Supplementary Figure 1 Abortive ligation (adenylation) of the gapped 5'-dRP-containing BER
|
|
|
- Darcy Joseph
- 5 years ago
- Views:
Transcription
1 Supplementary Figure 1 Abortive ligation (adenylation) of the gapped 5'-dRP-containing BER intermediate by human BER ligases. The ligation efficiency of DNA ligase I (a) and XRCC1/DNA ligase III complex (b). The migration positions of the DNA substrate (17 drp, Substrate 1 in Supplementary Table 1), ligation product and the adenylated DNA intermediate (17 drp +AMP) are indicated.
2 Supplementary Figure 2 The assessment of drp lyase activity of wild type and pol β mutants. (a) Uncropped image for the adenylated sugar phosphate removal by pol β from the gapped and nicked 5'-dRP and 5'-AMP-dRP-containing substrates. The final cropped image is presented in
3 Fig. 1. (b,c) The drp lyase assay for the adenylated sugar phosphate removal by the 3KΔ mutant of pol β from the gapped (b) and nicked (c) 5'-AMP-dRP-containing substrates. In both panels b and c, lane 1 is a minus enzyme control, and lane 2 is a reference reaction containing wild type pol β. The positions of the gapped (17 drp +AMP, Substrate 2 in Supplementary Table 1) and nicked (18 drp +AMP, Substrate 4 in Supplementary Table 1) DNA substrates and the pol β lyase products (17-mer in panel b, 18-mer in panel c) are indicated. The 3KΔ mutant was devoid of activity (lanes 3-6). (d,e) The drp lyase assay for the adenylated sugar phosphate removal by the K68A mutant of pol β from the gapped (d) and nicked (e) 5'-AMP-dRP-containing substrates. In both panels d and e, lane 1 is a minus enzyme control and lane 2 is a reference reaction containing wild type pol β. The positions of the gapped (17 drp +AMP, Substrate 2 in Supplementary Table 1) and nicked (18 drp +AMP, Substrate 4 in Supplementary Table 1) DNA substrates and the pol β lyase products (17-mer in panel d, 18-mer in panel e) after 5'-AMP-dRP removal by wild type pol β (lane 2), and K68A mutant (lanes 6-9) are indicated.
4 Supplementary Figure 3 The actions of APTX and pol β on different DNA substrates. The drp lyase and DNA deadenylation assays for the processing of the gapped and nicked 5'-AMP-dRP (a), the nicked 5'-AMP (b), and the gapped and nicked 5'-AMP-THF (c) containing DNA substrates. (a) Lane 1 and lane 4 are minus enzyme controls; lane 2 and lane 5 are the products of APTX 5'- AMP removal; lane 3 and lane 6 are the products of pol β lyase 5'-AMP-dRP removal. The positions of the gapped (17 drp +AMP, Substrate 2 in Supplementary Table 1) and nicked (18 drp +AMP, Substrate 4 in Supplementary Table 1) DNA substrates and the pol β lyase (17-mer or 18-mer) and APTX removal (17 drp or 18 drp ) products are indicated. (b) Lane 7 is a minus enzyme control, and lane 8 is the product of APTX 5'-AMP removal. The positions of the nicked DNA substrate (18+AMP, Substrate 5 in Supplementary Table 1), and APTX removal product (18-mer) are indicated. Pol β showed no activity with this substrate (lane 9). (c) Lane 10 and lane 13 are minus enzyme controls, and lane 11 and lane 14 are the products of APTX 5'-AMP removal. The positions for the gapped (17 THF +AMP, Substrate 6 in Supplementary Table 1) and nicked (18 THF +AMP, Substrate 7 in Supplementary Table 1) DNA substrates and APTX products (17 THF or 18 THF ) are indicated. Pol β showed no activity with these substrates (lanes 12 and 15).
5 Supplementary Figure 4 Transactions of pol β, APTX and FEN1 at the BER intermediates. (a,b,c) FEN1 excision assay for the nicked 5'-AMP-dRP (a), gapped 5'-AMP-dRP (b), the nicked 5'-AMP (c) containing DNA substrates (Supplementary Table 1). Lanes 1, 9 and 17 are minus enzyme controls, lanes 2 and 10 are reference reactions containing pol β. Lanes 3-8 in panel a, lanes in panel b, and lanes in panel c are FEN1 excision cleavage products. The positions of the DNA substrate for 18 drp +AMP (Substrate 4 in panel a), 17 drp +AMP (Substrate 2 in panel b), and 18+AMP (Substrate 5 in panel c) are indicated. The positions of the products for pol β lyase (18-mer in panel a, 17-mer in panel b), and FEN1 excision cleavage (17-, 16- or 15-mer in panel a; 16- or 15-mer in panel b; 17- or 16-mer in panel c) are indicated. Pol β showed no activity with Substrate 5 (lane 18). (d) The catalytic rates of pol β, FEN1, and APTX in the processing of 5'-AMP-dRP substrate. The rates of pol β ( ) FEN1 (u ), and APTX (n ) are as k obs of 0.05 s -1, 0.03 s -1, and 0.2 s -1, respectively. (e) A model proposing the fate of the adenylated BER
6 intermediate as mediated by pol β, FEN1, and APTX activities. The normal pol β lyase reaction illustrated at the top is biphasic and fast (>800 s -1 and 2 s -1 ).
7 Supplementary Figure 5 The fate of 5'-adenylated BER intermediate in the case of APTX and/or FEN1 deficiency. (a) The drp lyase, DNA adenylation and FEN1 excision assays with purified enzymes pol β, FEN1, and APTX. Lane 1 is a minus enzyme control, lane 2, 3, and 4 are reference reactions containing FEN1, pol β, and APTX, respectively. The migration position of the DNA substrate (17 drp +AMP, Substrate 2 in Supplementary Table 1) is indicated. (b-g) The drp lyase, DNA adenylation and FEN1 excision assays in the cell extracts prepared from the yeast strains (Supplementary Table 3). DNA substrate was not pretreated with UDG. The reaction products observed in the cell extracts from the wild-type (b, lanes 5-7), hnt3δ (c, lanes 8-10), rad27δ (d, lanes 11-13), hnt3δrad27δ (e, lanes 14-16), rad27δ::polb (f, lanes 17-19), hnt3δrad27δ::polb (g, lanes 20-22) yeast strains are indicated. In panels a-g, the positions of the products for FEN1 excision cleavage (16- or 15-mer), pol β lyase (17-mer), and APTX removal of AMP (17 drp ) are indicated. Three lanes in each set (in panels b-g) correspond to the time points 5, 15, or 30 min.
8 Supplementary Table 1 DNA substrates used in this study. FAM indicates the presence of a fluorescence tag. The abbreviations drp and THF represent a 5'-deoxyribose phosphate (5'-dRP) and synthetic abasic site analog (tetrahydrofuran), respectively. The presentations for DNA substrates 1-4 are designations after UDG treatment of the 36-mer double-stranded DNA substrates.
