Efficient and Accurate Analysis of non coding RNAs with InSyBio ncrnaseq
|
|
|
- Audra Moody
- 6 years ago
- Views:
Transcription
1 Efficient and Accurate Analysis of non coding RNAs with InSyBio ncrnaseq WHITE PAPER By InSyBio Ltd Aigli Korfiati Computer Engineer, MSc, PhD candidate InSyBio Product Development Manager October 2015 InSyBio ncrnaseq v2.0
2 INTRODUCTION This document is intended for researchers (molecular biologists, bioinformaticians, biostatisticians, etc...) working in academia or biopharma, biotechnology and health industries who want to analyze non coding RNAs, predict s and analyze and predict target sites. NCRNASEQ IS A NON-CODING RNA ANALYSIS TOOL FOR THE PREDICTION AND ANALYSIS OF non-coding RNAs, and target genes Non-coding RNA genes are RNA sequences transcribed from DNA, but not translated to proteins. Their identification as well as the identification of the genes they regulate is a promising research area. Traditionally, most RNA molecules were regarded as information-carrying intermediates from the gene to the translation machinery, with exceptions being transfer RNA (trna) and ribosomal RNA (rrna), both of which are infrastructural RNAs that are central to the protein synthesis process [1]. However, since the late 1990s, evidence suggests that other types of non-protein-coding RNA molecules are present in organisms ranging from bacteria to mammals, which are involved in a variety of cellular processes including plasmid replication, phage development, bacterial virulence, chromosome structure, DNA transcription, RNA processing and modification, development control, regulation of gene expression and others [2]. In humans, it is estimated that ~98% of the genome can be transcribed, of which only ~2% encodes protein genes, suggesting the possibility that a large percentage of the genome may encode ncrna genes [3]. Many of these RNAs, expressed in particular tissues and/or developmental stages, are associated with particular diseases including various cancers, neurodegenerative, inflammatory and cardiovascular diseases, schizophrenia, ataxia, cartilage-hair hypoplasia, DiGeorge syndrome and autism, and/or are involved in complex genetic phenomena such as imprinting and other forms of epigenetic control of gene expression [4, 5]. Although the vital importance of ncrna genes in cellular activities is well recognized and new members and classes of ncrnas are continuously being reported, our current knowledge about the collection of all ncrna genes encoded in a particular genome is very limited because of the lack of effective computational or experimental methods. Their computational identification represents one of the most important and challenging problems in computational biology. Existing methods for ncrna gene prediction rely mostly on homology information, thus limiting their applications to ncrna genes with known homologues. Evidence linking ncrnas with diseases seem to be attracting the attention of the pharmaceutical and biotechnology industries. Companies and institutions are developing ncrna-based strategies against cancer, cardiovascular, neurological and muscular diseases [5]. Thus, continued investigation of ncrna biogenesis and function will give us a new framework for a more comprehensive understanding of human disease.
3 InSyBio ncrnaseq enables users to analyze non-coding RNAs. Users can search and analyze the RNA sequence of their interest. They can also analyze a full sequences dataset derived from online available databases, experimental sequencing techniques or computational in silico techniques. With InSyBio ncrnaseq you can predict and analyze RNA genes and target genes combining a variety of sequential, structural and functional information, and using a high performance machine learning technique. The RNA analysis is conducted by the calculation of the 58 most informative features described in the literature, and the - targets analysis is conducted by the calculation of the 124 most informative ones. InSyBio ncrnaseq also provides results storage in its knowledge base, equipped with information retrieval tools, to allow users produce and extract their own datasets. INSYBIO NCRNASEQ KEY PROPERTIES: Non coding RNA analysis with the calculation of 58 sequential, thermodynamical and structural properties of the RNA sequence and -target site analysis with the calculation of 124 sequential, thermodynamical, structural and motif properties of the :mrna pair. MiRNAs and -target sites predictions with high accuracy. Integrated information on stem-loop and mature s from online available databases. Association of proteins to s participating to their regulatory mechanism. Batch computations of many sequences at once are allowed. No need for personal super computers to perform difficult computing tasks: they are now executed in our cloud infrastructure with minimum burden on the user s pc. WITH INSYBIO NCRNASEQ YOU CAN: a) Calculate 58 RNA genes-related features b) Predict s c) Calculate 124 target features d) Predict targets e) View integrated information about the of your interest f) Search stem-loop and mature s
4 INSYBIO NCRNASEQ SCENARIO: MIRNA PREDICTION WHITE PAPER In this scenario we have a set of sequences and want to predict which of them are pres and discriminate them from other RNA stem-loops or other RNAs which do not have the stem-loop structure. For this case study we constructed a dataset with 30 known pre-s, 15 snornas and 30 pseudo hairpins which look like pre-s but come from coding regions and thus we are sure that they are not real pre-s. STEPS 1. We created a text file and wrote the sequences in fasta format in a file editor. Since both lines of the fasta format (sequence description and sequence) are obligatory, we wrote >random_sequence_from_cds_x as the sequence description for the pseudo hairpins. 2. We then uploaded the file to ncrnaseq through Data Store selecting the category ncrna sequences. 3. The file type was ncrna sequences by default. So we just wrote a file title describing our file, selected the file prepared in step 1 and uploaded it by clicking the respective button. 4. After the file was automatically verified, we clicked the List all ncrna sequences button. 5. In the action selection dropdown list we selected Prediction at ncrnaseq. 6. We were redirected to the prediction page and we clicked the Start Calculation button. 7. The sequences were processed one by one and we could see in the resulting table or download in a tab-delimited file: their fasta format, their prediction ( (green) / pseudo (red) / other (blue)), their prediction score and 58 computed features. These features include sequential, thermodynamical and structural properties of the RNA sequence. The prediction score showed the strength/confidence of the prediction. Prediction scores close to 0 indicate weak predictions, while prediction scores far from 0 strong.
5 DISCUSSION WHITE PAPER Experimenting with a dataset with 30 known pre-s, 15 snornas and 30 pseudo hairpins which look like pre-s but come from coding regions led to the results summarized in the following table. total s other pseudo s pre-s snornas pseudo hairpins from coding regions From the total of 30 pre-s, they were all indeed predicted as s. Their prediction scores were also very high ranging from 0.71 to 1.27, values very far from the decision boundary of 0, indicating the confidence of the prediction. One of the pres prediction score was 0.24, but again this value is not close to the decision boundary. From the total of 15 snornas, 13 were indeed predicted as other, while 2 were predicted as pseudo s. Their prediction scores were and The criteria for a sequence to be predicted as other are to have a) a minimum free energy more than -15 kcal/mol, or b) less than 17 base pairs. The two snornas predicted as pseudo s had a minimum free energy approximately -17 kcal/mol and 19 and 20 base pairs, respectively. This fact indicates that these specific snornas structure is extremely close to the hairpin s structure and this is the reason they have been classified to the pseudo-hairpin class. The 30 pseudo hairpins from coding regions were all predicted as pseudo s. Their prediction scores were very far from the decision boundary of 0 ranging from to , values, indicating the confidence of the prediction. One pseudo hairpin prediction score was -0.45, but again this value is quite far from the decision boundary.
