Describing variants. recommendations for the description of DNA changes. Johan den Dunnen. HGVS.org. chair SVD-WG
|
|
|
- Barbra Chambers
- 6 years ago
- Views:
Transcription
1 Describing variants "mutation nomenclature" recommendations for the description of DNA changes Johan den Dunnen chair SVD-WG HGVS.org
2 HGVS / HVP / HUGO Sequence Variant Description working group Working Group Members: Anne-Francoise Roux (EGT) Donna Maglott (NCBI/EBI) Jean McGowan-Jordan (ISCN) Peter Taschner (LSDBs) Raymond Dalgleish (LSDBs) Reece Hart (industry) Johan den Dunnen (chair) HGVS - Marc Greenblatt HUGO - Stylianos Antonarakis
3 Nomenclature ( describing DNA variants ) Stable Meaningful Memorable Unequivocal
4 Definitions prevent confusion do not use "mutation" use variant, disease-associated variant do not use "polymorphism" use variant, not disease-associated variant do not use "pathogenic" use disease-associated, a disease-associated variant better use neutral terms sequence variant alteration CNV SNV (Copy Number Variant) (Single Nucleotide Variant, not SNP)
5 Variant description the basis / varnomen Hum Mutat (2016) 37:
6 Follow the recommendations when you disagree, start a debate -do not use private rules, this only causes confusion
7 facebook & twitter
8 Versioning version presented is (Nov.2015)
9 Variant types change in sequence ACATCAGGAGAAGATGTTC GAGACTTTGCCA ACATCAGGAGAAGATGTTT GAGACTTTGCCA ACATCAGGAGAAGATGTT GAGACTTTGCCA ACATCAGGAGAAGATGTTCCGAGACTTTGCCA change in amount change in position (Copy Number Variation) Structural Variation (SV)
10 DNA, RNA, protein unique descriptions prevent confusion DNA A, G, C, T g.957a>t, c.63-3t>c RNA a, g, c, u r.957a>u, r.(?), r.spl? protein ( mostly deduced ) three / one letter amino acid code * = stop codon p.(his78gln)
11 Reference sequence use official HGNC gene symbols provide reference sequence covering complete sequence largest transcript preferably a LRG e.g. LRG_123 give accession.version number e.g. NM_ indicate type of Reference Sequence DNA coding DNA c. genomic g. mitochondrial m. non-coding RNA n. RNA r. protein p.
12 The LRG EBI, NCBI, Gen2Phen
13 Numbering residues start with 1 genomic no +, - or other signs coding DNA 1 is first nucleotide of file 1 is A of ATG for introns refer to genomic Reference Sequence repeated segments (...CGTGTG TG A ) assume most 3' as changed coding DNA only 5' of ATG, -3, -2, -1, A, T, G, no nucleotide 0 3' of stop *1, *2, *3, no nucleotide 0 intron position between nt's 654 and 655 c.654+1, +2, +3,, -3, -2, c change + to - in middle
14 Numbering RNA ( deduced mostly ) like coding DNA protein ( deduced only ) from first to last amino acid rule of thumb: c. nucleotide position divided by 3 roughly gives amino acid residue description between parantheses
15 Reference Sequence coding DNA reference sequence (c.) non-coding DNA reference sequence (n.)
16 coding DNA or genomic? human genome sequence complete covers all transcripts different promoters, splice variants, diff. polya-addition, etc. but hg19 chr2:g _ del is long & complicated huge reference sequence files new builds follow each other regularly carries no understandable information coding DNA does not cover all variants but gives a clue towards position
17 Numbering - genomic3 g a>g g >c g g>a g c>g no relation to RNA & protein g a>t g t>g g g>c
18 Numbering - coding DNA c.1637a>g protein coding region c t>c in intron c g>a in intron (5' half) (3' half) relation to RNA & protein c.-23c>g 5' of protein coding region (5' of ATG) c.*143a>t 3' of protein coding region (3' of stop) c t>g intron in 5' UTR (5' of ATG) c g>c intron in 3' UTR (3' of stop)
19 Types of variation simple substitution deletion duplication insertion c.123a>g c.123dela c.123dupa c.123_124insc other conversion, inversion, translocation, transposition complex indel c.123delinsgtat combination of variants two alleles >1 per allele c.[123a>g;456c>t] c.[123a>g];[456c>t]