9 Supplementary Table 2 Data collection and refinement statistics.
10 Supplementary Table 3 Saccharomyces cerevisiae strains used in this study.
11 Supplementary Table 4 The sequences of oligonucleotides and primers used in this study. a F denotes THF, U represents a uracil base, and p stands for a phosphate group. The sequences of DNA oligonucleotides are shown in 5' to 3' orientation. b The downstream oligonucleotides include the 3'-FAM label. The downstream, upstream, and template oligonucleotides were used to construct the DNA substrates represented in Supplementary Table 1. For example, D Ugap (17-mer plus a uracil base) was used to construct either the gapped 5'-dRP-containing substrate (Substrate 1, Supplementary Table 1) or the gapped 5'-AMP-dRP-containing substrate (Substrate 2, Supplementary Table 1) after its annealing with the template b in the presence of the upstream b oligonucleotide. c Forward and reverse primers were used to construct the yeast strains used in this study. d Template, primer, and downstream oligonucleotides were used to construct the DNA substrate for crystallization of pol β-dna complex.
AP endonuclease 1 prevents trinucleotide repeat expansion via a novel mechanism during base excision repair
 Nucleic Acids Research Advance Access published May 18, 2015 Nucleic Acids Research, 2015 1 doi: 10.1093/nar/gkv530 AP endonuclease 1 prevents trinucleotide repeat expansion via a novel mechanism during
Nucleic Acids Research Advance Access published May 18, 2015 Nucleic Acids Research, 2015 1 doi: 10.1093/nar/gkv530 AP endonuclease 1 prevents trinucleotide repeat expansion via a novel mechanism during
CHAPTER 5. Reduced DNA gap repair in aging rat neuronal extracts and its restoration in vitro by DNA Polymerase p
 CHAPTER 5 Reduced DNA gap repair in aging rat neuronal extracts and its restoration in vitro by DNA Polymerase p Reduced DNA gap repair in aging rat neuronal extracts and its restoration in vitro by DNA
CHAPTER 5 Reduced DNA gap repair in aging rat neuronal extracts and its restoration in vitro by DNA Polymerase p Reduced DNA gap repair in aging rat neuronal extracts and its restoration in vitro by DNA
Roles of DNA Base Excision Repair in Maintaining the Integrity of DNA Methylation
 Florida International University FIU Digital Commons FIU Electronic Theses and Dissertations University Graduate School 11-15-2013 Roles of DNA Base Excision Repair in Maintaining the Integrity of DNA
Florida International University FIU Digital Commons FIU Electronic Theses and Dissertations University Graduate School 11-15-2013 Roles of DNA Base Excision Repair in Maintaining the Integrity of DNA
Supplementary Information for. Coordination of multiple enzyme activities by a single PCNA in archaeal Okazaki fragment.
 Supplementary Information for Coordination of multiple enzyme activities by a single PCNA in archaeal Okazaki fragment maturation Thomas R. Beattie and Stephen D. Bell Sir William Dunn School of Pathology,
Supplementary Information for Coordination of multiple enzyme activities by a single PCNA in archaeal Okazaki fragment maturation Thomas R. Beattie and Stephen D. Bell Sir William Dunn School of Pathology,
Please write your name or student ID number on every page.
 MCB 110 First Exam A TOTAL OF SIX PAGES NAME: Student ID Number: Question Maximum Points Your Points I 30 II 30 III 30 IV 38 V 22 150 Please write your name or student ID number on every page. This exam
MCB 110 First Exam A TOTAL OF SIX PAGES NAME: Student ID Number: Question Maximum Points Your Points I 30 II 30 III 30 IV 38 V 22 150 Please write your name or student ID number on every page. This exam
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11988 Supplementary Figure 1. Digestion of model DNA substrates. a, Linearized plasmid DNA (pik31- PstI, lanes 1 and 2), supercoiled plasmid (pik31, lanes 3 and 4), singly nicked plasmid
doi:10.1038/nature11988 Supplementary Figure 1. Digestion of model DNA substrates. a, Linearized plasmid DNA (pik31- PstI, lanes 1 and 2), supercoiled plasmid (pik31, lanes 3 and 4), singly nicked plasmid
4. Analysing genes II Isolate mutants*
 .. 4. Analysing s II Isolate mutants* Using the mutant to isolate the classify mutants by complementation analysis wild type study phenotype of mutants mutant 1 - use mutant to isolate sequence put individual
.. 4. Analysing s II Isolate mutants* Using the mutant to isolate the classify mutants by complementation analysis wild type study phenotype of mutants mutant 1 - use mutant to isolate sequence put individual
MCB 110 Spring 2017 Exam 1 SIX PAGES
 MCB 110 Spring 2017 Exam 1 SIX PAGES NAME: SID Number: Question Maximum Points Your Points I 28 II 32 III 32 IV 30 V 28 150 PLEASE WRITE your NAME or SID number on each page. This exam must be written
MCB 110 Spring 2017 Exam 1 SIX PAGES NAME: SID Number: Question Maximum Points Your Points I 28 II 32 III 32 IV 30 V 28 150 PLEASE WRITE your NAME or SID number on each page. This exam must be written
Reviewers' comments: Reviewer #1 (Remarks to the Author):
 Editorial Note: This manuscript has been previously reviewed at another journal that is not operating a transparent peer review scheme. This document only contains reviewer comments and rebuttal letters
Editorial Note: This manuscript has been previously reviewed at another journal that is not operating a transparent peer review scheme. This document only contains reviewer comments and rebuttal letters
Optimization of site-specific RNase H cutting of 1037 nt renilla luciferase mrna. The digestion
 SUPPLEMENTARY FIGURE 1. Optimization of site-specific RNase H cutting of 1037 nt renilla luciferase mrna. The digestion mixture was incubated at 37 C for 0 to 120 minutes. The cutting was nearly complete
SUPPLEMENTARY FIGURE 1. Optimization of site-specific RNase H cutting of 1037 nt renilla luciferase mrna. The digestion mixture was incubated at 37 C for 0 to 120 minutes. The cutting was nearly complete
Table 1. Primers, annealing temperatures, and product sizes for PCR amplification.
 Table 1. Primers, annealing temperatures, and product sizes for PCR amplification. Gene Direction Primer sequence (5 3 ) Annealing Temperature Size (bp) BRCA1 Forward TTGCGGGAGGAAAATGGGTAGTTA 50 o C 292
Table 1. Primers, annealing temperatures, and product sizes for PCR amplification. Gene Direction Primer sequence (5 3 ) Annealing Temperature Size (bp) BRCA1 Forward TTGCGGGAGGAAAATGGGTAGTTA 50 o C 292
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Genome-scale engineering of Saccharomyces cerevisiae with single nucleotide precision Zehua Bao, 1 Mohammad HamediRad, 2 Pu Xue, 2 Han Xiao, 1,7 Ipek Tasan, 3 Ran Chao, 2 Jing
SUPPLEMENTARY INFORMATION Genome-scale engineering of Saccharomyces cerevisiae with single nucleotide precision Zehua Bao, 1 Mohammad HamediRad, 2 Pu Xue, 2 Han Xiao, 1,7 Ipek Tasan, 3 Ran Chao, 2 Jing
substrate with ri was incubated with endonuclease V enzymes (25 nm) in buffers with varying ph (7.5-
 Supplementary information Effect of ph, Mg 2+ and Mn 2+ concentrations on endonuclease V activity. (a) Single-stranded RNA substrate with ri was incubated with endonuclease V enzymes (25 nm) in buffers
Supplementary information Effect of ph, Mg 2+ and Mn 2+ concentrations on endonuclease V activity. (a) Single-stranded RNA substrate with ri was incubated with endonuclease V enzymes (25 nm) in buffers
Please write your name or student ID number on every page.