6 INSYBIO NCRNASEQ SCENARIO: MIRNA TARGET PREDICTION In this scenario we have a set of s and mrnas and want to predict the mrna target sites of the s. InSyBio ncrnaseq takes as input s and potential mrna target sites (not full mrnas) and predicts which of them share a targeting relation. For this case study we want to predict the target sites of all s for the human mrna genes: CASC2, FECH, GKAP1 and TPR. PREPROCESSING STEPS 1. We firstly had to run a target site prediction program, like miranda [6] giving it as input a list of all human s from mirbase [7] and the sequences of the mrna genes. 2. Or, we could have directly downloaded from miranda ( the human target site predictions for the mrna genes: CASC2, FECH, GKAP1 and TPR. NCRNASEQ STEPS 3. We created two text files and wrote in a file editor the sequences in the one and the predicted mrna target sites from step 1 in the other, both in fasta format. Note that the first was tested against the first target site, the second against the second target site, etc. Since both lines of the fasta format (sequence description and sequence) are obligatory, we wrote the mrna gene symbol and the transcript id as a target site sequence description and the name and accession id as the sequence description. 4. We then uploaded the files to ncrnaseq through Data Store. We visited the Data Store dashboard and selected the category sequences for the s and mrna sequences for the mrna target sites. 5. We selected the corresponding file type (/mrna sequences), wrote the corresponding file titles and selected the files prepared in step 3. We uploaded them by clicking the respective button. 6. After the files were automatically verified, we clicked the List all /mrna sequences button. 7. In the action selection dropdown list we selected Target Prediction at ncrnaseq. 8. We were redirected to the target prediction page, where we also had to select an mrna/ file. Then we clicked the Start Calculation button. 9. The sequences were processed pair by pair and we could see in the resulting table or download in a tab-delimited file: the examined fasta format, the examined mrna target site fasta format, their prediction (Target (green) / no Target (red)), their prediction score and 124 computed features. These features include sequential, thermodynamical, structural and motif properties of the -target site pairs. The fifth column named Protein links the given to an Interact protein, indicating that the specific regulates the
7 expression of the respective protein. The link is supported if the :mrna pair shares a targeting relation and the mrna is related to an InSyBio Interact protein. Clicking on the protein, we were redirected to the Interact tool which presents all the existing information in the Interact database for the specific protein. The prediction score shows the strength/confidence of the prediction. Prediction scores close to 0 indicate weak predictions, while prediction scores far from 0 strong. DISCUSSION In this scenario we downloaded from miranda the human target site predictions for the mrna genes: CASC2, FECH, GKAP1 and TPR. In miranda downloads section the target site predictions are separated in four categories based on mirsvr, a down-regulation score at the mrna or protein levels and the conservation of the :mrna relationship across species: good mirsvr, good mirsvr non-, non-good mirsvr, non-good mirsvr non-. This resulted in a dataset with 1805 entries as described in the following table: good mirsvr good mirsvr non-good mirsvr non-good mirsvr sum non- non
8 We then created the respective files needed as input in ncrnaseq and performing the target prediction task we came up with the following results: good mirsvr good mirsvr non non-good mirsvr non-good mirsvr non- total samples predicted targets predicted non targets accuracy mean prediction score score standard deviation A total of target site pairs were predicted as true targets, forming a percentage over 94%. The ncrnaseq prediction score ranges from 0.58 to 0.64 for the first 3 categories: 1) good mirsvr, 2) good mirsvr non and 3) non-good mirsvr and is 0.5 for the category non-good mirsvr non-. We additionally performed random permutations of the -target site pairs for the same mrna gene and giving again the respective files as input in ncrnaseq and performing the target prediction task we came up with the following results: good mirsvr good mirsvr non non-good mirsvr non-good mirsvr non- total samples predicted targets predicted non targets accuracy
9 mean prediction score score standard deviation As expected, only 107 out of the 1805 samples were predicted as true targets indicating the specificity of ncrnaseq. The prediction score ranges from to -0.88, indicating the high confidence of the prediction results. To prove the performance of ncrnaseq we investigated two more datasets. The first one is a dataset with 462 experimentally verified target sites downloaded from TarBase version 5c [8] and mirecords release April 2013 [9] databases. The second one is a dataset with 1848 pseudo target sites generated giving as input to miranda artificial s with A, C, G, U frequencies that are not consistent with real s. They are not target sites, but they resemble much to -target site pairs. experimentally target sites verified pseudo target sites samples predicted targets predicted non targets accuracy mean prediction score score standard deviation From the results, we observe that only 59 experimentally verified target sites were predicted as non-targets and none pseudo target site was predicted as true target, thus yielding a total accuracy of over 97.4% and a geometric mean of sensitivity and specificity of over 93.3%. The input datasets of all experiments can be found in InSyBio evaluation and demo versions as pre-uploaded files.
10 REFERENCES [1] Eddy, S. R. (2001). Non coding RNA genes and the modern RNA world. Nature Reviews Genetics, 2(12), [2] Liu, C., Bai, B., Skogerbø, G., Cai, L., Deng, W., Zhang, Y.,... & Chen, R. (2005). NONCODE: an integrated knowledge database of non-coding RNAs. Nucleic acids research, 33(suppl 1), D112-D115. [3] Tran, T. T., Zhou, F., Marshburn, S., Stead, M., Kushner, S. R., & Xu, Y. (2009). De novo computational prediction of non-coding RNA genes in prokaryotic genomes. Bioinformatics, 25(22), [4] Pang, K. C., Stephen, S., Engström, P. G., Tajul-Arifin, K., Chen, W., Wahlestedt, C.,... & Mattick, J. S. (2005). RNAdb a comprehensive mammalian noncoding RNA database. Nucleic acids research, 33(suppl 1), D125-D130. [5] Esteller, M. (2011). Non-coding RNAs in human disease. Nature Reviews Genetics, 12(12), [6] mirsvr predicted target site scoring method: Comprehensive modeling of microrna targets predicts functional non- and non-canonical sites. Betel D, Koppal A, Agius P, Sander C, Leslie C., Genome Biology :R90 [7] mirbase: annotating high confidence micrornas using deep sequencing data. Kozomara A, Griffiths-Jones S. NAR :D68-D73 [8] Papadopoulos, G. L., Reczko, M., Simossis, V. A., Sethupathy, P., &Hatzigeorgiou, A. G. (2009). The database of experimentally supported targets: a functional update of TarBase. Nucleic acids research, 37(suppl 1), D155-D158. [9] Xiao, F., Zuo, Z., Cai, G., Kang, S., Gao, X., & Li, T. (2009). mirecords: an integrated resource for microrna target interactions. Nucleic acids research, 37(suppl 1), D105-D110.
11 ABOUT US InSyBio Ltd is a bioinformatics pioneer company ( in personalized healthcare, that focuses on developing computational frameworks and tools for the analysis of complex life-science and biological data in order to develop predictive integrated biomarkers (biomarkers of various categories) with increased prognostic and diagnostic aspects for the personalized Healthcare Industry. InSyBio Suite consists of tools for providing integrated biological information from various sources, while at the same time it is empowered with robust, user-friendly and installation-free bioinformatics tools based on intelligent algorithms and methods. HOW TO GET INSYBIO NCRNASEQ? A demo version of InSyBio ncrnaseq is freely available at To request a free one month full (evaluation) version of InSyBio ncrnaseq please us at info@insybio.com. To purchase InSyBio ncrnaseq commercial version 1.0 please contact us at sales@insybio.com. COPYRIGHT NOTICE External Publication of InSyBio Ltd - Any InSyBio information that is to be used in advertising, press releases, or promotional materials requires prior written approval from the InSyBio Ltd. A draft of the proposed document should accompany any such request. InSyBio Ltd reserves the right to deny approval of external usage for any reason. Copyright 2015 InSyBio Ltd. Reproduction without written permission is completely forbidden.