20 Substitution substitution designated by ">" > not used on protein level examples genomic cdna RNA protein g.54786a>t c.545a>t ( NM_ : c.546a>t ) r.545a>u p.(gln182leu)
21 Deletion deletion designated by "del" range indicated by "_" examples c.546del c.546delt c.586_591del c.586_591deltggtca, NOT c.586_591del6 c.(780+1_781-1)_(1392+1_1393-1)del exon 3 to 6 deletion, breakpoint not sequenced
22 Duplication duplication designated by "dup" range indicated by "_" examples c.546dup c.546dupt c.586_591dup c.586_591duptggtca, NOT c.586_591dup6 do not describe as insertion c.(780+1_781-1)_(1392+1_1393-1)dup exon 3 to 6 duplication, breakpoint not sequenced NOTE: dup should be in tandem
23 Insertion insertion designated by "ins" range indicated by "_"! give inserted sequence examples c.546_547inst NOT c.546inst or c.547inst c.1086_1087insgcgtga NOT c.1086_1087ins6 c.1086_1087insab :g.34_12567 when large insert submit to database and give database accession.version number
24 Inversion inversion affecting at least 2 nucleotides designated by "inv" range indicated by "_" example c.546_2031inv NOT c.2031_546inv
25 Conversion conversion affecting at least 2 nucleotides designated by "con" range indicated by "_" examples c.546_657con917_1028 c.546_2031connm_ :c.549_2034
26 Sequence repeats mono-nucleotide stretches g.8932a(18_23) c t(18_23) alleles T[18];[21] di-nucleotide stretches c cag(13_19) c _ (13_19) larger g.532_3886(20_45) 3.3 Kb repeat () = uncertain
27 SNVs (SNPs) SNV's at least once give description based on genome reference sequence hg19 chr9:g t>c rs :t>c dbsnp entry
28 Characters & codes codes used +, -, * > substitution (nucleotide) _ range ; separate changes (in/between alleles), more transcripts ( ) uncertain [ ] allele = equals reference sequence? unknown del deletion dup duplication ins insertion inv inversion con conversion ext extension fs frame shift
29 Uncertainty breakpoints Copy Number Variants ( last-normal_first-changed ) _ ( last-changed_first-normal ) del BAC / PAC probe chrx:g.( _ )_( _ )del hg19 SNP-array chrx:g.( _ )_( _ )del GRCh36.p2 (rs _rs )_(rs _rs947283)del
30 Uncertainty breakpoints2 whole exon changes c.(423+1_424-1)_(631+1_632-1)del intragenic deletion c.(?_-79)_(631+1_632-1)del deletion incl. 5' end c.(423+1_424-1)_(*763_?)del deletion incl. 3' end c.(?_-79)_(*763_?)del whole gene deletion, start/end undefined describe what was actually tested
31 Alleles allele indicated by "[ ]", separated by ";" 2 changes, 2 alleles c. [428A>G] ; [83dupG] 1 allele, several changes c. [12C>G ; 428A>G ; 983dupG] 2 changes, allele unknown c. 428A>G (;) 83dupG special cases mosaicism c. 428A= / A>G chimerism c. 428A= // A>G spaces in description used for clarity only
32 Complex deletion / insertions "indel" c.1166_1177delinsagt descriptions may become complex when only an expert understands the "code" consider database submission description: c.875_941delinsac
33 Changes in RNA description like DNA r. / a, g, c, u examples r.283c>u r.0 no RNA from allele r.? effect unknown r.spl affects RNA splicing r.(spl?) may affect splicing r.283= no change (equals reference sequence) r.[=, 436_456del] two transcripts from 1 allele
34 Changes in RNA2 one allele, 2 transcripts effect on splicing not 100% c.456+3g>c on RNA r.[=, 436_456del] > p.[=, Arg146_Lys152del]
35 Changes in protein description like DNA p. / Ala, Cys, Gly, His,, Ter p. / A, C, D, E, F, G, H,, * examples nonsense p.trp65* (p.w65* / p.trp65ter) no stop p.*1054glnext*31 p.0 - no protein p.met1? - likely, but unknown effect NOT p.met1val fs - frame shift no RNA data r.(?) p.(trp56*)
36 Frame shifts short form (sufficient) p.arg83fs long from (more detail) p.(arg83serfs*15) (no RNA analysis) indicate first amino acid changed do not try to include position changes at DNA level first changed amino acid length shifted frame (from first changed to * incl.) do not describe del, dup, ins, etc.
