SUPPLEMENTARY INFORMATION
|
|
|
- Violet Cross
- 5 years ago
- Views:
Transcription
1 doi: /nature11988 Supplementary Figure 1. Digestion of model DNA substrates. a, Linearized plasmid DNA (pik31- PstI, lanes 1 and 2), supercoiled plasmid (pik31, lanes 3 and 4), singly nicked plasmid (pik31- Nb.BvCI, lanes 5 and 6) and M13mp18 ssdna (lanes 7 and 8) was digested with S1 for 7.5 minutes at 37ºC. b, A 100-mer circular substrate was hybridized with complementary oligonucleotides to generate 20 nt, 10 nt or 0 nt (nicked substrate) ssdna gaps and then digested with S1 nuclease for the indicated times. 1
2 Supplementary Figure 2. Scheme of the polymerase release assay to monitor signal and collision release during replication. The lagging strand template (i.e. the leading strand product of a rolling circle reaction) is labeled using (α- 32 P)-dTTP and will be resistant to S1 (top diagram) unless a ssdna gap is produced by signal release (bottom diagram). The expected results for 100% collision release and for 100% signal release, are illustrated to the right. Analysis of S1 digested lagging strand template DNA on denaturing agarose gels digests the lagging strand template in the event of signal release, which produces ssdna gaps. In the case of 100% signal release, a ssdna gap will be present between each Okazaki fragment, and S1 digestion will yield products that equal the size of Okazaki fragments. The products of lagging strand Okazaki fragment synthesis are monitored in a separate reaction by addition of (α- 32 P)-dATP. In the case of 100% collision release, only nicks will separate Okazaki fragments, and the lagging strand template will be resistant to S1 cleavage. 2
3 Supplementary Figure 3. Determination of product molecular weights and calculation of frequency of signal release. a, Lane profiles from Fig. 2b were normalized to the corresponding molecular weight at each pixel in order to correct for the fact that longer products incorporate more radiolabel (unbroken lines), and were analyzed by fitting to a single Gaussian distribution (dashed lines). The peak of the fit corresponds to the average length of DNA, which enables the percent of signal release to be calculated (see Supplemental Methods). The fit derived from a single Gaussian analysis reveals a small subpopulation of slower migrating products (a, upper panel), which we presume derive from asynchronously fired replisomes or prematurely terminated forks. b, To ensure that this subpopulation does not affect the outcome of the results, we compared results in which we included this subpopulation by fitting the data to a double Gaussian distribution (upper panel). c, Signal release was calculated using the length (kb) obtained from either the single (1G) or double (2G) Gaussian distribution analysis. The result was essentially unchanged; n = 7 for 2G, and n = 3 for 1G analysis, respectively. d, Mean of the product size was used to calculate signal and collision release as described in Supplemental Methods; n = 5 ± SEM. 3
4 Supplementary Figure 4. Control reactions for the bead-based S1 assay. a, Replication reactions containing (α- 32 P)-dTTP to label the lagging strand template were performed as in Fig. 2b, in the absence of primase (lane 1). The ssdna product is degraded by S1 (lane 2). This control demonstrates that sufficient S1 is used to digest all ssdna that can possibly be produced in the assay in the absence of lagging strand synthesis. b, Replication reactions performed in the presence of (α- 32 P)-dATP to label the lagging strand Okazaki fragments were treated with or S1 (lane 2) as described in Fig. 2. This control shows that duplex regions, formed by Okazaki fragment synthesis, are not digested by S1. c, Replication reactions containing (α- 32 P)-dTTP were quenched, the beads were washed and then resuspended in replication buffer in the presence of 20 nm Pol II and dntps to fill any ssdna gaps prior to S1 analysis. The gap filling reaction shows that the lagging strand template becomes resistant to S1 when Okazaki fragments are complete, as the DNA products look the same in the absence (lane1) or presence (lane 2) of S1 treatment. d, Replication reactions performed in the presence of 5 phosphorylated oligonucleotides to initiate lagging strand synthesis and α- 32 P-dATP to label Okazaki fragments were quenched, the beads washed and incubated with a buffer control (lane 1) or T4 ligase (lane 2). The results show that a significant proportion of Okazaki fragments become ligated, demonstrating that they contain a nick produced by collision release. 4
5 Supplementary Figure 5. Polymerase release is independent of β clamp concentrations. a, Upper panel. S1 analysis of a β clamp titration into replication reactions as described in Fig. 2. Replication reactions contain either (α- 32 P)-dTTP (lanes 1-10), which labels the leading strand (lagging strand template), or (α- 32 P)-dATP (lanes 11-15), which labels Okazaki fragments. Lower panel. Quantification of percent signal release at different concentrations of β. b, Reactions were performed as in panel a, except equimolar amounts of both β clamp and γ-complex (γc) clamp loader were titrated into the replication reaction. 5
6 Supplementary Figure 6. Signal release occurs in the absence of primase. Quantification of the gel in Figure 2d. Percent signal release at different concentrations of DNA oligonucleotides; n = 3 ± SEM. Supplementary Figure 7. Polymerase signal release at different concentrations of primase. a, S1 analysis of replication reactions was performed at the indicated concentrations of primase. b, Quantification of percent signal release at different concentrations of primase; n = 4 ± SEM. 6
7 Supplementary Figure 8. Long Okazaki fragments are preferentially terminated by signal release. a, Schematic illustration of the fill-in reaction to label gaps produced by signal release. b, Alkaline gel analysis of Okazaki fragments produced during rolling circle replication in the presence of 320 nm primase. Reactions were performed in the presence (lanes 1 and 3) or absence (lane 2) of (α 32 P)-dATP to produce either radiolabeled or unlabeled Okazaki fragments, respectively. Quenched reactions were washed, and beads resuspended in replication buffer containing Pol II (lanes 2 and 3) and (α- 32 P)-dATP was added to label Okazaki fragments that are followed by a ssdna gap, and thus are extended by Pol II. Lane 2 shows that Okazaki fragments that are unlabeled in the initial reaction, become labeled by Pol II, and are significantly longer than the bulk of Okazaki fragments labeled during replication (compare with lane 1). Lane 3, with radiolabel in both Pol III and Poll II reactions, shows that the overall distribution of Okazaki fragments becomes lengthened relative to lane 1, but that shorter Okazaki fragments essentially remain unchanged. The scan profiles of each lane are shown to the right of the alkaline gel. The data enable an estimate of the average size of the gaps left during signal release by comparing the signal generated during the fill-in reaction, which reflects the presence of all gaps, to the Okazaki fragment lengths (lanes 1 and 2), using the following calculation: Gaps = Length (fill-in) Length (Okazaki fragments). The result of this calculation yields an average gap size of 800bp. The wide distribution of Okazaki fragment size is due to the fact that primase action is stochastic, resulting in a heterogeneous population of Okazaki fragment sizes. 7
8 Supplementary Figure 9. Representative line profiles of individual DNA strands as described in Fig. 4c and h. a, buffer flow was 10 µl/min. b, buffer flow was 100 µl/min. The data was averaged over a width of 3 pixels and 10 successive frames to minimize signal fluctuations due to lateral or vertical DNA strand motions in the flow. Therefore, the dips in the fluorescence intensity are directly reflecting a variation of the number of YoPro1 intercalators present on DNA, which correlates with the amount of dsdna present. 8
Single Molecule Studies Reveal the Function of a Third Polymerase in the Replisome
 SUPPLEMENTARY ONLINE MATERIAL For manuscript: Single Molecule Studies Reveal the Function of a Third Polymerase in the Replisome By Roxana E. Georgescu 1, Isabel Kurth 1 and Mike E. O Donnell 1 1 The Rockefeller
SUPPLEMENTARY ONLINE MATERIAL For manuscript: Single Molecule Studies Reveal the Function of a Third Polymerase in the Replisome By Roxana E. Georgescu 1, Isabel Kurth 1 and Mike E. O Donnell 1 1 The Rockefeller
Chapter 12. DNA Replication and Recombination
 Chapter 12 DNA Replication and Recombination I. DNA replication Three possible modes of replication A. Conservative entire original molecule maintained B. Semiconservative one strand is template for new
Chapter 12 DNA Replication and Recombination I. DNA replication Three possible modes of replication A. Conservative entire original molecule maintained B. Semiconservative one strand is template for new
DNA Replication II Biochemistry 302. Bob Kelm January 28, 2004
 DNA Replication II Biochemistry 302 Bob Kelm January 28, 2004 Conceptual model for proofreading based on kinetic considerations Fig. 24.44 stalling transient melting exonuclease site occupancy Following
DNA Replication II Biochemistry 302 Bob Kelm January 28, 2004 Conceptual model for proofreading based on kinetic considerations Fig. 24.44 stalling transient melting exonuclease site occupancy Following
Amplified segment of DNA can be purified from bacteria in sufficient quantity and quality for :
 Transformation Insertion of DNA of interest Amplification Amplified segment of DNA can be purified from bacteria in sufficient quantity and quality for : DNA Sequence. Understand relatedness of genes and
Transformation Insertion of DNA of interest Amplification Amplified segment of DNA can be purified from bacteria in sufficient quantity and quality for : DNA Sequence. Understand relatedness of genes and
NIH Public Access Author Manuscript Nature. Author manuscript; available in PMC 2013 October 04.
