Structure modeling and dynamics driven mutation and phosphorylation analysis of Beta-amyloid peptides
|
|
|
- Mariah Newman
- 6 years ago
- Views:
Transcription
1 Hypothesis Volume 10(9) Structure modeling and dynamics driven mutation and phosphorylation analysis of Beta-amyloid peptides Sunil kumar Singh, Ankita singh, Ved Prakash* & Selvaa kumar C Department of Biotechnology and Bioinformatics, padmashree Dr.D.Y.Patil University, Belapur , Navi Mumbai, India; Ved Prakash - Ved.prakash2012@vit.ac.in; Phone: ; *Corresponding author Received August 28, 2014; Accepted August 31, 2014; Published September 30, 2014 Abstract: The most common characteristics of diverse age-related neurodegenerative diseases are aggregation and accumulation of the misfolded protein in the brain. Alzheimer s disease (AD) is one of these protein conformational diseases. Extracellular accumulation of amyloid β (Aβ) is one the neuropathological hallmarks of Alzheimer disease. Various studies have shown that mutation in specific hydrophobic region of Aβ protein inhibit the formation of β sheet, thus aggregation of this protein is stalled. The identification of such mutation in Aβ protein can help us in elucidating the etiology of sporadic Aβ. In our study we have selected three positions: 19ILU, 21ALA and 41ILU in Aβ protein based on their hydrophobic nature and substituted them with PRO ( βsheet breaker). The effects of the substitutions were analysed using molecular dynamics simulation studies. The results validated that the mutations in the specified regions change the hydrophobicity of the protein and the βsheet formation was declined to zero per cent. Keywords: Alzheimer s disease, Amyloid-beta precursor protein, Amyloid-beta peptide, Gromos, PPI-Pred, Argus Lab, UCSF Chimera. Background: Age-related impairments in cognition and memory have been known since ancient times, but the clinical pathological features of the syndrome, now termed Alzheimer s disease (AD), were not documented in the medical literature until the first decade of this century. It is now established that AD is a complex and genetically heterogeneous, neurodegenerative disorder. It is the most common type of dementia. Dementia is derived from the Latin word, [de-=out from + mentia=the mind] means loss or impairment of mental powers due to a disease. There is no cure for AD till today, however promising research and development for early detection and treatment is underway. Plaques and Tangles: The Hallmarks of AD The brains of people with AD have an abundance of two abnormal structures: Bioinformation 10(9): (2014) Biomedical Informatics Amyloid Beta (Aβ) plaques build up between nerve cells. They contain deposits of (Aβ), a protein fragment. Neurofibrillary tangles, which are twisted fibers that build up inside the nerve cell. Amyloid Precursor Protein (APP) can be processed by at least three secretases: α-, β-, and ϒ-secretases [1, 2]. Cleavage of APP by an 'α-secretase' enzyme (a member of the ADAM family of metalloproteases) [3] occurs within the, Aβ sequence, and results in the secretion of an approx. 100 kda N-terminal fragment [1]. On the other hand, intact, Aβ is derived from APP by cellular processing pathways that involve the excision of the, Aβ region by the sequential action of 'β,-' and 'γ-secretase' enzymes, possibly in distinct subcellular compartments [4]. β- secretase (a membranebound aspartyl protease also called BACE) cleave the ectodomain of APP, resulting in the shedding
2 of APPsα and APPsβ (12 kda). The 12-kd fragment may then undergo γ -secretase cleavage within the hydrophobic transmembrane domain at either valine 710, alanine 712, or threonine 713 to release the 40, 42, or 43 residue Aβ peptides. The strongest evidence that abnormal proteolytic processing and increased Aβ generation are central to the disease process comes from studies of very rare inherited forms of AD [3]. Amyloid-β Peptide Conformations: Monomers The Aβ is a peptide that is ubiquitously and normally expressed in humans predominately in two forms, 39- and 43- amino acids in length (Aβ40 and Aβ42, respectively) [5, 6, 7]. Monomers are about 1.0 ± 0.3 nm in size, with a molecular weight of: Da for Aβ (1-40) and Da for Aβ (1-42). Monomers present mostly random coils and α-helix secondary structures. Two α-helical regions exist at residues 8-25 and 28-38, and these regions are separated by a flexible hinge. The rest of the peptide adopts random coil-like conformation [8]. conformational states of the Aβ peptides in aqueous media. MD studies have suggested that Aβ peptides, rather than being entirely disordered, are usually able to sample sequencespecific secondary structures. Using chemical shifts calculated from replica exchange molecular dynamics simulations, Wood and Rothlisberger investigated the differences observed between theoretical studies and experiments. They showed that the assigned random coil structures often derive from an averaging of β-sheet, α-helical and random coil structures [10]. Insertion of a bulky group or proline as β-sheet breaker within the self-recognition-derived peptide sequenc, KLVFF, has been demonstrated as effective in inhibiting amyloid aggregation. Furthermore, peptidic inhibitors were also reported that via stabilization of the helical structure in residues of Aβ prevented the formation of neurotoxic aggregates [10, 11]. Common Pathways of Aggregation amyloid-β peptide The earliest event in the process of aggregation is the formation of dimmers. Aβ derived from a segment of APP is believed to be predominantly α-helical in its native state. On the other hand, monomeric Aβ is largely conformed as random coil in solution. The formation of Aβ dimmers coincides with the adoption of partial β structure. The Aβ dimmers assemble into higher order aggregates or oligomers, since there is a remarkable increase in the amounts of higher order aggregates and a decrease in the amount of dimmers. These higher order Aβ aggregates are protein micelles with a hydrophobic core and a polar exterior that appear as 3 nm spherical particles determined by atomic force or electron microscopy These higher order Aβ aggregates are protein micelles with a hydrophobic core and a polar exterior that appear as 3 nm spherical particles determined by atomic force or electron microscopy. At later times, these spherical shape particles appear to form curvilinear strings or protofibrils. The protofibrils undergo a conformation change to form the straight, unbranched mature fibrils. Once the amyloid fibril lattice has been established, the fibril can grow by the addition of Aβ monomer or dimmer at the ends of the fibril. The higher order aggregates have molecular weights ranging from approximately 105 to 106 Da and with an average size corresponding to approximately 24 monomers. Aβ oligomers that appear as spherical particles are approximately 3 nm in diameter and have the characteristics of a protein amphipathic, since it lowers the surface tension of water. At longer incubation times, the oligomers also appear to co-aggregate to form curvilinear fibrils with a characteristic beaded appearance. The spherical oligomers and protofibrils appear to be intermediates in the pathway of fiber formation because they disappear as mature fibrils accumulate and the rate of monomer dissociation from them is too slow to account for fibril growth. The transition appears to involve a major conformational change, because fluorescence quenching analysis indicates that the carboxyl terminus is highly shielded from the aqueous solvent in the soluble oligomeric state, whereas it is exposed to the solvent in the fibrillar state. Once the amyloid fibril lattice has been established, it can grow by the addition of monomers onto the ends of the fibrils [9]. Molecular simulations offer a direct means of accessing the Figure 1: Amyloid Beta sequence and structure Methodology: Sequence & Structure Solution NMR structure PDB 1IYT was taken from the RCSB Protein Data and employed as starting configurations for the MD simulations and other analysis (Figure 1). Figure 2: In silico analysis of putative phosphorylation sites of Aβ: Protein sequences of human Aβ1-42 sequences were analyzed by using NetPhos 2.0 computational prediction tool Bioinformation 10(9): (2014) Biomedical Informatics
