Green Fluorescent Protein. Avinash Bayya Varun Maturi Nikhileswar Reddy Mukkamala Aravindh Subhramani
|
|
|
- Esther Caldwell
- 6 years ago
- Views:
Transcription
1 Green Fluorescent Protein Avinash Bayya Varun Maturi Nikhileswar Reddy Mukkamala Aravindh Subhramani
2 Introduction The Green fluorescent protein (GFP) was first isolated from the Jellyfish Aequorea victoria, widely found in North America. GFP has existed for more than one hundred and sixty million years in that species. GFP is a widely studied and highly exploited protein in the fields of biochemistry and cell biology. The Jellyfish also contains a bioluminescent protein called aequorin that emits blue light. When GFP is exposed to blue light, or ultraviolet light, it emits green fluorescent light. Therefore, when the protein aequorin binds with calcium and emits blue light, this blue light is in turn absorbed by GFP, which then emits bright green light. GFP has become an amazingly useful protein in scientific research because it allows us to examine the internal workings of the cell. It is easy to locate the GFP by just exposing it to ultraviolet light, since it then glows with a bright green fluorescent light. The trick lies in attaching the GFP to any object that you are interested in examining. For instance, you can attach the GFP with another protein and examine them through the microscope thanks to GFP s fluorescence. Exploring the Structure The structure of GFP was solved in 1996 by Omro et al and Yang et al. Their structures were included in the PDB with the accession codes 1EMA and 1GFL, respectively. GFP consists of 238 amino acids with a molecular weight of 27 or 30 kd. The protein fold contains 11 anti-parallel beta strands and alpha helices. These strands and helices form a classical beta barrel structure. (Figure 1), enclosing the Ser-Tyr-Gly (chromophore) residues in order to protect them from solvent interactions. Thus, GFP is often referred to as Light in the can. Chromophore The chromophore is formed by p-hydroxybenzylideneimidazolinone formed from residues 65-67, which are Ser-Tyr-Gly of the native protein. The green shade in Figure 2 indicates the structure of the chromophore.
3 The mechanism for the formation of the chromophore takes place in four steps; folding, cyclization, dehyration and oxidation. First the denatured GFP is folded to form the native conformation with a half time of 10 min. This is followed by the cyclization, where the imidazolinone is formed by nucleophilic interaction between the amide of Gly67 and the carbonyl of residue Ser65, and the dehydration. Then, the molecular oxygen dehydrogenates the bonds of the aromatic group of residue 66 in conjunction with imidazolinone (Figure 3). At this stage the chromophore is stable and can acquire visible absorbance and fluorescence. In the chromophore there is a special interaction between glycine and threonine creating a new bond, which in turn creates an unusual five-membered ring (Figure 4). Figure 4. Chromophore illustrated in SwissPDBViewer.
4 GFP, when viewed at high resolution, offers great explanation between the protein structure and spectroscopic functions. The chromophore is situated in the middle of the GFP, totally protected from the environment. This shielding is essential for the fluorescence. When the chromophore absorbs a photon, jostling water molecule would take away the energy of the chromophore. But inside the protein, the chromophore is protected by releasing a less energetic photon of light, instead of releasing energy. Some important polar groups buried beside the chromophore are Gln69, Arg96, His148, Thr203, Ser205 and Glu222. Also, GFP s dimeric form is highly influenced by the compulsory conditions for its expression. Changes in GFP results in the release of new colors Substitutions of the amino acids in the chromophore region results in change of GFP color when exposed to blue or ultraviolet light. There are three GFP variants; blue fluorescent protein (BFP), yellow fluorescent protein (YFP) and cyan fluorescent protein (CFP). The amino acid substitutions in the GFP variants are shown below (Table 1). Table 1. Amino acid substitutions in the GFP variants. Amino acids Color GFP Green Green wtgfp Phe Ser Tyr Gly Val Gln Ser Tyr Asn Met Val Thr EGFP Leu Thr Blue EBFP Leu Thr His Phe Yellow EYFP Phe Gly Leu Ala Tyr Cyan ECFP Leu Thr Trp Ile Thr Ala The above table represents the position of the amino acid in various GFP variants. These GFP variants are more enhanced than the wild type GFP and the result is that more fluorescence can be seen in mammalians. These variants can be used as reporter genes in order to learn about protein interactions, timing of cell cycle events, and much more.
5 GFP as a biosensor GFP is used within various fields of biology like molecular biology, neuroscience and cell biology. It can be used as a reporter gene, cell marker, fusion tag, and for quantitative monitoring of gene expression. GFP s fluorescence characteristic can also be used in methods designed for sensing various levels of ions or ph. As mentioned above, there have been many codon alterations performed in GFP in order to make the expression effective in mammalians. As an application in genetic engineering, fluorescent green rabbits, rats, mice, frogs, flies, worms, and other living organisms have been produced. Below, the use of GFP as a biosensor for measuring the concentration of lead in aquatic environments will be described. Here, the GFP gene is under the control of the lead resistance regulatory gene (PbrR) and its operator promoter (PbrO/P), originating from the plasmid pmol30 of Ralstonia metallidurans. The regulatory gene and its promotor were isolated by PCR using the PbrR/O/P forward and reverse primers carrying Nhe1 and Age1 sites, respectively. The amplified PCR product was cloned into the pdb402 plasmid containing the gfp gene. The entire PbrR/O/P/GFP was excised using Nhe1 and EcoRV and inserted into the pbr322 vector, and transformed into E. coli DH5α cells. A correctly cloned and confirmed sequence was named the Pbr-GFP biosensor (Figure 5). The Pbr GFP biosensor was grown in LB broth with 100 µg/ml ampicillin at 37 O C, and streaked on to LB-amp plates with various concentrations of lead ions (Pb 2+ ). The plates were incubated at 37 O C for 16h. The fluorescence was then directly measured using the Zeiss florescence stereozoom microscope at 25 at an excitation range of nm and an emission range of nm. The expression of GFP was evaluated at the following Pb 2+ concentrations: 50, 100, 150, 200, 250, 300, 350, and 400 µm. The fluorescence showed a steady increase from 50 µm and upwards, peaked at 250 µm, and then again decreased. The fluorescence was also highest after 12 h. The following formula (based on the 12 h measurements) was derived using multiple regressions, and it can be used in order to estimate the Pb 2+ concentration in an unknown water sample with 95% accuracy.