 MCB 110 First Exam A TOTAL OF SIX PAGES NAME: Student ID Number: Question Maximum Points Your Points I 36 II 35 III 27 IV 28 V 24 150 Please write your name or student ID number on every page. This exam
MCB 110 First Exam A TOTAL OF SIX PAGES NAME: Student ID Number: Question Maximum Points Your Points I 36 II 35 III 27 IV 28 V 24 150 Please write your name or student ID number on every page. This exam
American Society of Cytopathology Core Curriculum in Molecular Biology
 American Society of Cytopathology Core Curriculum in Molecular Biology American Society of Cytopathology Core Curriculum in Molecular Biology Chapter 3 Molecular Techniques Alternatives to PCR, Part I
American Society of Cytopathology Core Curriculum in Molecular Biology American Society of Cytopathology Core Curriculum in Molecular Biology Chapter 3 Molecular Techniques Alternatives to PCR, Part I
Mammalian Abasic Site Base Excision Repair
 THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 273, No. 33, Issue of August 14, pp. 21203 21209, 1998 Printed in U.S.A. Mammalian Abasic Site Base Excision Repair IDENTIFICATION OF THE REACTION SEQUENCE AND
THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 273, No. 33, Issue of August 14, pp. 21203 21209, 1998 Printed in U.S.A. Mammalian Abasic Site Base Excision Repair IDENTIFICATION OF THE REACTION SEQUENCE AND
Supplementary Figure 1. Analysis of replication rates by Pol. Nature Structural & Molecular Biology: doi: /nsmb.3207
 Supplementary Figure 1 Analysis of replication rates by Pol (a) Electrophoretic Mobility Shift Assay (EMSA) of Pol -DV binding to template-primer DNA (Fried, M. & Crothers, D.M. Equilibria and kinetics
Supplementary Figure 1 Analysis of replication rates by Pol (a) Electrophoretic Mobility Shift Assay (EMSA) of Pol -DV binding to template-primer DNA (Fried, M. & Crothers, D.M. Equilibria and kinetics
Methods (detailed). for all the experiments. Cells were grown in 50 ml YEA. The cells were harvested and
 Methods (detailed). Purification of yeast chromosomal DNA. Strain JZ105 (mat1m Δmat2,3::LEU2, ade6-210, leu1-32, ura4-d18, his2) was used for all the experiments. Cells were grown in 50 ml YEA. The cells
Methods (detailed). Purification of yeast chromosomal DNA. Strain JZ105 (mat1m Δmat2,3::LEU2, ade6-210, leu1-32, ura4-d18, his2) was used for all the experiments. Cells were grown in 50 ml YEA. The cells
RFC 38: Building Blocks - Standard Large DNA/Genome Construction. Jef Boeke, James DiCarlo. 7 October 2009
 RFC 38: Building Blocks - Standard Large DNA/Genome Construction Jef Boeke, James DiCarlo 7 October 2009 1. Purpose The physical assembly of standard parts is currently a non-standard process, which can
RFC 38: Building Blocks - Standard Large DNA/Genome Construction Jef Boeke, James DiCarlo 7 October 2009 1. Purpose The physical assembly of standard parts is currently a non-standard process, which can
* + * RecA * + + for RecA-dependent strand + + MIT Department of Biology 7.28, Spring Molecular Biology
 MIT Department of Biology 7.28, Spring 2005 - Molecular Biology 1) Question 1. 1A. You are studying the function of in vitro along with a lab partner. You run several reactions to examine the requirements
MIT Department of Biology 7.28, Spring 2005 - Molecular Biology 1) Question 1. 1A. You are studying the function of in vitro along with a lab partner. You run several reactions to examine the requirements
Supplemental Information
 Supplemental Information ATP-dependent unwinding of U4/U6 snrnas by the Brr2 helicase requires the C-terminus of Prp8 Corina Maeder 1,3, Alan K. Kutach 1,2,3, and Christine Guthrie 1 1 Department of Biochemistry
Supplemental Information ATP-dependent unwinding of U4/U6 snrnas by the Brr2 helicase requires the C-terminus of Prp8 Corina Maeder 1,3, Alan K. Kutach 1,2,3, and Christine Guthrie 1 1 Department of Biochemistry
Chapter 9 Preview - DNA
 Chapter 9 Preview - DNA Multiple Choice Identify the choice that best completes the statement or answers the question. 1. In order to show that DNA in cell extracts is responsible for genetic transformation
Chapter 9 Preview - DNA Multiple Choice Identify the choice that best completes the statement or answers the question. 1. In order to show that DNA in cell extracts is responsible for genetic transformation
DNA Replication in Prokaryotes and Eukaryotes
 DNA Replication in Prokaryotes and Eukaryotes 1. Overall mechanism 2. Roles of Polymerases & other proteins 3. More mechanism: Initiation and Termination 4. Mitochondrial DNA replication DNA replication
DNA Replication in Prokaryotes and Eukaryotes 1. Overall mechanism 2. Roles of Polymerases & other proteins 3. More mechanism: Initiation and Termination 4. Mitochondrial DNA replication DNA replication
III. Detailed Examination of the Mechanism of Replication A. Initiation B. Priming C. Elongation D. Proofreading and Termination
 Outline for Replication I. General Features of Replication A. Semi-Conservative B. Starts at Origin C. Bidirectional D. Semi-Discontinuous II. Proteins and Enzymes of Replication III. Detailed Examination
Outline for Replication I. General Features of Replication A. Semi-Conservative B. Starts at Origin C. Bidirectional D. Semi-Discontinuous II. Proteins and Enzymes of Replication III. Detailed Examination
Site-directed Mutagenesis
 Site-directed Mutagenesis Applications Subtilisin (Met à Ala mutation resistant to oxidation) Fluorescent proteins Protein structure-function Substrate trapping mutants Identify regulatory regions/sequences
Site-directed Mutagenesis Applications Subtilisin (Met à Ala mutation resistant to oxidation) Fluorescent proteins Protein structure-function Substrate trapping mutants Identify regulatory regions/sequences
Regulation of ARE transcript 3 end processing by the. yeast Cth2 mrna decay factor
 Regulation of ARE transcript 3 end processing by the yeast Cth2 mrna decay factor Manoël Prouteau, Marie-Claire Daugeron and Bertrand Séraphin Supplementary Information Material and Methods Plasmid construction
Regulation of ARE transcript 3 end processing by the yeast Cth2 mrna decay factor Manoël Prouteau, Marie-Claire Daugeron and Bertrand Séraphin Supplementary Information Material and Methods Plasmid construction
Vicinal abasic site Impair processing of Tg:G Mismatch and 8- Oxoguanine Lesions in Three-Component Bistranded Clustered DNA Damage
 Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2018 Vicinal abasic site Impair processing of Tg:G Mismatch and 8- Oxoguanine Lesions in Three-Component
Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2018 Vicinal abasic site Impair processing of Tg:G Mismatch and 8- Oxoguanine Lesions in Three-Component
SUPPLEMENTAL MATERIAL. Supplemental material contains Supplemental Figure Legends and Supplemental Figures 1 to
 SUPPLEMENTAL MATERIAL Supplemental material contains Supplemental Figure Legends and Supplemental Figures 1 to 6. SUPPLEMENTAL FIGURE LEGENDS Supplemental Figure 1. Overview of the mechanisms by which
SUPPLEMENTAL MATERIAL Supplemental material contains Supplemental Figure Legends and Supplemental Figures 1 to 6. SUPPLEMENTAL FIGURE LEGENDS Supplemental Figure 1. Overview of the mechanisms by which
Biochemical analyses indicate that binding and cleavage specificities define the ordered processing of human Okazaki fragments by Dna2 and FEN1
 6774 6786 Nucleic Acids Research, 2012, Vol. 40, No. 14 Published online 7 May 2012 doi:10.1093/nar/gks388 Biochemical analyses indicate that binding and cleavage specificities define the ordered processing
6774 6786 Nucleic Acids Research, 2012, Vol. 40, No. 14 Published online 7 May 2012 doi:10.1093/nar/gks388 Biochemical analyses indicate that binding and cleavage specificities define the ordered processing
CircLigase II ssdna Ligase
 Cat. Nos. CL9021K and CL9025K www.lucigen.com MA298E-CircLigase II ssdna Ligase 2/2018 1 1. Introduction CircLigase II ssdna Ligase is a thermostable ligase that catalyzes iintramolecular ligation (i.e.,
Cat. Nos. CL9021K and CL9025K www.lucigen.com MA298E-CircLigase II ssdna Ligase 2/2018 1 1. Introduction CircLigase II ssdna Ligase is a thermostable ligase that catalyzes iintramolecular ligation (i.e.,
Exam 2 Key - Spring 2008 A#: Please see us if you have any questions!