 NIH Public Access Author Manuscript Published in final edited form as: Nature. 2013 April 4; 496(7443): 119 122. doi:10.1038/nature11988. A solution to release twisted DNA during chromosome replication
NIH Public Access Author Manuscript Published in final edited form as: Nature. 2013 April 4; 496(7443): 119 122. doi:10.1038/nature11988. A solution to release twisted DNA during chromosome replication
DNA Replication II Biochemistry 302. Bob Kelm January 26, 2005
 DNA Replication II Biochemistry 302 Bob Kelm January 26, 2005 Following in Dad s footsteps Original A. Kornberg E. coli DNA Pol I is a lousy replicative enzyme. 400 molecules/cell but ~2 replication forks/cell
DNA Replication II Biochemistry 302 Bob Kelm January 26, 2005 Following in Dad s footsteps Original A. Kornberg E. coli DNA Pol I is a lousy replicative enzyme. 400 molecules/cell but ~2 replication forks/cell
DNA Replication II Biochemistry 302. January 25, 2006
 DNA Replication II Biochemistry 302 January 25, 2006 Following in Dad s footsteps Original A. Kornberg E. coli DNA Pol I is a lousy replicative enzyme. 400 molecules/cell but ~2 replication forks/cell
DNA Replication II Biochemistry 302 January 25, 2006 Following in Dad s footsteps Original A. Kornberg E. coli DNA Pol I is a lousy replicative enzyme. 400 molecules/cell but ~2 replication forks/cell
Supplementary Information for. Coordination of multiple enzyme activities by a single PCNA in archaeal Okazaki fragment.
 Supplementary Information for Coordination of multiple enzyme activities by a single PCNA in archaeal Okazaki fragment maturation Thomas R. Beattie and Stephen D. Bell Sir William Dunn School of Pathology,
Supplementary Information for Coordination of multiple enzyme activities by a single PCNA in archaeal Okazaki fragment maturation Thomas R. Beattie and Stephen D. Bell Sir William Dunn School of Pathology,
* + * RecA * + + for RecA-dependent strand + + MIT Department of Biology 7.28, Spring Molecular Biology
 MIT Department of Biology 7.28, Spring 2005 - Molecular Biology 1) Question 1. 1A. You are studying the function of in vitro along with a lab partner. You run several reactions to examine the requirements
MIT Department of Biology 7.28, Spring 2005 - Molecular Biology 1) Question 1. 1A. You are studying the function of in vitro along with a lab partner. You run several reactions to examine the requirements
Optimization of site-specific RNase H cutting of 1037 nt renilla luciferase mrna. The digestion
 SUPPLEMENTARY FIGURE 1. Optimization of site-specific RNase H cutting of 1037 nt renilla luciferase mrna. The digestion mixture was incubated at 37 C for 0 to 120 minutes. The cutting was nearly complete
SUPPLEMENTARY FIGURE 1. Optimization of site-specific RNase H cutting of 1037 nt renilla luciferase mrna. The digestion mixture was incubated at 37 C for 0 to 120 minutes. The cutting was nearly complete
The Size and Packaging of Genomes
 DNA Replication The Size and Packaging of Genomes Vary greatly in size Ø Smallest viruses- 4 or 5 genes Ø Escherichia coli- 4,288 genes Ø Human cell- 20,000 to 25,000 genes E. coli 4 million base pairs
DNA Replication The Size and Packaging of Genomes Vary greatly in size Ø Smallest viruses- 4 or 5 genes Ø Escherichia coli- 4,288 genes Ø Human cell- 20,000 to 25,000 genes E. coli 4 million base pairs
Lecture Four. Molecular Approaches I: Nucleic Acids
 Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
BIOCHEMISTRY REVIEW. Overview of Biomolecules. Chapter 11 DNA Replication
 BIOCHEMISTRY REVIEW Overview of Biomolecules Chapter 11 DNA Replication 2 3 4 5 6 7 8 9 Are You Getting It?? Which characteristics will be part of semi-conservative replication? (multiple answers) a) The
BIOCHEMISTRY REVIEW Overview of Biomolecules Chapter 11 DNA Replication 2 3 4 5 6 7 8 9 Are You Getting It?? Which characteristics will be part of semi-conservative replication? (multiple answers) a) The
Chapter 11 DNA Replication and Recombination
 Chapter 11 DNA Replication and Recombination Copyright Copyright 2009 Pearson 2009 Pearson Education, Education, Inc. Inc. 11.1 DNA is reproduced by Semiconservative Replication The complementarity of
Chapter 11 DNA Replication and Recombination Copyright Copyright 2009 Pearson 2009 Pearson Education, Education, Inc. Inc. 11.1 DNA is reproduced by Semiconservative Replication The complementarity of
Bootcamp: Molecular Biology Techniques and Interpretation
 Bootcamp: Molecular Biology Techniques and Interpretation Bi8 Winter 2016 Today s outline Detecting and quantifying nucleic acids and proteins: Basic nucleic acid properties Hybridization PCR and Designing
Bootcamp: Molecular Biology Techniques and Interpretation Bi8 Winter 2016 Today s outline Detecting and quantifying nucleic acids and proteins: Basic nucleic acid properties Hybridization PCR and Designing
Bacterial DNA replication
 Bacterial DNA replication Summary: What problems do these proteins solve? Tyr OH attacks PO4 and forms a covalent intermediate Structural changes in the protein open the gap by 20 Å! 1 Summary: What problems
Bacterial DNA replication Summary: What problems do these proteins solve? Tyr OH attacks PO4 and forms a covalent intermediate Structural changes in the protein open the gap by 20 Å! 1 Summary: What problems
Genetics and Genomics in Medicine Chapter 3. Questions & Answers
 Genetics and Genomics in Medicine Chapter 3 Multiple Choice Questions Questions & Answers Question 3.1 Which of the following statements, if any, is false? a) Amplifying DNA means making many identical
Genetics and Genomics in Medicine Chapter 3 Multiple Choice Questions Questions & Answers Question 3.1 Which of the following statements, if any, is false? a) Amplifying DNA means making many identical
CircLigase II ssdna Ligase
 Cat. Nos. CL9021K and CL9025K www.lucigen.com MA298E-CircLigase II ssdna Ligase 2/2018 1 1. Introduction CircLigase II ssdna Ligase is a thermostable ligase that catalyzes iintramolecular ligation (i.e.,
Cat. Nos. CL9021K and CL9025K www.lucigen.com MA298E-CircLigase II ssdna Ligase 2/2018 1 1. Introduction CircLigase II ssdna Ligase is a thermostable ligase that catalyzes iintramolecular ligation (i.e.,
Enzymatic assembly of DNA molecules up to several hundred kilobases
 nature methods Enzymatic assembly of DNA molecules up to several hundred kilobases Daniel G Gibson, Lei Young, Ray-Yuan Chuang, J Craig Venter, Clyde A Hutchison III & Hamilton O Smith Supplementary figures