6 C = ( ( D) + ( F) Where, C: Concentration of Pb 2+ in the sample Discussion D: Cell density F: GFP fluorescence in terms of RLU (Relative Light Unit) , and are constants. GFP can be used as a reporter gene; it shows high sensitivity, it is non-toxic, time efficient and cost effective when compared to the other classical reporter genes like lacz and GUS. Many spectral variants of GFP are now available, and therefore it is possible to label different proteins, in different colors, inside the same cell. GFP can also be used as an active tool in finding the dynamics of chromosomes in vivo. It can be used in the studies of active bacterial chromosome segregations, yeast mitosis, and centromere dynamics, by tagging it to specific chromosomes. As a last example, GFP can be used as a biosensor for detecting the lead ion concentration in water. It is easy to use this technique to estimate the concentration of any simple substance in the water that can be grown in media. References Virapong Prachayasittikul, Chartchalerm Isarankura-Na-Ayudhya, Yaneenart Suwanwong, Srisurang Tantimavanich., Construction of chimeric antibody binding green fluorescent protein for clinical application, EXCLI Journal 2005;4: Tapas Chakraborty, P. Gireesh Babu, Absar Alam and Aparna Chaudhari., GFP expressing bacterial biosensor to measure lead contamination in aquatic environment, CURRENT SCIENCE, VOL. 94, NO. 6, 25 MARCH Roger Y. Tsien., Annu. Rev. Biochem :
Janos Szabad Department of Biology University of Szeged 6720 Szeged, Somogyi str
 Janos Szabad Department of Biology University of Szeged 6720 Szeged, Somogyi str. 4. E-mail: szabad.janos@med.u-szeged.hu - Through the use of antibodies - against the protein - against a fusion partner
Janos Szabad Department of Biology University of Szeged 6720 Szeged, Somogyi str. 4. E-mail: szabad.janos@med.u-szeged.hu - Through the use of antibodies - against the protein - against a fusion partner
Bioinformatics of the Green Fluorescent Proteins
 Tested Studies for Laboratory Teaching Proceedings of the Association for Biology Laboratory Education Volume 39, Article 66, 2018 Bioinformatics of the Green Fluorescent Proteins Alma E. Rodriguez Estrada
Tested Studies for Laboratory Teaching Proceedings of the Association for Biology Laboratory Education Volume 39, Article 66, 2018 Bioinformatics of the Green Fluorescent Proteins Alma E. Rodriguez Estrada
Intracellular localization and trafficking of proteins or How (and why) to find a needle in a haystack
 Intracellular localization and trafficking of proteins or How (and why) to find a needle in a haystack :: The Structure of a Cell :: Relative sizes SUBUNITS 0.2 mm (200 μm) minimum resolvable by unaided
Intracellular localization and trafficking of proteins or How (and why) to find a needle in a haystack :: The Structure of a Cell :: Relative sizes SUBUNITS 0.2 mm (200 μm) minimum resolvable by unaided
ONTARIO SCIENCE CENTRE. Teacher Guide. Way to Glow Program
 ONTARIO SCIENCE CENTRE Teacher Guide Way to Glow Program Table of Contents Bacterial transformation background information 3 Experimental procedure 5 Expected results 7 Post-program activity sheet 8 Post-program
ONTARIO SCIENCE CENTRE Teacher Guide Way to Glow Program Table of Contents Bacterial transformation background information 3 Experimental procedure 5 Expected results 7 Post-program activity sheet 8 Post-program
ONTARIO SCIENCE CENTRE
 ONTARIO SCIENCE CENTRE Table of Contents Bacterial transformation background information 3 Experimental procedure 5 Expected results 8 Post-program activity sheet 9 Post-program activity sheet with answers
ONTARIO SCIENCE CENTRE Table of Contents Bacterial transformation background information 3 Experimental procedure 5 Expected results 8 Post-program activity sheet 9 Post-program activity sheet with answers
Comparing Sequences of Fluorescent Proteins Using BLAST (Basic Local Alignment Search Tool)
 Comparing Sequences of Fluorescent Proteins Using BLAST (Basic Local Alignment Search Tool) Mice expressing GFP under UV light (left & right), compared to normal mouse (center). Source: Wikipedia. Researcher
Comparing Sequences of Fluorescent Proteins Using BLAST (Basic Local Alignment Search Tool) Mice expressing GFP under UV light (left & right), compared to normal mouse (center). Source: Wikipedia. Researcher
Algorithms in Bioinformatics ONE Transcription Translation
 Algorithms in Bioinformatics ONE Transcription Translation Sami Khuri Department of Computer Science San José State University sami.khuri@sjsu.edu Biology Review DNA RNA Proteins Central Dogma Transcription
Algorithms in Bioinformatics ONE Transcription Translation Sami Khuri Department of Computer Science San José State University sami.khuri@sjsu.edu Biology Review DNA RNA Proteins Central Dogma Transcription
The Green Fluorescent Protein. w.chem.uwec.edu/chem412_s99/ppt/green.ppt
 The Green Fluorescent Protein w.chem.uwec.edu/chem412_s99/ppt/green.ppt www.chem.uwec.edu/chem412_s99/ppt/green.ppt Protein (gene) is from a jellyfish: Aequorea victoria www.chem.uwec.edu/chem412_s99/ppt/green.ppt
The Green Fluorescent Protein w.chem.uwec.edu/chem412_s99/ppt/green.ppt www.chem.uwec.edu/chem412_s99/ppt/green.ppt Protein (gene) is from a jellyfish: Aequorea victoria www.chem.uwec.edu/chem412_s99/ppt/green.ppt
Introduction to pglo lab
 Please take these notes carefully. You do not need to write anything in RED Introduction to pglo lab Bacteria Transformation What is a plasmid? A plasmid is a small circular piece of DNA (about 2,000 to
Please take these notes carefully. You do not need to write anything in RED Introduction to pglo lab Bacteria Transformation What is a plasmid? A plasmid is a small circular piece of DNA (about 2,000 to
Zool 3200: Cell Biology Exam 3 3/6/15
 Name: Trask Zool 3200: Cell Biology Exam 3 3/6/15 Answer each of the following questions in the space provided; circle the correct answer or answers for each multiple choice question and circle either
Name: Trask Zool 3200: Cell Biology Exam 3 3/6/15 Answer each of the following questions in the space provided; circle the correct answer or answers for each multiple choice question and circle either
Basic concepts of molecular biology
 Basic concepts of molecular biology Gabriella Trucco Email: gabriella.trucco@unimi.it Life The main actors in the chemistry of life are molecules called proteins nucleic acids Proteins: many different
Basic concepts of molecular biology Gabriella Trucco Email: gabriella.trucco@unimi.it Life The main actors in the chemistry of life are molecules called proteins nucleic acids Proteins: many different
Problem Set Unit The base ratios in the DNA and RNA for an onion (Allium cepa) are given below.
 Problem Set Unit 3 Name 1. Which molecule is found in both DNA and RNA? A. Ribose B. Uracil C. Phosphate D. Amino acid 2. Which molecules form the nucleotide marked in the diagram? A. phosphate, deoxyribose
Problem Set Unit 3 Name 1. Which molecule is found in both DNA and RNA? A. Ribose B. Uracil C. Phosphate D. Amino acid 2. Which molecules form the nucleotide marked in the diagram? A. phosphate, deoxyribose
Solutions to Problem Set 1
 MIT Department of Biology 7.014 Introductory Biology, Spring 004 Question 1 Solutions to 7.014 Problem Set 1 a) Describe the conditions of the atmosphere on prebiotic earth and how these conditions differ
MIT Department of Biology 7.014 Introductory Biology, Spring 004 Question 1 Solutions to 7.014 Problem Set 1 a) Describe the conditions of the atmosphere on prebiotic earth and how these conditions differ
How Do You Clone a Gene?
