEW CRITERIA Jeffrey W. Elias, PhD UCD SOM Office of
NIH IS CHANGING/ENHANCING THE REVIEW CRITERIA AND FOCUSING ON MISSING OPPORTUNITIES TO EXAMINE SEX DIFFERENCES JOURNALS ARE CHANGING/ENHANCING THE REVIEW CRITERIA Jeffrey W. Elias, PhD UCD SOM Office of
Efforts to Improve Data Transparency at the JBC
 Efforts to Improve Data Transparency at the JBC Roger J. Colbran Associate Editor Credit: Amanda Fosang, Associate Editor 2 Why Improve Transparency? Irreproducibility of pre-clinical studies Negative
Efforts to Improve Data Transparency at the JBC Roger J. Colbran Associate Editor Credit: Amanda Fosang, Associate Editor 2 Why Improve Transparency? Irreproducibility of pre-clinical studies Negative
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION DOI: 10.1038/ncb3575 In the format provided by the authors and unedited. Supplementary Figure 1 Validation of key reagents and assays a, top, IHC with antibody recognizing specifically
SUPPLEMENTARY INFORMATION DOI: 10.1038/ncb3575 In the format provided by the authors and unedited. Supplementary Figure 1 Validation of key reagents and assays a, top, IHC with antibody recognizing specifically
PROMOTING TRANSPARENT RESEARCH METHODS, PROTOCOLS AND DATA TO REDUCE IRREPRODUCIBILITY
 PROMOTING TRANSPARENT RESEARCH METHODS, PROTOCOLS AND DATA TO REDUCE IRREPRODUCIBILITY December 2015 STM Innovations Seminar Andrew L Hufton, PhD Managing Editor, Scientific Data Nature Publishing Group
PROMOTING TRANSPARENT RESEARCH METHODS, PROTOCOLS AND DATA TO REDUCE IRREPRODUCIBILITY December 2015 STM Innovations Seminar Andrew L Hufton, PhD Managing Editor, Scientific Data Nature Publishing Group
Reporting Summary. Statistical parameters. Software and code. nature research reporting summary April Corresponding author(s):
 Reporting Summary Corresponding author(s): Alexander Flügel 2018-07-09758C Nature Research wishes to improve the reproducibility of the work that we publish. This form provides structure for consistency
Reporting Summary Corresponding author(s): Alexander Flügel 2018-07-09758C Nature Research wishes to improve the reproducibility of the work that we publish. This form provides structure for consistency
Our web collection on statistics for biologists may be useful.
 Reporting Summary Corresponding author(s): Qi-jing Li, Lilin Ye and Bo Zhu Nature Research wishes to improve the reproducibility of the work that we publish. This form provides structure for consistency
Reporting Summary Corresponding author(s): Qi-jing Li, Lilin Ye and Bo Zhu Nature Research wishes to improve the reproducibility of the work that we publish. This form provides structure for consistency
Important Information for MCP Authors
 Guidelines to Authors for Publication of Manuscripts Describing Development and Application of Targeted Mass Spectrometry Measurements of Peptides and Proteins and Submission Checklist The following Guidelines
Guidelines to Authors for Publication of Manuscripts Describing Development and Application of Targeted Mass Spectrometry Measurements of Peptides and Proteins and Submission Checklist The following Guidelines
Revision Checklist for Science Signaling Research Manuscripts: Data Requirements and Style Guidelines
 Revision Checklist for Science Signaling Research Manuscripts: Data Requirements and Style Guidelines Further information can be found at: http://stke.sciencemag.org/sites/default/files/researcharticlerevmsinstructions_0.pdf.
Revision Checklist for Science Signaling Research Manuscripts: Data Requirements and Style Guidelines Further information can be found at: http://stke.sciencemag.org/sites/default/files/researcharticlerevmsinstructions_0.pdf.
BIOEQUIVALENCE TRIAL INFORMATION FORM (Medicines and Allied Substances Act [No. 3] of 2013 Part V Section 39)
![BIOEQUIVALENCE TRIAL INFORMATION FORM (Medicines and Allied Substances Act [No. 3] of 2013 Part V Section 39) BIOEQUIVALENCE TRIAL INFORMATION FORM (Medicines and Allied Substances Act [No. 3] of 2013 Part V Section 39)](/thumbs/90/102231627.jpg) ZAMRA BTIF BIOEQUIVALENCE TRIAL INFORMATION FORM (Medicines and Allied Substances Act [No. 3] of 2013 Part V Section 39) The Guidelines on Bioequivalence Studies to be consulted in completing this form.
ZAMRA BTIF BIOEQUIVALENCE TRIAL INFORMATION FORM (Medicines and Allied Substances Act [No. 3] of 2013 Part V Section 39) The Guidelines on Bioequivalence Studies to be consulted in completing this form.