 Page 1 of 5 Exam 2 Key - Spring 2008 A#: Please see us if you have any questions! 1. A mutation in which parts of two nonhomologous chromosomes change places is called a(n) A. translocation. B. transition.
Page 1 of 5 Exam 2 Key - Spring 2008 A#: Please see us if you have any questions! 1. A mutation in which parts of two nonhomologous chromosomes change places is called a(n) A. translocation. B. transition.
The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity
 Promega Notes Magazine Number 62, 1997, p. 02 The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity By Christine Andrews and Scott Lesley Promega
Promega Notes Magazine Number 62, 1997, p. 02 The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity By Christine Andrews and Scott Lesley Promega
A Guide to CRISPR/Cas9
 Genome editing and beyond freepik A Guide to CRISPR/Cas9 The latest advance in genomic DNA editing is the Clustered Regularly Interspaced Short Palindromic Repeat (CRISPR)/Cas9 system. This simple-touse
Genome editing and beyond freepik A Guide to CRISPR/Cas9 The latest advance in genomic DNA editing is the Clustered Regularly Interspaced Short Palindromic Repeat (CRISPR)/Cas9 system. This simple-touse
Lecture 22: Molecular techniques DNA cloning and DNA libraries
 Lecture 22: Molecular techniques DNA cloning and DNA libraries DNA cloning: general strategy -> to prepare large quantities of identical DNA Vector + DNA fragment Recombinant DNA (any piece of DNA derived
Lecture 22: Molecular techniques DNA cloning and DNA libraries DNA cloning: general strategy -> to prepare large quantities of identical DNA Vector + DNA fragment Recombinant DNA (any piece of DNA derived
T4 RNA Ligase 2 truncated active site mutants: improved tools for RNA analysis
 Viollet et al. BMC Biotechnology 211, 11:72 http://www.biomedcentral.com/1472-6/11/72 RESEARCH ARTICLE Open Access T4 RNA Ligase 2 truncated active site mutants: improved tools for RNA analysis Sebastien
Viollet et al. BMC Biotechnology 211, 11:72 http://www.biomedcentral.com/1472-6/11/72 RESEARCH ARTICLE Open Access T4 RNA Ligase 2 truncated active site mutants: improved tools for RNA analysis Sebastien
DNA Replication II Biochemistry 302. Bob Kelm January 28, 2004
 DNA Replication II Biochemistry 302 Bob Kelm January 28, 2004 Conceptual model for proofreading based on kinetic considerations Fig. 24.44 stalling transient melting exonuclease site occupancy Following
DNA Replication II Biochemistry 302 Bob Kelm January 28, 2004 Conceptual model for proofreading based on kinetic considerations Fig. 24.44 stalling transient melting exonuclease site occupancy Following
Computational Biology I LSM5191
 Computational Biology I LSM5191 Lecture 5 Notes: Genetic manipulation & Molecular Biology techniques Broad Overview of: Enzymatic tools in Molecular Biology Gel electrophoresis Restriction mapping DNA
Computational Biology I LSM5191 Lecture 5 Notes: Genetic manipulation & Molecular Biology techniques Broad Overview of: Enzymatic tools in Molecular Biology Gel electrophoresis Restriction mapping DNA
Proofreading, post-replication modification of DNA. Mitesh Shrestha
 Proofreading, post-replication modification of DNA Mitesh Shrestha Proofreading During DNA replication (copying), most DNA polymerases can check their work with each base that they add. This process is
Proofreading, post-replication modification of DNA Mitesh Shrestha Proofreading During DNA replication (copying), most DNA polymerases can check their work with each base that they add. This process is
DNA supercoiling, a critical signal regulating the basal expression of the lac operon in Escherichia coli
 Supplementary Information DNA supercoiling, a critical signal regulating the basal expression of the lac operon in Escherichia coli Geraldine Fulcrand 1,2, Samantha Dages 1,2, Xiaoduo Zhi 1,2, Prem Chapagain
Supplementary Information DNA supercoiling, a critical signal regulating the basal expression of the lac operon in Escherichia coli Geraldine Fulcrand 1,2, Samantha Dages 1,2, Xiaoduo Zhi 1,2, Prem Chapagain
Mechanisms of DNA Synthesis
 Mechanisms of DNA Synthesis Biochemistry 201 Advanced Molecular Biology January 19, 2000 Doug Brutlag Introduction Today's topic concerns the fundamental mechanism of DNA replication as exemplified by
Mechanisms of DNA Synthesis Biochemistry 201 Advanced Molecular Biology January 19, 2000 Doug Brutlag Introduction Today's topic concerns the fundamental mechanism of DNA replication as exemplified by
CircLigase II ssdna Ligase
 Cat. Nos. CL9021K and CL9025K Connect with Epicentre on our blog (epicentral.blogspot.com), Facebook (facebook.com/epicentrebio), and Twitter (@EpicentreBio). www.epicentre.com Lit. # 298 12/2012 1 EPILIT298
Cat. Nos. CL9021K and CL9025K Connect with Epicentre on our blog (epicentral.blogspot.com), Facebook (facebook.com/epicentrebio), and Twitter (@EpicentreBio). www.epicentre.com Lit. # 298 12/2012 1 EPILIT298
CircLigase ssdna Ligase
 Cat. Nos. CL4111K and CL4115K Connect with Epicentre on our blog (epicentral.blogspot.com), Facebook (facebook.com/epicentrebio), and Twitter (@EpicentreBio). www.epicentre.com Lit. # 222 10/2012 1 EPILIT222
Cat. Nos. CL4111K and CL4115K Connect with Epicentre on our blog (epicentral.blogspot.com), Facebook (facebook.com/epicentrebio), and Twitter (@EpicentreBio). www.epicentre.com Lit. # 222 10/2012 1 EPILIT222
Multiple choice questions (numbers in brackets indicate the number of correct answers)
 1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
PCR Amplifies Targeted Sequence
 PCR Amplifies Targeted Sequence Target Sequence Supercoiled DNA Strand DNA Strand Double Helix DNA Strand Chromosome P C R PCR PCR = Polymerase Chain Reaction. A primer directed-extension reaction for
PCR Amplifies Targeted Sequence Target Sequence Supercoiled DNA Strand DNA Strand Double Helix DNA Strand Chromosome P C R PCR PCR = Polymerase Chain Reaction. A primer directed-extension reaction for
CFTR-null wt CFTR-null 1.0. Probe: Neo R. Figure S1
 A. B. 4.0 3.0 2.0 1.0 4.0 3.0 2.0 1.0 1 2 3 4 5 6 7 8 9 10 11 12 13 14 Probe: Neo R CFTR-null wt CFTR-null Figure S1 A. 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 10kb 8kb CFTR-null wt B. Probe: CFTR
A. B. 4.0 3.0 2.0 1.0 4.0 3.0 2.0 1.0 1 2 3 4 5 6 7 8 9 10 11 12 13 14 Probe: Neo R CFTR-null wt CFTR-null Figure S1 A. 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 10kb 8kb CFTR-null wt B. Probe: CFTR
PDIP46 (DNA polymerase δ interacting protein 46) is an activating factor for human DNA polymerase δ
 PDIP46 (DNA polymerase δ interacting protein 46) is an activating factor for human DNA polymerase δ Supplementary Material Figure S1. PDIP46 is associated with Pol isolated by immunoaffinity chromatography.
