Editorial. Current Computational Models for Prediction of the Varied Interactions Related to Non-Coding RNAs
|
|
|
- Jane French
- 5 years ago
- Views:
Transcription
1 Editorial Current Computational Models for Prediction of the Varied Interactions Related to Non-Coding RNAs Xing Chen 1,*, Huiming Peng 2, Zheng Yin 3 1 School of Information and Electrical Engineering, China University of Mining and Technology, Xuzhou, , China 2 Department of Math, Science & Technologies, Forsyth Technical Community College, Winston-Salem, USA 3 Department of Systems Medicine and Bioengineering, Houston Methodist Research Institute, Houston, USA *Corresponding author Correspondence should be addressed to Xing Chen; xingchen@amss.ac.cn
2 Non-coding RNA (ncrna) refers to a kind of endogenous small RNA molecules that have no protein coding capacity. In recent years, extensive studies have been conducted to study the roles of ncrnas in cell biology, and accumulating evidences shows that these RNA molecules do not constitute transcriptional noise but play important roles in cellular functions, such as transcriptional and post-transcriptional regulation, epigenetic regulation, organ or tissue development, cell differentiation, cell cycle control, cellular transport, metabolic processes, chromosome dynamics, and so on. Their deregulation heavily contributes to various pathological conditions of human complex diseases, including breast cancer, hepatocellular cancer, prostate cancer, colon cancer, bladder cancer, thyroid cancer, lung cancer, ovarian cancer, leukemia, Alzheimer s disease, diabetes, and HIV. The development of computational models to predict the various complex ncrna-related interactions could significantly benefit the inference of ncrna function, the identification of ncrna-disease associations, the detection of ncrna biomarker, the identification of drug-target interactions, and drug design. The papers that follow in the next pages describe recent findings in the field of computational models for prediction of the varied interactions related to ncrnas. They represent only a fraction of the current research, the emerging trends, and applications of computational models for ncrnas. The special issue consists of eight research papers and one review paper.
3 The research papers include identification and annotation of ncrnas (four papers), mirna-target interaction prediction (one paper), ncrna function prediction (one paper), mirna-transcriptional factor interaction prediction (one paper), and protein-protein interaction prediction (one paper). A brief description of the papers follows. Guoqiang Wan et al. analyzed RNA-seq data and ChIP-seq histone modification data to identify the relationship between lncrna genes transcription and histone modification H3K4me3 or H3K27me3 in the paper Transcriptional Regulation of lncrna Genes by Histone Modification in Alzheimer s Disease. Jibin Qu et al. detected 254 small noncoding RNAs in genome of Pleurotus ostreatuin and analyzed the evolutionary conservation of them in the paper Identification and Characterization of Small Noncoding RNAs in Genome Sequences of the Edible Fungus Pleurotus ostreatus. A meta-path-based prediction method RMLM was developed in the paper A Meta-Path-Based Prediction Method for Human mirna-target Association by Jiawei Luo et al. for Predicting potential mirna-target interactions. The authors also developed RMLMSe, in which sequence information was utilized to improve the performance of RMLM. Limin Jiang et al. employed backpropagation (BP) neural network together with 98-dimensional novel features for microrna (mirna) precursor identification in the paper BP Neural Network Could Help
4 Improve Pre-miRNA Identification in Various Species. The authors demonstrated that the total prediction accuracy of this method was nearly 13.17% greater than the state-of-the-art mirna precursor prediction software tools. A framework named Gene2Function to annotate Gene Reference into Functions (GeneRIFs) was given by Yang Hu et al. in the paper Annotating the Function of the Human Genome with Gene Ontology and Disease Ontology. M-fold, TargetScan, GeneCoDia3 were used in the paper Human Ribosomal RNA-Derived Resident MicroRNAs as the Transmitter of Information upon the Cytoplasmic Cancer Stress for investigating RNA relationships between rrna and mirna against cellular stresses by Masaru Yoshikawa et al. The authors detected 17 RNA sequences identical with known mirnas in the human rrna and termed as rrna-hosted mirna analogs (rmirnas). The paper Ens-PPI: A Novel Ensemble Classifier for Predicting the Interactions of Proteins Using Autocovariance Transformation from PSSM by Zhen-Guo Gao et al. provided a novel predictor based on the Rotation Forest (RF) algorithm combined with Autocovariance (AC) features extracted from the Position-Specific Scoring Matrix. The method achieved promising prediction performance when implemented on the protein-protein interaction datasets of Yeast, H. pylori and independent datasets.
5 Qi Zhao et al. constructed a computational model of mirna-mediated feed-forward loops (FFLs) in the paper Effect of dynamic interaction between microrna and transcription factor on gene expression. The authors introduced four possible structural topologies of FFLs with two gate functions (AND gate and OR gate) based on the different dynamic interactions between mirna and TF on gene expression. Finally, in the review paper Long non-coding RNA identification: comparing machine learning based tools for long non-coding transcripts discrimination by Siyu Han et al., several popular methods for lncrna identification such as Coding Potential Calculator (CPC), Coding-Potential Assessment Tool (CPAT), Coding-Non-Coding Index (CNCI), Predictor of long non-coding RNAs and messenger RNAs based on an improved k-mer scheme (PLEK), Long non-coding RNA IDentification (LncRNA-ID), lncrscan-svm, lncrna-mfdl and LncRNApred were summarized with their advantages, disadvantages, and application scopes. ACKNOWLEDGMENTS We would like to thank the authors who submitted their work for consideration to this special issue as well as the reviewers for their efforts and constructive criticism. The work of Xing Chen was supported by National Natural Science Foundation of China under Grant No and Xing Chen
6 Huiming Peng Zheng Yin
RNA-SEQUENCING ANALYSIS
 RNA-SEQUENCING ANALYSIS Joseph Powell SISG- 2018 CONTENTS Introduction to RNA sequencing Data structure Analyses Transcript counting Alternative splicing Allele specific expression Discovery APPLICATIONS
RNA-SEQUENCING ANALYSIS Joseph Powell SISG- 2018 CONTENTS Introduction to RNA sequencing Data structure Analyses Transcript counting Alternative splicing Allele specific expression Discovery APPLICATIONS
Functional microrna targets in protein coding sequences. Merve Çakır
