CTX-M-27. Escherichia coli 312 Clinical and Laboratory Standards Institute double-disk synergy test 15 (4.8 ) extendedspectrum
|
|
|
- Edward Kristian Bryan
- 6 years ago
- Views:
Transcription
1 b- CTX-M-27 1) 1, 2) 1, 3) 1) 2) 3) Escherichia coli 312 Clinical and Laboratory Standards Institute double-disk synergy test 15 (4.8 ) extendedspectrum b-lactamase (ESBL) 13 levofloxacin (LVFX) 7 LVFX trimethoprim/sulfamethoxazole (TMP/SMX) ESBL ESBL 15 8 bla CTX-M-27,4 bla CTX-M-14 bla CTX-M-27 ceftazidime ESBL LVFX 13 7 O25 : H4 bla CTX-M cells/recipient TMP/SMX E. coli CTX-M- 27 O25 : H4 ESBL Key words: extended-spectrum b-lactamase, uropathogenic Escherichia coli, CTX-M-27, Extended-spectrum b-lactamase (ESBL) class A b ) ESBL TEM-, SHV-, CTX-M 1 3) 1990 TEM- SHV- ESBL Escherichia coli Klebsiella sp CTX-M ( ) TEL: FAX: kumiko@met.nagoya-u.ac.jp 1, 2) CTX-M-15 multilocus sequence typing (MLST) 131, O25 : H4 E. coli 2) CTX-M pandemic 2006 Infectious Diseases Society of America 4) CTX-M-15 b-lactamase CTX-M- 1 bla CTX-M (Aps Gly) b-lactamase IncFII 5, 6) CTX-M-15 b-lactamase E. coli TEM-1, OXA-1 25
2 26 b-lactamase qnr 5, 6) 7) CTX-M-15 E. coli ) ESBL E. coli ESBL ESBL E. coli ESBL ESBL ESBL E. coli , 161, 99, 213, CFU/ml E. coli E. coli ATCC ESBL E. coli CSH-2 E. coli MC4100 Salmonella H ESBL Clinical and Laboratory Standards Institute (CLSI) 9) cefotaxime (CTX), ceftazidime (CAZ), trimethoprim/sulfamethoxazole (TMP/ SMX), gentamicin (GM), imipenem (IPM), levofloxacin (LVFX), fosfomycin (FOM) KB ESBL CLSI 9) double-disk synergy test Etest ESBL CT/CTL, ESBL TZ/TZL, 3. ESBL ESBL Shibata 10) Yagi 11) PCR CTX-M-1, CTX-M-2, CTX-M-8, CTX-M-9, SHV- TEM- ESBL Table 1 DNA Wizard Genomic DNA Purification Kit (Promega) PCR DNA 1 ml (20 ng) PCR (1.0 U Taq polymerase, 0.2 mm dntp mixture, 20 mm Tris HCl (ph 8.4), 50 mm KCl, 2.5 mm MgCl 2 (TaKaRa), 0.3 mm primer set) 24 ml EtBr 2 QIAquick Gel Extraction Kit (QIA- GEN) BigDye Terminator v1.1 Cycle Sequencing Kit (Applied Biosystems) ABI PRISM TM 310 Genetic Analyzer (Applied Biosystems) BLAST program (DDBJ, Shizuoka, Japan) 4. qnra, qnrb qnrs Cattoir 12), aac(6 )-Ib-cr Park 13), qepa Yamane 14) PCR PCR ESBL Table 1 26
3 ESBL CTX-M Table 1. Primers used in this study for PCR and nucleotide sequencing PCR primers Annealing temperature ( ) Size of PCR product (bp) References CTX-M-1-F CTX-M-1-R CTX-M-2-F CTX-M-2-R CTX-M-8-F CTX-M-8-R CTX-M-9-F CTX-M-9-R TEM-specific-F (T1) a TEM-specific-R (T2) a SHV-specific-F (S1) a SHV-specific-R (S2) a QnrA-F QnrA-R QnrB-F QnrB-R QnrS-F QnrS-R Aac(6 )-Ib-cr-F Aac(6 )-Ib-cr-R QeqA-F QepA-R 5 -GCT GTT GTT AGG AAG TGT GC-3 5 -CCA TTG CCC GAG GTG AAG-3 5 -ACG CTA CCC CTG CTA TTT-3 5 -CCT TTC CGC CTT CTG CTC-3 5 -CGG ATG ATG CTA ATG ACA AC-3 5 -GTC AGA TTG CGA AGC GTC-3 5 -GCA GAT AAT ACG CAG GTG-3 5 -CGG CGT GGT GGT GTC TCT-3 5 -CCG TGT CGC CCT TAT TCC-3 5 -AGG CAC CTA TCT CAG CGA-3 5 -ATT TGT CGC TTC TTT ACT CGC-3 5 -TTT ATG GCG TTA CCT TTG ACC-3 5 -AGA GGA TTT CTC ACG CCA GG-3 5 -TGC CAG GCA CAG ATC TTG AC-3 5 -GGA ATA GAA ATT CGC CAC TG-3 5 -TTT GCT GTT CGC CAG TCG AA-3 5 -GCA AGT TCA TTG AAC AGG GT-3 5 -TCT AAA CCG TCG AGT TCG GCG-3 5 -TTG CGA TGC TCT ATG AGT GGC TA-3 5 -CTC GAA TGC CTG GCG TGT TT-3 5 -GCA GGT CCA GCA GCG GGT AG-3 5 -CTT CCT GCC CGA GTA TCG TG ) ) ) ) ) ) ) ) ) ) ) Sequencing primer for TEM and SHV b T3 T4 T5 T6 S3 S4 S5 S6 5 -CAG TCA CAG AAA AGC AT-3 5 -CTG GCG AAC TAC TTA CTC-3 5 -TAG GCG GAG GTA GGT CAG-3 5 -TAC GAA AAG ACA CTG ACC-3 5 -GCG GGT GGA TGC CGG TG-3 5 -CGG CGG GCT GGT TTA TCG-3 5 -TAA ATC ACC ACA ATG CCC-3 5 -GTC GGC AAG GTG TTT TTC-3 11) 11) a To determine the genotype of ESBL, PCR was performed using the TEM- and SHV-specific primers (T1 and T2 for TEM. S1 and S2 for SHV). b For nucleotide sequencing, the several sequencing primers (T3, T4, T5 and T6 for TEM; S3, S4, S5 and S6 for SHV) were used. 5. O H (PFGE) Barrett 15) XbaI 6 V/cm, s 19 CHEF-DRIII BIO-RAD PFGE Fingerprinting II (BIO RAD) 6. ESBL E. coli CSH-2 Broth Mating 16) Luria Bertani (LB) broth LB broth (OD 660 : ) ml 1:4 100 ml 400 ml 10,000 rpm 1 27
4 ml 20 mg/ml CTX 50 mg/ml rifampicin LB PFGE E. coli DNA QIAprep spin plasmid kit (QIAGEN) ESBL double-disk synergy test Etest TMP/ SMX CPFX Disk DNA PstI / E. coli LVFX (15.7 ), TMP/SMX (9.6 ), GM (9.3 ), CTX (4.8 ) CAZ FOM 1 IPM 1 LVFX (Table 2) 312 CTX disk Table 2. Antimicrobial susceptibility of 312 Escherichia coli stains Antimicrobial agents Number of resistant isolates Total Inpatient Outpatient CTX 15 ( 4.8 ) 8 (2.6 ) 7 (2.2 ) CAZ 2 ( 0.6 ) 0 (0.0 ) 2 (0.6 ) IPM 0 ( 0.0 ) 0 (0.0 ) 0 (0.0 ) LVFX 49 (15.7 ) 19 (6.1 ) 30 (9.6 ) GM 29 ( 9.3 ) 13 (4.2 ) 16 (5.1 ) TMP/SMX 30 ( 9.6 ) 12 (3.8 ) 18 (5.8 ) FOM 1 ( 0.3 ) 1 (0.3 ) 0 (0.0 ) 27 mm CAZ 22 mm ESBL 40 double-disk synergy test Etest ESBL 15/312 (4.8 ) 15 Table ESBL 4/99 (4.04 ), 11/213 (5.16 ) 8 7 ESBL E. coli LVFX 49 PCR qnrs 1 ESBL ESBL LVFX LVFX 13 7 TMP/SMX ESBL (Table 3) ESBL 15 bla CTX-M-1 bla CTX-M-2 1 CTX-M- 9 bla CTX-M 12 bla CTX-M-14 ; 4 bla CTX-M-27 ; 8 bla TEM-1 3 bla SHV-12 1 bla CTX-M-14 3 bla TEM-1 CTX-M-9 12 LVFX b- bla CTX-M-14 bla CTX-M-27 CAZ (Table 3) 6 CTX-M-9 O25 : H4 (Table 3) PFGE A 3 (No. 41, 45, 79), C 4 (No. 22, 70, 179, 186) 7 No O25 : H4 A 3 2 (No. 45, 79) C 4 3 (No. 22, 179, 186) (Table 3, Fig. 1) ESBL 15 Broth Mating CTX-M-9 4 (No. 70, 79, 179, 239) kbp, cells/recipient 28
5 ESBL CTX-M Table 3. Genetic and phenotypic traits of ESBL-producing Escherichia coli strains Strain number Age Sex (M/F) a Inpatient/ Outpatient ESBL genotype b MIC (mg/ml)c CTX CAZ Resistance phenotype to non-b-lactam d Serotype e Hospital a b c d e F Inpatient CTX-M O125 : H4 B M Outpatient CTX-M LVFX, TMP/SMX OUT : H4 C F Outpatient CTX-M LVFX, TMP/SMX OUT : H4 C F Inpatient CTX-M LVFX O25 : H4 C F Inpatient CTX-M14, TEM LVFX OUT : H4 A M Outpatient CTX-M LVFX O25 : H4 A M Inpatient CTX-M LVFX, TMP/SMX O25 : H4 C M Outpatient CTX-M-14, TEM LVFX O25 : H4 A F Inpatient CTX-M LVFX, TMP/SMX, GM O169 : HUT E F Outpatient SHV OUT : HUT B F Outpatient CTX-M LVFX, TMP/SMX O25 : H4 C F Outpatient CTX-M LVFX, TMP/SMX O25 : H4 C F Inpatient CTX-M14, TEM LVFX OUT : H4 D F Inpatient CTX-M LVFX, TMP/SMX OUT : H4 B F Inpatient CTX-M LVFX O25 : H4 B M, male; F, female b-lactamase genes were determined by PCR and by sequencing. MICs were deterimined using E test ESBL CT/CTL and ESBL TZ/TZL. LVFX, levofloxacin; TMP/SMX, trimethoprim/sulfamethoxazole; GM, gentamicin OUT, O-antigen untypeable; HUT, H-antigen untypeable (Fig. 2) 4 E. coli CSH-2 LVFX GM CTX double-disk synergy test ESBL 3 TMP/SMX E. coli TMP/SMX 17) E. coli TMP/SMX , 19) E. coli K. pneumoniae ESBL 6, 20) E. coli ESBL 4.8, 15/ Yamaguchi 4.3, 6/140 21) , 13/205 8) ESBL /15 LVFX /15 LVFX TMP/SMX ESBL ESBL 2) ESBL E. coli ESBL CTX-M ESBL ST 131 5, 22) (QRDR) 5, 22) 29
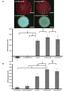
6 30 Fig. 1. Conjugation of plasmid and susceptibility profile of resulted transconjugant Transconjugation analysis was performed with E. coli CSH-2 as the recipient by the broth mating method. Transconjugants were selected on LB agar plates supplemented with rifampicin (50 mg/ml) and CTX (20 mg/ml). Plasmid DNA from recipient, donor, and the resulted transconjugant were purified using QIAprep spin plasmid kit (QIAGEN). Figure shows those of No. 179 among four isolates subjected to the transconjugation analysis. The susceptibility profiles of recipient, donor, and the resulted transconjugant were also shown. ESBL phenotypic tests were performed using E test (SYSMEX biomérieux). The susceptibilities of LVFX and TMP/SMX were determined by agar di#usion method according to CLSI recommendation, and categorized into susceptible (S), intermediate (I) and resistant (R) in accordance with CLSI criteria. 30
