INTRINSIC DISORDER WITHIN AND FLANKING THE DNA-BINDING DOMAINS OF HUMAN TRANSCRIPTION FACTORS
|
|
|
- Justin Baldwin
- 6 years ago
- Views:
Transcription
1 INTRINSIC DISORDER WITHIN AND FLANKING THE DNA-BINDING DOMAINS OF HUMAN TRANSCRIPTION FACTORS XIN GUO 1, MARTHA L. BULYK 2, and ALEXANDER J. HARTEMINK 1 1 Department of Computer Science, Duke University, Box 90129, Durham, NC , USA {xinguo,amink}@cs.duke.edu 2 Division of Genetics, Department of Medicine; Department of Pathology; Brigham and Women s Hospital and Harvard Medical School, Boston, MA 02115, USA Harvard-MIT Division of Health Sciences and Technology, Harvard Medical School, Boston, MA 02115, USA mlbulyk@receptor.med.harvard.edu Correspondence should be addressed to M.L.B. and A.J.H. While the term protein structure is commonplace, it is increasingly appreciated that proteins may not possess a single, well-defined structure: some regions of proteins are intrinsically disordered. The role these intrinsically disordered regions (IDRs) play in protein function is an area of significant interest. In particular, because proteins containing IDRs are largely involved in processes related to molecular recognition, the question arises whether IDRs are important in these recognition events. It has been observed that IDRs are enriched in transcription factors (TFs) in comparison with other proteins, and we sought to explore this enrichment more precisely, with an eye toward functional dissection of the prevalence and locations of IDRs in different classes of TFs. Specifically, we considered the occurrences of 76 classes of DNA-binding domains (DBDs) within a comprehensive set of 1,747 human, sequence-specific TFs. For each DBD class, we analyzed whether a significant level of disorder was present within the DBD itself, the N-terminal or C-terminal sequence flanking the DBD, or both flanking sequences. We found that although the DBDs themselves exhibit significant order, the regions flanking the DBDs exhibit significant disorder, which suggests a functional role for such IDRs in TF DNA binding. These results may have important implications for studies of TFs not just in human but across all eukaryotes, and suggest future studies focused on testing the roles of N- and C-terminal flanking regions in determining or modulating the DNA binding affinity and/or specificity of the associated TFs. Keywords: intrinsically disordered region, DNA-binding domain, transcription factor 1. Introduction The function of a protein is encoded in its amino acid sequence (i.e., primary structure). However, protein activity typically depends on the protein being folded properly into its component secondary structure elements (e.g., alpha helices, beta sheets) and the overall, global conformation of the protein (i.e., tertiary structure). Protein structure can be determined experimentally at high resolution either by X-ray crystallography or by nuclear magnetic resonance (NMR). X-ray crystallography is widely used, but cannot provide information on the conformation of regions that are either highly dynamic or unstructured in the crystal. In contrast, while NMR can provide information about flexibility and dynamics in proteins, it is currently limited to smaller proteins. Through a combination of structural and biochemical studies, it has become increasingly appreciated that a protein may not adopt a single, well-defined structure, a term connoting 104
2 a measure of rigidity. Rather, a protein may sample an ensemble of global conformations; parts of the protein may be largely constantly structured across this ensemble, while other parts may be quite variable or flexible across the ensemble. These latter regions are sometimes termed intrinsically disordered regions (IDRs), though they may adopt a more structured conformation upon interaction with another molecule, whether a protein, DNA, or other ligand [1]. Proteins are largely involved in processes related to molecular recognition (e.g., binding, signaling, complex formation, enzymatic catalysis), and IDRs may enable these recognition events either directly (e.g., serving as the recognition domain of a protein) or indirectly (e.g., serving as a hinge that allows two ordered regions of a protein to come together to effect recognition). For this reason, IDRs have been studied rather extensively over the past decade, and a large number of computational methods have been developed for the prediction of IDRs on the basis of amino acid sequence, though this remains an imperfect art (see [2] for a review). In this study, we were interested in exploring the role(s) that IDRs might play in the recognition tasks of transcription factors (TFs) in particular. Computational explorations have found that IDRs are generally more prevalent in TFs than would be expected by chance, especially in eukaryotes [3 5]. As a specific example, careful molecular studies have shown that a region of fifteen amino acids within the DNA-binding domain (DBD) of the estrogen receptor (ER) is disordered in solution, and makes contacts with DNA (and with another ER DBD monomer), as shown in a co-crystal structure of the ER DBD bound to DNA [6]. Moreover, IDRs outside the homeodomain DBD have also been found to impact the DNA-binding affinity of the Drosophila TF Ubx [7]. In addition, the region N-terminal to the proximal accessory region of the Saccharomyces cerevisiae C2H2 zinc finger TF Adr1 is disordered in solution (even after binding DNA) and increases the affinity for non-specific DNA, mainly by increasing the DNA association rate; increased affinity for non-specific DNA might allow a protein to find its specific sites more quickly after translocation from non-specific sites that are bound initially [8]. Finally, DBDs often have N- or C-terminal extensions, referred to as arms or tails, that bind DNA but are disordered when free in solution [9]. Intrigued by this ensemble of findings pointing to the importance of IDRs in TFs and their interactions with DNA, we sought to explore the connection between IDRs and TF function more precisely and systematically. We were particularly interested in determining whether IDRs were more prevalent in the regions flanking the DBDs that are responsible for the binding of sequence-specific TFs to DNA. 2. Materials and Methods 2.1. Constructing the TF and non-tf control sets of proteins We created two non-redundant datasets of human proteins: a TF set and a non-tf set for use as a control. The procedure for constructing these sets and ensuring their non-redundancy is described below and summarized in Figure 1A. We assembled the TF set from a published repertoire of human TFs [10]. In their study, Vaquerizas and colleagues manually curated and identified 1,987 TF-coding human genomic loci in the Ensembl database [11]; the list includes 1,960 high-confidence entries and 27 entries 105
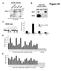
3 A all human TF loci in Ensembl (1,987) cross-reference all human protein entries in RefSeq (~35K) remove TFs B frequency (%) all TFs nr TFs non TF ctrl protein entries in RefSeq (2,362) remove redundancy: 1 isoform per locus non-redundant (nr) TFs (1,747) non-tf protein entries (~32.6K) sample w.r.t. sequence length distribution of nr TF set sampled non-tf ctrl set (1,747) C frequency (%) all TFs nr TFs non TF ctrl length (amino acids) W F Y I M L V N C T A G R D H Q K S E P Fig. 1. (A) A schematic of the pipeline for generating the TF set and the non-tf control set. (B) Sequence length distributions of the TF set (nr TFs), the non-tf control set, and the set of all human TFs (with redundancy). (C) The amino acid compositions of the TF set, the non-tf control set, and the set of all human TFs. Amino acids are listed from most order-promoting to most disorder-promoting, according to [13]. It is apparent from the histogram that compared to proteins in general, TFs have fewer order-promoting residues (e.g., W, F, Y, I, M, L, V) and more disorder-promoting residues (e.g., P, E, S, K, Q, H). curated as probable. We cross-referenced these Ensembl loci against the RefSeq database (release 47) [12] to obtain 2,362 protein isoforms associated with 1,747 genes. To reduce sequence redundancy and thus potential bias, if multiple isoforms were associated with the same gene, we selected only the longest. This resulted in a final total of 1,747 unique TF protein sequences, and in subsequent analysis, we call this our TF set. We assembled our non-tf control set by downloading all human proteins from RefSeq, and excluding the 2,362 TF-associated isoforms from above, which yielded a total of 32,567 non- TF proteins. To match the size and sequence length distribution of our TF set, we randomly sampled 1,747 proteins from the 32,567 according to the empirical sequence length distribution of the TF set; to ensure non-redundancy during this process, at each iteration we required that the sampled protein come from a locus not previously sampled. Therefore, the resulting control set contains 1,747 unique non-tf protein sequences Comparing the TF and non-tf control sets of proteins To ensure that the non-tf set represents a well-constructed control for the TF set, we compared various properties of the two sets. First, we compared the sequence length distributions of the TF set and the non-tf control set, in addition to the set of all human TFs (i.e., with redundancy). As shown in Figure 1B, no apparent differences exist between the sequence length distributions in the TF set, the non-tf control set, and the set of all human TFs. Next, we compared the amino acid compositions of the TF set, the non-tf control set, and the set of all human TFs (Figure 1C). The amino acid composition of sequences in IDRs has been shown to be significantly different from that in ordered regions [14], and IDRs have been shown to have high prevalence in TFs [4], so we might expect compositional differences between the TF sets and the non-tf control set. Indeed, compared to the non-tf control 106