RNA Structure and the Versatility of RNA. Mitesh Shrestha
 RNA Structure and the Versatility of RNA Mitesh Shrestha Ribonucleic Acid (RNA) Nitrogenous Bases (Adenine, Uracil, Guanine, Cytosine) Ribose Sugar Ribonucleic Acid (RNA) Phosphate Group RNA world Hypothesis
RNA Structure and the Versatility of RNA Mitesh Shrestha Ribonucleic Acid (RNA) Nitrogenous Bases (Adenine, Uracil, Guanine, Cytosine) Ribose Sugar Ribonucleic Acid (RNA) Phosphate Group RNA world Hypothesis
90 Algorithms in Bioinformatics I, WS 06, ZBIT, D. Huson, December 4, 2006
 90 Algorithms in Bioinformatics I, WS 06, ZBIT, D. Huson, December 4, 2006 8 RNA Secondary Structure Sources for this lecture: R. Durbin, S. Eddy, A. Krogh und G. Mitchison. Biological sequence analysis,
90 Algorithms in Bioinformatics I, WS 06, ZBIT, D. Huson, December 4, 2006 8 RNA Secondary Structure Sources for this lecture: R. Durbin, S. Eddy, A. Krogh und G. Mitchison. Biological sequence analysis,
RNA secondary structure prediction and analysis
 RNA secondary structure prediction and analysis 1 Resources Lecture Notes from previous years: Takis Benos Covariance algorithm: Eddy and Durbin, Nucleic Acids Research, v22: 11, 2079 Useful lecture slides
RNA secondary structure prediction and analysis 1 Resources Lecture Notes from previous years: Takis Benos Covariance algorithm: Eddy and Durbin, Nucleic Acids Research, v22: 11, 2079 Useful lecture slides
Themes: RNA and RNA Processing. Messenger RNA (mrna) What is a gene? RNA is very versatile! RNA-RNA interactions are very important!
 Themes: RNA is very versatile! RNA and RNA Processing Chapter 14 RNA-RNA interactions are very important! Prokaryotes and Eukaryotes have many important differences. Messenger RNA (mrna) Carries genetic
Themes: RNA is very versatile! RNA and RNA Processing Chapter 14 RNA-RNA interactions are very important! Prokaryotes and Eukaryotes have many important differences. Messenger RNA (mrna) Carries genetic
CHAPTER 21 LECTURE SLIDES
 CHAPTER 21 LECTURE SLIDES Prepared by Brenda Leady University of Toledo To run the animations you must be in Slideshow View. Use the buttons on the animation to play, pause, and turn audio/text on or off.
CHAPTER 21 LECTURE SLIDES Prepared by Brenda Leady University of Toledo To run the animations you must be in Slideshow View. Use the buttons on the animation to play, pause, and turn audio/text on or off.
AP Biology Gene Expression/Biotechnology REVIEW
 AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
Computational analysis of non-coding RNA. Andrew Uzilov BME110 Tue, Nov 16, 2010
 Computational analysis of non-coding RNA Andrew Uzilov auzilov@ucsc.edu BME110 Tue, Nov 16, 2010 1 Corrected/updated talk slides are here: http://tinyurl.com/uzilovrna redirects to: http://users.soe.ucsc.edu/~auzilov/bme110/fall2010/
Computational analysis of non-coding RNA Andrew Uzilov auzilov@ucsc.edu BME110 Tue, Nov 16, 2010 1 Corrected/updated talk slides are here: http://tinyurl.com/uzilovrna redirects to: http://users.soe.ucsc.edu/~auzilov/bme110/fall2010/
Name Class Date. Practice Test
 Name Class Date 12 DNA Practice Test Multiple Choice Write the letter that best answers the question or completes the statement on the line provided. 1. What do bacteriophages infect? a. mice. c. viruses.
Name Class Date 12 DNA Practice Test Multiple Choice Write the letter that best answers the question or completes the statement on the line provided. 1. What do bacteriophages infect? a. mice. c. viruses.
DNA Transcription. Dr Aliwaini
 DNA Transcription 1 DNA Transcription-Introduction The synthesis of an RNA molecule from DNA is called Transcription. All eukaryotic cells have five major classes of RNA: ribosomal RNA (rrna), messenger
DNA Transcription 1 DNA Transcription-Introduction The synthesis of an RNA molecule from DNA is called Transcription. All eukaryotic cells have five major classes of RNA: ribosomal RNA (rrna), messenger
Gene Expression: Transcription
 Gene Expression: Transcription The majority of genes are expressed as the proteins they encode. The process occurs in two steps: Transcription = DNA RNA Translation = RNA protein Taken together, they make
Gene Expression: Transcription The majority of genes are expressed as the proteins they encode. The process occurs in two steps: Transcription = DNA RNA Translation = RNA protein Taken together, they make
Protein Synthesis Transcription And Translation Lab Answers
 Lab Answers Free PDF ebook Download: Lab Answers Download or Read Online ebook protein synthesis transcription and translation lab answers in PDF Format From The Best User Guide Database 1.. Anatomy and
Lab Answers Free PDF ebook Download: Lab Answers Download or Read Online ebook protein synthesis transcription and translation lab answers in PDF Format From The Best User Guide Database 1.. Anatomy and
ChroMoS Guide (version 1.2)
 ChroMoS Guide (version 1.2) Background Genome-wide association studies (GWAS) reveal increasing number of disease-associated SNPs. Since majority of these SNPs are located in intergenic and intronic regions
ChroMoS Guide (version 1.2) Background Genome-wide association studies (GWAS) reveal increasing number of disease-associated SNPs. Since majority of these SNPs are located in intergenic and intronic regions
Chapter 13 - Concept Mapping
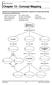 Chapter 13 - Concept Mapping Using the terms and phrases provided below, complete the concept map showing the discovery of DNA structure. amount of base pairs five-carbon sugar purine DNA polymerases Franklin
Chapter 13 - Concept Mapping Using the terms and phrases provided below, complete the concept map showing the discovery of DNA structure. amount of base pairs five-carbon sugar purine DNA polymerases Franklin
RNA-Sequencing analysis
 RNA-Sequencing analysis Markus Kreuz 25. 04. 2012 Institut für Medizinische Informatik, Statistik und Epidemiologie Content: Biological background Overview transcriptomics RNA-Seq RNA-Seq technology Challenges
RNA-Sequencing analysis Markus Kreuz 25. 04. 2012 Institut für Medizinische Informatik, Statistik und Epidemiologie Content: Biological background Overview transcriptomics RNA-Seq RNA-Seq technology Challenges
DNA and RNA. Chapter 12
 DNA and RNA Chapter 12 History of DNA Late 1800 s scientists discovered that DNA is in the nucleus of the cell 1902 Walter Sutton proposed that hereditary material resided in the chromosomes in the nucleus
DNA and RNA Chapter 12 History of DNA Late 1800 s scientists discovered that DNA is in the nucleus of the cell 1902 Walter Sutton proposed that hereditary material resided in the chromosomes in the nucleus
Modeling of Protein Production Process by Finite Automata (FA)
 Modeling of Protein Production Process by Finite Automata (FA) ISSAC BARJIS 1, JOE W. YEOL 2, YEONG SOON RYU 3 Physics and Biomedical Sciences 1, Mechanical Engineering Technology 3 City University of
Modeling of Protein Production Process by Finite Automata (FA) ISSAC BARJIS 1, JOE W. YEOL 2, YEONG SOON RYU 3 Physics and Biomedical Sciences 1, Mechanical Engineering Technology 3 City University of
RNA Genomics II. BME 110: CompBio Tools Todd Lowe & Andrew Uzilov May 17, 2011
 RNA Genomics II BME 110: CompBio Tools Todd Lowe & Andrew Uzilov May 17, 2011 1 TIME Why RNA? An evolutionary perspective The RNA World hypotheses: life arose as self-replicating non-coding RNA (ncrna)
RNA Genomics II BME 110: CompBio Tools Todd Lowe & Andrew Uzilov May 17, 2011 1 TIME Why RNA? An evolutionary perspective The RNA World hypotheses: life arose as self-replicating non-coding RNA (ncrna)
Protein Synthesis: Transcription and Translation
 Protein Synthesis: Transcription and Translation Proteins In living things, proteins are in charge of the expression of our traits (hair/eye color, ability to make insulin, predisposition for cancer, etc.)