 Functional microrna targets in protein coding sequences Martin Reczko, Manolis Maragkakis, Panagiotis Alexiou, Ivo Grosse, Artemis G. Hatzigeorgiou Merve Çakır 27.04.2012 microrna * micrornas are small
Functional microrna targets in protein coding sequences Martin Reczko, Manolis Maragkakis, Panagiotis Alexiou, Ivo Grosse, Artemis G. Hatzigeorgiou Merve Çakır 27.04.2012 microrna * micrornas are small
Data Mining for Biological Data Analysis
 Data Mining for Biological Data Analysis Data Mining and Text Mining (UIC 583 @ Politecnico di Milano) References Data Mining Course by Gregory-Platesky Shapiro available at www.kdnuggets.com Jiawei Han
Data Mining for Biological Data Analysis Data Mining and Text Mining (UIC 583 @ Politecnico di Milano) References Data Mining Course by Gregory-Platesky Shapiro available at www.kdnuggets.com Jiawei Han
Machine Learning. HMM applications in computational biology
 10-601 Machine Learning HMM applications in computational biology Central dogma DNA CCTGAGCCAACTATTGATGAA transcription mrna CCUGAGCCAACUAUUGAUGAA translation Protein PEPTIDE 2 Biological data is rapidly
10-601 Machine Learning HMM applications in computational biology Central dogma DNA CCTGAGCCAACTATTGATGAA transcription mrna CCUGAGCCAACUAUUGAUGAA translation Protein PEPTIDE 2 Biological data is rapidly
A Brief History. Bootstrapping. Bagging. Boosting (Schapire 1989) Adaboost (Schapire 1995)
 A Brief History Bootstrapping Bagging Boosting (Schapire 1989) Adaboost (Schapire 1995) What s So Good About Adaboost Improves classification accuracy Can be used with many different classifiers Commonly
A Brief History Bootstrapping Bagging Boosting (Schapire 1989) Adaboost (Schapire 1995) What s So Good About Adaboost Improves classification accuracy Can be used with many different classifiers Commonly
Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate
 Supplementary Figure Legends Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate BC041951 in gastric cancer. (A) The flow chart for selected candidate lncrnas in 660 up-regulated
Supplementary Figure Legends Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate BC041951 in gastric cancer. (A) The flow chart for selected candidate lncrnas in 660 up-regulated
ChIP-seq analysis 2/28/2018
 ChIP-seq analysis 2/28/2018 Acknowledgements Much of the content of this lecture is from: Furey (2012) ChIP-seq and beyond Park (2009) ChIP-seq advantages + challenges Landt et al. (2012) ChIP-seq guidelines
ChIP-seq analysis 2/28/2018 Acknowledgements Much of the content of this lecture is from: Furey (2012) ChIP-seq and beyond Park (2009) ChIP-seq advantages + challenges Landt et al. (2012) ChIP-seq guidelines
Efficient and Accurate Analysis of non coding RNAs with InSyBio ncrnaseq
 Efficient and Accurate Analysis of non coding RNAs with InSyBio ncrnaseq WHITE PAPER By InSyBio Ltd Aigli Korfiati Computer Engineer, MSc, PhD candidate InSyBio Product Development Manager October 2015
Efficient and Accurate Analysis of non coding RNAs with InSyBio ncrnaseq WHITE PAPER By InSyBio Ltd Aigli Korfiati Computer Engineer, MSc, PhD candidate InSyBio Product Development Manager October 2015
Introduction to human genomics and genome informatics
 Introduction to human genomics and genome informatics Session 1 Prince of Wales Clinical School Dr Jason Wong ARC Future Fellow Head, Bioinformatics & Integrative Genomics Adult Cancer Program, Lowy Cancer
Introduction to human genomics and genome informatics Session 1 Prince of Wales Clinical School Dr Jason Wong ARC Future Fellow Head, Bioinformatics & Integrative Genomics Adult Cancer Program, Lowy Cancer
IPA Advanced Training Course
 IPA Advanced Training Course Academia Sinica 2015 Oct Gene( 陳冠文 ) Supervisor and IPA certified analyst 1 Review for Introductory Training course Searching Building a Pathway Editing a Pathway for Publication
IPA Advanced Training Course Academia Sinica 2015 Oct Gene( 陳冠文 ) Supervisor and IPA certified analyst 1 Review for Introductory Training course Searching Building a Pathway Editing a Pathway for Publication
CS273B: Deep learning for Genomics and Biomedicine
 CS273B: Deep learning for Genomics and Biomedicine Lecture 2: Convolutional neural networks and applications to functional genomics 09/28/2016 Anshul Kundaje, James Zou, Serafim Batzoglou Outline Anatomy
CS273B: Deep learning for Genomics and Biomedicine Lecture 2: Convolutional neural networks and applications to functional genomics 09/28/2016 Anshul Kundaje, James Zou, Serafim Batzoglou Outline Anatomy
Transcription start site classification
 Transcription start site classification Max Libbrecht, Matt Fisher, Roy Frostig, Hrysoula Papadakis, Anshul Kundaje, Serafim Batzoglou December 11, 2009 Abstract Understanding the mechanisms of gene expression
Transcription start site classification Max Libbrecht, Matt Fisher, Roy Frostig, Hrysoula Papadakis, Anshul Kundaje, Serafim Batzoglou December 11, 2009 Abstract Understanding the mechanisms of gene expression
Our website:
 Biomedical Informatics Summer Internship Program (BMI SIP) The Department of Biomedical Informatics hosts an annual internship program each summer which provides high school, undergraduate, and graduate
Biomedical Informatics Summer Internship Program (BMI SIP) The Department of Biomedical Informatics hosts an annual internship program each summer which provides high school, undergraduate, and graduate
Prediction of noncoding RNAs with RNAz
 Prediction of noncoding RNAs with RNAz John Dzmil, III Steve Griesmer Philip Murillo April 4, 2007 What is non-coding RNA (ncrna)? RNA molecules that are not translated into proteins Size range from 20
Prediction of noncoding RNAs with RNAz John Dzmil, III Steve Griesmer Philip Murillo April 4, 2007 What is non-coding RNA (ncrna)? RNA molecules that are not translated into proteins Size range from 20
Bioinformatics & Protein Structural Analysis. Bioinformatics & Protein Structural Analysis. Learning Objective. Proteomics
 The molecular structures of proteins are complex and can be defined at various levels. These structures can also be predicted from their amino-acid sequences. Protein structure prediction is one of the
The molecular structures of proteins are complex and can be defined at various levels. These structures can also be predicted from their amino-acid sequences. Protein structure prediction is one of the
Basics of RNA-Seq. (With a Focus on Application to Single Cell RNA-Seq) Michael Kelly, PhD Team Lead, NCI Single Cell Analysis Facility
 2018 ABRF Meeting Satellite Workshop 4 Bridging the Gap: Isolation to Translation (Single Cell RNA-Seq) Sunday, April 22 Basics of RNA-Seq (With a Focus on Application to Single Cell RNA-Seq) Michael Kelly,
2018 ABRF Meeting Satellite Workshop 4 Bridging the Gap: Isolation to Translation (Single Cell RNA-Seq) Sunday, April 22 Basics of RNA-Seq (With a Focus on Application to Single Cell RNA-Seq) Michael Kelly,
