PIP-seq. Cells. Permanganate ChIP-Seq
|
|
|
- Miranda Houston
- 6 years ago
- Views:
Transcription
1 PIP-seq ells Formaldehyde Permanganate 5 Harvest Lyse Sonicate First dapter Ligation hip Elute Reverse rosslinks Piperidine cleavage Primer Extension Second dapter Ligation Deep Sequencing Permanganate hip-seq Supplementary Figure. PIP-seq protocol schema Shown is an illustration of the PIP-seq assay protocol as previously descried. In short, cells are cross-linked with formaldehyde and treated with potassium permanganate. hey are then lysed and sonicated to fragment sizes etween 2-5 p. n Illumina-compatile adapter is then ligated to the ends after end-repair and -tailing. he sample is then eluted from the antiody and crosslinks are reversed. Piperidine is then used to cleave oxidized thymines. Strands are denatured and a primer is annealed to the adapters, followed y second-strand synthesis (primer extension). second adapter is susequently ligated to the piperidine-cleaved end, via -tailing. he resulting lirary is then quantified and sequenced.
2 SS nucleotide frequency 26,63 Human SS FIIB op 25% Bottom 25%.4.4 solute Frequency solute Frequency c Log 2 ratio (normalized tag count) ll peaks (,) Pass threshold (95) Fail threshold (49) hip-exo PIP-seq RO-cap Supplementary Figure 2. Nucleotide frequencies surrounding annotated human SS (a) Heatmaps of nucleotide frequency within p of annotated SSs (N=26,63). Plots are sorted y FIIB PIP-seq tag counts located within 25 p of each SS. Only those tag 5 ends that mapped just 3 to a were counted, when sorting. () omposite plots from panel (a), separated out into the top 25% of FIIB PIP-seq peak tag counts versus the ottom 25%. (c) Log 2 ratio of the average called peak score for over the average random peak score for FIIB hip-exo, FIIB PIP-seq, and RO-cap
3 - position 5 end ag Reference genome FIIB PIP-seq c 5 Nucleotide occurence 26,63 Human SS 26,63 Human SS FIIB hip-exo d Relative Frequency Relative Frequency Normalized U.5 FIIB PIP-seq : FIIB hip-exo :. :.92 PIP-seq Nucleotide U :.48 :.4 :.2 :.9 :.69 hip-exo Supplementary Figure 3. FIIB PIP-seq validation (a) Schematic showing how reads were filtered y the specific - nucleotide 5 the to aligned sequence read () Heatmap distriution of FIIB PIP-seq (top) or hip-exo (ottom) tag 5 ends relative to all annotated human SSs 2, and separated out y the type of nucleotide (,,, ) occurring nucleotide upstream from the 5 end (i.e., - position). Plots are sorted y FIIB PIP-seq tag counts (that also have a - ) within 25 p of each SS. (c) omposite (average) of panel (a). Relative peak height values (mode) for each composite are reported. (d) rea under the curve (U) from panel normalized y the local nucleotide content, calculated as Σ(vg reads ±25 p window around SS) / (vg nucleotide frequency ±25 p window around SS)
4 - position 5 end ag Reference genome Pol II PIP-seq c 5 Nucleotide occurence 26,63 Human SS 26,63 Human SS Pol II hip-exo d Relative Frequency Relative Frequency Normalized U PIP-seq Pol II PIP-seq :. Pol II hip-exo :. : Nucleotide U :.49 :.39 :.7 :.45 :.42 hip-exo Supplementary Figure 4. Pol II PIP-seq validation (a-d) Same as Supplemental Figure 3, except the analysis was on Pol II instead of FIIB.
5 a FIIB FIIB signal Forward 25 Motif Occurence Reverse.5 Nucleosome Distance from FIIB PIP-seq peak (p) Motif Occurrence Nucleosome Randomized MNase-seq FIIB PIP-seq peaks (N=8,34) 25 RO-cap & MNase-seq signal Relative Motif Occurence FIIB PIP-seq peaks (N=8,34) Distance from FIIB PIP-seq peak (p) Motif Occurrence RO-cap PIP-seq Pol II Randomized RO-cap/Motif FIIB PIP-seq peaks (N=8,34) FIIB Distance from FIIB PIP-seq peak (p) Supplementary Figure 5. Motif enrichment relative to PI (a) omposite plots of FIIB PIP-seq, RO-cap, MNase-seq, and the enriched motif occurrence count and heatmaps of MNase-seq and enriched motif occurrence count relative to called FIIB PIP-seq peaks. JSPR 26 verterate motifs were scanned using FIMO in a 2 k window relative to FIIB PIP-seq peaks. ontrol data was generated y randomizing the sequence from each 2 k window. alled motifs were aligned relative to called peaks and sorted y distance of FIIB PIP-seq peak to called + nucleosome. () Heatmaps of PIP-seq data displaying the overlap of the motif occurrence and the RO-cap data. he lack line is the location of the called + nucleosome dyad and the grey line is exactly 73 p upstream representing the upper edge of the nucleosome.
6 Pol II PIP-seq ( / ) ( / ) 5 Nucleotide ratio FIIB PIP-seq 8 4 ( / ) ( / ) Pol II hip-exo FIIB hip-exo 8 Nucleotide ratio 5 ( / ) 4 ( / ) ( / ) ( / ) Supplementary Figure 6. PIP-seq strand normalization validation (a) Pol II and FIIB PIP-seq - tag counts were aligned relative to annotated human SSs (N=26,63), smoothed with a 7 p sliding window average, and then locally normalized (divided) y - tag counts treated similarly (/) so as to report on the relative -reactivity relative to the equally occurring nucleotide, as a readout of single-stranded DN. Similar control plots were made for and tags (/). () Pol II and FIIB hip-exo - N tags were normalized similarly as panel a.
7 c Proportion mrn Promoter ncrn Enhancer Insulator ranscription hromhmm states Repressed Heterochromatin O Biological Process mrn FIIB m RN m etaolic process translation nuclear-transcried m RN cataolic process m RN cataolic process translational initiation RN cataolic process translational term ination nuclear-transcried m RN cataolic process, nonsense-m ediated decay viral transcription viral gene expression Distal ncrn FIIB negative regulation of cytokine production regulation of adaptive immune response intrinsic apoptotic signaling pathway Fc-gamma receptor signaling pathway involved in phagocytosis response to type I interferon JK-S cascade regulation of D4-positive, alpha-eta cell differentiation negative regulation of stem cell differentiation growth horm one receptor signaling pathway regulat ion of D4-positive, alpha-eta cell activation p3 Occupancy 5 25 ncrn mrn Distance from FIIB (p) log(binom ial p value) Supplementary Figure 7. ncrn and mrn FIIB meta-analysis (a) Proportion of predicted regulatory functions ased on chromatin state maps for mrn and ncrn FIIB locations 3. () omposite plot of ENODE p3 hip-seq occupancy plotted relative to mrn and ncrn FIIB PIP-seq peaks 4. (c) O enrichment of closest annotated genes for mrn and distal ncrn FIIB locations 5.
