Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
|
|
|
- Abel Baldwin
- 5 years ago
- Views:
Transcription
1 Supplementary Figure 1 Multiple sequence alignments of four Swi2/Snf2 subfamily proteins, ScChd1, SsoRad54 and the RNA helicase Vasa. The sequence alignments of the Swi2/Snf2 subfamily proteins, ScChd1 and SsoRad54 were done with Clustal Omega, which were further aligned with the sequence of Vasa according to structural superposition (PDB code 2DB3). Secondary structural assignments labeled on the top are based on the structure determined in this study, and the helicase motifs are assigned as reported 23. The assignment of HSA domain is based on the reported structure (PDB code 4I6M) 20. The SnAc domain is labeled as an orange bar. The residues numbering is based on the sequence of MtSnf2. The suppressor mutants found in ScSth1 are labeled as magenta dots (in posthsa) and yellow dots (in supph) 18. Blue dots, the DNA-binding residues identified in this study; blue triangles, arginine fingers; red triangle, cancer-associated mutations found in human Brg1 gene 11, Bartlett, C., Orvis, T.J., Rosson, G.S. & Weissman, B.E. BRG1 mutations found in human cancer cell lines inactivate Rb-mediated cell-cycle
2 arrest. J. Cell Physiol. 226, (2011). 51. Medina, P.P. et al. Genetic and epigenetic screening for gene alterations of the chromatin-remodeling factor, SMARCA4/BRG1, in lung tumors. Genes Chromosomes Cancer 41, (2004). 52. Medina, P.P. et al. Frequent BRG1/SMARCA4-inactivating mutations in human lung cancer cell lines. Hum. Mutat. 29, (2008). 53. Wong, A.K. et al. BRG1, a component of the SWI-SNF complex, is mutated in multiple human tumor cell lines. Cancer Res. 60, (2000).
3 Supplementary Figure 2 SAXS measurements of MtSnf2. (a) Three different views of the overlay of the final average ab initio molecular envelope of MtSnf2 ( ) reconstructed from SAXS measurements (grey) with the docked crystal structure. The protein is colored as in Fig.1, and the N-and C-termini are labeled. Additional densities near the N- and C-termini can be attributed to the disordered residues at both ends (the first 13 and the last 48 residues, respectively). (b) Scattering intensity in arbitrary units versus momentum transfer q in Å -1 for MtSnf2. The linear fitting in the Guinier region of scattering curve (inset) indicates that MtSnf2 is monodisperse and homogenuous in solution. (b) The dimensionless Kratkyplot of MtSnf2 has typical feature for folded protein with disordered regions. (d) Pair distance distribution function (PDDF) of MtSnf2 with D max = 117 Å calculated using GNOM (q max =0.30 Å -1 ). (e) Fitting the theoretical scattering curve (red line) of MtSnf2 crystal structure to the experimental scattering curve (black circles), with a chi of 2.1. The theoretical SAXS curve agrees well with the SAXS measurement in solution, further validating the observed conformation in the crystals.
4 Supplementary Figure 3 Comparisons with the SF2 family proteins SsoRad54 and Vasa. (a) Structural alignment of the core1 domains of MtSnf2 (green) and SsoRad54 (grey). The DNA bound by SsoRad54 is shown as ribbon presentation. The DNA-binding sites of the core1 domains of MtSnf2 and SsoRad54 align very well, with K665 and K692 of MtSnf2 located at the same positions as R547 and K573 of SsoRad54. K687 of MtSnf2 is disordered in current structure, and aligned with K568 of SsoRad54 in the primary sequence (Supplementary Figure 1). (b) Structural alignment of the core2 domains of MtSnf2 (cyan) and SsoRad54 (grey). The arginine fingers of SsoRad54 are labeled (R840 and R843). The DNA binds to the surface of the core2 domain of SsoRad54 at a position in conflict with the SnAc domain (orange) of MtSnf2, suggesting MtSnf2 interacts with DNA in a different manner. Instead, biochemical analyses indicated that DNA contacts the core2 domains of both proteins in solution at motif V (around R950 of MtSnf2 and K808 of SsoRad54). R832 of MtSnf2 is also involved in DNA binding, which is absence in SsoRad54. (c) Structural alignment of the core1 domains of MtSnf2 (green) and Vasa helicase (grey). The bound RNA by Vasa is shown as stick presentation. Motif I of MtSnf2 is colored red, with bound sulfate ion as sphere presentation. The DNAbinding sites of MtSnf2 (K665, K687 and K692) identified in this study are in close proximity to the RNA bound by Vasa. (d) Structural alignment of the core2 domains of MtSnf2 (green) and Vasa (grey). Motif VI of MtSnf2 is colored red. The DNA-binding site of R950 in MtSnf2 is close to the RNA bound by Vasa.
5 Supplementary Figure 4 Interactions between post-hsa and supph of MtSnf2. (a) Structure of the posthsa-supph region of MtSnf2. The structure is colored as in Fig.1 with the supph helix in yellow. The suppressor mutants of ScSth1 are mapped to the corresponding positions of MtSnf2 (showed as stick models) 18. Residues are labeled based on the sequence of MtSnf2, and the residues in the parentheses are the corresponding suppressor mutants of ScSth1. Three suppressor mutations are located at posthsa (N384K, D385Yand L392V) and four at supph (E676Q, L680M/V, L681F and K688T). Two mutations at the HSA domain (T373P and K382N) are not present in the current structure. (b) Interactions between supph, posthsa and the core1 domain. The core1 domain is shown as surface presentation, and colored coded by the spectra of blue-to-white (decreasing conservation). The posthsa and supph helices bind to a conserved surface of the core1 domain through hydrophobic interactions, and they also interact with each other through specific H-bonds. (c) Multiple sequence alignments of the Swi2/Snf2 subfamily proteins around the posthsa and supph regions. The suppressor mutations of Sth1 are highlighted in magenta and yellow at the posthsa and supph helices, respectively.
6 Supplementary Figure 5 Chromatin-remodeling activities of various constructs used in this study. (a) Gels of the restriction enzyme-accessibility assays of four core1-core2 interface mutants of MtSnf2 ( ). The cut fractions were quantified and shown in Fig. 3d. Three independent assays were performed and one was showed. (b) Gels of the restriction enzyme-accessibility assays of MtSnf2 ( ) with the WT interface and five DNA-binding mutants. The cut fractions were shown in Fig. 4e. (c) Gels of the restriction enzyme-accessibility assays for ScSnf2 with different N-terminal boundaries. The catalytic core of ScSnf2 ( ), left panel; half HSAcontaining ScSnf2 ( ), middle panel; full HSA-containing ScSnf2 ( ) in complex with Arp7-Arp9-Rtt102 (Snf2 tetramer), right panel. The cut fractions were quantified and shown in Fig. 5c. (d) Gels of the restriction enzyme-accessibility assays of MtSnf2 with different N- terminal boundaries. The catalytic core MtSnf2 ( ), left panel; half HSA-containing MtSnf2 ( ), middle panel. The cut fractions were shown on the right. (e) The ATPase activities of MtSnf2 with different N-terminal boundaries in the absence (-) and presence of DNA/NCP. MtSnf2 ( ), black; MtSnf2 ( ), white. (f) Gels of the restriction enzyme-accessibility assays of MtSnf2 with truncation of the C- terminal flexible tail at different positions. The cut fractions were quantified and shown in Fig. 6e.