7 ESBL CTX-M Fig. 2. Dendrogram generated by Pulse-field gel electrophoresis For molecular typing, the chromosomal DNA of fifteen ESBL-producing isolates was obtained for comparison by Pulse-field gel electrophoresis (PFGE) using a CHEF-DRIII (BIO-RAD, Tokyo, Japan). The running parameters were as follows: initial pulse 2.2 s, final pulse 54.2 s, at 6 V/cm for 19 h at 14. The gel was stained with ethidium bromide and scanned. Both E. coli MC4100 and Salmonella H9812 were used as a DNA reference standard. The DNA pattern obtained by PFGE was analyzed with Fingerprinting II (BIO RAD). The unweighted pair group method (UPGMA) was used to generate a dendrogram with the position tolerances set at 1.0, and the Dice coe$cients were used for band similarity measurements to determine DNA relatedness. Isolates were considered to be clonally related if the Dice coe$cient correlation was over 80, which corresponds to the possibly related category according to the criteria suggested by Tenover et al. 26) OUT, outpatient; IN, inpatient. CTX-M ESBL LVFX ESBL O25 : H4 LVFX ST ST131 O25 : H4 5, 22) O25 : H4 ESBL E. coli ESBL 22) ESBL CTX-M-15, ST131 (O25 : H4) E. coli CTX-M-9 E. coli 1, 2, 16) ESBL E. coli CTX-M CTX-M-9 Suzuki 23) CTX-M-9 ESBL 73 11/15 Suzuki 65 84/142 23) Suzuki bla CTX-M-9 bla CTX-M-14 bla CTX-M-27 bla CTX-M-14 bla CTX-M-27 bla CTX-M CAZ 24) bla CTX-M-27 CAZ CAZ O25 : H4 bla CTX-M-27 CTX-M-9 ESBL E. coli 15 O25 : H4 A (No. 45, 79), C (No. 70, 179, 186) PFGE A C A 3 2 C
8 32 Suzuki 2000 CTX-M-9 O 25-ST 131 O 86-ST 38 ESBL E. coli 23) O25 : H4-ST131 PFGE 2 O25 : H4 ESBL ST ESBL ESBL TMP/SMX TMP/ SMX bla CTX-M 1 dfra 25) TMP/SMX 3 ESBL 5 ESBL CTX-M TMP/SMX ST O25 : H4 bla CTX-M-27 CAZ CTX-M ESBL ESBL 1) Bonnet, R Growing group of extendedspectrum b-lactamases: The CTX-M enzymes. Antimicrob. Agents Chemother. 48: ) Pitout, J. J. D., K. B. Laupland Extendedspectrum b-lactamase-producing Enterobacteriaceae: An emerging public-health concern. Lancet Infect. Dis. 8: ) Harada, S., Y. Ishii, K. Yamaguchi Extended-spectrum b-lactamases: Implications for the Clinical Laboratory and Therapy. Korean J. Lab. Med. 28: ) Talbot, G. H., J. Bradley, J. E. Edwards, et al Bad bugs need drugs: An update on the development pipeline from the Antimicrobial Availability Task Force of the Infectious Diseases Society of America. Clin. Infect. Dis. 42: ) Peirano, G., J. D. D. Pitout Molecular epidemiology of Escherichia coli producing CTX- M b-lactamases: The worldwide emergence of clone ST131 O25 : H4. Int. J. Antimicrob. Agents 35: ) Coque, T. M., F. Baquero, R. Canton Increasing prevalence of ESBL-producing Enterobacreriaceae in Europe. Eurosurveillance 13: ) Ensor, V. M., M. Shahid, J. T. Evans, et al Occurrence, prevalence and genetic environment of CTX-M b-lactamases in Enterobacteriaceae from Indian hospitals. J. Amtimcrob. Chemother. 25: ) ) Clinical and Laboratory Standards Institute. CLSI document M7-A7. 26: 2. Clinical and Laboratory Standards Institute, Wayne, PA. 10) Shibata, N., H. Kurokawa, Y. Doi, et al PCR classification of CTX-M-type b-lactamase genes identified in clinically isolated Gramnegative bacilli in Japan. Antimicrob. Agents Chemother. 50: ) Yagi, T., H. Kurokawa, N. Shibata, et al A preliminary survey of extended-spectrum b- lactamases (ESBLs) in clinical isolates of Klebsiella pneumoniae and Escherichia coli in Japan. FEMS Microbiol. Lett. 184:
9 ESBL CTX-M ) Cattoir, V., L. Poirel, V. Rotimi, et al Multiplex PCR for detection plasmid-mediated quinolone resistance qnr genes in ESBLproducing enterobacterial isolates. J. Antimicrob. Chemother. 60: ) Park, C. H., A. Robicsek, G. A. Jacoby, et al Prevalence in the United States of aac- (6 )-Ib-cr encoding a ciprofloxacin-modifying enzyme. Antimicrob. Agents Chemother. 50: ) Yamane, K., J. Wachino, S. Suzuki, et al Plasmid-mediated qepa gene among Escherichia coli clinical isolates from Japan. Antimicrob. Agents Chemother. 52: ) Barrett, T. J., H. Lior, J. H. Green, et al Laboratory investigation of a multistate foodborne outbreak of Escherichia coli O157 : H7 by using pulsed-field gel electrophoresis and phage typing. J. Clin. Microbiol. 32: ) Ray, J. L., K. M. Nielsen Experimental methods for assaying natural transformation and inferring horizontal gene transfer. Methods Enzymol. 395: ) David, N., M. D. Gilbert, C. Robert, et al p , 34 18) Paterson, D. L., R. A. Bonomo Extendedspectrum b-lactamase: A clinical update. Clin. Microbiol. Rev. 18: ) Nys, S., P. H. Terporten, A. A. Hoogkamp- Korstanje, et al Trends in antimicrobial susceptibility of Escherichia coli isolates from urology services in the Netherlands ( ). J. Amtimcrob. Chemother. 62: ) Hirakata, Y., J. Matsuda, Y. Miyazaki, et al Regional variation in the prevalence of extended-spectrum b-lactamase-producing clinical isolates in the Asia-Panpacific region (SENTRY ). Diagn. Microbiol. Infect. Dis. 52: ) Yamaguchi, K., Y. Ishii, M. Iwata, et al Nationwide surveillance of parenteral antibiotics containing meropenem activities against clinical isolates strains in Jpn. J. Antibiot. 60: (Japanese) 22) Lavilla, S., J. J. González-López, M. Sabaté, et al Prevalence of qnr genes among extendedspectrum b-lactamase-producing enterobacterial isolates in Barcelona, Spain. J. Amtimcrob. Chemother. 61: ) Suzuki, S. N. Shibata, K. Yamane, et al Change in the prevalence of extendedspectrum-b-lactamase-producing Escherichia coli in Japan by clonal spread. J. Antimicrob. Chemother. 63: ) Bonnet, R., C. Recule, R. Baraduc, et al E#ect of D240G substitution in a novel ESBL CTX-M-27. J. Antimicrob. Chemother. 52: ) Vinué, L., M. Lantero, Y. Saénz et al Characterization of extended-spectrum-blactamases and integrons in Escherichia coli isolates in a Spanish hospital. J. Med. Microbiol. 57: ) Tenover, F-C., R.-D. Arbeit, R.-V. Goering, et al Interpreting chromosomal DNA restriction patterns produced by pulsed-field gel electrophoresis: Criteria for bacterial strain typing. J. Clin. Microbiol. 33:
10 34 Multidrug-Resistant Phenotype and Prevalence of CTX-M-27 in Extended- Spectrum b-lactamase-producing Uropathogenic Escherichia coli Tatsuya Hattori, 1) Kunihiko Nakane, 1, 2) 1, 3) Kumiko Kawamura 1) Department of Pathophysiological Laboratory Science, Nagoya University Graduate School of Medicine 2) City of Okazaki Reseach Center, Hygiene Testing Group 3) Department of Medical Technology, Nagoya University School of Health Science The susceptibilities against 312 uropathogenic Escherichia coli strains isolated from clinical specimens were investigated, and 15 were extended-spectrum b-lactamase (ESBL)-producing isolates. Among the 15 ESBL-producing isolates, 13 were resistant to levofloxacin (LVFX) and 6 were resistant to both LVFX and trimethoprim/sulfamethoxazole (TMP/SMX), suggesting that ESBL-producing isolates tend to become multidrug-resistant (MDR) phenotypes. The genotypes of ESBL were determined by PCR and sequencing. Four isolates were CTX-M-14 producers and 8 isolates harbored bla CTX-M-27, which harbor the Asp240Gly substitution, have greater catalytic e$ciencies against ceftazidime than parental enzymes. None of these isolates harbored plasmid-mediated quinolone resistance genes, but 6 were serotype O25 : H4 among 13 ESBL-producing and LVFX-resistant isolates, suggesting fluoroquinolone-resistant phenotypes are associated with serotype O25. The CTX-M encoding plasmid was successfully transferred to recipient E. coli CSH-2 at a frequency of cells per recipient by conjugation in vitro, suggesting the easy dissemination of bla CTX-M -harboring plasmids. Moreover, the transconjugants obtained showed both ESBL-producing and TMP/SMX-resistant phenotypes. Among uropathogenic E. coli, the serotype O25 : H4 isolates harboring CTX-M-27 bla CTX-M have become prevalent strains, so due care should be taken in monitoring the epidemiology of ESBL-producing isolates. 34