Review Article Long Noncoding RNA Identification: Comparing Machine Learning Based Tools for Long Noncoding Transcripts Discrimination
 BioMed Research International Volume 2016, Article ID 8496165, 14 pages http://dx.doi.org/10.1155/2016/8496165 Review Article Long Noncoding RNA Identification: Comparing Machine Learning Based Tools for
BioMed Research International Volume 2016, Article ID 8496165, 14 pages http://dx.doi.org/10.1155/2016/8496165 Review Article Long Noncoding RNA Identification: Comparing Machine Learning Based Tools for
Title: Genome-Wide Predictions of Transcription Factor Binding Events using Multi- Dimensional Genomic and Epigenomic Features Background
 Title: Genome-Wide Predictions of Transcription Factor Binding Events using Multi- Dimensional Genomic and Epigenomic Features Team members: David Moskowitz and Emily Tsang Background Transcription factors
Title: Genome-Wide Predictions of Transcription Factor Binding Events using Multi- Dimensional Genomic and Epigenomic Features Team members: David Moskowitz and Emily Tsang Background Transcription factors
Introduction to BIOINFORMATICS
 COURSE OF BIOINFORMATICS a.a. 2016-2017 Introduction to BIOINFORMATICS What is Bioinformatics? (I) The sinergy between biology and informatics What is Bioinformatics? (II) From: http://www.bioteach.ubc.ca/bioinfo2010/
COURSE OF BIOINFORMATICS a.a. 2016-2017 Introduction to BIOINFORMATICS What is Bioinformatics? (I) The sinergy between biology and informatics What is Bioinformatics? (II) From: http://www.bioteach.ubc.ca/bioinfo2010/
The Function Transformation Omics Funomics. There are no two identical leaves in the world, so how to find effective
 The Function Transformation Omics Funomics Yongshuai Jiang 1,* Jing Xu 1, Simeng Hu 1, Di Liu 1, Linna Zhao 1, Xu Zhou 1 1.College of Bioinformatics Science and Technology, Harbin Medical University, Harbin,
The Function Transformation Omics Funomics Yongshuai Jiang 1,* Jing Xu 1, Simeng Hu 1, Di Liu 1, Linna Zhao 1, Xu Zhou 1 1.College of Bioinformatics Science and Technology, Harbin Medical University, Harbin,
Decoding Chromatin States with Epigenome Data Advanced Topics in Computa8onal Genomics
 Decoding Chromatin States with Epigenome Data 02-715 Advanced Topics in Computa8onal Genomics HMMs for Decoding Chromatin States Epigene8c modifica8ons of the genome have been associated with Establishing
Decoding Chromatin States with Epigenome Data 02-715 Advanced Topics in Computa8onal Genomics HMMs for Decoding Chromatin States Epigene8c modifica8ons of the genome have been associated with Establishing
Personalized Human Genome Sequencing
 Personalized Human Genome Sequencing Dr. Stefan Platz DABT, Global Head Drug Safety & Metabolism Biomedical research: strengths & limitations of non-animal alternatives 06 December 2016 The Human Genome
Personalized Human Genome Sequencing Dr. Stefan Platz DABT, Global Head Drug Safety & Metabolism Biomedical research: strengths & limitations of non-animal alternatives 06 December 2016 The Human Genome
Computational Genomics
 Computational Genomics http://www.cs.cmu.edu/~02710 Ziv Bar-Joseph zivbj@cs.cmu.edu GHC 8006 Chakra Chennubhotla chakracs@pitt.edu Suite 3064, BST3 Topics Introduction (1 Week) Sequence analysis(4 weeks)
Computational Genomics http://www.cs.cmu.edu/~02710 Ziv Bar-Joseph zivbj@cs.cmu.edu GHC 8006 Chakra Chennubhotla chakracs@pitt.edu Suite 3064, BST3 Topics Introduction (1 Week) Sequence analysis(4 weeks)
11/22/13. Proteomics, functional genomics, and systems biology. Biosciences 741: Genomics Fall, 2013 Week 11
 Proteomics, functional genomics, and systems biology Biosciences 741: Genomics Fall, 2013 Week 11 1 Figure 6.1 The future of genomics Functional Genomics The field of functional genomics represents the
Proteomics, functional genomics, and systems biology Biosciences 741: Genomics Fall, 2013 Week 11 1 Figure 6.1 The future of genomics Functional Genomics The field of functional genomics represents the
Understanding transcriptional regulation by integrative analysis of transcription factor binding data
 Understanding transcriptional regulation by integrative analysis of transcription factor binding data Cheng et al. 2012 Shu Yang Feb. 21, 2013 1 / 26 Introduction 2 / 26 DNA-binding Proteins sequence-specific
Understanding transcriptional regulation by integrative analysis of transcription factor binding data Cheng et al. 2012 Shu Yang Feb. 21, 2013 1 / 26 Introduction 2 / 26 DNA-binding Proteins sequence-specific
RNA genes. Functional non-coding RNAs (ncrna) Jan 31 st 2018.
 RNA genes Functional non-coding RNAs (ncrna) Jan 31 st 2018. After human genome sequencing it became obvious that human genome consists of many non-protein coding genes, genes that code for different RNAs
RNA genes Functional non-coding RNAs (ncrna) Jan 31 st 2018. After human genome sequencing it became obvious that human genome consists of many non-protein coding genes, genes that code for different RNAs
Supplementary Information
 Supplementary Information Integrative genomic analyses reveal clinically relevant long non-coding RNA in cancer Zhou Du 1#, Teng Fei 2,3,4,5#, Roel G.W. Verhaak 6, Zhen Su 7, Yong Zhang 1, Myles Brown
Supplementary Information Integrative genomic analyses reveal clinically relevant long non-coding RNA in cancer Zhou Du 1#, Teng Fei 2,3,4,5#, Roel G.W. Verhaak 6, Zhen Su 7, Yong Zhang 1, Myles Brown
BIOINFORMATICS THE MACHINE LEARNING APPROACH
 88 Proceedings of the 4 th International Conference on Informatics and Information Technology BIOINFORMATICS THE MACHINE LEARNING APPROACH A. Madevska-Bogdanova Inst, Informatics, Fac. Natural Sc. and
88 Proceedings of the 4 th International Conference on Informatics and Information Technology BIOINFORMATICS THE MACHINE LEARNING APPROACH A. Madevska-Bogdanova Inst, Informatics, Fac. Natural Sc. and
Higher Human Biology Unit 1: Human Cells Pupils Learning Outcomes
 Higher Human Biology Unit 1: Human Cells Pupils Learning Outcomes 1.1 Division and Differentiation in Human Cells I can state that cellular differentiation is the process by which a cell develops more
Higher Human Biology Unit 1: Human Cells Pupils Learning Outcomes 1.1 Division and Differentiation in Human Cells I can state that cellular differentiation is the process by which a cell develops more
Matrix Factorization-Based Data Fusion for Drug-Induced Liver Injury Prediction
 Matrix Factorization-Based Data Fusion for Drug-Induced Liver Injury Prediction Marinka Žitnik1 and Blaž Zupan 1,2 1 Faculty of Computer and Information Science, University of Ljubljana, Tržaška 25, 1000
Matrix Factorization-Based Data Fusion for Drug-Induced Liver Injury Prediction Marinka Žitnik1 and Blaž Zupan 1,2 1 Faculty of Computer and Information Science, University of Ljubljana, Tržaška 25, 1000
This place covers: Methods or systems for genetic or protein-related data processing in computational molecular biology.