4 set, both TF sets are enriched in disorder-promoting amino acids (e.g., P, E, S, K, Q, H), and depleted in order-promoting amino acids (e.g., W, F, Y, I, M, L, V) [13, 14], as expected. However, the amino acid compositions of our non-redundant TF set and the set of all human TFs are nearly identical, suggesting that our procedure for removing redundancy introduces no significant compositional bias Identifying DNA-binding domains (DBDs) and their locations within proteins Our goal is to investigate the prevalence and locations of IDRs within human TFs, and in particular, the spatial relationships between IDRs and DBDs in TFs. To identify all sequencespecific DBDs that occur within human TFs, we started with the entire set of human proteins from RefSeq and identified every Pfam domain [15] that was contained in a human protein with a p-value below We manually filtered for those domains whose text descriptions in the Pfam or InterPro [16] databases indicated that the domain mediates sequence-specific DNA binding, resulting in 76 domains which we henceforth call Pfam DBDs. Using HMMER [17] with default parameters, we searched for the locations of matches to Pfam DBDs within our TF set. We found 71 of the 76 Pfam DBDs matched to proteins in our TF set, with 32 DBDs appearing more than five times. Of the 1,747 proteins in our TF set, 669 contained only a single DBD, while another 642 contained multiple DBDs; proteins with multiple DBDs are typically those containing multiple zinc fingers, which are annotated as separate domains even if they occur in tandem within a protein. Indeed, the TF with the highest number of DBDs is zinc finger protein 91 (RefSeq: NP ), which contains 31 zf-c2h2 (zinc finger, C2H2-type) domains. The zf-c2h2 domain is interesting in its own right as it is by far the most prevalent domain in our TF set, appearing a total of 4,154 times, almost 20 times as often as the next most prevalent domain Predicting intrinsically disordered regions (IDRs) and their locations within proteins using multiple existing methods To perform our analysis, we first needed to predict the ordered and disordered regions within proteins using existing computational tools. Since this remains a bit of an imperfect art, we took care to ensure that our conclusions would not be overly dependent on the predictions of any single choice of method. Consequently, we chose to use three distinct disorder prediction tools, each demonstrated to perform with high accuracy [2]: PONDR VSL2 [18], DISOPRED2 [19], and PreDisorder 1.1 [20]. PONDR VSL2 (also called DisProt VSL2) was evaluated as the top-ranked disorder predictor in CASP7 in 2006 [21], and PreDisorder was ranked among the top methods in disorder prediction during CASP8 in 2008 and CASP9 in These methods employ a variety of techniques to analyze sequence and structural information for IDR prediction: PONDR VSL2 uses support vector machines (SVMs) to separately address prediction problems in short versus long sequence regions, and then merges the results using a logistic regression model; DISOPRED2 is also based on SVMs, and compared to other prediction methods, the main difference is that it is directly trained on the whole sequence using various combinations of 107
5 binary-encoded amino acid sequence, secondary structure predictions, and sequence profiles; and PreDisorder 1.1 is based on an ab initio prediction method along with a meta-prediction method Defining disorder features: spatial relationships of IDRs relative to DBDs within TFs Given the annotated DBDs and the predicted disorder regions in the TF set and the non- TF control set, we sought to systematically analyze the association between TF DBDs and predicted IDRs by testing for enrichment of IDRs at different locations relative to DBDs. Specifically, we were interested in IDRs within the DBD itself, as well as the regions flanking the DBD, and we developed five distinct disorder features : we say that a DBD is disordered if at least a fraction f of its residues are predicted to be disordered; we say that the N-terminal flank of a DBD is disordered if at least a fraction f of the 30 residues flanking the DBD in the N-terminal direction are predicted to be disordered; analogously, we say that the C-terminal flank of a DBD is disordered if at least a fraction f of the 30 residues flanking the DBD in the C-terminal direction are predicted to be disordered; we say that both flanks of a DBD are disordered if both the N-terminal and C-terminal flanks are disordered; and finally, we say that an entire TF is disordered if at least a fraction f of all of its residues are disordered. We wanted to be fairly stringent in identifying these disorder features, so that we could focus on those with the highest confidence; therefore, we chose the value of 0.8 for f Calculating statistical significance of disorder features To assess whether the prevalence of disorder features within and flanking DBDs was unusually high or low, we needed to determine a suitable measure of significance. Moreover, since different computational tools predict IDRs at different rates (see Section 3.2 below), our significance measure needed to enable the comparison of results across methods, and not be biased by methods that are systematically more or less likely to predict disorder within proteins. We thus developed two different null models to test for the significance of our disorder features (e.g., disordered DBD, N-terminal flank, or C-terminal flank). The first null model pretended that the location of a DBD occurred uniformly at random within each sequence, and was based on the TF set. The second null model also pretended that the location of a DBD occurred uniformly at random in each sequence, but was based on the non-tf control set. In summary, these two null models in which the location of a DBD was chosen uniformly at random were designed to test whether the spatial relationships between IDRs and DBDs were statistically significant or simply occurred by chance. With each null model providing a baseline expectation for how often a disorder feature might be found by chance, we could then compute a significance measure based on the p- value from a hypergeometric distribution (i.e., Fisher s exact test). For each disorder feature we considered, we computed two separate p-values, one for each null model. Consistency of significance across the two different null models thus gave us some confidence that our results were robust to the specific choice of null model. 108
6 A frequency (%) PONDR VSL2 DISOPRED2 PreDisorder1.1 B frequency (%) PONDR VSL2 DISOPRED2 PreDisorder fraction of a protein's residues predicted as disordered fraction of a protein's residues predicted as disordered Fig. 2. Distributions of the fraction of each protein s residues predicted as disordered by each method for the proteins in (A) the TF set and (B) the non-tf control set. Table 1. Statistics summarizing disorder predictions on all the residues of all the proteins in both the TF set and the non-tf control set using three different disorder prediction tools. TF set non-tf ctrl set % of residues average length % of residues average length predicted in IDRs of IDRs predicted in IDRs of IDRs PONDR VSL2 83.2% % 39 DISOPRED2 47.4% % 36 PreDisorder % % Results 3.1. Comparing the three methods for predicting IDRs within proteins We used three different disorder prediction tools to predict IDRs in both the TF set and the non-tf control set. Though the purpose of this paper is to make use of existing prediction methods and not to evaluate them (which has already been done by others, for example [2, 21]), it is important to have at least an overall sense of how each method is performing on our protein sets. A summary of the results of the three methods is shown in Figure 2 and listed in Table 1. In Figure 2, we compare the fraction of each protein s residues predicted as disordered by each method. In Table 1, we calculate the total percentage of protein residues predicted as disordered by each method, along with the average length of each predicted IDR. The figure and table reveal that all three methods consistently predict proteins in the TF set to have a greater fraction of disordered residues, more disordered residues, and longer IDRs than proteins in the non-tf control set, confirming earlier findings that IDRs are enriched in TFs. As an aside, it is apparent that PONDR VSL2 is far more likely than the other two methods to call a residue as disordered, in both the TF set and the non-tf control set, suggesting that the method is probably operating at a different point on its receiver operating characteristic (ROC) curve, with high sensitivity but also perhaps a relatively high false positive rate [21]. In addition, the average length of IDRs predicted by PONDR VSL2 is higher than the other two methods, which may be related to the previous point, but may also be because the method uses different SVMs to predict IDRs in short and long sequences separately. 109
7 3.2. Assessing significance of order or disorder within and flanking human TF DBDs To systematically study the associations between IDRs and DBDs, for each occurrence of a DBD class within a human TF, we calculated 30 different p-values: the significance under two different null models (based on the TF set and the non-tf control set) of five different kinds of disorder features (DBD, N-terminal flank, C-terminal flank, both flanks, and entire TF) as computed by three different prediction methods (PONDR VSL2, DISOPRED2, and PreDisorder 1.1). For each combination of null model and feature, we say that the feature exhibits significant disorder under that null model if at least two of the three prediction methods predict disorder at p-value 0.005; on the other hand, we say that the feature exhibits significant order under that null model if at least two of the three prediction methods predict disorder at p-value Note that it is certainly possible for a feature to be neither significantly ordered nor significantly disordered under a particular null model. Although we computed whether features exhibited significant order or disorder across all Pfam DBDs occurring in our TF set, to avoid artifacts due to small sample size, we restricted our subsequent analysis to the 32 DBD classes with at least five occurrences in the TF set. Many of the most frequent DBD classes, including the 10 most prevalent ones, are structurally similar and can be roughly classified into two groups: (1) those containing zinc fingers, and (2) those containing a basic helix-turn-helix type of domain, domains in which helices are separated by loops (e.g., Homeobox, HLH, Fork head, Ets). The enrichment analysis results for these 32 DBD classes are listed in Table 2; at the bottom of the table, we also included the Pfam domains Basic, AT hook, and P53 (Basic and AT hook are included because we mention them below in comparison to another study; P53 is a well-studied DBD included for general interest). The top 10 most frequently occurring DBD classes in human TFs all exhibit significant order within the DBD itself, suggesting that structural flexibility within these domains is rather limited. Strikingly, our results indicate that although the DBDs themselves exhibit significant order, the regions flanking the DBDs are likely to exhibit significant disorder. Only in the case of zf-c2h2 do the flanking regions exhibit significant order (this will be discussed further in the next section). In contrast, 26 of the other 31 DBDs exhibit significant disorder in either the N-terminal flank, the C-terminal flank, or both; and none of the other 31 DBDs exhibit significant order in either flank under either null model. This is consistent with prior studies in which it was found that DBDs are often separated by flexible linker regions, allowing TFs to bind DNA with fine control over DNA binding affinity [22, 23] Investigating detailed spatial relationships of IDRs relative to DBDs within TFs To further investigate the detailed spatial relationships of the IDR predictions of the three different methods to protein DBDs, we generated a meta-plot of the average predicted order/disorder in the vicinity of each Pfam DBD according to each prediction method. To do this, we first identified all occurrences of a Pfam DBD in the TF set, and then across all those occurrences, calculated the average (mean) order/disorder score predicted by each method at 110
