Supplementary Figure 1 Strategy for parallel detection of DHSs and adjacent nucleosomes
|
|
|
- Shauna Hoover
- 5 years ago
- Views:
Transcription
1 Supplementary Figure 1 Strategy for parallel detection of DHSs and adjacent nucleosomes DNase I cleavage DNase I DNase I digestion Sucrose gradient enrichment Small Large F1 F F9 F1 F1 F2 F3 F4 F5 F6 F7 F8 F9 F1 * * * 1-kb Purification of hypersensitive sites Sequencing library construction Massively parallel paired-end sequencing of two size ranges bp <2-bp Sequencing of both small and large Map to reference genome and infer actual DNase I In silico reconstruction of regulatory DNA architecture In silico 1- to 125-bp separation of based on length 126- to 185-bp Transcription factors Nucleosomes A standard DNase I digestion of intact human nuclei (John 29, Thurman 212, Neph 212) preferentially releases small pertaining to sites of transcription factor binding and nucleosome occupancy. DNase I hypersensitive sites (DHSs) are purified via ultracentrifugation of DNA layered on top of a 9% sucrose cushion. Small remained near the top of tube, while larger migrated to the bottom. The separation was monitored on a 1.5% agarose gel and <1-kb (red dashed line) were used to construct a DNase I library. Canonical DHS are small (<1-bp) while nucleosomal are larger (>125-bp), therefore two size ranges were selected from the same DNase I library via agarose gel extraction and sequenced independently in paired-end mode. The paired sequencing reads were mapped to the genome and used to reconstruct DNase I in silico. Stratification of the inferred on the basis on length revealed sites of transcription factor or nucleosome occupancy. Nature Methods: doi:1.138/nmeth.2713
2 Supplementary Figure 2 Size selection of DNase I 1-bp 5-bp 3-bp 2-bp 1-bp Sequenced libraries DNase I were purified and ligated with adapters compatible with the Illumina sequencing platform. The ligated were amplified with 8 cycles of PCR and loaded onto a 2% agarose gel and run for 1V for 1 hr in Tris-acetate/EDTA (TAE) buffer. DNA was purified from two independent regions of the gel and sequenced. Dotted lines indicate the excised gel slices. Nature Methods: doi:1.138/nmeth.2713
3 Supplementary Figure 3 Distribution of DNase I fragment lengths a bp bp Frequency (in millions) Class DHS Class 1.4 bp periodicity 1 Class Nucleosome ,861,593 DNase I DNase I fragment length (bp) b 1.4 bp Power ( 1 6 ) Frequency (bp) (a) A histogram of inferred DNase I fragment lengths from the >25 million sequenced. The shaded curves highlight the regions on the bimodal distribution that correspond to derived from cleavages in a DHS (orange) or around nucleosomes (blue). Grey dashed lines demarcate the size ranges used for the stratification of. (b) The periodicity of the histogram in Fig. 1c was analyzed by Fourier transformation and plotted as a power spectrum (x-axis, frequency in base-pairs; y-axis, magnitude of signal). Nature Methods: doi:1.138/nmeth.2713
4 Supplementary Figure 4 DNase I cleavage on the nucleosome core particle NCP DNase I DNase I cleavage (per-nt) n = 54,173 nucleosomes Distance to nucleoseome dyad (bp) Per nucleotide DNase I cleavage profile around the 1% (n core particle.top, localization of theoretical DNase I released from digestion of the nucleosome. Within Nature Methods: doi:1.138/nmeth.2713
5 Supplementary Figure 5 Detection of nucleosomes flanking DHSs a DHS vs. nucleosome fragment signal intensity log 2 (Normalized DHS density) Linear model r log 2 (Normalized nucleosome density) b DNase I density profiles surrounding DHS peaks (i) 1- to 125-bp (ii) 126- to 185-bp Mean DNase I fragment density Distance from DHS peak (bp) 8 1%-ile 9%-ile 8%-ile 7%-ile 6%-ile 5%-ile 4%-ile 2%-ile 1%-ile Distance from DHS peak (bp) Mean DNase I fragment density (a) Scatterplot of small (y-axis) vs. large fragment (x-axis) reveals a strong correlation (linear model; r 2 P < 2.2 x 1-6 ). (b-c) ( bp). Nature Methods: doi:1.138/nmeth.2713
6 9% Mean length of DNase I covering >5% of motif (bp) High DNase I cleavage Relative position from motif (bp) Low Low High 1% 5,92 NF-Y motif instances in DHS 4% 5% 6% 7% 8% 9% NF-Y binding site NF-Y binding site Relative position from motif (bp) Relative position from motif (bp) Low High DNase I cleavage Low High 1- to 125-bp fragment density Low 5 +5 Mean length of DNase I covering >5% of motif (bp) Relative position from motif (bp) 5 1% +5 5 Nucleosome derived 3% Least accessible TF derived 25 2 Mean length of all DNase I 2% h 5,92 Nf-Y motif instances in DHS ordered as in (a) Most accessible g 5 NF-Y f 1 Binding sites grouped by decile e High 126- to 185-bp fragment density 1- to 125-bp fragment density % 8% +5 7% 6% 5% Low Relative position from motif (bp) 4% Relative position from motif (bp) NRF1 binding site 2 3% Least accessible/ occupied NRF1 binding site 5 4,716 NRF1 motif instances in DHSs Mean length of all DNase I 2% Nucleosome derived +5 1% TF derived 4,716 NRF1 motif instances in DHSs ordered as in (a) Most accessible/ occupied 1 5 NRF1 Binding sites grouped by decile Supplementary Figure 6 DNase I fragment length parallels TF occupancy a b c d High 126- to 185-bp fragment density (a) Aggregate profile (top) and heatmap (bottom) of the per-nucleotide cleavages surrounding ±5-bp surrounding 5 4,716 predicted NRF1 binding sites in accessible chromatin. Each row of the heatmap corresponds to one predicted NRF1 binding site and the rows are sorted by the cleavage density ±25-bp surrounding the TF recongition sequence. (b) Distribution of the mean fragment lengths overlapping each NRF1 motif. Each box corresponds to 1% of the predicted NRF1 binding sites ordered as in (a). A dashed red line represents the mean fragment length of all detected irrespective of binding site motif. (c-d) Heatmaps of the density of TF-derived (c) and nucleosome derived (d) ±5-bp surrounding predicted NRF1 binding sites from (a). (e-h)same as (a-d) for the 5,92 predicted binding sites for the transcriptional regulator NF-Y in accessible chromatin. Nature Methods: doi:1.138/nmeth.2713
7 Supplementary Figure 7 Nucleotide conservation surrounding nucleosomes flanking DHSs NCP Linker Regulatory DNA Motif density (TRANSFAC per-nt) Conservation (phastcons 46-way vertebrate per-nt) Distance to nucleoseome dyad (bp) 5 Nucleotide conservation surrounding the 1% most accessible nucleosomes. Grey, density of predicted transcription factor binding sites (TRANSFAC, P < 1 5 ). Magenta, the average (mean) phastcons (46-way Nature Methods: doi:1.138/nmeth.2713