 G16B BIOINFORMATICS, i.e. INFORMATION AND COMMUNICATION TECHNOLOGY [ICT] SPECIALLY ADAPTED FOR GENETIC OR PROTEIN-RELATED DATA PROCESSING IN COMPUTATIONAL MOLECULAR BIOLOGY Methods or systems for genetic
G16B BIOINFORMATICS, i.e. INFORMATION AND COMMUNICATION TECHNOLOGY [ICT] SPECIALLY ADAPTED FOR GENETIC OR PROTEIN-RELATED DATA PROCESSING IN COMPUTATIONAL MOLECULAR BIOLOGY Methods or systems for genetic
Introduction to Bioinformatics
 Introduction to Bioinformatics If the 19 th century was the century of chemistry and 20 th century was the century of physic, the 21 st century promises to be the century of biology...professor Dr. Satoru
Introduction to Bioinformatics If the 19 th century was the century of chemistry and 20 th century was the century of physic, the 21 st century promises to be the century of biology...professor Dr. Satoru
Epigenetics, Environment and Human Health
 Epigenetics, Environment and Human Health A. Karim Ahmed National Council for Science and the Environment Washington, DC May, 2015 Epigenetics A New Biological Paradigm A Question about Cells: All cells
Epigenetics, Environment and Human Health A. Karim Ahmed National Council for Science and the Environment Washington, DC May, 2015 Epigenetics A New Biological Paradigm A Question about Cells: All cells
Making Deep Learning Understandable for Analyzing Sequen;al Data about Gene Regula;on. Dr. Yanjun Qi 2017/11/26
 Making Deep Learning Understandable for Analyzing Sequen;al Data about Gene Regula;on Dr. Yanjun Qi 2017/11/26 Roadmap ² Background of Machine Learning ² Background of Sequen?al Data about Gene Regula?on
Making Deep Learning Understandable for Analyzing Sequen;al Data about Gene Regula;on Dr. Yanjun Qi 2017/11/26 Roadmap ² Background of Machine Learning ² Background of Sequen?al Data about Gene Regula?on
Algorithms in Nature. (brief) introduction to biology
 Algorithms in Nature (brief) introduction to biology Organism, Organ, Cell Organism 2 Types of Cells Eukaryots: - Plants, animals, humans - DNA resides in the nucleus - Contain also other compartments
Algorithms in Nature (brief) introduction to biology Organism, Organ, Cell Organism 2 Types of Cells Eukaryots: - Plants, animals, humans - DNA resides in the nucleus - Contain also other compartments
Corporate Medical Policy
 Corporate Medical Policy Proteogenomic Testing for Patients with Cancer (GPS Cancer Test) File Name: Origination: Last CAP Review: Next CAP Review: Last Review: proteogenomic_testing_for_patients_with_cancer_gps_cancer_test
Corporate Medical Policy Proteogenomic Testing for Patients with Cancer (GPS Cancer Test) File Name: Origination: Last CAP Review: Next CAP Review: Last Review: proteogenomic_testing_for_patients_with_cancer_gps_cancer_test
Corporate Medical Policy
 Corporate Medical Policy Proteogenomic Testing for Patients with Cancer (GPS Cancer Test) File Name: Origination: Last CAP Review: Next CAP Review: Last Review: proteogenomic_testing_for_patients_with_cancer_gps_cancer_test
Corporate Medical Policy Proteogenomic Testing for Patients with Cancer (GPS Cancer Test) File Name: Origination: Last CAP Review: Next CAP Review: Last Review: proteogenomic_testing_for_patients_with_cancer_gps_cancer_test
2. Materials and Methods
 Identification of cancer-relevant Variations in a Novel Human Genome Sequence Robert Bruggner, Amir Ghazvinian 1, & Lekan Wang 1 CS229 Final Report, Fall 2009 1. Introduction Cancer affects people of all
Identification of cancer-relevant Variations in a Novel Human Genome Sequence Robert Bruggner, Amir Ghazvinian 1, & Lekan Wang 1 CS229 Final Report, Fall 2009 1. Introduction Cancer affects people of all
Polaris and Callisto: Advances in Cellular Biology. Jordan Moore Senior Field Applications Specialist
 Polaris and Callisto: Advances in Cellular Biology Jordan Moore Senior Field Applications Specialist Single-Cell Biology Number of cells The population average is a lie average Transcripts per cell Gene
Polaris and Callisto: Advances in Cellular Biology Jordan Moore Senior Field Applications Specialist Single-Cell Biology Number of cells The population average is a lie average Transcripts per cell Gene
What Can the Epigenome Teach Us About Cellular States and Diseases?
 What Can the Epigenome Teach Us About Cellular States and Diseases? (a computer scientist s view) Luca Pinello Outline Epigenetic: the code over the code What can we learn from epigenomic data? Resources
What Can the Epigenome Teach Us About Cellular States and Diseases? (a computer scientist s view) Luca Pinello Outline Epigenetic: the code over the code What can we learn from epigenomic data? Resources
RNA Structure and the Versatility of RNA. Mitesh Shrestha
 RNA Structure and the Versatility of RNA Mitesh Shrestha Ribonucleic Acid (RNA) Nitrogenous Bases (Adenine, Uracil, Guanine, Cytosine) Ribose Sugar Ribonucleic Acid (RNA) Phosphate Group RNA world Hypothesis
RNA Structure and the Versatility of RNA Mitesh Shrestha Ribonucleic Acid (RNA) Nitrogenous Bases (Adenine, Uracil, Guanine, Cytosine) Ribose Sugar Ribonucleic Acid (RNA) Phosphate Group RNA world Hypothesis
ChIP-seq and RNA-seq. Farhat Habib
 ChIP-seq and RNA-seq Farhat Habib fhabib@iiserpune.ac.in Biological Goals Learn how genomes encode the diverse patterns of gene expression that define each cell type and state. Protein-DNA interactions
ChIP-seq and RNA-seq Farhat Habib fhabib@iiserpune.ac.in Biological Goals Learn how genomes encode the diverse patterns of gene expression that define each cell type and state. Protein-DNA interactions
DNAFSMiner: A Web-Based Software Toolbox to Recognize Two Types of Functional Sites in DNA Sequences
 DNAFSMiner: A Web-Based Software Toolbox to Recognize Two Types of Functional Sites in DNA Sequences Huiqing Liu Hao Han Jinyan Li Limsoon Wong Institute for Infocomm Research, 21 Heng Mui Keng Terrace,
DNAFSMiner: A Web-Based Software Toolbox to Recognize Two Types of Functional Sites in DNA Sequences Huiqing Liu Hao Han Jinyan Li Limsoon Wong Institute for Infocomm Research, 21 Heng Mui Keng Terrace,
Learning Methods for DNA Binding in Computational Biology
 Learning Methods for DNA Binding in Computational Biology Mark Kon Dustin Holloway Yue Fan Chaitanya Sai Charles DeLisi Boston University IJCNN Orlando August 16, 2007 Outline Background on Transcription