7 Supplementary Figure 6 Nucleosome binding activities of various constructs used in this study. (a) EMSA of the nucleosome binding activities of MtSnf2 ( ) with the WT interface and five DNA-binding mutants. 2 nm Cy5-labelled 147 bp NCP was mixed with increasing amounts of proteins (in nm). The samples were resolved by electrophoresis on 4.5% native acrylamide gels, and the fluorescent bands were detected by Typhoon Trio+ imager. With higher concentrations of the proteins, the bound NCPs migrate more slowly, suggesting a heterogeneous population of the protein-bound NCP in the solution, and one NCP might bind more than one molecule of the protein. The star * sign indicates the position of free DNA. The bound fractions were quantified as the disappearance of free NCP relative to the intensity of the whole lane, and shown in Fig. 4c. (b) EMSA of nucleosome binding of three constructs of ScSnf2. The quantification of the bound fractions was showed in Fig. 5a. (c) EMSA of nucleosome binding of MtSnf2 ( ). EMSA of MtSnf2 ( ) was shown in Supplementary Fig. 6a, the left panel (WT interface). Quantification of the bound fractions is shown (right panel). The apparent Kds of the catalytic core MtSnf2 ( ) and the HSA-containing MtSnf2 ( ) are about 150 nm and 190 nm, respectively. (d) EMSA of nucleosome binding of MtSnf2 with truncation of the C-terminal flexible tail at different positions. EMSA of MtSnf2 ( ) was shown in Supplementary Fig. 6a, the left panel (WT interface). The quantification of the bound fractions is showed in Fig. 6c.
Supplementary Online Material. Structural mimicry in transcription regulation of human RNA polymerase II by the. DNA helicase RECQL5
 Supplementary Online Material Structural mimicry in transcription regulation of human RNA polymerase II by the DNA helicase RECQL5 Susanne A. Kassube, Martin Jinek, Jie Fang, Susan Tsutakawa and Eva Nogales
Supplementary Online Material Structural mimicry in transcription regulation of human RNA polymerase II by the DNA helicase RECQL5 Susanne A. Kassube, Martin Jinek, Jie Fang, Susan Tsutakawa and Eva Nogales
Supplemental Information Molecular Cell, Volume 41
 Supplemental Information Molecular Cell, Volume 41 Molecular Mechanisms for the RNA-Dependent ATPase Activity of Upf1 and Its Regulation by Upf2 Sutapa Chakrabarti, Uma Jayachandran, Fabien Bonneau, Francesca
Supplemental Information Molecular Cell, Volume 41 Molecular Mechanisms for the RNA-Dependent ATPase Activity of Upf1 and Its Regulation by Upf2 Sutapa Chakrabarti, Uma Jayachandran, Fabien Bonneau, Francesca
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Conserved arginines on the rim of Hfq catalyze base pair formation and exchange Subrata Panja and Sarah A. Woodson T.C. Jenkins Department of Biophysics, Johns Hopkins University,
SUPPLEMENTARY INFORMATION Conserved arginines on the rim of Hfq catalyze base pair formation and exchange Subrata Panja and Sarah A. Woodson T.C. Jenkins Department of Biophysics, Johns Hopkins University,
Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid
 Supplementary Figures Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid sequence of Drosophila RCC1. Same colors are for Figure 1 with sequence of β-wedge that interacts with Ran in
Supplementary Figures Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid sequence of Drosophila RCC1. Same colors are for Figure 1 with sequence of β-wedge that interacts with Ran in
SUPPLEMENTARY INFORMATION. Supplementary Figures 1-8
 SUPPLEMENTARY INFORMATION Supplementary Figures 1-8 Supplementary Figure 1. TFAM residues contacting the DNA minor groove (A) TFAM contacts on nonspecific DNA. Leu58, Ile81, Asn163, Pro178, and Leu182
SUPPLEMENTARY INFORMATION Supplementary Figures 1-8 Supplementary Figure 1. TFAM residues contacting the DNA minor groove (A) TFAM contacts on nonspecific DNA. Leu58, Ile81, Asn163, Pro178, and Leu182
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
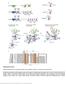 Supplementary Figure 1 Domain architecture and conformational states of the decapping complex, as revealed by structural studies. (a) Domain organization of Schizosaccharomyces pombe (Sp) and Saccharomyces
Supplementary Figure 1 Domain architecture and conformational states of the decapping complex, as revealed by structural studies. (a) Domain organization of Schizosaccharomyces pombe (Sp) and Saccharomyces
Nature Structural & Molecular Biology: doi: /nsmb.3428
 Supplementary Figure 1 Biochemical characterization of the monou and oligou activity switch of TUT4(7). (a) Mouse TUT4 and human TUT7 were assayed for monou and Lin28-dependent oligou addition activities
Supplementary Figure 1 Biochemical characterization of the monou and oligou activity switch of TUT4(7). (a) Mouse TUT4 and human TUT7 were assayed for monou and Lin28-dependent oligou addition activities
Supplementary Figure S1. Binding of HSA mutants to hfcrn. (a) The levels of titrated amounts of HSA
 Supplementary Figure S1. Binding of HSA mutants to hfcrn. (a) The levels of titrated amounts of HSA variants (5.0-0.002 μg/ml) directly coated in the wells at ph 6.0 were controlled using a horseradish
Supplementary Figure S1. Binding of HSA mutants to hfcrn. (a) The levels of titrated amounts of HSA variants (5.0-0.002 μg/ml) directly coated in the wells at ph 6.0 were controlled using a horseradish
Supplementary Figure 1 Two distinct conformational states of the HNH domain in crystal structures. a
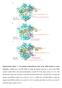 Supplementary Figure 1 Two distinct conformational states of the HNH domain in crystal structures. a HNH-state 1 in PDB 4OO8, in which the distance from the C atom of the HNH catalytic residue 840 to the
Supplementary Figure 1 Two distinct conformational states of the HNH domain in crystal structures. a HNH-state 1 in PDB 4OO8, in which the distance from the C atom of the HNH catalytic residue 840 to the
RNP purification, components and activity.
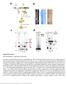 Supplementary Figure 1 RNP purification, components and activity. (a) Intein-mediated RNP production and purification. The Ll.LtrB intron RNA (red) (Exons E1 and E2 in green) and associated intron encoded
Supplementary Figure 1 RNP purification, components and activity. (a) Intein-mediated RNP production and purification. The Ll.LtrB intron RNA (red) (Exons E1 and E2 in green) and associated intron encoded
Supplementary Table 1. DNA sequence synthesized to express the Zika virus NS5 protein.
 Supplementary Table 1. DNA sequence synthesized to express the Zika virus NS5 protein. MR766 NS5 sequence 1 ACAGAGAACAGATTGGTGGTGGAGGTGGGACGGGAGAGACTCTGGGAGAGAAGTGGAAAG 61 CTCGTCTGAATCAGATGTCGGCCCTGGAGTTCTACTCTTATAAAAAGTCAGGTATCACTG
Supplementary Table 1. DNA sequence synthesized to express the Zika virus NS5 protein. MR766 NS5 sequence 1 ACAGAGAACAGATTGGTGGTGGAGGTGGGACGGGAGAGACTCTGGGAGAGAAGTGGAAAG 61 CTCGTCTGAATCAGATGTCGGCCCTGGAGTTCTACTCTTATAAAAAGTCAGGTATCACTG
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature08627 Supplementary Figure 1. DNA sequences used to construct nucleosomes in this work. a, DNA sequences containing the 601 positioning sequence (blue)24 with a PstI restriction site
doi: 10.1038/nature08627 Supplementary Figure 1. DNA sequences used to construct nucleosomes in this work. a, DNA sequences containing the 601 positioning sequence (blue)24 with a PstI restriction site
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature06147 SUPPLEMENTARY INFORMATION Figure S1 The genomic and domain structure of Dscam. The Dscam gene comprises 24 exons, encoding a signal peptide (SP), 10 IgSF domains, 6 fibronectin
doi: 10.1038/nature06147 SUPPLEMENTARY INFORMATION Figure S1 The genomic and domain structure of Dscam. The Dscam gene comprises 24 exons, encoding a signal peptide (SP), 10 IgSF domains, 6 fibronectin
Chapter 14 Regulation of Transcription
 Chapter 14 Regulation of Transcription Cis-acting sequences Distance-independent cis-acting elements Dissecting regulatory elements Transcription factors Overview transcriptional regulation Transcription
Chapter 14 Regulation of Transcription Cis-acting sequences Distance-independent cis-acting elements Dissecting regulatory elements Transcription factors Overview transcriptional regulation Transcription
Supplementary Information. Structural basis for duplex RNA recognition and cleavage by A.