PDIP46 (DNA polymerase δ interacting protein 46) is an activating factor for human DNA polymerase δ Supplementary Material Figure S1. PDIP46 is associated with Pol isolated by immunoaffinity chromatography.
DNA SEQUENCING BY SANGER METHOD
 DNA SEQUENCING BY SANGER METHOD First method described by Sanger and Coulson,1975 for DNA sequencing was called plus and minus. This method used E.coli DNA polymerase I and DNA ploymerase from bacteriophage
DNA SEQUENCING BY SANGER METHOD First method described by Sanger and Coulson,1975 for DNA sequencing was called plus and minus. This method used E.coli DNA polymerase I and DNA ploymerase from bacteriophage
CGTAAGAGTACGTCCAGCATCGGF ATCATTATCTACATCXXXXXTACCATTCATTCAGATCTCACTATCGCATTCTCATGCAGGTCGTAGCCXS
 5 CGTAAGAGTACGTCCAGCATCGGF ATCATTATCTACATCXXXXXTACCATTCATTCAGATCTCACTATCGCATTCTCATGCAGGTCGTAGCCXS 29 Supplementary Figure 1 Sequence of the DNA primer/template pair used in Figure 1bc. A 23nt primer is
5 CGTAAGAGTACGTCCAGCATCGGF ATCATTATCTACATCXXXXXTACCATTCATTCAGATCTCACTATCGCATTCTCATGCAGGTCGTAGCCXS 29 Supplementary Figure 1 Sequence of the DNA primer/template pair used in Figure 1bc. A 23nt primer is
Understanding the Cellular Mechanism of the Excess Microsporocytes I (EMSI) Gene. Andrew ElBardissi, The Pennsylvania State University
 Understanding the Cellular Mechanism of the Excess Microsporocytes I (EMSI) Gene Andrew ElBardissi, The Pennsylvania State University Abstract: Hong Ma, The Pennsylvania State University The Excess Microsporocytes
Understanding the Cellular Mechanism of the Excess Microsporocytes I (EMSI) Gene Andrew ElBardissi, The Pennsylvania State University Abstract: Hong Ma, The Pennsylvania State University The Excess Microsporocytes
University of Groningen
 University of Groningen Coupling dttp Hydrolysis with DNA Unwinding by the DNA Helicase of Bacteriophage T7 Satapathy, Ajit K.; Kulczyk, Arkadiusz W.; Ghosh, Sharmistha; van Oijen, Antonius; Richardson,
University of Groningen Coupling dttp Hydrolysis with DNA Unwinding by the DNA Helicase of Bacteriophage T7 Satapathy, Ajit K.; Kulczyk, Arkadiusz W.; Ghosh, Sharmistha; van Oijen, Antonius; Richardson,
Supplementary Information for:
 Supplementary Information for: A streamlined and high-throughput targeting approach for human germline and cancer genomes using Oligonucleotide-Selective Sequencing Samuel Myllykangas 1, Jason D. Buenrostro
Supplementary Information for: A streamlined and high-throughput targeting approach for human germline and cancer genomes using Oligonucleotide-Selective Sequencing Samuel Myllykangas 1, Jason D. Buenrostro
Chapter 11 DNA Replication and Recombination
 Chapter 11 DNA Replication and Recombination Copyright Copyright 2009 Pearson 2009 Pearson Education, Education, Inc. Inc. 11.1 DNA is reproduced by Semiconservative Replication The complementarity of
Chapter 11 DNA Replication and Recombination Copyright Copyright 2009 Pearson 2009 Pearson Education, Education, Inc. Inc. 11.1 DNA is reproduced by Semiconservative Replication The complementarity of
SUPPLEMENTARY INFORMATION
 doi:1.138/nature1895 i Core histones CAF-1 v ii Reb1 CAF-1 Fen1 vi Mature Okazaki fragment iii PCNA Reb1 iv vii Mature Okazaki fragment Reb1 Mature Okazaki fragments Figure S1. Model for nucleosome-mediated
doi:1.138/nature1895 i Core histones CAF-1 v ii Reb1 CAF-1 Fen1 vi Mature Okazaki fragment iii PCNA Reb1 iv vii Mature Okazaki fragment Reb1 Mature Okazaki fragments Figure S1. Model for nucleosome-mediated
Electronic Supplementary Information. Simultaneously sensitive detection of multiple DNA glycosylases from lung
 Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Simultaneously sensitive detection of multiple DNA
Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Simultaneously sensitive detection of multiple DNA
MIDTERM I NAME: Student ID Number: I 32 II 33 III 24 IV 30 V
 MIDTERM I NAME: Student ID Number: Question Maximum Points Your Points I 32 II 33 III 24 IV 30 V 31 150 Please write your name/student ID number on each of the following five pages. This exam must be written
MIDTERM I NAME: Student ID Number: Question Maximum Points Your Points I 32 II 33 III 24 IV 30 V 31 150 Please write your name/student ID number on each of the following five pages. This exam must be written
Chapter 16 DNA: The Genetic Material. The Nature of Genetic Material. Chemical Nature of Nucleic Acids. Chromosomes - DNA and protein
 Chapter 16 DNA: The Genetic Material The Nature of Genetic Material Chromosomes - DNA and protein Genes are subunits DNA = 4 similar nucleotides C(ytosine) A(denine) T(hymine) G(uanine) Proteins = 20 different
Chapter 16 DNA: The Genetic Material The Nature of Genetic Material Chromosomes - DNA and protein Genes are subunits DNA = 4 similar nucleotides C(ytosine) A(denine) T(hymine) G(uanine) Proteins = 20 different
BCMB Chapters 34 & 35 DNA Replication and Repair
 BCMB 3100 - Chapters 34 & 35 DNA Replication and Repair Semi-conservative DNA replication DNA polymerase DNA replication Replication fork; Okazaki fragments Sanger method for DNA sequencing DNA repair
BCMB 3100 - Chapters 34 & 35 DNA Replication and Repair Semi-conservative DNA replication DNA polymerase DNA replication Replication fork; Okazaki fragments Sanger method for DNA sequencing DNA repair
BCMB Chapters 34 & 35 DNA Replication and Repair
 BCMB 3100 - Chapters 34 & 35 DNA Replication and Repair Semi-conservative DNA replication DNA polymerase DNA replication Replication fork; Okazaki fragments Sanger method for DNA sequencing DNA repair
BCMB 3100 - Chapters 34 & 35 DNA Replication and Repair Semi-conservative DNA replication DNA polymerase DNA replication Replication fork; Okazaki fragments Sanger method for DNA sequencing DNA repair
The 5' cap (red) is added before synthesis of the primary transcript is complete. A non coding sequence following the last exon is shown in orange.