8 Supplementary ale : Sequencing Experiment Statistics!!! ssay! Replicate! otal!ags! otal!''!ags!! Pol$II$ PIP&seq$ $ $62,98,427$$ $26,593,623$$ Pol$II$ PIP&seq$ 2$ $,845,45$$ $4,949,54$$ FIIB$ PIP&seq$ $ $43,354,58$$ $5,486,376$$ FIIB$ PIP&seq$ 2$ $8,579,59$$ $7,258,22$$ FIIB$ PIP&seq$ 3$ $2,444,57$$ $4,89,$$ Input$ PIP&seq$ $ $4,23,956$$ $,39,29$$ Pol$II$ hip&exo$ $ $9,743,244$$ $&$$ FIIB$ hip&exo$ $ $4,723,536$$ $&$$
9 SUPPLEMENL*REFERENES*.# Li,#J.!et!al.#Kinetic#competition#etween#elongation#rate#and#inding#of#NELF# controls#promoter<proximal#pausing.#mol!ell#5,#7<22#(23).# 2.# Pruitt,#K.D.,#atusova,#.#&#Maglott,#D.R.#NBI#reference#sequences#(RefSeq):#a# curated#non<redundant#sequence#dataase#of#genomes,#transcripts#and# proteins.#nucleic!cids!res#35,#d6<5#(27).# 3.# Ernst,#J.!et!al.#Mapping#and#analysis#of#chromatin#state#dynamics#in#nine# human#cell#types.#nature#473,#43<9#(2).# 4.# onsortium,#e.p.#n#integrated#encyclopedia#of#dn#elements#in#the#human# genome.#nature#489,#57<74#(22).# 5.# McLean,#.Y.!et!al.#RE#improves#functional#interpretation#of#cis< regulatory#regions.#nat!biotechnol#28,#495<5#(2).# #
SUPPLEMENTARY INFORMATION
 SUPPLEMENRY INFORMION doi:.38/nature In vivo nucleosome mapping D4+ Lymphocytes radient-based and I-bead cell sorting D8+ Lymphocytes ranulocytes Lyse the cells Isolate and sequence mononucleosome cores
SUPPLEMENRY INFORMION doi:.38/nature In vivo nucleosome mapping D4+ Lymphocytes radient-based and I-bead cell sorting D8+ Lymphocytes ranulocytes Lyse the cells Isolate and sequence mononucleosome cores
SUPPLEMENTAL MATERIALS
 SUPPLEMENL MERILS Eh-seq: RISPR epitope tagging hip-seq of DN-binding proteins Daniel Savic, E. hristopher Partridge, Kimberly M. Newberry, Sophia. Smith, Sarah K. Meadows, rian S. Roberts, Mark Mackiewicz,
SUPPLEMENL MERILS Eh-seq: RISPR epitope tagging hip-seq of DN-binding proteins Daniel Savic, E. hristopher Partridge, Kimberly M. Newberry, Sophia. Smith, Sarah K. Meadows, rian S. Roberts, Mark Mackiewicz,
ChIP-Seq Data Analysis. J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015
 ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA
ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA
ChIP-Seq Tools. J Fass UCD Genome Center Bioinformatics Core Wednesday September 16, 2015
 ChIP-Seq Tools J Fass UCD Genome Center Bioinformatics Core Wednesday September 16, 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA or
ChIP-Seq Tools J Fass UCD Genome Center Bioinformatics Core Wednesday September 16, 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA or
ChIP-Seq Data Analysis. J Fass UCD Genome Center Bioinformatics Core Wednesday December 17, 2014
 ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday December 17, 2014 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind
ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday December 17, 2014 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind
Applied Bioinformatics - Lecture 16: Transcriptomics
 Applied Bioinformatics - Lecture 16: Transcriptomics David Hendrix Oregon State University Feb 15th 2016 Transcriptomics High-throughput Sequencing (deep sequencing) High-throughput sequencing (also
Applied Bioinformatics - Lecture 16: Transcriptomics David Hendrix Oregon State University Feb 15th 2016 Transcriptomics High-throughput Sequencing (deep sequencing) High-throughput sequencing (also
APPLICATION NOTE. Abstract. Introduction
 From minuscule amounts to magnificent results: reliable ChIP-seq data from 1, cells with the True MicroChIP and the MicroPlex Library Preparation kits Abstract Diagenode has developed groundbreaking solutions
From minuscule amounts to magnificent results: reliable ChIP-seq data from 1, cells with the True MicroChIP and the MicroPlex Library Preparation kits Abstract Diagenode has developed groundbreaking solutions
Supplementary Figure 1 Strategy for parallel detection of DHSs and adjacent nucleosomes
 Supplementary Figure 1 Strategy for parallel detection of DHSs and adjacent nucleosomes DNase I cleavage DNase I DNase I digestion Sucrose gradient enrichment Small Large F1 F2...... F9 F1 F1 F2 F3 F4
Supplementary Figure 1 Strategy for parallel detection of DHSs and adjacent nucleosomes DNase I cleavage DNase I DNase I digestion Sucrose gradient enrichment Small Large F1 F2...... F9 F1 F1 F2 F3 F4
Introduction to ChIP Seq data analyses. Acknowledgement: slides taken from Dr. H
 Introduction to ChIP Seq data analyses Acknowledgement: slides taken from Dr. H Wu @Emory ChIP seq: Chromatin ImmunoPrecipitation it ti + sequencing Same biological motivation as ChIP chip: measure specific
Introduction to ChIP Seq data analyses Acknowledgement: slides taken from Dr. H Wu @Emory ChIP seq: Chromatin ImmunoPrecipitation it ti + sequencing Same biological motivation as ChIP chip: measure specific
Novel methods for RNA and DNA- Seq analysis using SMART Technology. Andrew Farmer, D. Phil. Vice President, R&D Clontech Laboratories, Inc.
 Novel methods for RNA and DNA- Seq analysis using SMART Technology Andrew Farmer, D. Phil. Vice President, R&D Clontech Laboratories, Inc. Agenda Enabling Single Cell RNA-Seq using SMART Technology SMART
Novel methods for RNA and DNA- Seq analysis using SMART Technology Andrew Farmer, D. Phil. Vice President, R&D Clontech Laboratories, Inc. Agenda Enabling Single Cell RNA-Seq using SMART Technology SMART
ChIP. November 21, 2017
 ChIP November 21, 2017 functional signals: is DNA enough? what is the smallest number of letters used by a written language? DNA is only one part of the functional genome DNA is heavily bound by proteins,
ChIP November 21, 2017 functional signals: is DNA enough? what is the smallest number of letters used by a written language? DNA is only one part of the functional genome DNA is heavily bound by proteins,
ChIP-seq/Functional Genomics/Epigenomics. CBSU/3CPG/CVG Next-Gen Sequencing Workshop. Josh Waterfall. March 31, 2010
 ChIP-seq/Functional Genomics/Epigenomics CBSU/3CPG/CVG Next-Gen Sequencing Workshop Josh Waterfall March 31, 2010 Outline Introduction to ChIP-seq Control data sets Peak/enriched region identification
ChIP-seq/Functional Genomics/Epigenomics CBSU/3CPG/CVG Next-Gen Sequencing Workshop Josh Waterfall March 31, 2010 Outline Introduction to ChIP-seq Control data sets Peak/enriched region identification
ChIP-seq data analysis with Chipster. Eija Korpelainen CSC IT Center for Science, Finland
 ChIP-seq data analysis with Chipster Eija Korpelainen CSC IT Center for Science, Finland chipster@csc.fi What will I learn? Short introduction to ChIP-seq Analyzing ChIP-seq data Central concepts Analysis
ChIP-seq data analysis with Chipster Eija Korpelainen CSC IT Center for Science, Finland chipster@csc.fi What will I learn? Short introduction to ChIP-seq Analyzing ChIP-seq data Central concepts Analysis
Nature Methods: doi: /nmeth.4396
 Supplementary Figure 1 Comparison of technical replicate consistency between and across the standard ATAC-seq method, DNase-seq, and Omni-ATAC. (a) Heatmap-based representation of ATAC-seq quality control
Supplementary Figure 1 Comparison of technical replicate consistency between and across the standard ATAC-seq method, DNase-seq, and Omni-ATAC. (a) Heatmap-based representation of ATAC-seq quality control
ChIP-seq analysis 2/28/2018
 ChIP-seq analysis 2/28/2018 Acknowledgements Much of the content of this lecture is from: Furey (2012) ChIP-seq and beyond Park (2009) ChIP-seq advantages + challenges Landt et al. (2012) ChIP-seq guidelines
ChIP-seq analysis 2/28/2018 Acknowledgements Much of the content of this lecture is from: Furey (2012) ChIP-seq and beyond Park (2009) ChIP-seq advantages + challenges Landt et al. (2012) ChIP-seq guidelines
Analysis of ChIP-seq data with R / Bioconductor
 Analysis of ChIP-seq data with R / Bioconductor Martin Morgan Bioconductor / Fred Hutchinson Cancer Research Center Seattle, WA, USA 8-10 June 2009 ChIP-seq Chromatin immunopreciptation to enrich sample
Analysis of ChIP-seq data with R / Bioconductor Martin Morgan Bioconductor / Fred Hutchinson Cancer Research Center Seattle, WA, USA 8-10 June 2009 ChIP-seq Chromatin immunopreciptation to enrich sample
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Origin use and efficiency are similar among WT, rrm3, pif1-m2, and pif1-m2; rrm3 strains. A. Analysis of fork progression around confirmed and likely origins (from cerevisiae.oridb.org).