3 ( The result from NetPhosK contains three parts for each of the protein/peptide sequence analyzed. The first part indicates the name, length of the aa sequence and predicted phospho sites (a). The second part shows the predicted phospho residues, their positions in the sequence and the respective phospho prediction score (b). The third part shows the graphical illustrations of phosphorylation potential of predicted phospho sites (c). phosphorylation sites obtained from other protein homologues from higher eukaryotes. The NetPhos 2.0 server is a generic (non kinase specific) phosphorylation predictions server and perform the predictions for serine, threonine and tyrosine phosphorylation sites in protein/peptides. The input sequences of any protein/peptide in the one-letter amino acid code in FASTA format can be used for carrying out the predictions. The instructions for the usage of the programme are provided with the respective tools. In-silico phosphorylation of Aβ was done using ArgusLab and also by making changes in the coordinate file. Figure 3: In silico phosphorylation of Aβ. Phosphate group was added on serine 8 using ArgusLab and making changes in the coordinate file. The above figure shows phosphorylated and non phosphorylated Aβ. Figure 5: Hydrophobicity comparision: Phosphorylation of amyloid beta increases the hydrophobicity of the binding site (black circles) which means self aggregation of the peptide will be enhanced. RED and ORANGE regions are hydrophobic (UCSF Chimera Version ) Figure 4: PPI-Pred Analysis- Shows that the binding patch is increased (red) in case of paβ which depicts that paβ will interact more efficiently. NetPhos Server & Phosphorylation The identification of the phosphorylation sites of Aβ was carried out using NetPhosK 1.0 (Blom et al., 1999), and NetPhos 2.0 sever (Blom et al.,2004) (Figure 2). These two prediction programmes employ neural network based algorithms prediction processes which are based on the evolutionary information obtained from sequence similarity of the phosphorylation site and taxonomy. The NetPhosK 1.0 is a kinase specific eukaryotic phosphorylation site predictions server. The kinase predictions are verified with homologues Figure 6: Residues 19, 21, and 41 are replaced by PROLINE to study the effect of mutants (Arguslab ) and Comparing the hydrophobicity of the wildtype and mutants in UCSF Chimere Vers;ion revelated that hydrophobicity of self-recognition site of the peptide was reduced with might reduce their aggregative ability. Mutations & Hydrophobicity The Aβ was mutated at residues 19, 21 and 41 using Swiss- PdbViewer The energy of the mutant structure was minimised. The most probable binding sites for Aβ and its phosphorylated and mutant form was analysed using PPI-Pred ( The hydrophobicity Bioinformation 10(9): (2014) Biomedical Informatics
4 comparison between the three forms was carried out using chimera Simulation Three molecular dynamic simulations, one for each form of Aβ was performed for 25 ns using GROMOS96. GROMOS ffg53a6 force field was used. The peptide was cantered in a cubic simulation box with a 1 nm distance allowed between the peptide and the edges of the box treated with periodic boundary conditions. The particle mesh Ewald method was employed for treating long-range electrostatics with a 1.4 nm cutoff for calculating short-range forces. After steepest descent energy minimization in vacuo, the box was solvated with the SPC explicit water model, and Na+ and Cl ions were added to obtain a NaCl concentration of 150 mm and to achieve charge neutrality. The solvated peptide was then minimized using both steepest descent and conjugate gradient energy minimization methods. We performed a 100 ps equilibration dynamics (NVT/NPT) in which the water molecules were allowed to equilibrate around the restrained peptide atoms, in the process removing bad contacts and bringing the system near equilibrium conditions for the subsequent production MD run. For the equilibration step, the system was coupled to a Berendsen thermostat and barostat. The restraints were subsequently turned off, and a 25 ns production run was performed at 300 K in an NPT ensemble with temperature and pressure modulated by coupling to a Nose Hoover thermostat and a Parrinello Rahman barostat, respectively. The neighbor list was generated every 10 ps, with a cutoff of 1.4 nm, and coordinates were saved every 20 ps. Pymol was used for the visualization of the simulation results. Figure 7: Shows secondary structure transition in wild type, mutant F19P, mutant F21P, MUTANT 141P respectively during 25ns MD Simulations using the GROMS forcefiedl ffg53a6. Thr helical peptide is converting into parallel beat sheet. The three state Composition was predicted using DSSP server. ( Bioinformation 10(9): (2014) Biomedical Informatics
5 Figure 8: RMSD s of the wild type, mutant F19P, mutant F21P, mutant 141P respectively trajectories plotted vs time and RMSF plot for each residue of wild type, mutant F21P, mutant 141P respectively. Secondary Structure Dictionary of Secondary Structure of Proteins (DSSP) ( was used to analysise the secondary structure conformation of the wildtype and mutant Aβ after simulation. Results: Prediction of putative phosphorylation sites of Aβ The preliminary identification of the putative phosphorylation sites of Aβ and identification of the responsible kinases were carried out by in silico analysis using freely available world wide web (www) based Netphos2.0 computational tool ( The results from the in silico analysis indicate that Ser-8, Ser-26 and Tyr-10 residues might be potential phosphorylation sites in Aβ sequence. The serine at 8th position had the highest prediction score of The serine at 26 th position had a prediction score of The tyrosine at 10th position has a score of (Figure 2) Phosphorylation of Aβ The Aβ sequence contains two serine residues at 8th and 26th position, a tyrosine residue at 10th position which could possibly undergo phosphorylation. The primary goal of this work was to identify the role of phosphorylation in Aβ aggregation and pathogenesis of AD. Therefore, investigations were carried out to predict/identify/determine putative phosphorylation sites of Aβ. Phosphate group was added on serine 8 using ArgusLab (Figure 3). PPI-pred analysis shows that the binding patch is increased (red) in case of paβ which depicts that paβ will interact more efficiently (Figure 4). Bioinformation 10(9): (2014) Biomedical Informatics
6 Phosphorylation of amyloid beta increases the hydrophobicity of the binding site (black circles) which means self aggregation of the peptide will be enhanced. RED and ORANGE regions are hydrophobic (Figure 5). Discussion: The current investigation was aimed at understanding the role of extracellular phosphorylation and mutation of Aβ peptide in aggregation. The aggregation of Aβ peptides is significantly related to the pathogenesis of neuronal degeneration in AD. Despite many previous studies on the structural analysis of Aβ aggregates, the precise mechanism has not yet been clarified. To obtain information on the structure of Aβ42 fibrils, we performed in-silico phosphorylation as well as proline replacement in Aβ peptide. The aggregative ability of the modified forms was analysed by hydrophobicity comparison and molecular dynamics. The analysis gave an insight into the role of phosphorylation at serine 8 which is capable of enhancing the propensity of Aβ to adopt a β-sheet rich conformation due increase in the hydrophobicity in binding sites. The phosphorylation induced β-sheet rich structures may accelerate the formation of small oligomeric aggregates that might seed aggregation into larger oligomeric and fibrillar assemblies. Phosphorylation of Ser8 negatively regulates Aβ degradation. It decreases the clearance by microglial cells and thus promotes its aggregation. Thus the inhibition of extracellular Aβ phosphorylation might play a role in the therapy and/or prevention of AD. This analysis also sheds light on the effect of mutation in Aβ. Proline-substituted mutants of Aβ42 were formed in-silico and their aggregative ability was studies using MD simulation. The analysis revealed that F19P-, and F21P-Aβ42 did not form β- sheet therefore their aggregative ability was reduced. Whereas Aβ42 formed β-sheets thus they aggregated far more rapidly than the mutant forms. Previous studies revealed that the C-terminal two residues of Aβ42 play a critical role in its aggregative ability and neurotoxicity. Weinreb et al. proposed the hypothesis of hydrophobic cluster, stating that hydrophobic interaction among the side chains at the C terminus induces aggregation. In this hypothesis, Ile-41 is incorporated in the hydrophobic core formed by Leu-34 and Met-35. To confirm the role of the hydrophobic side chains at the C terminus of Aβ42, the hydrophilic threonine mutants at positions 41 or 42 (I41T- and A42T-A_42) were prepared and examined for their aggregative ability and neurotoxicity. Both I41T- and A42T-Aβ42 aggregated rapidly similar to wild-type Aβ42. Substitution with Thr did not abolish their cytotoxic effects [12]. Thus in this work C-terminal residue Ile 41 was also replaced by proline to find out whether C-terminal residues participates in the β-sheet formation. MD simulation was performed for the mutant to test the aggregative ability and neurotoxicity. The analysis revealed that the mutant did not form β-sheet therefore their aggregative ability was reduced. However, the C-terminal structure in the Aβ40 aggregation model is quite different from that of Aβ42. Our analysis data indicated that the C-terminal residues adopt a β-sheet structure thus helps in aggregation because of which Aβ40 aggregates far more slowly than Aβ42. Conclusion: Beta sheet is responsible for the formation of aggregates of Beta Amyloid (Aβ) in the Brain cells and since Proline (P) is a Beta sheet breaker, so Proline was used in order to reduce the Beta sheet formation and thus to reduce its aggregation too (Figure 6). Simulation of Wild type Beta amyloid at 25 nano second by using Gromacs shows that Beta sheet is formed at the rate of 9.5% while for mutant Beta Amyloid i.e. F19P, F21P and I41P, Beta sheet formation was decreased to Zero (Figure 7 & 8). Thus mutation (substitution of Proline) at position 19, 21 and 41 show the null Beta sheet formation thus resulting to no aggregation of Beta Amyloid and thus can be employed for the treatment of Alzheimer disease. References: [1] [2] [3] [4] / [5] [6] [7] [8] [9] [10] [11] BIOJ/185TOBIOJ.pdf [12] Edited by P Kangueane Citation: Singh et al. Bioinformation 10(9): (2014) License statement: This is an open-access article, which permits unrestricted use, distribution, and reproduction in any medium, for non-commercial purposes, provided the original author and source are credited Bioinformation 10(9): (2014) Biomedical Informatics