 Supplementary Information Structural asis for duplex RNA recognition and cleavage y A. fulgidus C3PO Eneida arizotto 1, Edward D Lowe 1 & James S Parker 1 1 Department of Biochemistry University of Oxford
Supplementary Information Structural asis for duplex RNA recognition and cleavage y A. fulgidus C3PO Eneida arizotto 1, Edward D Lowe 1 & James S Parker 1 1 Department of Biochemistry University of Oxford
Visualization of codon-dependent conformational rearrangements during translation termination
 Supplementary information for: Visualization of codon-dependent conformational rearrangements during translation termination Shan L. He 1 and Rachel Green 1 1 Howard Hughes Medical Institute, Department
Supplementary information for: Visualization of codon-dependent conformational rearrangements during translation termination Shan L. He 1 and Rachel Green 1 1 Howard Hughes Medical Institute, Department
Nature Structural & Molecular Biology: doi: /nsmb.2996
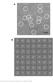 Supplementary Figure 1 Negative-stain EM of native cytoplasmic dynein. (a) A representative micrograph of negatively stained dynein, with the flexible dimeric molecules circled in white. (b) Reference-free
Supplementary Figure 1 Negative-stain EM of native cytoplasmic dynein. (a) A representative micrograph of negatively stained dynein, with the flexible dimeric molecules circled in white. (b) Reference-free
Supplementary Information for. Structure of human tyrosylprotein sulfotransferase-2 reveals the mechanism of protein tyrosine sulfation reaction
 Supplementary Information for Structure of human tyrosylprotein sulfotransferase-2 reveals the mechanism of protein tyrosine sulfation reaction Takamasa Teramoto, Yukari Fujikawa, Yoshirou Kawaguchi, Katsuhisa
Supplementary Information for Structure of human tyrosylprotein sulfotransferase-2 reveals the mechanism of protein tyrosine sulfation reaction Takamasa Teramoto, Yukari Fujikawa, Yoshirou Kawaguchi, Katsuhisa
HEK293T. Fig. 1 in the
 Supplementary Information Supplementary Figure 1 Zinc uptake assay of hzip4 and hzip4-δecd transiently expressed in HEK293T cells. The results of one representative e experiment are shown in Fig. 1 in
Supplementary Information Supplementary Figure 1 Zinc uptake assay of hzip4 and hzip4-δecd transiently expressed in HEK293T cells. The results of one representative e experiment are shown in Fig. 1 in
SUPPLEMENTARY INFORMATION
 Supplementary Table 1. Crystallographic statistics CRM1-SNUPN complex Space group P6 4 22 a=b=250.4, c=190.4 Data collection statistics: CRM1-selenomethionine SNUPN MAD data Peak Inflection Remote Native
Supplementary Table 1. Crystallographic statistics CRM1-SNUPN complex Space group P6 4 22 a=b=250.4, c=190.4 Data collection statistics: CRM1-selenomethionine SNUPN MAD data Peak Inflection Remote Native
SUPPLEMENTARY INFORMATION
 Supplementary Figure 1. Mechanism for signal-induced opening of the DNA box. a, An atomic model of the DNA box held closed by locks (orange and blue) that are double helices formed by two short strands
Supplementary Figure 1. Mechanism for signal-induced opening of the DNA box. a, An atomic model of the DNA box held closed by locks (orange and blue) that are double helices formed by two short strands
SUPPLEMENTARY INFORMATION
 Results Construct purification and coupling. Two A1-GP1bα ReaLiSM constructs, with and without cysteine residues near the N and C-termini (Fig. S2a), were expressed and purified by Ni affinity chromatography
Results Construct purification and coupling. Two A1-GP1bα ReaLiSM constructs, with and without cysteine residues near the N and C-termini (Fig. S2a), were expressed and purified by Ni affinity chromatography
Nature Structural & Molecular Biology: doi: /nsmb.2548
 Supplementary Figure 1. Structure of GltPhout. (a) Stereo view of a slice through a single GltPhout protomer shown in stick representation along with 2Fo-Fc and anomalous difference electron maps. The
Supplementary Figure 1. Structure of GltPhout. (a) Stereo view of a slice through a single GltPhout protomer shown in stick representation along with 2Fo-Fc and anomalous difference electron maps. The
Hmwk # 8 : DNA-Binding Proteins : Part II
 The purpose of this exercise is : Hmwk # 8 : DNA-Binding Proteins : Part II 1). to examine the case of a tandem head-to-tail homodimer binding to DNA 2). to view a Zn finger motif 3). to consider the case
The purpose of this exercise is : Hmwk # 8 : DNA-Binding Proteins : Part II 1). to examine the case of a tandem head-to-tail homodimer binding to DNA 2). to view a Zn finger motif 3). to consider the case
Sam68 STARR Sam68 QUA1- KH. p(r ) [Å] [Å] TSTAR STAR. Sam68 QUA1-KH and. constructs are
![Sam68 STARR Sam68 QUA1- KH. p(r ) [Å] [Å] TSTAR STAR. Sam68 QUA1-KH and. constructs are Sam68 STARR Sam68 QUA1- KH. p(r ) [Å] [Å] TSTAR STAR. Sam68 QUA1-KH and. constructs are](/thumbs/73/69166612.jpg) a b Sam68 STARR Sam68 QUA1- KH c d e ) p(r p(r ) r [Å] r [Å] Supplementary Figure 1: The QUA2 domain is not involved in i the overall conformation of the STAR domain (a) Overlay of T-STAR QUA1-KH in complex
a b Sam68 STARR Sam68 QUA1- KH c d e ) p(r p(r ) r [Å] r [Å] Supplementary Figure 1: The QUA2 domain is not involved in i the overall conformation of the STAR domain (a) Overlay of T-STAR QUA1-KH in complex
Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than. SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with
 Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with concentration of 800nM) were incubated with 1mM dgtp for the indicated
Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with concentration of 800nM) were incubated with 1mM dgtp for the indicated
SUPPLEMENTARY INFORMATION. Reengineering Protein Interfaces Yields Copper-Inducible Ferritin Cage Assembly
 SUPPLEMENTARY INFORMATION Reengineering Protein Interfaces Yields Copper-Inducible Ferritin Cage Assembly Dustin J. E. Huard, Kathleen M. Kane and F. Akif Tezcan* Department of Chemistry and Biochemistry,
SUPPLEMENTARY INFORMATION Reengineering Protein Interfaces Yields Copper-Inducible Ferritin Cage Assembly Dustin J. E. Huard, Kathleen M. Kane and F. Akif Tezcan* Department of Chemistry and Biochemistry,
Supplementary Figure 1. FRET probe labeling locations in the Cas9-RNA-DNA complex.