Supplementary Figure 1 Origin use and efficiency are similar among WT, rrm3, pif1-m2, and pif1-m2; rrm3 strains. A. Analysis of fork progression around confirmed and likely origins (from cerevisiae.oridb.org).
1 Najafabadi, H. S. et al. C2H2 zinc finger proteins greatly expand the human regulatory lexicon. Nat Biotechnol doi: /nbt.3128 (2015).
 F op-scoring motif Optimized motifs E Input sequences entral 1 bp region Dinucleotideshuffled seqs B D ll B1H-R predicted motifs Enriched B1H- R predicted motifs L!=!7! L!=!6! L!=5! L!=!4! L!=!3! L!=!2!
F op-scoring motif Optimized motifs E Input sequences entral 1 bp region Dinucleotideshuffled seqs B D ll B1H-R predicted motifs Enriched B1H- R predicted motifs L!=!7! L!=!6! L!=5! L!=!4! L!=!3! L!=!2!
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Endogenous gene tagging to study subcellular localization and chromatin binding. a, b, Schematic of experimental set-up to endogenously tag RNAi factors using the CRISPR Cas9 technology,
Supplementary Figure 1 Endogenous gene tagging to study subcellular localization and chromatin binding. a, b, Schematic of experimental set-up to endogenously tag RNAi factors using the CRISPR Cas9 technology,
Figure S1: NUN preparation yields nascent, unadenylated RNA with a different profile from Total RNA.
 Summary of Supplemental Information Figure S1: NUN preparation yields nascent, unadenylated RNA with a different profile from Total RNA. Figure S2: rrna removal procedure is effective for clearing out
Summary of Supplemental Information Figure S1: NUN preparation yields nascent, unadenylated RNA with a different profile from Total RNA. Figure S2: rrna removal procedure is effective for clearing out
Le proteine regolative variano nei vari tipi cellulari e in funzione degli stimoli ambientali
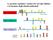 Le proteine regolative variano nei vari tipi cellulari e in funzione degli stimoli ambientali Tipo cellulare 1 Tipo cellulare 2 Tipo cellulare 3 DNA-protein Crosslink Lisi Frammentazione Immunopurificazione
Le proteine regolative variano nei vari tipi cellulari e in funzione degli stimoli ambientali Tipo cellulare 1 Tipo cellulare 2 Tipo cellulare 3 DNA-protein Crosslink Lisi Frammentazione Immunopurificazione
THE ANALYSIS OF CHROMATIN CONDENSATION STATE AND TRANSCRIPTIONAL ACTIVITY USING DNA MICROARRAYS 1. INTRODUCTION
 JOURNAL OF MEDICAL INFORMATICS & TECHNOLOGIES Vol.6/2003, ISSN 1642-6037 Piotr WIDŁAK *, Krzysztof FUJAREWICZ ** DNA microarrays, chromatin, transcription THE ANALYSIS OF CHROMATIN CONDENSATION STATE AND
JOURNAL OF MEDICAL INFORMATICS & TECHNOLOGIES Vol.6/2003, ISSN 1642-6037 Piotr WIDŁAK *, Krzysztof FUJAREWICZ ** DNA microarrays, chromatin, transcription THE ANALYSIS OF CHROMATIN CONDENSATION STATE AND
Computational Analysis of Ultra-high-throughput sequencing data: ChIP-Seq
 Computational Analysis of Ultra-high-throughput sequencing data: ChIP-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne Data flow in ChIP-Seq data analysis Level 1:
Computational Analysis of Ultra-high-throughput sequencing data: ChIP-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne Data flow in ChIP-Seq data analysis Level 1:
CELL BIOLOGY - CLUTCH CH. 7 - GENE EXPRESSION.
 !! www.clutchprep.com CONCEPT: CONTROL OF GENE EXPRESSION BASICS Gene expression is the process through which cells selectively to express some genes and not others Every cell in an organism is a clone
!! www.clutchprep.com CONCEPT: CONTROL OF GENE EXPRESSION BASICS Gene expression is the process through which cells selectively to express some genes and not others Every cell in an organism is a clone
SUPPLEMENTARY INFORMATION
 doi:1.138/nature1895 i Core histones CAF-1 v ii Reb1 CAF-1 Fen1 vi Mature Okazaki fragment iii PCNA Reb1 iv vii Mature Okazaki fragment Reb1 Mature Okazaki fragments Figure S1. Model for nucleosome-mediated
doi:1.138/nature1895 i Core histones CAF-1 v ii Reb1 CAF-1 Fen1 vi Mature Okazaki fragment iii PCNA Reb1 iv vii Mature Okazaki fragment Reb1 Mature Okazaki fragments Figure S1. Model for nucleosome-mediated
Supplementary Figure S1 Supplementary Figure S2 Supplementary Figure S3. Supplementary Figure S4
 Supplementary Figure S1 Supplementary Figure S2 Supplementary Figure S3 Supplementary Figure S4 Supplementary Figure S5 Supplementary Figure S6 Supplementary Figure S7 Supplementary Figure S8 Supplementary
Supplementary Figure S1 Supplementary Figure S2 Supplementary Figure S3 Supplementary Figure S4 Supplementary Figure S5 Supplementary Figure S6 Supplementary Figure S7 Supplementary Figure S8 Supplementary
TECH NOTE Ligation-Free ChIP-Seq Library Preparation
 TECH NOTE Ligation-Free ChIP-Seq Library Preparation The DNA SMART ChIP-Seq Kit Ligation-free template switching technology: Minimize sample handling in a single-tube workflow >> Simplified protocol with
TECH NOTE Ligation-Free ChIP-Seq Library Preparation The DNA SMART ChIP-Seq Kit Ligation-free template switching technology: Minimize sample handling in a single-tube workflow >> Simplified protocol with
Supplemental Figure 1 A
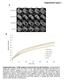 Supplemental Figure A prebleach postbleach 2 min 6 min 3 min mh2a.-gfp mh2a.2-gfp mh2a2-gfp GFP-H2A..9 Relative Intensity.8.7.6.5 mh2a. GFP n=8.4 mh2a.2 GFP n=4.3 mh2a2 GFP n=2.2 GFP H2A n=24. GFP n=7.