CFSSP: Chou and Fasman Secondary Structure Prediction server
 Wide Spectrum, Vol. 1, No. 9, (2013) pp 15-19 CFSSP: Chou and Fasman Secondary Structure Prediction server T. Ashok Kumar Department of Bioinformatics, Noorul Islam College of Arts and Science, Kumaracoil
Wide Spectrum, Vol. 1, No. 9, (2013) pp 15-19 CFSSP: Chou and Fasman Secondary Structure Prediction server T. Ashok Kumar Department of Bioinformatics, Noorul Islam College of Arts and Science, Kumaracoil
Comparative Modeling Part 1. Jaroslaw Pillardy Computational Biology Service Unit Cornell Theory Center
 Comparative Modeling Part 1 Jaroslaw Pillardy Computational Biology Service Unit Cornell Theory Center Function is the most important feature of a protein Function is related to structure Structure is
Comparative Modeling Part 1 Jaroslaw Pillardy Computational Biology Service Unit Cornell Theory Center Function is the most important feature of a protein Function is related to structure Structure is
Turn Plasticity Distinguishes Different Modes of Amyloid-β Aggregation
 SUPPORTING INFORMATION Turn Plasticity Distinguishes Different Modes of Amyloid-β Aggregation Nasrollah Rezaei-Ghaleh, Mehriar Amininasab, Karin Giller, Sathish Kumar, Anne Stündl, Anja Schneider, Stefan
SUPPORTING INFORMATION Turn Plasticity Distinguishes Different Modes of Amyloid-β Aggregation Nasrollah Rezaei-Ghaleh, Mehriar Amininasab, Karin Giller, Sathish Kumar, Anne Stündl, Anja Schneider, Stefan
Protein Structure Analysis
 BINF 731 Protein Structure Analysis http://binf.gmu.edu/vaisman/binf731/ Secondary Structure: Computational Problems Secondary structure characterization Secondary structure assignment Secondary structure
BINF 731 Protein Structure Analysis http://binf.gmu.edu/vaisman/binf731/ Secondary Structure: Computational Problems Secondary structure characterization Secondary structure assignment Secondary structure
Zool 3200: Cell Biology Exam 3 3/6/15
 Name: Trask Zool 3200: Cell Biology Exam 3 3/6/15 Answer each of the following questions in the space provided; circle the correct answer or answers for each multiple choice question and circle either
Name: Trask Zool 3200: Cell Biology Exam 3 3/6/15 Answer each of the following questions in the space provided; circle the correct answer or answers for each multiple choice question and circle either
Protein Folding Problem I400: Introduction to Bioinformatics
 Protein Folding Problem I400: Introduction to Bioinformatics November 29, 2004 Protein biomolecule, macromolecule more than 50% of the dry weight of cells is proteins polymer of amino acids connected into
Protein Folding Problem I400: Introduction to Bioinformatics November 29, 2004 Protein biomolecule, macromolecule more than 50% of the dry weight of cells is proteins polymer of amino acids connected into
Proteins Higher Order Structures
 Proteins Higher Order Structures Dr. Mohammad Alsenaidy Department of Pharmaceutics College of Pharmacy King Saud University Office: AA 101 msenaidy@ksu.edu.sa Previously on PHT 426!! Protein Structures
Proteins Higher Order Structures Dr. Mohammad Alsenaidy Department of Pharmaceutics College of Pharmacy King Saud University Office: AA 101 msenaidy@ksu.edu.sa Previously on PHT 426!! Protein Structures
Case 7 A Storage Protein From Seeds of Brassica nigra is a Serine Protease Inhibitor
 Case 7 A Storage Protein From Seeds of Brassica nigra is a Serine Protease Inhibitor Focus concept Purification of a novel seed storage protein allows sequence analysis and determination of the protein
Case 7 A Storage Protein From Seeds of Brassica nigra is a Serine Protease Inhibitor Focus concept Purification of a novel seed storage protein allows sequence analysis and determination of the protein
Ch Biophysical Chemistry
 Ch 247.53. Biophysical Chemistry Nina Rosario L. Rojas 2012-2013 sem 1 Review of Protein Structure Why structure? Primary, secondary, tertiary structure Disulfide bonds scheme 2 STRUCTURE- REGULAR STRUCTURE
Ch 247.53. Biophysical Chemistry Nina Rosario L. Rojas 2012-2013 sem 1 Review of Protein Structure Why structure? Primary, secondary, tertiary structure Disulfide bonds scheme 2 STRUCTURE- REGULAR STRUCTURE
Case 7 A Storage Protein From Seeds of Brassica nigra is a Serine Protease Inhibitor Last modified 29 September 2005
 Case 7 A Storage Protein From Seeds of Brassica nigra is a Serine Protease Inhibitor Last modified 9 September 005 Focus concept Purification of a novel seed storage protein allows sequence analysis and
Case 7 A Storage Protein From Seeds of Brassica nigra is a Serine Protease Inhibitor Last modified 9 September 005 Focus concept Purification of a novel seed storage protein allows sequence analysis and
CS273: Algorithms for Structure Handout # 5 and Motion in Biology Stanford University Tuesday, 13 April 2004
 CS273: Algorithms for Structure Handout # 5 and Motion in Biology Stanford University Tuesday, 13 April 2004 Lecture #5: 13 April 2004 Topics: Sequence motif identification Scribe: Samantha Chui 1 Introduction
CS273: Algorithms for Structure Handout # 5 and Motion in Biology Stanford University Tuesday, 13 April 2004 Lecture #5: 13 April 2004 Topics: Sequence motif identification Scribe: Samantha Chui 1 Introduction
Multi-compartmental modeling of SORLA s influence on amyloidogenic processing in Alzheimer s disease
 Lao et al. BMC Systems Biology 2012, 6:74 RESEARCH ARTICLE Multi-compartmental modeling of SORLA s influence on amyloidogenic processing in Alzheimer s disease Angelyn Lao 1, Vanessa Schmidt 2, Yvonne
Lao et al. BMC Systems Biology 2012, 6:74 RESEARCH ARTICLE Multi-compartmental modeling of SORLA s influence on amyloidogenic processing in Alzheimer s disease Angelyn Lao 1, Vanessa Schmidt 2, Yvonne
6-Foot Mini Toober Activity
 Big Idea The interaction between the substrate and enzyme is highly specific. Even a slight change in shape of either the substrate or the enzyme may alter the efficient and selective ability of the enzyme
Big Idea The interaction between the substrate and enzyme is highly specific. Even a slight change in shape of either the substrate or the enzyme may alter the efficient and selective ability of the enzyme
Programme Good morning and summary of last week Levels of Protein Structure - I Levels of Protein Structure - II
 Programme 8.00-8.10 Good morning and summary of last week 8.10-8.30 Levels of Protein Structure - I 8.30-9.00 Levels of Protein Structure - II 9.00-9.15 Break 9.15-11.15 Exercise: Building a protein model
Programme 8.00-8.10 Good morning and summary of last week 8.10-8.30 Levels of Protein Structure - I 8.30-9.00 Levels of Protein Structure - II 9.00-9.15 Break 9.15-11.15 Exercise: Building a protein model
Alzheimer s Disease. Table of Contents. Differential processing of APP Hyperphosphorylation of Tau
 Alzheimer s Disease Alzheimer s disease (AD) is one of the most common neurodegenerative diseases worldwide. Clinically, it is characterized by the presence of extracellular amyloid plaques and intracellular
Alzheimer s Disease Alzheimer s disease (AD) is one of the most common neurodegenerative diseases worldwide. Clinically, it is characterized by the presence of extracellular amyloid plaques and intracellular
Solutions to Problem Set 1
 MIT Department of Biology 7.014 Introductory Biology, Spring 004 Question 1 Solutions to 7.014 Problem Set 1 a) Describe the conditions of the atmosphere on prebiotic earth and how these conditions differ
MIT Department of Biology 7.014 Introductory Biology, Spring 004 Question 1 Solutions to 7.014 Problem Set 1 a) Describe the conditions of the atmosphere on prebiotic earth and how these conditions differ
Protein Folding and Misfolding
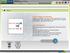 Folding of proteins into their native conformations occurs spontaneously under physiological conditions and is dictated by the primary structure of the protein. Learning Objective After interacting with
Folding of proteins into their native conformations occurs spontaneously under physiological conditions and is dictated by the primary structure of the protein. Learning Objective After interacting with
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature06147 SUPPLEMENTARY INFORMATION Figure S1 The genomic and domain structure of Dscam. The Dscam gene comprises 24 exons, encoding a signal peptide (SP), 10 IgSF domains, 6 fibronectin
doi: 10.1038/nature06147 SUPPLEMENTARY INFORMATION Figure S1 The genomic and domain structure of Dscam. The Dscam gene comprises 24 exons, encoding a signal peptide (SP), 10 IgSF domains, 6 fibronectin
Fig. 1. A schematic illustration of pipeline from gene to drug : integration of virtual and real experiments.