37 Recent additions added versioning to support users easier to find latest changes allows statement "following HGVS version 2.0" stricter definitions separate different classes added hierarchy computer-generated description automated error-checking ( Mutalyzer ) simplified use special characters "_", ";", "+", "*", improved consistency
38 Acknowledgement Presentation prepared by: Johan den Dunnen Human Genetics & Clinical Genetics Leiden University Medical Center Leiden, Nederland chair SVD-WG date: April 2017
Q & A. scan QR code. or go to
 Describing variants "mutation nomenclature" recommendations for the description of DNA changes tinyurl.com/ VEPTC-4 http://varnomen.hgvs.org Johan den Dunnen chair SVD-WG Q & A scan QR code Human and Clinical
Describing variants "mutation nomenclature" recommendations for the description of DNA changes tinyurl.com/ VEPTC-4 http://varnomen.hgvs.org Johan den Dunnen chair SVD-WG Q & A scan QR code Human and Clinical
HGVS Mutation Nomenclature Queries
 Molecular Genetics Participants Meeting 2014 HGVS Mutation Nomenclature Queries UK NATIONAL EXTERNAL QUALITY ASSESSMENT SERVICES General Remark HGVS nomenclature aim: To provide unambiguous descriptions
Molecular Genetics Participants Meeting 2014 HGVS Mutation Nomenclature Queries UK NATIONAL EXTERNAL QUALITY ASSESSMENT SERVICES General Remark HGVS nomenclature aim: To provide unambiguous descriptions
Mutation entries in SMA databases Guidelines for national curators
 1 Mutation entries in SMA databases Guidelines for national curators GENERAL CONSIDERATIONS Role of the curator(s) of a national database Molecular data can be collected by many different ways. There are
1 Mutation entries in SMA databases Guidelines for national curators GENERAL CONSIDERATIONS Role of the curator(s) of a national database Molecular data can be collected by many different ways. There are
Understanding Genes & Mutations. John A Phillips III May 16, 2005
 Understanding Genes & Mutations John A Phillips III May 16, 2005 Learning Objectives Understand gene structure Become familiar with genetic & mutation databases Be able to find information on genetic variation
Understanding Genes & Mutations John A Phillips III May 16, 2005 Learning Objectives Understand gene structure Become familiar with genetic & mutation databases Be able to find information on genetic variation
In silico variant analysis: Challenges and Pitfalls
 In silico variant analysis: Challenges and Pitfalls Fiona Cunningham Variation annotation coordinator EMBL-EBI www.ensembl.org Sequencing -> Variants -> Interpretation Structural variants SNP? In-dels
In silico variant analysis: Challenges and Pitfalls Fiona Cunningham Variation annotation coordinator EMBL-EBI www.ensembl.org Sequencing -> Variants -> Interpretation Structural variants SNP? In-dels
For more information about how to cite these materials visit
 Author(s): David Ginsburg, M.D., 2012 License: Unless otherwise noted, this material is made available under the terms of the Creative Commons Attribution Non-commercial Share Alike 3.0 License: http://creativecommons.org/licenses/by-nc-sa/3.0/
Author(s): David Ginsburg, M.D., 2012 License: Unless otherwise noted, this material is made available under the terms of the Creative Commons Attribution Non-commercial Share Alike 3.0 License: http://creativecommons.org/licenses/by-nc-sa/3.0/
BS 50 Genetics and Genomics Week of Oct 24
 BS 50 Genetics and Genomics Week of Oct 24 Additional Practice Problems for Section Question 1: The following table contains a list of statements that apply to replication, transcription, both, or neither.
BS 50 Genetics and Genomics Week of Oct 24 Additional Practice Problems for Section Question 1: The following table contains a list of statements that apply to replication, transcription, both, or neither.
Introduction to Next Generation Sequencing (NGS) Andrew Parrish Exeter, 2 nd November 2017
 Introduction to Next Generation Sequencing (NGS) Andrew Parrish Exeter, 2 nd November 2017 Topics to cover today What is Next Generation Sequencing (NGS)? Why do we need NGS? Common approaches to NGS NGS
Introduction to Next Generation Sequencing (NGS) Andrew Parrish Exeter, 2 nd November 2017 Topics to cover today What is Next Generation Sequencing (NGS)? Why do we need NGS? Common approaches to NGS NGS
Mutations. Lecture 15
 Mutations Lecture 15 Objectives 1: Mutation Define mutation. Describe the types of mutations and their effects on the protein that is produced Distinguishing between spontaneous and induced mutations.
Mutations Lecture 15 Objectives 1: Mutation Define mutation. Describe the types of mutations and their effects on the protein that is produced Distinguishing between spontaneous and induced mutations.
Structural Variation CYP2D6
 reference gene locus The gene locus contains three genes,, CYP and CYP, of which only is functional. CYP and CYP are considered pseudogenes. All three genes are composed of nine exons and share a high
reference gene locus The gene locus contains three genes,, CYP and CYP, of which only is functional. CYP and CYP are considered pseudogenes. All three genes are composed of nine exons and share a high
SNP calling and VCF format
 SNP calling and VCF format Laurent Falquet, Oct 12 SNP? What is this? A type of genetic variation, among others: Family of Single Nucleotide Aberrations Single Nucleotide Polymorphisms (SNPs) Single Nucleotide
SNP calling and VCF format Laurent Falquet, Oct 12 SNP? What is this? A type of genetic variation, among others: Family of Single Nucleotide Aberrations Single Nucleotide Polymorphisms (SNPs) Single Nucleotide
Jumping Into Your Gene Pool: Understanding Genetic Test Results
 Jumping Into Your Gene Pool: Understanding Genetic Test Results Susan A. Berry, MD Professor and Director Division of Genetics and Metabolism Department of Pediatrics University of Minnesota Acknowledgment
Jumping Into Your Gene Pool: Understanding Genetic Test Results Susan A. Berry, MD Professor and Director Division of Genetics and Metabolism Department of Pediatrics University of Minnesota Acknowledgment
Annotating your variants: Ensembl Variant Effect Predictor (VEP) Helen Sparrow Ensembl EMBL-EBI 2nd November 2016
 Training materials Ensembl training materials are protected by a CC BY license http://creativecommons.org/licenses/by/4.0/ If you wish to re-use these materials, please credit Ensembl for their creation
Training materials Ensembl training materials are protected by a CC BY license http://creativecommons.org/licenses/by/4.0/ If you wish to re-use these materials, please credit Ensembl for their creation
Investigating Inherited Diseases
 Investigating Inherited Diseases The purpose of these exercises is to introduce bioinformatics databases and tools. We investigate an important human gene and see how mutations give rise to inherited diseases.