 S-20 Edvo-Kit #S-20 How Do You Clone a Gene? Experiment Objective: The objective of this experiment is to gain an understanding of the structure of DNA, a genetically engineered clone, and how genes are
S-20 Edvo-Kit #S-20 How Do You Clone a Gene? Experiment Objective: The objective of this experiment is to gain an understanding of the structure of DNA, a genetically engineered clone, and how genes are
Bacterial Transformation with pglo, Part 1
 Biology 211 Bacterial Transformation with pglo, Part 1 OBJECTIVES: Practice formulating predictions Describe the principles of bacterial transformation. Explain the procedure for gene transfer using plasmid
Biology 211 Bacterial Transformation with pglo, Part 1 OBJECTIVES: Practice formulating predictions Describe the principles of bacterial transformation. Explain the procedure for gene transfer using plasmid
7.014 Solution Set 4
 7.014 Solution Set 4 Question 1 Shown below is a fragment of the sequence of a hypothetical bacterial gene. This gene encodes production of HWDWN, protein essential for metabolizing sugar yummose. The
7.014 Solution Set 4 Question 1 Shown below is a fragment of the sequence of a hypothetical bacterial gene. This gene encodes production of HWDWN, protein essential for metabolizing sugar yummose. The
Transformation of E. coli with the Rainbow of BioBridge Fluroescent Plasmids
 Transformation of E. coli with the Rainbow of BioBridge Fluroescent Plasmids Introduction: Green fluorescent protein (GFP) was the first fluorescent protein to be discovered and manipulated for use in
Transformation of E. coli with the Rainbow of BioBridge Fluroescent Plasmids Introduction: Green fluorescent protein (GFP) was the first fluorescent protein to be discovered and manipulated for use in
11 questions for a total of 120 points
 Your Name: BYS 201, Final Exam, May 3, 2010 11 questions for a total of 120 points 1. 25 points Take a close look at these tables of amino acids. Some of them are hydrophilic, some hydrophobic, some positive
Your Name: BYS 201, Final Exam, May 3, 2010 11 questions for a total of 120 points 1. 25 points Take a close look at these tables of amino acids. Some of them are hydrophilic, some hydrophobic, some positive
In order to do transformation, the gene to be transferred is placed into a plasmid. This is done with the help of restriction enzymes, 7
 Fluorescent Protein Transformation Student Background Genetic transformation occurs when a cell takes up (i.e. takes inside) and expresses a new piece of genetic material DNA. Genetic transformation literally
Fluorescent Protein Transformation Student Background Genetic transformation occurs when a cell takes up (i.e. takes inside) and expresses a new piece of genetic material DNA. Genetic transformation literally
466 Asn (N) to Ala (A) Generate beta dimer Interface
 Table S1: Amino acid changes to the HexA α-subunit to convert the dimer interface from α to β and to introduce the putative GM2A binding surface from β- onto the α- subunit Residue position (α-numbering)
Table S1: Amino acid changes to the HexA α-subunit to convert the dimer interface from α to β and to introduce the putative GM2A binding surface from β- onto the α- subunit Residue position (α-numbering)
Lecture 25 (11/15/17)
 Lecture 25 (11/15/17) Reading: Ch9; 328-332 Ch25; 990-995, 1005-1012 Problems: Ch9 (study-guide: applying); 1,2 Ch9 (study-guide: facts); 7,8 Ch25 (text); 1-3,5-7,9,10,13-15 Ch25 (study-guide: applying);
Lecture 25 (11/15/17) Reading: Ch9; 328-332 Ch25; 990-995, 1005-1012 Problems: Ch9 (study-guide: applying); 1,2 Ch9 (study-guide: facts); 7,8 Ch25 (text); 1-3,5-7,9,10,13-15 Ch25 (study-guide: applying);
Computational Methods for Protein Structure Prediction
 Computational Methods for Protein Structure Prediction Ying Xu 2017/12/6 1 Outline introduction to protein structures the problem of protein structure prediction why it is possible to predict protein structures
Computational Methods for Protein Structure Prediction Ying Xu 2017/12/6 1 Outline introduction to protein structures the problem of protein structure prediction why it is possible to predict protein structures
Programme Good morning and summary of last week Levels of Protein Structure - I Levels of Protein Structure - II
 Programme 8.00-8.10 Good morning and summary of last week 8.10-8.30 Levels of Protein Structure - I 8.30-9.00 Levels of Protein Structure - II 9.00-9.15 Break 9.15-11.15 Exercise: Building a protein model
Programme 8.00-8.10 Good morning and summary of last week 8.10-8.30 Levels of Protein Structure - I 8.30-9.00 Levels of Protein Structure - II 9.00-9.15 Break 9.15-11.15 Exercise: Building a protein model
Alpha-helices, beta-sheets and U-turns within a protein are stabilized by (hint: two words).
 1 Quiz1 Q1 2011 Alpha-helices, beta-sheets and U-turns within a protein are stabilized by (hint: two words) Value Correct Answer 1 noncovalent interactions 100% Equals hydrogen bonds (100%) Equals H-bonds
1 Quiz1 Q1 2011 Alpha-helices, beta-sheets and U-turns within a protein are stabilized by (hint: two words) Value Correct Answer 1 noncovalent interactions 100% Equals hydrogen bonds (100%) Equals H-bonds
03-511/711 Computational Genomics and Molecular Biology, Fall
 03-511/711 Computational Genomics and Molecular Biology, Fall 2011 1 Problem Set 0 Due Tuesday, September 6th This homework is intended to be a self-administered placement quiz, to help you (and me) determine
03-511/711 Computational Genomics and Molecular Biology, Fall 2011 1 Problem Set 0 Due Tuesday, September 6th This homework is intended to be a self-administered placement quiz, to help you (and me) determine
DNA.notebook March 08, DNA Overview
 DNA Overview Deoxyribonucleic Acid, or DNA, must be able to do 2 things: 1) give instructions for building and maintaining cells. 2) be copied each time a cell divides. DNA is made of subunits called nucleotides
DNA Overview Deoxyribonucleic Acid, or DNA, must be able to do 2 things: 1) give instructions for building and maintaining cells. 2) be copied each time a cell divides. DNA is made of subunits called nucleotides
 Problem: The GC base pairs are more stable than AT base pairs. Why? 5. Triple-stranded DNA was first observed in 1957. Scientists later discovered that the formation of triplestranded DNA involves a type
Problem: The GC base pairs are more stable than AT base pairs. Why? 5. Triple-stranded DNA was first observed in 1957. Scientists later discovered that the formation of triplestranded DNA involves a type
7.014 Problem Set 4 Answers to this problem set are to be turned in. Problem sets will not be accepted late. Solutions will be posted on the web.