8 Table 2. Enrichment analysis of significantly occurring ordered and disordered regions within and flanking human TF DBDs. DBD N-terminal flank C-terminal flank both flanks whole TF sequence No. DBD (Pfam) TF family average DBD length (res.) number of DBDs in TFs TF set non-tf ctrl set TF set non-tf ctrl set TF set non-tf ctrl set TF set non-tf ctrl set TF set non-tf ctrl set 1 PF00096 zf-c2h OR OR OR OR OR OR OR OR OR OR 2 PF00046 Homeobox OR OR ID ID ID ID ID ID ID ID 3 PF00010 HLH OR OR ID ID ID ID ID ID ID 4 PF00505 HMG box OR OR ID ID ID ID ID ID ID 5 PF00250 Fork head OR OR ID ID ID ID ID ID ID ID 6 PF00105 zf-c OR OR ID ID ID ID ID ID OR 7 PF00249 Myb DNA-binding OR OR ID 8 PF00170 bzip OR OR ID ID ID ID ID ID 9 PF00178 Ets OR OR ID ID ID ID ID ID 10 PF00320 GATA OR ID ID ID ID ID ID ID 11 PF00907 T-box ID ID ID ID ID ID 12 PF01530 zf-c2hc ID ID ID ID ID ID ID ID ID ID 13 PF02319 E2F TDP ID ID ID 14 PF00313 CSD PF05485 THAP ID ID ID ID ID ID 16 PF01422 zf-nf-x PF03165 MH ID ID ID ID ID ID 18 PF07716 bzip ID ID ID ID 19 PF00292 PAX ID ID ID ID ID ID 20 PF00098 zf-cchc ID ID ID 21 PF00808 CBFD NFYB HMF ID 22 PF04218 CENP-B N PF00751 DM ID ID ID ID ID ID ID 24 PF01342 SAND ID ID ID ID ID ID 25 PF02257 RFX DNA binding ID ID 26 PF02864 STAT bind PF02892 zf-bed ID ID ID ID ID ID 28 PF10401 IRF ID 29 PF00447 HSF DNA-bind ID ID ID ID 30 PF04516 CP PF03299 TF AP ID ID ID 32 PF05044 Prox ID ID ID ID PF01586 Basic ID ID ID ID ID ID ID ID ID ID PF00870 P ID ID ID ID ID ID PF02178 AT hook ID ID ID ID ID ID ID ID ID ID Notes: The DBDs with at least 5 occurrences in the TF set are listed in the table, together with Basic, P53, and AT hook. ID indicates significant disordered DBDs, DBD flanks, or TFs (in at least two of three methods, p-value 0.005). OR indicates significant ordered DBDs, DBD flanks, or TFs (in at least two of three methods, p-value 0.005). A dash ( ) indicates entries that are neither significantly ordered nor significantly disordered. The DBDs with fewer than 5 occurrences in the TF set include: Runt, TEA, Basic, HALZ, z-alpha, FYRN, FYRC, P53, ARID, DMA, AKAP95, GATA-N, P53 tetramer, Homez, XPA N, zf-dhhc, GCM, CG-1, Vert HS TF, SIM C, Rad51, HAND, Beta-trefoil, LAG1-DNAbind, PWI, zf-mynd, SAP, GCR, Oest recep, Prog receptor, zf-traf, zf-chy, Vert IL3-reg TF, HSA, Rio2 N, BrkDBD, zf-rag1, AT hook, and TMF DNA bd. 111
9 each residue within the DBD match and both of its flanks (up to 30 amino acids). In cases where a TF contained only a partial DBD match and not a full domain according to the HMMER alignment, we considered only the aligned region in our calculations. We normalized the resulting scores for the purpose of comparison across methods, and for uniformity in scale across plots for different DBD classes (Figure 3). Figure 3 displays meta-plots for five of the ten Pfam DBDs most prevalent in human TFs. Results from DISOPRED2 and PreDisorder 1.1 are fairly consistent across all five domain classes. Moreover, all three methods are in good agreement in zf-c2hc and demonstrate similar prediction trends in zf-c4, Homeobox, and HLH. Extended to all the DBDs listed in Table 2, over 67.2% of the DBD classes that are found to exhibit either significant disorder or significant order are identified as such by all three methods. Nevertheless, some discrepancies in the results from the different methods are evident, such as zf-c2h2. The C2H2-type zinc finger domain is the most prevalent DBD class found in metazoan TFs, including in human [24]. It is also one of the most highly ordered DBDs; however, the linker regions between these C2H2 zinc finger domains are often disordered [25]. As shown in Figure 3A, PONDR VSL2 reports that the C2H2 domain occurrences in human TFs exhibit significant disorder in both the C2H2 domain itself and the adjacent N- and C-terminal flanks; however, DISOPRED2 and PreDisorder both report the opposite, namely that zf-c2h2 and its flanks exhibit significant order. Liu and colleagues carefully analyzed the difficulties of predicting intrinsic disorder in the zf-c2h2 domains and their linker regions [4]. They concluded that because many linker regions between C2H2 zinc fingers are quite short, the windowing procedures employed by some IDR prediction algorithms prevent them from being detected as disordered; the result is an artifact in which linker regions between C2H2 zinc fingers are over-predicted as being ordered Analyzing spatial relationships for some DBD classes prevalent in human TFs Zinc fingers Zinc fingers are small structural motifs whose folds are stabilized by coordination of one or more zinc ions. Zinc fingers can be classified according to their zinc-coordinating residues and folds. In Figures 3A-C, we show our IDR prediction results for the three major zinc finger domain classes found in human TFs: zf-c2h2 (the most prevalent DBD class in human TFs), zf-c4 (also referred to as nuclear receptors), and zf-c2hc. Although all three classes contain zinc fingers, we find variability in their regions of order and disorder. As discussed above, the C2H2 zinc finger domain is itself believed to be highly ordered, with individual ordered zinc fingers separated by highly flexible linker regions [25]. We find that the C4 domain exhibits significant order within the DBD itself, but significant disorder in flanking regions. In contrast, we find that the C2HC domain exhibits significant disorder in both the DBD and flanking regions. 112
10 disordered N-term flank zf-c2h2 C-term flank N-term flank zf-c2hc C-term flank N-term flank zf-c4 C-term flank ordered (A) (B) (C) disordered N-term flank Homeobox C-term flank N-term flank HLH C-term flank PONDR VSL2 DISOPRED2 PreDisorder1.1 ordered (D) (E) Fig. 3. Shown are meta-plots for five prevalent DBDs in human TFs. (A) zinc-finger C2H2-type (length: 23 amino acids), (B) zinc-finger C2HC-type (length: 31 amino acids), (C) zinc-finger C4-type (length: 70 amino acids), (D) homeodomain fold (length: 58 amino acids), and (E) helix-loop-helix (length: 53 amino acids) Homeobox Homeobox (homeodomain fold) is the second-most abundant DBD class within human TFs. The homeodomain fold consists of an approximately 60 amino acid helix-turn-helix structure in which three alpha helices are connected by short loop regions. Our results (Figure 3D) extend the results of a prior study [7] that found multiple intrinsically disordered sequences located outside the homeodomain DBD of the Drosophila TF Ubx, that allow Hox family members (i.e., a subclass of TFs with Homeobox DBDs) to bind DNA with high affinity but relatively low specificity [26, 27] HLH HLH (basic helix-loop-helix) is the third-most abundant DBD class within human TFs, and is characterized by two α-helices connected by a loop. TFs that have this domain typically bind DNA as either homo- or hetero-dimers, with each monomer contacting DNA through a helix containing basic residues that facilitate DNA binding [28]. As shown in Figure 3E, all three methods report that HLH exhibits significant order within the domain itself, but significant disorder in both the N- and C-terminal flanking regions. Our results also indicate that a short but highly disordered region may frequently occur in the middle of the HLH domain, consistent with prior observations that the linker regions and the loop region of HLH 113
11 proteins are of higher flexibility, allowing dimerization by folding and packing one smaller helix against the other one [28]. 