 RNA PROCESSING The 5' cap (red) is added before synthesis of the primary transcript is complete. A non coding sequence following the last exon is shown in orange. Splicing can occur either before or after
RNA PROCESSING The 5' cap (red) is added before synthesis of the primary transcript is complete. A non coding sequence following the last exon is shown in orange. Splicing can occur either before or after
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature18622 Supplementary Discussion Discussion of prior work on NCIN hypothesis. An early study reported incorporation of NAD + into initial products by E. coli RNAP
SUPPLEMENTARY INFORMATION doi:10.1038/nature18622 Supplementary Discussion Discussion of prior work on NCIN hypothesis. An early study reported incorporation of NAD + into initial products by E. coli RNAP
MCB 110 SP 2005 Second Midterm
 MCB110 SP 2005 Midterm II Page 1 ANSWER KEY MCB 110 SP 2005 Second Midterm Question Maximum Points Your Points I 40 II 40 III 36 IV 34 150 NOTE: This is my answer key (KC). The graders were given disgression
MCB110 SP 2005 Midterm II Page 1 ANSWER KEY MCB 110 SP 2005 Second Midterm Question Maximum Points Your Points I 40 II 40 III 36 IV 34 150 NOTE: This is my answer key (KC). The graders were given disgression
Chapter 20 Biotechnology
 Chapter 20 Biotechnology Manipulation of DNA In 2007, the first entire human genome had been sequenced. The ability to sequence an organisms genomes were made possible by advances in biotechnology, (the
Chapter 20 Biotechnology Manipulation of DNA In 2007, the first entire human genome had been sequenced. The ability to sequence an organisms genomes were made possible by advances in biotechnology, (the
An overhang-based DNA block shuffling method for creating a customized random library
 An overhang-based DNA block shuffling method for creating a customized random library Kosuke Fujishima, Chris Venter, Kendrick Wang, Raphael Ferreira and Lynn J. Rothschild Supplementary figures and text:
An overhang-based DNA block shuffling method for creating a customized random library Kosuke Fujishima, Chris Venter, Kendrick Wang, Raphael Ferreira and Lynn J. Rothschild Supplementary figures and text:
Fundamentals of Real Time PCR
 Fundamentals of Real Time PCR Mohamed Abdel Fattah Senior Technical Specialist Scientific Support - Molecular Biology Dept. AnalysisAB Co. Mobile: 012 27 906 74 E-mail: analysis@analysis-ab.com What is
Fundamentals of Real Time PCR Mohamed Abdel Fattah Senior Technical Specialist Scientific Support - Molecular Biology Dept. AnalysisAB Co. Mobile: 012 27 906 74 E-mail: analysis@analysis-ab.com What is
Construct Design and Cloning Guide for Cas9-triggered homologous recombination
 Construct Design and Cloning Guide for Cas9-triggered homologous recombination Written by Dan Dickinson (ddickins@live.unc.edu) and last updated December 2013. Reference: Dickinson DJ, Ward JD, Reiner
Construct Design and Cloning Guide for Cas9-triggered homologous recombination Written by Dan Dickinson (ddickins@live.unc.edu) and last updated December 2013. Reference: Dickinson DJ, Ward JD, Reiner
DNA REPLICATION. Anna Onofri Liceo «I.Versari»
 DNA REPLICATION Anna Onofri Liceo «I.Versari» Learning objectives 1. Understand the basic rules governing DNA replication 2. Understand the function of key proteins involved in a generalised replication
DNA REPLICATION Anna Onofri Liceo «I.Versari» Learning objectives 1. Understand the basic rules governing DNA replication 2. Understand the function of key proteins involved in a generalised replication
Supplementary Figure 1 Two distinct conformational states of the HNH domain in crystal structures. a
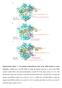 Supplementary Figure 1 Two distinct conformational states of the HNH domain in crystal structures. a HNH-state 1 in PDB 4OO8, in which the distance from the C atom of the HNH catalytic residue 840 to the
Supplementary Figure 1 Two distinct conformational states of the HNH domain in crystal structures. a HNH-state 1 in PDB 4OO8, in which the distance from the C atom of the HNH catalytic residue 840 to the
Kinetic Mechanism of Human DNA Ligase I Reveals Magnesium-dependent Changes in the Rate-limiting Step That Compromise Ligation Efficiency*
 THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 286, NO. 26, pp. 23054 23062, July 1, 2011 2011 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in the U.S.A. Kinetic Mechanism of
THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 286, NO. 26, pp. 23054 23062, July 1, 2011 2011 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in the U.S.A. Kinetic Mechanism of
Basics of Recombinant DNA Technology Biochemistry 302. March 5, 2004 Bob Kelm
 Basics of Recombinant DNA Technology Biochemistry 302 March 5, 2004 Bob Kelm Applications of recombinant DNA technology Mapping and identifying genes (DNA cloning) Propagating genes (DNA subcloning) Modifying
Basics of Recombinant DNA Technology Biochemistry 302 March 5, 2004 Bob Kelm Applications of recombinant DNA technology Mapping and identifying genes (DNA cloning) Propagating genes (DNA subcloning) Modifying
Welcome to Class 18! Lecture 18: Outline and Objectives. Replication is semiconservative! Replication: DNA DNA! Introductory Biochemistry!