Supplemental Figure A prebleach postbleach 2 min 6 min 3 min mh2a.-gfp mh2a.2-gfp mh2a2-gfp GFP-H2A..9 Relative Intensity.8.7.6.5 mh2a. GFP n=8.4 mh2a.2 GFP n=4.3 mh2a2 GFP n=2.2 GFP H2A n=24. GFP n=7.
Considerations for Illumina library preparation. Henriette O Geen June 20, 2014 UCD Genome Center
 Considerations for Illumina library preparation Henriette O Geen June 20, 2014 UCD Genome Center Diversity of applications De novo genome Sequencing ranscriptome Expression Splice Isoform bundance Genotyping
Considerations for Illumina library preparation Henriette O Geen June 20, 2014 UCD Genome Center Diversity of applications De novo genome Sequencing ranscriptome Expression Splice Isoform bundance Genotyping
Supplemental Figure 1.
 Supplemental Data. Charron et al. Dynamic landscapes of four histone modifications during de-etiolation in Arabidopsis. Plant Cell (2009). 10.1105/tpc.109.066845 Supplemental Figure 1. Immunodetection
Supplemental Data. Charron et al. Dynamic landscapes of four histone modifications during de-etiolation in Arabidopsis. Plant Cell (2009). 10.1105/tpc.109.066845 Supplemental Figure 1. Immunodetection
Supplementary Table 1: Oligo designs. A list of ATAC-seq oligos used for PCR.
 Ad1_noMX: Ad2.1_TAAGGCGA Ad2.2_CGTACTAG Ad2.3_AGGCAGAA Ad2.4_TCCTGAGC Ad2.5_GGACTCCT Ad2.6_TAGGCATG Ad2.7_CTCTCTAC Ad2.8_CAGAGAGG Ad2.9_GCTACGCT Ad2.10_CGAGGCTG Ad2.11_AAGAGGCA Ad2.12_GTAGAGGA Ad2.13_GTCGTGAT
Ad1_noMX: Ad2.1_TAAGGCGA Ad2.2_CGTACTAG Ad2.3_AGGCAGAA Ad2.4_TCCTGAGC Ad2.5_GGACTCCT Ad2.6_TAGGCATG Ad2.7_CTCTCTAC Ad2.8_CAGAGAGG Ad2.9_GCTACGCT Ad2.10_CGAGGCTG Ad2.11_AAGAGGCA Ad2.12_GTAGAGGA Ad2.13_GTCGTGAT
Deep Sequencing technologies
 Deep Sequencing technologies Gabriela Salinas 30 October 2017 Transcriptome and Genome Analysis Laboratory http://www.uni-bc.gwdg.de/index.php?id=709 Microarray and Deep-Sequencing Core Facility University
Deep Sequencing technologies Gabriela Salinas 30 October 2017 Transcriptome and Genome Analysis Laboratory http://www.uni-bc.gwdg.de/index.php?id=709 Microarray and Deep-Sequencing Core Facility University
Charles Girardot, Furlong Lab. MACS, CisGenome, SISSRs and other peak calling algorithms: differences and practical use
 Charles Girardot, Furlong Lab MACS, CisGenome, SISSRs and other peak calling algorithms: differences and practical use ChIP-Seq signal properties Only 5 ends of ChIPed fragments are sequenced Shifted read
Charles Girardot, Furlong Lab MACS, CisGenome, SISSRs and other peak calling algorithms: differences and practical use ChIP-Seq signal properties Only 5 ends of ChIPed fragments are sequenced Shifted read
Supplementary Information for:
 Supplementary Information for: A streamlined and high-throughput targeting approach for human germline and cancer genomes using Oligonucleotide-Selective Sequencing Samuel Myllykangas 1, Jason D. Buenrostro
Supplementary Information for: A streamlined and high-throughput targeting approach for human germline and cancer genomes using Oligonucleotide-Selective Sequencing Samuel Myllykangas 1, Jason D. Buenrostro
Supplement to: The Genomic Sequence of the Chinese Hamster Ovary (CHO)-K1 cell line
 Supplement to: The Genomic Sequence of the Chinese Hamster Ovary (CHO)-K1 cell line Table of Contents SUPPLEMENTARY TEXT:... 2 FILTERING OF RAW READS PRIOR TO ASSEMBLY:... 2 COMPARATIVE ANALYSIS... 2 IMMUNOGENIC
Supplement to: The Genomic Sequence of the Chinese Hamster Ovary (CHO)-K1 cell line Table of Contents SUPPLEMENTARY TEXT:... 2 FILTERING OF RAW READS PRIOR TO ASSEMBLY:... 2 COMPARATIVE ANALYSIS... 2 IMMUNOGENIC
File S1. Program overview and features
 File S1 Program overview and features Query list filtering. Further filtering may be applied through user selected query lists (Figure. 2B, Table S3) that restrict the results and/or report specifically
File S1 Program overview and features Query list filtering. Further filtering may be applied through user selected query lists (Figure. 2B, Table S3) that restrict the results and/or report specifically
TECH NOTE Pushing the Limit: A Complete Solution for Generating Stranded RNA Seq Libraries from Picogram Inputs of Total Mammalian RNA
 TECH NOTE Pushing the Limit: A Complete Solution for Generating Stranded RNA Seq Libraries from Picogram Inputs of Total Mammalian RNA Stranded, Illumina ready library construction in
TECH NOTE Pushing the Limit: A Complete Solution for Generating Stranded RNA Seq Libraries from Picogram Inputs of Total Mammalian RNA Stranded, Illumina ready library construction in
Integrated NGS Sample Preparation Solutions for Limiting Amounts of RNA and DNA. March 2, Steven R. Kain, Ph.D. ABRF 2013
 Integrated NGS Sample Preparation Solutions for Limiting Amounts of RNA and DNA March 2, 2013 Steven R. Kain, Ph.D. ABRF 2013 NuGEN s Core Technologies Selective Sequence Priming Nucleic Acid Amplification
Integrated NGS Sample Preparation Solutions for Limiting Amounts of RNA and DNA March 2, 2013 Steven R. Kain, Ph.D. ABRF 2013 NuGEN s Core Technologies Selective Sequence Priming Nucleic Acid Amplification
Supplementary Information
 Supplementary Information Table of contents: Pages: Supplementary Methods 2-3 Supplementary Figure 1 4 Supplementary Figure 2 5 Supplementary Figure 3 6 Supplementary Figure 4 7 Supplementary Figure 5
Supplementary Information Table of contents: Pages: Supplementary Methods 2-3 Supplementary Figure 1 4 Supplementary Figure 2 5 Supplementary Figure 3 6 Supplementary Figure 4 7 Supplementary Figure 5
Discovering gene regulatory control using ChIP-chip and ChIP-seq. Part 1. An introduction to gene regulatory control, concepts and methodologies
 Discovering gene regulatory control using ChIP-chip and ChIP-seq Part 1 An introduction to gene regulatory control, concepts and methodologies Ian Simpson ian.simpson@.ed.ac.uk http://bit.ly/bio2links
Discovering gene regulatory control using ChIP-chip and ChIP-seq Part 1 An introduction to gene regulatory control, concepts and methodologies Ian Simpson ian.simpson@.ed.ac.uk http://bit.ly/bio2links
Epigenetics and DNase-Seq
 Epigenetics and DNase-Seq BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2018 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material, are licensed under CC BY-NC 4.0 by Anthony
Epigenetics and DNase-Seq BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2018 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material, are licensed under CC BY-NC 4.0 by Anthony
Lateral DNA Transfer MECHANISMS AND CONSEQUENCES Frederic Bushman
 Lateral DN ransfer MECHNISMS ND CONSEQUENCES y1 mrn Intron ranscription gag pol Integrated y1 DN Yeast cell Excised intron Reverse RN splicing transcription Integration C Old copy of y1 New copy lacking
Lateral DN ransfer MECHNISMS ND CONSEQUENCES y1 mrn Intron ranscription gag pol Integrated y1 DN Yeast cell Excised intron Reverse RN splicing transcription Integration C Old copy of y1 New copy lacking
of NOBOX. Supplemental Figure S1F shows the staining profile of LHX8 antibody (Abcam, ab41519), which is a