8 Supplementary Figure 8 Sequence composition surrounding nucleosomes flanking DHSs a Dinucleotide pattern associated with well-positioned nucleosomes (Valouev et al., 211) Motif density GC dinucleotide GG dinucleotide AA dinucleotide NCP Linker Regulatory DNA Motif density (TRANSFAC per-nt) Dinucleotide frequency ( 1 2 ) (21-bp avg.) Distance to nucleoseome dyad (bp) 5 b 1 2 Shuffle sequence maintaining dinucleotide composition within region bp from nucleosome dyad Scan for TRANSFAC motifs (P < 1 5 ) Motif density (TRANSFAC per-nt) Not Dinucleotides shuffled shuffled (a) Sequence features surrounding the 1% most accessible nucleosomes. Grey, density of predicted transcription factor binding sites (TRANSFAC, P < 1 5 ). Colored curves show the dinucletide frequency for GC dinucleotides linker DNA. (b) Analysis of TRANSFAC motif density controlling for sequence dinucleotide composition. Nature Methods: doi:1.138/nmeth.2713
9 Supplementary Figure 9 Nucleosome organization surrounding TF binding sites b USF Nucleosome dyad distance to motif (bp).12 Nucleosome dyad distance to motif (bp).25 Density Density Density c AP-1 2 CTCF 2 a Nucleosome dyad distance to motif (bp) (a-c) The nucleosome profiles surrounding the top 1% most highly occupied transcription factor binding sites of (a) CTCF, (b) AP-1 and (c) USF-1. Nature Methods: doi:1.138/nmeth.2713
10 Supplementary Figure 1 DNase I reveals a diversity of promoter types Total unidirectional promoter DHSs (n = 18,454) Promoter with 1 TSS 12 No. DHS No. +1 nucleosomes in promoter DHSs Promoter with 2 TSSs utilizing distinct +1 nucleosomes 18,454 DHS scorresponding to unidirectional promoters were scanned for the presence of one or more +1 nucleosomes. Nature Methods: doi:1.138/nmeth.2713
11 Supplementary Figure 11 Fig. 5a Fig. 5a Fig. 5a Nature Methods: doi:1.138/nmeth.2713
12 Supplementary Table 1 Summary of sequencing statistics corresponding to the two size fractions gel purifed. DNase I library Total mapped (Q > 1) Proper mapping (-chrm) % unique mapping No. cleavages detected <2 bp 119,623,788 89,763, % 179,527, bp 29,192,86 162,97, % 324,195,288 Combined total 328,8, ,861, % 53,723,186 Nature Methods: doi:1.138/nmeth.2713
SUPPLEMENTARY INFORMATION
 SUPPLEMENRY INFORMION doi:.38/nature In vivo nucleosome mapping D4+ Lymphocytes radient-based and I-bead cell sorting D8+ Lymphocytes ranulocytes Lyse the cells Isolate and sequence mononucleosome cores
SUPPLEMENRY INFORMION doi:.38/nature In vivo nucleosome mapping D4+ Lymphocytes radient-based and I-bead cell sorting D8+ Lymphocytes ranulocytes Lyse the cells Isolate and sequence mononucleosome cores
Nature Methods: doi: /nmeth.4396
 Supplementary Figure 1 Comparison of technical replicate consistency between and across the standard ATAC-seq method, DNase-seq, and Omni-ATAC. (a) Heatmap-based representation of ATAC-seq quality control
Supplementary Figure 1 Comparison of technical replicate consistency between and across the standard ATAC-seq method, DNase-seq, and Omni-ATAC. (a) Heatmap-based representation of ATAC-seq quality control
Supplementary Table 1: Oligo designs. A list of ATAC-seq oligos used for PCR.
 Ad1_noMX: Ad2.1_TAAGGCGA Ad2.2_CGTACTAG Ad2.3_AGGCAGAA Ad2.4_TCCTGAGC Ad2.5_GGACTCCT Ad2.6_TAGGCATG Ad2.7_CTCTCTAC Ad2.8_CAGAGAGG Ad2.9_GCTACGCT Ad2.10_CGAGGCTG Ad2.11_AAGAGGCA Ad2.12_GTAGAGGA Ad2.13_GTCGTGAT
Ad1_noMX: Ad2.1_TAAGGCGA Ad2.2_CGTACTAG Ad2.3_AGGCAGAA Ad2.4_TCCTGAGC Ad2.5_GGACTCCT Ad2.6_TAGGCATG Ad2.7_CTCTCTAC Ad2.8_CAGAGAGG Ad2.9_GCTACGCT Ad2.10_CGAGGCTG Ad2.11_AAGAGGCA Ad2.12_GTAGAGGA Ad2.13_GTCGTGAT
DNA Visualizer Extraction Kit
 DNA Visualizer Extraction Kit Catalog Number D0006 50 reactions Version: 03 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Intended Use... 3 Background... 3 General Information...
DNA Visualizer Extraction Kit Catalog Number D0006 50 reactions Version: 03 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Intended Use... 3 Background... 3 General Information...
Figure S1. Replication initiation sites at efficient ORIs do not coincide with nucleosomedepleted
 Figure S1. Replication initiation sites at efficient ORIs do not coincide with nucleosomedepleted regions in the mouse genome. Typical examples of SNS profiles (Cayrou et al., 11) and nucleosome occupancies
Figure S1. Replication initiation sites at efficient ORIs do not coincide with nucleosomedepleted regions in the mouse genome. Typical examples of SNS profiles (Cayrou et al., 11) and nucleosome occupancies
Deep learning frameworks for regulatory genomics and epigenomics
 Deep learning frameworks for regulatory genomics and epigenomics Chuan Sheng Foo Avanti Shrikumar Nicholas Sinnott- Armstrong ANSHUL KUNDAJE Genetics, Computer science Stanford University Johnny Israeli
Deep learning frameworks for regulatory genomics and epigenomics Chuan Sheng Foo Avanti Shrikumar Nicholas Sinnott- Armstrong ANSHUL KUNDAJE Genetics, Computer science Stanford University Johnny Israeli
Intracellular receptors specify complex patterns of gene expression that are cell and gene
 SUPPLEMENTAL RESULTS AND DISCUSSION Some HPr-1AR ARE-containing Genes Are Unresponsive to Androgen Intracellular receptors specify complex patterns of gene expression that are cell and gene specific. For
SUPPLEMENTAL RESULTS AND DISCUSSION Some HPr-1AR ARE-containing Genes Are Unresponsive to Androgen Intracellular receptors specify complex patterns of gene expression that are cell and gene specific. For
Integrated NGS Sample Preparation Solutions for Limiting Amounts of RNA and DNA. March 2, Steven R. Kain, Ph.D. ABRF 2013
 Integrated NGS Sample Preparation Solutions for Limiting Amounts of RNA and DNA March 2, 2013 Steven R. Kain, Ph.D. ABRF 2013 NuGEN s Core Technologies Selective Sequence Priming Nucleic Acid Amplification
Integrated NGS Sample Preparation Solutions for Limiting Amounts of RNA and DNA March 2, 2013 Steven R. Kain, Ph.D. ABRF 2013 NuGEN s Core Technologies Selective Sequence Priming Nucleic Acid Amplification
Figure S1. nuclear extracts. HeLa cell nuclear extract. Input IgG IP:ORC2 ORC2 ORC2. MCM4 origin. ORC2 occupancy
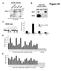 A nuclear extracts B HeLa cell nuclear extract Figure S1 ORC2 (in kda) 21 132 7 ORC2 Input IgG IP:ORC2 32 ORC C D PRKDC ORC2 occupancy Directed against ORC2 C-terminus (sc-272) MCM origin 2 2 1-1 -1kb