Investigating Inherited Diseases The purpose of these exercises is to introduce bioinformatics databases and tools. We investigate an important human gene and see how mutations give rise to inherited diseases.
Single Nucleotide Variant Analysis. H3ABioNet May 14, 2014
 Single Nucleotide Variant Analysis H3ABioNet May 14, 2014 Outline What are SNPs and SNVs? How do we identify them? How do we call them? SAMTools GATK VCF File Format Let s call variants! Single Nucleotide
Single Nucleotide Variant Analysis H3ABioNet May 14, 2014 Outline What are SNPs and SNVs? How do we identify them? How do we call them? SAMTools GATK VCF File Format Let s call variants! Single Nucleotide
Disease and selection in the human genome 3
 Disease and selection in the human genome 3 Ka/Ks revisited Please sit in row K or forward RBFD: human populations, adaptation and immunity Neandertal Museum, Mettman Germany Sequence genome Measure expression
Disease and selection in the human genome 3 Ka/Ks revisited Please sit in row K or forward RBFD: human populations, adaptation and immunity Neandertal Museum, Mettman Germany Sequence genome Measure expression
Lecture 2: Biology Basics Continued
 Lecture 2: Biology Basics Continued Central Dogma DNA: The Code of Life The structure and the four genomic letters code for all living organisms Adenine, Guanine, Thymine, and Cytosine which pair A-T and
Lecture 2: Biology Basics Continued Central Dogma DNA: The Code of Life The structure and the four genomic letters code for all living organisms Adenine, Guanine, Thymine, and Cytosine which pair A-T and
Molecular Genetics of Disease and the Human Genome Project
 9 Molecular Genetics of Disease and the Human Genome Project Fig. 1. The 23 chromosomes in the human genome. There are 22 autosomes (chromosomes 1 to 22) and two sex chromosomes (X and Y). Females inherit
9 Molecular Genetics of Disease and the Human Genome Project Fig. 1. The 23 chromosomes in the human genome. There are 22 autosomes (chromosomes 1 to 22) and two sex chromosomes (X and Y). Females inherit
Section 10.3 Outline 10.3 How Is the Base Sequence of a Messenger RNA Molecule Translated into Protein?
 Section 10.3 Outline 10.3 How Is the Base Sequence of a Messenger RNA Molecule Translated into Protein? Messenger RNA Carries Information for Protein Synthesis from the DNA to Ribosomes Ribosomes Consist
Section 10.3 Outline 10.3 How Is the Base Sequence of a Messenger RNA Molecule Translated into Protein? Messenger RNA Carries Information for Protein Synthesis from the DNA to Ribosomes Ribosomes Consist
Gene mutation and DNA polymorphism
 Gene mutation and DNA polymorphism Outline of this chapter Gene Mutation DNA Polymorphism Gene Mutation Definition Major Types Definition A gene mutation is a change in the nucleotide sequence that composes
Gene mutation and DNA polymorphism Outline of this chapter Gene Mutation DNA Polymorphism Gene Mutation Definition Major Types Definition A gene mutation is a change in the nucleotide sequence that composes
Genes are coded DNA instructions that control the production of proteins within a cell. The first step in decoding genetic messages is to copy a part
 Genes are coded DNA instructions that control the production of proteins within a cell. The first step in decoding genetic messages is to copy a part of the nucleotide sequence of the DNA into RNA. RNA
Genes are coded DNA instructions that control the production of proteins within a cell. The first step in decoding genetic messages is to copy a part of the nucleotide sequence of the DNA into RNA. RNA
Cover Page. The handle holds various files of this Leiden University dissertation.