nature methods Enzymatic assembly of DNA molecules up to several hundred kilobases Daniel G Gibson, Lei Young, Ray-Yuan Chuang, J Craig Venter, Clyde A Hutchison III & Hamilton O Smith Supplementary figures
PDIP46 (DNA polymerase δ interacting protein 46) is an activating factor for human DNA polymerase δ
 PDIP46 (DNA polymerase δ interacting protein 46) is an activating factor for human DNA polymerase δ Supplementary Material Figure S1. PDIP46 is associated with Pol isolated by immunoaffinity chromatography.
PDIP46 (DNA polymerase δ interacting protein 46) is an activating factor for human DNA polymerase δ Supplementary Material Figure S1. PDIP46 is associated with Pol isolated by immunoaffinity chromatography.
You Should Be Able To
 DNA Replica,on You Should Be Able To 1. Describe the func9on of: DNA POL1, DNA POL3, Sliding Clamp, SSBPs, Ligase, Topoisomerase, Helicase, Primase 2. Describe DNA synthesis on the leading and lagging
DNA Replica,on You Should Be Able To 1. Describe the func9on of: DNA POL1, DNA POL3, Sliding Clamp, SSBPs, Ligase, Topoisomerase, Helicase, Primase 2. Describe DNA synthesis on the leading and lagging
Molecular Biology: General Theory
 Molecular Biology: General Theory Author: Dr Darshana Morar Licensed under a Creative Commons Attribution license. DNA REPLICATION DNA replication is the process of duplicating the DNA sequence in the
Molecular Biology: General Theory Author: Dr Darshana Morar Licensed under a Creative Commons Attribution license. DNA REPLICATION DNA replication is the process of duplicating the DNA sequence in the
Molecular Biology: General Theory
 Molecular Biology: General Theory Author: Dr Darshana Morar Licensed under a Creative Commons Attribution license. DNA REPLICATION DNA replication is the process of duplicating the DNA sequence in the
Molecular Biology: General Theory Author: Dr Darshana Morar Licensed under a Creative Commons Attribution license. DNA REPLICATION DNA replication is the process of duplicating the DNA sequence in the
DNA supercoiling, a critical signal regulating the basal expression of the lac operon in Escherichia coli
 Supplementary Information DNA supercoiling, a critical signal regulating the basal expression of the lac operon in Escherichia coli Geraldine Fulcrand 1,2, Samantha Dages 1,2, Xiaoduo Zhi 1,2, Prem Chapagain
Supplementary Information DNA supercoiling, a critical signal regulating the basal expression of the lac operon in Escherichia coli Geraldine Fulcrand 1,2, Samantha Dages 1,2, Xiaoduo Zhi 1,2, Prem Chapagain
Computational Biology I LSM5191
 Computational Biology I LSM5191 Lecture 5 Notes: Genetic manipulation & Molecular Biology techniques Broad Overview of: Enzymatic tools in Molecular Biology Gel electrophoresis Restriction mapping DNA
Computational Biology I LSM5191 Lecture 5 Notes: Genetic manipulation & Molecular Biology techniques Broad Overview of: Enzymatic tools in Molecular Biology Gel electrophoresis Restriction mapping DNA
BCMB Chapters 34 & 35 DNA Replication and Repair
 BCMB 3100 - Chapters 34 & 35 DNA Replication and Repair Semi-conservative DNA replication DNA polymerase DNA replication Replication fork; Okazaki fragments Sanger method for DNA sequencing DNA repair
BCMB 3100 - Chapters 34 & 35 DNA Replication and Repair Semi-conservative DNA replication DNA polymerase DNA replication Replication fork; Okazaki fragments Sanger method for DNA sequencing DNA repair
BCMB Chapters 34 & 35 DNA Replication and Repair
 BCMB 3100 - Chapters 34 & 35 DNA Replication and Repair Semi-conservative DNA replication DNA polymerase DNA replication Replication fork; Okazaki fragments Sanger method for DNA sequencing DNA repair
BCMB 3100 - Chapters 34 & 35 DNA Replication and Repair Semi-conservative DNA replication DNA polymerase DNA replication Replication fork; Okazaki fragments Sanger method for DNA sequencing DNA repair
Lecture 1 Sunday, 4 March :24 pm
 Lecture 1 Sunday, 4 March 2018 10:24 pm Amino acid side chains can be Hydrophobic, hydrophilic Positive, negatively charged Movement of information OH removed from 2' carbon to make the end more stable
Lecture 1 Sunday, 4 March 2018 10:24 pm Amino acid side chains can be Hydrophobic, hydrophilic Positive, negatively charged Movement of information OH removed from 2' carbon to make the end more stable
Requirements for the Genetic Material
 Requirements for the Genetic Material 1. Replication Reproduced and transmitted faithfully from cell to cell-generation to generation. 2. Information Storage Biologically useful information in a stable
Requirements for the Genetic Material 1. Replication Reproduced and transmitted faithfully from cell to cell-generation to generation. 2. Information Storage Biologically useful information in a stable
BIOL3090 Final Exam Summary, Pete s Lectures. Lecture 1: DNA Replication
 BIOL3090 Final Exam Summary, Pete s Lectures Lecture 1: DNA Replication - The five rules of DNA replication: 1. DNA is made of two complementary strands (always) 2. DNA replication is semiconservative
BIOL3090 Final Exam Summary, Pete s Lectures Lecture 1: DNA Replication - The five rules of DNA replication: 1. DNA is made of two complementary strands (always) 2. DNA replication is semiconservative
 The replication of DNA Kornberg 1957 Meselson and Stahl 1958 Cairns 1963 Okazaki 1968 DNA Replication The driving force for DNA synthesis. The addition of a nucleotide to a growing polynucleotide
The replication of DNA Kornberg 1957 Meselson and Stahl 1958 Cairns 1963 Okazaki 1968 DNA Replication The driving force for DNA synthesis. The addition of a nucleotide to a growing polynucleotide
Replication. Obaidur Rahman
 Replication Obaidur Rahman DIRCTION OF DNA SYNTHESIS How many reactions can a DNA polymerase catalyze? So how many reactions can it catalyze? So 4 is one answer, right, 1 for each nucleotide. But what
Replication Obaidur Rahman DIRCTION OF DNA SYNTHESIS How many reactions can a DNA polymerase catalyze? So how many reactions can it catalyze? So 4 is one answer, right, 1 for each nucleotide. But what
CircLigase II ssdna Ligase
 Cat. Nos. CL9021K and CL9025K Connect with Epicentre on our blog (epicentral.blogspot.com), Facebook (facebook.com/epicentrebio), and Twitter (@EpicentreBio). www.epicentre.com Lit. # 298 12/2012 1 EPILIT298