A nuclear extracts B HeLa cell nuclear extract Figure S1 ORC2 (in kda) 21 132 7 ORC2 Input IgG IP:ORC2 32 ORC C D PRKDC ORC2 occupancy Directed against ORC2 C-terminus (sc-272) MCM origin 2 2 1-1 -1kb
Next-generation sequencing technologies
 Next-generation sequencing technologies NGS applications Illumina sequencing workflow Overview Sequencing by ligation Short-read NGS Sequencing by synthesis Illumina NGS Single-molecule approach Long-read
Next-generation sequencing technologies NGS applications Illumina sequencing workflow Overview Sequencing by ligation Short-read NGS Sequencing by synthesis Illumina NGS Single-molecule approach Long-read
Performance characteristics of the High Sensitivity DNA kit for the Agilent 2100 Bioanalyzer
 Performance characteristics of the High Sensitivity DNA kit for the Agilent 2100 Bioanalyzer Technical Note 10 Measured conc. [ng/µl] 1 Y intercept = 0.09 r 2 = 0.993 0.1 0.1 1 10 Reference concentration
Performance characteristics of the High Sensitivity DNA kit for the Agilent 2100 Bioanalyzer Technical Note 10 Measured conc. [ng/µl] 1 Y intercept = 0.09 r 2 = 0.993 0.1 0.1 1 10 Reference concentration
Computational Analysis of Ultra-high-throughput sequencing data: ChIP-Seq
 Computational Analysis of Ultra-high-throughput sequencing data: ChIP-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne Data flow in ChIP-Seq data analysis Level 1:
Computational Analysis of Ultra-high-throughput sequencing data: ChIP-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne Data flow in ChIP-Seq data analysis Level 1:
Suppl. Table S1. Characteristics of DHS regions analyzed by bisulfite sequencing. No. CpGs analyzed in the amplicon. Genomic location specificity
 Suppl. Table S1. Characteristics of DHS regions analyzed by bisulfite sequencing. DHS/GRE Genomic location Tissue specificity DHS type CpG density (per 100 bp) No. CpGs analyzed in the amplicon CpG within
Suppl. Table S1. Characteristics of DHS regions analyzed by bisulfite sequencing. DHS/GRE Genomic location Tissue specificity DHS type CpG density (per 100 bp) No. CpGs analyzed in the amplicon CpG within
Library construction for nextgeneration sequencing: Overviews. and challenges
 Library construction for nextgeneration sequencing: Overviews and challenges During this time, as sequencing technologies have improved and evolved, so too have methods for preparing nucleic acids for
Library construction for nextgeneration sequencing: Overviews and challenges During this time, as sequencing technologies have improved and evolved, so too have methods for preparing nucleic acids for
Gel/PCR Extraction Kit
 Gel/PCR Extraction Kit Item No: EX-GP200 (200rxns) Content Content Binding Buffer BD Wash Buffer PE Elution Buffer (10 mm Tris-HCl, ph 8.5) Spin Columns EX-GP200 80 ml 20 mlx3 10 ml 200 each Description
Gel/PCR Extraction Kit Item No: EX-GP200 (200rxns) Content Content Binding Buffer BD Wash Buffer PE Elution Buffer (10 mm Tris-HCl, ph 8.5) Spin Columns EX-GP200 80 ml 20 mlx3 10 ml 200 each Description
Supplementary Figures
 Supplementary Figures A B Supplementary Figure 1. Examples of discrepancies in predicted and validated breakpoint coordinates. A) Most frequently, predicted breakpoints were shifted relative to those derived
Supplementary Figures A B Supplementary Figure 1. Examples of discrepancies in predicted and validated breakpoint coordinates. A) Most frequently, predicted breakpoints were shifted relative to those derived
Personal and population genomics of human regulatory variation
 Personal and population genomics of human regulatory variation Benjamin Vernot,, and Joshua M. Akey Department of Genome Sciences, University of Washington, Seattle, Washington 98195, USA Ting WANG 1 Brief
Personal and population genomics of human regulatory variation Benjamin Vernot,, and Joshua M. Akey Department of Genome Sciences, University of Washington, Seattle, Washington 98195, USA Ting WANG 1 Brief
Nature Biotechnology: doi: /nbt Supplementary Figure 1. sndrop-seq overview.
 Supplementary Figure 1 sndrop-seq overview. A. sndrop-seq method showing modifications needed to process nuclei, including bovine serum albumin (BSA) coating and droplet heating to ensure complete nuclear
Supplementary Figure 1 sndrop-seq overview. A. sndrop-seq method showing modifications needed to process nuclei, including bovine serum albumin (BSA) coating and droplet heating to ensure complete nuclear
DNA sequence and chromatin structure. Mapping nucleosome positioning using high-throughput sequencing
 DNA sequence and chromatin structure Mapping nucleosome positioning using high-throughput sequencing DNA sequence and chromatin structure Higher-order 30 nm fibre Mapping nucleosome positioning using high-throughput
DNA sequence and chromatin structure Mapping nucleosome positioning using high-throughput sequencing DNA sequence and chromatin structure Higher-order 30 nm fibre Mapping nucleosome positioning using high-throughput
BunDLE-seq (Binding to Designed Library, Extracting and Sequencing) -
 Protocol BunDLE-seq (Binding to Designed Library, Extracting and Sequencing) - A quantitative investigation of various determinants of TF binding; going beyond the characterization of core site Einat Zalckvar*
Protocol BunDLE-seq (Binding to Designed Library, Extracting and Sequencing) - A quantitative investigation of various determinants of TF binding; going beyond the characterization of core site Einat Zalckvar*
SUPPLEMENTARY INFORMATION
 doi:1.138/nature1895 i Core histones CAF-1 v ii Reb1 CAF-1 Fen1 vi Mature Okazaki fragment iii PCNA Reb1 iv vii Mature Okazaki fragment Reb1 Mature Okazaki fragments Figure S1. Model for nucleosome-mediated
doi:1.138/nature1895 i Core histones CAF-1 v ii Reb1 CAF-1 Fen1 vi Mature Okazaki fragment iii PCNA Reb1 iv vii Mature Okazaki fragment Reb1 Mature Okazaki fragments Figure S1. Model for nucleosome-mediated
PIP-seq. Cells. Permanganate ChIP-Seq
 PIP-seq ells Formaldehyde Permanganate 5 Harvest Lyse Sonicate First dapter Ligation 3 3 5 hip Elute Reverse rosslinks Piperidine cleavage 5 3 3 5 Primer Extension Second dapter Ligation 5 3 3 5 Deep Sequencing
PIP-seq ells Formaldehyde Permanganate 5 Harvest Lyse Sonicate First dapter Ligation 3 3 5 hip Elute Reverse rosslinks Piperidine cleavage 5 3 3 5 Primer Extension Second dapter Ligation 5 3 3 5 Deep Sequencing
Zymo-Spin Technology. The microcentrifuge column that revolutionized nucleic acid clean-up