 Cover Page The handle http://hdl.handle.net/1887/45045 holds various files of this Leiden University dissertation. Author: Vis, J.K. Title: Algorithms for the description of molecular sequences Issue Date:
Cover Page The handle http://hdl.handle.net/1887/45045 holds various files of this Leiden University dissertation. Author: Vis, J.K. Title: Algorithms for the description of molecular sequences Issue Date:
Unit 6 DNA ppt 3 Gene Expression and Mutations Chapter 8.6 & 8.7 pg
 Unit 6 DNA ppt 3 Gene Expression and Mutations Chapter 8.6 & 8.7 pg 248-255 Which genes are transcribed on the chromosomes are carefully regulated at many points. Watch this! https://www.youtube.com/watch?v=oewozs_jtgk
Unit 6 DNA ppt 3 Gene Expression and Mutations Chapter 8.6 & 8.7 pg 248-255 Which genes are transcribed on the chromosomes are carefully regulated at many points. Watch this! https://www.youtube.com/watch?v=oewozs_jtgk
VARIANT TERMINOLOGY AND EXON NUMBERING
 HVP/GL/003-01/EN 2016-01-28 G/DSDBAC HVP GUIDELINE VARIANT TERMINOLOGY AND EXON NUMBERING Authors Raymond Dalgleish, Mauno Vihinen Editor Timothy D. Smith Published by: Human Variome Project International
HVP/GL/003-01/EN 2016-01-28 G/DSDBAC HVP GUIDELINE VARIANT TERMINOLOGY AND EXON NUMBERING Authors Raymond Dalgleish, Mauno Vihinen Editor Timothy D. Smith Published by: Human Variome Project International
 www.lessonplansinc.com Topic: Gene Mutations WS Summary: Students will learn about frame shift mutations and base substitution mutations. Goals & Objectives: Students will be able to demonstrate how mutations
www.lessonplansinc.com Topic: Gene Mutations WS Summary: Students will learn about frame shift mutations and base substitution mutations. Goals & Objectives: Students will be able to demonstrate how mutations
Genetics 101: What does it mean to have an LBSL-specific mutation
 Genetics 101: What does it mean to have an LBSL-specific mutation Rebecca McClellan, MGC, CGC Johns Hopkins Center for Inherited Heart Disease Kennedy Krieger Metabolism Clinic Genetics 101 A gene is the
Genetics 101: What does it mean to have an LBSL-specific mutation Rebecca McClellan, MGC, CGC Johns Hopkins Center for Inherited Heart Disease Kennedy Krieger Metabolism Clinic Genetics 101 A gene is the
Gene Identification in silico
 Gene Identification in silico Nita Parekh, IIIT Hyderabad Presented at National Seminar on Bioinformatics and Functional Genomics, at Bioinformatics centre, Pondicherry University, Feb 15 17, 2006. Introduction
Gene Identification in silico Nita Parekh, IIIT Hyderabad Presented at National Seminar on Bioinformatics and Functional Genomics, at Bioinformatics centre, Pondicherry University, Feb 15 17, 2006. Introduction
Hands-On Four Investigating Inherited Diseases
 Hands-On Four Investigating Inherited Diseases The purpose of these exercises is to introduce bioinformatics databases and tools. We investigate an important human gene and see how mutations give rise
Hands-On Four Investigating Inherited Diseases The purpose of these exercises is to introduce bioinformatics databases and tools. We investigate an important human gene and see how mutations give rise
Genes and Proteins. Objectives
 Genes and Proteins Lecture 15 Objectives At the end of this series of lectures, you should be able to: Define terms. Explain the central dogma of molecular biology. Describe the locations, reactants, and
Genes and Proteins Lecture 15 Objectives At the end of this series of lectures, you should be able to: Define terms. Explain the central dogma of molecular biology. Describe the locations, reactants, and
Assignment 9: Genetic Variation
 Assignment 9: Genetic Variation Due Date: Friday, March 30 th, 2018, 10 am In this assignment, you will profile genome variation information and attempt to answer biologically relevant questions. The variant
Assignment 9: Genetic Variation Due Date: Friday, March 30 th, 2018, 10 am In this assignment, you will profile genome variation information and attempt to answer biologically relevant questions. The variant
Overview of Emerging Clinical Genomic Standards, from Healthcare IT Standards Organizations
 Overview of Emerging Clinical Genomic Standards, from Healthcare IT Standards Organizations October 11, 2009 Notes assembled by Mollie Ullman-Cullere, co-chair HL7 Clinical Genomics Workgroup for use by
Overview of Emerging Clinical Genomic Standards, from Healthcare IT Standards Organizations October 11, 2009 Notes assembled by Mollie Ullman-Cullere, co-chair HL7 Clinical Genomics Workgroup for use by
GENETICS and the DNA code NOTES
 GENETICS and the DNA code NOTES BACKGROUND DNA is the hereditary material of most organisms. It is an organic compound made of two strands, twisted around one another to form a double helix. Each strand
GENETICS and the DNA code NOTES BACKGROUND DNA is the hereditary material of most organisms. It is an organic compound made of two strands, twisted around one another to form a double helix. Each strand
Training materials.
 Training materials Ensembl training materials are protected by a CC BY license http://creativecommons.org/licenses/by/4.0/ If you wish to re-use these materials, please credit Ensembl for their creation
Training materials Ensembl training materials are protected by a CC BY license http://creativecommons.org/licenses/by/4.0/ If you wish to re-use these materials, please credit Ensembl for their creation
Map-Based Cloning of Qualitative Plant Genes
 Map-Based Cloning of Qualitative Plant Genes Map-based cloning using the genetic relationship between a gene and a marker as the basis for beginning a search for a gene Chromosome walking moving toward
Map-Based Cloning of Qualitative Plant Genes Map-based cloning using the genetic relationship between a gene and a marker as the basis for beginning a search for a gene Chromosome walking moving toward
MOLECULAR GENETICS PROTEIN SYNTHESIS. Molecular Genetics Activity #2 page 1
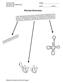 AP BIOLOGY MOLECULAR GENETICS ACTIVITY #2 NAME DATE HOUR PROTEIN SYNTHESIS Molecular Genetics Activity #2 page 1 GENETIC CODE PROTEIN SYNTHESIS OVERVIEW Molecular Genetics Activity #2 page 2 PROTEIN SYNTHESIS
AP BIOLOGY MOLECULAR GENETICS ACTIVITY #2 NAME DATE HOUR PROTEIN SYNTHESIS Molecular Genetics Activity #2 page 1 GENETIC CODE PROTEIN SYNTHESIS OVERVIEW Molecular Genetics Activity #2 page 2 PROTEIN SYNTHESIS
Introduction to Genomic Medicine Exeter Expert Series: Cardiology
 Introduction to Genomic Medicine Exeter Expert Series: Cardiology 2-3 November, 2017 Session I. Introductory concepts of genomics: gene, genome and variants Presented by Júlia Baptista PhD Clinical Scientist
Introduction to Genomic Medicine Exeter Expert Series: Cardiology 2-3 November, 2017 Session I. Introductory concepts of genomics: gene, genome and variants Presented by Júlia Baptista PhD Clinical Scientist
Gene Prediction. Mario Stanke. Institut für Mikrobiologie und Genetik Abteilung Bioinformatik. Gene Prediction p.