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
Y-chromosomal haplogroup typing Using SBE reaction
 Schematic of multiplex PCR followed by SBE reaction Multiplex PCR Exo SAP purification SBE reaction 5 A 3 ddatp ddgtp 3 T 5 A G 3 T 5 3 5 G C 5 3 3 C 5 ddttp ddctp 5 T 3 T C 3 A 5 3 A 5 5 C 3 3 G 5 3 G
Schematic of multiplex PCR followed by SBE reaction Multiplex PCR Exo SAP purification SBE reaction 5 A 3 ddatp ddgtp 3 T 5 A G 3 T 5 3 5 G C 5 3 3 C 5 ddttp ddctp 5 T 3 T C 3 A 5 3 A 5 5 C 3 3 G 5 3 G
Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR
 Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR 1 The problem We wish to clone a yet unknown gene from a known
Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR 1 The problem We wish to clone a yet unknown gene from a known
Supporting Information. Copyright Wiley-VCH Verlag GmbH & Co. KGaA, Weinheim, 2006
 Supporting Information Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Supporting Information for Expanding the Genetic
Supporting Information Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Supporting Information for Expanding the Genetic
Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH).
 Bisulfite Treatment of DNA Dilute DNA sample to 2µg DNA in 50µl ddh 2 O. Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH). Incubate in a 37ºC water bath for 30 minutes. To 55µl samples
Bisulfite Treatment of DNA Dilute DNA sample to 2µg DNA in 50µl ddh 2 O. Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH). Incubate in a 37ºC water bath for 30 minutes. To 55µl samples
Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis
 1 2 3 4 5 6 7 8 9 10 11 12 Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis Information Research). Exons
1 2 3 4 5 6 7 8 9 10 11 12 Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis Information Research). Exons
Arabidopsis actin depolymerizing factor AtADF4 mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB
 Arabidopsis actin depolymerizing factor mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB Files in this Data Supplement: Supplemental Table S1 Supplemental Table
Arabidopsis actin depolymerizing factor mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB Files in this Data Supplement: Supplemental Table S1 Supplemental Table
Supplementary. Table 1: Oligonucleotides and Plasmids. complementary to positions from 77 of the SRα '- GCT CTA GAG AAC TTG AAG TAC AGA CTG C
 Supplementary Table 1: Oligonucleotides and Plasmids 913954 5'- GCT CTA GAG AAC TTG AAG TAC AGA CTG C 913955 5'- CCC AAG CTT ACA GTG TGG CCA TTC TGC TG 223396 5'- CGA CGC GTA CAG TGT GGC CAT TCT GCT G
Supplementary Table 1: Oligonucleotides and Plasmids 913954 5'- GCT CTA GAG AAC TTG AAG TAC AGA CTG C 913955 5'- CCC AAG CTT ACA GTG TGG CCA TTC TGC TG 223396 5'- CGA CGC GTA CAG TGT GGC CAT TCT GCT G
Supplemental Data Supplemental Figure 1.
 Supplemental Data Supplemental Figure 1. Silique arrangement in the wild-type, jhs, and complemented lines. Wild-type (WT) (A), the jhs1 mutant (B,C), and the jhs1 mutant complemented with JHS1 (Com) (D)
Supplemental Data Supplemental Figure 1. Silique arrangement in the wild-type, jhs, and complemented lines. Wild-type (WT) (A), the jhs1 mutant (B,C), and the jhs1 mutant complemented with JHS1 (Com) (D)
Cat. # Product Size DS130 DynaExpress TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1
 Product Name: Kit Component TA PCR Cloning Kit (ptakn-2) Cat. # Product Size DS130 TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1 2 Ligation Buffer
Product Name: Kit Component TA PCR Cloning Kit (ptakn-2) Cat. # Product Size DS130 TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1 2 Ligation Buffer
Table S1. Bacterial strains (Related to Results and Experimental Procedures)
 Table S1. Bacterial strains (Related to Results and Experimental Procedures) Strain number Relevant genotype Source or reference 1045 AB1157 Graham Walker (Donnelly and Walker, 1989) 2458 3084 (MG1655)
Table S1. Bacterial strains (Related to Results and Experimental Procedures) Strain number Relevant genotype Source or reference 1045 AB1157 Graham Walker (Donnelly and Walker, 1989) 2458 3084 (MG1655)
Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC
 Supplementary Appendixes Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC ACG TAG CTC CGG CTG GA-3 for vimentin, /5AmMC6/TCC CTC GCG CGT GGC TTC CGC
Supplementary Appendixes Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC ACG TAG CTC CGG CTG GA-3 for vimentin, /5AmMC6/TCC CTC GCG CGT GGC TTC CGC
Supporting information for Biochemistry, 1995, 34(34), , DOI: /bi00034a013
 Supporting information for Biochemistry, 1995, 34(34), 10807 10815, DOI: 10.1021/bi00034a013 LESNIK 10807-1081 Terms & Conditions Electronic Supporting Information files are available without a subscription
Supporting information for Biochemistry, 1995, 34(34), 10807 10815, DOI: 10.1021/bi00034a013 LESNIK 10807-1081 Terms & Conditions Electronic Supporting Information files are available without a subscription
Supplemental material
 Supplemental material Diversity of O-antigen repeat-unit structures can account for the substantial sequence variation of Wzx translocases Yaoqin Hong and Peter R. Reeves School of Molecular Bioscience,
Supplemental material Diversity of O-antigen repeat-unit structures can account for the substantial sequence variation of Wzx translocases Yaoqin Hong and Peter R. Reeves School of Molecular Bioscience,
Hes6. PPARα. PPARγ HNF4 CD36
 SUPPLEMENTARY INFORMATION Supplementary Table Positions and Sequences of ChIP primers -63 AGGTCACTGCCA -79 AGGTCTGCTGTG Hes6-0067 GGGCAaAGTTCA ACOT -395 GGGGCAgAGTTCA PPARα -309 GGCTCAaAGTTCAaGTTCA CPTa
SUPPLEMENTARY INFORMATION Supplementary Table Positions and Sequences of ChIP primers -63 AGGTCACTGCCA -79 AGGTCTGCTGTG Hes6-0067 GGGCAaAGTTCA ACOT -395 GGGGCAgAGTTCA PPARα -309 GGCTCAaAGTTCAaGTTCA CPTa
Gene synthesis by circular assembly amplification
 Gene synthesis by circular assembly amplification Duhee Bang & George M Church Supplementary figures and text: Supplementary Figure 1. Dpo4 gene (1.05kb) construction by various methods. Supplementary
Gene synthesis by circular assembly amplification Duhee Bang & George M Church Supplementary figures and text: Supplementary Figure 1. Dpo4 gene (1.05kb) construction by various methods. Supplementary
strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular
![strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular](/thumbs/72/66361212.jpg) Additional file 2 Identification of AOX1 in P. pastoris GS115 with a Mut s phenotype Results and Discussion The HBsAg producing strain was originally identified as a Mut s (methanol utilization slow) strain
Additional file 2 Identification of AOX1 in P. pastoris GS115 with a Mut s phenotype Results and Discussion The HBsAg producing strain was originally identified as a Mut s (methanol utilization slow) strain
Disease and selection in the human genome 3
 Disease and selection in the human genome 3 Ka/Ks revisited Please sit in row K or forward RBFD: human populations, adaptation and immunity Neandertal Museum, Mettman Germany Sequence genome Measure expression
Disease and selection in the human genome 3 Ka/Ks revisited Please sit in row K or forward RBFD: human populations, adaptation and immunity Neandertal Museum, Mettman Germany Sequence genome Measure expression
PCR analysis was performed to show the presence and the integrity of the var1csa and var-
 Supplementary information: Methods: Table S1: Primer Name Nucleotide sequence (5-3 ) DBL3-F tcc ccg cgg agt gaa aca tca tgt gac tg DBL3-R gac tag ttt ctt tca ata aat cac tcg c DBL5-F cgc cct agg tgc ttc
Supplementary information: Methods: Table S1: Primer Name Nucleotide sequence (5-3 ) DBL3-F tcc ccg cgg agt gaa aca tca tgt gac tg DBL3-R gac tag ttt ctt tca ata aat cac tcg c DBL5-F cgc cct agg tgc ttc
PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells
 Supplementary Information for: PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells Ju Hye Jang 1, Hyun Kim 2, Mi Jung Jang 2, Ju Hyun Cho 1,2,* 1 Research Institute
Supplementary Information for: PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells Ju Hye Jang 1, Hyun Kim 2, Mi Jung Jang 2, Ju Hyun Cho 1,2,* 1 Research Institute
Supplementary Figure 1A A404 Cells +/- Retinoic Acid
 Supplementary Figure 1A A44 Cells +/- Retinoic Acid 1 1 H3 Lys4 di-methylation SM-actin VEC cfos (-) RA (+) RA 14 1 1 8 6 4 H3 Lys79 di-methylation SM-actin VEC cfos (-) RA (+) RA Supplementary Figure
Supplementary Figure 1A A44 Cells +/- Retinoic Acid 1 1 H3 Lys4 di-methylation SM-actin VEC cfos (-) RA (+) RA 14 1 1 8 6 4 H3 Lys79 di-methylation SM-actin VEC cfos (-) RA (+) RA Supplementary Figure
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/10/494/eaan6284/dc1 Supplementary Materials for Activation of master virulence regulator PhoP in acidic ph requires the Salmonella-specific protein UgtL Jeongjoon
www.sciencesignaling.org/cgi/content/full/10/494/eaan6284/dc1 Supplementary Materials for Activation of master virulence regulator PhoP in acidic ph requires the Salmonella-specific protein UgtL Jeongjoon
II 0.95 DM2 (RPP1) DM3 (At3g61540) b
 Table S2. F 2 Segregation Ratios at 16 C, Related to Figure 2 Cross n c Phenotype Model e 2 Locus A Locus B Normal F 1 -like Enhanced d Uk-1/Uk-3 149 64 36 49 DM2 (RPP1) DM1 (SSI4) a Bla-1/Hh-0 F 3 111
Table S2. F 2 Segregation Ratios at 16 C, Related to Figure 2 Cross n c Phenotype Model e 2 Locus A Locus B Normal F 1 -like Enhanced d Uk-1/Uk-3 149 64 36 49 DM2 (RPP1) DM1 (SSI4) a Bla-1/Hh-0 F 3 111
Supplemental Data. mir156-regulated SPL Transcription. Factors Define an Endogenous Flowering. Pathway in Arabidopsis thaliana
 Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
ORFs and genes. Please sit in row K or forward
 ORFs and genes Please sit in row K or forward https://www.flickr.com/photos/teseum/3231682806/in/photostream/ Question: why do some strains of Vibrio cause cholera and others don t? Methods Mechanisms
ORFs and genes Please sit in row K or forward https://www.flickr.com/photos/teseum/3231682806/in/photostream/ Question: why do some strains of Vibrio cause cholera and others don t? Methods Mechanisms
Materials Protein synthesis kit. This kit consists of 24 amino acids, 24 transfer RNAs, four messenger RNAs and one ribosome (see below).
 Protein Synthesis Instructions The purpose of today s lab is to: Understand how a cell manufactures proteins from amino acids, using information stored in the genetic code. Assemble models of four very
Protein Synthesis Instructions The purpose of today s lab is to: Understand how a cell manufactures proteins from amino acids, using information stored in the genetic code. Assemble models of four very
ΔPDD1 x ΔPDD1. ΔPDD1 x wild type. 70 kd Pdd1. Pdd3
 Supplemental Fig. S1 ΔPDD1 x wild type ΔPDD1 x ΔPDD1 70 kd Pdd1 50 kd 37 kd Pdd3 Supplemental Fig. S1. ΔPDD1 strains express no detectable Pdd1 protein. Western blot analysis of whole-protein extracts
Supplemental Fig. S1 ΔPDD1 x wild type ΔPDD1 x ΔPDD1 70 kd Pdd1 50 kd 37 kd Pdd3 Supplemental Fig. S1. ΔPDD1 strains express no detectable Pdd1 protein. Western blot analysis of whole-protein extracts
Multiplexing Genome-scale Engineering
 Multiplexing Genome-scale Engineering Harris Wang, Ph.D. Department of Systems Biology Department of Pathology & Cell Biology http://wanglab.c2b2.columbia.edu Rise of Genomics An Expanding Toolbox Esvelt
Multiplexing Genome-scale Engineering Harris Wang, Ph.D. Department of Systems Biology Department of Pathology & Cell Biology http://wanglab.c2b2.columbia.edu Rise of Genomics An Expanding Toolbox Esvelt
Overexpression Normal expression Overexpression Normal expression. 26 (21.1%) N (%) P-value a N (%)
 SUPPLEMENTARY TABLES Table S1. Alteration of ZNF322A protein expression levels in relation to clinicopathological parameters in 123 Asian and 74 Caucasian lung cancer patients. Asian patients Caucasian
SUPPLEMENTARY TABLES Table S1. Alteration of ZNF322A protein expression levels in relation to clinicopathological parameters in 123 Asian and 74 Caucasian lung cancer patients. Asian patients Caucasian
Supplementary Information. Construction of Lasso Peptide Fusion Proteins
 Supplementary Information Construction of Lasso Peptide Fusion Proteins Chuhan Zong 1, Mikhail O. Maksimov 2, A. James Link 2,3 * Departments of 1 Chemistry, 2 Chemical and Biological Engineering, and
Supplementary Information Construction of Lasso Peptide Fusion Proteins Chuhan Zong 1, Mikhail O. Maksimov 2, A. James Link 2,3 * Departments of 1 Chemistry, 2 Chemical and Biological Engineering, and
Supplemental Data. Bennett et al. (2010). Plant Cell /tpc
 BRN1 ---------MSSSNGGVPPGFRFHPTDEELLHYYLKKKISYEKFEMEVIKEVDLNKIEPWDLQDRCKIGSTPQNEWYFFSHKDRKYPTGS 81 BRN2 --------MGSSSNGGVPPGFRFHPTDEELLHYYLKKKISYQKFEMEVIREVDLNKLEPWDLQERCKIGSTPQNEWYFFSHKDRKYPTGS 82 SMB
BRN1 ---------MSSSNGGVPPGFRFHPTDEELLHYYLKKKISYEKFEMEVIKEVDLNKIEPWDLQDRCKIGSTPQNEWYFFSHKDRKYPTGS 81 BRN2 --------MGSSSNGGVPPGFRFHPTDEELLHYYLKKKISYQKFEMEVIREVDLNKLEPWDLQERCKIGSTPQNEWYFFSHKDRKYPTGS 82 SMB
PCR-based Markers and Cut Flower Longevity in Carnation
 PCRbased Markers and Cut Flower Longevity in Carnation Laura De Benedetti, Luca Braglia, Simona Bruna, Gianluca Burchi *, Antonio Mercuri and Tito Schiva Istituto Sperimentale per la Floricoltura, Corso
PCRbased Markers and Cut Flower Longevity in Carnation Laura De Benedetti, Luca Braglia, Simona Bruna, Gianluca Burchi *, Antonio Mercuri and Tito Schiva Istituto Sperimentale per la Floricoltura, Corso
MacBlunt PCR Cloning Kit Manual
 MacBlunt PCR Cloning Kit Manual Shipping and Storage MacBlunt PCR Cloning Kits are shipped on dry ice. Each kit contains a box with cloning reagents and an attached bag with Eco-Blue Competent Cells (optional).