Learning Methods for DNA Binding in Computational Biology Mark Kon Dustin Holloway Yue Fan Chaitanya Sai Charles DeLisi Boston University IJCNN Orlando August 16, 2007 Outline Background on Transcription
A Supervised Learning Approach to the Prediction of Hi-C Data
 A Supervised Learning Approach to the Prediction of Hi-C Data Tyler Derr YUE Lab The Department of Biochemistry & Molecular Biology Institute for Personalized Medicine Pennsylvania State University College
A Supervised Learning Approach to the Prediction of Hi-C Data Tyler Derr YUE Lab The Department of Biochemistry & Molecular Biology Institute for Personalized Medicine Pennsylvania State University College
Feature Selection of Gene Expression Data for Cancer Classification: A Review
 Available online at www.sciencedirect.com ScienceDirect Procedia Computer Science 50 (2015 ) 52 57 2nd International Symposium on Big Data and Cloud Computing (ISBCC 15) Feature Selection of Gene Expression
Available online at www.sciencedirect.com ScienceDirect Procedia Computer Science 50 (2015 ) 52 57 2nd International Symposium on Big Data and Cloud Computing (ISBCC 15) Feature Selection of Gene Expression
Smart India Hackathon
 TM Persistent and Hackathons Smart India Hackathon 2017 i4c www.i4c.co.in Digital Transformation 25% of India between age of 16-25 Our country needs audacious digital transformation to reach its potential
TM Persistent and Hackathons Smart India Hackathon 2017 i4c www.i4c.co.in Digital Transformation 25% of India between age of 16-25 Our country needs audacious digital transformation to reach its potential
Corporate Medical Policy
 Corporate Medical Policy Proteogenomic Testing for Patients with Cancer (GPS Cancer Test) File Name: Origination: Last CAP Review: Next CAP Review: Last Review: proteogenomic_testing_for_patients_with_cancer_gps_cancer_test
Corporate Medical Policy Proteogenomic Testing for Patients with Cancer (GPS Cancer Test) File Name: Origination: Last CAP Review: Next CAP Review: Last Review: proteogenomic_testing_for_patients_with_cancer_gps_cancer_test
A STUDY ON STATISTICAL BASED FEATURE SELECTION METHODS FOR CLASSIFICATION OF GENE MICROARRAY DATASET
 A STUDY ON STATISTICAL BASED FEATURE SELECTION METHODS FOR CLASSIFICATION OF GENE MICROARRAY DATASET 1 J.JEYACHIDRA, M.PUNITHAVALLI, 1 Research Scholar, Department of Computer Science and Applications,
A STUDY ON STATISTICAL BASED FEATURE SELECTION METHODS FOR CLASSIFICATION OF GENE MICROARRAY DATASET 1 J.JEYACHIDRA, M.PUNITHAVALLI, 1 Research Scholar, Department of Computer Science and Applications,
Comparison of Commercial Transfection Reagents: Cell line optimized transfection kits for in vitro cancer research.
 Comparison of Commercial Transfection Reagents: Cell line optimized transfection kits for in vitro cancer research. by Altogen Labs, 11200 Manchaca Road, Suite 203 Austin TX 78748 USA Tel. (512) 433-6177
Comparison of Commercial Transfection Reagents: Cell line optimized transfection kits for in vitro cancer research. by Altogen Labs, 11200 Manchaca Road, Suite 203 Austin TX 78748 USA Tel. (512) 433-6177
Mapping strategies for sequence reads
 Mapping strategies for sequence reads Ernest Turro University of Cambridge 21 Oct 2013 Quantification A basic aim in genomics is working out the contents of a biological sample. 1. What distinct elements
Mapping strategies for sequence reads Ernest Turro University of Cambridge 21 Oct 2013 Quantification A basic aim in genomics is working out the contents of a biological sample. 1. What distinct elements
Introduction to Microarray Data Analysis and Gene Networks. Alvis Brazma European Bioinformatics Institute
 Introduction to Microarray Data Analysis and Gene Networks Alvis Brazma European Bioinformatics Institute A brief outline of this course What is gene expression, why it s important Microarrays and how
Introduction to Microarray Data Analysis and Gene Networks Alvis Brazma European Bioinformatics Institute A brief outline of this course What is gene expression, why it s important Microarrays and how
NGS to address ncrna and viruses
 NGS to address ncrna and viruses Introduction & TRON Next generation sequencing transcriptomics ncrnas vrna June 30, 2010 John Castle Institute for Translational Oncology and Immunology (TRON) Mainz, Germany
NGS to address ncrna and viruses Introduction & TRON Next generation sequencing transcriptomics ncrnas vrna June 30, 2010 John Castle Institute for Translational Oncology and Immunology (TRON) Mainz, Germany
Genome-Wide Survey of MicroRNA - Transcription Factor Feed-Forward Regulatory Circuits in Human. Supporting Information
 Genome-Wide Survey of MicroRNA - Transcription Factor Feed-Forward Regulatory Circuits in Human Angela Re #, Davide Corá #, Daniela Taverna and Michele Caselle # equal contribution * corresponding author,
Genome-Wide Survey of MicroRNA - Transcription Factor Feed-Forward Regulatory Circuits in Human Angela Re #, Davide Corá #, Daniela Taverna and Michele Caselle # equal contribution * corresponding author,
A New Algorithm for Protein-Protein Interaction Prediction
 A New Algorithm for Protein-Protein Interaction Prediction Yiwei Li The University of Western Ontario Supervisor: Dr. Lucian Ilie Overview 1 Background 2 Previous approaches Experimental approaches Computational
A New Algorithm for Protein-Protein Interaction Prediction Yiwei Li The University of Western Ontario Supervisor: Dr. Lucian Ilie Overview 1 Background 2 Previous approaches Experimental approaches Computational
Bioinformatics. Outline of lecture
 Bioinformatics Uma Chandran, MSIS, PhD Department of Biomedical Informatics University of Pittsburgh chandran@pitt.edu 412 648 9326 07/08/2014 Outline of lecture What is Bioinformatics? Examples of bioinformatics
Bioinformatics Uma Chandran, MSIS, PhD Department of Biomedical Informatics University of Pittsburgh chandran@pitt.edu 412 648 9326 07/08/2014 Outline of lecture What is Bioinformatics? Examples of bioinformatics
DOUBLE-STRAND DNA BREAK PREDICTION USING EPIGENOME MARKS AT KILOBASE RESOLUTION