BIOEQUIVALENCE TRIAL INFORMATION
 PRESENTATION OF BIOEQUIVALENCE TRIAL INFORMATION BIOEQUIVALENCE TRIAL INFORMATION GENERAL INSTRUCTIONS: Please review all the instructions thoroughly and carefully prior to completing the Bioequivalence
PRESENTATION OF BIOEQUIVALENCE TRIAL INFORMATION BIOEQUIVALENCE TRIAL INFORMATION GENERAL INSTRUCTIONS: Please review all the instructions thoroughly and carefully prior to completing the Bioequivalence
ENCODE RBP Antibody Characterization Guidelines
 ENCODE RBP Antibody Characterization Guidelines Approved on November 18, 2016 Background An integral part of the ENCODE Project is to characterize the antibodies used in the experiments. This document
ENCODE RBP Antibody Characterization Guidelines Approved on November 18, 2016 Background An integral part of the ENCODE Project is to characterize the antibodies used in the experiments. This document
Medicines Control Authority Of Zimbabwe
 Medicines Control Authority Of Zimbabwe BIOEQUIVALENCE APPLICATION FORM Form: EVF03 This application form is designed to facilitate information exchange between the Applicant and the MCAZ for bioequivalence
Medicines Control Authority Of Zimbabwe BIOEQUIVALENCE APPLICATION FORM Form: EVF03 This application form is designed to facilitate information exchange between the Applicant and the MCAZ for bioequivalence
Microarray Gene Expression Analysis at CNIO
 Microarray Gene Expression Analysis at CNIO Orlando Domínguez Genomics Unit Biotechnology Program, CNIO 8 May 2013 Workflow, from samples to Gene Expression data Experimental design user/gu/ubio Samples
Microarray Gene Expression Analysis at CNIO Orlando Domínguez Genomics Unit Biotechnology Program, CNIO 8 May 2013 Workflow, from samples to Gene Expression data Experimental design user/gu/ubio Samples
Life Sciences Reporting Summary
 Life Sciences Reporting Summary Corresponding author(s): Christer Betsholtz Initial submission Revised version Final submission Nature Research wishes to improve the reproducibility of the work that we
Life Sciences Reporting Summary Corresponding author(s): Christer Betsholtz Initial submission Revised version Final submission Nature Research wishes to improve the reproducibility of the work that we
Cerebrospinal fluid tau, neurogranin and neurofilament light in Alzheimerís disease
 Cerebrospinal fluid tau, neurogranin and neurofilament light in Alzheimerís disease Niklas Mattsson, Philip Insel, Sebastian Palmqvist, Erik Portelius, Henrik Zetterberg, Michael Weiner, Kaj Blennow, Oskar
Cerebrospinal fluid tau, neurogranin and neurofilament light in Alzheimerís disease Niklas Mattsson, Philip Insel, Sebastian Palmqvist, Erik Portelius, Henrik Zetterberg, Michael Weiner, Kaj Blennow, Oskar
Precision in Quantitative Imaging: Trial Development and Quality Assurance
 Precision in Quantitative Imaging: Trial Development and Quality Assurance Susanna I Lee MD, PhD Thanks to: Mitchell Schnall, Mark Rosen. Dan Sullivan, Patrick Bossuyt Imaging Chain: Patient Data Raw data
Precision in Quantitative Imaging: Trial Development and Quality Assurance Susanna I Lee MD, PhD Thanks to: Mitchell Schnall, Mark Rosen. Dan Sullivan, Patrick Bossuyt Imaging Chain: Patient Data Raw data
Edinburgh Research Explorer
 Edinburgh Research Explorer Protocol for a retrospective, controlled cohort study of the impact of a change in Nature journals' editorial policy for life sciences research on the completeness of reporting
Edinburgh Research Explorer Protocol for a retrospective, controlled cohort study of the impact of a change in Nature journals' editorial policy for life sciences research on the completeness of reporting
Methods in Clinical Cancer Research Workshop Format for Protocol Concept Synopsis Sheet (blank) SYNOPSIS
 B Methods in Clinical Cancer Research Workshop Format for Protocol Concept Synopsis Sheet (blank) elow is the format to follow for the concept sheet for protocols. It is based on the International Committee
B Methods in Clinical Cancer Research Workshop Format for Protocol Concept Synopsis Sheet (blank) elow is the format to follow for the concept sheet for protocols. It is based on the International Committee
Life Sciences Reporting Summary
 Life Sciences Reporting Summary Corresponding author(s): Stein Aerts Initial submission Revised version Final submission Nature Research wishes to improve the reproducibility of the work that we publish.
Life Sciences Reporting Summary Corresponding author(s): Stein Aerts Initial submission Revised version Final submission Nature Research wishes to improve the reproducibility of the work that we publish.