 Supplementary Figure 1. FRET probe labeling locations in the Cas9-RNA-DNA complex. (a) Cy3 and Cy5 labeling locations shown in the crystal structure of Cas9-RNA bound to a cognate DNA target (PDB ID: 4UN3)
Supplementary Figure 1. FRET probe labeling locations in the Cas9-RNA-DNA complex. (a) Cy3 and Cy5 labeling locations shown in the crystal structure of Cas9-RNA bound to a cognate DNA target (PDB ID: 4UN3)
SUPPLEMENTAL MATERIAL BIOCHEMICAL AND STRUCTURAL STUDIES ON THE M. TUBERCULOSIS O 6 -METHYLGUANINE METHYLTRANSFERASE AND MUTATED VARIANTS
 SUPPLEMENTAL MATERIAL BIOCHEMICAL AND STRUCTURAL STUDIES ON THE M. TUBERCULOSIS O 6 -METHYLGUANINE METHYLTRANSFERASE AND MUTATED VARIANTS Riccardo Miggiano 1, Valentina Casazza 1, Silvia Garavaglia 1,
SUPPLEMENTAL MATERIAL BIOCHEMICAL AND STRUCTURAL STUDIES ON THE M. TUBERCULOSIS O 6 -METHYLGUANINE METHYLTRANSFERASE AND MUTATED VARIANTS Riccardo Miggiano 1, Valentina Casazza 1, Silvia Garavaglia 1,
SUPPLEMENTARY INFORMATION. Chemical modulation of Chaperone-mediated autophagy by novel
 SUPPLEMENTARY INFORMATION Chemical modulation of Chaperone-mediated autophagy by novel retinoic acid derivatives Jaime Anguiano 1, Thomas P Garner 2, Murugesan Mahalingam 1, Bhaskar C. Das 1, Evripidis
SUPPLEMENTARY INFORMATION Chemical modulation of Chaperone-mediated autophagy by novel retinoic acid derivatives Jaime Anguiano 1, Thomas P Garner 2, Murugesan Mahalingam 1, Bhaskar C. Das 1, Evripidis
has only one nucleotide, U20, between G19 and A21, while trna Glu CUC has two
 SPPLEMENTRY INFORMTION doi:1.138/nature9411 Supplementary Discussion The structural characteristics of trn Gln G in comparison to trn Glu 16 in trn Gln G is directed towards the G19 56 pair, or the outer
SPPLEMENTRY INFORMTION doi:1.138/nature9411 Supplementary Discussion The structural characteristics of trn Gln G in comparison to trn Glu 16 in trn Gln G is directed towards the G19 56 pair, or the outer
Supplementatry Fig 1. Domain structure, biophysical characterisation and electron microscopy of a TD. (a) XTACC3/Maskin and XMAP215/chTOG domain
 Supplementatry Fig 1. Domain structure, biophysical characterisation and electron microscopy of a TD. (a) XTACC3/Maskin and XMAP215/chTOG domain architecture. Various C-terminal fragments were cloned and
Supplementatry Fig 1. Domain structure, biophysical characterisation and electron microscopy of a TD. (a) XTACC3/Maskin and XMAP215/chTOG domain architecture. Various C-terminal fragments were cloned and
Why does this matter?
 Background Why does this matter? Better understanding of how the nucleosome affects transcription Important for understanding the nucleosome s role in gene expression Treats each component and region of
Background Why does this matter? Better understanding of how the nucleosome affects transcription Important for understanding the nucleosome s role in gene expression Treats each component and region of
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature10258 Supplementary Figure 1 Reconstitution of the CENP-A nucleosome with recombinant human histones H2A, H2B, H4, and CENP-A. a, Purified recombinant human
SUPPLEMENTARY INFORMATION doi:10.1038/nature10258 Supplementary Figure 1 Reconstitution of the CENP-A nucleosome with recombinant human histones H2A, H2B, H4, and CENP-A. a, Purified recombinant human
Molecular design principles underlying β-strand swapping. in the adhesive dimerization of cadherins
 Supplementary information for: Molecular design principles underlying β-strand swapping in the adhesive dimerization of cadherins Jeremie Vendome 1,2,3,5, Shoshana Posy 1,2,3,5,6, Xiangshu Jin, 1,3 Fabiana
Supplementary information for: Molecular design principles underlying β-strand swapping in the adhesive dimerization of cadherins Jeremie Vendome 1,2,3,5, Shoshana Posy 1,2,3,5,6, Xiangshu Jin, 1,3 Fabiana
Chapter 8 DNA Recognition in Prokaryotes by Helix-Turn-Helix Motifs
 Chapter 8 DNA Recognition in Prokaryotes by Helix-Turn-Helix Motifs 1. Helix-turn-helix proteins 2. Zinc finger proteins 3. Leucine zipper proteins 4. Beta-scaffold factors 5. Others λ-repressor AND CRO
Chapter 8 DNA Recognition in Prokaryotes by Helix-Turn-Helix Motifs 1. Helix-turn-helix proteins 2. Zinc finger proteins 3. Leucine zipper proteins 4. Beta-scaffold factors 5. Others λ-repressor AND CRO
Nature Structural & Molecular Biology: doi: /nsmb.3175
 Supplementary Figure 1 Medium- and low-fret groups probably representing varying positions on the DNA show similar changes between H3 and CENP-A nucleosomes with and without CENP-C CD, as does the high-fret
Supplementary Figure 1 Medium- and low-fret groups probably representing varying positions on the DNA show similar changes between H3 and CENP-A nucleosomes with and without CENP-C CD, as does the high-fret
Nature Methods: doi: /nmeth Supplementary Figure 1. Construction of a sensitive TetR mediated auxotrophic off-switch.
 Supplementary Figure 1 Construction of a sensitive TetR mediated auxotrophic off-switch. A Production of the Tet repressor in yeast when conjugated to either the LexA4 or LexA8 promoter DNA binding sequences.
Supplementary Figure 1 Construction of a sensitive TetR mediated auxotrophic off-switch. A Production of the Tet repressor in yeast when conjugated to either the LexA4 or LexA8 promoter DNA binding sequences.
SUPPLEMENTAL FIGURE LEGENDS. Figure S1: Homology alignment of DDR2 amino acid sequence. Shown are
 SUPPLEMENTAL FIGURE LEGENDS Figure S1: Homology alignment of DDR2 amino acid sequence. Shown are the amino acid sequences of human DDR2, mouse DDR2 and the closest homologs in zebrafish and C. Elegans.
SUPPLEMENTAL FIGURE LEGENDS Figure S1: Homology alignment of DDR2 amino acid sequence. Shown are the amino acid sequences of human DDR2, mouse DDR2 and the closest homologs in zebrafish and C. Elegans.