 Zymo-Spin Technology The microcentrifuge column that revolutionized nucleic acid clean-up Overview A Comprehensive Solution for DNA and RNA Clean-Up High-quality, inhibitor-free DNA and RNA is crucial
Zymo-Spin Technology The microcentrifuge column that revolutionized nucleic acid clean-up Overview A Comprehensive Solution for DNA and RNA Clean-Up High-quality, inhibitor-free DNA and RNA is crucial
High Resolution LabChip XT Fractionation of Illumina Compatible Small RNA Libraries using the DNA 300 Assay Kit
 application Note Next Generation Sequencing Authors: Lucy Sun Danh Tran Donna Fong Ryan Swenerton Elaine Wong-Ho Rajendra Singh Josh Molho Caliper Life Sciences a PerkinElmer Company Alameda, California
application Note Next Generation Sequencing Authors: Lucy Sun Danh Tran Donna Fong Ryan Swenerton Elaine Wong-Ho Rajendra Singh Josh Molho Caliper Life Sciences a PerkinElmer Company Alameda, California
Genome 373: High- Throughput DNA Sequencing. Doug Fowler
 Genome 373: High- Throughput DNA Sequencing Doug Fowler Tasks give ML unity We learned about three tasks that are commonly encountered in ML Models/Algorithms Give ML Diversity Classification Regression
Genome 373: High- Throughput DNA Sequencing Doug Fowler Tasks give ML unity We learned about three tasks that are commonly encountered in ML Models/Algorithms Give ML Diversity Classification Regression
INDEX KIT COMPONENTS 1 STORAGE AND STABILITY 1 INTRODUCTION 1 BENEFITS
 INDEX KIT COMPONENTS 1 STORAGE AND STABILITY 1 INTRODUCTION 1 BENEFITS ERRORE. IL SEGNALIBRO NON È DEFINITO. IMPORTANT NOTES 1 METHOD FOR DNA EXTRACTION FROM AGAROSE GELS 2 METHOD FOR DNA EXTRACTION FROM
INDEX KIT COMPONENTS 1 STORAGE AND STABILITY 1 INTRODUCTION 1 BENEFITS ERRORE. IL SEGNALIBRO NON È DEFINITO. IMPORTANT NOTES 1 METHOD FOR DNA EXTRACTION FROM AGAROSE GELS 2 METHOD FOR DNA EXTRACTION FROM
Mate-pair library data improves genome assembly
 De Novo Sequencing on the Ion Torrent PGM APPLICATION NOTE Mate-pair library data improves genome assembly Highly accurate PGM data allows for de Novo Sequencing and Assembly For a draft assembly, generate
De Novo Sequencing on the Ion Torrent PGM APPLICATION NOTE Mate-pair library data improves genome assembly Highly accurate PGM data allows for de Novo Sequencing and Assembly For a draft assembly, generate
Parts of a standard FastQC report
 FastQC FastQC, written by Simon Andrews of Babraham Bioinformatics, is a very popular tool used to provide an overview of basic quality control metrics for raw next generation sequencing data. There are
FastQC FastQC, written by Simon Andrews of Babraham Bioinformatics, is a very popular tool used to provide an overview of basic quality control metrics for raw next generation sequencing data. There are
Single-molecule imaging of DNA curtains reveals intrinsic energy landscapes for nucleosome deposition
 SUPPLEMENTARY INFORMATION Single-molecule imaging of DNA curtains reveals intrinsic energy landscapes for nucleosome deposition Mari-Liis Visnapuu 1 and Eric C. Greene 1 1 Department of Biochemistry &
SUPPLEMENTARY INFORMATION Single-molecule imaging of DNA curtains reveals intrinsic energy landscapes for nucleosome deposition Mari-Liis Visnapuu 1 and Eric C. Greene 1 1 Department of Biochemistry &
Mapping and quantifying mammalian transcriptomes by RNA-Seq. Ali Mortazavi, Brian A Williams, Kenneth McCue, Lorian Schaeffer & Barbara Wold
 Mapping and quantifying mammalian transcriptomes by RNA-Seq Ali Mortazavi, Brian A Williams, Kenneth McCue, Lorian Schaeffer & Barbara Wold Supplementary figures and text: Supplementary Figure 1 RNA shatter
Mapping and quantifying mammalian transcriptomes by RNA-Seq Ali Mortazavi, Brian A Williams, Kenneth McCue, Lorian Schaeffer & Barbara Wold Supplementary figures and text: Supplementary Figure 1 RNA shatter
Guide-it sgrna In Vitro Transcription and Screening Systems User Manual
 Clontech Laboratories, Inc. Guide-it sgrna In Vitro Transcription and Screening Systems User Manual Cat. Nos. 631438, 631439 & 631440 (042114) Clontech Laboratories, Inc. A Takara Bio Company 1290 Terra
Clontech Laboratories, Inc. Guide-it sgrna In Vitro Transcription and Screening Systems User Manual Cat. Nos. 631438, 631439 & 631440 (042114) Clontech Laboratories, Inc. A Takara Bio Company 1290 Terra
TECH NOTE Ligation-Free ChIP-Seq Library Preparation
 TECH NOTE Ligation-Free ChIP-Seq Library Preparation The DNA SMART ChIP-Seq Kit Ligation-free template switching technology: Minimize sample handling in a single-tube workflow >> Simplified protocol with
TECH NOTE Ligation-Free ChIP-Seq Library Preparation The DNA SMART ChIP-Seq Kit Ligation-free template switching technology: Minimize sample handling in a single-tube workflow >> Simplified protocol with
Introduction to genome biology
 Introduction to genome biology Lisa Stubbs Deep transcritpomes for traditional model species from ENCODE (and modencode) Deep RNA-seq and chromatin analysis on 147 human cell types, as well as tissues,
Introduction to genome biology Lisa Stubbs Deep transcritpomes for traditional model species from ENCODE (and modencode) Deep RNA-seq and chromatin analysis on 147 human cell types, as well as tissues,
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Multiplet rate and sensitivity of the GemCode single cell platform from scrna-seq of 50:50 mixing of 293Ts and 3T3s. (a) Inferred multiplet rate as a function
Supplementary Figures Supplementary Figure 1. Multiplet rate and sensitivity of the GemCode single cell platform from scrna-seq of 50:50 mixing of 293Ts and 3T3s. (a) Inferred multiplet rate as a function
Nature Structural & Molecular Biology: doi: /nsmb.3175
 Supplementary Figure 1 Medium- and low-fret groups probably representing varying positions on the DNA show similar changes between H3 and CENP-A nucleosomes with and without CENP-C CD, as does the high-fret
Supplementary Figure 1 Medium- and low-fret groups probably representing varying positions on the DNA show similar changes between H3 and CENP-A nucleosomes with and without CENP-C CD, as does the high-fret
SUPPLEMENTARY INFORMATION SUPPLEMENTARY FIGURE LEGENDS
 1 2 3 4 6 7 8 9 11 12 13 14 1 16 17 18 19 21 22 23 24 2 26 27 28 29 3 31 32 33 34 3 36 37 38 39 4 41 42 43 44 4 46 47 48 49 1 SUPPLEMENTRY INFORMTION SUPPLEMENTRY FIURE LEENDS Supplementary Figure S1.
1 2 3 4 6 7 8 9 11 12 13 14 1 16 17 18 19 21 22 23 24 2 26 27 28 29 3 31 32 33 34 3 36 37 38 39 4 41 42 43 44 4 46 47 48 49 1 SUPPLEMENTRY INFORMTION SUPPLEMENTRY FIURE LEENDS Supplementary Figure S1.