Using Excel s Analysis ToolPak Add-In
 Using Excel s Analysis ToolPak Add-In Bijay Lal Pradhan, PhD Introduction I have a strong opinions that we can perform different quantitative analysis, including statistical analysis, in Excel. It is powerful,
Using Excel s Analysis ToolPak Add-In Bijay Lal Pradhan, PhD Introduction I have a strong opinions that we can perform different quantitative analysis, including statistical analysis, in Excel. It is powerful,
EECS730: Introduction to Bioinformatics
 EECS730: Introduction to Bioinformatics Lecture 14: Microarray Some slides were adapted from Dr. Luke Huan (University of Kansas), Dr. Shaojie Zhang (University of Central Florida), and Dr. Dong Xu and
EECS730: Introduction to Bioinformatics Lecture 14: Microarray Some slides were adapted from Dr. Luke Huan (University of Kansas), Dr. Shaojie Zhang (University of Central Florida), and Dr. Dong Xu and
Practical What & Where of Rigor and Reproducibility in the NIH Application. SIU School of Medicine Office of Grants & Contracts with Sophia Ran, PhD
 Practical What & Where of Rigor and Reproducibility in the NIH Application SIU School of Medicine Office of Grants & Contracts with Sophia Ran, PhD Recent Training Sessions NIH Regional Seminar Rigor and
Practical What & Where of Rigor and Reproducibility in the NIH Application SIU School of Medicine Office of Grants & Contracts with Sophia Ran, PhD Recent Training Sessions NIH Regional Seminar Rigor and
Reporting Checklist for Nature Neuroscience
 Coesponding Autho: Manuscipt Numbe: Manuscipt Type: Yuki Oka NNA5991A Aticle Repoting Checklist fo Natue Neuoscience # Main Figues: 7 # Supplementay Figues: # Supplementay Tables: 0 # Supplementay Videos:
Coesponding Autho: Manuscipt Numbe: Manuscipt Type: Yuki Oka NNA5991A Aticle Repoting Checklist fo Natue Neuoscience # Main Figues: 7 # Supplementay Figues: # Supplementay Tables: 0 # Supplementay Videos:
iphemap: An atlas of Phenotype to genotype relationships of human ipsc models of neurological diseases
 iphemap: An atlas of Phenotype to genotype relationships of human ipsc models of neurological diseases Ethan W. Hollingsworth, Jacob E. Vaughn, Josh C. Orack, Chelsea Skinner, Jamil Khouri, Sofia B. Lizarraga,
iphemap: An atlas of Phenotype to genotype relationships of human ipsc models of neurological diseases Ethan W. Hollingsworth, Jacob E. Vaughn, Josh C. Orack, Chelsea Skinner, Jamil Khouri, Sofia B. Lizarraga,
Supplementary Figure 1. APP cleavage assay. HEK293 cells were transfected with various
 Supplementary Figure 1. APP cleavage assay. HEK293 cells were transfected with various GST-tagged N-terminal truncated APP fragments including GST-APP full-length (FL), APP (123-695), APP (189-695), or
Supplementary Figure 1. APP cleavage assay. HEK293 cells were transfected with various GST-tagged N-terminal truncated APP fragments including GST-APP full-length (FL), APP (123-695), APP (189-695), or
Estoril Education Day
 Estoril Education Day -Experimental design in Proteomics October 23rd, 2010 Peter James Note Taking All the Powerpoint slides from the Talks are available for download from: http://www.immun.lth.se/education/
Estoril Education Day -Experimental design in Proteomics October 23rd, 2010 Peter James Note Taking All the Powerpoint slides from the Talks are available for download from: http://www.immun.lth.se/education/
Identification of biological themes in microarray data from a mouse heart development time series using GeneSifter
 Identification of biological themes in microarray data from a mouse heart development time series using GeneSifter VizX Labs, LLC Seattle, WA 98119 Abstract Oligonucleotide microarrays were used to study
Identification of biological themes in microarray data from a mouse heart development time series using GeneSifter VizX Labs, LLC Seattle, WA 98119 Abstract Oligonucleotide microarrays were used to study
Animal Research Ethics Committee. Guidelines for Submitting Protocols for ethical approval of research or teaching involving live animals
 Animal Research Ethics Committee Guidelines for Submitting Protocols for ethical approval of research or teaching involving live animals AREC Document No: 3 Revised Guidelines for Submitting Protocols
Animal Research Ethics Committee Guidelines for Submitting Protocols for ethical approval of research or teaching involving live animals AREC Document No: 3 Revised Guidelines for Submitting Protocols
Supplementary Figure 1. Utilization of publicly available antibodies in different applications.