Cat. Nos. CL9021K and CL9025K Connect with Epicentre on our blog (epicentral.blogspot.com), Facebook (facebook.com/epicentrebio), and Twitter (@EpicentreBio). www.epicentre.com Lit. # 298 12/2012 1 EPILIT298
XactEdit Cas9 Nuclease with NLS User Manual
 XactEdit Cas9 Nuclease with NLS User Manual An RNA-guided recombinant endonuclease for efficient targeted DNA cleavage Catalog Numbers CE1000-50K, CE1000-50, CE1000-250, CE1001-250, CE1001-1000 Table of
XactEdit Cas9 Nuclease with NLS User Manual An RNA-guided recombinant endonuclease for efficient targeted DNA cleavage Catalog Numbers CE1000-50K, CE1000-50, CE1000-250, CE1001-250, CE1001-1000 Table of
The replication forks Summarising what we know:
 When does replication occur? MBLG1001 lecture 10 Replication the once in a lifetime event! Full blown replication only occurs once, just before cell division BUT the DNA template is constantly being repaired.
When does replication occur? MBLG1001 lecture 10 Replication the once in a lifetime event! Full blown replication only occurs once, just before cell division BUT the DNA template is constantly being repaired.
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Reconfigurable, Braced, Three-Dimensional DNA Nanostructures Russell P. Goodman 1, Mike Heilemann 1, 2, Sören Doose 1, 2, Christoph M. Erben 1, Achillefs N. Kapanidis 1, and Andrew
SUPPLEMENTARY INFORMATION Reconfigurable, Braced, Three-Dimensional DNA Nanostructures Russell P. Goodman 1, Mike Heilemann 1, 2, Sören Doose 1, 2, Christoph M. Erben 1, Achillefs N. Kapanidis 1, and Andrew
CSIR UGC NET, GATE (ENGINEERING), GATE (Science), IIT-JAM, UGC NET, TIFR, IISc, NIMCET, JEST etc. JNU CEEB SAMPLE THEORY
 JNU CEEB SAMPLE THEORY REPLICATION TYPE OF DNA REPLICATION REPLICATION IN PROKARYOTES REPLICATION IN EUKARYOTES For IIT-JAM, JNU, GATE, NET, NIMCET and Other Entrance Exams 1-C-8, Sheela Chowdhary Road,
JNU CEEB SAMPLE THEORY REPLICATION TYPE OF DNA REPLICATION REPLICATION IN PROKARYOTES REPLICATION IN EUKARYOTES For IIT-JAM, JNU, GATE, NET, NIMCET and Other Entrance Exams 1-C-8, Sheela Chowdhary Road,
DNA Replication in Prokaryotes and Eukaryotes
 DNA Replication in Prokaryotes and Eukaryotes 1. Overall mechanism 2. Roles of Polymerases & other proteins 3. More mechanism: Initiation and Termination 4. Mitochondrial DNA replication DNA replication
DNA Replication in Prokaryotes and Eukaryotes 1. Overall mechanism 2. Roles of Polymerases & other proteins 3. More mechanism: Initiation and Termination 4. Mitochondrial DNA replication DNA replication
CGTAAGAGTACGTCCAGCATCGGF ATCATTATCTACATCXXXXXTACCATTCATTCAGATCTCACTATCGCATTCTCATGCAGGTCGTAGCCXS
 5 CGTAAGAGTACGTCCAGCATCGGF ATCATTATCTACATCXXXXXTACCATTCATTCAGATCTCACTATCGCATTCTCATGCAGGTCGTAGCCXS 29 Supplementary Figure 1 Sequence of the DNA primer/template pair used in Figure 1bc. A 23nt primer is
5 CGTAAGAGTACGTCCAGCATCGGF ATCATTATCTACATCXXXXXTACCATTCATTCAGATCTCACTATCGCATTCTCATGCAGGTCGTAGCCXS 29 Supplementary Figure 1 Sequence of the DNA primer/template pair used in Figure 1bc. A 23nt primer is
Welcome to Class 18! Lecture 18: Outline and Objectives. Replication is semiconservative! Replication: DNA DNA! Introductory Biochemistry!
 Lecture 18: Outline and Objectives Welcome to Class 18! Introductory Biochemistry! l DNA Replication! l DNA polymerase! l the enzymatic reaction! l proofreading and accuracy! l DNA synthesis! l origins
Lecture 18: Outline and Objectives Welcome to Class 18! Introductory Biochemistry! l DNA Replication! l DNA polymerase! l the enzymatic reaction! l proofreading and accuracy! l DNA synthesis! l origins
Fidelity of DNA polymerase
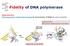 Fidelity of DNA polymerase Shape selectivity: DNA polymerase's conformational change for determination of fidelity for each nucleotide Induced fit: Structure determines function Matched nucleotide Fidelity
Fidelity of DNA polymerase Shape selectivity: DNA polymerase's conformational change for determination of fidelity for each nucleotide Induced fit: Structure determines function Matched nucleotide Fidelity
DNA Structure. DNA: The Genetic Material. Chapter 14
 DNA: The Genetic Material Chapter 14 DNA Structure DNA is a nucleic acid. The building blocks of DNA are nucleotides, each composed of: a 5-carbon sugar called deoxyribose a phosphate group (PO 4 ) a nitrogenous
DNA: The Genetic Material Chapter 14 DNA Structure DNA is a nucleic acid. The building blocks of DNA are nucleotides, each composed of: a 5-carbon sugar called deoxyribose a phosphate group (PO 4 ) a nitrogenous
ZR-96 Oligo Clean & Concentrator Catalog Nos. D4062 & D4063
 INSTRUCTION MANUAL ZR-96 Oligo Clean & Concentrator Catalog Nos. D4062 & D4063 Highlights Quick, high-throughput (96-well) recovery of ultra-pure DNA and RNA oligonucleotides. Complete removal of dyes,
INSTRUCTION MANUAL ZR-96 Oligo Clean & Concentrator Catalog Nos. D4062 & D4063 Highlights Quick, high-throughput (96-well) recovery of ultra-pure DNA and RNA oligonucleotides. Complete removal of dyes,
Biochemical analyses indicate that binding and cleavage specificities define the ordered processing of human Okazaki fragments by Dna2 and FEN1
 6774 6786 Nucleic Acids Research, 2012, Vol. 40, No. 14 Published online 7 May 2012 doi:10.1093/nar/gks388 Biochemical analyses indicate that binding and cleavage specificities define the ordered processing
6774 6786 Nucleic Acids Research, 2012, Vol. 40, No. 14 Published online 7 May 2012 doi:10.1093/nar/gks388 Biochemical analyses indicate that binding and cleavage specificities define the ordered processing
Molecular Cell Biology - Problem Drill 11: Recombinant DNA
 Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
Chapter 3. Enzyme manipulation of DNA and RNA
 Chapter 3 Enzyme manipulation of DNA and RNA To measure incorporation of radioactivity (to see if the probe is good or not for hybridization) Acid precipitation method: - Add sonicated salmon sperm DNA
Chapter 3 Enzyme manipulation of DNA and RNA To measure incorporation of radioactivity (to see if the probe is good or not for hybridization) Acid precipitation method: - Add sonicated salmon sperm DNA
Genetic Information: DNA replication
 Genetic Information: DNA replication Umut Fahrioglu, PhD MSc DNA Replication Replication of DNA is vital to the transmission of genomes and the genes they contain from one cell generation to the other.