Introduction to genome biology
 Introduction to genome biology Lisa Stubbs We ve found most genes; but what about the rest of the genome? Genome size* 12 Mb 95 Mb 170 Mb 1500 Mb 2700 Mb 3200 Mb #coding genes ~7000 ~20000 ~14000 ~26000
Introduction to genome biology Lisa Stubbs We ve found most genes; but what about the rest of the genome? Genome size* 12 Mb 95 Mb 170 Mb 1500 Mb 2700 Mb 3200 Mb #coding genes ~7000 ~20000 ~14000 ~26000
Figure S1. Unrearranged locus. Rearranged locus. Concordant read pairs. Region1. Region2. Cluster of discordant read pairs, bundle
 Figure S1 a Unrearranged locus Rearranged locus Concordant read pairs Region1 Concordant read pairs Cluster of discordant read pairs, bundle Region2 Concordant read pairs b Physical coverage 5 4 3 2 1
Figure S1 a Unrearranged locus Rearranged locus Concordant read pairs Region1 Concordant read pairs Cluster of discordant read pairs, bundle Region2 Concordant read pairs b Physical coverage 5 4 3 2 1
Supplementary Figure 2
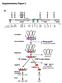 Supplementary Figure 2 a SBS-C1 SBS-C2 SBS-C3 SBS-C4 SBS-C5 SBS-C6 SBS-C7 SBS-C8 SBS-C9 LCR CNS-1 CNS-2 Il5 Rad50 Il13 Il4 Kif3a Sept8 0 50 100 150 200kb Sau3AI 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17
Supplementary Figure 2 a SBS-C1 SBS-C2 SBS-C3 SBS-C4 SBS-C5 SBS-C6 SBS-C7 SBS-C8 SBS-C9 LCR CNS-1 CNS-2 Il5 Rad50 Il13 Il4 Kif3a Sept8 0 50 100 150 200kb Sau3AI 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17
Lecture 7. Next-generation sequencing technologies
 Lecture 7 Next-generation sequencing technologies Next-generation sequencing technologies General principles of short-read NGS Construct a library of fragments Generate clonal template populations Massively
Lecture 7 Next-generation sequencing technologies Next-generation sequencing technologies General principles of short-read NGS Construct a library of fragments Generate clonal template populations Massively
Genetics and Genomics in Medicine Chapter 3. Questions & Answers
 Genetics and Genomics in Medicine Chapter 3 Multiple Choice Questions Questions & Answers Question 3.1 Which of the following statements, if any, is false? a) Amplifying DNA means making many identical
Genetics and Genomics in Medicine Chapter 3 Multiple Choice Questions Questions & Answers Question 3.1 Which of the following statements, if any, is false? a) Amplifying DNA means making many identical
TIANgel Mini DNA Purification Kit
 TIANgel Mini DNA Purification Kit For DNA purification from agarose and polyacrylamide gels www.tiangen.com/en DP130419 TIANgel Mini DNA Purification Kit Kit Contents (Spin column) Cat. no. DP208 Contents
TIANgel Mini DNA Purification Kit For DNA purification from agarose and polyacrylamide gels www.tiangen.com/en DP130419 TIANgel Mini DNA Purification Kit Kit Contents (Spin column) Cat. no. DP208 Contents
Shin Lin CS229 Final Project Identifying Transcription Factor Binding by the DNase Hypersensitivity Assay
 BACKGROUND In the DNA of cell nuclei, transcription factors (TF) bind regulatory regions throughout the genome in tissue specific patterns. The binding occurs for three reasons: 1) the TF is present, 2)
BACKGROUND In the DNA of cell nuclei, transcription factors (TF) bind regulatory regions throughout the genome in tissue specific patterns. The binding occurs for three reasons: 1) the TF is present, 2)
THE ANALYSIS OF CHROMATIN CONDENSATION STATE AND TRANSCRIPTIONAL ACTIVITY USING DNA MICROARRAYS 1. INTRODUCTION
 JOURNAL OF MEDICAL INFORMATICS & TECHNOLOGIES Vol.6/2003, ISSN 1642-6037 Piotr WIDŁAK *, Krzysztof FUJAREWICZ ** DNA microarrays, chromatin, transcription THE ANALYSIS OF CHROMATIN CONDENSATION STATE AND
JOURNAL OF MEDICAL INFORMATICS & TECHNOLOGIES Vol.6/2003, ISSN 1642-6037 Piotr WIDŁAK *, Krzysztof FUJAREWICZ ** DNA microarrays, chromatin, transcription THE ANALYSIS OF CHROMATIN CONDENSATION STATE AND
of NOBOX. Supplemental Figure S1F shows the staining profile of LHX8 antibody (Abcam, ab41519), which is a
 SUPPLEMENTAL MATERIALS AND METHODS LHX8 Staining During the course of this work, it came to our attention that the NOBOX antibody used in this study had been discontinued. There are a number of other nuclear
SUPPLEMENTAL MATERIALS AND METHODS LHX8 Staining During the course of this work, it came to our attention that the NOBOX antibody used in this study had been discontinued. There are a number of other nuclear
Chromatin Structure. a basic discussion of protein-nucleic acid binding
 Chromatin Structure 1 Chromatin DNA packaging g First a basic discussion of protein-nucleic acid binding Questions to answer: How do proteins bind DNA / RNA? How do proteins recognize a specific nucleic
Chromatin Structure 1 Chromatin DNA packaging g First a basic discussion of protein-nucleic acid binding Questions to answer: How do proteins bind DNA / RNA? How do proteins recognize a specific nucleic
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Origin use and efficiency are similar among WT, rrm3, pif1-m2, and pif1-m2; rrm3 strains. A. Analysis of fork progression around confirmed and likely origins (from cerevisiae.oridb.org).
Supplementary Figure 1 Origin use and efficiency are similar among WT, rrm3, pif1-m2, and pif1-m2; rrm3 strains. A. Analysis of fork progression around confirmed and likely origins (from cerevisiae.oridb.org).
The Agilent Total RNA Isolation Kit
 Better RNA purity. Better data. Without DNase treatment. The Agilent Total RNA Isolation Kit Now you can isolate highly purified, intact RNA without DNase treatment Introducing the Agilent Total RNA Isolation
Better RNA purity. Better data. Without DNase treatment. The Agilent Total RNA Isolation Kit Now you can isolate highly purified, intact RNA without DNase treatment Introducing the Agilent Total RNA Isolation
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Number and length distributions of the inferred fosmids.
 Supplementary Figure 1 Number and length distributions of the inferred fosmids. Fosmid were inferred by mapping each pool s sequence reads to hg19. We retained only those reads that mapped to within a
Supplementary Figure 1 Number and length distributions of the inferred fosmids. Fosmid were inferred by mapping each pool s sequence reads to hg19. We retained only those reads that mapped to within a
NB536: Bioinformatics
 NB536: Bioinformatics Instructor Prof. Jong Kyoung Kim Department of New Biology Office: E4-613 E-mail: jkkim@dgist.ac.kr Homepage: https://scg.dgist.ac.kr Course website https://scg.dgist.ac.kr/index.php/courses
NB536: Bioinformatics Instructor Prof. Jong Kyoung Kim Department of New Biology Office: E4-613 E-mail: jkkim@dgist.ac.kr Homepage: https://scg.dgist.ac.kr Course website https://scg.dgist.ac.kr/index.php/courses
Impact of gdna Integrity on the Outcome of DNA Methylation Studies
 Impact of gdna Integrity on the Outcome of DNA Methylation Studies Application Note Nucleic Acid Analysis Authors Emily Putnam, Keith Booher, and Xueguang Sun Zymo Research Corporation, Irvine, CA, USA
Impact of gdna Integrity on the Outcome of DNA Methylation Studies Application Note Nucleic Acid Analysis Authors Emily Putnam, Keith Booher, and Xueguang Sun Zymo Research Corporation, Irvine, CA, USA
Supplemental Figure 1.