MacBlunt PCR Cloning Kit Manual Shipping and Storage MacBlunt PCR Cloning Kits are shipped on dry ice. Each kit contains a box with cloning reagents and an attached bag with Eco-Blue Competent Cells (optional).
NAME:... MODEL ANSWER... STUDENT NUMBER:... Maximum marks: 50. Internal Examiner: Hugh Murrell, Computer Science, UKZN
 COMP710, Bioinformatics with Julia, Test One, Thursday the 20 th of April, 2017, 09h30-11h30 1 NAME:...... MODEL ANSWER... STUDENT NUMBER:...... Maximum marks: 50 Internal Examiner: Hugh Murrell, Computer
COMP710, Bioinformatics with Julia, Test One, Thursday the 20 th of April, 2017, 09h30-11h30 1 NAME:...... MODEL ANSWER... STUDENT NUMBER:...... Maximum marks: 50 Internal Examiner: Hugh Murrell, Computer
Supporting Information
 Supporting Information Table S1. Oligonucleotide sequences used in this work Oligo DNA A B C D CpG-A CpG-B CpG-C CpG-D Sequence 5 ACA TTC CTA AGT CTG AAA CAT TAC AGC TTG CTA CAC GAG AAG AGC CGC CAT AGT
Supporting Information Table S1. Oligonucleotide sequences used in this work Oligo DNA A B C D CpG-A CpG-B CpG-C CpG-D Sequence 5 ACA TTC CTA AGT CTG AAA CAT TAC AGC TTG CTA CAC GAG AAG AGC CGC CAT AGT
Supporting Online Information
 Supporting Online Information Isolation of Human Genomic DNA Sequences with Expanded Nucleobase Selectivity Preeti Rathi, Sara Maurer, Grzegorz Kubik and Daniel Summerer* Department of Chemistry and Chemical
Supporting Online Information Isolation of Human Genomic DNA Sequences with Expanded Nucleobase Selectivity Preeti Rathi, Sara Maurer, Grzegorz Kubik and Daniel Summerer* Department of Chemistry and Chemical
Legends for supplementary figures 1-3
 High throughput resistance profiling of Plasmodium falciparum infections based on custom dual indexing and Illumina next generation sequencing-technology Sidsel Nag 1,2 *, Marlene D. Dalgaard 3, Poul-Erik
High throughput resistance profiling of Plasmodium falciparum infections based on custom dual indexing and Illumina next generation sequencing-technology Sidsel Nag 1,2 *, Marlene D. Dalgaard 3, Poul-Erik
SUPPLEMENTARY MATERIALS AND METHODS. E. coli strains, plasmids, and growth conditions. Escherichia coli strain P90C (1)
 SUPPLEMENTARY MATERIALS AND METHODS E. coli strains, plasmids, and growth conditions. Escherichia coli strain P90C (1) dinb::kan (lab stock) derivative was used as wild-type. MG1655 alka tag dinb (2) is
SUPPLEMENTARY MATERIALS AND METHODS E. coli strains, plasmids, and growth conditions. Escherichia coli strain P90C (1) dinb::kan (lab stock) derivative was used as wild-type. MG1655 alka tag dinb (2) is
evaluated with UAS CLB eliciting UAS CIT -N Libraries increase in the
 Supplementary Figures Supplementary Figure 1: Promoter scaffold library assemblies. Many ensembless of libraries were evaluated in this work. As a legend, the box outline color in top half of the figure
Supplementary Figures Supplementary Figure 1: Promoter scaffold library assemblies. Many ensembless of libraries were evaluated in this work. As a legend, the box outline color in top half of the figure
SUPPORTING INFORMATION
 SUPPORTING INFORMATION Investigation of the Biosynthesis of the Lasso Peptide Chaxapeptin Using an E. coli-based Production System Helena Martin-Gómez, Uwe Linne, Fernando Albericio, Judit Tulla-Puche,*
SUPPORTING INFORMATION Investigation of the Biosynthesis of the Lasso Peptide Chaxapeptin Using an E. coli-based Production System Helena Martin-Gómez, Uwe Linne, Fernando Albericio, Judit Tulla-Puche,*
Dierks Supplementary Fig. S1
 Dierks Supplementary Fig. S1 ITK SYK PH TH K42R wt K42R (kinase deficient) R29C E42K Y323F R29C E42K Y323F (reduced phospholipid binding) (enhanced phospholipid binding) (reduced Cbl binding) E42K Y323F
Dierks Supplementary Fig. S1 ITK SYK PH TH K42R wt K42R (kinase deficient) R29C E42K Y323F R29C E42K Y323F (reduced phospholipid binding) (enhanced phospholipid binding) (reduced Cbl binding) E42K Y323F
Search for and Analysis of Single Nucleotide Polymorphisms (SNPs) in Rice (Oryza sativa, Oryza rufipogon) and Establishment of SNP Markers
 DNA Research 9, 163 171 (2002) Search for and Analysis of Single Nucleotide Polymorphisms (SNPs) in Rice (Oryza sativa, Oryza rufipogon) and Establishment of SNP Markers Shinobu Nasu, Junko Suzuki, Rieko
DNA Research 9, 163 171 (2002) Search for and Analysis of Single Nucleotide Polymorphisms (SNPs) in Rice (Oryza sativa, Oryza rufipogon) and Establishment of SNP Markers Shinobu Nasu, Junko Suzuki, Rieko
Supplementary Figures
 Supplementary Figures Supplementary Fig. 1 Characterization of GSCs. a. Immunostaining of primary GSC spheres from GSC lines. Nestin (neural progenitor marker, red), TLX (green). Merged images of nestin,
Supplementary Figures Supplementary Fig. 1 Characterization of GSCs. a. Immunostaining of primary GSC spheres from GSC lines. Nestin (neural progenitor marker, red), TLX (green). Merged images of nestin,
Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were
 1 Supplemental methods 2 3 4 5 6 7 8 9 1 11 12 13 14 15 16 17 18 19 21 22 23 Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were monitored by quantitative reverse-transcription
1 Supplemental methods 2 3 4 5 6 7 8 9 1 11 12 13 14 15 16 17 18 19 21 22 23 Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were monitored by quantitative reverse-transcription
Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system
 Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system Dr. Tim Welsink Molecular Biology Transient Gene Expression OUTLINE Short
Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system Dr. Tim Welsink Molecular Biology Transient Gene Expression OUTLINE Short
11th Meeting of the Science Working Group. Lima, Peru, October 2012 SWG-11-JM-11
 11th Meeting of the Science Working Group Lima, Peru, 15-19 October 2012 Russian population genetics studies of jack mackerel in the South Pacific P.K.Afanasiev M.A.Rabchun A.I.Glubokov Introduction. In
11th Meeting of the Science Working Group Lima, Peru, 15-19 October 2012 Russian population genetics studies of jack mackerel in the South Pacific P.K.Afanasiev M.A.Rabchun A.I.Glubokov Introduction. In
NESTED Sequence-based Typing (SBT) protocol for epidemiological typing of Legionella pneumophila directly from clinical samples
 NESTED Sequence-based Typing (SBT) protocol for epidemiological typing of Legionella pneumophila directly from clinical samples VERSION 2.0 SUMMARY This procedure describes the use of nested Sequence-Based
NESTED Sequence-based Typing (SBT) protocol for epidemiological typing of Legionella pneumophila directly from clinical samples VERSION 2.0 SUMMARY This procedure describes the use of nested Sequence-Based
Supplemental Table 1. Primers used for PCR.
 Supplemental Table 1. Primers used for PCR. Gene Type Primer Sequence Genotyping and semi-quantitative RT-PCR F 5 -TTG CCC GAT CAC CAT CTG TA-3 rwa1-1 R 5 -TGT AGC GAT CAA GGC CTG ATC TAA-3 LB 5 -TAG CAT
Supplemental Table 1. Primers used for PCR. Gene Type Primer Sequence Genotyping and semi-quantitative RT-PCR F 5 -TTG CCC GAT CAC CAT CTG TA-3 rwa1-1 R 5 -TGT AGC GAT CAA GGC CTG ATC TAA-3 LB 5 -TAG CAT
Supporting Information
 Supporting Information Transfection of DNA Cages into Mammalian Cells Email: a.turberfield@physics.ox.ac.uk Table of Contents Supporting Figure 1 DNA tetrahedra used in transfection experiments 2 Supporting
Supporting Information Transfection of DNA Cages into Mammalian Cells Email: a.turberfield@physics.ox.ac.uk Table of Contents Supporting Figure 1 DNA tetrahedra used in transfection experiments 2 Supporting
S4B fluorescence (AU)
 A S4B fluorescence (AU) S4B fluorescence (AU) dsbb csgba csgd dsbb csgba bcsa 5000 * NS NS 4000 * 3000 2000 1000 0 ΔcsgBAΔbcsA ΔcsgDΔdsbBΔbcsA ΔcsgBA ΔdsbBΔcsgBA ΔcsgDΔdsbB B -1000 4000 * * NS 3500 * 3000
A S4B fluorescence (AU) S4B fluorescence (AU) dsbb csgba csgd dsbb csgba bcsa 5000 * NS NS 4000 * 3000 2000 1000 0 ΔcsgBAΔbcsA ΔcsgDΔdsbBΔbcsA ΔcsgBA ΔdsbBΔcsgBA ΔcsgDΔdsbB B -1000 4000 * * NS 3500 * 3000
Lecture 11: Gene Prediction
 Lecture 11: Gene Prediction Study Chapter 6.11-6.14 1 Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and end positions of genes in a genome Where are
Lecture 11: Gene Prediction Study Chapter 6.11-6.14 1 Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and end positions of genes in a genome Where are
RPA-AB RPA-C Supplemental Figure S1: SDS-PAGE stained with Coomassie Blue after protein purification.