Genetic Information: DNA replication Umut Fahrioglu, PhD MSc DNA Replication Replication of DNA is vital to the transmission of genomes and the genes they contain from one cell generation to the other.
Molecular Biology (2)
 Molecular Biology (2) DNA replication Mamoun Ahram, PhD Second semester, 2018-2019 Resources This lecture Cooper, pp. 191-207 2 Some basic information The entire DNA content of the cell is known as genome.
Molecular Biology (2) DNA replication Mamoun Ahram, PhD Second semester, 2018-2019 Resources This lecture Cooper, pp. 191-207 2 Some basic information The entire DNA content of the cell is known as genome.
For Research Use Only Ver
 INSTRUCTION MANUAL Oligo Clean & Concentrator Catalog Nos. D4060 & D4061 Highlights Quick (2 minute) recovery of ultra-pure DNA and RNA oligonucleotides. Complete removal of dyes, salts, enzymes, nucleotides,
INSTRUCTION MANUAL Oligo Clean & Concentrator Catalog Nos. D4060 & D4061 Highlights Quick (2 minute) recovery of ultra-pure DNA and RNA oligonucleotides. Complete removal of dyes, salts, enzymes, nucleotides,
SUPPLEMENTARY INFORMATION
 doi:1.138/nature1895 i Core histones CAF-1 v ii Reb1 CAF-1 Fen1 vi Mature Okazaki fragment iii PCNA Reb1 iv vii Mature Okazaki fragment Reb1 Mature Okazaki fragments Figure S1. Model for nucleosome-mediated
doi:1.138/nature1895 i Core histones CAF-1 v ii Reb1 CAF-1 Fen1 vi Mature Okazaki fragment iii PCNA Reb1 iv vii Mature Okazaki fragment Reb1 Mature Okazaki fragments Figure S1. Model for nucleosome-mediated
CircLigase ssdna Ligase
 Cat. Nos. CL4111K and CL4115K Connect with Epicentre on our blog (epicentral.blogspot.com), Facebook (facebook.com/epicentrebio), and Twitter (@EpicentreBio). www.epicentre.com Lit. # 222 10/2012 1 EPILIT222
Cat. Nos. CL4111K and CL4115K Connect with Epicentre on our blog (epicentral.blogspot.com), Facebook (facebook.com/epicentrebio), and Twitter (@EpicentreBio). www.epicentre.com Lit. # 222 10/2012 1 EPILIT222
Supplementary Information for
 Supplementary Information for Conformational landscapes of DNA polymerase I and mutator derivatives establish fidelity checkpoints for nucleotide insertion Hohlbein et al. 1 Supplementary Figure S1. Wt
Supplementary Information for Conformational landscapes of DNA polymerase I and mutator derivatives establish fidelity checkpoints for nucleotide insertion Hohlbein et al. 1 Supplementary Figure S1. Wt
DNA Metabolism. I. DNA Replication. A. Template concept: 1. How can you make a copy of a molecule? 2. Complementary Hydrogen bonding
 DNA Metabolism I. DNA Replication A. Template concept: 1. How can you make a copy of a molecule? 2. Complementary Hydrogen bonding B. DNA replication follows a set of fundamental rules 1. Semiconservative
DNA Metabolism I. DNA Replication A. Template concept: 1. How can you make a copy of a molecule? 2. Complementary Hydrogen bonding B. DNA replication follows a set of fundamental rules 1. Semiconservative
DNA replication. - proteins for initiation of replication; - proteins for polymerization of nucleotides.
 DNA replication Replication represents the duplication of the genetic information encoded in DNA that is the crucial step in the reproduction of living organisms and the growth of multicellular organisms.
DNA replication Replication represents the duplication of the genetic information encoded in DNA that is the crucial step in the reproduction of living organisms and the growth of multicellular organisms.