 DOUBLE-STRAND DNA BREAK PREDICTION USING EPIGENOME MARKS AT KILOBASE RESOLUTION Raphaël MOURAD, Assist. Prof. Centre de Biologie Intégrative Université Paul Sabatier, Toulouse III INTRODUCTION Double-strand
DOUBLE-STRAND DNA BREAK PREDICTION USING EPIGENOME MARKS AT KILOBASE RESOLUTION Raphaël MOURAD, Assist. Prof. Centre de Biologie Intégrative Université Paul Sabatier, Toulouse III INTRODUCTION Double-strand
The Major Function Of Rna Is To Carry Out The Genetic Instructions For Protein Synthesis
 The Major Function Of Rna Is To Carry Out The Genetic Instructions For Protein Synthesis For example, protein synthesis in human mitochondria relies on a genetic code that Leder and Nirenberg were able
The Major Function Of Rna Is To Carry Out The Genetic Instructions For Protein Synthesis For example, protein synthesis in human mitochondria relies on a genetic code that Leder and Nirenberg were able
Introduction to genome biology
 Introduction to genome biology Lisa Stubbs Deep transcritpomes for traditional model species from ENCODE (and modencode) Deep RNA-seq and chromatin analysis on 147 human cell types, as well as tissues,
Introduction to genome biology Lisa Stubbs Deep transcritpomes for traditional model species from ENCODE (and modencode) Deep RNA-seq and chromatin analysis on 147 human cell types, as well as tissues,
2/19/13. Contents. Applications of HMMs in Epigenomics
 2/19/13 I529: Machine Learning in Bioinformatics (Spring 2013) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Background:
2/19/13 I529: Machine Learning in Bioinformatics (Spring 2013) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Background:
Il trascrittoma dei mammiferi
 29 Novembre 2005 Il trascrittoma dei mammiferi dott. Manuela Gariboldi Gruppo di ricerca IFOM: Genetica molecolare dei tumori (responsabile dott. Paolo Radice) Copyright 2005 IFOM Fondazione Istituto FIRC
29 Novembre 2005 Il trascrittoma dei mammiferi dott. Manuela Gariboldi Gruppo di ricerca IFOM: Genetica molecolare dei tumori (responsabile dott. Paolo Radice) Copyright 2005 IFOM Fondazione Istituto FIRC
Satellite Education Workshop (SW4): Epigenomics: Design, Implementation and Analysis for RNA-seq and Methyl-seq Experiments
 Satellite Education Workshop (SW4): Epigenomics: Design, Implementation and Analysis for RNA-seq and Methyl-seq Experiments Saturday March 17, 2012 Orlando, Florida Workshop Description: This full day
Satellite Education Workshop (SW4): Epigenomics: Design, Implementation and Analysis for RNA-seq and Methyl-seq Experiments Saturday March 17, 2012 Orlando, Florida Workshop Description: This full day
Transcription factor binding site prediction in vivo using DNA sequence and shape features
 Transcription factor binding site prediction in vivo using DNA sequence and shape features Anthony Mathelier, Lin Yang, Tsu-Pei Chiu, Remo Rohs, and Wyeth Wasserman anthony.mathelier@gmail.com @AMathelier
Transcription factor binding site prediction in vivo using DNA sequence and shape features Anthony Mathelier, Lin Yang, Tsu-Pei Chiu, Remo Rohs, and Wyeth Wasserman anthony.mathelier@gmail.com @AMathelier
GATCGTGCACGATCTCGGCAATTCGGGATGCCGGCTCGTCACCGGTCGCT
 Problem. (pts) A. (5pts) Your colleague professor Eugene Mathew Lateed generated a genome-wide DNA methylation map for normal colon cells using MRE-seq and MeDIP-seq. In an intergenic region, he found
Problem. (pts) A. (5pts) Your colleague professor Eugene Mathew Lateed generated a genome-wide DNA methylation map for normal colon cells using MRE-seq and MeDIP-seq. In an intergenic region, he found
Control of Eukaryotic Gene Expression (Learning Objectives)
 Control of Eukaryotic Gene Expression (Learning Objectives) 1. Compare and contrast chromatin and chromosome: composition, proteins involved and level of packing. Explain the structure and function of
Control of Eukaryotic Gene Expression (Learning Objectives) 1. Compare and contrast chromatin and chromosome: composition, proteins involved and level of packing. Explain the structure and function of
Genetics and Bioinformatics
 Genetics and Bioinformatics Kristel Van Steen, PhD 2 Montefiore Institute - Systems and Modeling GIGA - Bioinformatics ULg kristel.vansteen@ulg.ac.be Lecture 1: Setting the pace 1 Bioinformatics what s
Genetics and Bioinformatics Kristel Van Steen, PhD 2 Montefiore Institute - Systems and Modeling GIGA - Bioinformatics ULg kristel.vansteen@ulg.ac.be Lecture 1: Setting the pace 1 Bioinformatics what s
Applications of HMMs in Epigenomics
 I529: Machine Learning in Bioinformatics (Spring 2013) Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Contents Background:
I529: Machine Learning in Bioinformatics (Spring 2013) Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Contents Background:
Random matrix analysis for gene co-expression experiments in cancer cells
 Random matrix analysis for gene co-expression experiments in cancer cells OIST-iTHES-CTSR 2016 July 9 th, 2016 Ayumi KIKKAWA (MTPU, OIST) Introduction : What is co-expression of genes? There are 20~30k
Random matrix analysis for gene co-expression experiments in cancer cells OIST-iTHES-CTSR 2016 July 9 th, 2016 Ayumi KIKKAWA (MTPU, OIST) Introduction : What is co-expression of genes? There are 20~30k
Characterizing DNA binding sites high throughput approaches Biol4230 Tues, April 24, 2018 Bill Pearson Pinn 6-057
 Characterizing DNA binding sites high throughput approaches Biol4230 Tues, April 24, 2018 Bill Pearson wrp@virginia.edu 4-2818 Pinn 6-057 Reviewing sites: affinity and specificity representation binding
Characterizing DNA binding sites high throughput approaches Biol4230 Tues, April 24, 2018 Bill Pearson wrp@virginia.edu 4-2818 Pinn 6-057 Reviewing sites: affinity and specificity representation binding
Advanced Algorithms and Models for Computational Biology
 10-810 Advanced Algorithms and Models for Computational Biology Ziv Bar-Joseph zivbj@cs.cmu.edu WeH 4107 Eric Xing epxing@cs.cmu.edu WeH 4127 http://www.cs.cmu.edu/~epxing/class/10810-06/ Topics Introduction
10-810 Advanced Algorithms and Models for Computational Biology Ziv Bar-Joseph zivbj@cs.cmu.edu WeH 4107 Eric Xing epxing@cs.cmu.edu WeH 4127 http://www.cs.cmu.edu/~epxing/class/10810-06/ Topics Introduction
Ingenuity Pathway Analysis (IPA )