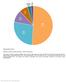 Supplementary Figure 1 Utilization of publicly available antibodies in different applications. The fraction of publicly available antibodies toward human protein targets is shown with data for the following
Supplementary Figure 1 Utilization of publicly available antibodies in different applications. The fraction of publicly available antibodies toward human protein targets is shown with data for the following
MRI Protocols in Experimental Stroke. Xenios Milidonis Centre for Clinical Brain Sciences The University of Edinburgh
 MRI Protocols in Experimental Stroke Xenios Milidonis Centre for Clinical Brain Sciences The University of Edinburgh MultiPART Meeting, Barcelona, 2 October 204 Objective In MultiPART magnetic resonance
MRI Protocols in Experimental Stroke Xenios Milidonis Centre for Clinical Brain Sciences The University of Edinburgh MultiPART Meeting, Barcelona, 2 October 204 Objective In MultiPART magnetic resonance
Psy 420 Midterm 2 Part 2 (Version A) In lab (50 points total)
 Psy 40 Midterm Part (Version A) In lab (50 points total) A researcher wants to know if memory is improved by repetition (Duh!). So he shows a group of five participants a list of 0 words at for different
Psy 40 Midterm Part (Version A) In lab (50 points total) A researcher wants to know if memory is improved by repetition (Duh!). So he shows a group of five participants a list of 0 words at for different
A normal genetic variation modulates synaptic MMP-9 protein levels and the severity of schizophrenia symptoms
 A normal genetic variation modulates synaptic MMP-9 protein levels and the severity of schizophrenia symptoms Katarzyna Lepeta, Katarzyna J. Purzycka, Katarzyna Pachulska-Wieczorek, Marina Mitjans, Martin
A normal genetic variation modulates synaptic MMP-9 protein levels and the severity of schizophrenia symptoms Katarzyna Lepeta, Katarzyna J. Purzycka, Katarzyna Pachulska-Wieczorek, Marina Mitjans, Martin
FFGWAS. Fast Functional Genome Wide Association AnalysiS of Surface-based Imaging Genetic Data
 FFGWAS Fast Functional Genome Wide Association AnalysiS of Surface-based Imaging Genetic Data Chao Huang Department of Biostatistics Biomedical Research Imaging Center The University of North Carolina
FFGWAS Fast Functional Genome Wide Association AnalysiS of Surface-based Imaging Genetic Data Chao Huang Department of Biostatistics Biomedical Research Imaging Center The University of North Carolina
CHAPTER FIVE CROSSTABS PROCEDURE
 CHAPTER FIVE CROSSTABS PROCEDURE 5.0 Introduction This chapter focuses on how to compare groups when the outcome is categorical (nominal or ordinal) by using SPSS. The aim of the series of exercise is
CHAPTER FIVE CROSSTABS PROCEDURE 5.0 Introduction This chapter focuses on how to compare groups when the outcome is categorical (nominal or ordinal) by using SPSS. The aim of the series of exercise is
Gene Expression Data Analysis
 Gene Expression Data Analysis Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu BMIF 310, Fall 2009 Gene expression technologies (summary) Hybridization-based
Gene Expression Data Analysis Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu BMIF 310, Fall 2009 Gene expression technologies (summary) Hybridization-based
Real-time RT PCR. for identification of differentially expressed genes. (with Schizophrenia application) Rolf Sundberg, Stockholm Univ.
 Real-time RT PCR for identification of differentially expressed genes. (with Schizophrenia application) Rolf Sundberg, Stockholm Univ. Göteborg, May 2006 Real-time PCR for mrna quantitation Review paper
Real-time RT PCR for identification of differentially expressed genes. (with Schizophrenia application) Rolf Sundberg, Stockholm Univ. Göteborg, May 2006 Real-time PCR for mrna quantitation Review paper
BIOSTATISTICS AND MEDICAL INFORMATICS (B M I)
 Biostatistics and Medical Informatics (B M I) 1 BIOSTATISTICS AND MEDICAL INFORMATICS (B M I) B M I/POP HLTH 451 INTRODUCTION TO SAS PROGRAMMING FOR 2 credits. Use of the SAS programming language for the
Biostatistics and Medical Informatics (B M I) 1 BIOSTATISTICS AND MEDICAL INFORMATICS (B M I) B M I/POP HLTH 451 INTRODUCTION TO SAS PROGRAMMING FOR 2 credits. Use of the SAS programming language for the
Jiali Li, Sukhvir P Mahal, Cheryl A Demczyk and Charles Weissmann
 Manuscript EMBOR-2011-35144 Mutability of prions Jiali Li, Sukhvir P Mahal, Cheryl A Demczyk and Charles Weissmann Corresponding author: Charles Weissmann, Scripps Florida Review timeline: Submission date:
Manuscript EMBOR-2011-35144 Mutability of prions Jiali Li, Sukhvir P Mahal, Cheryl A Demczyk and Charles Weissmann Corresponding author: Charles Weissmann, Scripps Florida Review timeline: Submission date:
SUPPLEMENTARY INFORMATION
 DOI:.38/ncb327 a b Sequence coverage (%) 4 3 2 IP: -GFP isoform IP: GFP IP: -GFP IP: GFP Sequence coverage (%) 4 3 2 IP: -GFP IP: GFP 33 52 58 isoform 2 33 49 47 IP: Control IP: Peptide Sequence Start