 MIT Department of Biology 7.014 Introductory Biology, Spring 2005 Name: Section : 7.014 Problem Set 4 Answers to this problem set are to be turned in. Problem sets will not be accepted late. Solutions
MIT Department of Biology 7.014 Introductory Biology, Spring 2005 Name: Section : 7.014 Problem Set 4 Answers to this problem set are to be turned in. Problem sets will not be accepted late. Solutions
Solutions to 7.02 Quiz II 10/27/05
 Solutions to 7.02 Quiz II 10/27/05 Class Average = 83 Standard Deviation = 9 Range Grade % 87-100 A 43 74-86 B 39 55-73 C 17 > 54 D 1 Question 1 (56 points) While studying deep sea bacteria, you discover
Solutions to 7.02 Quiz II 10/27/05 Class Average = 83 Standard Deviation = 9 Range Grade % 87-100 A 43 74-86 B 39 55-73 C 17 > 54 D 1 Question 1 (56 points) While studying deep sea bacteria, you discover
Supporting Information
 Supporting Information Wiley-VCH 2007 69451 Weinheim, Germany A Biosynthetic Route to Dehydroalanine Containing Proteins Jiangyun Wang, Stefan M. Schiller and Peter G. Schultz* Department of Chemistry
Supporting Information Wiley-VCH 2007 69451 Weinheim, Germany A Biosynthetic Route to Dehydroalanine Containing Proteins Jiangyun Wang, Stefan M. Schiller and Peter G. Schultz* Department of Chemistry
Illumatool ΤΜ Tunable Light System: A Non-Destructive Light Source For Molecular And Cellular Biology Applications. John Fox, Lightools Research.
 Illumatool ΤΜ Tunable Light System: A Non-Destructive Light Source For Molecular And Cellular Biology Applications. John Fox, Lightools Research. Fluorescent dyes and proteins are basic analytical tools
Illumatool ΤΜ Tunable Light System: A Non-Destructive Light Source For Molecular And Cellular Biology Applications. John Fox, Lightools Research. Fluorescent dyes and proteins are basic analytical tools
In silico measurements of twist and bend. moduli for beta solenoid protein self-
 In silico measurements of twist and bend moduli for beta solenoid protein self- assembly units Leonard P. Heinz, Krishnakumar M. Ravikumar, and Daniel L. Cox Department of Physics and Institute for Complex
In silico measurements of twist and bend moduli for beta solenoid protein self- assembly units Leonard P. Heinz, Krishnakumar M. Ravikumar, and Daniel L. Cox Department of Physics and Institute for Complex
Packing of Secondary Structures
 7.88 Lecture Notes - 5 7.24/7.88J/5.48J The Protein Folding and Human Disease Packing of Secondary Structures Packing of Helices against sheets Packing of sheets against sheets Parallel Orthogonal Table:
7.88 Lecture Notes - 5 7.24/7.88J/5.48J The Protein Folding and Human Disease Packing of Secondary Structures Packing of Helices against sheets Packing of sheets against sheets Parallel Orthogonal Table:
Green Fluorescent Protein (GFP) Purification. Hydrophobic Interaction Chromatography
 Green Fluorescent Protein (GFP) Purification Hydrophobic Interaction Chromatography What is the GFP gene? GFP is a green fluorescent protein that is normally found in jellyfish. It has been engineered
Green Fluorescent Protein (GFP) Purification Hydrophobic Interaction Chromatography What is the GFP gene? GFP is a green fluorescent protein that is normally found in jellyfish. It has been engineered
Assignment 13. In the Griffith experiment, why did mice die when injected with live R bacteria plus heatkilled
 Assignment 13 1. Multiple-choice (1 point) In the Griffith experiment, why did mice die when injected with live R bacteria plus heatkilled S bacteria? Some of the S bacteria were still alive. The R bacteria
Assignment 13 1. Multiple-choice (1 point) In the Griffith experiment, why did mice die when injected with live R bacteria plus heatkilled S bacteria? Some of the S bacteria were still alive. The R bacteria
6-Foot Mini Toober Activity
 Big Idea The interaction between the substrate and enzyme is highly specific. Even a slight change in shape of either the substrate or the enzyme may alter the efficient and selective ability of the enzyme
Big Idea The interaction between the substrate and enzyme is highly specific. Even a slight change in shape of either the substrate or the enzyme may alter the efficient and selective ability of the enzyme
7.014 Quiz II Handout
 7.014 Quiz II Handout Quiz II: Wednesday, March 17 12:05-12:55 54-100 **This will be a closed book exam** Quiz Review Session: Friday, March 12 7:00-9:00 pm room 54-100 Open Tutoring Session: Tuesday,
7.014 Quiz II Handout Quiz II: Wednesday, March 17 12:05-12:55 54-100 **This will be a closed book exam** Quiz Review Session: Friday, March 12 7:00-9:00 pm room 54-100 Open Tutoring Session: Tuesday,
Name Section Problem Set 3
 Name Section 7.013 Problem Set 3 The completed problem must be turned into the wood box outside 68120 by 4:40 pm, Thursday, March 13. Problem sets will not be accepted late. Question 1 Based upon your
Name Section 7.013 Problem Set 3 The completed problem must be turned into the wood box outside 68120 by 4:40 pm, Thursday, March 13. Problem sets will not be accepted late. Question 1 Based upon your
Fundamentals of Protein Structure
 Outline Fundamentals of Protein Structure Yu (Julie) Chen and Thomas Funkhouser Princeton University CS597A, Fall 2005 Protein structure Primary Secondary Tertiary Quaternary Forces and factors Levels
Outline Fundamentals of Protein Structure Yu (Julie) Chen and Thomas Funkhouser Princeton University CS597A, Fall 2005 Protein structure Primary Secondary Tertiary Quaternary Forces and factors Levels
CHAPTER 20 DNA TECHNOLOGY AND GENOMICS. Section A: DNA Cloning
 Section A: DNA Cloning 1. DNA technology makes it possible to clone genes for basic research and commercial applications: an overview 2. Restriction enzymes are used to make recombinant DNA 3. Genes can
Section A: DNA Cloning 1. DNA technology makes it possible to clone genes for basic research and commercial applications: an overview 2. Restriction enzymes are used to make recombinant DNA 3. Genes can
TECHNICAL BULLETIN. In Vitro Bacterial Split Fluorescent Protein Fold n Glow Solubility Assay Kits
 In Vitro Bacterial Split Fluorescent Protein Fold n Glow Solubility Assay Kits Catalog Numbers APPA001 In Vitro Bacterial Split GFP "Fold 'n' Glow" Solubility Assay Kit (Green) APPA008 In Vitro Bacterial
In Vitro Bacterial Split Fluorescent Protein Fold n Glow Solubility Assay Kits Catalog Numbers APPA001 In Vitro Bacterial Split GFP "Fold 'n' Glow" Solubility Assay Kit (Green) APPA008 In Vitro Bacterial
Supporting information
 Supporting information Construction of strains and plasmids To create ptc67, a PCR product obtained with primers cc2570-162f (gcatgggcaagcttgaggacggcgtcatgt) and cc2570+512f (gaggccgtggtaccatagaggcgggcg),
Supporting information Construction of strains and plasmids To create ptc67, a PCR product obtained with primers cc2570-162f (gcatgggcaagcttgaggacggcgtcatgt) and cc2570+512f (gaggccgtggtaccatagaggcgggcg),
Nucleic acid and protein Flow of genetic information
 Nucleic acid and protein Flow of genetic information References: Glick, BR and JJ Pasternak, 2003, Molecular Biotechnology: Principles and Applications of Recombinant DNA, ASM Press, Washington DC, pages.