 1 Integration of Structure-Based Drug Design and SPR Biosensor Technology in Discovery of New Lead Compounds Alexis S. Ivanov Institute of Biomedical Chemistry RAMS, 10, Pogodinskaya str., Moscow, 119121,
1 Integration of Structure-Based Drug Design and SPR Biosensor Technology in Discovery of New Lead Compounds Alexis S. Ivanov Institute of Biomedical Chemistry RAMS, 10, Pogodinskaya str., Moscow, 119121,
Journal of Chemical and Pharmaceutical Research
 Available online www.jocpr.com Journal of Chemical and Pharmaceutical Research ISSN No: 0975-7384 CODEN(USA): JCPRC5 J. Chem. Pharm. Res., 2011, 3(6):66-72 Molecular Dynamics Simulation of a NaK channel
Available online www.jocpr.com Journal of Chemical and Pharmaceutical Research ISSN No: 0975-7384 CODEN(USA): JCPRC5 J. Chem. Pharm. Res., 2011, 3(6):66-72 Molecular Dynamics Simulation of a NaK channel
NNvPDB: Neural Network based Protein Secondary Structure Prediction with PDB Validation
 www.bioinformation.net Web server Volume 11(8) NNvPDB: Neural Network based Protein Secondary Structure Prediction with PDB Validation Seethalakshmi Sakthivel, Habeeb S.K.M* Department of Bioinformatics,
www.bioinformation.net Web server Volume 11(8) NNvPDB: Neural Network based Protein Secondary Structure Prediction with PDB Validation Seethalakshmi Sakthivel, Habeeb S.K.M* Department of Bioinformatics,
Peptoid-Based KLVFF Mimics: A Unique Approach to Alzheimer's Disease
 University of Arkansas, Fayetteville ScholarWorks@UARK Theses and Dissertations 7-2015 Peptoid-Based KLVFF Mimics: A Unique Approach to Alzheimer's Disease James Phillip Turner University of Arkansas,
University of Arkansas, Fayetteville ScholarWorks@UARK Theses and Dissertations 7-2015 Peptoid-Based KLVFF Mimics: A Unique Approach to Alzheimer's Disease James Phillip Turner University of Arkansas,
Interaction of Amyloid Inhibitor Proteins with Amyloid Beta Peptides: Insight from Molecular Dynamics Simulations
 RESEARCH ARTICLE Interaction of Amyloid Inhibitor Proteins with Amyloid Beta Peptides: Insight from Molecular Dynamics Simulations Payel Das 1 *, Seung-gu Kang 1, Sally Temple 2, Georges Belfort 3 OPEN
RESEARCH ARTICLE Interaction of Amyloid Inhibitor Proteins with Amyloid Beta Peptides: Insight from Molecular Dynamics Simulations Payel Das 1 *, Seung-gu Kang 1, Sally Temple 2, Georges Belfort 3 OPEN
Experimental Techniques 2
 Experimental Techniques 2 High-throughput interaction detection Yeast two-hybrid - pairwise organisms as machines to learn about organisms yeast, worm, fly, human,... low intersection between repeated
Experimental Techniques 2 High-throughput interaction detection Yeast two-hybrid - pairwise organisms as machines to learn about organisms yeast, worm, fly, human,... low intersection between repeated
Protein Structure/Function Relationships
 Protein Structure/Function Relationships W. M. Grogan, Ph.D. OBJECTIVES 1. Describe and cite examples of fibrous and globular proteins. 2. Describe typical tertiary structural motifs found in proteins.
Protein Structure/Function Relationships W. M. Grogan, Ph.D. OBJECTIVES 1. Describe and cite examples of fibrous and globular proteins. 2. Describe typical tertiary structural motifs found in proteins.
Homology modeling and structural validation of tissue factor pathway inhibitor
 www.bioinformation.net Hypothesis Volume 9(16) Homology modeling and structural validation of tissue factor pathway inhibitor Piyush Agrawal, Zoozeal Thakur & Mahesh Kulharia* Centre for Bioinformatics,
www.bioinformation.net Hypothesis Volume 9(16) Homology modeling and structural validation of tissue factor pathway inhibitor Piyush Agrawal, Zoozeal Thakur & Mahesh Kulharia* Centre for Bioinformatics,
SECONDARY STRUCTURE AND OTHER 1D PREDICTION MICHAEL TRESS, CNIO
 SECONDARY STRUCTURE AND OTHER 1D PREDICTION MICHAEL TRESS, CNIO Amino acids have characteristic local features Protein structures have limited room for manoeuvre because the peptide bond is planar, there
SECONDARY STRUCTURE AND OTHER 1D PREDICTION MICHAEL TRESS, CNIO Amino acids have characteristic local features Protein structures have limited room for manoeuvre because the peptide bond is planar, there
Structure formation and association of biomolecules. Prof. Dr. Martin Zacharias Lehrstuhl für Molekulardynamik (T38) Technische Universität München
 Structure formation and association of biomolecules Prof. Dr. Martin Zacharias Lehrstuhl für Molekulardynamik (T38) Technische Universität München Motivation Many biomolecules are chemically synthesized
Structure formation and association of biomolecules Prof. Dr. Martin Zacharias Lehrstuhl für Molekulardynamik (T38) Technische Universität München Motivation Many biomolecules are chemically synthesized
LIST OF ACRONYMS & ABBREVIATIONS
 LIST OF ACRONYMS & ABBREVIATIONS ALA ARG ASN ATD CRD CYS GLN GLU GLY GPCR HIS hstr ILE LEU LYS MET mglur1 NHDC PDB PHE PRO SER T1R2 T1R3 TMD TRP TYR THR 7-TM VFTM ZnSO 4 Alanine, A Arginine, R Asparagines,
LIST OF ACRONYMS & ABBREVIATIONS ALA ARG ASN ATD CRD CYS GLN GLU GLY GPCR HIS hstr ILE LEU LYS MET mglur1 NHDC PDB PHE PRO SER T1R2 T1R3 TMD TRP TYR THR 7-TM VFTM ZnSO 4 Alanine, A Arginine, R Asparagines,
Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid
 Supplementary Figures Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid sequence of Drosophila RCC1. Same colors are for Figure 1 with sequence of β-wedge that interacts with Ran in
Supplementary Figures Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid sequence of Drosophila RCC1. Same colors are for Figure 1 with sequence of β-wedge that interacts with Ran in
Algorithms in Bioinformatics ONE Transcription Translation
 Algorithms in Bioinformatics ONE Transcription Translation Sami Khuri Department of Computer Science San José State University sami.khuri@sjsu.edu Biology Review DNA RNA Proteins Central Dogma Transcription
Algorithms in Bioinformatics ONE Transcription Translation Sami Khuri Department of Computer Science San José State University sami.khuri@sjsu.edu Biology Review DNA RNA Proteins Central Dogma Transcription
MATH 5610, Computational Biology
 MATH 5610, Computational Biology Lecture 2 Intro to Molecular Biology (cont) Stephen Billups University of Colorado at Denver MATH 5610, Computational Biology p.1/24 Announcements Error on syllabus Class
MATH 5610, Computational Biology Lecture 2 Intro to Molecular Biology (cont) Stephen Billups University of Colorado at Denver MATH 5610, Computational Biology p.1/24 Announcements Error on syllabus Class
Structural Bioinformatics (C3210) Conformational Analysis Protein Folding Protein Structure Prediction
 Structural Bioinformatics (C3210) Conformational Analysis Protein Folding Protein Structure Prediction Conformational Analysis 2 Conformational Analysis Properties of molecules depend on their three-dimensional
Structural Bioinformatics (C3210) Conformational Analysis Protein Folding Protein Structure Prediction Conformational Analysis 2 Conformational Analysis Properties of molecules depend on their three-dimensional
11 questions for a total of 120 points
 Your Name: BYS 201, Final Exam, May 3, 2010 11 questions for a total of 120 points 1. 25 points Take a close look at these tables of amino acids. Some of them are hydrophilic, some hydrophobic, some positive
Your Name: BYS 201, Final Exam, May 3, 2010 11 questions for a total of 120 points 1. 25 points Take a close look at these tables of amino acids. Some of them are hydrophilic, some hydrophobic, some positive
Comparative Simulation Studies of Native and Single-Site Mutant Human Beta-Defensin-1 Peptides
 Comparative Simulation Studies of Native and Single-Site Mutant Human Beta-Defensin-1 Peptides Rabab A. Toubar, Artem Zhmurov, Valeri Barsegov, * and Kenneth A. Marx * Department of Chemistry, University
Comparative Simulation Studies of Native and Single-Site Mutant Human Beta-Defensin-1 Peptides Rabab A. Toubar, Artem Zhmurov, Valeri Barsegov, * and Kenneth A. Marx * Department of Chemistry, University
Basic concepts of molecular biology
 Basic concepts of molecular biology Gabriella Trucco Email: gabriella.trucco@unimi.it Life The main actors in the chemistry of life are molecules called proteins nucleic acids Proteins: many different
Basic concepts of molecular biology Gabriella Trucco Email: gabriella.trucco@unimi.it Life The main actors in the chemistry of life are molecules called proteins nucleic acids Proteins: many different
Bioinformatics Prof. M. Michael Gromiha Department of Biotechnology Indian Institute of Technology, Madras. Lecture - 5a Protein sequence databases
 Bioinformatics Prof. M. Michael Gromiha Department of Biotechnology Indian Institute of Technology, Madras Lecture - 5a Protein sequence databases In this lecture, we will mainly discuss on Protein Sequence
Bioinformatics Prof. M. Michael Gromiha Department of Biotechnology Indian Institute of Technology, Madras Lecture - 5a Protein sequence databases In this lecture, we will mainly discuss on Protein Sequence
Introduction to Proteins
 Introduction to Proteins Lecture 4 Module I: Molecular Structure & Metabolism Molecular Cell Biology Core Course (GSND5200) Matthew Neiditch - Room E450U ICPH matthew.neiditch@umdnj.edu What is a protein?