 SUPPLEMENTAL MATERIALS AND METHODS LHX8 Staining During the course of this work, it came to our attention that the NOBOX antibody used in this study had been discontinued. There are a number of other nuclear
SUPPLEMENTAL MATERIALS AND METHODS LHX8 Staining During the course of this work, it came to our attention that the NOBOX antibody used in this study had been discontinued. There are a number of other nuclear
Section C: The Control of Gene Expression
 Section C: The Control of Gene Expression 1. Each cell of a multicellular eukaryote expresses only a small fraction of its genes 2. The control of gene expression can occur at any step in the pathway from
Section C: The Control of Gene Expression 1. Each cell of a multicellular eukaryote expresses only a small fraction of its genes 2. The control of gene expression can occur at any step in the pathway from
Gene Expression Microarrays. For microarrays, purity of the RNA was further assessed by
 Supplemental Methods Gene Expression Microarrays. For microarrays, purity of the RNA was further assessed by an Agilent 2100 Bioanalyzer. 500 ng of RNA was reverse transcribed into crna and biotin-utp
Supplemental Methods Gene Expression Microarrays. For microarrays, purity of the RNA was further assessed by an Agilent 2100 Bioanalyzer. 500 ng of RNA was reverse transcribed into crna and biotin-utp
Parts of a standard FastQC report
 FastQC FastQC, written by Simon Andrews of Babraham Bioinformatics, is a very popular tool used to provide an overview of basic quality control metrics for raw next generation sequencing data. There are
FastQC FastQC, written by Simon Andrews of Babraham Bioinformatics, is a very popular tool used to provide an overview of basic quality control metrics for raw next generation sequencing data. There are
Many transcription factors! recognize DNA shape
 Many transcription factors! recognize DN shape Katie Pollard! Gladstone Institutes USF Division of Biostatistics, Institute for Human Genetics, and Institute for omputational Health Sciences ENODE Users
Many transcription factors! recognize DN shape Katie Pollard! Gladstone Institutes USF Division of Biostatistics, Institute for Human Genetics, and Institute for omputational Health Sciences ENODE Users
Data and Metadata Models Recommendations Version 1.2 Developed by the IHEC Metadata Standards Workgroup
 Data and Metadata Models Recommendations Version 1.2 Developed by the IHEC Metadata Standards Workgroup 1. Introduction The data produced by IHEC is illustrated in Figure 1. Figure 1. The space of epigenomic
Data and Metadata Models Recommendations Version 1.2 Developed by the IHEC Metadata Standards Workgroup 1. Introduction The data produced by IHEC is illustrated in Figure 1. Figure 1. The space of epigenomic
Microarrays: since we use probes we obviously must know the sequences we are looking at!
 These background are needed: 1. - Basic Molecular Biology & Genetics DNA replication Transcription Post-transcriptional RNA processing Translation Post-translational protein modification Gene expression
These background are needed: 1. - Basic Molecular Biology & Genetics DNA replication Transcription Post-transcriptional RNA processing Translation Post-translational protein modification Gene expression
Control of Eukaryotic Gene Expression (Learning Objectives)
 Control of Eukaryotic Gene Expression (Learning Objectives) 1. Compare and contrast chromatin and chromosome: composition, proteins involved and level of packing. Explain the structure and function of
Control of Eukaryotic Gene Expression (Learning Objectives) 1. Compare and contrast chromatin and chromosome: composition, proteins involved and level of packing. Explain the structure and function of
The ChIP-Seq project. Giovanna Ambrosini, Philipp Bucher. April 19, 2010 Lausanne. EPFL-SV Bucher Group
 The ChIP-Seq project Giovanna Ambrosini, Philipp Bucher EPFL-SV Bucher Group April 19, 2010 Lausanne Overview Focus on technical aspects Description of applications (C programs) Where to find binaries,
The ChIP-Seq project Giovanna Ambrosini, Philipp Bucher EPFL-SV Bucher Group April 19, 2010 Lausanne Overview Focus on technical aspects Description of applications (C programs) Where to find binaries,
Deep learning frameworks for regulatory genomics and epigenomics
 Deep learning frameworks for regulatory genomics and epigenomics Chuan Sheng Foo Avanti Shrikumar Nicholas Sinnott- Armstrong ANSHUL KUNDAJE Genetics, Computer science Stanford University Johnny Israeli
Deep learning frameworks for regulatory genomics and epigenomics Chuan Sheng Foo Avanti Shrikumar Nicholas Sinnott- Armstrong ANSHUL KUNDAJE Genetics, Computer science Stanford University Johnny Israeli
CS273B: Deep learning for Genomics and Biomedicine
 CS273B: Deep learning for Genomics and Biomedicine Lecture 2: Convolutional neural networks and applications to functional genomics 09/28/2016 Anshul Kundaje, James Zou, Serafim Batzoglou Outline Anatomy
CS273B: Deep learning for Genomics and Biomedicine Lecture 2: Convolutional neural networks and applications to functional genomics 09/28/2016 Anshul Kundaje, James Zou, Serafim Batzoglou Outline Anatomy
Recombinant DNA Technology
 Recombinant DNA Technology Common General Cloning Strategy Target DNA from donor organism extracted, cut with restriction endonuclease and ligated into a cloning vector cut with compatible restriction
Recombinant DNA Technology Common General Cloning Strategy Target DNA from donor organism extracted, cut with restriction endonuclease and ligated into a cloning vector cut with compatible restriction
Galaxy Platform For NGS Data Analyses
 Galaxy Platform For NGS Data Analyses Weihong Yan wyan@chem.ucla.edu Collaboratory Web Site http://qcb.ucla.edu/collaboratory http://collaboratory.lifesci.ucla.edu Workshop Outline ü Day 1 UCLA galaxy
Galaxy Platform For NGS Data Analyses Weihong Yan wyan@chem.ucla.edu Collaboratory Web Site http://qcb.ucla.edu/collaboratory http://collaboratory.lifesci.ucla.edu Workshop Outline ü Day 1 UCLA galaxy
Chapter 17 Lecture. Concepts of Genetics. Tenth Edition. Regulation of Gene Expression in Eukaryotes
 Chapter 17 Lecture Concepts of Genetics Tenth Edition Regulation of Gene Expression in Eukaryotes Chapter Contents 17.1 Eukaryotic Gene Regulation Can Occur at Any of the Steps Leading from DNA to Protein
Chapter 17 Lecture Concepts of Genetics Tenth Edition Regulation of Gene Expression in Eukaryotes Chapter Contents 17.1 Eukaryotic Gene Regulation Can Occur at Any of the Steps Leading from DNA to Protein
Measuring gene expression
 Measuring gene expression Grundlagen der Bioinformatik SS2018 https://www.youtube.com/watch?v=v8gh404a3gg Agenda Organization Gene expression Background Technologies FISH Nanostring Microarrays RNA-seq
Measuring gene expression Grundlagen der Bioinformatik SS2018 https://www.youtube.com/watch?v=v8gh404a3gg Agenda Organization Gene expression Background Technologies FISH Nanostring Microarrays RNA-seq
Selected Techniques Part I