3-Input Majority Logic Gate and Multiple Input Logic Circuit Based on DNA Strand Displacement
 Supporting Information for 3-Input Majority Logic Gate and Multiple Input Logic Circuit Based on DNA Strand Displacement Wei Li, Yang Yang, Hao Yan, Yan Liu Department of Chemistry and Biochemistry and
Supporting Information for 3-Input Majority Logic Gate and Multiple Input Logic Circuit Based on DNA Strand Displacement Wei Li, Yang Yang, Hao Yan, Yan Liu Department of Chemistry and Biochemistry and
III. Detailed Examination of the Mechanism of Replication A. Initiation B. Priming C. Elongation D. Proofreading and Termination
 Outline for Replication I. General Features of Replication A. Semi-Conservative B. Starts at Origin C. Bidirectional D. Semi-Discontinuous II. Proteins and Enzymes of Replication III. Detailed Examination
Outline for Replication I. General Features of Replication A. Semi-Conservative B. Starts at Origin C. Bidirectional D. Semi-Discontinuous II. Proteins and Enzymes of Replication III. Detailed Examination
University of Groningen. Single-molecule studies of DNA replication Geertsema, Hylkje
 University of Groningen Single-molecule studies of DNA replication Geertsema, Hylkje IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's PDF) if you wish to cite from it. Please
University of Groningen Single-molecule studies of DNA replication Geertsema, Hylkje IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's PDF) if you wish to cite from it. Please
University of Groningen. Single-molecule studies of DNA replication Geertsema, Hylkje
 University of Groningen Single-molecule studies of DNA replication Geertsema, Hylkje IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's PDF) if you wish to cite from it. Please
University of Groningen Single-molecule studies of DNA replication Geertsema, Hylkje IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's PDF) if you wish to cite from it. Please
Tutorial Week #9 Page 1 of 11
 Tutorial Week #9 Page 1 of 11 Tutorial Week #9 DNA Replication Before the tutorial: Read ECB Chapter 6 p195-207, and review your lecture notes Read this tutorial and create a table of definitions and functions
Tutorial Week #9 Page 1 of 11 Tutorial Week #9 DNA Replication Before the tutorial: Read ECB Chapter 6 p195-207, and review your lecture notes Read this tutorial and create a table of definitions and functions
Bi 8 Lecture 4. Ellen Rothenberg 14 January Reading: from Alberts Ch. 8
 Bi 8 Lecture 4 DNA approaches: How we know what we know Ellen Rothenberg 14 January 2016 Reading: from Alberts Ch. 8 Central concept: DNA or RNA polymer length as an identifying feature RNA has intrinsically
Bi 8 Lecture 4 DNA approaches: How we know what we know Ellen Rothenberg 14 January 2016 Reading: from Alberts Ch. 8 Central concept: DNA or RNA polymer length as an identifying feature RNA has intrinsically
Design. Construction. Characterization
 Design Construction Characterization DNA mrna (messenger) A C C transcription translation C A C protein His A T G C T A C G Plasmids replicon copy number incompatibility selection marker origin of replication
Design Construction Characterization DNA mrna (messenger) A C C transcription translation C A C protein His A T G C T A C G Plasmids replicon copy number incompatibility selection marker origin of replication
Supplemental Materials and Methods
 Supplemental Materials and Methods Proteins and reagents Proteins were purified as described previously: RecA, RecQ, and SSB proteins (Harmon and Kowalczykowski 1998); RecF protein (Morimatsu and Kowalczykowski
Supplemental Materials and Methods Proteins and reagents Proteins were purified as described previously: RecA, RecQ, and SSB proteins (Harmon and Kowalczykowski 1998); RecF protein (Morimatsu and Kowalczykowski
REPLICATION AND POLYMERASE CHAIN REACTION (PCR)
 REPLICATION AND POLYMERASE CHAIN REACTION (PCR) Debbie S. Retnoningrum School of Pharmacy, ITB References: 1. Glick, BR and JJ Pasternak, 2003, pages 27 28; 110 120 2. Groves MJ, 2006, pages 40 44 3. Brown
REPLICATION AND POLYMERASE CHAIN REACTION (PCR) Debbie S. Retnoningrum School of Pharmacy, ITB References: 1. Glick, BR and JJ Pasternak, 2003, pages 27 28; 110 120 2. Groves MJ, 2006, pages 40 44 3. Brown
Certificate of Analysis. Quality Control Parameters. 2 -deoxyadenosine-5 -triphosphate. MW = g /mol
 COA No: CA BN-0006 datp 100mM Storage Conditions: Lot number: -20 C DA-516110 For Research Use Only Expiry date: November 2018 Quality Control Parameters 2 -deoxyadenosine-5 -triphosphate C10H12N5O12P3Li4
COA No: CA BN-0006 datp 100mM Storage Conditions: Lot number: -20 C DA-516110 For Research Use Only Expiry date: November 2018 Quality Control Parameters 2 -deoxyadenosine-5 -triphosphate C10H12N5O12P3Li4
DNA Replication I Biochemistry 302. Bob Kelm January 24, 2005
 DNA Replication I Biochemistry 302 Bob Kelm January 24, 2005 Watson Crick prediction: Each stand of parent DNA serves as a template for synthesis of a new complementary daughter strand Fig. 4.12 Proof
DNA Replication I Biochemistry 302 Bob Kelm January 24, 2005 Watson Crick prediction: Each stand of parent DNA serves as a template for synthesis of a new complementary daughter strand Fig. 4.12 Proof
Recitation CHAPTER 9 DNA Technologies
 Recitation CHAPTER 9 DNA Technologies DNA Cloning: General Scheme A cloning vector and eukaryotic chromosomes are separately cleaved with the same restriction endonuclease. (A single chromosome is shown
Recitation CHAPTER 9 DNA Technologies DNA Cloning: General Scheme A cloning vector and eukaryotic chromosomes are separately cleaved with the same restriction endonuclease. (A single chromosome is shown
4. Analysing genes II Isolate mutants*
 .. 4. Analysing s II Isolate mutants* Using the mutant to isolate the classify mutants by complementation analysis wild type study phenotype of mutants mutant 1 - use mutant to isolate sequence put individual
.. 4. Analysing s II Isolate mutants* Using the mutant to isolate the classify mutants by complementation analysis wild type study phenotype of mutants mutant 1 - use mutant to isolate sequence put individual
Methods (detailed). for all the experiments. Cells were grown in 50 ml YEA. The cells were harvested and
 Methods (detailed). Purification of yeast chromosomal DNA. Strain JZ105 (mat1m Δmat2,3::LEU2, ade6-210, leu1-32, ura4-d18, his2) was used for all the experiments. Cells were grown in 50 ml YEA. The cells
Methods (detailed). Purification of yeast chromosomal DNA. Strain JZ105 (mat1m Δmat2,3::LEU2, ade6-210, leu1-32, ura4-d18, his2) was used for all the experiments. Cells were grown in 50 ml YEA. The cells
#FD µl (for 200 rxns) Expiry Date: Description. 1 ml of 10X FastDigest Green Buffer. Store at -20 C
 PRODUCT INFORMATION Thermo Scientific FastDigest SalI #FD0644 Lot: 5'...G T C G A C...3' 3'...C A G C T G...5' Supplied with: Store at -20 C 200 µl (for 200 rxns) Expiry Date: BSA included www.thermoscientific.com/onebio
PRODUCT INFORMATION Thermo Scientific FastDigest SalI #FD0644 Lot: 5'...G T C G A C...3' 3'...C A G C T G...5' Supplied with: Store at -20 C 200 µl (for 200 rxns) Expiry Date: BSA included www.thermoscientific.com/onebio
N173A. residue number
 a 0.3 Δω 1 ( 1 H Ν ) [ppm] 0-0.3 N173A -0.6 Y177A b Δω 3 ( 13 C α ) [ppm] 2 0-2 c Δω 3 ( 13 C β ) [ppm] 2 0-2 -4 Supplementary Figure 1 140 150 160 170 180 190 200 residue number Comparison of chemical
a 0.3 Δω 1 ( 1 H Ν ) [ppm] 0-0.3 N173A -0.6 Y177A b Δω 3 ( 13 C α ) [ppm] 2 0-2 c Δω 3 ( 13 C β ) [ppm] 2 0-2 -4 Supplementary Figure 1 140 150 160 170 180 190 200 residue number Comparison of chemical
The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity
 Promega Notes Magazine Number 62, 1997, p. 02 The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity By Christine Andrews and Scott Lesley Promega
Promega Notes Magazine Number 62, 1997, p. 02 The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity By Christine Andrews and Scott Lesley Promega
DNA: Structure & Replication
 DNA Form & Function DNA: Structure & Replication Understanding DNA replication and the resulting transmission of genetic information from cell to cell, and generation to generation lays the groundwork
DNA Form & Function DNA: Structure & Replication Understanding DNA replication and the resulting transmission of genetic information from cell to cell, and generation to generation lays the groundwork
BIO 311C Spring Lecture 34 Friday 23 Apr.