DOI:.38/ncb327 a b Sequence coverage (%) 4 3 2 IP: -GFP isoform IP: GFP IP: -GFP IP: GFP Sequence coverage (%) 4 3 2 IP: -GFP IP: GFP 33 52 58 isoform 2 33 49 47 IP: Control IP: Peptide Sequence Start
Integrative Genomics 1a. Introduction
 2016 Course Outline Integrative Genomics 1a. Introduction ggibson.gt@gmail.com http://www.cig.gatech.edu 1a. Experimental Design and Hypothesis Testing (GG) 1b. Normalization (GG) 2a. RNASeq (MI) 2b. Clustering
2016 Course Outline Integrative Genomics 1a. Introduction ggibson.gt@gmail.com http://www.cig.gatech.edu 1a. Experimental Design and Hypothesis Testing (GG) 1b. Normalization (GG) 2a. RNASeq (MI) 2b. Clustering
SECTION 11 ACUTE TOXICITY DATA ANALYSIS
 SECTION 11 ACUTE TOXICITY DATA ANALYSIS 11.1 INTRODUCTION 11.1.1 The objective of acute toxicity tests with effluents and receiving waters is to identify discharges of toxic effluents in acutely toxic
SECTION 11 ACUTE TOXICITY DATA ANALYSIS 11.1 INTRODUCTION 11.1.1 The objective of acute toxicity tests with effluents and receiving waters is to identify discharges of toxic effluents in acutely toxic
Searching for Truth = Experiment
 STATISTICS AND ANIMAL EXPERIMENTATION: THE GOOD, THE BAD AND THE UGL Dr. Manuel Berdoy Oxford University Veterinary Services. OSLO June 09 The Essence of Statistical Thinking, Common Mistakes and Solutions:
STATISTICS AND ANIMAL EXPERIMENTATION: THE GOOD, THE BAD AND THE UGL Dr. Manuel Berdoy Oxford University Veterinary Services. OSLO June 09 The Essence of Statistical Thinking, Common Mistakes and Solutions:
Supplementary Material Cohort Description
 Supplementary Material Cohort Description The following longitudinal twin cohorts were included: BrainSCALE: Brain Structure and Cognition: an Adolescent Longitudinal Twin Study into genetic etiology University
Supplementary Material Cohort Description The following longitudinal twin cohorts were included: BrainSCALE: Brain Structure and Cognition: an Adolescent Longitudinal Twin Study into genetic etiology University
AP Statistics Scope & Sequence
 AP Statistics Scope & Sequence Grading Period Unit Title Learning Targets Throughout the School Year First Grading Period *Apply mathematics to problems in everyday life *Use a problem-solving model that
AP Statistics Scope & Sequence Grading Period Unit Title Learning Targets Throughout the School Year First Grading Period *Apply mathematics to problems in everyday life *Use a problem-solving model that
David M. Rocke Division of Biostatistics and Department of Biomedical Engineering University of California, Davis
 David M. Rocke Division of Biostatistics and Department of Biomedical Engineering University of California, Davis Outline RNA-Seq for differential expression analysis Statistical methods for RNA-Seq: Structure
David M. Rocke Division of Biostatistics and Department of Biomedical Engineering University of California, Davis Outline RNA-Seq for differential expression analysis Statistical methods for RNA-Seq: Structure
Ngf (Rat) ELISA Kit. Catalog Number KA assays Version: 33. Intended for research use only.
 Ngf (Rat) ELISA Kit Catalog Number KA0401 96 assays Version: 33 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Intended Use... 3 Background... 3 Principle of the Assay...
Ngf (Rat) ELISA Kit Catalog Number KA0401 96 assays Version: 33 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Intended Use... 3 Background... 3 Principle of the Assay...
New NIH 2-page Rigor and Transparency Document
 New NIH 2-page Rigor and Transparency Document Grants affected in 2016 include: NOT-OD-16-081 NOT-OD-16-058 NOT-OD-16-034 NOT-OD-16-011 NOT-OD-16-031 NOT-OD-16-012 NOT-OD-16-005 NOT-OD-16-004 NOT-OD-15-103
New NIH 2-page Rigor and Transparency Document Grants affected in 2016 include: NOT-OD-16-081 NOT-OD-16-058 NOT-OD-16-034 NOT-OD-16-011 NOT-OD-16-031 NOT-OD-16-012 NOT-OD-16-005 NOT-OD-16-004 NOT-OD-15-103
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for Integrative Biology. This journal is The Royal Society of Chemistry 2015 Electronic Supplementary Information Table S1. Definition of quantitative cellular features
Electronic Supplementary Material (ESI) for Integrative Biology. This journal is The Royal Society of Chemistry 2015 Electronic Supplementary Information Table S1. Definition of quantitative cellular features
QTL mapping in mice. Karl W Broman. Department of Biostatistics Johns Hopkins University Baltimore, Maryland, USA.
 QTL mapping in mice Karl W Broman Department of Biostatistics Johns Hopkins University Baltimore, Maryland, USA www.biostat.jhsph.edu/ kbroman Outline Experiments, data, and goals Models ANOVA at marker
QTL mapping in mice Karl W Broman Department of Biostatistics Johns Hopkins University Baltimore, Maryland, USA www.biostat.jhsph.edu/ kbroman Outline Experiments, data, and goals Models ANOVA at marker
Validation of Analytical Methods used for the Characterization, Physicochemical and Functional Analysis and of Biopharmaceuticals.