 Lecture 18: Outline and Objectives Welcome to Class 18! Introductory Biochemistry! l DNA Replication! l DNA polymerase! l the enzymatic reaction! l proofreading and accuracy! l DNA synthesis! l origins
Lecture 18: Outline and Objectives Welcome to Class 18! Introductory Biochemistry! l DNA Replication! l DNA polymerase! l the enzymatic reaction! l proofreading and accuracy! l DNA synthesis! l origins
(56) References cited:
 (19) TEPZZ 4488B_T (11) EP 2 4 488 B1 (12) EUROPEAN PATENT SPECIFICATION (4) Date of publication and mention of the grant of the patent: 07.06.17 Bulletin 17/23 (21) Application number: 1726. (1) Int Cl.:
(19) TEPZZ 4488B_T (11) EP 2 4 488 B1 (12) EUROPEAN PATENT SPECIFICATION (4) Date of publication and mention of the grant of the patent: 07.06.17 Bulletin 17/23 (21) Application number: 1726. (1) Int Cl.:
Genetics and Genomics in Medicine Chapter 3. Questions & Answers
 Genetics and Genomics in Medicine Chapter 3 Multiple Choice Questions Questions & Answers Question 3.1 Which of the following statements, if any, is false? a) Amplifying DNA means making many identical
Genetics and Genomics in Medicine Chapter 3 Multiple Choice Questions Questions & Answers Question 3.1 Which of the following statements, if any, is false? a) Amplifying DNA means making many identical
The Leu22Pro tumor-associated variant of DNA polymerase beta is drp lyase deficient
 Published online 26 November 2007 Nucleic Acids Research, 2008, Vol. 36, No. 2 411 422 doi:10.1093/nar/gkm1053 The Leu22Pro tumor-associated variant of DNA polymerase beta is drp lyase deficient Shibani
Published online 26 November 2007 Nucleic Acids Research, 2008, Vol. 36, No. 2 411 422 doi:10.1093/nar/gkm1053 The Leu22Pro tumor-associated variant of DNA polymerase beta is drp lyase deficient Shibani
human Cdc45 Figure 1c. (c)
 1 Details of the refined crystallographic model of human Cdc45 and comparison of its active-site region with that of bacterial RecJ. (a) Stereo view of a representative example of the final 2F o -F c electron
1 Details of the refined crystallographic model of human Cdc45 and comparison of its active-site region with that of bacterial RecJ. (a) Stereo view of a representative example of the final 2F o -F c electron
MCB 110 First Midterm SIX PAGES
 MCB 110 First Midterm SIX PAGES NAME: SID Number: Question Maximum Points Your Points I 27 II 32 III 35 IV 33 V 24 151 This exam must be written in PEN if you want the option of a regrade. DO NOT USE WHITE
MCB 110 First Midterm SIX PAGES NAME: SID Number: Question Maximum Points Your Points I 27 II 32 III 35 IV 33 V 24 151 This exam must be written in PEN if you want the option of a regrade. DO NOT USE WHITE
Recitation CHAPTER 9 DNA Technologies
 Recitation CHAPTER 9 DNA Technologies DNA Cloning: General Scheme A cloning vector and eukaryotic chromosomes are separately cleaved with the same restriction endonuclease. (A single chromosome is shown
Recitation CHAPTER 9 DNA Technologies DNA Cloning: General Scheme A cloning vector and eukaryotic chromosomes are separately cleaved with the same restriction endonuclease. (A single chromosome is shown
BEST QUALITY HIGHEST PURITY. Recombinant ENZYMES & PROTEINS
 BEST QUALITY HIGHEST PURITY Recombinant ENZYMES & PROTEINS We offer a wide range of highest quality enzymes and proteins for molecular biology including DNA polymerases, reverse transcriptases, DNA ligases,
BEST QUALITY HIGHEST PURITY Recombinant ENZYMES & PROTEINS We offer a wide range of highest quality enzymes and proteins for molecular biology including DNA polymerases, reverse transcriptases, DNA ligases,
Bacterial DNA replication
 Bacterial DNA replication Summary: What problems do these proteins solve? Tyr OH attacks PO4 and forms a covalent intermediate Structural changes in the protein open the gap by 20 Å! 1 Summary: What problems
Bacterial DNA replication Summary: What problems do these proteins solve? Tyr OH attacks PO4 and forms a covalent intermediate Structural changes in the protein open the gap by 20 Å! 1 Summary: What problems
DNA vs. RNA DNA: deoxyribonucleic acid (double stranded) RNA: ribonucleic acid (single stranded) Both found in most bacterial and eukaryotic cells RNA
 DNA Replication DNA vs. RNA DNA: deoxyribonucleic acid (double stranded) RNA: ribonucleic acid (single stranded) Both found in most bacterial and eukaryotic cells RNA molecule can assume different structures
DNA Replication DNA vs. RNA DNA: deoxyribonucleic acid (double stranded) RNA: ribonucleic acid (single stranded) Both found in most bacterial and eukaryotic cells RNA molecule can assume different structures
4) separates the DNA strands during replication a. A b. B c. C d. D e. E. 5) covalently connects segments of DNA a. A b. B c. C d. D e.
 1) Chargaff's analysis of the relative base composition of DNA was significant because he was able to show that a. the relative proportion of each of the four bases differs from species to species. b.
1) Chargaff's analysis of the relative base composition of DNA was significant because he was able to show that a. the relative proportion of each of the four bases differs from species to species. b.
All This For Four Letters!?! DNA and Its Role in Heredity
 All This For Four Letters!?! DNA and Its Role in Heredity What Is the Evidence that the Gene Is DNA? By the 1920s, it was known that chromosomes consisted of DNA and proteins. A new dye stained DNA and
All This For Four Letters!?! DNA and Its Role in Heredity What Is the Evidence that the Gene Is DNA? By the 1920s, it was known that chromosomes consisted of DNA and proteins. A new dye stained DNA and
Thebiotutor.com A2 Biology OCR Unit F215: Control, genomes and environment Module 2.3 Genomes and gene technologies Answers
 Thebiotutor.com A2 Biology OCR Unit F215: Control, genomes and environment Module 2.3 Genomes and gene technologies Answers Andy Todd 1 1. (i) plasmid cut by restriction enzyme; at specific sequence; same
Thebiotutor.com A2 Biology OCR Unit F215: Control, genomes and environment Module 2.3 Genomes and gene technologies Answers Andy Todd 1 1. (i) plasmid cut by restriction enzyme; at specific sequence; same
DNA polymerase proofreading: active site switching catalyzed by the bacteriophage T4 DNA polymerase
 5452 5463 Nucleic Acids Research, 2007, Vol. 35, No. 16 Published online 15 August 2007 doi:10.1093/nar/gkm591 DNA polymerase proofreading: active site switching catalyzed by the bacteriophage T4 DNA polymerase
5452 5463 Nucleic Acids Research, 2007, Vol. 35, No. 16 Published online 15 August 2007 doi:10.1093/nar/gkm591 DNA polymerase proofreading: active site switching catalyzed by the bacteriophage T4 DNA polymerase
Combinatorial Evolution of Enzymes and Synthetic pathways Using
 Supplementary data Combinatorial Evolution of Enzymes and Synthetic pathways Using One-Step PCR Peng Jin,, Zhen Kang,,, *, Junli Zhang,, Linpei Zhang,, Guocheng Du,, and Jian Chen, The Key Laboratory of
Supplementary data Combinatorial Evolution of Enzymes and Synthetic pathways Using One-Step PCR Peng Jin,, Zhen Kang,,, *, Junli Zhang,, Linpei Zhang,, Guocheng Du,, and Jian Chen, The Key Laboratory of
Chapter 6 - Molecular Genetic Techniques
 Chapter 6 - Molecular Genetic Techniques Two objects of molecular & genetic technologies For analysis For generation Molecular genetic technologies! For analysis DNA gel electrophoresis Southern blotting
Chapter 6 - Molecular Genetic Techniques Two objects of molecular & genetic technologies For analysis For generation Molecular genetic technologies! For analysis DNA gel electrophoresis Southern blotting
In vitro mutagenesis
 Core course BMS361N Genetic Engineering In vitro mutagenesis Prof. Narkunaraja Shanmugam Dept. Of Biomedical Science School of Basic Medical Sciences Bharathidasan University In vitro mutagenesis Many
Core course BMS361N Genetic Engineering In vitro mutagenesis Prof. Narkunaraja Shanmugam Dept. Of Biomedical Science School of Basic Medical Sciences Bharathidasan University In vitro mutagenesis Many
XXII DNA cloning and sequencing. Outline
 XXII DNA cloning and sequencing 1) Deriving DNA for cloning Outline 2) Vectors; forming recombinant DNA; cloning DNA; and screening for clones containing recombinant DNA [replica plating and autoradiography;
XXII DNA cloning and sequencing 1) Deriving DNA for cloning Outline 2) Vectors; forming recombinant DNA; cloning DNA; and screening for clones containing recombinant DNA [replica plating and autoradiography;
Mycobacterium tuberculosis (Mtb) and Mycobacterium smegmatis (Msmeg) contain two ε (dnaq) exonuclease homologs.