Protein Synthesis: Transcription and Translation Proteins In living things, proteins are in charge of the expression of our traits (hair/eye color, ability to make insulin, predisposition for cancer, etc.)
Name 10 Molecular Biology of the Gene Test Date Study Guide You must know: The structure of DNA. The major steps to replication.
 Name 10 Molecular Biology of the Gene Test Date Study Guide You must know: The structure of DNA. The major steps to replication. The difference between replication, transcription, and translation. How
Name 10 Molecular Biology of the Gene Test Date Study Guide You must know: The structure of DNA. The major steps to replication. The difference between replication, transcription, and translation. How
2. Outline the levels of DNA packing in the eukaryotic nucleus below next to the diagram provided.
 AP Biology Reading Packet 6- Molecular Genetics Part 2 Name Chapter 19: Eukaryotic Genomes 1. Define the following terms: a. Euchromatin b. Heterochromatin c. Nucleosome 2. Outline the levels of DNA packing
AP Biology Reading Packet 6- Molecular Genetics Part 2 Name Chapter 19: Eukaryotic Genomes 1. Define the following terms: a. Euchromatin b. Heterochromatin c. Nucleosome 2. Outline the levels of DNA packing
Introduction to Microarray Data Analysis and Gene Networks. Alvis Brazma European Bioinformatics Institute
 Introduction to Microarray Data Analysis and Gene Networks Alvis Brazma European Bioinformatics Institute A brief outline of this course What is gene expression, why it s important Microarrays and how
Introduction to Microarray Data Analysis and Gene Networks Alvis Brazma European Bioinformatics Institute A brief outline of this course What is gene expression, why it s important Microarrays and how
Lecture 2: Central Dogma of Molecular Biology & Intro to Programming
 Lecture 2: Central Dogma of Molecular Biology & Intro to Programming Central Dogma of Molecular Biology Proteins: workhorse molecules of biological systems Proteins are synthesized from the genetic blueprints
Lecture 2: Central Dogma of Molecular Biology & Intro to Programming Central Dogma of Molecular Biology Proteins: workhorse molecules of biological systems Proteins are synthesized from the genetic blueprints
Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit
 Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
Computational aspects of ncrna research. Mihaela Zavolan Biozentrum, Basel Swiss Institute of Bioinformatics
 Computational aspects of ncrna research Mihaela Zavolan Biozentrum, Basel Swiss Institute of Bioinformatics Computational aspects on ncrna Bacterial ncrnas research Gene discovery Target discovery Discovery
Computational aspects of ncrna research Mihaela Zavolan Biozentrum, Basel Swiss Institute of Bioinformatics Computational aspects on ncrna Bacterial ncrnas research Gene discovery Target discovery Discovery
DNA Transcription. Visualizing Transcription. The Transcription Process
 DNA Transcription By: Suzanne Clancy, Ph.D. 2008 Nature Education Citation: Clancy, S. (2008) DNA transcription. Nature Education 1(1) If DNA is a book, then how is it read? Learn more about the DNA transcription
DNA Transcription By: Suzanne Clancy, Ph.D. 2008 Nature Education Citation: Clancy, S. (2008) DNA transcription. Nature Education 1(1) If DNA is a book, then how is it read? Learn more about the DNA transcription
IPA Advanced Training Course
 IPA Advanced Training Course Academia Sinica 2015 Oct Gene( 陳冠文 ) Supervisor and IPA certified analyst 1 Review for Introductory Training course Searching Building a Pathway Editing a Pathway for Publication
IPA Advanced Training Course Academia Sinica 2015 Oct Gene( 陳冠文 ) Supervisor and IPA certified analyst 1 Review for Introductory Training course Searching Building a Pathway Editing a Pathway for Publication
DNA RNA PROTEIN. Professor Andrea Garrison Biology 11 Illustrations 2010 Pearson Education, Inc. unless otherwise noted
 DNA RNA PROTEIN Professor Andrea Garrison Biology 11 Illustrations 2010 Pearson Education, Inc. unless otherwise noted DNA Molecule of heredity Contains all the genetic info our cells inherit Determines
DNA RNA PROTEIN Professor Andrea Garrison Biology 11 Illustrations 2010 Pearson Education, Inc. unless otherwise noted DNA Molecule of heredity Contains all the genetic info our cells inherit Determines
Bundle 5 Test Review
 Bundle 5 Test Review DNA vs. RNA DNA Replication Gene Mutations- Protein Synthesis 1. Label the different components and complete the complimentary base pairing. What is this molecule called? _Nucleic
Bundle 5 Test Review DNA vs. RNA DNA Replication Gene Mutations- Protein Synthesis 1. Label the different components and complete the complimentary base pairing. What is this molecule called? _Nucleic
Fig Ch 17: From Gene to Protein
 Fig. 17-1 Ch 17: From Gene to Protein Basic Principles of Transcription and Translation RNA is the intermediate between genes and the proteins for which they code Transcription is the synthesis of RNA
Fig. 17-1 Ch 17: From Gene to Protein Basic Principles of Transcription and Translation RNA is the intermediate between genes and the proteins for which they code Transcription is the synthesis of RNA
DNA Structure and Analysis. Chapter 4: Background
 DNA Structure and Analysis Chapter 4: Background Molecular Biology Three main disciplines of biotechnology Biochemistry Genetics Molecular Biology # Biotechnology: A Laboratory Skills Course explorer.bio-rad.com
DNA Structure and Analysis Chapter 4: Background Molecular Biology Three main disciplines of biotechnology Biochemistry Genetics Molecular Biology # Biotechnology: A Laboratory Skills Course explorer.bio-rad.com
Molecular Genetics Student Objectives
 Molecular Genetics Student Objectives Exam 1: Enduring understanding 3.A: Heritable information provides for continuity of life. Essential knowledge 3.A.1: DNA, and in some cases RNA, is the primary source
Molecular Genetics Student Objectives Exam 1: Enduring understanding 3.A: Heritable information provides for continuity of life. Essential knowledge 3.A.1: DNA, and in some cases RNA, is the primary source
Fundamentals of Bioinformatics: computation, biology, computational biology
 Fundamentals of Bioinformatics: computation, biology, computational biology Vasilis J. Promponas Bioinformatics Research Laboratory Department of Biological Sciences University of Cyprus A short self-introduction
Fundamentals of Bioinformatics: computation, biology, computational biology Vasilis J. Promponas Bioinformatics Research Laboratory Department of Biological Sciences University of Cyprus A short self-introduction
M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION
 M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION Chapter 7 Microbial Genetics Lecture prepared by Mindy Miller-Kittrell, University of Tennessee, Knoxville The Structure and Replication
M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION Chapter 7 Microbial Genetics Lecture prepared by Mindy Miller-Kittrell, University of Tennessee, Knoxville The Structure and Replication
Epigenetics, Environment and Human Health
 Epigenetics, Environment and Human Health A. Karim Ahmed National Council for Science and the Environment Washington, DC May, 2015 Epigenetics A New Biological Paradigm A Question about Cells: All cells
Epigenetics, Environment and Human Health A. Karim Ahmed National Council for Science and the Environment Washington, DC May, 2015 Epigenetics A New Biological Paradigm A Question about Cells: All cells
If Dna Has The Instructions For Building Proteins Why Is Mrna Needed
 If Dna Has The Instructions For Building Proteins Why Is Mrna Needed if a strand of DNA has the sequence CGGTATATC, then the complementary each strand of DNA contains the info needed to produce the complementary
If Dna Has The Instructions For Building Proteins Why Is Mrna Needed if a strand of DNA has the sequence CGGTATATC, then the complementary each strand of DNA contains the info needed to produce the complementary
Chapter 12. DNA TRANSCRIPTION and TRANSLATION
 Chapter 12 DNA TRANSCRIPTION and TRANSLATION 12-3 RNA and Protein Synthesis WARM UP What are proteins? Where do they come from? From DNA to RNA to Protein DNA in our cells carry the instructions for making
Chapter 12 DNA TRANSCRIPTION and TRANSLATION 12-3 RNA and Protein Synthesis WARM UP What are proteins? Where do they come from? From DNA to RNA to Protein DNA in our cells carry the instructions for making
What is RNA? Another type of nucleic acid A working copy of DNA Does not matter if it is damaged or destroyed
 RNA Section 3.1 What is RNA? Another type of nucleic acid A working copy of DNA Does not matter if it is damaged or destroyed Used to direct the production of proteins that determines an organisms characteristics
RNA Section 3.1 What is RNA? Another type of nucleic acid A working copy of DNA Does not matter if it is damaged or destroyed Used to direct the production of proteins that determines an organisms characteristics
RNA Secondary Structure Prediction Computational Genomics Seyoung Kim
 RNA Secondary Structure Prediction 02-710 Computational Genomics Seyoung Kim Outline RNA folding Dynamic programming for RNA secondary structure prediction Covariance model for RNA structure prediction
RNA Secondary Structure Prediction 02-710 Computational Genomics Seyoung Kim Outline RNA folding Dynamic programming for RNA secondary structure prediction Covariance model for RNA structure prediction
DNA Structure & the Genome. Bio160 General Biology
 DNA Structure & the Genome Bio160 General Biology Lecture Outline I. DNA A nucleic acid II. Chromosome Structure III. Chromosomes and Genes IV. DNA vs. RNA I. DNA A Nucleic Acid Structure of DNA: Remember:
DNA Structure & the Genome Bio160 General Biology Lecture Outline I. DNA A nucleic acid II. Chromosome Structure III. Chromosomes and Genes IV. DNA vs. RNA I. DNA A Nucleic Acid Structure of DNA: Remember:
RNA is a single strand molecule composed of subunits called nucleotides joined by phosphodiester bonds.