 1 Selected Techniques Part I Gel Electrophoresis Can be both qualitative and quantitative Qualitative About what size is the fragment? How many fragments are present? Is there in insert or not? Quantitative
1 Selected Techniques Part I Gel Electrophoresis Can be both qualitative and quantitative Qualitative About what size is the fragment? How many fragments are present? Is there in insert or not? Quantitative
Poly A + RNA polya 3' XXXXX. SMART CDS primer First-strand synthesis & tailing by SMARTScribe RT. SMARTer II A Oligonucleotide. Single step 5' XXXXX
 5' 5' XXXXX SMARTer II A Oligonucleotide XXXXX 5' 5' XXXXX Poly A + RNA polya 3' SMART CDS primer First-strand synthesis & tailing y SMARTScrie RT polya Template switching and extension y SMARTScrie RT
5' 5' XXXXX SMARTer II A Oligonucleotide XXXXX 5' 5' XXXXX Poly A + RNA polya 3' SMART CDS primer First-strand synthesis & tailing y SMARTScrie RT polya Template switching and extension y SMARTScrie RT
Gene expression analysis. Biosciences 741: Genomics Fall, 2013 Week 5. Gene expression analysis
 Gene expression analysis Biosciences 741: Genomics Fall, 2013 Week 5 Gene expression analysis From EST clusters to spotted cdna microarrays Long vs. short oligonucleotide microarrays vs. RT-PCR Methods
Gene expression analysis Biosciences 741: Genomics Fall, 2013 Week 5 Gene expression analysis From EST clusters to spotted cdna microarrays Long vs. short oligonucleotide microarrays vs. RT-PCR Methods
SO YOU WANT TO DO A: RNA-SEQ EXPERIMENT MATT SETTLES, PHD UNIVERSITY OF CALIFORNIA, DAVIS
 SO YOU WANT TO DO A: RNA-SEQ EXPERIMENT MATT SETTLES, PHD UNIVERSITY OF CALIFORNIA, DAVIS SETTLES@UCDAVIS.EDU Bioinformatics Core Genome Center UC Davis BIOINFORMATICS.UCDAVIS.EDU DISCLAIMER This talk/workshop
SO YOU WANT TO DO A: RNA-SEQ EXPERIMENT MATT SETTLES, PHD UNIVERSITY OF CALIFORNIA, DAVIS SETTLES@UCDAVIS.EDU Bioinformatics Core Genome Center UC Davis BIOINFORMATICS.UCDAVIS.EDU DISCLAIMER This talk/workshop
Mapping long-range promoter contacts in human cells with high-resolution capture Hi-C
 CORRECTION NOTICE Nat. Genet. 47, 598 606 (2015) Mapping long-range promoter contacts in human cells with high-resolution capture Hi-C Borbala Mifsud, Filipe Tavares-Cadete, Alice N Young, Robert Sugar,
CORRECTION NOTICE Nat. Genet. 47, 598 606 (2015) Mapping long-range promoter contacts in human cells with high-resolution capture Hi-C Borbala Mifsud, Filipe Tavares-Cadete, Alice N Young, Robert Sugar,
Discovering gene regulatory control using ChIP-chip and ChIP-seq. An introduction to gene regulatory control, concepts and methodologies
 Discovering gene regulatory control using ChIP-chip and ChIP-seq An introduction to gene regulatory control, concepts and methodologies Ian Simpson ian.simpson@.ed.ac.uk bit.ly/bio2_2012 The Central Dogma
Discovering gene regulatory control using ChIP-chip and ChIP-seq An introduction to gene regulatory control, concepts and methodologies Ian Simpson ian.simpson@.ed.ac.uk bit.ly/bio2_2012 The Central Dogma
pej605 pej414 containing 81 bp downstream and 579 bp This study
 SUPPLEMENTARY DATA Table S. Details of plasmids used in this study. Plasmid Description Reference or source pfm8 Protein expression vector based on pet5b containing (0) His-tagged lexa. pcr 4-TOPO Cloning
SUPPLEMENTARY DATA Table S. Details of plasmids used in this study. Plasmid Description Reference or source pfm8 Protein expression vector based on pet5b containing (0) His-tagged lexa. pcr 4-TOPO Cloning
(A) Extrachromosomal DNA (B) RNA found in bacterial cells (C) Is part of the bacterial chromosome (D) Is part of the eukaryote chromosome
 Microbiology - Problem Drill 07: Microbial Genetics and Biotechnology No. 1 of 10 1. A plasmid is? (A) Extrachromosomal DNA (B) RNA found in bacterial cells (C) Is part of the bacterial chromosome (D)
Microbiology - Problem Drill 07: Microbial Genetics and Biotechnology No. 1 of 10 1. A plasmid is? (A) Extrachromosomal DNA (B) RNA found in bacterial cells (C) Is part of the bacterial chromosome (D)
Caroline Townsend December 2012 Biochem 218 A critical review of ChIP-seq enrichment analysis tools
 Caroline Townsend December 2012 Biochem 218 A critical review of ChIP-seq enrichment analysis tools Introduction Transcriptional regulation, chromatin states, and genome stability pathways are largely
Caroline Townsend December 2012 Biochem 218 A critical review of ChIP-seq enrichment analysis tools Introduction Transcriptional regulation, chromatin states, and genome stability pathways are largely
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Processing of mutations and generation of simulated controls. On the left, a diagram illustrates the manner in which covariate-matched simulated mutations were obtained, filtered
Supplementary Figure 1 Processing of mutations and generation of simulated controls. On the left, a diagram illustrates the manner in which covariate-matched simulated mutations were obtained, filtered
Whole Transcriptome Analysis of Illumina RNA- Seq Data. Ryan Peters Field Application Specialist
 Whole Transcriptome Analysis of Illumina RNA- Seq Data Ryan Peters Field Application Specialist Partek GS in your NGS Pipeline Your Start-to-Finish Solution for Analysis of Next Generation Sequencing Data
Whole Transcriptome Analysis of Illumina RNA- Seq Data Ryan Peters Field Application Specialist Partek GS in your NGS Pipeline Your Start-to-Finish Solution for Analysis of Next Generation Sequencing Data
Characterizing DNA binding sites high throughput approaches Biol4230 Tues, April 24, 2018 Bill Pearson Pinn 6-057
 Characterizing DNA binding sites high throughput approaches Biol4230 Tues, April 24, 2018 Bill Pearson wrp@virginia.edu 4-2818 Pinn 6-057 Reviewing sites: affinity and specificity representation binding
Characterizing DNA binding sites high throughput approaches Biol4230 Tues, April 24, 2018 Bill Pearson wrp@virginia.edu 4-2818 Pinn 6-057 Reviewing sites: affinity and specificity representation binding
Nature Structural and Molecular Biology: doi: /nsmb Supplementary Figure 1. Validation of CDK9-inhibitor treatment.
 Supplementary Figure 1 Validation of CDK9-inhibitor treatment. (a) Schematic of GAPDH with the middle of the amplicons indicated in base pairs. The transcription start site (TSS) and the terminal polyadenylation
Supplementary Figure 1 Validation of CDK9-inhibitor treatment. (a) Schematic of GAPDH with the middle of the amplicons indicated in base pairs. The transcription start site (TSS) and the terminal polyadenylation
Genome 373: High- Throughput DNA Sequencing. Doug Fowler
 Genome 373: High- Throughput DNA Sequencing Doug Fowler Tasks give ML unity We learned about three tasks that are commonly encountered in ML Models/Algorithms Give ML Diversity Classification Regression
Genome 373: High- Throughput DNA Sequencing Doug Fowler Tasks give ML unity We learned about three tasks that are commonly encountered in ML Models/Algorithms Give ML Diversity Classification Regression
Nature Methods: doi: /nmeth Supplementary Figure 1. Construction of a sensitive TetR mediated auxotrophic off-switch.