The molecular basis of lysine 48 ubiquitin chain synthesis by Ube2K
 Supplementary Information The molecular basis of lysine 48 ubiquitin chain synthesis by Adam J. Middleton, Catherine L. Day* Department of Biochemistry, Otago School of Medical Sciences, University of
Supplementary Information The molecular basis of lysine 48 ubiquitin chain synthesis by Adam J. Middleton, Catherine L. Day* Department of Biochemistry, Otago School of Medical Sciences, University of
Solutions to 7.02 Quiz II 10/27/05
 Solutions to 7.02 Quiz II 10/27/05 Class Average = 83 Standard Deviation = 9 Range Grade % 87-100 A 43 74-86 B 39 55-73 C 17 > 54 D 1 Question 1 (56 points) While studying deep sea bacteria, you discover
Solutions to 7.02 Quiz II 10/27/05 Class Average = 83 Standard Deviation = 9 Range Grade % 87-100 A 43 74-86 B 39 55-73 C 17 > 54 D 1 Question 1 (56 points) While studying deep sea bacteria, you discover
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11726 Supplementary Figure 1 SRP RNA constructs used in single molecule experiments. a, Secondary structure of wildtype E. coli SRP RNA, which forms an elongated hairpin structure capped
doi:10.1038/nature11726 Supplementary Figure 1 SRP RNA constructs used in single molecule experiments. a, Secondary structure of wildtype E. coli SRP RNA, which forms an elongated hairpin structure capped
Sequence Determinants of a Conformational Switch in a Protein
 Sequence Determinants of a Conformational Switch in a Protein Thomas A. Anderson, Matthew H. J. Cordes, and Robert T. Sauer Kyle Skalenko 4/17/09 Introduction Protein folding is guided by the following
Sequence Determinants of a Conformational Switch in a Protein Thomas A. Anderson, Matthew H. J. Cordes, and Robert T. Sauer Kyle Skalenko 4/17/09 Introduction Protein folding is guided by the following
X-ray structures of fructosyl peptide oxidases revealing residues responsible for gating oxygen access in the oxidative half reaction
 X-ray structures of fructosyl peptide oxidases revealing residues responsible for gating oxygen access in the oxidative half reaction Tomohisa Shimasaki 1, Hiromi Yoshida 2, Shigehiro Kamitori 2 & Koji
X-ray structures of fructosyl peptide oxidases revealing residues responsible for gating oxygen access in the oxidative half reaction Tomohisa Shimasaki 1, Hiromi Yoshida 2, Shigehiro Kamitori 2 & Koji
Plant Molecular and Cellular Biology Lecture 9: Nuclear Genome Organization: Chromosome Structure, Chromatin, DNA Packaging, Mitosis Gary Peter
 Plant Molecular and Cellular Biology Lecture 9: Nuclear Genome Organization: Chromosome Structure, Chromatin, DNA Packaging, Mitosis Gary Peter 9/16/2008 1 Learning Objectives 1. List and explain how DNA
Plant Molecular and Cellular Biology Lecture 9: Nuclear Genome Organization: Chromosome Structure, Chromatin, DNA Packaging, Mitosis Gary Peter 9/16/2008 1 Learning Objectives 1. List and explain how DNA
Supplementary Figure 1 Telomerase RNA fragments used in single-molecule FRET experiments. A pseudoknot fragment (nts ) labeled at position U42
 Supplementary Figure 1 Telomerase RNA fragments used in single-molecule FRET experiments. A pseudoknot fragment (nts 32-195) labeled at position U42 with Cy3 (green circle) was constructed by a two piece
Supplementary Figure 1 Telomerase RNA fragments used in single-molecule FRET experiments. A pseudoknot fragment (nts 32-195) labeled at position U42 with Cy3 (green circle) was constructed by a two piece
Supplemental Figure 1
 Supplemental Figure 1 Supplemental Fig. 1. (A) The binding avidity between Fz4-FcH6 and MBP-Norrin dimers measured by biolayer interferometry. 5 µg/ml Fz4-Fc protein was loaded onto anti human Fc capture
Supplemental Figure 1 Supplemental Fig. 1. (A) The binding avidity between Fz4-FcH6 and MBP-Norrin dimers measured by biolayer interferometry. 5 µg/ml Fz4-Fc protein was loaded onto anti human Fc capture
(a) Overview of the 2-helix bundle (2HB) nanospring design used in this study. The
 1 Supplementary Figure 1 Design of the DNA origami spring (nanospring). (a) Overview of the 2-helix bundle (2HB) nanospring design used in this study. The scheme was produced by cadnano software 1. Scaffold,
1 Supplementary Figure 1 Design of the DNA origami spring (nanospring). (a) Overview of the 2-helix bundle (2HB) nanospring design used in this study. The scheme was produced by cadnano software 1. Scaffold,
PDIP46 (DNA polymerase δ interacting protein 46) is an activating factor for human DNA polymerase δ
 PDIP46 (DNA polymerase δ interacting protein 46) is an activating factor for human DNA polymerase δ Supplementary Material Figure S1. PDIP46 is associated with Pol isolated by immunoaffinity chromatography.
PDIP46 (DNA polymerase δ interacting protein 46) is an activating factor for human DNA polymerase δ Supplementary Material Figure S1. PDIP46 is associated with Pol isolated by immunoaffinity chromatography.
Structure and Possible Mechanism of the CcbJ Methyltransferase from Streptomyces caelestis
 Supplemental material to accompany Structure and Possible Mechanism of the CcbJ Methyltransferase from Streptomyces caelestis Jacob Bauer, a Gabriela Ondrovičová, a Lucie Najmanová, b Vladimír Pevala,
Supplemental material to accompany Structure and Possible Mechanism of the CcbJ Methyltransferase from Streptomyces caelestis Jacob Bauer, a Gabriela Ondrovičová, a Lucie Najmanová, b Vladimír Pevala,
SUPPLEMENTARY INFORMATION Figures. Supplementary Figure 1 a. Page 1 of 30. Nature Chemical Biology: doi: /nchembio.2528
 SUPPLEMENTARY INFORMATION Figures Supplementary Figure 1 a b c Page 1 of 0 11 Supplementary Figure 1: Biochemical characterisation and binding validation of the reversible USP inhibitor 1. a, Biochemical
SUPPLEMENTARY INFORMATION Figures Supplementary Figure 1 a b c Page 1 of 0 11 Supplementary Figure 1: Biochemical characterisation and binding validation of the reversible USP inhibitor 1. a, Biochemical
Site-specific time-resolved FRET reveals local variations in the unfolding mechanism in an apparently two-state protein unfolding transition
 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2017 Supplementary information for Site-specific time-resolved FRET reveals local variations
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2017 Supplementary information for Site-specific time-resolved FRET reveals local variations
Supplemental Information. Structural Basis for Guide RNA Processing and. Seed-Dependent DNA Targeting by CRISPR-Cas12a
 Molecular Cell, Volume 66 Supplemental Information Structural Basis for Guide RNA Processing and Seed-Dependent DNA Targeting by CRISPR-Cas12a Daan C. Swarts, John van der Oost, and Martin Jinek Figure
Molecular Cell, Volume 66 Supplemental Information Structural Basis for Guide RNA Processing and Seed-Dependent DNA Targeting by CRISPR-Cas12a Daan C. Swarts, John van der Oost, and Martin Jinek Figure
human Cdc45 Figure 1c. (c)
 1 Details of the refined crystallographic model of human Cdc45 and comparison of its active-site region with that of bacterial RecJ. (a) Stereo view of a representative example of the final 2F o -F c electron
1 Details of the refined crystallographic model of human Cdc45 and comparison of its active-site region with that of bacterial RecJ. (a) Stereo view of a representative example of the final 2F o -F c electron
Mycobacterium tuberculosis (Mtb) and Mycobacterium smegmatis (Msmeg) contain two ε (dnaq) exonuclease homologs.