4. Discussion In this study, we used three different computational disorder prediction methods to investigate the prevalence of IDRs within DBDs and in their flanking regions across essentially the entire repertoire of human, sequence-specific TFs and their associated Pfam DBDs. Our choice of multiple prediction methods was motivated by a desire to be able to draw robust conclusions that were not dependent on any one particular method. Previously it was found that TFs are enriched for IDRs [3, 4]. At the same time, DBDs responsible for TF binding did not seem themselves to be particularly enriched for IDRs. For example, of the 25 DBDs studied in [4], only the Basic and AT hook domains exhibited high amounts of disorder; however, those domains are not particularly prevalent in human TFs, occurring in our TF set just four times and one time, respectively. a We were intrigued by the possibility that the enrichment of IDRs observed in TFs might be at least partly due to disorder in the regions flanking DBDs; under such a hypothesis, DBDs can be thought of as islands of order flanked by regions of disorder. Our results support exactly such a hypothesis: the most prevalent DBDs in human TFs exhibit significant order, but the flanking regions of these DBDs generally exhibit significant disorder. Similarly, although DBDs of intermediate prevalence (occurring between 5 and 20 times in our TF set) do not appear often enough to exhibit either significant order or disorder within the domains themselves, most of them still exhibit significant disorder in one or both flanking regions. The functional role played by the significant prevalence of disorder in the regions flanking DBDs of human TFs is unclear. However, we can speculate that the increased flexibility afforded by these flanking IDRs might contribute to the ability of TFs to 1) recognize target sequences in the DNA appropriately, 2) bind to a wider diversity of DNA target sequences, 3) be anchored with higher affinity to the DNA after recognizing target sequences, 4) bind to other factors and complexes positioned on the DNA or involved in transcriptional regulation, or 5) present activation domains to downstream transcriptional regulatory machinery. It should be emphasized that these possibilities are speculative; however, the results of this study suggest numerous testable hypotheses regarding the roles of N- and C-terminal regions flanking DBDs for many frequently occurring DBDs in hundreds of human TFs. For example, the importance of the predicted disorder in these flanking regions in determining or modulating the DNA binding affinity and/or specificity of the associated TFs could be investigated with protein binding microarrays (PBMs) [29, 30]. PBMs could assay the affinity and/or specificity of proteins representing the DBDs with their flanking regions, as compared to either the DBDs alone or the DBDs with mutant flanking regions predicted not to be significantly disordered. If found to contribute to the DNA binding affinity and/or specificity of TFs, IDRs that flank DBDs would broaden the scope of functional domains to be considered when evaluating the a Though they do not occur often, where they do occur, they exhibit significant disorder in our results as well, corroborating the results in [4]; see Table
12 potential impact of mutations or natural polymorphisms within exomes, such as in medical sequencing projects. This study was focused on human TFs; however, since these DBD classes are the predominant DBD classes not just in human TFs but throughout eukaryotes, the results of this study may have important implications for studies of TFs across all eukaryotes. 5. Acknowledgments We are grateful to Lizzy Rossin for creating the initial list of Pfam DBDs in human TFs. This work was funded by grants from NIH (R01 HG003985) to M.L.B., and from NIH (P50 GM ) and DARPA (HR ) to A.J.H. References [1] D. Eliezer, et al., Curr. Opin. Struct. Biol. 19, 23 (Feb 2009). [2] B. He, et al., Cell Res. 19, 929 (Aug 2009). [3] Y. Minezaki, et al., J. Mol. Biol. 359, 1137 (Jun 2006). [4] J. Liu, et al., Biochemistry 45, 6873 (Jun 2006). [5] M. Fuxreiter, et al., Trends Biochem. Sci. 36, 415 (Aug 2011). [6] J. W. Schwabe, et al., Structure 1, 187 (Nov 1993). [7] Y. Liu, et al., J. Biol. Chem. 283, (Jul 2008). [8] L. E. Schaufler, et al., J. Mol. Biol. 329, 931 (Jun 2003). [9] C. Crane-Robinson, et al., Trends Biochem. Sci. 31, 547 (Oct 2006). [10] J. M. Vaquerizas, et al., Nat. Rev. Genet. 10, 252 (Apr 2009). [11] P. Flicek, et al., Nucleic Acids Res. 39, D800 (Jan 2011). [12] K. D. Pruitt, et al., Nucleic Acids Res. 37, D32 (Jan 2009). [13] A. Campen, et al., Protein Pept. Lett. 15, 956 (2008). [14] A. K. Dunker, et al., J. Mol. Graph. Model. 19, 26 (2001). [15] R. D. Finn, et al., Nucleic Acids Res. 38, D211 (Jan 2010). [16] S. Hunter, et al., Nucleic Acids Res. 37, D211 (Jan 2009). [17] S. R. Eddy, et al., Genome Inform 23, 205 (Oct 2009). [18] K. Peng, et al., BMC Bioinformatics 7, 208 (2006). [19] J. J. Ward, et al., J. Mol. Biol. 337, 635 (Mar 2004). [20] X. Deng, et al., BMC Bioinformatics 10, 436 (2009). [21] L. Bordoli, et al., Proteins 69 Suppl 8, 129 (2007). [22] H. X. Zhou, et al., Biochemistry 40, (Dec 2001). [23] S. Fukuchi, et al., J. Mol. Biol. 355, 845 (Jan 2006). [24] R. Tupler, et al., Nature 409, 832 (Feb 2001). [25] C. O. Pabo, et al., Annu. Rev. Biochem. 70, 313 (2001). [26] W. J. Gehring, et al., Cell 78, 211 (Jul 1994). [27] T. Hoey, et al., Nature 332, 858 (Apr 1988). [28] T. D. Littlewood, et al., Protein Profile 2, 621 (1995). [29] S. Mukherjee, et al., Nat. Genet. 36, 1331 (Dec 2004). [30] M. F. Berger, et al., Cell 133, 1266 (Jun 2008). 115
Evaluation of Disorder Predictions in CASP5
 PROTEINS: Structure, Function, and Genetics 53:561 565 (2003) Evaluation of Disorder Predictions in CASP5 Eugene Melamud 1,2 and John Moult 1 * 1 Center for Advanced Research in Biotechnology, University
PROTEINS: Structure, Function, and Genetics 53:561 565 (2003) Evaluation of Disorder Predictions in CASP5 Eugene Melamud 1,2 and John Moult 1 * 1 Center for Advanced Research in Biotechnology, University
Chapter 25: Regulating Eukaryotic Transcription The Ligand Responsive Activators
 Chapter 25: Regulating Eukaryotic Transcription The Ligand Responsive Activators At least 5 potential gene expression control points Superfamily of Gene Regulators Activation of gene structure Initiation
Chapter 25: Regulating Eukaryotic Transcription The Ligand Responsive Activators At least 5 potential gene expression control points Superfamily of Gene Regulators Activation of gene structure Initiation
Prediction of protein disorder Zsuzsanna Dosztányi
 Prediction of protein disorder Zsuzsanna Dosztányi MTA-ELTE Momentum Bioinformatics Group Department of Biochemistry, Eotvos Lorand University, Budapest, Hungary dosztanyi@caeser.elte.hu IDPs Intrinsically
Prediction of protein disorder Zsuzsanna Dosztányi MTA-ELTE Momentum Bioinformatics Group Department of Biochemistry, Eotvos Lorand University, Budapest, Hungary dosztanyi@caeser.elte.hu IDPs Intrinsically
Chapter 14 Regulation of Transcription
 Chapter 14 Regulation of Transcription Cis-acting sequences Distance-independent cis-acting elements Dissecting regulatory elements Transcription factors Overview transcriptional regulation Transcription
Chapter 14 Regulation of Transcription Cis-acting sequences Distance-independent cis-acting elements Dissecting regulatory elements Transcription factors Overview transcriptional regulation Transcription
Title: Genome-Wide Predictions of Transcription Factor Binding Events using Multi- Dimensional Genomic and Epigenomic Features Background
 Title: Genome-Wide Predictions of Transcription Factor Binding Events using Multi- Dimensional Genomic and Epigenomic Features Team members: David Moskowitz and Emily Tsang Background Transcription factors
Title: Genome-Wide Predictions of Transcription Factor Binding Events using Multi- Dimensional Genomic and Epigenomic Features Team members: David Moskowitz and Emily Tsang Background Transcription factors
2. Materials and Methods
 Identification of cancer-relevant Variations in a Novel Human Genome Sequence Robert Bruggner, Amir Ghazvinian 1, & Lekan Wang 1 CS229 Final Report, Fall 2009 1. Introduction Cancer affects people of all
Identification of cancer-relevant Variations in a Novel Human Genome Sequence Robert Bruggner, Amir Ghazvinian 1, & Lekan Wang 1 CS229 Final Report, Fall 2009 1. Introduction Cancer affects people of all
Structural Bioinformatics (C3210) Conformational Analysis Protein Folding Protein Structure Prediction
 Structural Bioinformatics (C3210) Conformational Analysis Protein Folding Protein Structure Prediction Conformational Analysis 2 Conformational Analysis Properties of molecules depend on their three-dimensional
Structural Bioinformatics (C3210) Conformational Analysis Protein Folding Protein Structure Prediction Conformational Analysis 2 Conformational Analysis Properties of molecules depend on their three-dimensional
Structure/function relationship in DNA-binding proteins
 PHRM 836 September 22, 2015 Structure/function relationship in DNA-binding proteins Devlin Chapter 8.8-9 u General description of transcription factors (TFs) u Sequence-specific interactions between DNA
PHRM 836 September 22, 2015 Structure/function relationship in DNA-binding proteins Devlin Chapter 8.8-9 u General description of transcription factors (TFs) u Sequence-specific interactions between DNA
Enhancers. Activators and repressors of transcription
 Enhancers Can be >50 kb away from the gene they regulate. Can be upstream from a promoter, downstream from a promoter, within an intron, or even downstream of the final exon of a gene. Are often cell type
Enhancers Can be >50 kb away from the gene they regulate. Can be upstream from a promoter, downstream from a promoter, within an intron, or even downstream of the final exon of a gene. Are often cell type
Transcription factors
 Atlas of Genetics and Cytogenetics in Oncology and Haematology Transcription factors I Introduction * II Initiation of transcription III Transcription factors family pdf version I Introduction III.1 Helix-Turn-Helix
Atlas of Genetics and Cytogenetics in Oncology and Haematology Transcription factors I Introduction * II Initiation of transcription III Transcription factors family pdf version I Introduction III.1 Helix-Turn-Helix
Intrinsic disorder in putative protein sequences
 Intrinsic disorder in putative protein sequences Uros Midic, Zoran Obradovic Center for Data Analytics and Biomedical Informatics Temple University Philadelphia, PA, USA uros@temple.edu, zoran.obradovic@temple.edu
Intrinsic disorder in putative protein sequences Uros Midic, Zoran Obradovic Center for Data Analytics and Biomedical Informatics Temple University Philadelphia, PA, USA uros@temple.edu, zoran.obradovic@temple.edu
Annotating 7G24-63 Justin Richner May 4, Figure 1: Map of my sequence
 Annotating 7G24-63 Justin Richner May 4, 2005 Zfh2 exons Thd1 exons Pur-alpha exons 0 40 kb 8 = 1 kb = LINE, Penelope = DNA/Transib, Transib1 = DINE = Novel Repeat = LTR/PAO, Diver2 I = LTR/Gypsy, Invader
Annotating 7G24-63 Justin Richner May 4, 2005 Zfh2 exons Thd1 exons Pur-alpha exons 0 40 kb 8 = 1 kb = LINE, Penelope = DNA/Transib, Transib1 = DINE = Novel Repeat = LTR/PAO, Diver2 I = LTR/Gypsy, Invader
Supplementary Fig. S1. Building a training set of cardiac enhancers. (A-E) Empirical validation of candidate enhancers containing matches to Twi and
 Supplementary Fig. S1. Building a training set of cardiac enhancers. (A-E) Empirical validation of candidate enhancers containing matches to Twi and Tin TFBS motifs and located in the flanking or intronic
Supplementary Fig. S1. Building a training set of cardiac enhancers. (A-E) Empirical validation of candidate enhancers containing matches to Twi and Tin TFBS motifs and located in the flanking or intronic
A Hidden Markov Model for Identification of Helix-Turn-Helix Motifs