 Supplementary Figure 1 Construction of a sensitive TetR mediated auxotrophic off-switch. A Production of the Tet repressor in yeast when conjugated to either the LexA4 or LexA8 promoter DNA binding sequences.
Supplementary Figure 1 Construction of a sensitive TetR mediated auxotrophic off-switch. A Production of the Tet repressor in yeast when conjugated to either the LexA4 or LexA8 promoter DNA binding sequences.
Analysis of Microarray Data
 Analysis of Microarray Data Lecture 3: Visualization and Functional Analysis George Bell, Ph.D. Senior Bioinformatics Scientist Bioinformatics and Research Computing Whitehead Institute Outline Review
Analysis of Microarray Data Lecture 3: Visualization and Functional Analysis George Bell, Ph.D. Senior Bioinformatics Scientist Bioinformatics and Research Computing Whitehead Institute Outline Review
Technical Review. Real time PCR
 Technical Review Real time PCR Normal PCR: Analyze with agarose gel Normal PCR vs Real time PCR Real-time PCR, also known as quantitative PCR (qpcr) or kinetic PCR Key feature: Used to amplify and simultaneously
Technical Review Real time PCR Normal PCR: Analyze with agarose gel Normal PCR vs Real time PCR Real-time PCR, also known as quantitative PCR (qpcr) or kinetic PCR Key feature: Used to amplify and simultaneously
Recitation CHAPTER 9 DNA Technologies
 Recitation CHAPTER 9 DNA Technologies DNA Cloning: General Scheme A cloning vector and eukaryotic chromosomes are separately cleaved with the same restriction endonuclease. (A single chromosome is shown
Recitation CHAPTER 9 DNA Technologies DNA Cloning: General Scheme A cloning vector and eukaryotic chromosomes are separately cleaved with the same restriction endonuclease. (A single chromosome is shown
Supplemental Information. Autoregulatory Feedback Controls. Sequential Action of cis-regulatory Modules. at the brinker Locus
 Developmental Cell, Volume 26 Supplemental Information Autoregulatory Feedback Controls Sequential Action of cis-regulatory Modules at the brinker Locus Leslie Dunipace, Abbie Saunders, Hilary L. Ashe,
Developmental Cell, Volume 26 Supplemental Information Autoregulatory Feedback Controls Sequential Action of cis-regulatory Modules at the brinker Locus Leslie Dunipace, Abbie Saunders, Hilary L. Ashe,
Simultaneous multiplexed amplicon sequencing and transcriptome profiling in single cells
 SUPPLEMENTARY Brief Communication INFORMATION https://doi.org/10.1038/s41592-018-0259-9 In the format provided by the authors and unedited. Simultaneous multiplexed amplicon sequencing and transcriptome
SUPPLEMENTARY Brief Communication INFORMATION https://doi.org/10.1038/s41592-018-0259-9 In the format provided by the authors and unedited. Simultaneous multiplexed amplicon sequencing and transcriptome
Genes found in the genome include protein-coding genes and non-coding RNA genes. Which nucleotide is not normally found in non-coding RNA genes?
 Midterm Q Genes found in the genome include protein-coding genes and non-coding RNA genes Which nucleotide is not normally found in non-coding RNA genes? G T 3 A 4 C 5 U 00% Midterm Q Which of the following
Midterm Q Genes found in the genome include protein-coding genes and non-coding RNA genes Which nucleotide is not normally found in non-coding RNA genes? G T 3 A 4 C 5 U 00% Midterm Q Which of the following
nature methods A paired-end sequencing strategy to map the complex landscape of transcription initiation
 nature methods A paired-end sequencing strategy to map the complex landscape of transcription initiation Ting Ni, David L Corcoran, Elizabeth A Rach, Shen Song, Eric P Spana, Yuan Gao, Uwe Ohler & Jun
nature methods A paired-end sequencing strategy to map the complex landscape of transcription initiation Ting Ni, David L Corcoran, Elizabeth A Rach, Shen Song, Eric P Spana, Yuan Gao, Uwe Ohler & Jun
Introduction to genome biology
 Introduction to genome biology Lisa Stubbs We ve found most genes; but what about the rest of the genome? Genome size* 12 Mb 95 Mb 170 Mb 1500 Mb 2700 Mb 3200 Mb #coding genes ~7000 ~20000 ~14000 ~26000
Introduction to genome biology Lisa Stubbs We ve found most genes; but what about the rest of the genome? Genome size* 12 Mb 95 Mb 170 Mb 1500 Mb 2700 Mb 3200 Mb #coding genes ~7000 ~20000 ~14000 ~26000
Figure S1. nuclear extracts. HeLa cell nuclear extract. Input IgG IP:ORC2 ORC2 ORC2. MCM4 origin. ORC2 occupancy
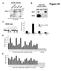 A nuclear extracts B HeLa cell nuclear extract Figure S1 ORC2 (in kda) 21 132 7 ORC2 Input IgG IP:ORC2 32 ORC C D PRKDC ORC2 occupancy Directed against ORC2 C-terminus (sc-272) MCM origin 2 2 1-1 -1kb
A nuclear extracts B HeLa cell nuclear extract Figure S1 ORC2 (in kda) 21 132 7 ORC2 Input IgG IP:ORC2 32 ORC C D PRKDC ORC2 occupancy Directed against ORC2 C-terminus (sc-272) MCM origin 2 2 1-1 -1kb
Identification of individual motifs on the genome scale. Some slides are from Mayukh Bhaowal
 Identification of individual motifs on the genome scale Some slides are from Mayukh Bhaowal Two papers Nature 423, 241-254 (15 May 2003) Sequencing and comparison of yeast species to identify genes and
Identification of individual motifs on the genome scale Some slides are from Mayukh Bhaowal Two papers Nature 423, 241-254 (15 May 2003) Sequencing and comparison of yeast species to identify genes and
MODULE TSS1: TRANSCRIPTION START SITES INTRODUCTION (BASIC)
 MODULE TSS1: TRANSCRIPTION START SITES INTRODUCTION (BASIC) Lesson Plan: Title JAMIE SIDERS, MEG LAAKSO & WILSON LEUNG Identifying transcription start sites for Peaked promoters using chromatin landscape,
MODULE TSS1: TRANSCRIPTION START SITES INTRODUCTION (BASIC) Lesson Plan: Title JAMIE SIDERS, MEG LAAKSO & WILSON LEUNG Identifying transcription start sites for Peaked promoters using chromatin landscape,
Unit IX Problem 3 Genetics: Basic Concepts in Molecular Biology
 Unit IX Problem 3 Genetics: Basic Concepts in Molecular Biology - The central dogma (principle) of molecular biology: Information from DNA are transcribed to mrna which will be further translated to synthesize
Unit IX Problem 3 Genetics: Basic Concepts in Molecular Biology - The central dogma (principle) of molecular biology: Information from DNA are transcribed to mrna which will be further translated to synthesize
C24. ros1-1. ros1-1 rdm18-1. ros1-1 rdm18-2. ros1-1 nrpe1
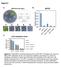 Figure S1 Methylation level 1 0.8 0.6 0.4 0.2 0 p35s methylation levels 24 ros1-1 ros1-1 rdm18-1 ros1-1 rdm18-2 ros1-1 nrpe1 mg mhg mhh Figure S1. RM18/PKL promotes silencing at the p35s-npt II transgene
Figure S1 Methylation level 1 0.8 0.6 0.4 0.2 0 p35s methylation levels 24 ros1-1 ros1-1 rdm18-1 ros1-1 rdm18-2 ros1-1 nrpe1 mg mhg mhh Figure S1. RM18/PKL promotes silencing at the p35s-npt II transgene
Exercises: Analysing ChIP-Seq data
 Exercises: Analysing ChIP-Seq data Version 2018-03-2 Exercises: Analysing ChIP-Seq data 2 Licence This manual is 2018, Simon Andrews. This manual is distributed under the creative commons Attribution-Non-Commercial-Share
Exercises: Analysing ChIP-Seq data Version 2018-03-2 Exercises: Analysing ChIP-Seq data 2 Licence This manual is 2018, Simon Andrews. This manual is distributed under the creative commons Attribution-Non-Commercial-Share
Nature Genetics: doi: /ng Supplementary Figure 1. H3K27ac HiChIP enriches enhancer promoter-associated chromatin contacts.