 The Versatility of RNA Primary structure of RNA RNA is a single strand molecule composed of subunits called nucleotides joined by phosphodiester bonds. Each nucleotide subunit is composed of a ribose sugar,
The Versatility of RNA Primary structure of RNA RNA is a single strand molecule composed of subunits called nucleotides joined by phosphodiester bonds. Each nucleotide subunit is composed of a ribose sugar,
DNA, RNA and Protein Synthesis
 By the end of this lesson, I can Relate how Griffith s bacterial experiments showed that a hereditary factor was involved in transformation. Summarize how Avery s experiments led his group to conclude
By the end of this lesson, I can Relate how Griffith s bacterial experiments showed that a hereditary factor was involved in transformation. Summarize how Avery s experiments led his group to conclude
DNA is the genetic material. DNA structure. Chapter 7: DNA Replication, Transcription & Translation; Mutations & Ames test
 DNA is the genetic material Chapter 7: DNA Replication, Transcription & Translation; Mutations & Ames test Dr. Amy Rogers Bio 139 General Microbiology Hereditary information is carried by DNA Griffith/Avery
DNA is the genetic material Chapter 7: DNA Replication, Transcription & Translation; Mutations & Ames test Dr. Amy Rogers Bio 139 General Microbiology Hereditary information is carried by DNA Griffith/Avery
Gene Expression and Heritable Phenotype. CBS520 Eric Nabity
 Gene Expression and Heritable Phenotype CBS520 Eric Nabity DNA is Just the Beginning DNA was determined to be the genetic material, and the structure was identified as a (double stranded) double helix.
Gene Expression and Heritable Phenotype CBS520 Eric Nabity DNA is Just the Beginning DNA was determined to be the genetic material, and the structure was identified as a (double stranded) double helix.
Gene Expression. Student:
 Gene Expression Student: 1. A ribozyme is A. a section of the DNA that is expressed in the mrna. B. a self-splicing intron that acts like an enzyme. C. a complex made up of many ribosomes replicating the
Gene Expression Student: 1. A ribozyme is A. a section of the DNA that is expressed in the mrna. B. a self-splicing intron that acts like an enzyme. C. a complex made up of many ribosomes replicating the
Long and short/small RNA-seq data analysis
 Long and short/small RNA-seq data analysis GEF5, 4.9.2015 Sami Heikkinen, PhD, Dos. Topics 1. RNA-seq in a nutshell 2. Long vs short/small RNA-seq 3. Bioinformatic analysis work flows GEF5 / Heikkinen
Long and short/small RNA-seq data analysis GEF5, 4.9.2015 Sami Heikkinen, PhD, Dos. Topics 1. RNA-seq in a nutshell 2. Long vs short/small RNA-seq 3. Bioinformatic analysis work flows GEF5 / Heikkinen
CHAPTER 11 DNA NOTES PT. 4: PROTEIN SYNTHESIS TRANSCRIPTION & TRANSLATION
 CHAPTER 11 DNA NOTES PT. 4: PROTEIN SYNTHESIS TRANSCRIPTION & TRANSLATION DNA and the Language of Life RECAP Synthesis= Making something Protein Synthesis= Making Proteins Three steps in Protein Synthesis
CHAPTER 11 DNA NOTES PT. 4: PROTEIN SYNTHESIS TRANSCRIPTION & TRANSLATION DNA and the Language of Life RECAP Synthesis= Making something Protein Synthesis= Making Proteins Three steps in Protein Synthesis
Webquest: From DNA to Protein A Review of DNA and Gene Expression Concepts
 MARINE BIOTECHNOLOGY & BIOINFORMATICS an NSF ITEST Grant A lesson plan for Webquest: From DNA to Protein A Review of DNA and Gene Expression Concepts Designed by Elisabeth Childers (echilders@nhusd.k12.ca.us)
MARINE BIOTECHNOLOGY & BIOINFORMATICS an NSF ITEST Grant A lesson plan for Webquest: From DNA to Protein A Review of DNA and Gene Expression Concepts Designed by Elisabeth Childers (echilders@nhusd.k12.ca.us)
AGRO/ANSC/BIO/GENE/HORT 305 Fall, 2016 Overview of Genetics Lecture outline (Chpt 1, Genetics by Brooker) #1
 AGRO/ANSC/BIO/GENE/HORT 305 Fall, 2016 Overview of Genetics Lecture outline (Chpt 1, Genetics by Brooker) #1 - Genetics: Progress from Mendel to DNA: Gregor Mendel, in the mid 19 th century provided the
AGRO/ANSC/BIO/GENE/HORT 305 Fall, 2016 Overview of Genetics Lecture outline (Chpt 1, Genetics by Brooker) #1 - Genetics: Progress from Mendel to DNA: Gregor Mendel, in the mid 19 th century provided the
STUDY GUIDE SECTION 10-1 Discovery of DNA
 STUDY GUIDE SECTION 10-1 Discovery of DNA Name Period Date Multiple Choice-Write the correct letter in the blank. 1. The virulent strain of the bacterium S. pneumoniae causes disease because it a. has
STUDY GUIDE SECTION 10-1 Discovery of DNA Name Period Date Multiple Choice-Write the correct letter in the blank. 1. The virulent strain of the bacterium S. pneumoniae causes disease because it a. has
O C. 5 th C. 3 rd C. the national health museum
 Elements of Molecular Biology Cells Cells is a basic unit of all living organisms. It stores all information to replicate itself Nucleus, chromosomes, genes, All living things are made of cells Prokaryote,
Elements of Molecular Biology Cells Cells is a basic unit of all living organisms. It stores all information to replicate itself Nucleus, chromosomes, genes, All living things are made of cells Prokaryote,
Genomics and Gene Recognition Genes and Blue Genes
 Genomics and Gene Recognition Genes and Blue Genes November 3, 2004 Eukaryotic Gene Structure eukaryotic genomes are considerably more complex than those of prokaryotes eukaryotic cells have organelles
Genomics and Gene Recognition Genes and Blue Genes November 3, 2004 Eukaryotic Gene Structure eukaryotic genomes are considerably more complex than those of prokaryotes eukaryotic cells have organelles
PROTEIN SYNTHESIS Flow of Genetic Information The flow of genetic information can be symbolized as: DNA RNA Protein
 PROTEIN SYNTHESIS Flow of Genetic Information The flow of genetic information can be symbolized as: DNA RNA Protein This is also known as: The central dogma of molecular biology Protein Proteins are made
PROTEIN SYNTHESIS Flow of Genetic Information The flow of genetic information can be symbolized as: DNA RNA Protein This is also known as: The central dogma of molecular biology Protein Proteins are made
 Genome Annotation Genome annotation What is the function of each part of the genome? Where are the genes? What is the mrna sequence (transcription, splicing) What is the protein sequence? What does
Genome Annotation Genome annotation What is the function of each part of the genome? Where are the genes? What is the mrna sequence (transcription, splicing) What is the protein sequence? What does