 A Hidden Markov Model for Identification of Helix-Turn-Helix Motifs CHANGHUI YAN and JING HU Department of Computer Science Utah State University Logan, UT 84341 USA cyan@cc.usu.edu http://www.cs.usu.edu/~cyan
A Hidden Markov Model for Identification of Helix-Turn-Helix Motifs CHANGHUI YAN and JING HU Department of Computer Science Utah State University Logan, UT 84341 USA cyan@cc.usu.edu http://www.cs.usu.edu/~cyan
The relationship between relative solvent accessible surface area (rasa) and irregular structures in protean segments (ProSs)
 www.bioinformation.net Volume 12(9) Hypothesis The relationship between relative solvent accessible surface area (rasa) and irregular structures in protean segments (ProSs) Divya Shaji Graduate School
www.bioinformation.net Volume 12(9) Hypothesis The relationship between relative solvent accessible surface area (rasa) and irregular structures in protean segments (ProSs) Divya Shaji Graduate School
Solution Structure of the DNA-binding Domain of GAL4 from Saccharomyces cerevisiae
 Vol. 14, No. 1-4 175 Solution Structure of the DNA-binding Domain of GAL4 from Saccharomyces cerevisiae James D. Baleja, V. Thanabal, Ted Mau, and Gerhard Wagner Department of Biological Chemistry and
Vol. 14, No. 1-4 175 Solution Structure of the DNA-binding Domain of GAL4 from Saccharomyces cerevisiae James D. Baleja, V. Thanabal, Ted Mau, and Gerhard Wagner Department of Biological Chemistry and
Correcting Sampling Bias in Structural Genomics through Iterative Selection of Underrepresented Targets
 Correcting Sampling Bias in Structural Genomics through Iterative Selection of Underrepresented Targets Kang Peng Slobodan Vucetic Zoran Obradovic Abstract In this study we proposed an iterative procedure
Correcting Sampling Bias in Structural Genomics through Iterative Selection of Underrepresented Targets Kang Peng Slobodan Vucetic Zoran Obradovic Abstract In this study we proposed an iterative procedure
Bioinformatics & Protein Structural Analysis. Bioinformatics & Protein Structural Analysis. Learning Objective. Proteomics
 The molecular structures of proteins are complex and can be defined at various levels. These structures can also be predicted from their amino-acid sequences. Protein structure prediction is one of the
The molecular structures of proteins are complex and can be defined at various levels. These structures can also be predicted from their amino-acid sequences. Protein structure prediction is one of the
11/22/13. Proteomics, functional genomics, and systems biology. Biosciences 741: Genomics Fall, 2013 Week 11
 Proteomics, functional genomics, and systems biology Biosciences 741: Genomics Fall, 2013 Week 11 1 Figure 6.1 The future of genomics Functional Genomics The field of functional genomics represents the
Proteomics, functional genomics, and systems biology Biosciences 741: Genomics Fall, 2013 Week 11 1 Figure 6.1 The future of genomics Functional Genomics The field of functional genomics represents the
ChIP-Seq Data Analysis. J Fass UCD Genome Center Bioinformatics Core Wednesday December 17, 2014
 ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday December 17, 2014 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind
ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday December 17, 2014 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind
Assemblytics: a web analytics tool for the detection of assembly-based variants Maria Nattestad and Michael C. Schatz
 Assemblytics: a web analytics tool for the detection of assembly-based variants Maria Nattestad and Michael C. Schatz Table of Contents Supplementary Note 1: Unique Anchor Filtering Supplementary Figure
Assemblytics: a web analytics tool for the detection of assembly-based variants Maria Nattestad and Michael C. Schatz Table of Contents Supplementary Note 1: Unique Anchor Filtering Supplementary Figure
CS273: Algorithms for Structure Handout # 5 and Motion in Biology Stanford University Tuesday, 13 April 2004
 CS273: Algorithms for Structure Handout # 5 and Motion in Biology Stanford University Tuesday, 13 April 2004 Lecture #5: 13 April 2004 Topics: Sequence motif identification Scribe: Samantha Chui 1 Introduction
CS273: Algorithms for Structure Handout # 5 and Motion in Biology Stanford University Tuesday, 13 April 2004 Lecture #5: 13 April 2004 Topics: Sequence motif identification Scribe: Samantha Chui 1 Introduction
In 1996, the genome of Saccharomyces cerevisiae was completed due to the work of
 Summary: Kellis, M. et al. Nature 423,241-253. Background In 1996, the genome of Saccharomyces cerevisiae was completed due to the work of approximately 600 scientists world-wide. This group of researchers
Summary: Kellis, M. et al. Nature 423,241-253. Background In 1996, the genome of Saccharomyces cerevisiae was completed due to the work of approximately 600 scientists world-wide. This group of researchers
DNA. Branden & Tooze, Ch. 7 Deoxyribose nucleic acids are made of three parts
 DNA Branden & Tooze, Ch. 7 Deoxyribose nucleic acids are made of three parts base: adenine, cytosine, guanine, thymine sugar: deoxyribose phosphate: will form the phosphate backbone wide narrow DNA binding
DNA Branden & Tooze, Ch. 7 Deoxyribose nucleic acids are made of three parts base: adenine, cytosine, guanine, thymine sugar: deoxyribose phosphate: will form the phosphate backbone wide narrow DNA binding
Figure S1. nuclear extracts. HeLa cell nuclear extract. Input IgG IP:ORC2 ORC2 ORC2. MCM4 origin. ORC2 occupancy
 A nuclear extracts B HeLa cell nuclear extract Figure S1 ORC2 (in kda) 21 132 7 ORC2 Input IgG IP:ORC2 32 ORC C D PRKDC ORC2 occupancy Directed against ORC2 C-terminus (sc-272) MCM origin 2 2 1-1 -1kb
A nuclear extracts B HeLa cell nuclear extract Figure S1 ORC2 (in kda) 21 132 7 ORC2 Input IgG IP:ORC2 32 ORC C D PRKDC ORC2 occupancy Directed against ORC2 C-terminus (sc-272) MCM origin 2 2 1-1 -1kb
ChIP-seq and RNA-seq. Farhat Habib
 ChIP-seq and RNA-seq Farhat Habib fhabib@iiserpune.ac.in Biological Goals Learn how genomes encode the diverse patterns of gene expression that define each cell type and state. Protein-DNA interactions
ChIP-seq and RNA-seq Farhat Habib fhabib@iiserpune.ac.in Biological Goals Learn how genomes encode the diverse patterns of gene expression that define each cell type and state. Protein-DNA interactions
Computational Methods for Protein Structure Prediction and Fold Recognition... 1 I. Cymerman, M. Feder, M. PawŁowski, M.A. Kurowski, J.M.
 Contents Computational Methods for Protein Structure Prediction and Fold Recognition........................... 1 I. Cymerman, M. Feder, M. PawŁowski, M.A. Kurowski, J.M. Bujnicki 1 Primary Structure Analysis...................
Contents Computational Methods for Protein Structure Prediction and Fold Recognition........................... 1 I. Cymerman, M. Feder, M. PawŁowski, M.A. Kurowski, J.M. Bujnicki 1 Primary Structure Analysis...................
Chapter 8 DNA Recognition in Prokaryotes by Helix-Turn-Helix Motifs
 Chapter 8 DNA Recognition in Prokaryotes by Helix-Turn-Helix Motifs 1. Helix-turn-helix proteins 2. Zinc finger proteins 3. Leucine zipper proteins 4. Beta-scaffold factors 5. Others λ-repressor AND CRO
Chapter 8 DNA Recognition in Prokaryotes by Helix-Turn-Helix Motifs 1. Helix-turn-helix proteins 2. Zinc finger proteins 3. Leucine zipper proteins 4. Beta-scaffold factors 5. Others λ-repressor AND CRO
Prediction of DNA-Binding Proteins from Structural Features
 Prediction of DNA-Binding Proteins from Structural Features Andrea Szabóová 1, Ondřej Kuželka 1, Filip Železný1, and Jakub Tolar 2 1 Czech Technical University, Prague, Czech Republic 2 University of Minnesota,
Prediction of DNA-Binding Proteins from Structural Features Andrea Szabóová 1, Ondřej Kuželka 1, Filip Železný1, and Jakub Tolar 2 1 Czech Technical University, Prague, Czech Republic 2 University of Minnesota,
ChIP-Seq Tools. J Fass UCD Genome Center Bioinformatics Core Wednesday September 16, 2015
 ChIP-Seq Tools J Fass UCD Genome Center Bioinformatics Core Wednesday September 16, 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA or
ChIP-Seq Tools J Fass UCD Genome Center Bioinformatics Core Wednesday September 16, 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA or
Representation in Supervised Machine Learning Application to Biological Problems
 Representation in Supervised Machine Learning Application to Biological Problems Frank Lab Howard Hughes Medical Institute & Columbia University 2010 Robert Howard Langlois Hughes Medical Institute What
Representation in Supervised Machine Learning Application to Biological Problems Frank Lab Howard Hughes Medical Institute & Columbia University 2010 Robert Howard Langlois Hughes Medical Institute What
ChIP-Seq Data Analysis. J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015
 ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA
ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA
Homework 4. Due in class, Wednesday, November 10, 2004
 1 GCB 535 / CIS 535 Fall 2004 Homework 4 Due in class, Wednesday, November 10, 2004 Comparative genomics 1. (6 pts) In Loots s paper (http://www.seas.upenn.edu/~cis535/lab/sciences-loots.pdf), the authors
1 GCB 535 / CIS 535 Fall 2004 Homework 4 Due in class, Wednesday, November 10, 2004 Comparative genomics 1. (6 pts) In Loots s paper (http://www.seas.upenn.edu/~cis535/lab/sciences-loots.pdf), the authors
DNA Binding Domains: Structural Motifs. Effector Domain. Zinc Fingers. Zinc Fingers, continued. Zif268
 DNA Binding Domains: Structural Motifs Studies of known transcription factors have found several motifs of protein design to allow sequence-specific binding of DNA. We will cover only three of these motifs:
DNA Binding Domains: Structural Motifs Studies of known transcription factors have found several motifs of protein design to allow sequence-specific binding of DNA. We will cover only three of these motifs:
Methods and tools for exploring functional genomics data
 Methods and tools for exploring functional genomics data William Stafford Noble Department of Genome Sciences Department of Computer Science and Engineering University of Washington Outline Searching for
Methods and tools for exploring functional genomics data William Stafford Noble Department of Genome Sciences Department of Computer Science and Engineering University of Washington Outline Searching for
Question 2: There are 5 retroelements (2 LINEs and 3 LTRs), 6 unclassified elements (XDMR and XDMR_DM), and 7 satellite sequences.