 Validation of Analytical Methods used for the Characterization, Physicochemical and Functional Analysis and of Biopharmaceuticals. 1 Analytical Method Validation: 1..1 Philosophy: Method validation is
Validation of Analytical Methods used for the Characterization, Physicochemical and Functional Analysis and of Biopharmaceuticals. 1 Analytical Method Validation: 1..1 Philosophy: Method validation is
Alternative TSSs are co-regulated in single cells in the mouse brain
 Alternative TSSs are co-regulated in single cells in the mouse brain Kasper Karlsson, Peter Lönnerberg, Sten Linnarsson Corresponding author: Sten Linnarsson, Karolinska Institutet Review timeline: Submission
Alternative TSSs are co-regulated in single cells in the mouse brain Kasper Karlsson, Peter Lönnerberg, Sten Linnarsson Corresponding author: Sten Linnarsson, Karolinska Institutet Review timeline: Submission
Mouse TNF alpha ELISA Kit
 Mouse TNF alpha ELISA Kit Catalog No. GWB-ZZD049 Size 96 wells/kit Sandwich ELISA kit for quantitative detection of mouse TNF alpha in cell culture supernates, serum and plasma(heparin, EDTA). Typical
Mouse TNF alpha ELISA Kit Catalog No. GWB-ZZD049 Size 96 wells/kit Sandwich ELISA kit for quantitative detection of mouse TNF alpha in cell culture supernates, serum and plasma(heparin, EDTA). Typical
Aims and challenges of clinical trial data sharing
 Aims and challenges of clinical trial data sharing Facilitating data access to non-industry funded research UCL and Yale meeting: 9 October 2015 London Dr Trish Groves Head of research,
Aims and challenges of clinical trial data sharing Facilitating data access to non-industry funded research UCL and Yale meeting: 9 October 2015 London Dr Trish Groves Head of research,
PROPENSITY SCORE MATCHING A PRACTICAL TUTORIAL
 PROPENSITY SCORE MATCHING A PRACTICAL TUTORIAL Cody Chiuzan, PhD Biostatistics, Epidemiology and Research Design (BERD) Lecture March 19, 2018 1 Outline Experimental vs Non-Experimental Study WHEN and
PROPENSITY SCORE MATCHING A PRACTICAL TUTORIAL Cody Chiuzan, PhD Biostatistics, Epidemiology and Research Design (BERD) Lecture March 19, 2018 1 Outline Experimental vs Non-Experimental Study WHEN and
STAT 2300: Unit 1 Learning Objectives Spring 2019
 STAT 2300: Unit 1 Learning Objectives Spring 2019 Unit tests are written to evaluate student comprehension, acquisition, and synthesis of these skills. The problems listed as Assigned MyStatLab Problems
STAT 2300: Unit 1 Learning Objectives Spring 2019 Unit tests are written to evaluate student comprehension, acquisition, and synthesis of these skills. The problems listed as Assigned MyStatLab Problems
Papers for 11 September
 Papers for 11 September v Kreitman M (1983) Nucleotide polymorphism at the alcohol-dehydrogenase locus of Drosophila melanogaster. Nature 304, 412-417. v Hishimoto et al. (2010) Alcohol and aldehyde dehydrogenase
Papers for 11 September v Kreitman M (1983) Nucleotide polymorphism at the alcohol-dehydrogenase locus of Drosophila melanogaster. Nature 304, 412-417. v Hishimoto et al. (2010) Alcohol and aldehyde dehydrogenase
RIGOR AND REPRODUCIBILITY: BACK TO BASICS
 RIGOR AND REPRODUCIBILITY: BACK TO BASICS OCT 27, 2016 NIH REGIONAL SEMINAR - CHICAGO OVERVIEW Why the concern about reproducibility? The NIH response Updates to grant applications Training and Resources
RIGOR AND REPRODUCIBILITY: BACK TO BASICS OCT 27, 2016 NIH REGIONAL SEMINAR - CHICAGO OVERVIEW Why the concern about reproducibility? The NIH response Updates to grant applications Training and Resources
For quantitative detection of mouse EPO in cell culture supernates, serum, plasma (heparin, EDTA) and urine.