 Supplementary Figure 1 Mycobacterium tuberculosis (Mtb) and Mycobacterium smegmatis (Msmeg) contain two ε (dnaq) exonuclease homologs. (a) Sequence alignment of the ε-exonuclease homologs from four different
Supplementary Figure 1 Mycobacterium tuberculosis (Mtb) and Mycobacterium smegmatis (Msmeg) contain two ε (dnaq) exonuclease homologs. (a) Sequence alignment of the ε-exonuclease homologs from four different
Chromatin Structure. a basic discussion of protein-nucleic acid binding
 Chromatin Structure 1 Chromatin DNA packaging g First a basic discussion of protein-nucleic acid binding Questions to answer: How do proteins bind DNA / RNA? How do proteins recognize a specific nucleic
Chromatin Structure 1 Chromatin DNA packaging g First a basic discussion of protein-nucleic acid binding Questions to answer: How do proteins bind DNA / RNA? How do proteins recognize a specific nucleic
Oligonucleotides were purchased from Eurogentec, purified by denaturing gel electrophoresis
 SUPPLEMENRY INFORMION Purification of probes and Oligonucleotides sequence Oligonucleotides were purchased from Eurogentec, purified by denaturing gel electrophoresis and recovered by electroelution. Labelling
SUPPLEMENRY INFORMION Purification of probes and Oligonucleotides sequence Oligonucleotides were purchased from Eurogentec, purified by denaturing gel electrophoresis and recovered by electroelution. Labelling
Enhancers. Activators and repressors of transcription
 Enhancers Can be >50 kb away from the gene they regulate. Can be upstream from a promoter, downstream from a promoter, within an intron, or even downstream of the final exon of a gene. Are often cell type
Enhancers Can be >50 kb away from the gene they regulate. Can be upstream from a promoter, downstream from a promoter, within an intron, or even downstream of the final exon of a gene. Are often cell type
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/2/11/e1601625/dc1 Supplementary Materials for A molecular mechanism of chaperone-client recognition This PDF file includes: Lichun He, Timothy Sharpe, Adam Mazur,
advances.sciencemag.org/cgi/content/full/2/11/e1601625/dc1 Supplementary Materials for A molecular mechanism of chaperone-client recognition This PDF file includes: Lichun He, Timothy Sharpe, Adam Mazur,
Structural basis of a novel PD-L1 nanobody for immune checkpoint blockade
 Structural basis of a novel PD-L1 nanobody for immune checkpoint blockade Supplementary Materials Supplementary methods Table S1-S Figure S1-S 1 1 1 1 1 1 1 1 1 0 1 0 Supplementary Methods Competitive
Structural basis of a novel PD-L1 nanobody for immune checkpoint blockade Supplementary Materials Supplementary methods Table S1-S Figure S1-S 1 1 1 1 1 1 1 1 1 0 1 0 Supplementary Methods Competitive
AP Biology. The BIG Questions. Chapter 19. Prokaryote vs. eukaryote genome. Prokaryote vs. eukaryote genome. Why turn genes on & off?
 The BIG Questions Chapter 19. Control of Eukaryotic Genome How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions?
The BIG Questions Chapter 19. Control of Eukaryotic Genome How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions?
Ensemble refinement shows conformational flexibility in crystal structures of human complement factor D
 Supplementary Information for Ensemble refinement shows conformational flexibility in crystal structures of human complement factor D Federico Forneris a,b, B. Tom Burnley a,b,c and Piet Gros a * a Crystal
Supplementary Information for Ensemble refinement shows conformational flexibility in crystal structures of human complement factor D Federico Forneris a,b, B. Tom Burnley a,b,c and Piet Gros a * a Crystal
DNA supercoiling, a critical signal regulating the basal expression of the lac operon in Escherichia coli
 Supplementary Information DNA supercoiling, a critical signal regulating the basal expression of the lac operon in Escherichia coli Geraldine Fulcrand 1,2, Samantha Dages 1,2, Xiaoduo Zhi 1,2, Prem Chapagain
Supplementary Information DNA supercoiling, a critical signal regulating the basal expression of the lac operon in Escherichia coli Geraldine Fulcrand 1,2, Samantha Dages 1,2, Xiaoduo Zhi 1,2, Prem Chapagain
Modulating the Cascade architecture of a minimal Type I-F CRISPR-Cas system
 SUPPLEMENTARY DATA Modulating the Cascade architecture of a minimal Type I-F CRISPR-Cas system Daniel Gleditzsch 1, Hanna Müller-Esparza 1, Patrick Pausch 2,3, Kundan Sharma 4, Srivatsa Dwarakanath 1,
SUPPLEMENTARY DATA Modulating the Cascade architecture of a minimal Type I-F CRISPR-Cas system Daniel Gleditzsch 1, Hanna Müller-Esparza 1, Patrick Pausch 2,3, Kundan Sharma 4, Srivatsa Dwarakanath 1,
Molecular Biology (BIOL 4320) Exam #1 March 12, 2002
 Molecular Biology (BIOL 4320) Exam #1 March 12, 2002 Name KEY SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number.
Molecular Biology (BIOL 4320) Exam #1 March 12, 2002 Name KEY SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number.
Supplementary Figure 1 PZA inhibits root hair formation as well as cell elongation in the maturation zone of eto1-2 roots. (A) The PI staining of the
 Supplementary Figure 1 PZA inhibits root hair formation as well as cell elongation in the maturation zone of eto1-2 roots. (A) The PI staining of the roots of three-day-old etiolated seedlings of Col-0
Supplementary Figure 1 PZA inhibits root hair formation as well as cell elongation in the maturation zone of eto1-2 roots. (A) The PI staining of the roots of three-day-old etiolated seedlings of Col-0
Supplementary Fig. 1. Initial electron density maps for the NOX-D20:mC5a complex obtained after SAD-phasing. (a) Initial experimental electron
 Supplementary Fig. 1. Initial electron density maps for the NOX-D20:mC5a complex obtained after SAD-phasing. (a) Initial experimental electron density map obtained after SAD-phasing and density modification
Supplementary Fig. 1. Initial electron density maps for the NOX-D20:mC5a complex obtained after SAD-phasing. (a) Initial experimental electron density map obtained after SAD-phasing and density modification
SUPPLEMENTARY INFORMATION
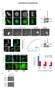 SUPPLEMENTARY INFORMATION Supplementary Figure 1: Function of MICAL1 and dmical in cytokinesis (a) HeLa transfected with GFP- MICAL1 (green) were stained with Aurora B (red). Scale bars, 10 µm. (b) Western
SUPPLEMENTARY INFORMATION Supplementary Figure 1: Function of MICAL1 and dmical in cytokinesis (a) HeLa transfected with GFP- MICAL1 (green) were stained with Aurora B (red). Scale bars, 10 µm. (b) Western
Molecular basis for H3K36me3 recognition by the Tudor domain of PHF1
 Supplementary information Molecular basis for H3K36me3 recognition by the Tudor domain of PHF1 Catherine A. Musselman 1, Nikita Avvakumov 2, Reiko Watanabe 3, Christopher G. Abraham 4, Marie-Eve Lalonde
Supplementary information Molecular basis for H3K36me3 recognition by the Tudor domain of PHF1 Catherine A. Musselman 1, Nikita Avvakumov 2, Reiko Watanabe 3, Christopher G. Abraham 4, Marie-Eve Lalonde
Phi29 Scaffold Has a Helix-Loop-Helix Motif and a Disordered Tail. Marc C Morais et al., Nature Structural Biology 2003
 Phi29 Scaffold Has a Helix-Loop-Helix Motif and a Disordered Tail Marc C Morais et al., Nature Structural Biology 2003 Free scaffold 13+ 10 min 3 min 1 min 40 sec 100 % 0 100 % 0 100 % 0 100 % 0 Partially
Phi29 Scaffold Has a Helix-Loop-Helix Motif and a Disordered Tail Marc C Morais et al., Nature Structural Biology 2003 Free scaffold 13+ 10 min 3 min 1 min 40 sec 100 % 0 100 % 0 100 % 0 100 % 0 Partially
Supporting Information Defined Bilayer Interactions of DNA Nanopores Revealed with a Nuclease-Based Nanoprobe Strategy
 Supporting Information Defined Bilayer Interactions of DNA Nanopores Revealed with a Nuclease-Based Nanoprobe Strategy Jonathan R. Burns* & Stefan Howorka* 1 Contents 1. Design of DNA nanopores... 3 1.1.