Introduction to Proteins Lecture 4 Module I: Molecular Structure & Metabolism Molecular Cell Biology Core Course (GSND5200) Matthew Neiditch - Room E450U ICPH matthew.neiditch@umdnj.edu What is a protein?
Protein Structure Databases, cont. 11/09/05
 11/9/05 Protein Structure Databases (continued) Prediction & Modeling Bioinformatics Seminars Nov 10 Thurs 3:40 Com S Seminar in 223 Atanasoff Computational Epidemiology Armin R. Mikler, Univ. North Texas
11/9/05 Protein Structure Databases (continued) Prediction & Modeling Bioinformatics Seminars Nov 10 Thurs 3:40 Com S Seminar in 223 Atanasoff Computational Epidemiology Armin R. Mikler, Univ. North Texas
Translation. Protein Synthesis
 Protein Structure Translation Protein Synthesis Size and Shape Comparison of Proteins Levels of Protein Structure 1 o 2 o 3 o 4 o Amino Acids Peptide Bonds Proteins are formed by creating peptide bonds
Protein Structure Translation Protein Synthesis Size and Shape Comparison of Proteins Levels of Protein Structure 1 o 2 o 3 o 4 o Amino Acids Peptide Bonds Proteins are formed by creating peptide bonds
BC 367, Exam 2 November 13, Part I. Multiple Choice (3 pts each)- Please circle the single best answer.
 Name BC 367, Exam 2 November 13, 2008 Part I. Multiple Choice (3 pts each)- Please circle the single best answer. 1. The enzyme pyruvate dehydrogenase catalyzes the following reaction. What kind of enzyme
Name BC 367, Exam 2 November 13, 2008 Part I. Multiple Choice (3 pts each)- Please circle the single best answer. 1. The enzyme pyruvate dehydrogenase catalyzes the following reaction. What kind of enzyme
Docking studies on short and ultra-short peptides against beta-secretase (BACE1) enzyme
 Research Article Docking studies on short and ultra-short peptides against beta-secretase (BACE1) enzyme Pandurangan Perumal 1, Liew Chun Lap 1, Ling Yu Luong 1, Tan Ling Hoi 1, Pavadai Parasuraman 2 ABSTRACT
Research Article Docking studies on short and ultra-short peptides against beta-secretase (BACE1) enzyme Pandurangan Perumal 1, Liew Chun Lap 1, Ling Yu Luong 1, Tan Ling Hoi 1, Pavadai Parasuraman 2 ABSTRACT
Protein Structure. Protein Structure Tertiary & Quaternary
 Lecture 4 Protein Structure Protein Structure Tertiary & Quaternary Dr. Sameh Sarray Hlaoui Primary structure: The linear sequence of amino acids held together by peptide bonds. Secondary structure: The
Lecture 4 Protein Structure Protein Structure Tertiary & Quaternary Dr. Sameh Sarray Hlaoui Primary structure: The linear sequence of amino acids held together by peptide bonds. Secondary structure: The
Supplementary Information
 Supplementary Information Rationally Designed Peptidomimetic Modulators of A Toxicity in Alzheimer s Disease K. Rajasekhar, 1 S. N. Suresh, 2 Ravi Manjithaya* 2 and T. Govindaraju* 1 1 Bioorganic Chemistry
Supplementary Information Rationally Designed Peptidomimetic Modulators of A Toxicity in Alzheimer s Disease K. Rajasekhar, 1 S. N. Suresh, 2 Ravi Manjithaya* 2 and T. Govindaraju* 1 1 Bioorganic Chemistry
The relationship between relative solvent accessible surface area (rasa) and irregular structures in protean segments (ProSs)
 www.bioinformation.net Volume 12(9) Hypothesis The relationship between relative solvent accessible surface area (rasa) and irregular structures in protean segments (ProSs) Divya Shaji Graduate School
www.bioinformation.net Volume 12(9) Hypothesis The relationship between relative solvent accessible surface area (rasa) and irregular structures in protean segments (ProSs) Divya Shaji Graduate School
LUPAS Luminescent Polymers for in vivo Imaging of Amyloid Signatures
 LUPAS Luminescent Polymers for in vivo Imaging of Amyloid Signatures A research project for innovative diagnostics for neurodegenerative disorders Funded by the European Union under the 7 th Framework
LUPAS Luminescent Polymers for in vivo Imaging of Amyloid Signatures A research project for innovative diagnostics for neurodegenerative disorders Funded by the European Union under the 7 th Framework
Biochemistry Prof. S. DasGupta Department of Chemistry Indian Institute of Technology Kharagpur. Lecture - 5 Protein Structure - III
 Biochemistry Prof. S. DasGupta Department of Chemistry Indian Institute of Technology Kharagpur Lecture - 5 Protein Structure - III This is lecture number three on protein structure. (Refer Slide Time:
Biochemistry Prof. S. DasGupta Department of Chemistry Indian Institute of Technology Kharagpur Lecture - 5 Protein Structure - III This is lecture number three on protein structure. (Refer Slide Time:
Compound Re-Profiling. Dr Robert Scoffin CEO, Cresset
 Compound Re-Profiling Dr Robert Scoffin CEO, Cresset What is it? F F F N N H 2 N S O O Br > Compound Re-Profiling or Re-Purposing is the process of finding a new clinical use for an existing treatment
Compound Re-Profiling Dr Robert Scoffin CEO, Cresset What is it? F F F N N H 2 N S O O Br > Compound Re-Profiling or Re-Purposing is the process of finding a new clinical use for an existing treatment
STRUCTURAL BIOLOGY. α/β structures Closed barrels Open twisted sheets Horseshoe folds
 STRUCTURAL BIOLOGY α/β structures Closed barrels Open twisted sheets Horseshoe folds The α/β domains Most frequent domain structures are α/β domains: A central parallel or mixed β sheet Surrounded by α
STRUCTURAL BIOLOGY α/β structures Closed barrels Open twisted sheets Horseshoe folds The α/β domains Most frequent domain structures are α/β domains: A central parallel or mixed β sheet Surrounded by α
Supplementary Figure 1. APP cleavage assay. HEK293 cells were transfected with various
 Supplementary Figure 1. APP cleavage assay. HEK293 cells were transfected with various GST-tagged N-terminal truncated APP fragments including GST-APP full-length (FL), APP (123-695), APP (189-695), or
Supplementary Figure 1. APP cleavage assay. HEK293 cells were transfected with various GST-tagged N-terminal truncated APP fragments including GST-APP full-length (FL), APP (123-695), APP (189-695), or
Secondary Structure Prediction. Michael Tress, CNIO
 Secondary Structure Prediction Michael Tress, CNIO Why do we need to know about secondary structure? Secondary structure prediction is one important step towards deducing the 3D structure of a protein.