 Ingenuity Pathway Analysis (IPA ) For the analysis and interpretation of omics data IPA is a web-based software application for the analysis, integration, and interpretation of data derived from omics
Ingenuity Pathway Analysis (IPA ) For the analysis and interpretation of omics data IPA is a web-based software application for the analysis, integration, and interpretation of data derived from omics
AIT - Austrian Institute of Technology
 BIOMARKER DISCOVERY, BIOINFORMATICS, AND BIOSENSOR DEVELOPMENT Technology Experience AIT Austrian Institute of Technology Low-Emission Transport AIT - Austrian Institute of Technology Energy Health & Bioresources
BIOMARKER DISCOVERY, BIOINFORMATICS, AND BIOSENSOR DEVELOPMENT Technology Experience AIT Austrian Institute of Technology Low-Emission Transport AIT - Austrian Institute of Technology Energy Health & Bioresources
I. Gene Expression Figure 1: Central Dogma of Molecular Biology
 I. Gene Expression Figure 1: Central Dogma of Molecular Biology Central Dogma: Gene Expression: RNA Structure RNA nucleotides contain the pentose sugar Ribose instead of deoxyribose. Contain the bases
I. Gene Expression Figure 1: Central Dogma of Molecular Biology Central Dogma: Gene Expression: RNA Structure RNA nucleotides contain the pentose sugar Ribose instead of deoxyribose. Contain the bases
Gene Selection in Cancer Classification using PSO/SVM and GA/SVM Hybrid Algorithms
 Laboratoire d Informatique Fondamentale de Lille Gene Selection in Cancer Classification using PSO/SVM and GA/SVM Hybrid Algorithms Enrique Alba, José GarcíaNieto, Laetitia Jourdan and ElGhazali Talbi
Laboratoire d Informatique Fondamentale de Lille Gene Selection in Cancer Classification using PSO/SVM and GA/SVM Hybrid Algorithms Enrique Alba, José GarcíaNieto, Laetitia Jourdan and ElGhazali Talbi
Lecture 5: Regulation
 Machine Learning in Computational Biology CSC 2431 Lecture 5: Regulation Instructor: Anna Goldenberg Central Dogma of Biology Transcription DNA RNA protein Process of producing RNA from DNA Constitutive
Machine Learning in Computational Biology CSC 2431 Lecture 5: Regulation Instructor: Anna Goldenberg Central Dogma of Biology Transcription DNA RNA protein Process of producing RNA from DNA Constitutive
Comprehensive assembly of novel transcripts from unmapped human RNA-Seq data and their association with cancer
 Comprehensive assembly of novel transcripts from unmapped human RNA-Seq data and their association with cancer Majid Kazemian, Min Ren, Jian-Xin Lin, Wei Liao, Rosanne Spolski, Warren J. Leonard Corresponding
Comprehensive assembly of novel transcripts from unmapped human RNA-Seq data and their association with cancer Majid Kazemian, Min Ren, Jian-Xin Lin, Wei Liao, Rosanne Spolski, Warren J. Leonard Corresponding
Control of Eukaryotic Genes. AP Biology
 Control of Eukaryotic Genes The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions? Evolution
Control of Eukaryotic Genes The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions? Evolution
Year III Pharm.D Dr. V. Chitra
 Year III Pharm.D Dr. V. Chitra 1 Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Only one strand of DNA serves
Year III Pharm.D Dr. V. Chitra 1 Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Only one strand of DNA serves
Textbook Reading Guidelines
 Understanding Bioinformatics by Marketa Zvelebil and Jeremy Baum Last updated: January 16, 2013 Textbook Reading Guidelines Preface: Read the whole preface, and especially: For the students with Life Science
Understanding Bioinformatics by Marketa Zvelebil and Jeremy Baum Last updated: January 16, 2013 Textbook Reading Guidelines Preface: Read the whole preface, and especially: For the students with Life Science
A Protein Secondary Structure Prediction Method Based on BP Neural Network Ru-xi YIN, Li-zhen LIU*, Wei SONG, Xin-lei ZHAO and Chao DU
 2017 2nd International Conference on Artificial Intelligence: Techniques and Applications (AITA 2017 ISBN: 978-1-60595-491-2 A Protein Secondary Structure Prediction Method Based on BP Neural Network Ru-xi
2017 2nd International Conference on Artificial Intelligence: Techniques and Applications (AITA 2017 ISBN: 978-1-60595-491-2 A Protein Secondary Structure Prediction Method Based on BP Neural Network Ru-xi
CURRICULUM VITAE February Xiao-Ou Zhang
 CURRICULUM VITAE February 2019 Xiao-Ou Zhang Address: University of Massachusetts Medical School 368 Plantation St. Albert Sherman Center Worcester, MA 01605, USA Telephone: +1 (774) 301 5131 Email: XiaoOu.Zhang@umassmed.edu
CURRICULUM VITAE February 2019 Xiao-Ou Zhang Address: University of Massachusetts Medical School 368 Plantation St. Albert Sherman Center Worcester, MA 01605, USA Telephone: +1 (774) 301 5131 Email: XiaoOu.Zhang@umassmed.edu
QIAGEN s NGS Solutions for Biomarkers NGS & Bioinformatics team QIAGEN (Suzhou) Translational Medicine Co.,Ltd
 QIAGEN s NGS Solutions for Biomarkers NGS & Bioinformatics team QIAGEN (Suzhou) Translational Medicine Co.,Ltd 1 Our current NGS & Bioinformatics Platform 2 Our NGS workflow and applications 3 QIAGEN s
QIAGEN s NGS Solutions for Biomarkers NGS & Bioinformatics team QIAGEN (Suzhou) Translational Medicine Co.,Ltd 1 Our current NGS & Bioinformatics Platform 2 Our NGS workflow and applications 3 QIAGEN s
Introduction to Bioinformatics
 Introduction to Bioinformatics Dr. Taysir Hassan Abdel Hamid Lecturer, Information Systems Department Faculty of Computer and Information Assiut University taysirhs@aun.edu.eg taysir_soliman@hotmail.com
Introduction to Bioinformatics Dr. Taysir Hassan Abdel Hamid Lecturer, Information Systems Department Faculty of Computer and Information Assiut University taysirhs@aun.edu.eg taysir_soliman@hotmail.com
Name Class Date. Practice Test
 Name Class Date 12 DNA Practice Test Multiple Choice Write the letter that best answers the question or completes the statement on the line provided. 1. What do bacteriophages infect? a. mice. c. viruses.
Name Class Date 12 DNA Practice Test Multiple Choice Write the letter that best answers the question or completes the statement on the line provided. 1. What do bacteriophages infect? a. mice. c. viruses.
Gene expression DNA RNA. Protein DNA. Replication. Initiation Elongation Processing Export. DNA RNA Protein. Transcription. Degradation.