 BIO 311C Spring 2010 1 Lecture 34 Friday 23 Apr. Summary of DNA Replication in Prokaryotes origin of replication initial double helix origin of replication new growing polynucleotide chains Circular molecule
BIO 311C Spring 2010 1 Lecture 34 Friday 23 Apr. Summary of DNA Replication in Prokaryotes origin of replication initial double helix origin of replication new growing polynucleotide chains Circular molecule
DNA. Chapter 1. Molecular Diagnostics Fundamentals, Methods and Clinical Applications Second Edition 1/29/2013. Copyright 2012 F.A.
 DNA Chapter 1 1 Deoxyribonucleic acid (DNA) is a genetic information storage system. A T G C T A C G DNA is a polymer of nucleotides. Nucleotides are phosphorylated nucleosides. DNA Nucleosides comprise
DNA Chapter 1 1 Deoxyribonucleic acid (DNA) is a genetic information storage system. A T G C T A C G DNA is a polymer of nucleotides. Nucleotides are phosphorylated nucleosides. DNA Nucleosides comprise
Chapter 4. Recombinant DNA Technology
 Chapter 4 Recombinant DNA Technology 5. Plasmid Cloning Vectors Plasmid Plasmids Self replicating Double-stranded Mostly circular DNA ( 500 kb) Linear : Streptomyces, Borrelia burgdorferi Replicon
Chapter 4 Recombinant DNA Technology 5. Plasmid Cloning Vectors Plasmid Plasmids Self replicating Double-stranded Mostly circular DNA ( 500 kb) Linear : Streptomyces, Borrelia burgdorferi Replicon
Storage and Expression of Genetic Information
 Storage and Expression of Genetic Information 29. DNA structure, Replication and Repair ->Ch 25. DNA metabolism 30. RNA Structure, Synthesis and Processing ->Ch 26. RNA metabolism 31. Protein Synthesis
Storage and Expression of Genetic Information 29. DNA structure, Replication and Repair ->Ch 25. DNA metabolism 30. RNA Structure, Synthesis and Processing ->Ch 26. RNA metabolism 31. Protein Synthesis
DNA Replication and Repair
 DN Replication and Repair http://hyperphysics.phy-astr.gsu.edu/hbase/organic/imgorg/cendog.gif DN Replication genetic information is passed on to the next generation semi-conservative Parent molecule with
DN Replication and Repair http://hyperphysics.phy-astr.gsu.edu/hbase/organic/imgorg/cendog.gif DN Replication genetic information is passed on to the next generation semi-conservative Parent molecule with
Supplementary Information. Arrays of Individual DNA Molecules on Nanopatterned Substrates
 Supplementary Information Arrays of Individual DNA Molecules on Nanopatterned Substrates Roland Hager, Alma Halilovic, Jonathan R. Burns, Friedrich Schäffler, Stefan Howorka S1 Figure S-1. Characterization
Supplementary Information Arrays of Individual DNA Molecules on Nanopatterned Substrates Roland Hager, Alma Halilovic, Jonathan R. Burns, Friedrich Schäffler, Stefan Howorka S1 Figure S-1. Characterization
ZR-96 Genomic DNA Clean & Concentrator -5 Catalog Nos. D4066 & D4067
 INSTRUCTION MANUAL ZR-96 Genomic DNA Clean & Concentrator -5 Catalog Nos. D4066 & D4067 Highlights 96-well plate recovery of large-sized DNA (e.g., genomic, mitochondrial, plasmid (BAC/PAC), viral, phage,
INSTRUCTION MANUAL ZR-96 Genomic DNA Clean & Concentrator -5 Catalog Nos. D4066 & D4067 Highlights 96-well plate recovery of large-sized DNA (e.g., genomic, mitochondrial, plasmid (BAC/PAC), viral, phage,
The flow of Genetic information
 The flow of Genetic information http://highered.mcgrawhill.com/sites/0072507470/student_view0/chapter3/animation dna_replication quiz_1_.html 1 DNA Replication DNA is a double-helical molecule Watson and
The flow of Genetic information http://highered.mcgrawhill.com/sites/0072507470/student_view0/chapter3/animation dna_replication quiz_1_.html 1 DNA Replication DNA is a double-helical molecule Watson and
Supplemental Information. Single-Molecule Imaging Reveals How. Mre11-Rad50-Nbs1 Initiates DNA Break Repair
 Molecular Cell, Volume 67 Supplemental Information Single-Molecule Imaging Reveals How Mre11-Rad50-Nbs1 Initiates DNA Break Repair Logan R. Myler, Ignacio F. Gallardo, Michael M. Soniat, Rajashree A. Deshpande,
Molecular Cell, Volume 67 Supplemental Information Single-Molecule Imaging Reveals How Mre11-Rad50-Nbs1 Initiates DNA Break Repair Logan R. Myler, Ignacio F. Gallardo, Michael M. Soniat, Rajashree A. Deshpande,
Mutagenesis PCR I (Multiple Site Directed Mutagenesis)
 Mutagenesis Mutagenesis PCR I (Multiple Site Directed Mutagenesis) Mixture 25µl total reaction volume : 1. 2.5 µl of 10X Taq lligase buffer (need the NAD for Taq ligase) 2. 0.5 µl 100mM ATP 3. X µl (50-100
Mutagenesis Mutagenesis PCR I (Multiple Site Directed Mutagenesis) Mixture 25µl total reaction volume : 1. 2.5 µl of 10X Taq lligase buffer (need the NAD for Taq ligase) 2. 0.5 µl 100mM ATP 3. X µl (50-100
Table 1. Primers, annealing temperatures, and product sizes for PCR amplification.
 Table 1. Primers, annealing temperatures, and product sizes for PCR amplification. Gene Direction Primer sequence (5 3 ) Annealing Temperature Size (bp) BRCA1 Forward TTGCGGGAGGAAAATGGGTAGTTA 50 o C 292
Table 1. Primers, annealing temperatures, and product sizes for PCR amplification. Gene Direction Primer sequence (5 3 ) Annealing Temperature Size (bp) BRCA1 Forward TTGCGGGAGGAAAATGGGTAGTTA 50 o C 292
DNA REPLICATION. DNA structure. Semiconservative replication. DNA structure. Origin of replication. Replication bubbles and forks.