Secondary Structure Prediction Michael Tress, CNIO Why do we need to know about secondary structure? Secondary structure prediction is one important step towards deducing the 3D structure of a protein.
Supplementary Information
 Supplementary Information Probing Medin Monomer Structure and its Amyloid Nucleation Using 13 C-Direct Detection NMR in Combination with Structural Bioinformatics Hannah A. Davies, Daniel J. Rigden, Marie
Supplementary Information Probing Medin Monomer Structure and its Amyloid Nucleation Using 13 C-Direct Detection NMR in Combination with Structural Bioinformatics Hannah A. Davies, Daniel J. Rigden, Marie
STRUCTURE, DYNAMICS AND INTERACTIONS OF PROTEINS BY NMR SPECTROSCOPY
 STRUCTURE, DYNAMICS AND INTERACTIONS OF PROTEINS BY NMR SPECTROSCOPY Constantin T. Craescu INSERM & Institut Curie - Recherche Orsay, France A SHORT INTRODUCTION TO PROTEIN STRUCTURE Chemical composition
STRUCTURE, DYNAMICS AND INTERACTIONS OF PROTEINS BY NMR SPECTROSCOPY Constantin T. Craescu INSERM & Institut Curie - Recherche Orsay, France A SHORT INTRODUCTION TO PROTEIN STRUCTURE Chemical composition
Sequence Analysis '17 -- lecture Secondary structure 3. Sequence similarity and homology 2. Secondary structure prediction
 Sequence Analysis '17 -- lecture 16 1. Secondary structure 3. Sequence similarity and homology 2. Secondary structure prediction Alpha helix Right-handed helix. H-bond is from the oxygen at i to the nitrogen
Sequence Analysis '17 -- lecture 16 1. Secondary structure 3. Sequence similarity and homology 2. Secondary structure prediction Alpha helix Right-handed helix. H-bond is from the oxygen at i to the nitrogen
Molecular design principles underlying β-strand swapping. in the adhesive dimerization of cadherins
 Supplementary information for: Molecular design principles underlying β-strand swapping in the adhesive dimerization of cadherins Jeremie Vendome 1,2,3,5, Shoshana Posy 1,2,3,5,6, Xiangshu Jin, 1,3 Fabiana
Supplementary information for: Molecular design principles underlying β-strand swapping in the adhesive dimerization of cadherins Jeremie Vendome 1,2,3,5, Shoshana Posy 1,2,3,5,6, Xiangshu Jin, 1,3 Fabiana
From mechanism to medicne
 From mechanism to medicne a look at proteins and drug design Chem 342 δ δ δ+ M 2009 δ+ δ+ δ M Drug Design - an Iterative Approach @ DSU Structural Analysis of Receptor Structural Analysis of Ligand-Receptor
From mechanism to medicne a look at proteins and drug design Chem 342 δ δ δ+ M 2009 δ+ δ+ δ M Drug Design - an Iterative Approach @ DSU Structural Analysis of Receptor Structural Analysis of Ligand-Receptor
Bioinformatics. ONE Introduction to Biology. Sami Khuri Department of Computer Science San José State University Biology/CS 123A Fall 2012
 Bioinformatics ONE Introduction to Biology Sami Khuri Department of Computer Science San José State University Biology/CS 123A Fall 2012 Biology Review DNA RNA Proteins Central Dogma Transcription Translation
Bioinformatics ONE Introduction to Biology Sami Khuri Department of Computer Science San José State University Biology/CS 123A Fall 2012 Biology Review DNA RNA Proteins Central Dogma Transcription Translation
Increasing Protein Stability
 Increasing Protein Stability Stability depends on the extent of disulfide bond the presence of certain amino acids at the N terminus (Amino acids added to N terminus of β-galactosidase) (Table 6.4) Met,
Increasing Protein Stability Stability depends on the extent of disulfide bond the presence of certain amino acids at the N terminus (Amino acids added to N terminus of β-galactosidase) (Table 6.4) Met,
Homology Modelling. Thomas Holberg Blicher NNF Center for Protein Research University of Copenhagen
 Homology Modelling Thomas Holberg Blicher NNF Center for Protein Research University of Copenhagen Why are Protein Structures so Interesting? They provide a detailed picture of interesting biological features,
Homology Modelling Thomas Holberg Blicher NNF Center for Protein Research University of Copenhagen Why are Protein Structures so Interesting? They provide a detailed picture of interesting biological features,
Secondary Structure Prediction. Michael Tress CNIO
 Secondary Structure Prediction Michael Tress CNIO Why do we Need to Know About Secondary Structure? Secondary structure prediction is an important step towards deducing protein 3D structure. Secondary
Secondary Structure Prediction Michael Tress CNIO Why do we Need to Know About Secondary Structure? Secondary structure prediction is an important step towards deducing protein 3D structure. Secondary
Near-Native Protein Folding
 Near-Native Protein Folding Stefka Fidanova Institute for Parallel Processing at Bulgarian Academy of Science, Sofia, Bulgaria Abstract: The protein folding problem is a fundamental problem in computational
Near-Native Protein Folding Stefka Fidanova Institute for Parallel Processing at Bulgarian Academy of Science, Sofia, Bulgaria Abstract: The protein folding problem is a fundamental problem in computational
Basic concepts of molecular biology
 Basic concepts of molecular biology Gabriella Trucco Email: gabriella.trucco@unimi.it What is life made of? 1665: Robert Hooke discovered that organisms are composed of individual compartments called cells
Basic concepts of molecular biology Gabriella Trucco Email: gabriella.trucco@unimi.it What is life made of? 1665: Robert Hooke discovered that organisms are composed of individual compartments called cells
Giri Narasimhan. CAP 5510: Introduction to Bioinformatics. ECS 254; Phone: x3748
 CAP 5510: Introduction to Bioinformatics Giri Narasimhan ECS 254; Phone: x3748 giri@cis.fiu.edu www.cis.fiu.edu/~giri/teach/bioinfs07.html 2/19/07 CAP5510 1 HMM for Sequence Alignment 2/19/07 CAP5510 2
CAP 5510: Introduction to Bioinformatics Giri Narasimhan ECS 254; Phone: x3748 giri@cis.fiu.edu www.cis.fiu.edu/~giri/teach/bioinfs07.html 2/19/07 CAP5510 1 HMM for Sequence Alignment 2/19/07 CAP5510 2
Chapter Twelve Protein Synthesis: Translation of the Genetic Message
 Mary K. Campbell Shawn O. Farrell international.cengage.com/ Chapter Twelve Protein Synthesis: Translation of the Genetic Message Paul D. Adams University of Arkansas 1 Translating the Genetic Message
Mary K. Campbell Shawn O. Farrell international.cengage.com/ Chapter Twelve Protein Synthesis: Translation of the Genetic Message Paul D. Adams University of Arkansas 1 Translating the Genetic Message
Nucleic acid and protein Flow of genetic information
 Nucleic acid and protein Flow of genetic information References: Glick, BR and JJ Pasternak, 2003, Molecular Biotechnology: Principles and Applications of Recombinant DNA, ASM Press, Washington DC, pages.
Nucleic acid and protein Flow of genetic information References: Glick, BR and JJ Pasternak, 2003, Molecular Biotechnology: Principles and Applications of Recombinant DNA, ASM Press, Washington DC, pages.