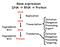 Gene expression DNA RNA Protein DNA DNA Degradation RNA Degradation Protein Replication Transcription Translation Initiation Elongation Processing Export Initiation Elongation Processing Targeting Chapter
Gene expression DNA RNA Protein DNA DNA Degradation RNA Degradation Protein Replication Transcription Translation Initiation Elongation Processing Export Initiation Elongation Processing Targeting Chapter
Molecular Cell Biology - Problem Drill 06: Genes and Chromosomes
 Molecular Cell Biology - Problem Drill 06: Genes and Chromosomes Question No. 1 of 10 1. Which of the following statements about genes is correct? Question #1 (A) Genes carry the information for protein
Molecular Cell Biology - Problem Drill 06: Genes and Chromosomes Question No. 1 of 10 1. Which of the following statements about genes is correct? Question #1 (A) Genes carry the information for protein
Integrating Diverse Datasets Improves Developmental Enhancer Prediction
 Integrating Diverse Datasets Improves Developmental Enhancer Prediction Genevieve D. Erwin 1,2, Nir Oksenberg 2,3, Rebecca M. Truty 1, Dennis Kostka 4, Karl K. Murphy 2,3, Nadav Ahituv 2,3, Katherine S.
Integrating Diverse Datasets Improves Developmental Enhancer Prediction Genevieve D. Erwin 1,2, Nir Oksenberg 2,3, Rebecca M. Truty 1, Dennis Kostka 4, Karl K. Murphy 2,3, Nadav Ahituv 2,3, Katherine S.
resequencing storage SNP ncrna metagenomics private trio de novo exome ncrna RNA DNA bioinformatics RNA-seq comparative genomics
 RNA Sequencing T TM variation genetics validation SNP ncrna metagenomics private trio de novo exome mendelian ChIP-seq RNA DNA bioinformatics custom target high-throughput resequencing storage ncrna comparative
RNA Sequencing T TM variation genetics validation SNP ncrna metagenomics private trio de novo exome mendelian ChIP-seq RNA DNA bioinformatics custom target high-throughput resequencing storage ncrna comparative
Beyond Genomics: Detecting Codes and Signals in the Cellular Transcriptome
 Beyond Genomics: Detecting Codes and Signals in the Cellular Transcriptome Brendan J. Frey University of Toronto Purpose of my talk To identify aspects of bioinformatics in which attendees of ISIT may
Beyond Genomics: Detecting Codes and Signals in the Cellular Transcriptome Brendan J. Frey University of Toronto Purpose of my talk To identify aspects of bioinformatics in which attendees of ISIT may
6.047 / Computational Biology: Genomes, Networks, Evolution Fall 2008
 MIT OpenCourseWare http://ocw.mit.edu 6.047 / 6.878 Computational Biology: Genomes, Networks, Evolution Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu 6.047 / 6.878 Computational Biology: Genomes, Networks, Evolution Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
Genomics and Gene Recognition Genes and Blue Genes
 Genomics and Gene Recognition Genes and Blue Genes November 3, 2004 Eukaryotic Gene Structure eukaryotic genomes are considerably more complex than those of prokaryotes eukaryotic cells have organelles
Genomics and Gene Recognition Genes and Blue Genes November 3, 2004 Eukaryotic Gene Structure eukaryotic genomes are considerably more complex than those of prokaryotes eukaryotic cells have organelles
ChIP-seq and RNA-seq
 ChIP-seq and RNA-seq Biological Goals Learn how genomes encode the diverse patterns of gene expression that define each cell type and state. Protein-DNA interactions (ChIPchromatin immunoprecipitation)
ChIP-seq and RNA-seq Biological Goals Learn how genomes encode the diverse patterns of gene expression that define each cell type and state. Protein-DNA interactions (ChIPchromatin immunoprecipitation)
Gene expression analysis. Biosciences 741: Genomics Fall, 2013 Week 5. Gene expression analysis
 Gene expression analysis Biosciences 741: Genomics Fall, 2013 Week 5 Gene expression analysis From EST clusters to spotted cdna microarrays Long vs. short oligonucleotide microarrays vs. RT-PCR Methods
Gene expression analysis Biosciences 741: Genomics Fall, 2013 Week 5 Gene expression analysis From EST clusters to spotted cdna microarrays Long vs. short oligonucleotide microarrays vs. RT-PCR Methods
RNA-Sequencing analysis
 RNA-Sequencing analysis Markus Kreuz 25. 04. 2012 Institut für Medizinische Informatik, Statistik und Epidemiologie Content: Biological background Overview transcriptomics RNA-Seq RNA-Seq technology Challenges
RNA-Sequencing analysis Markus Kreuz 25. 04. 2012 Institut für Medizinische Informatik, Statistik und Epidemiologie Content: Biological background Overview transcriptomics RNA-Seq RNA-Seq technology Challenges
ChIP-seq data analysis with Chipster. Eija Korpelainen CSC IT Center for Science, Finland
 ChIP-seq data analysis with Chipster Eija Korpelainen CSC IT Center for Science, Finland chipster@csc.fi What will I learn? Short introduction to ChIP-seq Analyzing ChIP-seq data Central concepts Analysis
ChIP-seq data analysis with Chipster Eija Korpelainen CSC IT Center for Science, Finland chipster@csc.fi What will I learn? Short introduction to ChIP-seq Analyzing ChIP-seq data Central concepts Analysis
earray 5.0 Create your own Custom Microarray Design
 earray 5.0 Create your own Custom Microarray Design http://earray.chem.agilent.com earray 5.x Overview Session Summary Session Summary Agilent Genomics Microarray Solution earray Functional Overview Gene
earray 5.0 Create your own Custom Microarray Design http://earray.chem.agilent.com earray 5.x Overview Session Summary Session Summary Agilent Genomics Microarray Solution earray Functional Overview Gene
Textbook Reading Guidelines
 Understanding Bioinformatics by Marketa Zvelebil and Jeremy Baum Last updated: May 1, 2009 Textbook Reading Guidelines Preface: Read the whole preface, and especially: For the students with Life Science
Understanding Bioinformatics by Marketa Zvelebil and Jeremy Baum Last updated: May 1, 2009 Textbook Reading Guidelines Preface: Read the whole preface, and especially: For the students with Life Science
Value Correct Answer Feedback. Student Response. A. Dicer enzyme. complex. C. the Dicer-RISC complex D. none of the above
 1 RNA mediated interference is a post-transcriptional gene silencing mechanism Which component of the RNAi pathway have been implicated in cleavage of the target mrna? A Dicer enzyme B the RISC-siRNA complex
1 RNA mediated interference is a post-transcriptional gene silencing mechanism Which component of the RNAi pathway have been implicated in cleavage of the target mrna? A Dicer enzyme B the RISC-siRNA complex
Reading Between the Genes: Computational Models to Discover Function from Noncoding DNA
 Reading Between the Genes: Computational Models to Discover Function from Noncoding DNA Yves A. Lussier, Joanne Berghout, Francesca Vitali, Kenneth S. Ramos Center for Biomedical Informatics and Biostatistics,
Reading Between the Genes: Computational Models to Discover Function from Noncoding DNA Yves A. Lussier, Joanne Berghout, Francesca Vitali, Kenneth S. Ramos Center for Biomedical Informatics and Biostatistics,