Developmental Biology BY1101 P. Murphy
 Developmental Biology BY1101 P. Murphy Lecture 7 Cellular differentiation and the regulation of gene expression. In this lecture we looked at two main questions: How is gene expression regulated? (revision
Developmental Biology BY1101 P. Murphy Lecture 7 Cellular differentiation and the regulation of gene expression. In this lecture we looked at two main questions: How is gene expression regulated? (revision
Chapter 15 Gene Technologies and Human Applications
 Chapter Outline Chapter 15 Gene Technologies and Human Applications Section 1: The Human Genome KEY IDEAS > Why is the Human Genome Project so important? > How do genomics and gene technologies affect
Chapter Outline Chapter 15 Gene Technologies and Human Applications Section 1: The Human Genome KEY IDEAS > Why is the Human Genome Project so important? > How do genomics and gene technologies affect
Controlling DNA. Ethical guidelines for the use of DNA technology. Module Type: Discussion, literature review, and debate
 Ethical guidelines for the use of DNA technology Author: Tara Cornelisse, Ph.D. Candidate, Environmental Studies, University of California Santa Cruz. Field-tested with: 11 th -12 th grade students in
Ethical guidelines for the use of DNA technology Author: Tara Cornelisse, Ph.D. Candidate, Environmental Studies, University of California Santa Cruz. Field-tested with: 11 th -12 th grade students in
8/21/2014. From Gene to Protein
 From Gene to Protein Chapter 17 Objectives Describe the contributions made by Garrod, Beadle, and Tatum to our understanding of the relationship between genes and enzymes Briefly explain how information
From Gene to Protein Chapter 17 Objectives Describe the contributions made by Garrod, Beadle, and Tatum to our understanding of the relationship between genes and enzymes Briefly explain how information
Product Applications for the Sequence Analysis Collection
 Product Applications for the Sequence Analysis Collection Pipeline Pilot Contents Introduction... 1 Pipeline Pilot and Bioinformatics... 2 Sequence Searching with Profile HMM...2 Integrating Data in a
Product Applications for the Sequence Analysis Collection Pipeline Pilot Contents Introduction... 1 Pipeline Pilot and Bioinformatics... 2 Sequence Searching with Profile HMM...2 Integrating Data in a
DNA. translation. base pairing rules for DNA Replication. thymine. cytosine. amino acids. The building blocks of proteins are?
 2 strands, has the 5-carbon sugar deoxyribose, and has the nitrogen base Thymine. The actual process of assembling the proteins on the ribosome is called? DNA translation Adenine pairs with Thymine, Thymine
2 strands, has the 5-carbon sugar deoxyribose, and has the nitrogen base Thymine. The actual process of assembling the proteins on the ribosome is called? DNA translation Adenine pairs with Thymine, Thymine
Chapter 8: DNA and RNA
 Chapter 8: DNA and RNA Lecture Outline Enger, E. D., Ross, F. C., & Bailey, D. B. (2012). Concepts in biology (14th ed.). New York: McGraw- Hill. 1 8-1 DNA and the Importance of Proteins Proteins play
Chapter 8: DNA and RNA Lecture Outline Enger, E. D., Ross, F. C., & Bailey, D. B. (2012). Concepts in biology (14th ed.). New York: McGraw- Hill. 1 8-1 DNA and the Importance of Proteins Proteins play
Nucleic Acids And Protein Synthesis Answer Key
 And Protein Answer Key Free PDF ebook Download: And Answer Key Download or Read Online ebook nucleic acids and protein synthesis answer key in PDF Format From The Best User Guide Database Key ~. ' Date.
And Protein Answer Key Free PDF ebook Download: And Answer Key Download or Read Online ebook nucleic acids and protein synthesis answer key in PDF Format From The Best User Guide Database Key ~. ' Date.
Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017
 Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017 Agenda What is Functional Genomics? RNA Transcription/Gene Expression Measuring Gene
Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017 Agenda What is Functional Genomics? RNA Transcription/Gene Expression Measuring Gene
RNA and Protein Synthesis
 RNA and Protein Synthesis CTE: Agriculture and Natural Resources: C5.3 Understand various cell actions, such as osmosis and cell division. C5.4 Compare and contrast plant and animal cells, bacteria, and
RNA and Protein Synthesis CTE: Agriculture and Natural Resources: C5.3 Understand various cell actions, such as osmosis and cell division. C5.4 Compare and contrast plant and animal cells, bacteria, and
Self-test Quiz for Chapter 12 (From DNA to Protein: Genotype to Phenotype)
 Self-test Quiz for Chapter 12 (From DNA to Protein: Genotype to Phenotype) Question#1: One-Gene, One-Polypeptide The figure below shows the results of feeding trials with one auxotroph strain of Neurospora
Self-test Quiz for Chapter 12 (From DNA to Protein: Genotype to Phenotype) Question#1: One-Gene, One-Polypeptide The figure below shows the results of feeding trials with one auxotroph strain of Neurospora
TaqMan Advanced mirna Assays
 PRODUCT BULLETIN TaqMan Advanced mirna Assays TaqMan Advanced mirna Assays Key features Universal reverse transcription (RT) one RT step for all Applied Biosystems TaqMan Advanced mirna Assays Sensitive
PRODUCT BULLETIN TaqMan Advanced mirna Assays TaqMan Advanced mirna Assays Key features Universal reverse transcription (RT) one RT step for all Applied Biosystems TaqMan Advanced mirna Assays Sensitive
Nucleic Acids: DNA and RNA
 Nucleic Acids: DNA and RNA Living organisms are complex systems. Hundreds of thousands of proteins exist inside each one of us to help carry out our daily functions. These proteins are produced locally,
Nucleic Acids: DNA and RNA Living organisms are complex systems. Hundreds of thousands of proteins exist inside each one of us to help carry out our daily functions. These proteins are produced locally,
The Integrated Biomedical Sciences Graduate Program
 The Integrated Biomedical Sciences Graduate Program at the university of notre dame Cutting-edge biomedical research and training that transcends traditional departmental and disciplinary boundaries to
The Integrated Biomedical Sciences Graduate Program at the university of notre dame Cutting-edge biomedical research and training that transcends traditional departmental and disciplinary boundaries to
Synthetic Biology. Sustainable Energy. Therapeutics Industrial Enzymes. Agriculture. Accelerating Discoveries, Expanding Possibilities. Design.