 Bio4342 Exercise 1 Answers: Detecting and Interpreting Genetic Homology (Answers prepared by Wilson Leung) Question 1: Low complexity DNA can be described as sequences that consist primarily of one or
Bio4342 Exercise 1 Answers: Detecting and Interpreting Genetic Homology (Answers prepared by Wilson Leung) Question 1: Low complexity DNA can be described as sequences that consist primarily of one or
Textbook Reading Guidelines
 Understanding Bioinformatics by Marketa Zvelebil and Jeremy Baum Last updated: May 1, 2009 Textbook Reading Guidelines Preface: Read the whole preface, and especially: For the students with Life Science
Understanding Bioinformatics by Marketa Zvelebil and Jeremy Baum Last updated: May 1, 2009 Textbook Reading Guidelines Preface: Read the whole preface, and especially: For the students with Life Science
C2H2 zinc finger proteins greatly expand the human regulatory lexicon
 Supplementary Information for: C2H2 zinc finger proteins greatly expand the human regulatory lexicon Hamed S. Najafabadi*,1, Sanie Mnaimneh*,1, Frank W. Schmitges*,1, Michael Garton 1, Kathy N. Lam 2,
Supplementary Information for: C2H2 zinc finger proteins greatly expand the human regulatory lexicon Hamed S. Najafabadi*,1, Sanie Mnaimneh*,1, Frank W. Schmitges*,1, Michael Garton 1, Kathy N. Lam 2,
Sequence Analysis '17 -- lecture Secondary structure 3. Sequence similarity and homology 2. Secondary structure prediction
 Sequence Analysis '17 -- lecture 16 1. Secondary structure 3. Sequence similarity and homology 2. Secondary structure prediction Alpha helix Right-handed helix. H-bond is from the oxygen at i to the nitrogen
Sequence Analysis '17 -- lecture 16 1. Secondary structure 3. Sequence similarity and homology 2. Secondary structure prediction Alpha helix Right-handed helix. H-bond is from the oxygen at i to the nitrogen
CSE : Computational Issues in Molecular Biology. Lecture 19. Spring 2004
 CSE 397-497: Computational Issues in Molecular Biology Lecture 19 Spring 2004-1- Protein structure Primary structure of protein is determined by number and order of amino acids within polypeptide chain.
CSE 397-497: Computational Issues in Molecular Biology Lecture 19 Spring 2004-1- Protein structure Primary structure of protein is determined by number and order of amino acids within polypeptide chain.
ChroMoS Guide (version 1.2)
 ChroMoS Guide (version 1.2) Background Genome-wide association studies (GWAS) reveal increasing number of disease-associated SNPs. Since majority of these SNPs are located in intergenic and intronic regions
ChroMoS Guide (version 1.2) Background Genome-wide association studies (GWAS) reveal increasing number of disease-associated SNPs. Since majority of these SNPs are located in intergenic and intronic regions
Expression Analysis Systematic Explorer (EASE)
 Expression Analysis Systematic Explorer (EASE) EASE is a customizable software application for rapid biological interpretation of gene lists that result from the analysis of microarray, proteomics, SAGE,
Expression Analysis Systematic Explorer (EASE) EASE is a customizable software application for rapid biological interpretation of gene lists that result from the analysis of microarray, proteomics, SAGE,
Protein Folding Problem I400: Introduction to Bioinformatics
 Protein Folding Problem I400: Introduction to Bioinformatics November 29, 2004 Protein biomolecule, macromolecule more than 50% of the dry weight of cells is proteins polymer of amino acids connected into
Protein Folding Problem I400: Introduction to Bioinformatics November 29, 2004 Protein biomolecule, macromolecule more than 50% of the dry weight of cells is proteins polymer of amino acids connected into
CFSSP: Chou and Fasman Secondary Structure Prediction server
 Wide Spectrum, Vol. 1, No. 9, (2013) pp 15-19 CFSSP: Chou and Fasman Secondary Structure Prediction server T. Ashok Kumar Department of Bioinformatics, Noorul Islam College of Arts and Science, Kumaracoil
Wide Spectrum, Vol. 1, No. 9, (2013) pp 15-19 CFSSP: Chou and Fasman Secondary Structure Prediction server T. Ashok Kumar Department of Bioinformatics, Noorul Islam College of Arts and Science, Kumaracoil
Annotating Fosmid 14p24 of D. Virilis chromosome 4
 Lo 1 Annotating Fosmid 14p24 of D. Virilis chromosome 4 Lo, Louis April 20, 2006 Annotation Report Introduction In the first half of Research Explorations in Genomics I finished a 38kb fragment of chromosome
Lo 1 Annotating Fosmid 14p24 of D. Virilis chromosome 4 Lo, Louis April 20, 2006 Annotation Report Introduction In the first half of Research Explorations in Genomics I finished a 38kb fragment of chromosome
Alexander Statnikov, Ph.D.
 Alexander Statnikov, Ph.D. Director, Computational Causal Discovery Laboratory Benchmarking Director, Best Practices Integrative Informatics Consultation Service Assistant Professor, Department of Medicine,
Alexander Statnikov, Ph.D. Director, Computational Causal Discovery Laboratory Benchmarking Director, Best Practices Integrative Informatics Consultation Service Assistant Professor, Department of Medicine,
Gene regulation V Biochemistry 302. March 6, 2006
 Gene regulation V Biochemistry 302 March 6, 2006 Common structural motifs associated with transcriptional regulatory proteins Helix-turn-helix Prokaryotic repressors and activators Eukaryotic homeodomain
Gene regulation V Biochemistry 302 March 6, 2006 Common structural motifs associated with transcriptional regulatory proteins Helix-turn-helix Prokaryotic repressors and activators Eukaryotic homeodomain
Characterizing DNA binding sites high throughput approaches Biol4230 Tues, April 24, 2018 Bill Pearson Pinn 6-057
 Characterizing DNA binding sites high throughput approaches Biol4230 Tues, April 24, 2018 Bill Pearson wrp@virginia.edu 4-2818 Pinn 6-057 Reviewing sites: affinity and specificity representation binding
Characterizing DNA binding sites high throughput approaches Biol4230 Tues, April 24, 2018 Bill Pearson wrp@virginia.edu 4-2818 Pinn 6-057 Reviewing sites: affinity and specificity representation binding
A Protein Secondary Structure Prediction Method Based on BP Neural Network Ru-xi YIN, Li-zhen LIU*, Wei SONG, Xin-lei ZHAO and Chao DU
 2017 2nd International Conference on Artificial Intelligence: Techniques and Applications (AITA 2017 ISBN: 978-1-60595-491-2 A Protein Secondary Structure Prediction Method Based on BP Neural Network Ru-xi
2017 2nd International Conference on Artificial Intelligence: Techniques and Applications (AITA 2017 ISBN: 978-1-60595-491-2 A Protein Secondary Structure Prediction Method Based on BP Neural Network Ru-xi
Chimp Sequence Annotation: Region 2_3
 Chimp Sequence Annotation: Region 2_3 Jeff Howenstein March 30, 2007 BIO434W Genomics 1 Introduction We received region 2_3 of the ChimpChunk sequence, and the first step we performed was to run RepeatMasker
Chimp Sequence Annotation: Region 2_3 Jeff Howenstein March 30, 2007 BIO434W Genomics 1 Introduction We received region 2_3 of the ChimpChunk sequence, and the first step we performed was to run RepeatMasker
Practice Exam A. Briefly describe how IL-25 treatment might be able to help this responder subgroup of liver cancer patients.
 Practice Exam 2007 1. A special JAK-STAT signaling system (JAK5-STAT5) was recently identified in which a gene called TS5 becomes selectively transcribed and expressed in the liver upon induction by a
Practice Exam 2007 1. A special JAK-STAT signaling system (JAK5-STAT5) was recently identified in which a gene called TS5 becomes selectively transcribed and expressed in the liver upon induction by a
Data Retrieval from GenBank
 Data Retrieval from GenBank Peter J. Myler Bioinformatics of Intracellular Pathogens JNU, Feb 7-0, 2009 http://www.ncbi.nlm.nih.gov (January, 2007) http://ncbi.nlm.nih.gov/sitemap/resourceguide.html Accessing
Data Retrieval from GenBank Peter J. Myler Bioinformatics of Intracellular Pathogens JNU, Feb 7-0, 2009 http://www.ncbi.nlm.nih.gov (January, 2007) http://ncbi.nlm.nih.gov/sitemap/resourceguide.html Accessing
PREDICTING PREVENTABLE ADVERSE EVENTS USING INTEGRATED SYSTEMS PHARMACOLOGY
 PREDICTING PREVENTABLE ADVERSE EVENTS USING INTEGRATED SYSTEMS PHARMACOLOGY GUY HASKIN FERNALD 1, DORNA KASHEF 2, NICHOLAS P. TATONETTI 1 Center for Biomedical Informatics Research 1, Department of Computer
PREDICTING PREVENTABLE ADVERSE EVENTS USING INTEGRATED SYSTEMS PHARMACOLOGY GUY HASKIN FERNALD 1, DORNA KASHEF 2, NICHOLAS P. TATONETTI 1 Center for Biomedical Informatics Research 1, Department of Computer
Bioinformatics Tools. Stuart M. Brown, Ph.D Dept of Cell Biology NYU School of Medicine
 Bioinformatics Tools Stuart M. Brown, Ph.D Dept of Cell Biology NYU School of Medicine Bioinformatics Tools Stuart M. Brown, Ph.D Dept of Cell Biology NYU School of Medicine Overview This lecture will
Bioinformatics Tools Stuart M. Brown, Ph.D Dept of Cell Biology NYU School of Medicine Bioinformatics Tools Stuart M. Brown, Ph.D Dept of Cell Biology NYU School of Medicine Overview This lecture will
Non-coding Function & Variation, MPRAs II. Mike White Bio /5/18
 Non-coding Function & Variation, MPRAs II Mike White Bio 5488 3/5/18 MPRA Review Problem 1: Where does your CRE DNA come from? DNA synthesis Genomic fragments Targeted regulome capture Problem 2: How do
Non-coding Function & Variation, MPRAs II Mike White Bio 5488 3/5/18 MPRA Review Problem 1: Where does your CRE DNA come from? DNA synthesis Genomic fragments Targeted regulome capture Problem 2: How do
Feature-based classification of human transcription factors into hypothetical subclasses related to regulatory function
 Ehsani et al. BMC Bioinformatics (2016) 17:459 DOI 10.1186/s12859-016-1349-2 RESEARCH ARTICLE Open Access Feature-based classification of human transcription factors into hypothetical subclasses related
Ehsani et al. BMC Bioinformatics (2016) 17:459 DOI 10.1186/s12859-016-1349-2 RESEARCH ARTICLE Open Access Feature-based classification of human transcription factors into hypothetical subclasses related
Array-Ready Oligo Set for the Rat Genome Version 3.0
 Array-Ready Oligo Set for the Rat Genome Version 3.0 We are pleased to announce Version 3.0 of the Rat Genome Oligo Set containing 26,962 longmer probes representing 22,012 genes and 27,044 gene transcripts.