 RPA-AB RPA-C (a) (b) (c) (d) (e) (f) Supplemental Figure S: SDS-PAGE stained with Coomassie Blue after protein purification. (a) RPA; (b) RPA-AB; (c) RPA-CDE; (d) RPA-CDE core; (e) RPA-DE; and (f) RPA-C
RPA-AB RPA-C (a) (b) (c) (d) (e) (f) Supplemental Figure S: SDS-PAGE stained with Coomassie Blue after protein purification. (a) RPA; (b) RPA-AB; (c) RPA-CDE; (d) RPA-CDE core; (e) RPA-DE; and (f) RPA-C
G+C content. 1 Introduction. 2 Chromosomes Topology & Counts. 3 Genome size. 4 Replichores and gene orientation. 5 Chirochores.
 1 Introduction 2 Chromosomes Topology & Counts 3 Genome size 4 Replichores and gene orientation 5 Chirochores 6 7 Codon usage 121 marc.bailly-bechet@univ-lyon1.fr Bacterial genome structures Introduction
1 Introduction 2 Chromosomes Topology & Counts 3 Genome size 4 Replichores and gene orientation 5 Chirochores 6 7 Codon usage 121 marc.bailly-bechet@univ-lyon1.fr Bacterial genome structures Introduction
Primer Design Workshop. École d'été en géné-que des champignons 2012 Dr. Will Hintz University of Victoria
 Primer Design Workshop École d'été en géné-que des champignons 2012 Dr. Will Hintz University of Victoria Scenario You have discovered the presence of a novel endophy5c organism living inside the cells
Primer Design Workshop École d'été en géné-que des champignons 2012 Dr. Will Hintz University of Victoria Scenario You have discovered the presence of a novel endophy5c organism living inside the cells
hcd1tg/hj1tg/ ApoE-/- hcd1tg/hj1tg/ ApoE+/+
 ApoE+/+ ApoE-/- ApoE-/- H&E (1x) Supplementary Figure 1. No obvious pathology is observed in the colon of diseased ApoE-/me. Colon samples were fixed in 1% formalin and laid out in Swiss rolls for paraffin
ApoE+/+ ApoE-/- ApoE-/- H&E (1x) Supplementary Figure 1. No obvious pathology is observed in the colon of diseased ApoE-/me. Colon samples were fixed in 1% formalin and laid out in Swiss rolls for paraffin
Supplemental Table 1. Mutant ADAMTS3 alleles detected in HEK293T clone 4C2. WT CCTGTCACTTTGGTTGATAGC MVLLSLWLIAAALVEVR
 Supplemental Dataset Supplemental Table 1. Mutant ADAMTS3 alleles detected in HEK293T clone 4C2. DNA sequence Amino acid sequence WT CCTGTCACTTTGGTTGATAGC MVLLSLWLIAAALVEVR Allele 1 CCTGTC------------------GATAGC
Supplemental Dataset Supplemental Table 1. Mutant ADAMTS3 alleles detected in HEK293T clone 4C2. DNA sequence Amino acid sequence WT CCTGTCACTTTGGTTGATAGC MVLLSLWLIAAALVEVR Allele 1 CCTGTC------------------GATAGC
Project 07/111 Final Report October 31, Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines
 Project 07/111 Final Report October 31, 2007. Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines Project Leader: Dr Douglas C. Hodgins (519-824-4120 Ex 54758, fax 519-824-5930)
Project 07/111 Final Report October 31, 2007. Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines Project Leader: Dr Douglas C. Hodgins (519-824-4120 Ex 54758, fax 519-824-5930)
2
 1 2 3 4 5 6 7 Supplemental Table 1. Magnaporthe oryzae strains generated in this study. Strain background Genotype Strain name Description Guy-11 H1:RFP H1:RFP Strain expressing Histone H1- encoding gene
1 2 3 4 5 6 7 Supplemental Table 1. Magnaporthe oryzae strains generated in this study. Strain background Genotype Strain name Description Guy-11 H1:RFP H1:RFP Strain expressing Histone H1- encoding gene
Supplemental Information. Human Senataxin Resolves RNA/DNA Hybrids. Formed at Transcriptional Pause Sites. to Promote Xrn2-Dependent Termination
 Supplemental Information Molecular Cell, Volume 42 Human Senataxin Resolves RNA/DNA Hybrids Formed at Transcriptional Pause Sites to Promote Xrn2-Dependent Termination Konstantina Skourti-Stathaki, Nicholas
Supplemental Information Molecular Cell, Volume 42 Human Senataxin Resolves RNA/DNA Hybrids Formed at Transcriptional Pause Sites to Promote Xrn2-Dependent Termination Konstantina Skourti-Stathaki, Nicholas
SAY IT WITH DNA: Protein Synthesis Activity by Larry Flammer
 TEACHER S GUIDE SAY IT WITH DNA: Protein Synthesis Activity by Larry Flammer SYNOPSIS This activity uses the metaphor of decoding a secret message for the Protein Synthesis process. Students teach themselves
TEACHER S GUIDE SAY IT WITH DNA: Protein Synthesis Activity by Larry Flammer SYNOPSIS This activity uses the metaphor of decoding a secret message for the Protein Synthesis process. Students teach themselves
Keywords: qepa, qnrs1, rmtb, bla LAP-1, ISCR3C, gene co-transferance. Introduction
 Available online at www.annclinlabsci.org Annals of Clinical & Laboratory Science, vol. 39, no. 1, 2009 55 Accumulation of Plasmid-Mediated Fluoroquinolone Resistance Genes, qepa and qnrs1, in Enterobacter
Available online at www.annclinlabsci.org Annals of Clinical & Laboratory Science, vol. 39, no. 1, 2009 55 Accumulation of Plasmid-Mediated Fluoroquinolone Resistance Genes, qepa and qnrs1, in Enterobacter
MCB421 FALL2005 EXAM#3 ANSWERS Page 1 of 12. ANSWER: Both transposon types form small duplications of adjacent host DNA sequences.