Nucleic acid and protein Flow of genetic information References: Glick, BR and JJ Pasternak, 2003, Molecular Biotechnology: Principles and Applications of Recombinant DNA, ASM Press, Washington DC, pages.
Student Manual. pglo Transformation
 Student Manual pglo Transformation STUDENT MANUAL LESSON 1 Lesson 1 Introduction to Transformation In this lab you will perform a procedure known as genetic transformation. Remember that a gene is a piece
Student Manual pglo Transformation STUDENT MANUAL LESSON 1 Lesson 1 Introduction to Transformation In this lab you will perform a procedure known as genetic transformation. Remember that a gene is a piece
Basic concepts of molecular biology
 Basic concepts of molecular biology Gabriella Trucco Email: gabriella.trucco@unimi.it What is life made of? 1665: Robert Hooke discovered that organisms are composed of individual compartments called cells
Basic concepts of molecular biology Gabriella Trucco Email: gabriella.trucco@unimi.it What is life made of? 1665: Robert Hooke discovered that organisms are composed of individual compartments called cells
Protein Structure Analysis
 BINF 731 Protein Structure Analysis http://binf.gmu.edu/vaisman/binf731/ Secondary Structure: Computational Problems Secondary structure characterization Secondary structure assignment Secondary structure
BINF 731 Protein Structure Analysis http://binf.gmu.edu/vaisman/binf731/ Secondary Structure: Computational Problems Secondary structure characterization Secondary structure assignment Secondary structure
Expression of Recombinant Proteins
 Expression of Recombinant Proteins Uses of Cloned Genes sequencing reagents (eg, probes) protein production insufficient natural quantities modify/mutagenesis library screening Expression Vector Features
Expression of Recombinant Proteins Uses of Cloned Genes sequencing reagents (eg, probes) protein production insufficient natural quantities modify/mutagenesis library screening Expression Vector Features
1/4/18 NUCLEIC ACIDS. Nucleic Acids. Nucleic Acids. ECS129 Instructor: Patrice Koehl
 NUCLEIC ACIDS ECS129 Instructor: Patrice Koehl Nucleic Acids Nucleotides DNA Structure RNA Synthesis Function Secondary structure Tertiary interactions Wobble hypothesis DNA RNA Replication Transcription
NUCLEIC ACIDS ECS129 Instructor: Patrice Koehl Nucleic Acids Nucleotides DNA Structure RNA Synthesis Function Secondary structure Tertiary interactions Wobble hypothesis DNA RNA Replication Transcription
NUCLEIC ACIDS. ECS129 Instructor: Patrice Koehl
 NUCLEIC ACIDS ECS129 Instructor: Patrice Koehl Nucleic Acids Nucleotides DNA Structure RNA Synthesis Function Secondary structure Tertiary interactions Wobble hypothesis DNA RNA Replication Transcription
NUCLEIC ACIDS ECS129 Instructor: Patrice Koehl Nucleic Acids Nucleotides DNA Structure RNA Synthesis Function Secondary structure Tertiary interactions Wobble hypothesis DNA RNA Replication Transcription
Case 7 A Storage Protein From Seeds of Brassica nigra is a Serine Protease Inhibitor Last modified 29 September 2005
 Case 7 A Storage Protein From Seeds of Brassica nigra is a Serine Protease Inhibitor Last modified 9 September 005 Focus concept Purification of a novel seed storage protein allows sequence analysis and
Case 7 A Storage Protein From Seeds of Brassica nigra is a Serine Protease Inhibitor Last modified 9 September 005 Focus concept Purification of a novel seed storage protein allows sequence analysis and
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Figure 1. Analysis of unique sequences isolated in Sort 14. Unique sequences are listed next to their frequency of occurrence, in parenthesis. Dithiol resistance is
Supplementary Figure 1 Supplementary Figure 1. Analysis of unique sequences isolated in Sort 14. Unique sequences are listed next to their frequency of occurrence, in parenthesis. Dithiol resistance is
Bioinformatics. ONE Introduction to Biology. Sami Khuri Department of Computer Science San José State University Biology/CS 123A Fall 2012
 Bioinformatics ONE Introduction to Biology Sami Khuri Department of Computer Science San José State University Biology/CS 123A Fall 2012 Biology Review DNA RNA Proteins Central Dogma Transcription Translation
Bioinformatics ONE Introduction to Biology Sami Khuri Department of Computer Science San José State University Biology/CS 123A Fall 2012 Biology Review DNA RNA Proteins Central Dogma Transcription Translation
CFSSP: Chou and Fasman Secondary Structure Prediction server
 Wide Spectrum, Vol. 1, No. 9, (2013) pp 15-19 CFSSP: Chou and Fasman Secondary Structure Prediction server T. Ashok Kumar Department of Bioinformatics, Noorul Islam College of Arts and Science, Kumaracoil
Wide Spectrum, Vol. 1, No. 9, (2013) pp 15-19 CFSSP: Chou and Fasman Secondary Structure Prediction server T. Ashok Kumar Department of Bioinformatics, Noorul Islam College of Arts and Science, Kumaracoil
First&year&tutorial&in&Chemical&Biology&(amino&acids,&peptide&and&proteins)&! 1.&!
 First&year&tutorial&in&Chemical&Biology&(amino&acids,&peptide&and&proteins& 1.& a. b. c. d. e. 2.& a. b. c. d. e. f. & UsingtheCahn Ingold Prelogsystem,assignstereochemicaldescriptorstothe threeaminoacidsshownbelow.
First&year&tutorial&in&Chemical&Biology&(amino&acids,&peptide&and&proteins& 1.& a. b. c. d. e. 2.& a. b. c. d. e. f. & UsingtheCahn Ingold Prelogsystem,assignstereochemicaldescriptorstothe threeaminoacidsshownbelow.
7.013 Problem Set 3 FRIDAY October 8th, 2004
 MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert. Weinberg, Dr. laudette ardel Name: T: 7.013 Problem Set 3 FRIDY October 8th, 2004 Problem
MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert. Weinberg, Dr. laudette ardel Name: T: 7.013 Problem Set 3 FRIDY October 8th, 2004 Problem
Supplementary Table 1: List of CH3 domain interface residues in the first chain (A) and
 Supplementary Tables Supplementary Table 1: List of CH3 domain interface residues in the first chain (A) and their side chain contacting residues in the second chain (B) a Interface Res. in Contacting
Supplementary Tables Supplementary Table 1: List of CH3 domain interface residues in the first chain (A) and their side chain contacting residues in the second chain (B) a Interface Res. in Contacting
Thr Gly Tyr. Gly Lys Asn
 Your unique body characteristics (traits), such as hair color or blood type, are determined by the proteins your body produces. Proteins are the building blocks of life - in fact, about 45% of the human
Your unique body characteristics (traits), such as hair color or blood type, are determined by the proteins your body produces. Proteins are the building blocks of life - in fact, about 45% of the human
Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid
 Supplementary Figures Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid sequence of Drosophila RCC1. Same colors are for Figure 1 with sequence of β-wedge that interacts with Ran in
Supplementary Figures Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid sequence of Drosophila RCC1. Same colors are for Figure 1 with sequence of β-wedge that interacts with Ran in
NAME TA SEC Problem Set 3 FRIDAY March 5, Problem sets will NOT be accepted late.