 DNA REPLICATION 5 4 Phosphate 3 DNA structure Nitrogenous base 1 Deoxyribose 2 Nucleotide DNA strand = DNA polynucleotide 2004 Biology Olympiad Preparation Program 2 2004 Biology Olympiad Preparation Program
DNA REPLICATION 5 4 Phosphate 3 DNA structure Nitrogenous base 1 Deoxyribose 2 Nucleotide DNA strand = DNA polynucleotide 2004 Biology Olympiad Preparation Program 2 2004 Biology Olympiad Preparation Program
Supplementary Figure 1. Analysis of replication rates by Pol. Nature Structural & Molecular Biology: doi: /nsmb.3207
 Supplementary Figure 1 Analysis of replication rates by Pol (a) Electrophoretic Mobility Shift Assay (EMSA) of Pol -DV binding to template-primer DNA (Fried, M. & Crothers, D.M. Equilibria and kinetics
Supplementary Figure 1 Analysis of replication rates by Pol (a) Electrophoretic Mobility Shift Assay (EMSA) of Pol -DV binding to template-primer DNA (Fried, M. & Crothers, D.M. Equilibria and kinetics
Genomic DNA Clean & Concentrator -25 Catalog Nos. D4064 & D4065
 INSTRUCTION MANUAL Genomic DNA Clean & Concentrator -25 Catalog Nos. D4064 & D4065 Highlights Quick (5 minute) spin column recovery of large-sized DNA (e.g., genomic, mitochondrial, plasmid (BAC/PAC),
INSTRUCTION MANUAL Genomic DNA Clean & Concentrator -25 Catalog Nos. D4064 & D4065 Highlights Quick (5 minute) spin column recovery of large-sized DNA (e.g., genomic, mitochondrial, plasmid (BAC/PAC),
Purification and Sequencing of DNA Guides from Prokaryotic Argonaute Daan C. Swarts *, Edze R. Westra, Stan J. J. Brouns and John van der Oost
 Purification and Sequencing of DNA Guides from Prokaryotic Argonaute Daan C. Swarts *, Edze R. Westra, Stan J. J. Brouns and John van der Oost Department of Agrotechnology and Food Sciences, Wageningen
Purification and Sequencing of DNA Guides from Prokaryotic Argonaute Daan C. Swarts *, Edze R. Westra, Stan J. J. Brouns and John van der Oost Department of Agrotechnology and Food Sciences, Wageningen
Fig. 16-7a. 5 end Hydrogen bond 3 end. 1 nm. 3.4 nm nm
 Fig. 16-7a end Hydrogen bond end 1 nm 3.4 nm 0.34 nm (a) Key features of DNA structure end (b) Partial chemical structure end Fig. 16-8 Adenine (A) Thymine (T) Guanine (G) Cytosine (C) Concept 16.2: Many
Fig. 16-7a end Hydrogen bond end 1 nm 3.4 nm 0.34 nm (a) Key features of DNA structure end (b) Partial chemical structure end Fig. 16-8 Adenine (A) Thymine (T) Guanine (G) Cytosine (C) Concept 16.2: Many
Enzymes used in DNA Replication
 Enzymes used in DNA Replication This document holds the enzymes used in DNA replication, their pictorial representation and functioning. DNA polymerase: DNA polymerase is the chief enzyme of DNA replication.
Enzymes used in DNA Replication This document holds the enzymes used in DNA replication, their pictorial representation and functioning. DNA polymerase: DNA polymerase is the chief enzyme of DNA replication.
University of Groningen. Single-molecule studies of DNA replication Geertsema, Hylkje
 University of Groningen Single-molecule studies of DNA replication Geertsema, Hylkje IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's PDF) if you wish to cite from it. Please
University of Groningen Single-molecule studies of DNA replication Geertsema, Hylkje IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's PDF) if you wish to cite from it. Please
Multi-Amplified Sensing of MicroRNA by a Small DNA Fragment-Driven Enzymatic Cascade Reaction
 Supporting Information for Multi-Amplified Sensing of MicroRNA by a Small DNA Fragment-Driven Enzymatic Cascade Reaction Eunjung Kim, Philip D. Howes, Spencer W. Crowder, and Molly M. Stevens* Department
Supporting Information for Multi-Amplified Sensing of MicroRNA by a Small DNA Fragment-Driven Enzymatic Cascade Reaction Eunjung Kim, Philip D. Howes, Spencer W. Crowder, and Molly M. Stevens* Department
Supplementary Information
 Supplementary Information Self-assembled Messenger RNA Nanoparticles (mrna-nps) for Efficient Gene Expression Hyejin Kim 1, Yongkuk Park 1 and Jong Bum Lee 1, * 1 Department of Chemical Engineering, University
Supplementary Information Self-assembled Messenger RNA Nanoparticles (mrna-nps) for Efficient Gene Expression Hyejin Kim 1, Yongkuk Park 1 and Jong Bum Lee 1, * 1 Department of Chemical Engineering, University
DNA Replication in Eukaryotes
 OpenStax-CNX module: m44517 1 DNA Replication in Eukaryotes OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section,
OpenStax-CNX module: m44517 1 DNA Replication in Eukaryotes OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section,
Intercalation-based single-molecule fluorescence assay to study DNA supercoil
 Intercalation-based single-molecule fluorescence assay to study DNA supercoil dynamics Mahipal Ganji, Sung Hyun Kim, Jaco van der Torre, Elio Abbondanzieri*, Cees Dekker* Supplementary information Supplementary
Intercalation-based single-molecule fluorescence assay to study DNA supercoil dynamics Mahipal Ganji, Sung Hyun Kim, Jaco van der Torre, Elio Abbondanzieri*, Cees Dekker* Supplementary information Supplementary
BEST QUALITY HIGHEST PURITY. Recombinant ENZYMES & PROTEINS
 BEST QUALITY HIGHEST PURITY Recombinant ENZYMES & PROTEINS We offer a wide range of highest quality enzymes and proteins for molecular biology including DNA polymerases, reverse transcriptases, DNA ligases,
BEST QUALITY HIGHEST PURITY Recombinant ENZYMES & PROTEINS We offer a wide range of highest quality enzymes and proteins for molecular biology including DNA polymerases, reverse transcriptases, DNA ligases,
The Biotechnology Toolbox
 Chapter 15 The Biotechnology Toolbox Cutting and Pasting DNA Cutting DNA Restriction endonuclease or restriction enzymes Cellular protection mechanism for infected foreign DNA Recognition and cutting specific
Chapter 15 The Biotechnology Toolbox Cutting and Pasting DNA Cutting DNA Restriction endonuclease or restriction enzymes Cellular protection mechanism for infected foreign DNA Recognition and cutting specific
ARUNAI ACADEMY FOR PG TRB-BOTANY DHARMAPURI REPLICATION - ENZYMES.
 ARUNAI ACADEMY FOR PG TRB-BOTANY DHARMAPURI.9500244679 REPLICATION - ENZYMES DNA HELICASE Sparation of two strands- DNA helicase enzyme functions Unwinds DNA. DNA double helix by breaking the hydrogen
ARUNAI ACADEMY FOR PG TRB-BOTANY DHARMAPURI.9500244679 REPLICATION - ENZYMES DNA HELICASE Sparation of two strands- DNA helicase enzyme functions Unwinds DNA. DNA double helix by breaking the hydrogen
Replication of DNA Virus Genomes. Lecture 7 Virology W3310/4310 Spring 2013
 Replication of DNA Virus Genomes Lecture 7 Virology W3310/4310 Spring 2013 It s all about Initiation Problems faced by DNA replication machinery Viruses must replicate their genomes to make new progeny
Replication of DNA Virus Genomes Lecture 7 Virology W3310/4310 Spring 2013 It s all about Initiation Problems faced by DNA replication machinery Viruses must replicate their genomes to make new progeny
Plant Molecular and Cellular Biology Lecture 4: E. coli DNA Replicase Structure & Function. Gary Peter
 Plant Molecular and Cellular Biology Lecture 4: E. coli DNA Replicase Structure & Function Gary Peter Learning Objectives 1. List and explain the mechanisms by which E. coli DNA is replicated 2. Describe
Plant Molecular and Cellular Biology Lecture 4: E. coli DNA Replicase Structure & Function Gary Peter Learning Objectives 1. List and explain the mechanisms by which E. coli DNA is replicated 2. Describe