Array-Ready Oligo Set for the Rat Genome Version 3.0 We are pleased to announce Version 3.0 of the Rat Genome Oligo Set containing 26,962 longmer probes representing 22,012 genes and 27,044 gene transcripts.
Protein Sequence Analysis. BME 110: CompBio Tools Todd Lowe April 19, 2007 (Slide Presentation: Carol Rohl)
 Protein Sequence Analysis BME 110: CompBio Tools Todd Lowe April 19, 2007 (Slide Presentation: Carol Rohl) Linear Sequence Analysis What can you learn from a (single) protein sequence? Calculate it s physical
Protein Sequence Analysis BME 110: CompBio Tools Todd Lowe April 19, 2007 (Slide Presentation: Carol Rohl) Linear Sequence Analysis What can you learn from a (single) protein sequence? Calculate it s physical
ChIP-seq and RNA-seq
 ChIP-seq and RNA-seq Biological Goals Learn how genomes encode the diverse patterns of gene expression that define each cell type and state. Protein-DNA interactions (ChIPchromatin immunoprecipitation)
ChIP-seq and RNA-seq Biological Goals Learn how genomes encode the diverse patterns of gene expression that define each cell type and state. Protein-DNA interactions (ChIPchromatin immunoprecipitation)
Homology Modelling. Thomas Holberg Blicher NNF Center for Protein Research University of Copenhagen
 Homology Modelling Thomas Holberg Blicher NNF Center for Protein Research University of Copenhagen Why are Protein Structures so Interesting? They provide a detailed picture of interesting biological features,
Homology Modelling Thomas Holberg Blicher NNF Center for Protein Research University of Copenhagen Why are Protein Structures so Interesting? They provide a detailed picture of interesting biological features,
Gene Signal Estimates from Exon Arrays
 Gene Signal Estimates from Exon Arrays I. Introduction: With exon arrays like the GeneChip Human Exon 1.0 ST Array, researchers can examine the transcriptional profile of an entire gene (Figure 1). Being
Gene Signal Estimates from Exon Arrays I. Introduction: With exon arrays like the GeneChip Human Exon 1.0 ST Array, researchers can examine the transcriptional profile of an entire gene (Figure 1). Being
Background Analysis and Cross Hybridization. Application
 Background Analysis and Cross Hybridization Application Pius Brzoska, Ph.D. Abstract Microarray technology provides a powerful tool with which to study the coordinate expression of thousands of genes in
Background Analysis and Cross Hybridization Application Pius Brzoska, Ph.D. Abstract Microarray technology provides a powerful tool with which to study the coordinate expression of thousands of genes in
Homework Project Protein Structure Prediction Contact me if you have any questions/problems with the assignment:
 Homework Project Protein Structure Prediction Contact me if you have any questions/problems with the assignment: Roland.Dunbrack@gmail.com For this homework, each of you will be assigned a particular human
Homework Project Protein Structure Prediction Contact me if you have any questions/problems with the assignment: Roland.Dunbrack@gmail.com For this homework, each of you will be assigned a particular human
Database Searching and BLAST Dannie Durand
 Computational Genomics and Molecular Biology, Fall 2013 1 Database Searching and BLAST Dannie Durand Tuesday, October 8th Review: Karlin-Altschul Statistics Recall that a Maximal Segment Pair (MSP) is
Computational Genomics and Molecular Biology, Fall 2013 1 Database Searching and BLAST Dannie Durand Tuesday, October 8th Review: Karlin-Altschul Statistics Recall that a Maximal Segment Pair (MSP) is
Protein Structure Function
 Protein Engineering 15hp Protein Structure Function christofer@xray.bmc.uu.se The Protein Domain 2 What is a domain? An evolutionary conserved, independently folding unit within a protein. A domain often,
Protein Engineering 15hp Protein Structure Function christofer@xray.bmc.uu.se The Protein Domain 2 What is a domain? An evolutionary conserved, independently folding unit within a protein. A domain often,
user s guide Question 1
 Question 1 How does one find a gene of interest and determine that gene s structure? Once the gene has been located on the map, how does one easily examine other genes in that same region? doi:10.1038/ng966
Question 1 How does one find a gene of interest and determine that gene s structure? Once the gene has been located on the map, how does one easily examine other genes in that same region? doi:10.1038/ng966
BIOLOGY 200 Molecular Biology Students registered for the 9:30AM lecture should NOT attend the 4:30PM lecture.
 BIOLOGY 200 Molecular Biology Students registered for the 9:30AM lecture should NOT attend the 4:30PM lecture. Midterm date change! The midterm will be held on October 19th (likely 6-8PM). Contact Kathy
BIOLOGY 200 Molecular Biology Students registered for the 9:30AM lecture should NOT attend the 4:30PM lecture. Midterm date change! The midterm will be held on October 19th (likely 6-8PM). Contact Kathy
Introduction to Proteins
 Introduction to Proteins Lecture 4 Module I: Molecular Structure & Metabolism Molecular Cell Biology Core Course (GSND5200) Matthew Neiditch - Room E450U ICPH matthew.neiditch@umdnj.edu What is a protein?
Introduction to Proteins Lecture 4 Module I: Molecular Structure & Metabolism Molecular Cell Biology Core Course (GSND5200) Matthew Neiditch - Room E450U ICPH matthew.neiditch@umdnj.edu What is a protein?
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature21393 SUPPLEMENTARY TEXT SpMED Immuno-purification and subunit localization Wild-type and subunit deletion mutant SpMEDs (Extended Data Table 1) were immunopurified through TAP-tagged
doi:10.1038/nature21393 SUPPLEMENTARY TEXT SpMED Immuno-purification and subunit localization Wild-type and subunit deletion mutant SpMEDs (Extended Data Table 1) were immunopurified through TAP-tagged
Prediction of Transcription Factors that Regulate Common Binding Motifs Dana Wyman and Emily Alsentzer CS 229, Fall 2014
 Prediction of Transcription Factors that Regulate Common Binding Motifs Dana Wyman and Emily Alsentzer CS 229, Fall 2014 Introduction A. Background Proper regulation of mrna levels is essential to nearly
Prediction of Transcription Factors that Regulate Common Binding Motifs Dana Wyman and Emily Alsentzer CS 229, Fall 2014 Introduction A. Background Proper regulation of mrna levels is essential to nearly
BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers
 BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers Web resources: NCBI database: http://www.ncbi.nlm.nih.gov/ Ensembl database: http://useast.ensembl.org/index.html UCSC
BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers Web resources: NCBI database: http://www.ncbi.nlm.nih.gov/ Ensembl database: http://useast.ensembl.org/index.html UCSC
Using Mapmaker/QTL for QTL mapping
 Using Mapmaker/QTL for QTL mapping M. Maheswaran Tamil Nadu Agriculture University, Coimbatore Mapmaker/QTL overview A number of methods have been developed to map genes controlling quantitatively measured
Using Mapmaker/QTL for QTL mapping M. Maheswaran Tamil Nadu Agriculture University, Coimbatore Mapmaker/QTL overview A number of methods have been developed to map genes controlling quantitatively measured
Project Category: Life Sciences SUNet ID: bentyeh
 Project Category: Life Sciences SUNet ID: bentyeh Predicting Protein Interactions of Intrinsically Disordered Protein Regions Benjamin Yeh, Department of Computer Science, Stanford University CS 229 Project,
Project Category: Life Sciences SUNet ID: bentyeh Predicting Protein Interactions of Intrinsically Disordered Protein Regions Benjamin Yeh, Department of Computer Science, Stanford University CS 229 Project,
Just the Facts: A Basic Introduction to the Science Underlying NCBI Resources
 National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools News About NCBI Site Map
National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools News About NCBI Site Map
: Genomic Regions Enrichment of Annotations Tool
 http://great.stanford.edu/ : Genomic Regions Enrichment of Annotations Tool Gill Bejerano Dept. of Developmental Biology & Dept. of Computer Science Stanford University 1 Human Gene Regulation 10 13 different
http://great.stanford.edu/ : Genomic Regions Enrichment of Annotations Tool Gill Bejerano Dept. of Developmental Biology & Dept. of Computer Science Stanford University 1 Human Gene Regulation 10 13 different
Introduction to NGS analyses
 Introduction to NGS analyses Giorgio L Papadopoulos Institute of Molecular Biology and Biotechnology Bioinformatics Support Group 04/12/2015 Papadopoulos GL (IMBB, FORTH) IMBB NGS Seminar 04/12/2015 1
Introduction to NGS analyses Giorgio L Papadopoulos Institute of Molecular Biology and Biotechnology Bioinformatics Support Group 04/12/2015 Papadopoulos GL (IMBB, FORTH) IMBB NGS Seminar 04/12/2015 1
Differential Gene Expression
 Biology 4361 Developmental Biology Differential Gene Expression September 28, 2006 Chromatin Structure ~140 bp ~60 bp Transcriptional Regulation: 1. Packing prevents access CH 3 2. Acetylation ( C O )
Biology 4361 Developmental Biology Differential Gene Expression September 28, 2006 Chromatin Structure ~140 bp ~60 bp Transcriptional Regulation: 1. Packing prevents access CH 3 2. Acetylation ( C O )
NGS Approaches to Epigenomics
 I519 Introduction to Bioinformatics, 2013 NGS Approaches to Epigenomics Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Background: chromatin structure & DNA methylation Epigenomic
I519 Introduction to Bioinformatics, 2013 NGS Approaches to Epigenomics Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Background: chromatin structure & DNA methylation Epigenomic
EECS 730 Introduction to Bioinformatics Sequence Alignment. Luke Huan Electrical Engineering and Computer Science
 EECS 730 Introduction to Bioinformatics Sequence Alignment Luke Huan Electrical Engineering and Computer Science http://people.eecs.ku.edu/~jhuan/ Database What is database An organized set of data Can
EECS 730 Introduction to Bioinformatics Sequence Alignment Luke Huan Electrical Engineering and Computer Science http://people.eecs.ku.edu/~jhuan/ Database What is database An organized set of data Can
Microarray Data Analysis in GeneSpring GX 11. Month ##, 200X
 Microarray Data Analysis in GeneSpring GX 11 Month ##, 200X Agenda Genome Browser GO GSEA Pathway Analysis Network building Find significant pathways Extract relations via NLP Data Visualization Options
Microarray Data Analysis in GeneSpring GX 11 Month ##, 200X Agenda Genome Browser GO GSEA Pathway Analysis Network building Find significant pathways Extract relations via NLP Data Visualization Options
TALENs (Transcription Activator-Like Effector Nucleases)
 TALENs (Transcription Activator-Like Effector Nucleases) The fundamental rationale between TALENs and ZFNs is similar, namely, combine a sequencespecific DNA-binding peptide domain with a nuclease domain
TALENs (Transcription Activator-Like Effector Nucleases) The fundamental rationale between TALENs and ZFNs is similar, namely, combine a sequencespecific DNA-binding peptide domain with a nuclease domain
Nature Methods: doi: /nmeth.4396
 Supplementary Figure 1 Comparison of technical replicate consistency between and across the standard ATAC-seq method, DNase-seq, and Omni-ATAC. (a) Heatmap-based representation of ATAC-seq quality control
Supplementary Figure 1 Comparison of technical replicate consistency between and across the standard ATAC-seq method, DNase-seq, and Omni-ATAC. (a) Heatmap-based representation of ATAC-seq quality control
Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid
 Supplementary Figures Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid sequence of Drosophila RCC1. Same colors are for Figure 1 with sequence of β-wedge that interacts with Ran in
Supplementary Figures Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid sequence of Drosophila RCC1. Same colors are for Figure 1 with sequence of β-wedge that interacts with Ran in
Conformational properties of enzymes
 Conformational properties of enzymes; Physics of enzyme substrate interactions; Electronic conformational interactions, cooperative properties of enzymes Mitesh Shrestha Conformational properties of enzymes
Conformational properties of enzymes; Physics of enzyme substrate interactions; Electronic conformational interactions, cooperative properties of enzymes Mitesh Shrestha Conformational properties of enzymes
Supplementary Data for DNA sequence+shape kernel enables alignment-free modeling of transcription factor binding.