 MIT Department of Biology 7.013: Introductory Biology - Spring 2004 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. laudette ardel NME T SE 7.013 Problem Set 3 FRIDY March 5, 2004 Problem
MIT Department of Biology 7.013: Introductory Biology - Spring 2004 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. laudette ardel NME T SE 7.013 Problem Set 3 FRIDY March 5, 2004 Problem
MBMB451A Section1 Fall 2008 KEY These questions may have more than one correct answer
 MBMB451A Section1 Fall 2008 KEY These questions may have more than one correct answer 1. In a double stranded molecule of DNA, the ratio of purines : pyrimidines is (a) variable (b) determined by the base
MBMB451A Section1 Fall 2008 KEY These questions may have more than one correct answer 1. In a double stranded molecule of DNA, the ratio of purines : pyrimidines is (a) variable (b) determined by the base
Supplementary Information
 Supplementary Information Fluorescent protein choice The dual fluorescence channel ratiometric promoter characterization plasmid we designed for this study required a pair of fluorescent proteins with
Supplementary Information Fluorescent protein choice The dual fluorescence channel ratiometric promoter characterization plasmid we designed for this study required a pair of fluorescent proteins with
NAD + + H 2 O C (+1) 2 e -
 Bioluminescence Bioluminescence is the biochemical emission of visible light by living organisms. As such, it is a special category of chemiluminescence. (Do not confuse bioluminescence with phosphorescence,
Bioluminescence Bioluminescence is the biochemical emission of visible light by living organisms. As such, it is a special category of chemiluminescence. (Do not confuse bioluminescence with phosphorescence,
Bi Lecture 3 Loss-of-function (Ch. 4A) Monday, April 8, 13
 Bi190-2013 Lecture 3 Loss-of-function (Ch. 4A) Infer Gene activity from type of allele Loss-of-Function alleles are Gold Standard If organism deficient in gene A fails to accomplish process B, then gene
Bi190-2013 Lecture 3 Loss-of-function (Ch. 4A) Infer Gene activity from type of allele Loss-of-Function alleles are Gold Standard If organism deficient in gene A fails to accomplish process B, then gene
Case 7 A Storage Protein From Seeds of Brassica nigra is a Serine Protease Inhibitor
 Case 7 A Storage Protein From Seeds of Brassica nigra is a Serine Protease Inhibitor Focus concept Purification of a novel seed storage protein allows sequence analysis and determination of the protein
Case 7 A Storage Protein From Seeds of Brassica nigra is a Serine Protease Inhibitor Focus concept Purification of a novel seed storage protein allows sequence analysis and determination of the protein
Structure formation and association of biomolecules. Prof. Dr. Martin Zacharias Lehrstuhl für Molekulardynamik (T38) Technische Universität München
 Structure formation and association of biomolecules Prof. Dr. Martin Zacharias Lehrstuhl für Molekulardynamik (T38) Technische Universität München Motivation Many biomolecules are chemically synthesized
Structure formation and association of biomolecules Prof. Dr. Martin Zacharias Lehrstuhl für Molekulardynamik (T38) Technische Universität München Motivation Many biomolecules are chemically synthesized
36. The double bonds in naturally-occuring fatty acids are usually isomers. A. cis B. trans C. both cis and trans D. D- E. L-
 36. The double bonds in naturally-occuring fatty acids are usually isomers. A. cis B. trans C. both cis and trans D. D- E. L- 37. The essential fatty acids are A. palmitic acid B. linoleic acid C. linolenic
36. The double bonds in naturally-occuring fatty acids are usually isomers. A. cis B. trans C. both cis and trans D. D- E. L- 37. The essential fatty acids are A. palmitic acid B. linoleic acid C. linolenic
Product Manual. GFP ELISA Kit. Catalog Number. FOR RESEARCH USE ONLY Not for use in diagnostic procedures
 Product Manual GFP ELISA Kit Catalog Number AKR-121 96 assays FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Green fluorescent protein (GFP) is a spontaneously fluorescent protein
Product Manual GFP ELISA Kit Catalog Number AKR-121 96 assays FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Green fluorescent protein (GFP) is a spontaneously fluorescent protein
Figure 1. Map of cloning vector pgem T-Easy (bacterial plasmid DNA)
 Texas A&M University-Corpus Christi CHEM4402 Biochemistry II Laboratory Laboratory 6: Ligation & Bacterial Transformation (Bring your text and laptop to class if you wish to work on your assignment during
Texas A&M University-Corpus Christi CHEM4402 Biochemistry II Laboratory Laboratory 6: Ligation & Bacterial Transformation (Bring your text and laptop to class if you wish to work on your assignment during
MOLEBIO LAB #3: Electrophoretic Separation of Proteins
 MOLEBIO LAB #3: Electrophoretic Separation of Proteins Introduction: Proteins occupy a central position in the structure and function of all living organisms. Some proteins serve as structural components
MOLEBIO LAB #3: Electrophoretic Separation of Proteins Introduction: Proteins occupy a central position in the structure and function of all living organisms. Some proteins serve as structural components
Dr. R. Sankar, BSE 631 (2018)
 Pauling, Corey and Branson Diffraction of DNA http://www.nature.com/scitable/topicpage/dna-is-a-structure-that-encodes-biological-6493050 In short, stereochemistry is important in determining which helices
Pauling, Corey and Branson Diffraction of DNA http://www.nature.com/scitable/topicpage/dna-is-a-structure-that-encodes-biological-6493050 In short, stereochemistry is important in determining which helices
Module Code: BIO00007C
 Examination Candidate Number: Desk Number: BSc and MSc Degree Examinations 2018-9 Department : BIOLOGY Title of Exam: Genetics Time Allowed: 1 Hour 30 Minutes Marking Scheme: Total marks available for
Examination Candidate Number: Desk Number: BSc and MSc Degree Examinations 2018-9 Department : BIOLOGY Title of Exam: Genetics Time Allowed: 1 Hour 30 Minutes Marking Scheme: Total marks available for
Fluorescent protein. Origin of fluorescent proteins. Structural and biophysical analysis of FPs. Application in molecular biology and biotechnology
 Origin of fluorescent proteins types of FPs mechanism of fluorescence Fluorescent protein Structural and biophysical analysis of FPs Application in molecular biology and biotechnology Variety of organisms
Origin of fluorescent proteins types of FPs mechanism of fluorescence Fluorescent protein Structural and biophysical analysis of FPs Application in molecular biology and biotechnology Variety of organisms
Chapter 20 Recombinant DNA Technology. Copyright 2009 Pearson Education, Inc.