Supporting Information Defined Bilayer Interactions of DNA Nanopores Revealed with a Nuclease-Based Nanoprobe Strategy Jonathan R. Burns* & Stefan Howorka* 1 Contents 1. Design of DNA nanopores... 3 1.1.
A quantitative investigation of linker histone interactions with nucleosomes and chromatin
 White et al Supplementary figures and figure legends A quantitative investigation of linker histone interactions with nucleosomes and chromatin 1 Alison E. White, Aaron R. Hieb, and 1 Karolin Luger 1 White
White et al Supplementary figures and figure legends A quantitative investigation of linker histone interactions with nucleosomes and chromatin 1 Alison E. White, Aaron R. Hieb, and 1 Karolin Luger 1 White
Supplementary Information. Single-molecule analysis reveals multi-state folding of a guanine. riboswitch
 Supplementary Information Single-molecule analysis reveals multi-state folding of a guanine riboswitch Vishnu Chandra 1,4,#, Zain Hannan 1,5,#, Huizhong Xu 2,# and Maumita Mandal 1,2,3,6* Department of
Supplementary Information Single-molecule analysis reveals multi-state folding of a guanine riboswitch Vishnu Chandra 1,4,#, Zain Hannan 1,5,#, Huizhong Xu 2,# and Maumita Mandal 1,2,3,6* Department of
Supplementary Figure 1. Nature Structural & Molecular Biology: doi: /nsmb.3494
 Supplementary Figure 1 Pol structure-function analysis (a) Inactivating polymerase and helicase mutations do not alter the stability of Pol. Flag epitopes were introduced using CRISPR/Cas9 gene targeting
Supplementary Figure 1 Pol structure-function analysis (a) Inactivating polymerase and helicase mutations do not alter the stability of Pol. Flag epitopes were introduced using CRISPR/Cas9 gene targeting
SUPPLEMENTARY INFORMATION. Determinants of laminin polymerization revealed. by the structure of the α5 chain amino-terminal region
 SUPPLEMENTARY INFORMATION Determinants of laminin polymerization revealed by the structure of the α5 chain amino-terminal region Sadaf-Ahmahni Hussain 1, Federico Carafoli 1 & Erhard Hohenester 1 1 Department
SUPPLEMENTARY INFORMATION Determinants of laminin polymerization revealed by the structure of the α5 chain amino-terminal region Sadaf-Ahmahni Hussain 1, Federico Carafoli 1 & Erhard Hohenester 1 1 Department
Engineering splicing factors with designed specificities
 nature methods Engineering splicing factors with designed specificities Yang Wang, Cheom-Gil Cheong, Traci M Tanaka Hall & Zefeng Wang Supplementary figures and text: Supplementary Figure 1 Supplementary
nature methods Engineering splicing factors with designed specificities Yang Wang, Cheom-Gil Cheong, Traci M Tanaka Hall & Zefeng Wang Supplementary figures and text: Supplementary Figure 1 Supplementary
Zhang et al., RepID facilitates replication Initiation. Supplemental Information:
 Supplemental Information: a b 1 Supplementary Figure 1 (a) DNA sequence of all the oligonucleotides used in this study. Only one strand is shown. The unshaded nucleotide sequences show changes from the
Supplemental Information: a b 1 Supplementary Figure 1 (a) DNA sequence of all the oligonucleotides used in this study. Only one strand is shown. The unshaded nucleotide sequences show changes from the
Practice Exam A. Briefly describe how IL-25 treatment might be able to help this responder subgroup of liver cancer patients.
 Practice Exam 2007 1. A special JAK-STAT signaling system (JAK5-STAT5) was recently identified in which a gene called TS5 becomes selectively transcribed and expressed in the liver upon induction by a
Practice Exam 2007 1. A special JAK-STAT signaling system (JAK5-STAT5) was recently identified in which a gene called TS5 becomes selectively transcribed and expressed in the liver upon induction by a
Supplementary Note 1. Enzymatic properties of the purified Syn BVR
 Supplementary Note 1. Enzymatic properties of the purified Syn BVR The expression vector pet15b-syn bvr allowed us to routinely prepare 15 mg of electrophoretically homogenous Syn BVR from 2.5 L of TB-medium
Supplementary Note 1. Enzymatic properties of the purified Syn BVR The expression vector pet15b-syn bvr allowed us to routinely prepare 15 mg of electrophoretically homogenous Syn BVR from 2.5 L of TB-medium
Supplementary Figure 1. Supporting data for Figure 1
 Supplementary Figure 1 Supporting data for Figure 1 (A) Schematic of BLI assay used to measure dissociation. 5 monobiotinylated substrate DNA (identical to λ1, Figure 2) is associated with streptavidin-coated
Supplementary Figure 1 Supporting data for Figure 1 (A) Schematic of BLI assay used to measure dissociation. 5 monobiotinylated substrate DNA (identical to λ1, Figure 2) is associated with streptavidin-coated
Complex Transcription Machinery
 Complex Transcription Machinery Subunits of the Basal Txn Apparatus Stepwise Assembly of the Pre-initiation Complex Interplay of Activators, Co-regulators and RNA Polymerase at the Promoter Divide and
Complex Transcription Machinery Subunits of the Basal Txn Apparatus Stepwise Assembly of the Pre-initiation Complex Interplay of Activators, Co-regulators and RNA Polymerase at the Promoter Divide and
CSE : Computational Issues in Molecular Biology. Lecture 19. Spring 2004
 CSE 397-497: Computational Issues in Molecular Biology Lecture 19 Spring 2004-1- Protein structure Primary structure of protein is determined by number and order of amino acids within polypeptide chain.
CSE 397-497: Computational Issues in Molecular Biology Lecture 19 Spring 2004-1- Protein structure Primary structure of protein is determined by number and order of amino acids within polypeptide chain.