 Supplemental Data. Charron et al. Dynamic landscapes of four histone modifications during de-etiolation in Arabidopsis. Plant Cell (2009). 10.1105/tpc.109.066845 Supplemental Figure 1. Immunodetection
Supplemental Data. Charron et al. Dynamic landscapes of four histone modifications during de-etiolation in Arabidopsis. Plant Cell (2009). 10.1105/tpc.109.066845 Supplemental Figure 1. Immunodetection
Zhang et al., RepID facilitates replication Initiation. Supplemental Information:
 Supplemental Information: a b 1 Supplementary Figure 1 (a) DNA sequence of all the oligonucleotides used in this study. Only one strand is shown. The unshaded nucleotide sequences show changes from the
Supplemental Information: a b 1 Supplementary Figure 1 (a) DNA sequence of all the oligonucleotides used in this study. Only one strand is shown. The unshaded nucleotide sequences show changes from the
Conserved elements with potential to form polymorphic G-quadruplex structures in the first intron of human genes
 Published online 10 January 2008 Nucleic Acids Research, 2008, Vol. 36, No. 4 1321 1333 doi:10.1093/nar/gkm1138 Conserved elements with potential to form polymorphic G-quadruplex structures in the first
Published online 10 January 2008 Nucleic Acids Research, 2008, Vol. 36, No. 4 1321 1333 doi:10.1093/nar/gkm1138 Conserved elements with potential to form polymorphic G-quadruplex structures in the first
Roche Molecular Biochemicals Technical Note No. LC 10/2000
 Roche Molecular Biochemicals Technical Note No. LC 10/2000 LightCycler Overview of LightCycler Quantification Methods 1. General Introduction Introduction Content Definitions This Technical Note will introduce
Roche Molecular Biochemicals Technical Note No. LC 10/2000 LightCycler Overview of LightCycler Quantification Methods 1. General Introduction Introduction Content Definitions This Technical Note will introduce
Illumina Sequencing Sample Preparation for use with CRISPRi/a-v2 Libraries
 Illumina Sequencing Sample Preparation for use with CRISPRi/a-v2 Libraries Overview This protocol describes the four steps required for generating sequencing samples from cells harvested from screens conducted
Illumina Sequencing Sample Preparation for use with CRISPRi/a-v2 Libraries Overview This protocol describes the four steps required for generating sequencing samples from cells harvested from screens conducted
Overview of Next Generation Sequencing technologies. Céline Keime
 Overview of Next Generation Sequencing technologies Céline Keime keime@igbmc.fr Next Generation Sequencing < Second generation sequencing < General principle < Sequencing by synthesis - Illumina < Sequencing
Overview of Next Generation Sequencing technologies Céline Keime keime@igbmc.fr Next Generation Sequencing < Second generation sequencing < General principle < Sequencing by synthesis - Illumina < Sequencing
MODULE TSS1: TRANSCRIPTION START SITES INTRODUCTION (BASIC)
 MODULE TSS1: TRANSCRIPTION START SITES INTRODUCTION (BASIC) Lesson Plan: Title JAMIE SIDERS, MEG LAAKSO & WILSON LEUNG Identifying transcription start sites for Peaked promoters using chromatin landscape,
MODULE TSS1: TRANSCRIPTION START SITES INTRODUCTION (BASIC) Lesson Plan: Title JAMIE SIDERS, MEG LAAKSO & WILSON LEUNG Identifying transcription start sites for Peaked promoters using chromatin landscape,
The ChIP-Seq project. Giovanna Ambrosini, Philipp Bucher. April 19, 2010 Lausanne. EPFL-SV Bucher Group
 The ChIP-Seq project Giovanna Ambrosini, Philipp Bucher EPFL-SV Bucher Group April 19, 2010 Lausanne Overview Focus on technical aspects Description of applications (C programs) Where to find binaries,
The ChIP-Seq project Giovanna Ambrosini, Philipp Bucher EPFL-SV Bucher Group April 19, 2010 Lausanne Overview Focus on technical aspects Description of applications (C programs) Where to find binaries,
Simultaneous profiling of transcriptome and DNA methylome from a single cell
 Additional file 1: Supplementary materials Simultaneous profiling of transcriptome and DNA methylome from a single cell Youjin Hu 1, 2, Kevin Huang 1, 3, Qin An 1, Guizhen Du 1, Ganlu Hu 2, Jinfeng Xue
Additional file 1: Supplementary materials Simultaneous profiling of transcriptome and DNA methylome from a single cell Youjin Hu 1, 2, Kevin Huang 1, 3, Qin An 1, Guizhen Du 1, Ganlu Hu 2, Jinfeng Xue
Mapping long-range promoter contacts in human cells with high-resolution capture Hi-C
 CORRECTION NOTICE Nat. Genet. 47, 598 606 (2015) Mapping long-range promoter contacts in human cells with high-resolution capture Hi-C Borbala Mifsud, Filipe Tavares-Cadete, Alice N Young, Robert Sugar,
CORRECTION NOTICE Nat. Genet. 47, 598 606 (2015) Mapping long-range promoter contacts in human cells with high-resolution capture Hi-C Borbala Mifsud, Filipe Tavares-Cadete, Alice N Young, Robert Sugar,
Get to Know Your DNA. Every Single Fragment.
 HaloPlex HS NGS Target Enrichment System Get to Know Your DNA. Every Single Fragment. High sensitivity detection of rare variants using molecular barcodes How Does Molecular Barcoding Work? HaloPlex HS
HaloPlex HS NGS Target Enrichment System Get to Know Your DNA. Every Single Fragment. High sensitivity detection of rare variants using molecular barcodes How Does Molecular Barcoding Work? HaloPlex HS
Understanding transcriptional regulation by integrative analysis of transcription factor binding data
 Understanding transcriptional regulation by integrative analysis of transcription factor binding data Cheng et al. 2012 Shu Yang Feb. 21, 2013 1 / 26 Introduction 2 / 26 DNA-binding Proteins sequence-specific
Understanding transcriptional regulation by integrative analysis of transcription factor binding data Cheng et al. 2012 Shu Yang Feb. 21, 2013 1 / 26 Introduction 2 / 26 DNA-binding Proteins sequence-specific
Nextera DNA Sample Prep Kit (Roche FLX-compatible)
 Cat. Nos. FL09115, FL091120, and FLBC0950 The is designed to prepare genomic DNA libraries compatible with the Roche 454 Genome Sequencer FLX System, with standard FLX chemistry. Nextera technology* employs
Cat. Nos. FL09115, FL091120, and FLBC0950 The is designed to prepare genomic DNA libraries compatible with the Roche 454 Genome Sequencer FLX System, with standard FLX chemistry. Nextera technology* employs
Supplementary Information. Arrays of Individual DNA Molecules on Nanopatterned Substrates
 Supplementary Information Arrays of Individual DNA Molecules on Nanopatterned Substrates Roland Hager, Alma Halilovic, Jonathan R. Burns, Friedrich Schäffler, Stefan Howorka S1 Figure S-1. Characterization
Supplementary Information Arrays of Individual DNA Molecules on Nanopatterned Substrates Roland Hager, Alma Halilovic, Jonathan R. Burns, Friedrich Schäffler, Stefan Howorka S1 Figure S-1. Characterization
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/2/2/e1501254/dc1 Supplementary Materials for Dramatic performance of Clostridium thermocellum explained by its wide range of cellulase modalities Qi Xu, Michael
advances.sciencemag.org/cgi/content/full/2/2/e1501254/dc1 Supplementary Materials for Dramatic performance of Clostridium thermocellum explained by its wide range of cellulase modalities Qi Xu, Michael
Description...1 Components...1 Storage... 1 Technical Information...1 Protocol...2 Examples using the kit...4 Troubleshooting...