 Supplementary Figure 1 Mycobacterium tuberculosis (Mtb) and Mycobacterium smegmatis (Msmeg) contain two ε (dnaq) exonuclease homologs. (a) Sequence alignment of the ε-exonuclease homologs from four different
Supplementary Figure 1 Mycobacterium tuberculosis (Mtb) and Mycobacterium smegmatis (Msmeg) contain two ε (dnaq) exonuclease homologs. (a) Sequence alignment of the ε-exonuclease homologs from four different
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Schwarz UI, Ritchie MD, Bradford Y, et al. Genetic determinants
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Schwarz UI, Ritchie MD, Bradford Y, et al. Genetic determinants
Supplementary Information
 Supplementary Information Vidigal and Ventura a wt locus 5 region 3 region CCTCTGCCACTGCGAGGGCGTCCAATGGTGCTTG(...)AACAGGTGGAATATCCCTACTCTA predicted deletion clone 1 clone 2 clone 3 CCTCTGCCACTGCGAGGGCGTC-AGGTGGAATATCCCTACTCTA
Supplementary Information Vidigal and Ventura a wt locus 5 region 3 region CCTCTGCCACTGCGAGGGCGTCCAATGGTGCTTG(...)AACAGGTGGAATATCCCTACTCTA predicted deletion clone 1 clone 2 clone 3 CCTCTGCCACTGCGAGGGCGTC-AGGTGGAATATCCCTACTCTA
Electronic Supplementary Information
 Electronic Supplementary Information Ultrasensitive quantification of mature micrornas by real-time PCR based on ligation of ribonucleotide-modified DNA probe Jiangyan Zhang, Zhengping Li,* Hui Wang, Yucong
Electronic Supplementary Information Ultrasensitive quantification of mature micrornas by real-time PCR based on ligation of ribonucleotide-modified DNA probe Jiangyan Zhang, Zhengping Li,* Hui Wang, Yucong
DNA. Is a molecule that encodes the genetic instructions used in the development and functioning of all known living organisms and many viruses.
 Is a molecule that encodes the genetic instructions used in the development and functioning of all known living organisms and many viruses. Genetic information is encoded as a sequence of nucleotides (guanine,
Is a molecule that encodes the genetic instructions used in the development and functioning of all known living organisms and many viruses. Genetic information is encoded as a sequence of nucleotides (guanine,
Influence of tlp5, che1p, tlp4a, and che4stas Promoters on Chemotaxis in Azospirillum brasilense
 University of Tennessee, Knoxville Trace: Tennessee Research and Creative Exchange University of Tennessee Honors Thesis Projects University of Tennessee Honors Program 5-2015 Influence of tlp5, che1p,
University of Tennessee, Knoxville Trace: Tennessee Research and Creative Exchange University of Tennessee Honors Thesis Projects University of Tennessee Honors Program 5-2015 Influence of tlp5, che1p,
Directe d Mutagenesis
 Directe d Mutagenesis A Practical Approac h M. J. McPHERSON 1. Mutagenesis facilitated by the removal or introduction of unique restriction sites 1 P. Carte r 1. Introduction to site-directed mutagenesis
Directe d Mutagenesis A Practical Approac h M. J. McPHERSON 1. Mutagenesis facilitated by the removal or introduction of unique restriction sites 1 P. Carte r 1. Introduction to site-directed mutagenesis
supplementary information
 DOI: 10.1038/ncb2007 Figure S1 Temperature sensitivity of DNA ligase I alleles. a, SSL204 (CDC9), SSL612 (cdc9-1) and SSL613 (cdc9-2) cells were spotted in ten-fold serial dilutions on YPD plates and incubated
DOI: 10.1038/ncb2007 Figure S1 Temperature sensitivity of DNA ligase I alleles. a, SSL204 (CDC9), SSL612 (cdc9-1) and SSL613 (cdc9-2) cells were spotted in ten-fold serial dilutions on YPD plates and incubated
Sequencing of DNA lesions facilitated by site-specific excision via base. excision repair DNA glycosylases yielding ligatable gaps
 Supporting information Sequencing of DNA lesions facilitated by site-specific excision via base excision repair DNA glycosylases yielding ligatable gaps Jan Riedl, Aaron M. Fleming, and Cynthia J. Burrows*
Supporting information Sequencing of DNA lesions facilitated by site-specific excision via base excision repair DNA glycosylases yielding ligatable gaps Jan Riedl, Aaron M. Fleming, and Cynthia J. Burrows*
Recombinant DNA Technology
 Recombinant DNA Technology Common General Cloning Strategy Target DNA from donor organism extracted, cut with restriction endonuclease and ligated into a cloning vector cut with compatible restriction
Recombinant DNA Technology Common General Cloning Strategy Target DNA from donor organism extracted, cut with restriction endonuclease and ligated into a cloning vector cut with compatible restriction
Supporting Information
 Supporting Information Deng et al. 10.1073/pnas.1102117108 Fig. S1. Predicted structure of Arabidopsis bzip60 RNA. Lowest free energy form (ΔG = 309.72 (initially 343.10) of bzip60 mrna folded by M-Fold
Supporting Information Deng et al. 10.1073/pnas.1102117108 Fig. S1. Predicted structure of Arabidopsis bzip60 RNA. Lowest free energy form (ΔG = 309.72 (initially 343.10) of bzip60 mrna folded by M-Fold
TwistAmp DNA Amplification Kits Assay Design Manual
 TM Unwind DNA s possibilities TwistAmp DNA Amplification Kits Assay Design Manual Contents 1. Assay Considerations 3 1.1 Reaction temperature 3 1.2 Quantification and amplification onset 3 1.3 Preventing
TM Unwind DNA s possibilities TwistAmp DNA Amplification Kits Assay Design Manual Contents 1. Assay Considerations 3 1.1 Reaction temperature 3 1.2 Quantification and amplification onset 3 1.3 Preventing
Information on barcode decoding
 Information on barcode decoding The Bioneer version 1.0 haploid deletion library (Bioneer catalog number M-1030H) was supplied as glycerol frozen stocks in thirty-one 96-well plates. According to the information
Information on barcode decoding The Bioneer version 1.0 haploid deletion library (Bioneer catalog number M-1030H) was supplied as glycerol frozen stocks in thirty-one 96-well plates. According to the information