Homology Modelling. Thomas Holberg Blicher NNF Center for Protein Research University of Copenhagen
 Homology Modelling Thomas Holberg Blicher NNF Center for Protein Research University of Copenhagen Why are Protein Structures so Interesting? They provide a detailed picture of interesting biological features,
Homology Modelling Thomas Holberg Blicher NNF Center for Protein Research University of Copenhagen Why are Protein Structures so Interesting? They provide a detailed picture of interesting biological features,
Protein 3D Structure Prediction
 Protein 3D Structure Prediction Michael Tress CNIO ?? MREYKLVVLGSGGVGKSALTVQFVQGIFVDE YDPTIEDSYRKQVEVDCQQCMLEILDTAGTE QFTAMRDLYMKNGQGFALVYSITAQSTFNDL QDLREQILRVKDTEDVPMILVGNKCDLEDER VVGKEQGQNLARQWCNCAFLESSAKSKINVN
Protein 3D Structure Prediction Michael Tress CNIO ?? MREYKLVVLGSGGVGKSALTVQFVQGIFVDE YDPTIEDSYRKQVEVDCQQCMLEILDTAGTE QFTAMRDLYMKNGQGFALVYSITAQSTFNDL QDLREQILRVKDTEDVPMILVGNKCDLEDER VVGKEQGQNLARQWCNCAFLESSAKSKINVN
Gene function at the level of traits Gene function at the molecular level
 Gene expression Gene function at the level of traits Gene function at the molecular level Two levels tied together since the molecular level affects the structure and function of cells which determines
Gene expression Gene function at the level of traits Gene function at the molecular level Two levels tied together since the molecular level affects the structure and function of cells which determines
BRITISH BIOMEDICAL BULLETIN
 Journal Home Page www.bbbulletin.org BRITISH BIOMEDICAL BULLETIN Original Bioinformatics Analysis of α-amylase Three-Dimensional Structure in Aspergillus oryzae Javad Sharifi-Rad,2, Fatemeh Taktaz* 3 and
Journal Home Page www.bbbulletin.org BRITISH BIOMEDICAL BULLETIN Original Bioinformatics Analysis of α-amylase Three-Dimensional Structure in Aspergillus oryzae Javad Sharifi-Rad,2, Fatemeh Taktaz* 3 and
Protein Folding in vivo. Biochemistry 530 David Baker
 Protein Folding in vivo Biochemistry 530 David Baker I. Protein folding in vivo: Molecular Chaperones II. Protein unfolding in vivo: ATP driven unfolding in mitochondrial import and the proteasome. III.
Protein Folding in vivo Biochemistry 530 David Baker I. Protein folding in vivo: Molecular Chaperones II. Protein unfolding in vivo: ATP driven unfolding in mitochondrial import and the proteasome. III.
Dr. R. Sankar, BSE 631 (2018)
 Pauling, Corey and Branson Diffraction of DNA http://www.nature.com/scitable/topicpage/dna-is-a-structure-that-encodes-biological-6493050 In short, stereochemistry is important in determining which helices
Pauling, Corey and Branson Diffraction of DNA http://www.nature.com/scitable/topicpage/dna-is-a-structure-that-encodes-biological-6493050 In short, stereochemistry is important in determining which helices
Module 3. Lecture 5. Regulation of Gene Expression in Prokaryotes
 Module 3 Lecture 5 Regulation of Gene Expression in Prokaryotes Recap So far, we have looked at prokaryotic gene regulation using 3 operon models. lac: a catabolic operon which displays induction via negative
Module 3 Lecture 5 Regulation of Gene Expression in Prokaryotes Recap So far, we have looked at prokaryotic gene regulation using 3 operon models. lac: a catabolic operon which displays induction via negative
Lac Operon contains three structural genes and is controlled by the lac repressor: (1) LacY protein transports lactose into the cell.
 Regulation of gene expression a. Expression of most genes can be turned off and on, usually by controlling the initiation of transcription. b. Lactose degradation in E. coli (Negative Control) Lac Operon
Regulation of gene expression a. Expression of most genes can be turned off and on, usually by controlling the initiation of transcription. b. Lactose degradation in E. coli (Negative Control) Lac Operon
Proteins and folding
 Proteins and folding October 8th, 2013. Kinga Futó Biopolymers Mitotic spindle of a dividing cell Actin filament network in the epidermal cell of Tobacco leaf. DNA strand released from bacteriophage 1
Proteins and folding October 8th, 2013. Kinga Futó Biopolymers Mitotic spindle of a dividing cell Actin filament network in the epidermal cell of Tobacco leaf. DNA strand released from bacteriophage 1
Alzheimer s Disease. Joachim Herz. STARS January 11, 2010
 Alzheimer s Disease Joachim Herz STARS January 11, 2010 1 The Hallmarks of Alzheimer Disease Amyloid Plaques Neurofibrillary Tangles From: Medical Library of Utah 1906 First description of Alzheimer s
Alzheimer s Disease Joachim Herz STARS January 11, 2010 1 The Hallmarks of Alzheimer Disease Amyloid Plaques Neurofibrillary Tangles From: Medical Library of Utah 1906 First description of Alzheimer s
EE550 Computational Biology
 EE550 Computational Biology Week 1 Course Notes Instructor: Bilge Karaçalı, PhD Syllabus Schedule : Thursday 13:30, 14:30, 15:30 Text : Paul G. Higgs, Teresa K. Attwood, Bioinformatics and Molecular Evolution,
EE550 Computational Biology Week 1 Course Notes Instructor: Bilge Karaçalı, PhD Syllabus Schedule : Thursday 13:30, 14:30, 15:30 Text : Paul G. Higgs, Teresa K. Attwood, Bioinformatics and Molecular Evolution,
Proteins: Wide range of func2ons. Polypep2des. Amino Acid Monomers
 Proteins: Wide range of func2ons Proteins coded in DNA account for more than 50% of the dry mass of most cells Protein func9ons structural support storage transport cellular communica9ons movement defense
Proteins: Wide range of func2ons Proteins coded in DNA account for more than 50% of the dry mass of most cells Protein func9ons structural support storage transport cellular communica9ons movement defense
Ali Yaghi. Tamara Wahbeh. Mamoun Ahram
 28 Ali Yaghi Tamara Wahbeh Mamoun Ahram This sheet is a continuation of protein purification methods. Isoelectric focusing Separation of proteins based on Isoelectric points(charge),and it is a horizontal
28 Ali Yaghi Tamara Wahbeh Mamoun Ahram This sheet is a continuation of protein purification methods. Isoelectric focusing Separation of proteins based on Isoelectric points(charge),and it is a horizontal
NORWEGIAN UNIVERSITY OF SCIENCE AND TECHNOLOGY DEPARTMENT OF BIOTECHNOLOGY Professor Bjørn E. Christensen, Department of Biotechnology
 Page 1 NRWEGIAN UNIVERSITY F SCIENCE AND TECNLGY DEPARTMENT F BITECNLGY Professor Bjørn E. Christensen, Department of Biotechnology Contact during the exam: phone: 73593327, 92634016 EXAM TBT4135 BIPLYMERS
Page 1 NRWEGIAN UNIVERSITY F SCIENCE AND TECNLGY DEPARTMENT F BITECNLGY Professor Bjørn E. Christensen, Department of Biotechnology Contact during the exam: phone: 73593327, 92634016 EXAM TBT4135 BIPLYMERS
Protein interaction network for Alzheimer's disease using computational approach
 www.bioinformation.net Hypothesis Volume 9(19) Protein interaction network for Alzheimer's disease using computational approach V Srinivasa Rao 1 *, K Srinivas 1, GN Sunand Kumar 1 & GN Sujini 2 1Department
www.bioinformation.net Hypothesis Volume 9(19) Protein interaction network for Alzheimer's disease using computational approach V Srinivasa Rao 1 *, K Srinivas 1, GN Sunand Kumar 1 & GN Sujini 2 1Department
From RNA To Protein
 From RNA To Protein 22-11-2016 Introduction mrna Processing heterogeneous nuclear RNA (hnrna) RNA that comprises transcripts of nuclear genes made by RNA polymerase II; it has a wide size distribution
From RNA To Protein 22-11-2016 Introduction mrna Processing heterogeneous nuclear RNA (hnrna) RNA that comprises transcripts of nuclear genes made by RNA polymerase II; it has a wide size distribution
Bi 8 Lecture 7. Ellen Rothenberg 26 January Reading: Ch. 3, pp ; panel 3-1
 Bi 8 Lecture 7 PROTEIN STRUCTURE, Functional analysis, and evolution Ellen Rothenberg 26 January 2016 Reading: Ch. 3, pp. 109-134; panel 3-1 (end with free amine) aromatic, hydrophobic small, hydrophilic
Bi 8 Lecture 7 PROTEIN STRUCTURE, Functional analysis, and evolution Ellen Rothenberg 26 January 2016 Reading: Ch. 3, pp. 109-134; panel 3-1 (end with free amine) aromatic, hydrophobic small, hydrophilic
Learning to Use PyMOL (includes instructions for PS #2)
 Learning to Use PyMOL (includes instructions for PS #2) To begin, download the saved PyMOL session file, 4kyz.pse from the Chem 391 Assignments web page: http://people.reed.edu/~glasfeld/chem391/assign.html