 Gene Prediction Mario Stanke mstanke@gwdg.de Institut für Mikrobiologie und Genetik Abteilung Bioinformatik Gene Prediction p.1/23 Why Predict Genes with a Computer? tons of data 39/250 eukaryotic/prokaryotic
Gene Prediction Mario Stanke mstanke@gwdg.de Institut für Mikrobiologie und Genetik Abteilung Bioinformatik Gene Prediction p.1/23 Why Predict Genes with a Computer? tons of data 39/250 eukaryotic/prokaryotic
Text Reference, Campbell v.8, chapter 17 PROTEIN SYNTHESIS
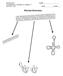 AP BIOLOGY Text Reference, Campbell v.8, chapter 17 ACTIVITY 1.22 NAME DATE HOUR PROTEIN SYNTHESIS GENETIC CODE PROTEIN SYNTHESIS OVERVIEW PROTEIN SYNTHESIS TRANSCRIPTION PROTEIN SYNTHESIS TRANSLATION
AP BIOLOGY Text Reference, Campbell v.8, chapter 17 ACTIVITY 1.22 NAME DATE HOUR PROTEIN SYNTHESIS GENETIC CODE PROTEIN SYNTHESIS OVERVIEW PROTEIN SYNTHESIS TRANSCRIPTION PROTEIN SYNTHESIS TRANSLATION
Structural variation. Marta Puig Institut de Biotecnologia i Biomedicina Universitat Autònoma de Barcelona
 Structural variation Marta Puig Institut de Biotecnologia i Biomedicina Universitat Autònoma de Barcelona Genetic variation How much genetic variation is there between individuals? What type of variants
Structural variation Marta Puig Institut de Biotecnologia i Biomedicina Universitat Autònoma de Barcelona Genetic variation How much genetic variation is there between individuals? What type of variants
Genetic Testing and Analysis. (858) MRN: Specimen: Saliva Received: 07/26/2016 GENETIC ANALYSIS REPORT
 GBinsight Sample Name: GB4408 Race: East Asian Gender: Female Reason for Testing: Family history of premature CAD MRN: 0123456790 Specimen: Saliva Received: 07/26/2016 Test ID: 113-1487118782-1 Test: Dyslipidemia
GBinsight Sample Name: GB4408 Race: East Asian Gender: Female Reason for Testing: Family history of premature CAD MRN: 0123456790 Specimen: Saliva Received: 07/26/2016 Test ID: 113-1487118782-1 Test: Dyslipidemia
Introduction to human genomics and genome informatics
 Introduction to human genomics and genome informatics Session 1 Prince of Wales Clinical School Dr Jason Wong ARC Future Fellow Head, Bioinformatics & Integrative Genomics Adult Cancer Program, Lowy Cancer
Introduction to human genomics and genome informatics Session 1 Prince of Wales Clinical School Dr Jason Wong ARC Future Fellow Head, Bioinformatics & Integrative Genomics Adult Cancer Program, Lowy Cancer
Human Genetic Variation. Ricardo Lebrón Dpto. Genética UGR
 Human Genetic Variation Ricardo Lebrón rlebron@ugr.es Dpto. Genética UGR What is Genetic Variation? Origins of Genetic Variation Genetic Variation is the difference in DNA sequences between individuals.
Human Genetic Variation Ricardo Lebrón rlebron@ugr.es Dpto. Genética UGR What is Genetic Variation? Origins of Genetic Variation Genetic Variation is the difference in DNA sequences between individuals.
Niemann-Pick Type C Disease Gene Variation Database ( )
 NPC-db (vs. 1.1) User Manual An introduction to the Niemann-Pick Type C Disease Gene Variation Database ( http://npc.fzk.de ) curated 2007/2008 by Dirk Dolle and Heiko Runz, Institute of Human Genetics,
NPC-db (vs. 1.1) User Manual An introduction to the Niemann-Pick Type C Disease Gene Variation Database ( http://npc.fzk.de ) curated 2007/2008 by Dirk Dolle and Heiko Runz, Institute of Human Genetics,
Training materials.