 QuickClean II Gel Extraction Kit Cat. No. L00418 Technical Manual No. TM0594 Version: 03042011 I II III IV V VI VII VIII Description......1 Components.....1 Storage.... 1 Technical Information....1 Protocol.....2
QuickClean II Gel Extraction Kit Cat. No. L00418 Technical Manual No. TM0594 Version: 03042011 I II III IV V VI VII VIII Description......1 Components.....1 Storage.... 1 Technical Information....1 Protocol.....2
Nature Methods: doi: /nmeth Supplementary Figure 1. Retention of RNA with LabelX.
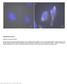 Supplementary Figure 1 Retention of RNA with LabelX. (a) Epi-fluorescence image of single molecule FISH (smfish) against GAPDH on HeLa cells expanded without LabelX treatment. (b) Epi-fluorescence image
Supplementary Figure 1 Retention of RNA with LabelX. (a) Epi-fluorescence image of single molecule FISH (smfish) against GAPDH on HeLa cells expanded without LabelX treatment. (b) Epi-fluorescence image
2/10/17. Contents. Applications of HMMs in Epigenomics
 2/10/17 I529: Machine Learning in Bioinformatics (Spring 2017) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2017 Background:
2/10/17 I529: Machine Learning in Bioinformatics (Spring 2017) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2017 Background:
EZ Vision DNA Dye as Loading Buffer
 EZ Vision DNA Dye as Loading Buffer Code Description Size N472-1ML N472-KIT N650-1ML N650-KIT N313-1ML N313-KIT N473-2PK N473-3PK EZ Vision One DNA Dye as Loading Buffer, 6X EZ Vision Two, DNA Dye as Loading
EZ Vision DNA Dye as Loading Buffer Code Description Size N472-1ML N472-KIT N650-1ML N650-KIT N313-1ML N313-KIT N473-2PK N473-3PK EZ Vision One DNA Dye as Loading Buffer, 6X EZ Vision Two, DNA Dye as Loading
Data Sheet Quick PCR Cloning Kit
 Data Sheet Quick PCR Cloning Kit 6044 Cornerstone Ct. West, Ste. E DESCRIPTION: The Quick PCR Cloning Kit is a simple and highly efficient method to insert any gene or DNA fragment into a vector, without
Data Sheet Quick PCR Cloning Kit 6044 Cornerstone Ct. West, Ste. E DESCRIPTION: The Quick PCR Cloning Kit is a simple and highly efficient method to insert any gene or DNA fragment into a vector, without
Result Tables The Result Table, which indicates chromosomal positions and annotated gene names, promoter regions and CpG islands, is the best way for
 Result Tables The Result Table, which indicates chromosomal positions and annotated gene names, promoter regions and CpG islands, is the best way for you to discover methylation changes at specific genomic
Result Tables The Result Table, which indicates chromosomal positions and annotated gene names, promoter regions and CpG islands, is the best way for you to discover methylation changes at specific genomic
Bi 8 Lecture 4. Ellen Rothenberg 14 January Reading: from Alberts Ch. 8
 Bi 8 Lecture 4 DNA approaches: How we know what we know Ellen Rothenberg 14 January 2016 Reading: from Alberts Ch. 8 Central concept: DNA or RNA polymer length as an identifying feature RNA has intrinsically
Bi 8 Lecture 4 DNA approaches: How we know what we know Ellen Rothenberg 14 January 2016 Reading: from Alberts Ch. 8 Central concept: DNA or RNA polymer length as an identifying feature RNA has intrinsically
Targeted RNA sequencing reveals the deep complexity of the human transcriptome.
 Targeted RNA sequencing reveals the deep complexity of the human transcriptome. Tim R. Mercer 1, Daniel J. Gerhardt 2, Marcel E. Dinger 1, Joanna Crawford 1, Cole Trapnell 3, Jeffrey A. Jeddeloh 2,4, John
Targeted RNA sequencing reveals the deep complexity of the human transcriptome. Tim R. Mercer 1, Daniel J. Gerhardt 2, Marcel E. Dinger 1, Joanna Crawford 1, Cole Trapnell 3, Jeffrey A. Jeddeloh 2,4, John
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature08627 Supplementary Figure 1. DNA sequences used to construct nucleosomes in this work. a, DNA sequences containing the 601 positioning sequence (blue)24 with a PstI restriction site
doi: 10.1038/nature08627 Supplementary Figure 1. DNA sequences used to construct nucleosomes in this work. a, DNA sequences containing the 601 positioning sequence (blue)24 with a PstI restriction site
APPLICATION NOTE. Abstract. Introduction
 From minuscule amounts to magnificent results: reliable ChIP-seq data from 1, cells with the True MicroChIP and the MicroPlex Library Preparation kits Abstract Diagenode has developed groundbreaking solutions
From minuscule amounts to magnificent results: reliable ChIP-seq data from 1, cells with the True MicroChIP and the MicroPlex Library Preparation kits Abstract Diagenode has developed groundbreaking solutions
ChIP-Seq Tools. J Fass UCD Genome Center Bioinformatics Core Wednesday September 16, 2015
 ChIP-Seq Tools J Fass UCD Genome Center Bioinformatics Core Wednesday September 16, 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA or
ChIP-Seq Tools J Fass UCD Genome Center Bioinformatics Core Wednesday September 16, 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA or
ChIP-Seq Data Analysis. J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015
 ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA
ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA
Supplementary Methods pcfd5 cloning protocol
 Supplementary Methods cloning protocol vermilion trna grna trna grna U6:3 Terminator AmpR attb is a vector for expressing one or multiple trna-flanked Cas9 grnas under the control of the strong, ubiquitous
Supplementary Methods cloning protocol vermilion trna grna trna grna U6:3 Terminator AmpR attb is a vector for expressing one or multiple trna-flanked Cas9 grnas under the control of the strong, ubiquitous
TaKaRa MiniBEST Agarose Gel DNA Extraction Kit Ver.4.0
 Cat. # 9762 For Research Use TaKaRa MiniBEST Agarose Gel Product Manual Table of Contents I. Description... 3 II. Kit Components... 3 III. Shipping and Storage... 3 IV. Preparation before Use... 4 V. Protocol...
Cat. # 9762 For Research Use TaKaRa MiniBEST Agarose Gel Product Manual Table of Contents I. Description... 3 II. Kit Components... 3 III. Shipping and Storage... 3 IV. Preparation before Use... 4 V. Protocol...