 Mouse EPO ELISA Kit Catalog No. EK0333 Size 96T(8 12 divisible strips) For quantitative detection of mouse EPO in cell culture supernates, serum, plasma (heparin, EDTA) and urine. Typical Data Obtained
Mouse EPO ELISA Kit Catalog No. EK0333 Size 96T(8 12 divisible strips) For quantitative detection of mouse EPO in cell culture supernates, serum, plasma (heparin, EDTA) and urine. Typical Data Obtained
Discussion Solution Mollusks and Litter Decomposition
 Discussion Solution Mollusks and Litter Decomposition. Is the rate of litter decomposition affected by the presence of mollusks? 2. Does the effect of mollusks on litter decomposition differ among the
Discussion Solution Mollusks and Litter Decomposition. Is the rate of litter decomposition affected by the presence of mollusks? 2. Does the effect of mollusks on litter decomposition differ among the
Archives of Scientific Psychology Reporting Questionnaire for Manuscripts Describing Primary Data Collections
 (Based on APA Journal Article Reporting Standards JARS Questionnaire) 1 Archives of Scientific Psychology Reporting Questionnaire for Manuscripts Describing Primary Data Collections JARS: ALL: These questions
(Based on APA Journal Article Reporting Standards JARS Questionnaire) 1 Archives of Scientific Psychology Reporting Questionnaire for Manuscripts Describing Primary Data Collections JARS: ALL: These questions
Seven Keys to Successful Microarray Data Analysis
 Seven Keys to Successful Microarray Data Analysis Experiment Design Platform Selection Data Management System Access Differential Expression Biological Significance Data Publication Type of experiment
Seven Keys to Successful Microarray Data Analysis Experiment Design Platform Selection Data Management System Access Differential Expression Biological Significance Data Publication Type of experiment
Lab 1: A review of linear models
 Lab 1: A review of linear models The purpose of this lab is to help you review basic statistical methods in linear models and understanding the implementation of these methods in R. In general, we need
Lab 1: A review of linear models The purpose of this lab is to help you review basic statistical methods in linear models and understanding the implementation of these methods in R. In general, we need
QTL mapping in mice. Karl W Broman. Department of Biostatistics Johns Hopkins University Baltimore, Maryland, USA.
 QTL mapping in mice Karl W Broman Department of Biostatistics Johns Hopkins University Baltimore, Maryland, USA www.biostat.jhsph.edu/ kbroman Outline Experiments, data, and goals Models ANOVA at marker
QTL mapping in mice Karl W Broman Department of Biostatistics Johns Hopkins University Baltimore, Maryland, USA www.biostat.jhsph.edu/ kbroman Outline Experiments, data, and goals Models ANOVA at marker
Supplementary Figure 1. α-synuclein is truncated in PD and LBD brains. Nature Structural & Molecular Biology: doi: /nsmb.
 Supplementary Figure 1 α-synuclein is truncated in PD and LBD brains. (a) Specificity of anti-n103 antibody. Anti-N103 antibody was coated on an ELISA plate and different concentrations of full-length
Supplementary Figure 1 α-synuclein is truncated in PD and LBD brains. (a) Specificity of anti-n103 antibody. Anti-N103 antibody was coated on an ELISA plate and different concentrations of full-length
Common Issues in Qualification and Validation of Analytical Procedures
 Common Issues in Qualification and Validation of Analytical Procedures Alexey Khrenov, PhD OTAT/CBER/FDA CMC Strategy Forum January 29, 2018 - Washington, DC Disclaimer These comments are an informal communication
Common Issues in Qualification and Validation of Analytical Procedures Alexey Khrenov, PhD OTAT/CBER/FDA CMC Strategy Forum January 29, 2018 - Washington, DC Disclaimer These comments are an informal communication
Phosphoproteome dynamics reveal novel ERK1/2 MAP kinase substrates with broad spectrum of functions
 Molecular Systems Biology Peer Review Process File Phosphoproteome dynamics reveal novel ERK1/2 MAP kinase substrates with broad spectrum of functions Mathieu Courcelles, Christophe Frémin, Laure Voisin,
Molecular Systems Biology Peer Review Process File Phosphoproteome dynamics reveal novel ERK1/2 MAP kinase substrates with broad spectrum of functions Mathieu Courcelles, Christophe Frémin, Laure Voisin,
Expanded View Figures
 MO reports Phenotype of 1T human tau transgenic mice Sumihiro Maeda et al xpanded View igures Western lotting LIS HT7/GPH 3 1 HT7/ctin 3 1 HT7-77G7P htau Level (ng/mg) 1 1 htau-wt (L1) htau-1t (L3) htau
MO reports Phenotype of 1T human tau transgenic mice Sumihiro Maeda et al xpanded View igures Western lotting LIS HT7/GPH 3 1 HT7/ctin 3 1 HT7-77G7P htau Level (ng/mg) 1 1 htau-wt (L1) htau-1t (L3) htau
Supplementary Figure 1. TSA (10 nmol/l), non-class-selective HDAC inhibitor, potentiates
 Supplementary Figure 1. TSA (10 nmol/l), non-class-selective HDAC inhibitor, potentiates vascular calcification (VC). (a) Von Kossa staining shows that TSA potentiated the Pi-induced VC. Scale bar, 100
Supplementary Figure 1. TSA (10 nmol/l), non-class-selective HDAC inhibitor, potentiates vascular calcification (VC). (a) Von Kossa staining shows that TSA potentiated the Pi-induced VC. Scale bar, 100
Laboratory Developed Tests. William Castellani, MD Inter-regional Commissioner Clinical Pathologist, Penn State Hershey Medical Center
 Laboratory Developed Tests William Castellani, MD Inter-regional Commissioner Clinical Pathologist, Penn State Hershey Medical Center Objectives Compare and contrast new test validation requirements for
Laboratory Developed Tests William Castellani, MD Inter-regional Commissioner Clinical Pathologist, Penn State Hershey Medical Center Objectives Compare and contrast new test validation requirements for
Experimental Design and Statistical Methods. Workshop GENERAL OVERVIEW. Jesús Piedrafita Arilla.