Learning to Use PyMOL (includes instructions for PS #2) To begin, download the saved PyMOL session file, 4kyz.pse from the Chem 391 Assignments web page: http://people.reed.edu/~glasfeld/chem391/assign.html
Fundamentals of Protein Structure
 Outline Fundamentals of Protein Structure Yu (Julie) Chen and Thomas Funkhouser Princeton University CS597A, Fall 2005 Protein structure Primary Secondary Tertiary Quaternary Forces and factors Levels
Outline Fundamentals of Protein Structure Yu (Julie) Chen and Thomas Funkhouser Princeton University CS597A, Fall 2005 Protein structure Primary Secondary Tertiary Quaternary Forces and factors Levels
DNA Binding Domains: Structural Motifs. Effector Domain. Zinc Fingers. Zinc Fingers, continued. Zif268
 DNA Binding Domains: Structural Motifs Studies of known transcription factors have found several motifs of protein design to allow sequence-specific binding of DNA. We will cover only three of these motifs:
DNA Binding Domains: Structural Motifs Studies of known transcription factors have found several motifs of protein design to allow sequence-specific binding of DNA. We will cover only three of these motifs:
RNA Part I: Chemical Structure of RNA
 RA Part I: Chemical Structure of RA Structural differences between RA and DA Resistance of phosphate esters to basic hydrolysis The 2 - group of RA facilitates chemical cleavage in aqueous a by forming
RA Part I: Chemical Structure of RA Structural differences between RA and DA Resistance of phosphate esters to basic hydrolysis The 2 - group of RA facilitates chemical cleavage in aqueous a by forming
IN SILICO STUDY OF MOLECULAR DYNAMICS OF HUMAN TRANSTHYRETIN VARIANTS
 Digest Journal of Nanomaterials and Biostructures Vol. 6, No 2, April - June 2011, p. 879-887 IN SILICO STUDY OF MOLECULAR DYNAMICS OF HUMAN TRANSTHYRETIN VARIANTS a, b, * MARIAN BUŢU, ALINA BUŢU a a National
Digest Journal of Nanomaterials and Biostructures Vol. 6, No 2, April - June 2011, p. 879-887 IN SILICO STUDY OF MOLECULAR DYNAMICS OF HUMAN TRANSTHYRETIN VARIANTS a, b, * MARIAN BUŢU, ALINA BUŢU a a National
Turning λ Cro into a
 Turning λ Cro into a 3 Transcriptional Activator Figure by MIT pencourseware. Fred Bushman and Mark Ptashne Cell (1988) 54:191-197 Presented by atalie Kuldell for 0.90 February 4th, 009 Small patch of
Turning λ Cro into a 3 Transcriptional Activator Figure by MIT pencourseware. Fred Bushman and Mark Ptashne Cell (1988) 54:191-197 Presented by atalie Kuldell for 0.90 February 4th, 009 Small patch of
1-D Predictions. Prediction of local features: Secondary structure & surface exposure
 Programme 8.00-8.30 Last week s quiz results 8.30-9.00 Prediction of secondary structure & surface exposure 9.00-9.20 Protein disorder prediction 9.20-9.30 Break get computers upstairs 9.30-11.00 Ex.:
Programme 8.00-8.30 Last week s quiz results 8.30-9.00 Prediction of secondary structure & surface exposure 9.00-9.20 Protein disorder prediction 9.20-9.30 Break get computers upstairs 9.30-11.00 Ex.:
Do anti-amyloid beta protein antibody cross reactivities confound Alzheimer disease research?
 Hunter and Brayne Journal of Negative Results in BioMedicine (2017) 16:1 DOI 10.1186/s12952-017-0066-3 MINI-REVIEW Open Access Do anti-amyloid beta protein antibody cross reactivities confound Alzheimer
Hunter and Brayne Journal of Negative Results in BioMedicine (2017) 16:1 DOI 10.1186/s12952-017-0066-3 MINI-REVIEW Open Access Do anti-amyloid beta protein antibody cross reactivities confound Alzheimer
from α- to β-cleavage
 Supplementary Information Specific antibody binding to the APP 672-699 region shifts APP processing from α- to β-cleavage Song Li 1,6,8, Juan Deng 1,7,8, Huayan Hou 1,8, Jun Tian 1, Brian Giunta 2,3, Yanjiang
Supplementary Information Specific antibody binding to the APP 672-699 region shifts APP processing from α- to β-cleavage Song Li 1,6,8, Juan Deng 1,7,8, Huayan Hou 1,8, Jun Tian 1, Brian Giunta 2,3, Yanjiang
Molecular Modeling Lecture 8. Local structure Database search Multiple alignment Automated homology modeling
 Molecular Modeling 2018 -- Lecture 8 Local structure Database search Multiple alignment Automated homology modeling An exception to the no-insertions-in-helix rule Actual structures (myosin)! prolines
Molecular Modeling 2018 -- Lecture 8 Local structure Database search Multiple alignment Automated homology modeling An exception to the no-insertions-in-helix rule Actual structures (myosin)! prolines
BIRKBECK COLLEGE (University of London)
 BIRKBECK COLLEGE (University of London) SCHOOL OF BIOLOGICAL SCIENCES M.Sc. EXAMINATION FOR INTERNAL STUDENTS ON: Postgraduate Certificate in Principles of Protein Structure MSc Structural Molecular Biology
BIRKBECK COLLEGE (University of London) SCHOOL OF BIOLOGICAL SCIENCES M.Sc. EXAMINATION FOR INTERNAL STUDENTS ON: Postgraduate Certificate in Principles of Protein Structure MSc Structural Molecular Biology
BACE1 ELISA Kit. Cat. No.:DEIA3191 Pkg.Size:96T. Intended use. General Description. Principle Of The Test. Reagents And Materials Provided
 BACE1 ELISA Kit Cat. No.:DEIA3191 Pkg.Size:96T Intended use This immunoassay kit allows for the use in quantitative determination of human β-site APP-Cleaving Enzyme 1, BACE1 concentrations in cell culture
BACE1 ELISA Kit Cat. No.:DEIA3191 Pkg.Size:96T Intended use This immunoassay kit allows for the use in quantitative determination of human β-site APP-Cleaving Enzyme 1, BACE1 concentrations in cell culture
About OMICS Group Conferences
 About OMICS Group OMICS Group International is an amalgamation of Open Access publications and worldwide international science conferences and events. Established in the year 2007 with the sole aim of
About OMICS Group OMICS Group International is an amalgamation of Open Access publications and worldwide international science conferences and events. Established in the year 2007 with the sole aim of
GENE EXPRESSION AT THE MOLECULAR LEVEL. Copyright (c) The McGraw-Hill Companies, Inc. Permission required for reproduction or display.
 GENE EXPRESSION AT THE MOLECULAR LEVEL Copyright (c) The McGraw-Hill Companies, Inc. Permission required for reproduction or display. 1 Gene expression Gene function at the level of traits Gene function
GENE EXPRESSION AT THE MOLECULAR LEVEL Copyright (c) The McGraw-Hill Companies, Inc. Permission required for reproduction or display. 1 Gene expression Gene function at the level of traits Gene function
Prediction of Protein-Protein Binding Sites and Epitope Mapping. Copyright 2017 Chemical Computing Group ULC All Rights Reserved.
 Prediction of Protein-Protein Binding Sites and Epitope Mapping Epitope Mapping Antibodies interact with antigens at epitopes Epitope is collection residues on antigen Continuous (sequence) or non-continuous
Prediction of Protein-Protein Binding Sites and Epitope Mapping Epitope Mapping Antibodies interact with antigens at epitopes Epitope is collection residues on antigen Continuous (sequence) or non-continuous
Bioinformatics of the Green Fluorescent Proteins
 Tested Studies for Laboratory Teaching Proceedings of the Association for Biology Laboratory Education Volume 39, Article 66, 2018 Bioinformatics of the Green Fluorescent Proteins Alma E. Rodriguez Estrada
Tested Studies for Laboratory Teaching Proceedings of the Association for Biology Laboratory Education Volume 39, Article 66, 2018 Bioinformatics of the Green Fluorescent Proteins Alma E. Rodriguez Estrada
The mechanism(s) of protein folding. What is meant by mechanism. Experimental approaches
 The mechanism(s) of protein folding What is meant by mechanism Computational approaches Experimental approaches Questions: What events occur and in what time sequence when a protein folds Is there a specified
The mechanism(s) of protein folding What is meant by mechanism Computational approaches Experimental approaches Questions: What events occur and in what time sequence when a protein folds Is there a specified
Innovative therapies for Alzheimer s disease
 WHITE PAPER IMS DISEASE INSIGHTS Innovative therapies for Alzheimer s disease New treatments will drive significant market value growth over the next ten years Author: Peter Alfinito, Ph.D., lead AnAlyst,
WHITE PAPER IMS DISEASE INSIGHTS Innovative therapies for Alzheimer s disease New treatments will drive significant market value growth over the next ten years Author: Peter Alfinito, Ph.D., lead AnAlyst,