Reduced structural flexibility for exonuclease deficient DNA polymerase III mutant
 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2018 Reduced structural flexibility for exonuclease deficient DNA polymerase III mutant
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2018 Reduced structural flexibility for exonuclease deficient DNA polymerase III mutant
Molecular Genetics of Disease and the Human Genome Project
 9 Molecular Genetics of Disease and the Human Genome Project Fig. 1. The 23 chromosomes in the human genome. There are 22 autosomes (chromosomes 1 to 22) and two sex chromosomes (X and Y). Females inherit
9 Molecular Genetics of Disease and the Human Genome Project Fig. 1. The 23 chromosomes in the human genome. There are 22 autosomes (chromosomes 1 to 22) and two sex chromosomes (X and Y). Females inherit
Biol 321 Spring 2013 Quiz 4 25 pts NAME
 Biol 321 Spring 2013 Quiz 4 25 pts NAME 1. (3 pts.) a. What is the name of this compound? BE EXPLICIT deoxyribose 5 b. Number the carbons on this structure: 4 1 3 2 2. (4 pts.) Circle True or False. If
Biol 321 Spring 2013 Quiz 4 25 pts NAME 1. (3 pts.) a. What is the name of this compound? BE EXPLICIT deoxyribose 5 b. Number the carbons on this structure: 4 1 3 2 2. (4 pts.) Circle True or False. If
Supplementary Figure 1 Strategy for parallel detection of DHSs and adjacent nucleosomes
 Supplementary Figure 1 Strategy for parallel detection of DHSs and adjacent nucleosomes DNase I cleavage DNase I DNase I digestion Sucrose gradient enrichment Small Large F1 F2...... F9 F1 F1 F2 F3 F4
Supplementary Figure 1 Strategy for parallel detection of DHSs and adjacent nucleosomes DNase I cleavage DNase I DNase I digestion Sucrose gradient enrichment Small Large F1 F2...... F9 F1 F1 F2 F3 F4
Supplementary Information Crystal structure, biochemical and cellular activities demonstrate separate functions of MTH1 and MTH2
 Supplementary Information Crystal structure, biochemical and cellular activities demonstrate separate functions of MTH1 and MTH2 Supplementary Figure 1 Comparison of activities of NUDIX proteins with nucleotide
Supplementary Information Crystal structure, biochemical and cellular activities demonstrate separate functions of MTH1 and MTH2 Supplementary Figure 1 Comparison of activities of NUDIX proteins with nucleotide
Supplementary information to eif4b stimulates eif4a ATPase and unwinding activities by direct
 Supplementary information to eif4b stimulates eif4a ATPase and unwinding activities by direct interaction through its 7-repeats region, Alexandra Z. Andreou, Ulf Harms & Dagmar Klostermeier. Supplementary
Supplementary information to eif4b stimulates eif4a ATPase and unwinding activities by direct interaction through its 7-repeats region, Alexandra Z. Andreou, Ulf Harms & Dagmar Klostermeier. Supplementary
Viral RNAi suppressor reversibly binds sirna to. outcompete Dicer and RISC via multiple-turnover
 Supplementary Data Viral RNAi suppressor reversibly binds sirna to outcompete Dicer and RISC via multiple-turnover Renata A. Rawlings 1,2, Vishalakshi Krishnan 2 and Nils G. Walter 2 * 1 Biophysics and
Supplementary Data Viral RNAi suppressor reversibly binds sirna to outcompete Dicer and RISC via multiple-turnover Renata A. Rawlings 1,2, Vishalakshi Krishnan 2 and Nils G. Walter 2 * 1 Biophysics and
Supplemental Information. Single-Molecule Imaging Reveals How. Mre11-Rad50-Nbs1 Initiates DNA Break Repair
 Molecular Cell, Volume 67 Supplemental Information Single-Molecule Imaging Reveals How Mre11-Rad50-Nbs1 Initiates DNA Break Repair Logan R. Myler, Ignacio F. Gallardo, Michael M. Soniat, Rajashree A. Deshpande,
Molecular Cell, Volume 67 Supplemental Information Single-Molecule Imaging Reveals How Mre11-Rad50-Nbs1 Initiates DNA Break Repair Logan R. Myler, Ignacio F. Gallardo, Michael M. Soniat, Rajashree A. Deshpande,
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Origin use and efficiency are similar among WT, rrm3, pif1-m2, and pif1-m2; rrm3 strains. A. Analysis of fork progression around confirmed and likely origins (from cerevisiae.oridb.org).
Supplementary Figure 1 Origin use and efficiency are similar among WT, rrm3, pif1-m2, and pif1-m2; rrm3 strains. A. Analysis of fork progression around confirmed and likely origins (from cerevisiae.oridb.org).
Supplementary Figure 1. Antibody-induced cargo release studied by native PAGE. A clear band corresponding to the cargo strand (lane 1) is visible.
 Supplementary Figure 1. Antibody-induced cargo release studied by native PAGE. A clear band corresponding to the cargo strand (lane 1) is visible. Because SYBR Gold is less sensitive to single stranded
Supplementary Figure 1. Antibody-induced cargo release studied by native PAGE. A clear band corresponding to the cargo strand (lane 1) is visible. Because SYBR Gold is less sensitive to single stranded
Supplementary Figure 1.
 Supplementary Figure 1. Assessment of quaternary structure of soluble RSV F proteins. Soluble variants of F proteins from A2 and B1 RSV strains were expressed in HEK293 cells. The cell culture supernatants
Supplementary Figure 1. Assessment of quaternary structure of soluble RSV F proteins. Soluble variants of F proteins from A2 and B1 RSV strains were expressed in HEK293 cells. The cell culture supernatants
SUPPLEMENTARY MATERIAL
 SUPPLEMENTARY MATERIAL 1 Table 1. Comparison of experimental and theoretical nucleosomal arrangements on selected sequences* Sequence High-affinity 601 synthetic sequence experimental method site-directed
SUPPLEMENTARY MATERIAL 1 Table 1. Comparison of experimental and theoretical nucleosomal arrangements on selected sequences* Sequence High-affinity 601 synthetic sequence experimental method site-directed
Gene Expression - Transcription
 DNA Gene Expression - Transcription Genes are expressed as encoded proteins in a 2 step process: transcription + translation Central dogma of biology: DNA RNA protein Transcription: copy DNA strand making
DNA Gene Expression - Transcription Genes are expressed as encoded proteins in a 2 step process: transcription + translation Central dogma of biology: DNA RNA protein Transcription: copy DNA strand making
Supplementary Information. Small Molecule-Induced Domain Swapping as a Mechanism for Controlling Protein Function and Assembly
 Supplementary Information Small Molecule-Induced Domain Swapping as a Mechanism for Controlling Protein Function and Assembly Joshua M. Karchin, Jeung-Hoi Ha, Kevin E. Namitz, Michael S. Cosgrove, and
Supplementary Information Small Molecule-Induced Domain Swapping as a Mechanism for Controlling Protein Function and Assembly Joshua M. Karchin, Jeung-Hoi Ha, Kevin E. Namitz, Michael S. Cosgrove, and
Challenges to measuring intracellular Ca 2+ Calmodulin: nature s Ca 2+ sensor
 Calcium Signals in Biological Systems Lecture 3 (2/9/0) Measuring intracellular Ca 2+ signals II: Genetically encoded Ca 2+ sensors Henry M. Colecraft, Ph.D. Challenges to measuring intracellular Ca 2+
Calcium Signals in Biological Systems Lecture 3 (2/9/0) Measuring intracellular Ca 2+ signals II: Genetically encoded Ca 2+ sensors Henry M. Colecraft, Ph.D. Challenges to measuring intracellular Ca 2+
Biochemistry 111. Carl Parker x A Braun
 Biochemistry 111 Carl Parker x6368 101A Braun csp@caltech.edu Central Dogma of Molecular Biology DNA-Dependent RNA Polymerase Requires a DNA Template Synthesizes RNA in a 5 to 3 direction Requires ribonucleoside
Biochemistry 111 Carl Parker x6368 101A Braun csp@caltech.edu Central Dogma of Molecular Biology DNA-Dependent RNA Polymerase Requires a DNA Template Synthesizes RNA in a 5 to 3 direction Requires ribonucleoside
Nucleic acids. How DNA works. DNA RNA Protein. DNA (deoxyribonucleic acid) RNA (ribonucleic acid) Central Dogma of Molecular Biology
 Nucleic acid chemistry and basic molecular theory Nucleic acids DNA (deoxyribonucleic acid) RNA (ribonucleic acid) Central Dogma of Molecular Biology Cell cycle DNA RNA Protein Transcription Translation
Nucleic acid chemistry and basic molecular theory Nucleic acids DNA (deoxyribonucleic acid) RNA (ribonucleic acid) Central Dogma of Molecular Biology Cell cycle DNA RNA Protein Transcription Translation
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Detection of MCM-subunit SUMOylation under normal growth conditions. a. Sumoylated forms of MCM subunits show differential shifts when SUMO is attached to differently sized tags.
Supplementary Figure 1 Detection of MCM-subunit SUMOylation under normal growth conditions. a. Sumoylated forms of MCM subunits show differential shifts when SUMO is attached to differently sized tags.