 Experimental Design and Statistical Methods Workshop GENERAL OVERVIEW Jesús Piedrafita Arilla jesus.piedrafita@uab.cat Departament de Ciència Animal i dels Aliments TWO TYPES OF STUDIES EXPERIMENT: it
Experimental Design and Statistical Methods Workshop GENERAL OVERVIEW Jesús Piedrafita Arilla jesus.piedrafita@uab.cat Departament de Ciència Animal i dels Aliments TWO TYPES OF STUDIES EXPERIMENT: it
Designing a Complex-Omics Experiments. Xiangqin Cui. Section on Statistical Genetics Department of Biostatistics University of Alabama at Birmingham
 Designing a Complex-Omics Experiments Xiangqin Cui Section on Statistical Genetics Department of Biostatistics University of Alabama at Birmingham 1/7/2015 Some slides are from previous lectures of Grier
Designing a Complex-Omics Experiments Xiangqin Cui Section on Statistical Genetics Department of Biostatistics University of Alabama at Birmingham 1/7/2015 Some slides are from previous lectures of Grier
Design of Experiments (DOE) Instructor: Thomas Oesterle
 1 Design of Experiments (DOE) Instructor: Thomas Oesterle 2 Instructor Thomas Oesterle thomas.oest@gmail.com 3 Agenda Introduction Planning the Experiment Selecting a Design Matrix Analyzing the Data Modeling
1 Design of Experiments (DOE) Instructor: Thomas Oesterle 2 Instructor Thomas Oesterle thomas.oest@gmail.com 3 Agenda Introduction Planning the Experiment Selecting a Design Matrix Analyzing the Data Modeling
CHAPTER-IV DATA ANALYSIS AND INTERPRETATION
 CHAPTER-IV DATA ANALYSIS AND INTERPRETATION 4.1 Introduction Data analysis is a process of assigning meaning to collected data, analysing significance and determination of findings and conclusions. Data
CHAPTER-IV DATA ANALYSIS AND INTERPRETATION 4.1 Introduction Data analysis is a process of assigning meaning to collected data, analysing significance and determination of findings and conclusions. Data
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Design and sequence of system 1 for targeted demethylation.
 Supplementary Figure 1 Design and sequence of system 1 for targeted demethylation. (a) Design of system 1 for targeted demethylation. TET1CD was fused to a catalytic inactive Cas9 nuclease (dcas9) and
Supplementary Figure 1 Design and sequence of system 1 for targeted demethylation. (a) Design of system 1 for targeted demethylation. TET1CD was fused to a catalytic inactive Cas9 nuclease (dcas9) and
Crowe Critical Appraisal Tool (CCAT) User Guide
 Crowe Critical Appraisal Tool (CCAT) User Guide Version 1.4 (19 November 2013) Use with the CCAT Form version 1.4 only Michael Crowe, PhD michael.crowe@my.jcu.edu.au This work is licensed under the Creative
Crowe Critical Appraisal Tool (CCAT) User Guide Version 1.4 (19 November 2013) Use with the CCAT Form version 1.4 only Michael Crowe, PhD michael.crowe@my.jcu.edu.au This work is licensed under the Creative
Figure S1. TT and MZ-CRC-1 cell viability and proliferation after Palbociclib treatment. Cell viability assays (A) and cell counting assays (B) were
 Figure S1. TT and MZ-CRC-1 cell viability and proliferation after Palbociclib treatment. Cell viability assays (A) and cell counting assays (B) were performed with TT and MZ-CRC-1 cells treated with Palbociclib
Figure S1. TT and MZ-CRC-1 cell viability and proliferation after Palbociclib treatment. Cell viability assays (A) and cell counting assays (B) were performed with TT and MZ-CRC-1 cells treated with Palbociclib
Statistical Methods for Quantitative Trait Loci (QTL) Mapping
 Statistical Methods for Quantitative Trait Loci (QTL) Mapping Lectures 4 Oct 10, 011 CSE 57 Computational Biology, Fall 011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday 1:00-1:0 Johnson
Statistical Methods for Quantitative Trait Loci (QTL) Mapping Lectures 4 Oct 10, 011 CSE 57 Computational Biology, Fall 011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday 1:00-1:0 Johnson
ProteinPilot Report for ProteinPilot Software
 ProteinPilot Report for ProteinPilot Software Detailed Analysis of Protein Identification / Quantitation Results Automatically Sean L Seymour, Christie Hunter SCIEX, USA Powerful mass spectrometers like
ProteinPilot Report for ProteinPilot Software Detailed Analysis of Protein Identification / Quantitation Results Automatically Sean L Seymour, Christie Hunter SCIEX, USA Powerful mass spectrometers like