 Training materials - Ensembl training materials are protected by a CC BY license - http://creativecommons.org/licenses/by/4.0/ - If you wish to re-use these materials, please credit Ensembl for their creation
Training materials - Ensembl training materials are protected by a CC BY license - http://creativecommons.org/licenses/by/4.0/ - If you wish to re-use these materials, please credit Ensembl for their creation
Mapping strategies for sequence reads
 Mapping strategies for sequence reads Ernest Turro University of Cambridge 21 Oct 2013 Quantification A basic aim in genomics is working out the contents of a biological sample. 1. What distinct elements
Mapping strategies for sequence reads Ernest Turro University of Cambridge 21 Oct 2013 Quantification A basic aim in genomics is working out the contents of a biological sample. 1. What distinct elements
BIOL 300 Foundations of Biology Summer 2017 Telleen Lecture Outline
 BIOL 300 Foundations of Biology Summer 2017 Telleen Lecture Outline RNA, the Genetic Code, Proteins I. How RNA differs from DNA A. The sugar ribose replaces deoxyribose. The presence of the oxygen on the
BIOL 300 Foundations of Biology Summer 2017 Telleen Lecture Outline RNA, the Genetic Code, Proteins I. How RNA differs from DNA A. The sugar ribose replaces deoxyribose. The presence of the oxygen on the
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature13127 Factors to consider in assessing candidate pathogenic mutations in presumed monogenic conditions The questions itemized below expand upon the definitions in Table 1 and are provided
doi:10.1038/nature13127 Factors to consider in assessing candidate pathogenic mutations in presumed monogenic conditions The questions itemized below expand upon the definitions in Table 1 and are provided
Chapter 5. Structural Genomics
 Chapter 5. Structural Genomics Contents 5. Structural Genomics 5.1. DNA Sequencing Strategies 5.1.1. Map-based Strategies 5.1.2. Whole Genome Shotgun Sequencing 5.2. Genome Annotation 5.2.1. Using Bioinformatic
Chapter 5. Structural Genomics Contents 5. Structural Genomics 5.1. DNA Sequencing Strategies 5.1.1. Map-based Strategies 5.1.2. Whole Genome Shotgun Sequencing 5.2. Genome Annotation 5.2.1. Using Bioinformatic
Release Notes for Genomes Processed Using Complete Genomics Software
 Release Notes for Genomes Processed Using Complete Genomics Software Version 1.11.0 Related Documents... 1 Changes to Version 1.11.0... 2 Changes to Version 1.10.0... 6 Changes to Version 1.9.0... 10 Changes
Release Notes for Genomes Processed Using Complete Genomics Software Version 1.11.0 Related Documents... 1 Changes to Version 1.11.0... 2 Changes to Version 1.10.0... 6 Changes to Version 1.9.0... 10 Changes
Standards for Pathology Informatics in Australia (SPIA)
 Standards for Pathology Informatics in Australia (SPIA) Reporting Terminology and Codes Genetic Pathology (v3.0) Superseding and incorporating the Australian Pathology Units and Terminology Standards and
Standards for Pathology Informatics in Australia (SPIA) Reporting Terminology and Codes Genetic Pathology (v3.0) Superseding and incorporating the Australian Pathology Units and Terminology Standards and
MATH 5610, Computational Biology
 MATH 5610, Computational Biology Lecture 2 Intro to Molecular Biology (cont) Stephen Billups University of Colorado at Denver MATH 5610, Computational Biology p.1/24 Announcements Error on syllabus Class
MATH 5610, Computational Biology Lecture 2 Intro to Molecular Biology (cont) Stephen Billups University of Colorado at Denver MATH 5610, Computational Biology p.1/24 Announcements Error on syllabus Class
Lecture 2: Biology Basics Continued. Fall 2018 August 23, 2018
 Lecture 2: Biology Basics Continued Fall 2018 August 23, 2018 Genetic Material for Life Central Dogma DNA: The Code of Life The structure and the four genomic letters code for all living organisms Adenine,
Lecture 2: Biology Basics Continued Fall 2018 August 23, 2018 Genetic Material for Life Central Dogma DNA: The Code of Life The structure and the four genomic letters code for all living organisms Adenine,
Aligning GENCODE and RefSeq transcripts By EMBL-EBI and NCBI
 Aligning GENCODE and RefSeq transcripts By EMBL-EBI and NCBI Joannella Morales, Ph.D. LRG Project Manager jmorales@ebi.ac.uk contact@lrg-sequence.org https://www.lrg-sequence.org https://www.ensembl.org
Aligning GENCODE and RefSeq transcripts By EMBL-EBI and NCBI Joannella Morales, Ph.D. LRG Project Manager jmorales@ebi.ac.uk contact@lrg-sequence.org https://www.lrg-sequence.org https://www.ensembl.org
13.1 RNA Lesson Objectives Contrast RNA and DNA. Explain the process of transcription.
 13.1 RNA Lesson Objectives Contrast RNA and DNA. Explain the process of transcription. The Role of RNA 1. Complete the table to contrast the structures of DNA and RNA. DNA Sugar Number of Strands Bases
13.1 RNA Lesson Objectives Contrast RNA and DNA. Explain the process of transcription. The Role of RNA 1. Complete the table to contrast the structures of DNA and RNA. DNA Sugar Number of Strands Bases
BIOSTAT516 Statistical Methods in Genetic Epidemiology Autumn 2005 Handout1, prepared by Kathleen Kerr and Stephanie Monks
 Rationale of Genetic Studies Some goals of genetic studies include: to identify the genetic causes of phenotypic variation develop genetic tests o benefits to individuals and to society are still uncertain
Rationale of Genetic Studies Some goals of genetic studies include: to identify the genetic causes of phenotypic variation develop genetic tests o benefits to individuals and to society are still uncertain
Genetics Transcription Translation Replication
 Genetics Transcription Translation Replication 1. Which statement best describes the relationship between an allele and a gene? A. An allele is a variation of a gene that can be expressed as a phenotype.