Nextera DNA Sample Prep Kit
 Nextera DNA Sample Prep Kit (Roche FLX-compatible) Cat. Nos. FL09115, FL091120, and FLBC0950 * Covered by patents issued and pending. Connect with Epicentre on our blog (epicentral.blogspot.com), Facebook
Nextera DNA Sample Prep Kit (Roche FLX-compatible) Cat. Nos. FL09115, FL091120, and FLBC0950 * Covered by patents issued and pending. Connect with Epicentre on our blog (epicentral.blogspot.com), Facebook
ChIP-Seq Data Analysis. J Fass UCD Genome Center Bioinformatics Core Wednesday December 17, 2014
 ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday December 17, 2014 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind
ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday December 17, 2014 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind
Huijuan Feng, Shining Ma,Chao Ye & Zhixing Feng
 Huijuan Feng, Shining Ma,Chao Ye & Zhixing Feng Background-Author introduction Research interest: Methods for gene mapping of complex traits Inference of population structure from genetic data Genome variation
Huijuan Feng, Shining Ma,Chao Ye & Zhixing Feng Background-Author introduction Research interest: Methods for gene mapping of complex traits Inference of population structure from genetic data Genome variation
Chapter 7. DNA Microarrays
 Bioinformatics III Structural Bioinformatics and Genome Analysis Chapter 7. DNA Microarrays 7.9 Next Generation Sequencing 454 Sequencing Solexa Illumina Solid TM System Sequencing Process of determining
Bioinformatics III Structural Bioinformatics and Genome Analysis Chapter 7. DNA Microarrays 7.9 Next Generation Sequencing 454 Sequencing Solexa Illumina Solid TM System Sequencing Process of determining
ChIP. November 21, 2017
 ChIP November 21, 2017 functional signals: is DNA enough? what is the smallest number of letters used by a written language? DNA is only one part of the functional genome DNA is heavily bound by proteins,
ChIP November 21, 2017 functional signals: is DNA enough? what is the smallest number of letters used by a written language? DNA is only one part of the functional genome DNA is heavily bound by proteins,
XactEdit Cas9 Nuclease with NLS User Manual
 XactEdit Cas9 Nuclease with NLS User Manual An RNA-guided recombinant endonuclease for efficient targeted DNA cleavage Catalog Numbers CE1000-50K, CE1000-50, CE1000-250, CE1001-250, CE1001-1000 Table of
XactEdit Cas9 Nuclease with NLS User Manual An RNA-guided recombinant endonuclease for efficient targeted DNA cleavage Catalog Numbers CE1000-50K, CE1000-50, CE1000-250, CE1001-250, CE1001-1000 Table of
Bead Type (NaI) Gel Extraction Kits
 Bead Type (NaI) Contents Kit Contents Principle Important Notes Bead Type (NaI) Gel Extraction Kit Protocol Troubleshooting Guide 2 3 3 4 5 Ordering Information 6 Kit Contents Catalog No. Number of preparations
Bead Type (NaI) Contents Kit Contents Principle Important Notes Bead Type (NaI) Gel Extraction Kit Protocol Troubleshooting Guide 2 3 3 4 5 Ordering Information 6 Kit Contents Catalog No. Number of preparations
Complete protocol in 110 minutes Enzymatic fragmentation without sonication One-step fragmentation/tagging to save time
 Molecular Cloning Laboratories Manual Version 1.2 Product name: MCNext UT DNA Sample Prep Kit Cat #: MCUDS-4, MCUDS-24, MCUDS-96 Description: This protocol explains how to prepare up to 96 pooled indexed
Molecular Cloning Laboratories Manual Version 1.2 Product name: MCNext UT DNA Sample Prep Kit Cat #: MCUDS-4, MCUDS-24, MCUDS-96 Description: This protocol explains how to prepare up to 96 pooled indexed
Epigenetics and DNase-Seq
 Epigenetics and DNase-Seq BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2018 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material, are licensed under CC BY-NC 4.0 by Anthony
Epigenetics and DNase-Seq BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2018 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material, are licensed under CC BY-NC 4.0 by Anthony
Nature Genetics: doi: /ng Supplementary Figure 1. ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets.
 Supplementary Figure 1 ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets. Gene structures are shown underneath each panel. Supplementary Figure 2 pref6::ref6-gfp complements
Supplementary Figure 1 ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets. Gene structures are shown underneath each panel. Supplementary Figure 2 pref6::ref6-gfp complements
Supplementary Figure 1 An overview of pirna biogenesis during fetal mouse reprogramming. (a) (b)
 Supplementary Figure 1 An overview of pirna biogenesis during fetal mouse reprogramming. (a) A schematic overview of the production and amplification of a single pirna from a transposon transcript. The
Supplementary Figure 1 An overview of pirna biogenesis during fetal mouse reprogramming. (a) A schematic overview of the production and amplification of a single pirna from a transposon transcript. The
Analysis of ChIP-seq data with R / Bioconductor
 Analysis of ChIP-seq data with R / Bioconductor Martin Morgan Bioconductor / Fred Hutchinson Cancer Research Center Seattle, WA, USA 8-10 June 2009 ChIP-seq Chromatin immunopreciptation to enrich sample
Analysis of ChIP-seq data with R / Bioconductor Martin Morgan Bioconductor / Fred Hutchinson Cancer Research Center Seattle, WA, USA 8-10 June 2009 ChIP-seq Chromatin immunopreciptation to enrich sample
Supporting Information
 Supporting Information Gómez-Marín et al. 10.1073/pnas.1505463112 SI Materials and Methods Generation of BAC DK74B2-six2a::GFP-six3a::mCherry-iTol2. BAC clone (number 74B2) from DanioKey zebrafish BAC
Supporting Information Gómez-Marín et al. 10.1073/pnas.1505463112 SI Materials and Methods Generation of BAC DK74B2-six2a::GFP-six3a::mCherry-iTol2. BAC clone (number 74B2) from DanioKey zebrafish BAC
Factors affecting PCR
 Lec. 11 Dr. Ahmed K. Ali Factors affecting PCR The sequences of the primers are critical to the success of the experiment, as are the precise temperatures used in the heating and cooling stages of the
Lec. 11 Dr. Ahmed K. Ali Factors affecting PCR The sequences of the primers are critical to the success of the experiment, as are the precise temperatures used in the heating and cooling stages of the
The preparation of native chromatin from cultured human cells.
 Native chromatin immunoprecipitation protocol The preparation of native chromatin from cultured human cells. All solutions need to be ice cold. Sucrose containing solutions must be made up fresh on the
Native chromatin immunoprecipitation protocol The preparation of native chromatin from cultured human cells. All solutions need to be ice cold. Sucrose containing solutions must be made up fresh on the
Supplementary information Bi-specific Aptamers mediating Tumour Cell Lysis
 Supplementary information Bi-specific Aptamers mediating Tumour Cell Lysis Figure S1 SELEX scheme of different CD16 DNA selection approaches. Nine rounds of filter SELEX were conducted in parallel, only
Supplementary information Bi-specific Aptamers mediating Tumour Cell Lysis Figure S1 SELEX scheme of different CD16 DNA selection approaches. Nine rounds of filter SELEX were conducted in parallel, only
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Endogenous gene tagging to study subcellular localization and chromatin binding. a, b, Schematic of experimental set-up to endogenously tag RNAi factors using the CRISPR Cas9 technology,
Supplementary Figure 1 Endogenous gene tagging to study subcellular localization and chromatin binding. a, b, Schematic of experimental set-up to endogenously tag RNAi factors using the CRISPR Cas9 technology,
Gene expression analysis. Biosciences 741: Genomics Fall, 2013 Week 5. Gene expression analysis
 Gene expression analysis Biosciences 741: Genomics Fall, 2013 Week 5 Gene expression analysis From EST clusters to spotted cdna microarrays Long vs. short oligonucleotide microarrays vs. RT-PCR Methods
Gene expression analysis Biosciences 741: Genomics Fall, 2013 Week 5 Gene expression analysis From EST clusters to spotted cdna microarrays Long vs. short oligonucleotide microarrays vs. RT-PCR Methods
Information on barcode decoding
 Information on barcode decoding The Bioneer version 1.0 haploid deletion library (Bioneer catalog number M-1030H) was supplied as glycerol frozen stocks in thirty-one 96-well plates. According to the information
Information on barcode decoding The Bioneer version 1.0 haploid deletion library (Bioneer catalog number M-1030H) was supplied as glycerol frozen stocks in thirty-one 96-well plates. According to the information