 Page 1 of 12 (10pts) 1. There are two mechanisms for transposition used by bacterial transposable elements: replicative (Tn3) and non-replicative (Tn5 and Tn10). Compare and contrast the two mechanisms
Page 1 of 12 (10pts) 1. There are two mechanisms for transposition used by bacterial transposable elements: replicative (Tn3) and non-replicative (Tn5 and Tn10). Compare and contrast the two mechanisms
SUPPLEMENTAL MATERIAL GENOTYPING WITH MULTIPLEXING TARGETED RESEQUENCING
 SUPPLEMENTAL MATERIAL GENOTYPING WITH MULTIPLEXING TARGETED RESEQUENCING All of the patients and control subjects were sequenced and genotyped in the same way. Shotgun libraries of approximately 250 bp
SUPPLEMENTAL MATERIAL GENOTYPING WITH MULTIPLEXING TARGETED RESEQUENCING All of the patients and control subjects were sequenced and genotyped in the same way. Shotgun libraries of approximately 250 bp
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature07182 SUPPLEMENTAL FIGURES AND TABLES Fig. S1. myf5-expressing cells give rise to brown fat depots and skeletal muscle (a) Perirenal BAT from control (cre negative) and myf5-cre:r26r3-yfp
doi: 10.1038/nature07182 SUPPLEMENTAL FIGURES AND TABLES Fig. S1. myf5-expressing cells give rise to brown fat depots and skeletal muscle (a) Perirenal BAT from control (cre negative) and myf5-cre:r26r3-yfp
Supplementary Methods Quantitative RT-PCR. For mrna, total RNA was prepared using TRIzol reagent (Invitrogen) and genomic DNA was eliminated with TURB
 Supplementary Methods Quantitative RT-PCR. For mrna, total RNA was prepared using TRIzol reagent (Invitrogen) and genomic DNA was eliminated with TURBO DNA-free Kit (Ambion). One µg of total RNA was reverse
Supplementary Methods Quantitative RT-PCR. For mrna, total RNA was prepared using TRIzol reagent (Invitrogen) and genomic DNA was eliminated with TURBO DNA-free Kit (Ambion). One µg of total RNA was reverse
SUMMARY. Key words: antibioticresistance, Enterobacteriaceae, ESBL, CTX-M,
 SUMMARY Key words: antibioticresistance, Enterobacteriaceae, ESBL, CTX-M, The doctoral thesis entitled Prevalence of Enterobacteriaceae producing extendedspectrum beta-lactamases (ESBL) isolated from broilers
SUMMARY Key words: antibioticresistance, Enterobacteriaceae, ESBL, CTX-M, The doctoral thesis entitled Prevalence of Enterobacteriaceae producing extendedspectrum beta-lactamases (ESBL) isolated from broilers
Appendix 1a. Microsatellite analysis of P1-hyg, P2-neo and their. Amplified cdna (base pairs) P1-hyg P2-neo All progeny A A
 Supplementary information Appendix 1a. Microsatellite analysis of P1-hyg, P2-neo and their double drug resistant progeny (ABI 377). Primer set a,b Amplified cdna (base pairs) P1-hyg P2-neo All progeny
Supplementary information Appendix 1a. Microsatellite analysis of P1-hyg, P2-neo and their double drug resistant progeny (ABI 377). Primer set a,b Amplified cdna (base pairs) P1-hyg P2-neo All progeny
Lecture 19A. DNA computing
 Lecture 19A. DNA computing What exactly is DNA (deoxyribonucleic acid)? DNA is the material that contains codes for the many physical characteristics of every living creature. Your cells use different
Lecture 19A. DNA computing What exactly is DNA (deoxyribonucleic acid)? DNA is the material that contains codes for the many physical characteristics of every living creature. Your cells use different
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Xie YL, Chakravorty S, Armstrong DT, et al. Evaluation of a
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Xie YL, Chakravorty S, Armstrong DT, et al. Evaluation of a
Complexity of the Ruminococcus flavefaciens FD-1 cellulosome reflects an expansion of family-related protein-protein interactions
 Complexity of the Ruminococcus flavefaciens FD-1 cellulosome reflects an expansion of family-related protein-protein interactions Vered Israeli-Ruimy 1,*, Pedro Bule 2,*, Sadanari Jindou 3, Bareket Dassa
Complexity of the Ruminococcus flavefaciens FD-1 cellulosome reflects an expansion of family-related protein-protein interactions Vered Israeli-Ruimy 1,*, Pedro Bule 2,*, Sadanari Jindou 3, Bareket Dassa
Homework. A bit about the nature of the atoms of interest. Project. The role of electronega<vity
 Homework Why cited articles are especially useful. citeulike science citation index When cutting and pasting less is more. Project Your protein: I will mail these out this weekend If you haven t gotten
Homework Why cited articles are especially useful. citeulike science citation index When cutting and pasting less is more. Project Your protein: I will mail these out this weekend If you haven t gotten
Supplementary Information
 Supplementary Information A general solution for opening double-stranded DNA for isothermal amplification Gangyi Chen, Juan Dong, Yi Yuan, Na Li, Xin Huang, Xin Cui* and Zhuo Tang* Supplementary Materials
Supplementary Information A general solution for opening double-stranded DNA for isothermal amplification Gangyi Chen, Juan Dong, Yi Yuan, Na Li, Xin Huang, Xin Cui* and Zhuo Tang* Supplementary Materials
Supporting Information
 Supporting Information Barderas et al. 10.1073/pnas.0801221105 SI Text: Docking of gastrin to Constructed scfv Models Interactive predocking of the 4-WL-5 motif into the central pocket observed in the
Supporting Information Barderas et al. 10.1073/pnas.0801221105 SI Text: Docking of gastrin to Constructed scfv Models Interactive predocking of the 4-WL-5 motif into the central pocket observed in the
SUPPLEMENTAL TABLE S1. Additional descriptions of plasmid constructions and the oligonucleotides used Plasmid or Oligonucleotide
 SUPPLEMENTAL TABLE S1. Additional descriptions of plasmid constructions and the oligonucleotides used Plasmid or Oligonucleotide former/ working Description a designation Plasmids pes213a b pes213-tn5
SUPPLEMENTAL TABLE S1. Additional descriptions of plasmid constructions and the oligonucleotides used Plasmid or Oligonucleotide former/ working Description a designation Plasmids pes213a b pes213-tn5
for Programmed Chemo-enzymatic Synthesis of Antigenic Oligosaccharides
 Supporting Information Design of α-transglucosidases of Controlled Specificity for Programmed Chemo-enzymatic Synthesis of Antigenic Oligosaccharides Elise Champion ±,,,, Isabelle André ±,,, Claire Moulis
Supporting Information Design of α-transglucosidases of Controlled Specificity for Programmed Chemo-enzymatic Synthesis of Antigenic Oligosaccharides Elise Champion ±,,,, Isabelle André ±,,, Claire Moulis
An engineered tryptophan zipper-type peptide as a molecular recognition scaffold
 SUPPLEMENTARY MATERIAL An engineered tryptophan zipper-type peptide as a molecular recognition scaffold Zihao Cheng and Robert E. Campbell* Supplementary Methods Library construction for FRET-based screening
SUPPLEMENTARY MATERIAL An engineered tryptophan zipper-type peptide as a molecular recognition scaffold Zihao Cheng and Robert E. Campbell* Supplementary Methods Library construction for FRET-based screening
Det matematisk-naturvitenskapelige fakultet
 UNIVERSITETET I OSLO Det matematisk-naturvitenskapelige fakultet Exam in: MBV4010 Arbeidsmetoder i molekylærbiologi og biokjemi I MBV4010 Methods in molecular biology and biochemistry I Day of exam: Friday
UNIVERSITETET I OSLO Det matematisk-naturvitenskapelige fakultet Exam in: MBV4010 Arbeidsmetoder i molekylærbiologi og biokjemi I MBV4010 Methods in molecular biology and biochemistry I Day of exam: Friday
การสอบสวนโรคต ดเช อในโรงพยาบาล
 การสอบสวนโรคต ดเช อในโรงพยาบาล MRSA Model in Rajavithi Hospital by Chanwit Tribuddharat, M.D., Ph.D. Department of Microbiology Faculty of Medicine Siriraj Hospital Mahidol University E-mail sictb@mahidol.ac.th
การสอบสวนโรคต ดเช อในโรงพยาบาล MRSA Model in Rajavithi Hospital by Chanwit Tribuddharat, M.D., Ph.D. Department of Microbiology Faculty of Medicine Siriraj Hospital Mahidol University E-mail sictb@mahidol.ac.th
Occurrence of viruses in apple and pear orchards in Latvia.
 INTERNATIONAL SCIENTIFIC CONFERENCE Sustainable Fruit Growing: From Plant to Product Occurrence of viruses in apple and pear orchards in Latvia. Neda P pola, Anna K le, Inga Moro ko Bi evska. The economically
INTERNATIONAL SCIENTIFIC CONFERENCE Sustainable Fruit Growing: From Plant to Product Occurrence of viruses in apple and pear orchards in Latvia. Neda P pola, Anna K le, Inga Moro ko Bi evska. The economically
Codon Bias with PRISM. 2IM24/25, Fall 2007
 Codon Bias with PRISM 2IM24/25, Fall 2007 from RNA to protein mrna vs. trna aminoacid trna anticodon mrna codon codon-anticodon matching Watson-Crick base pairing A U and C G binding first two nucleotide
Codon Bias with PRISM 2IM24/25, Fall 2007 from RNA to protein mrna vs. trna aminoacid trna anticodon mrna codon codon-anticodon matching Watson-Crick base pairing A U and C G binding first two nucleotide
Protein Structure Analysis
 BINF 731 Protein Structure Analysis http://binf.gmu.edu/vaisman/binf731/ Iosif Vaisman COMPUTATIONAL BIOLOGY COMPUTATIONAL STRUCTURAL BIOLOGY COMPUTATIONAL MOLECULAR BIOLOGY BIOINFORMATICS STRUCTURAL BIOINFORMATICS
BINF 731 Protein Structure Analysis http://binf.gmu.edu/vaisman/binf731/ Iosif Vaisman COMPUTATIONAL BIOLOGY COMPUTATIONAL STRUCTURAL BIOLOGY COMPUTATIONAL MOLECULAR BIOLOGY BIOINFORMATICS STRUCTURAL BIOINFORMATICS
National PHL TB DST Reference Center PSQ Reporting Language Table of Contents
 PSQ Reporting Language Table of Contents Document Page Number PSQ for Rifampin 2-6 Comparison table for rpob Codon Numbering 2 rpob mutation list (new numbering system) 3-5 rpob interpretations 6 PSQ for
PSQ Reporting Language Table of Contents Document Page Number PSQ for Rifampin 2-6 Comparison table for rpob Codon Numbering 2 rpob mutation list (new numbering system) 3-5 rpob interpretations 6 PSQ for
I. Things to keep in mind before you start this protocol
 Inverse PCR and Sequencing Protocol on 5 Fly Preps For recovery of sequences flanking piggybac elements This protocol is an adaptation of "Inverse PCR and Cycle Sequencing Protocols" by E. Jay Rehm Berkeley
Inverse PCR and Sequencing Protocol on 5 Fly Preps For recovery of sequences flanking piggybac elements This protocol is an adaptation of "Inverse PCR and Cycle Sequencing Protocols" by E. Jay Rehm Berkeley
Supplemental Information. Target-Mediated Protection of Endogenous. MicroRNAs in C. elegans. Inventory of Supplementary Information
 Developmental Cell, Volume 20 Supplemental Information Target-Mediated Protection of Endogenous MicroRNAs in C. elegans Saibal Chatterjee, Monika Fasler, Ingo Büssing, and Helge Großhans Inventory of Supplementary
Developmental Cell, Volume 20 Supplemental Information Target-Mediated Protection of Endogenous MicroRNAs in C. elegans Saibal Chatterjee, Monika Fasler, Ingo Büssing, and Helge Großhans Inventory of Supplementary
Expression of Recombinant Proteins
 Expression of Recombinant Proteins Uses of Cloned Genes sequencing reagents (eg, probes) protein production insufficient natural quantities modify/mutagenesis library screening Expression Vector Features
Expression of Recombinant Proteins Uses of Cloned Genes sequencing reagents (eg, probes) protein production insufficient natural quantities modify/mutagenesis library screening Expression Vector Features
Supplementary Figure 1. The level of pri-mir-8 gradually decreases while those of BR-C and E74 increase during 3rd instar larval development.