 Chapter 20 Recombinant DNA Technology Copyright 2009 Pearson Education, Inc. 20.1 Recombinant DNA Technology Began with Two Key Tools: Restriction Enzymes and DNA Cloning Vectors Recombinant DNA refers
Chapter 20 Recombinant DNA Technology Copyright 2009 Pearson Education, Inc. 20.1 Recombinant DNA Technology Began with Two Key Tools: Restriction Enzymes and DNA Cloning Vectors Recombinant DNA refers
Ch Biophysical Chemistry
 Ch 247.53. Biophysical Chemistry Nina Rosario L. Rojas 2012-2013 sem 1 Review of Protein Structure Why structure? Primary, secondary, tertiary structure Disulfide bonds scheme 2 STRUCTURE- REGULAR STRUCTURE
Ch 247.53. Biophysical Chemistry Nina Rosario L. Rojas 2012-2013 sem 1 Review of Protein Structure Why structure? Primary, secondary, tertiary structure Disulfide bonds scheme 2 STRUCTURE- REGULAR STRUCTURE
7.1 Techniques for Producing and Analyzing DNA. SBI4U Ms. Ho-Lau
 7.1 Techniques for Producing and Analyzing DNA SBI4U Ms. Ho-Lau What is Biotechnology? From Merriam-Webster: the manipulation of living organisms or their components to produce useful usually commercial
7.1 Techniques for Producing and Analyzing DNA SBI4U Ms. Ho-Lau What is Biotechnology? From Merriam-Webster: the manipulation of living organisms or their components to produce useful usually commercial
FLUORESCENCE. Matyas Molnar and Dirk Pacholsky
 FLUORESCENCE Matyas Molnar and Dirk Pacholsky 1 Information This lecture contains images and information from the following internet homepages http://micro.magnet.fsu.edu/primer/index.html http://www.microscopyu.com/
FLUORESCENCE Matyas Molnar and Dirk Pacholsky 1 Information This lecture contains images and information from the following internet homepages http://micro.magnet.fsu.edu/primer/index.html http://www.microscopyu.com/
1. DNA replication. (a) Why is DNA replication an essential process?
 ame Section 7.014 Problem Set 3 Please print out this problem set and record your answers on the printed copy. Answers to this problem set are to be turned in to the box outside 68120 by 5:00pm on Friday
ame Section 7.014 Problem Set 3 Please print out this problem set and record your answers on the printed copy. Answers to this problem set are to be turned in to the box outside 68120 by 5:00pm on Friday
Texas A&M University-Corpus Christi CHEM4402 Biochemistry II Laboratory Laboratory 4 - Polymerase Chain Reaction (PCR)
 Texas A&M University-Corpus Christi CHEM4402 Biochemistry II Laboratory Laboratory 4 - Polymerase Chain Reaction (PCR) Progressing with the sequence of experiments, we are now ready to amplify the green
Texas A&M University-Corpus Christi CHEM4402 Biochemistry II Laboratory Laboratory 4 - Polymerase Chain Reaction (PCR) Progressing with the sequence of experiments, we are now ready to amplify the green
Molecular Biology. Biology Review ONE. Protein Factory. Genotype to Phenotype. From DNA to Protein. DNA à RNA à Protein. June 2016
 Molecular Biology ONE Sami Khuri Department of Computer Science San José State University Biology Review DNA RNA Proteins Central Dogma Transcription Translation Genotype to Phenotype Protein Factory DNA
Molecular Biology ONE Sami Khuri Department of Computer Science San José State University Biology Review DNA RNA Proteins Central Dogma Transcription Translation Genotype to Phenotype Protein Factory DNA
THE GREEN FLUORESCENT PROTEIN
 Annu. Rev. Biochem. 1998. 67:509 44 Copyright c 1998 by Annual Reviews. All rights reserved THE GREEN FLUORESCENT PROTEIN Roger Y. Tsien Howard Hughes Medical Institute; University of California, San Diego;
Annu. Rev. Biochem. 1998. 67:509 44 Copyright c 1998 by Annual Reviews. All rights reserved THE GREEN FLUORESCENT PROTEIN Roger Y. Tsien Howard Hughes Medical Institute; University of California, San Diego;
The Central Dogma. 1) A simplified model. 2) Played out across life. 3) Many points for control.
 Molecular Biology The Central Dogma 1) A simplified model. 2) Played out across life. 3) Many dis@nct points for control. Molecular Biology DNA: four nucleo@de bases (GC,AT) (2 bits) gene@c code in 3 base
Molecular Biology The Central Dogma 1) A simplified model. 2) Played out across life. 3) Many dis@nct points for control. Molecular Biology DNA: four nucleo@de bases (GC,AT) (2 bits) gene@c code in 3 base
DNA/Protein Binding, Molecular Docking and in Vitro Anti-cancer Activity of some Thioether-Dipyrrinato Complexes
 DNA/Protein Binding, Molecular Docking and in Vitro Anti-cancer Activity of some Thioether-Dipyrrinato Complexes Rakesh Kumar Gupta, Gunjan Sharma, ξ Rampal Pandey, Amit Kumar, Biplob Koch, ξ Pei- Zhou
DNA/Protein Binding, Molecular Docking and in Vitro Anti-cancer Activity of some Thioether-Dipyrrinato Complexes Rakesh Kumar Gupta, Gunjan Sharma, ξ Rampal Pandey, Amit Kumar, Biplob Koch, ξ Pei- Zhou
Challenges to measuring intracellular Ca 2+ Calmodulin: nature s Ca 2+ sensor
 Calcium Signals in Biological Systems Lecture 3 (2/9/0) Measuring intracellular Ca 2+ signals II: Genetically encoded Ca 2+ sensors Henry M. Colecraft, Ph.D. Challenges to measuring intracellular Ca 2+
Calcium Signals in Biological Systems Lecture 3 (2/9/0) Measuring intracellular Ca 2+ signals II: Genetically encoded Ca 2+ sensors Henry M. Colecraft, Ph.D. Challenges to measuring intracellular Ca 2+
Classroom Tested Lesson
 Classroom Tested Lesson Video Description Secrets of the Sequence, Show 124, Episode 2 A Green Light for Biology approximately 10 minutes viewing time This discovery known as Green Fluorescent Protein
Classroom Tested Lesson Video Description Secrets of the Sequence, Show 124, Episode 2 A Green Light for Biology approximately 10 minutes viewing time This discovery known as Green Fluorescent Protein
Supplementary Materials: A Toolbox of Genetically Encoded FRET-Based Biosensors for Rapid L-Lysine Analysis
 S1 of S9 Supplementary Materials: A Toolbox of Genetically Encoded FRET-Based Biosensors for Rapid L-Lysine Analysis Victoria Steffen, Julia Otten, Susann Engelmann, Andreas Radek, Michael Limberg, Bernd
S1 of S9 Supplementary Materials: A Toolbox of Genetically Encoded FRET-Based Biosensors for Rapid L-Lysine Analysis Victoria Steffen, Julia Otten, Susann Engelmann, Andreas Radek, Michael Limberg, Bernd
7.013 Spring 2005 Problem Set 1
 MIT Department of Biology 7.013: Introductory Biology Spring 005 Instructors: rofessor azel Sive, rofessor Tyler Jacks, Dr. laudette Gardel AME TA Section # 7.013 Spring 005 roblem Set 1 FRIDAY February
MIT Department of Biology 7.013: Introductory Biology Spring 005 Instructors: rofessor azel Sive, rofessor Tyler Jacks, Dr. laudette Gardel AME TA Section # 7.013 Spring 005 roblem Set 1 FRIDAY February
1.1 Photosensing in biological systems
 Chapter 1 Introduction 1.1 Photosensing in biological systems The absorption of visible light and its conversion to other forms of energy is at the core of some of the most fundamental processes in biology.