 Supplementary Figure 1 H3K27ac HiChIP enriches enhancer promoter-associated chromatin contacts. (a) Schematic of chromatin contacts captured in H3K27ac HiChIP. (b) Loop call overlap for cohesin HiChIP
Supplementary Figure 1 H3K27ac HiChIP enriches enhancer promoter-associated chromatin contacts. (a) Schematic of chromatin contacts captured in H3K27ac HiChIP. (b) Loop call overlap for cohesin HiChIP
Supplementary Figures
 Supplementary Figures A B Supplementary Figure 1. Examples of discrepancies in predicted and validated breakpoint coordinates. A) Most frequently, predicted breakpoints were shifted relative to those derived
Supplementary Figures A B Supplementary Figure 1. Examples of discrepancies in predicted and validated breakpoint coordinates. A) Most frequently, predicted breakpoints were shifted relative to those derived
Genome 541 Gene regulation and epigenomics Lecture 3 Integrative analysis of genomics assays
 Genome 541 Gene regulation and epigenomics Lecture 3 Integrative analysis of genomics assays Please consider both the forward and reverse strands (i.e. reverse compliment sequence). You do not need to
Genome 541 Gene regulation and epigenomics Lecture 3 Integrative analysis of genomics assays Please consider both the forward and reverse strands (i.e. reverse compliment sequence). You do not need to
Introduction to Next Generation Sequencing
 The Sequencing Revolution Introduction to Next Generation Sequencing Dena Leshkowitz,WIS 1 st BIOmics Workshop High throughput Short Read Sequencing Technologies Highly parallel reactions (millions to
The Sequencing Revolution Introduction to Next Generation Sequencing Dena Leshkowitz,WIS 1 st BIOmics Workshop High throughput Short Read Sequencing Technologies Highly parallel reactions (millions to
Experimental Design. Dr. Matthew L. Settles. Genome Center University of California, Davis
 Experimental Design Dr. Matthew L. Settles Genome Center University of California, Davis settles@ucdavis.edu What is Differential Expression Differential expression analysis means taking normalized sequencing
Experimental Design Dr. Matthew L. Settles Genome Center University of California, Davis settles@ucdavis.edu What is Differential Expression Differential expression analysis means taking normalized sequencing
Flow through. lgg. 7mIPA1-GFP. ChIP-SA rep 1. Flow through. lgg. 7mIPA1-GFP. ChIP-SA rep 2. Flow through. Input. ChIP-YP rep 1.
 Supplemental Data. Lu et al. Plant Cell. (213). 1.115/tpc.113.113639 Number of reads A Kd 8 lgg Flow through agfp lgg kb 2. 7mIPA1-GFP 1. B 6 Kd 8 6 Kd 8 agfp agfp lgg agfp lgg Flow through Flow through
Supplemental Data. Lu et al. Plant Cell. (213). 1.115/tpc.113.113639 Number of reads A Kd 8 lgg Flow through agfp lgg kb 2. 7mIPA1-GFP 1. B 6 Kd 8 6 Kd 8 agfp agfp lgg agfp lgg Flow through Flow through
Non-coding Function & Variation, MPRAs II. Mike White Bio /5/18
 Non-coding Function & Variation, MPRAs II Mike White Bio 5488 3/5/18 MPRA Review Problem 1: Where does your CRE DNA come from? DNA synthesis Genomic fragments Targeted regulome capture Problem 2: How do
Non-coding Function & Variation, MPRAs II Mike White Bio 5488 3/5/18 MPRA Review Problem 1: Where does your CRE DNA come from? DNA synthesis Genomic fragments Targeted regulome capture Problem 2: How do
Transcriptomics analysis with RNA seq: an overview Frederik Coppens
 Transcriptomics analysis with RNA seq: an overview Frederik Coppens Platforms Applications Analysis Quantification RNA content Platforms Platforms Short (few hundred bases) Long reads (multiple kilobases)
Transcriptomics analysis with RNA seq: an overview Frederik Coppens Platforms Applications Analysis Quantification RNA content Platforms Platforms Short (few hundred bases) Long reads (multiple kilobases)
Lecture Summary: Regulation of transcription. General mechanisms-what are the major regulatory points?
 BCH 401G Lecture 37 Andres Lecture Summary: Regulation of transcription. General mechanisms-what are the major regulatory points? RNA processing: Capping, polyadenylation, splicing. Why process mammalian
BCH 401G Lecture 37 Andres Lecture Summary: Regulation of transcription. General mechanisms-what are the major regulatory points? RNA processing: Capping, polyadenylation, splicing. Why process mammalian
High-quality stranded RNA-seq libraries from single cells using the SMART-Seq Stranded Kit Product highlights:
 TECH NOTE High-quality stranded RNA-seq libraries from single cells using the SMART-Seq Stranded Kit Product highlights: Simple workflow starts directly from 1 1,000 cells or 10 pg 10 ng total RNA to generate
TECH NOTE High-quality stranded RNA-seq libraries from single cells using the SMART-Seq Stranded Kit Product highlights: Simple workflow starts directly from 1 1,000 cells or 10 pg 10 ng total RNA to generate
NAME TA SEC Problem Set 3 FRIDAY March 5, Problem sets will NOT be accepted late.
 MIT Department of Biology 7.013: Introductory Biology - Spring 2004 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. laudette ardel NME T SE 7.013 Problem Set 3 FRIDY March 5, 2004 Problem
MIT Department of Biology 7.013: Introductory Biology - Spring 2004 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. laudette ardel NME T SE 7.013 Problem Set 3 FRIDY March 5, 2004 Problem
Polymerase Chain Reaction
 Polymerase Chain Reaction Problem Suppose you have a patient with an infection or a heritable disease. You want to know which infection or disease it is and.. you want to know it fast and... from as little
Polymerase Chain Reaction Problem Suppose you have a patient with an infection or a heritable disease. You want to know which infection or disease it is and.. you want to know it fast and... from as little
Green Center Computational Core ChIP- Seq Pipeline, Just a Click Away
 Green Center Computational Core ChIP- Seq Pipeline, Just a Click Away Venkat Malladi Computational Biologist Computational Core Cecil H. and Ida Green Center for Reproductive Biology Science Introduc
Green Center Computational Core ChIP- Seq Pipeline, Just a Click Away Venkat Malladi Computational Biologist Computational Core Cecil H. and Ida Green Center for Reproductive Biology Science Introduc
Variant calling workflow for the Oncomine Comprehensive Assay using Ion Reporter Software v4.4
 WHITE PAPER Oncomine Comprehensive Assay Variant calling workflow for the Oncomine Comprehensive Assay using Ion Reporter Software v4.4 Contents Scope and purpose of document...2 Content...2 How Torrent
WHITE PAPER Oncomine Comprehensive Assay Variant calling workflow for the Oncomine Comprehensive Assay using Ion Reporter Software v4.4 Contents Scope and purpose of document...2 Content...2 How Torrent