 qrt-pcr RT-PCR Relative pri-mir-8 level 1.2 1.0 0.8 0.6 0.4 0.2 0.0 Early 3rd (72h) Mid 3rd (96h) Late 3rd (119h) BR-C E74 mtl rrna Early 3rd (72h) Mid 3rd (96h) Late 3rd (119h) Supplementary Figure 1.
qrt-pcr RT-PCR Relative pri-mir-8 level 1.2 1.0 0.8 0.6 0.4 0.2 0.0 Early 3rd (72h) Mid 3rd (96h) Late 3rd (119h) BR-C E74 mtl rrna Early 3rd (72h) Mid 3rd (96h) Late 3rd (119h) Supplementary Figure 1.
Engineering D66N mutant using quick change site directed mutagenesis. Harkewal Singh 09/01/2010
 Engineering D66N mutant using quick change site directed mutagenesis Harkewal Singh 09/01/2010 1 1- What is quick change site directed mutagenesis? 2- An overview of the kit contents. 3- A brief information
Engineering D66N mutant using quick change site directed mutagenesis Harkewal Singh 09/01/2010 1 1- What is quick change site directed mutagenesis? 2- An overview of the kit contents. 3- A brief information
DNA sentences. How are proteins coded for by DNA? Materials. Teacher instructions. Student instructions. Reflection
 DNA sentences How are proteins coded for by DNA? Deoxyribonucleic acid (DNA) is the molecule of life. DNA is one of the most recognizable nucleic acids, a double-stranded helix. The process by which DNA
DNA sentences How are proteins coded for by DNA? Deoxyribonucleic acid (DNA) is the molecule of life. DNA is one of the most recognizable nucleic acids, a double-stranded helix. The process by which DNA
Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of
 Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of the cell line) were immunostained for HA, acetylated
Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of the cell line) were immunostained for HA, acetylated
SUPPLEMENTARY INFORMATION
 doi:1.138/nature11496 Cl. 8 Cl. E93 Rag1 -/- 3H9 + BM Rag1 -/- BM CD CD c-kit c-kit c-kit wt Spleen c-kit B22 B22 IgM IgM IgM Supplementary Figure 1. FACS analysis of single-cell-derived pre-b cell clones.
doi:1.138/nature11496 Cl. 8 Cl. E93 Rag1 -/- 3H9 + BM Rag1 -/- BM CD CD c-kit c-kit c-kit wt Spleen c-kit B22 B22 IgM IgM IgM Supplementary Figure 1. FACS analysis of single-cell-derived pre-b cell clones.
Sequencing of DNA lesions facilitated by site-specific excision via base. excision repair DNA glycosylases yielding ligatable gaps
 Supporting information Sequencing of DNA lesions facilitated by site-specific excision via base excision repair DNA glycosylases yielding ligatable gaps Jan Riedl, Aaron M. Fleming, and Cynthia J. Burrows*
Supporting information Sequencing of DNA lesions facilitated by site-specific excision via base excision repair DNA glycosylases yielding ligatable gaps Jan Riedl, Aaron M. Fleming, and Cynthia J. Burrows*
SUPPLEMENTARY MATERIALS
 SUPPLEMENTARY MATERIALS Supplementary Table S1: Water sampling sites in Brisbane River and their characteristics Sampling sites GPS coordinates Site characteristics Suspected source of fecal pollution
SUPPLEMENTARY MATERIALS Supplementary Table S1: Water sampling sites in Brisbane River and their characteristics Sampling sites GPS coordinates Site characteristics Suspected source of fecal pollution
Multiplex PCR Study of Plasmid-Mediated AmpC Beta-Lactamase Genes in Clinical Isolates of Escherichia coli
 J Med Bacteriol. Vol. 5, No. 5, 6 (216): pp.21-28 jmb.tums.ac.ir Multiplex PCR Study of Plasmid-Mediated AmpC Beta-Lactamase Genes in Clinical Isolates of Escherichia coli Maryam Dehghani 1, Azam Haddadi
J Med Bacteriol. Vol. 5, No. 5, 6 (216): pp.21-28 jmb.tums.ac.ir Multiplex PCR Study of Plasmid-Mediated AmpC Beta-Lactamase Genes in Clinical Isolates of Escherichia coli Maryam Dehghani 1, Azam Haddadi
Yeast BioBrick Assembly (YBA)
 Yeast BioBrick Assembly (YBA) Standardized method for vector assembly of BioBrick devices via homologous recombination in Saccharomyces cerevisiae Martin Schneider, Leonard Fresenborg, Virginia Schadeweg
Yeast BioBrick Assembly (YBA) Standardized method for vector assembly of BioBrick devices via homologous recombination in Saccharomyces cerevisiae Martin Schneider, Leonard Fresenborg, Virginia Schadeweg
SUPPORTING INFORMATION FILE
 Intrinsic and extrinsic connections of Tet3 dioxygenase with CXXC zinc finger modules Nan Liu, Mengxi Wang, Wen Deng, Christine S. Schmidt, Weihua Qin, Heinrich Leonhardt and Fabio Spada Department of
Intrinsic and extrinsic connections of Tet3 dioxygenase with CXXC zinc finger modules Nan Liu, Mengxi Wang, Wen Deng, Christine S. Schmidt, Weihua Qin, Heinrich Leonhardt and Fabio Spada Department of
Supporting Information. Trifluoroacetophenone-Linked Nucleotides and DNA for Studying of DNA-protein Interactions by 19 F NMR Spectroscopy
 Supporting Information Trifluoroacetophenone-Linked Nucleotides and DNA for Studying of DNA-protein Interactions by 19 F NMR Spectroscopy Agata Olszewska, Radek Pohl and Michal Hocek # * Institute of Organic
Supporting Information Trifluoroacetophenone-Linked Nucleotides and DNA for Studying of DNA-protein Interactions by 19 F NMR Spectroscopy Agata Olszewska, Radek Pohl and Michal Hocek # * Institute of Organic
CHE-3H84: Protein Engineering Past Exam Papers
 CHE-3H84: Protein Engineering Past Exam Papers Sorted by Topic then Year Knowledge-Based Engineering of Proteins, Large Scale Production of Recombinant Proteins, and Protein Purification Dr. Hemmings 2006/7
CHE-3H84: Protein Engineering Past Exam Papers Sorted by Topic then Year Knowledge-Based Engineering of Proteins, Large Scale Production of Recombinant Proteins, and Protein Purification Dr. Hemmings 2006/7
Genomics and Gene Recognition Genes and Blue Genes
 Genomics and Gene Recognition Genes and Blue Genes November 1, 2004 Prokaryotic Gene Structure prokaryotes are simplest free-living organisms studying prokaryotes can give us a sense what is the minimum
Genomics and Gene Recognition Genes and Blue Genes November 1, 2004 Prokaryotic Gene Structure prokaryotes are simplest free-living organisms studying prokaryotes can give us a sense what is the minimum
High-throughput cloning and expression in recalcitrant bacteria
 High-throughput cloning and expression in recalcitrant bacteria Eric R Geertsma & Bert Poolman Supplementary text and figures: Supplementary Figure 1 Frequency of SfiI sites yielding identical 3 extensions
High-throughput cloning and expression in recalcitrant bacteria Eric R Geertsma & Bert Poolman Supplementary text and figures: Supplementary Figure 1 Frequency of SfiI sites yielding identical 3 extensions
FAT10 and NUB1L bind the VWA domain of Rpn10 and Rpn1 to enable proteasome-mediated proteolysis
 SUPPLEMENTARY INFORMATION FAT10 and NUB1L bind the VWA domain of Rpn10 and Rpn1 to enable proteasome-mediated proteolysis Neha Rani, Annette Aichem, Gunter Schmidtke, Stefan Kreft, and Marcus Groettrup
SUPPLEMENTARY INFORMATION FAT10 and NUB1L bind the VWA domain of Rpn10 and Rpn1 to enable proteasome-mediated proteolysis Neha Rani, Annette Aichem, Gunter Schmidtke, Stefan Kreft, and Marcus Groettrup