Chapter 1 Introduction 1.1 Photosensing in biological systems The absorption of visible light and its conversion to other forms of energy is at the core of some of the most fundamental processes in biology.
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature06147 SUPPLEMENTARY INFORMATION Figure S1 The genomic and domain structure of Dscam. The Dscam gene comprises 24 exons, encoding a signal peptide (SP), 10 IgSF domains, 6 fibronectin
doi: 10.1038/nature06147 SUPPLEMENTARY INFORMATION Figure S1 The genomic and domain structure of Dscam. The Dscam gene comprises 24 exons, encoding a signal peptide (SP), 10 IgSF domains, 6 fibronectin
STRUCTURAL BIOLOGY. α/β structures Closed barrels Open twisted sheets Horseshoe folds
 STRUCTURAL BIOLOGY α/β structures Closed barrels Open twisted sheets Horseshoe folds The α/β domains Most frequent domain structures are α/β domains: A central parallel or mixed β sheet Surrounded by α
STRUCTURAL BIOLOGY α/β structures Closed barrels Open twisted sheets Horseshoe folds The α/β domains Most frequent domain structures are α/β domains: A central parallel or mixed β sheet Surrounded by α
Comparison of different methods for purification analysis of a green fluorescent Strep-tag fusion protein. Application
 Comparison of different methods for purification analysis of a green fluorescent Strep-tag fusion protein Application Petra Sebastian Meike Kuschel Stefan Schmidt Abstract This Application Note describes
Comparison of different methods for purification analysis of a green fluorescent Strep-tag fusion protein Application Petra Sebastian Meike Kuschel Stefan Schmidt Abstract This Application Note describes
5. Which of the following enzymes catalyze the attachment of an amino acid to trna in the formation of aminoacyl trna?
 Sample Examination Questions for Exam 3 Material Biology 3300 / Dr. Jerald Hendrix Warning! These questions are posted solely to provide examples of past test questions. There is no guarantee that any
Sample Examination Questions for Exam 3 Material Biology 3300 / Dr. Jerald Hendrix Warning! These questions are posted solely to provide examples of past test questions. There is no guarantee that any
蛋白質體學. Proteomics Amino acids, Peptides and Proteins 陳威戎 & 21
 蛋白質體學 Proteomics 2015 Amino acids, Peptides and Proteins 陳威戎 2015. 09. 14 & 21 Outline 1. Amino Acids 2. Peptides and Proteins 3. Covalent Structure of Proteins Amino Acids Proteins are polymers of amino
蛋白質體學 Proteomics 2015 Amino acids, Peptides and Proteins 陳威戎 2015. 09. 14 & 21 Outline 1. Amino Acids 2. Peptides and Proteins 3. Covalent Structure of Proteins Amino Acids Proteins are polymers of amino
Student Manual. pglo Transformation
 Student Manual pglo Transformation Lesson 1 Introduction to Transformation In this lab you will perform a procedure known as genetic transformation. Remember that a gene is a piece of DNA which provides
Student Manual pglo Transformation Lesson 1 Introduction to Transformation In this lab you will perform a procedure known as genetic transformation. Remember that a gene is a piece of DNA which provides
Materials Protein synthesis kit. This kit consists of 24 amino acids, 24 transfer RNAs, four messenger RNAs and one ribosome (see below).
 Protein Synthesis Instructions The purpose of today s lab is to: Understand how a cell manufactures proteins from amino acids, using information stored in the genetic code. Assemble models of four very
Protein Synthesis Instructions The purpose of today s lab is to: Understand how a cell manufactures proteins from amino acids, using information stored in the genetic code. Assemble models of four very
Supplementary Figure 1
 Supplementary Figure 1 2 Supplementary Figure 1: Sequence alignment of HsHSD17B8 and HsCBR4 of with KAR orthologs. The secondary structure elements as calculated by DSSP and residue numbers are displayed
Supplementary Figure 1 2 Supplementary Figure 1: Sequence alignment of HsHSD17B8 and HsCBR4 of with KAR orthologs. The secondary structure elements as calculated by DSSP and residue numbers are displayed
Supplementary Online Material. An Expanded Eukaryotic Genetic Code 2QH, UK. * To whom correspondence should be addressed.
 Supplementary Online Material An Expanded Eukaryotic Genetic Code Jason W. Chin 1, T. Ashton Cropp, J. Christopher Anderson, Mridul Mukherji, Zhiwen Zhang and Peter G. Schultz* Department of Chemistry
Supplementary Online Material An Expanded Eukaryotic Genetic Code Jason W. Chin 1, T. Ashton Cropp, J. Christopher Anderson, Mridul Mukherji, Zhiwen Zhang and Peter G. Schultz* Department of Chemistry
Unit 1. DNA and the Genome
 Unit 1 DNA and the Genome Gene Expression Key Area 3 Vocabulary 1: Transcription Translation Phenotype RNA (mrna, trna, rrna) Codon Anticodon Ribosome RNA polymerase RNA splicing Introns Extrons Gene Expression
Unit 1 DNA and the Genome Gene Expression Key Area 3 Vocabulary 1: Transcription Translation Phenotype RNA (mrna, trna, rrna) Codon Anticodon Ribosome RNA polymerase RNA splicing Introns Extrons Gene Expression
Fluorescence Light Microscopy for Cell Biology
 Fluorescence Light Microscopy for Cell Biology Why use light microscopy? Traditional questions that light microscopy has addressed: Structure within a cell Locations of specific molecules within a cell
Fluorescence Light Microscopy for Cell Biology Why use light microscopy? Traditional questions that light microscopy has addressed: Structure within a cell Locations of specific molecules within a cell
1) The penicillin family of antibiotics, discovered by Alexander Fleming in 1928, has the following general structure: O O
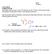 ame: TF ame: LS1a Fall 06 Problem Set #3 Due Friday 10/13 at noon in your TF s drop box on the 2 nd floor of the Science Center All questions including the (*extra*) ones should be turned in 1) The penicillin
ame: TF ame: LS1a Fall 06 Problem Set #3 Due Friday 10/13 at noon in your TF s drop box on the 2 nd floor of the Science Center All questions including the (*extra*) ones should be turned in 1) The penicillin
DNA strands and label their 5 and 3 Top strand. Direction of
 Question 1 (20 points) a) In the table below, name the sub-cellular location or organelle(s) of the eukaryotic cell that will fluoresce when the following macromolecules are tagged with a fluorescent dye.
Question 1 (20 points) a) In the table below, name the sub-cellular location or organelle(s) of the eukaryotic cell that will fluoresce when the following macromolecules are tagged with a fluorescent dye.