 Supplementary Data for DNA sequence+shape kernel enables alignment-free modeling of transcription factor binding. Wenxiu Ma 1, Lin Yang 2, Remo Rohs 2, and William Stafford Noble 3 1 Department of Statistics,
Supplementary Data for DNA sequence+shape kernel enables alignment-free modeling of transcription factor binding. Wenxiu Ma 1, Lin Yang 2, Remo Rohs 2, and William Stafford Noble 3 1 Department of Statistics,
Machine learning applications in genomics: practical issues & challenges. Yuzhen Ye School of Informatics and Computing, Indiana University
 Machine learning applications in genomics: practical issues & challenges Yuzhen Ye School of Informatics and Computing, Indiana University Reference Machine learning applications in genetics and genomics
Machine learning applications in genomics: practical issues & challenges Yuzhen Ye School of Informatics and Computing, Indiana University Reference Machine learning applications in genetics and genomics
Problem Set #2
 20.320 Problem Set #2 Due on September 30rd, 2011 at 11:59am. No extensions will be granted. General Instructions: 1. You are expected to state all of your assumptions, and provide step-by-step solutions
20.320 Problem Set #2 Due on September 30rd, 2011 at 11:59am. No extensions will be granted. General Instructions: 1. You are expected to state all of your assumptions, and provide step-by-step solutions
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature16191 SUPPLEMENTARY DISCUSSION We analyzed the current database of solved protein structures to assess the uniqueness of our designed toroids in terms of global similarity and bundle
doi:10.1038/nature16191 SUPPLEMENTARY DISCUSSION We analyzed the current database of solved protein structures to assess the uniqueness of our designed toroids in terms of global similarity and bundle
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Multiple sequence alignments of four Swi2/Snf2 subfamily proteins, ScChd1, SsoRad54 and the RNA helicase Vasa. The sequence alignments of the Swi2/Snf2 subfamily proteins, ScChd1
Supplementary Figure 1 Multiple sequence alignments of four Swi2/Snf2 subfamily proteins, ScChd1, SsoRad54 and the RNA helicase Vasa. The sequence alignments of the Swi2/Snf2 subfamily proteins, ScChd1
Analyzing Gene Set Enrichment
 Analyzing Gene Set Enrichment BaRC Hot Topics June 20, 2016 Yanmei Huang Bioinformatics and Research Computing Whitehead Institute http://barc.wi.mit.edu/hot_topics/ Purpose of Gene Set Enrichment Analysis
Analyzing Gene Set Enrichment BaRC Hot Topics June 20, 2016 Yanmei Huang Bioinformatics and Research Computing Whitehead Institute http://barc.wi.mit.edu/hot_topics/ Purpose of Gene Set Enrichment Analysis
Bacterial Genome Annotation
 Bacterial Genome Annotation Bacterial Genome Annotation For an annotation you want to predict from the sequence, all of... protein-coding genes their stop-start the resulting protein the function the control
Bacterial Genome Annotation Bacterial Genome Annotation For an annotation you want to predict from the sequence, all of... protein-coding genes their stop-start the resulting protein the function the control
Outline. Introduction to ab initio and evidence-based gene finding. Prokaryotic gene predictions
 Outline Introduction to ab initio and evidence-based gene finding Overview of computational gene predictions Different types of eukaryotic gene predictors Common types of gene prediction errors Wilson
Outline Introduction to ab initio and evidence-based gene finding Overview of computational gene predictions Different types of eukaryotic gene predictors Common types of gene prediction errors Wilson
Statistical Methods for Network Analysis of Biological Data
 The Protein Interaction Workshop, 8 12 June 2015, IMS Statistical Methods for Network Analysis of Biological Data Minghua Deng, dengmh@pku.edu.cn School of Mathematical Sciences Center for Quantitative
The Protein Interaction Workshop, 8 12 June 2015, IMS Statistical Methods for Network Analysis of Biological Data Minghua Deng, dengmh@pku.edu.cn School of Mathematical Sciences Center for Quantitative
Transcription Gene regulation
 Transcription Gene regulation The machine that transcribes a gene is composed of perhaps 50 proteins, including RNA polymerase, the enzyme that converts DNA code into RNA code. A crew of transcription
Transcription Gene regulation The machine that transcribes a gene is composed of perhaps 50 proteins, including RNA polymerase, the enzyme that converts DNA code into RNA code. A crew of transcription
Regulation of gene expression. (Lehninger pg )
 Regulation of gene expression (Lehninger pg. 1072-1085) Today s lecture Gene expression Constitutive, inducible, repressible genes Specificity factors, activators, repressors Negative and positive gene
Regulation of gene expression (Lehninger pg. 1072-1085) Today s lecture Gene expression Constitutive, inducible, repressible genes Specificity factors, activators, repressors Negative and positive gene
A property-based analysis of human transcription factors
 Bahrami et al. BMC Research Notes (2015) 8:82 DOI 10.1186/s13104-015-1039-6 RESEARCH ARTICLE Open Access A property-based analysis of human transcription factors Shahram Bahrami 1,2, Rezvan Ehsani 1 and
Bahrami et al. BMC Research Notes (2015) 8:82 DOI 10.1186/s13104-015-1039-6 RESEARCH ARTICLE Open Access A property-based analysis of human transcription factors Shahram Bahrami 1,2, Rezvan Ehsani 1 and
Week 1 BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers
 Week 1 BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers Web resources: NCBI database: http://www.ncbi.nlm.nih.gov/ Ensembl database: http://useast.ensembl.org/index.html
Week 1 BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers Web resources: NCBI database: http://www.ncbi.nlm.nih.gov/ Ensembl database: http://useast.ensembl.org/index.html
Bi 8 Lecture 7. Ellen Rothenberg 26 January Reading: Ch. 3, pp ; panel 3-1
 Bi 8 Lecture 7 PROTEIN STRUCTURE, Functional analysis, and evolution Ellen Rothenberg 26 January 2016 Reading: Ch. 3, pp. 109-134; panel 3-1 (end with free amine) aromatic, hydrophobic small, hydrophilic
Bi 8 Lecture 7 PROTEIN STRUCTURE, Functional analysis, and evolution Ellen Rothenberg 26 January 2016 Reading: Ch. 3, pp. 109-134; panel 3-1 (end with free amine) aromatic, hydrophobic small, hydrophilic
Comparative Modeling Part 1. Jaroslaw Pillardy Computational Biology Service Unit Cornell Theory Center
 Comparative Modeling Part 1 Jaroslaw Pillardy Computational Biology Service Unit Cornell Theory Center Function is the most important feature of a protein Function is related to structure Structure is
Comparative Modeling Part 1 Jaroslaw Pillardy Computational Biology Service Unit Cornell Theory Center Function is the most important feature of a protein Function is related to structure Structure is
Complex Transcription Machinery
 Complex Transcription Machinery Subunits of the Basal Txn Apparatus Stepwise Assembly of the Pre-initiation Complex Interplay of Activators, Co-regulators and RNA Polymerase at the Promoter Divide and
Complex Transcription Machinery Subunits of the Basal Txn Apparatus Stepwise Assembly of the Pre-initiation Complex Interplay of Activators, Co-regulators and RNA Polymerase at the Promoter Divide and