 Synthetic Biology Accelerating Discoveries, Expanding Possibilities Sustainable Energy Therapeutics Industrial Enzymes Agriculture Design Build Generate Solutions to Advance Synthetic Biology Research
Synthetic Biology Accelerating Discoveries, Expanding Possibilities Sustainable Energy Therapeutics Industrial Enzymes Agriculture Design Build Generate Solutions to Advance Synthetic Biology Research
Assembling Protein Molecules
 How Does Dna Provide Instructions For Assembling Protein Molecules What does the information in DNA molecules provide instructions for? A. Assembling B. Assembling protein molecules into amino acids. C.
How Does Dna Provide Instructions For Assembling Protein Molecules What does the information in DNA molecules provide instructions for? A. Assembling B. Assembling protein molecules into amino acids. C.
Higher Human Biology Unit 1: Human Cells Pupils Learning Outcomes
 Higher Human Biology Unit 1: Human Cells Pupils Learning Outcomes 1.1 Division and Differentiation in Human Cells I can state that cellular differentiation is the process by which a cell develops more
Higher Human Biology Unit 1: Human Cells Pupils Learning Outcomes 1.1 Division and Differentiation in Human Cells I can state that cellular differentiation is the process by which a cell develops more
Feature Selection of Gene Expression Data for Cancer Classification: A Review
 Available online at www.sciencedirect.com ScienceDirect Procedia Computer Science 50 (2015 ) 52 57 2nd International Symposium on Big Data and Cloud Computing (ISBCC 15) Feature Selection of Gene Expression
Available online at www.sciencedirect.com ScienceDirect Procedia Computer Science 50 (2015 ) 52 57 2nd International Symposium on Big Data and Cloud Computing (ISBCC 15) Feature Selection of Gene Expression
CHAPTER 20 DNA TECHNOLOGY AND GENOMICS. Section A: DNA Cloning
 Section A: DNA Cloning 1. DNA technology makes it possible to clone genes for basic research and commercial applications: an overview 2. Restriction enzymes are used to make recombinant DNA 3. Genes can
Section A: DNA Cloning 1. DNA technology makes it possible to clone genes for basic research and commercial applications: an overview 2. Restriction enzymes are used to make recombinant DNA 3. Genes can
Sequence Databases and database scanning
 Sequence Databases and database scanning Marjolein Thunnissen Lund, 2012 Types of databases: Primary sequence databases (proteins and nucleic acids). Composite protein sequence databases. Secondary databases.
Sequence Databases and database scanning Marjolein Thunnissen Lund, 2012 Types of databases: Primary sequence databases (proteins and nucleic acids). Composite protein sequence databases. Secondary databases.
DNA vs. RNA B-4.1. Compare DNA and RNA in terms of structure, nucleotides and base pairs.
 DNA vs. RNA B-4.1 Compare DNA and RNA in terms of structure, nucleotides and base pairs. Key Concepts l Nucleic Acids: l deoxyribonucleic acid (DNA) l ribonucleic acid (RNA) l Nucleotides: l nitrogen base,
DNA vs. RNA B-4.1 Compare DNA and RNA in terms of structure, nucleotides and base pairs. Key Concepts l Nucleic Acids: l deoxyribonucleic acid (DNA) l ribonucleic acid (RNA) l Nucleotides: l nitrogen base,
What happens after DNA Replication??? Transcription, translation, gene expression/protein synthesis!!!!
 What happens after DNA Replication??? Transcription, translation, gene expression/protein synthesis!!!! Protein Synthesis/Gene Expression Why do we need to make proteins? To build parts for our body as
What happens after DNA Replication??? Transcription, translation, gene expression/protein synthesis!!!! Protein Synthesis/Gene Expression Why do we need to make proteins? To build parts for our body as
Chapter 8 Lecture Outline. Transcription, Translation, and Bioinformatics
 Chapter 8 Lecture Outline Transcription, Translation, and Bioinformatics Replication, Transcription, Translation n Repetitive processes Build polymers of nucleotides or amino acids n All have 3 major steps
Chapter 8 Lecture Outline Transcription, Translation, and Bioinformatics Replication, Transcription, Translation n Repetitive processes Build polymers of nucleotides or amino acids n All have 3 major steps
Genetics Lecture 21 Recombinant DNA
 Genetics Lecture 21 Recombinant DNA Recombinant DNA In 1971, a paper published by Kathleen Danna and Daniel Nathans marked the beginning of the recombinant DNA era. The paper described the isolation of
Genetics Lecture 21 Recombinant DNA Recombinant DNA In 1971, a paper published by Kathleen Danna and Daniel Nathans marked the beginning of the recombinant DNA era. The paper described the isolation of
Chapter 10 - Molecular Biology of the Gene
 Bio 100 - Molecular Genetics 1 A. Bacterial Transformation Chapter 10 - Molecular Biology of the Gene Researchers found that they could transfer an inherited characteristic (e.g. the ability to cause pneumonia),
Bio 100 - Molecular Genetics 1 A. Bacterial Transformation Chapter 10 - Molecular Biology of the Gene Researchers found that they could transfer an inherited characteristic (e.g. the ability to cause pneumonia),
Thermo Scientific Dharmacon SMARTvector 2.0 Lentiviral shrna Particles
 Thermo Scientific Dharmacon SMARTvector 2.0 Lentiviral shrna Particles Long-term gene silencing shrna-specific design algorithm High titer, purified particles Thermo Scientific Dharmacon SMARTvector shrna
Thermo Scientific Dharmacon SMARTvector 2.0 Lentiviral shrna Particles Long-term gene silencing shrna-specific design algorithm High titer, purified particles Thermo Scientific Dharmacon SMARTvector shrna
Ch. 10 Notes DNA: Transcription and Translation
 Ch. 10 Notes DNA: Transcription and Translation GOALS Compare the structure of RNA with that of DNA Summarize the process of transcription Relate the role of codons to the sequence of amino acids that
Ch. 10 Notes DNA: Transcription and Translation GOALS Compare the structure of RNA with that of DNA Summarize the process of transcription Relate the role of codons to the sequence of amino acids that
FACULTY OF BIOCHEMISTRY AND MOLECULAR MEDICINE
 FACULTY OF BIOCHEMISTRY AND MOLECULAR MEDICINE BIOMOLECULES COURSE: COMPUTER PRACTICAL 1 Author of the exercise: Prof. Lloyd Ruddock Edited by Dr. Leila Tajedin 2017-2018 Assistant: Leila Tajedin (leila.tajedin@oulu.fi)
FACULTY OF BIOCHEMISTRY AND MOLECULAR MEDICINE BIOMOLECULES COURSE: COMPUTER PRACTICAL 1 Author of the exercise: Prof. Lloyd Ruddock Edited by Dr. Leila Tajedin 2017-2018 Assistant: Leila Tajedin (leila.tajedin@oulu.fi)
1. Mitosis = growth, repair, asexual reproduc4on
 Places Muta4ons get passed on: Cell Reproduc4on: 2 types of cell reproduc4on: 1. Mitosis = growth, repair, asexual reproduc4on Photocopy machine Growth/Repair Passed on in the same body 2. Meiosis = sexual
Places Muta4ons get passed on: Cell Reproduc4on: 2 types of cell reproduc4on: 1. Mitosis = growth, repair, asexual reproduc4on Photocopy machine Growth/Repair Passed on in the same body 2. Meiosis = sexual
MicroSEQ Rapid Microbial Identification System
 MicroSEQ Rapid Microbial Identification System Giving you complete control over microbial identification using the gold-standard genotypic method The MicroSEQ ID microbial identification system, based
MicroSEQ Rapid Microbial Identification System Giving you complete control over microbial identification using the gold-standard genotypic method The MicroSEQ ID microbial identification system, based
From Gene to Protein via Transcription and Translation i
 How do genes influence our characteristics? From Gene to Protein via Transcription and Translation i A gene is a segment of DNA that provides the instructions for making a protein. Proteins have many different
How do genes influence our characteristics? From Gene to Protein via Transcription and Translation i A gene is a segment of DNA that provides the instructions for making a protein. Proteins have many different
Bio11 Announcements. Ch 21: DNA Biology and Technology. DNA Functions. DNA and RNA Structure. How do DNA and RNA differ? What are genes?
 Bio11 Announcements TODAY Genetics (review) and quiz (CP #4) Structure and function of DNA Extra credit due today Next week in lab: Case study presentations Following week: Lab Quiz 2 Ch 21: DNA Biology
Bio11 Announcements TODAY Genetics (review) and quiz (CP #4) Structure and function of DNA Extra credit due today Next week in lab: Case study presentations Following week: Lab Quiz 2 Ch 21: DNA Biology
Click here to read the case study about protein synthesis.
 Click here to read the case study about protein synthesis. Big Question: How do cells use the genetic information stored in DNA to make millions of different proteins the body needs? Key Concept: Genetics
Click here to read the case study about protein synthesis. Big Question: How do cells use the genetic information stored in DNA to make millions of different proteins the body needs? Key Concept: Genetics
Protein Synthesis. Lab Exercise 12. Introduction. Contents. Objectives
 Lab Exercise Protein Synthesis Contents Objectives 1 Introduction 1 Activity.1 Overview of Process 2 Activity.2 Transcription 2 Activity.3 Translation 3 Resutls Section 4 Introduction Having information
Lab Exercise Protein Synthesis Contents Objectives 1 Introduction 1 Activity.1 Overview of Process 2 Activity.2 Transcription 2 Activity.3 Translation 3 Resutls Section 4 Introduction Having information
Study Guide for Chapter 12 Exam DNA, RNA, & Protein Synthesis
 Name: Date: Period: Study Guide for Chapter 12 Exam DNA, RNA, & Protein Synthesis ***Completing this study guide in its entirety will result in extra credit on the exam. You must show me the DAY OF the
Name: Date: Period: Study Guide for Chapter 12 Exam DNA, RNA, & Protein Synthesis ***Completing this study guide in its entirety will result in extra credit on the exam. You must show me the DAY OF the
RNA Secondary Structure Prediction
 RNA Secondary Structure Prediction Outline 1) Introduction: RNA structure basics 2) Dynamic programming for RNA secondary structure prediction The Central Dogma of Molecular Biology DNA CCTGAGCCAACTATTGATGAA
RNA Secondary Structure Prediction Outline 1) Introduction: RNA structure basics 2) Dynamic programming for RNA secondary structure prediction The Central Dogma of Molecular Biology DNA CCTGAGCCAACTATTGATGAA
TRANSCRIPTION AND PROCESSING OF RNA
 TRANSCRIPTION AND PROCESSING OF RNA 1. The steps of gene expression. 2. General characterization of transcription: steps, components of transcription apparatus. 3. Transcription of eukaryotic structural
TRANSCRIPTION AND PROCESSING OF RNA 1. The steps of gene expression. 2. General characterization of transcription: steps, components of transcription apparatus. 3. Transcription of eukaryotic structural
DNA: The Genetic Material. Chapter 10
 DNA: The Genetic Material Chapter 10 DNA as the Genetic Material DNA was first extracted from nuclei in 1870 named nuclein after their source. Chemical analysis determined that DNA was a weak acid rich
DNA: The Genetic Material Chapter 10 DNA as the Genetic Material DNA was first extracted from nuclei in 1870 named nuclein after their source. Chemical analysis determined that DNA was a weak acid rich
Genes and human health - the science and ethics
 Deoxyribonucleic acid (DNA) - why is it so important? Genes and human health - the science and ethics DNA is essential to all living organisms, from bacteria to man, as it contains a code which specifies
Deoxyribonucleic acid (DNA) - why is it so important? Genes and human health - the science and ethics DNA is essential to all living organisms, from bacteria to man, as it contains a code which specifies
Year III Pharm.D Dr. V. Chitra
 Year III Pharm.D Dr. V. Chitra 1 Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Only one strand of DNA serves
Year III Pharm.D Dr. V. Chitra 1 Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Only one strand of DNA serves
Protein Synthesis. OpenStax College
 OpenStax-CNX module: m46032 1 Protein Synthesis OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section, you will
OpenStax-CNX module: m46032 1 Protein Synthesis OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section, you will
Independent Study Guide The Blueprint of Life, from DNA to Protein (Chapter 7)
 Independent Study Guide The Blueprint of Life, from DNA to Protein (Chapter 7) I. General Principles (Chapter 7 introduction) a. Morse code distinct series of dots and dashes encode the 26 letters of the
Independent Study Guide The Blueprint of Life, from DNA to Protein (Chapter 7) I. General Principles (Chapter 7 introduction) a. Morse code distinct series of dots and dashes encode the 26 letters of the
Prokaryotic Transcription
 Prokaryotic Transcription Transcription Basics DNA is the genetic material Nucleic acid Capable of self-replication and synthesis of RNA RNA is the middle man Nucleic acid Structure and base sequence are
Prokaryotic Transcription Transcription Basics DNA is the genetic material Nucleic acid Capable of self-replication and synthesis of RNA RNA is the middle man Nucleic acid Structure and base sequence are
BIOLOGY LTF DIAGNOSTIC TEST DNA to PROTEIN & BIOTECHNOLOGY
 Biology Multiple Choice 016074 BIOLOGY LTF DIAGNOSTIC TEST DNA to PROTEIN & BIOTECHNOLOGY Test Code: 016074 Directions: Each of the questions or incomplete statements below is followed by five suggested
Biology Multiple Choice 016074 BIOLOGY LTF DIAGNOSTIC TEST DNA to PROTEIN & BIOTECHNOLOGY Test Code: 016074 Directions: Each of the questions or incomplete statements below is followed by five suggested
Standards: SC.912.L.16.5: Explain the basic processes of transcription and translation, and how they result in the expression of genes.
 Delicious DNA: Transcription and Translation Simulation Using an Edible Model Authors: Darcy Holoweski and Catherine Quist Standards: SC.912.L.16.5: Explain the basic processes of transcription and translation,
Delicious DNA: Transcription and Translation Simulation Using an Edible Model Authors: Darcy Holoweski and Catherine Quist Standards: SC.912.L.16.5: Explain the basic processes of transcription and translation,
Replication Review. 1. What is DNA Replication? 2. Where does DNA Replication take place in eukaryotic cells?
 Replication Review 1. What is DNA Replication? 2. Where does DNA Replication take place in eukaryotic cells? 3. Where does DNA Replication take place in the cell cycle? 4. 4. What guides DNA Replication?
Replication Review 1. What is DNA Replication? 2. Where does DNA Replication take place in eukaryotic cells? 3. Where does DNA Replication take place in the cell cycle? 4. 4. What guides DNA Replication?
Introduction to Bioinformatics
 Introduction to Bioinformatics Alla L Lapidus, Ph.D. SPbSU St. Petersburg Term Bioinformatics Term Bioinformatics was invented by Paulien Hogeweg (Полина Хогевег) and Ben Hesper in 1970 as "the study of
Introduction to Bioinformatics Alla L Lapidus, Ph.D. SPbSU St. Petersburg Term Bioinformatics Term Bioinformatics was invented by Paulien Hogeweg (Полина Хогевег) and Ben Hesper in 1970 as "the study of