Genetics Transcription Translation Replication 1. Which statement best describes the relationship between an allele and a gene? A. An allele is a variation of a gene that can be expressed as a phenotype.
BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers
 BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers Web resources: NCBI database: http://www.ncbi.nlm.nih.gov/ Ensembl database: http://useast.ensembl.org/index.html UCSC
BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers Web resources: NCBI database: http://www.ncbi.nlm.nih.gov/ Ensembl database: http://useast.ensembl.org/index.html UCSC
Degenerate site - twofold degenerate site - fourfold degenerate site
 Genetic code Codon: triple base pairs defining each amino acid. Why genetic code is triple? double code represents 4 2 = 16 different information triple code: 4 3 = 64 (two much to represent 20 amino acids)
Genetic code Codon: triple base pairs defining each amino acid. Why genetic code is triple? double code represents 4 2 = 16 different information triple code: 4 3 = 64 (two much to represent 20 amino acids)
Genome annotation & EST
 Genome annotation & EST What is genome annotation? The process of taking the raw DNA sequence produced by the genome sequence projects and adding the layers of analysis and interpretation necessary
Genome annotation & EST What is genome annotation? The process of taking the raw DNA sequence produced by the genome sequence projects and adding the layers of analysis and interpretation necessary
Product Description SALSA MLPA Probemix P099-C3 GCH1-TH-SGCE To be used with the MLPA General Protocol.
 Product Description SALSA Probemix P099-C3 GCH1-TH-SGCE To be used with the MLPA General Protocol. Version C3. As compared to version C2, one reference probe has been replaced and one probe has a small
Product Description SALSA Probemix P099-C3 GCH1-TH-SGCE To be used with the MLPA General Protocol. Version C3. As compared to version C2, one reference probe has been replaced and one probe has a small
Question 2: There are 5 retroelements (2 LINEs and 3 LTRs), 6 unclassified elements (XDMR and XDMR_DM), and 7 satellite sequences.
 Bio4342 Exercise 1 Answers: Detecting and Interpreting Genetic Homology (Answers prepared by Wilson Leung) Question 1: Low complexity DNA can be described as sequences that consist primarily of one or
Bio4342 Exercise 1 Answers: Detecting and Interpreting Genetic Homology (Answers prepared by Wilson Leung) Question 1: Low complexity DNA can be described as sequences that consist primarily of one or
Chapter 14: Genes in Action
 Chapter 14: Genes in Action Section 1: Mutation and Genetic Change Mutation: Nondisjuction: a failure of homologous chromosomes to separate during meiosis I or the failure of sister chromatids to separate
Chapter 14: Genes in Action Section 1: Mutation and Genetic Change Mutation: Nondisjuction: a failure of homologous chromosomes to separate during meiosis I or the failure of sister chromatids to separate
14 March, 2016: Introduction to Genomics
 14 March, 2016: Introduction to Genomics Genome Genome within Ensembl browser http://www.ensembl.org/homo_sapiens/location/view?db=core;g=ensg00000139618;r=13:3231547432400266 Genome within Ensembl browser
14 March, 2016: Introduction to Genomics Genome Genome within Ensembl browser http://www.ensembl.org/homo_sapiens/location/view?db=core;g=ensg00000139618;r=13:3231547432400266 Genome within Ensembl browser
DNA polymorphisms and RNA-Seq alternative splicing blow bubbles in de Bruijn Graphs
 DNA polymorphisms and RNA-Seq alternative splicing blow bubbles in de Bruijn Graphs Nadia Pisanti University of Pisa & Leiden University Outline New Generation Sequencing (NGS), and the importance of detecting
DNA polymorphisms and RNA-Seq alternative splicing blow bubbles in de Bruijn Graphs Nadia Pisanti University of Pisa & Leiden University Outline New Generation Sequencing (NGS), and the importance of detecting
In addition, 10 reference probes are included in this probemix, detecting several different autosomal chromosomal locations.
 SALSA MLPA probemix P179-B1 Limb Malformations-1 Lot B1-1014, B1-0611. As compared to the previous lot A2 (lot A2-0311), one GLI3 and three ROR2 probes have been replaced. Four extra GLI3 probes and one
SALSA MLPA probemix P179-B1 Limb Malformations-1 Lot B1-1014, B1-0611. As compared to the previous lot A2 (lot A2-0311), one GLI3 and three ROR2 probes have been replaced. Four extra GLI3 probes and one
Week 1 BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers
 Week 1 BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers Web resources: NCBI database: http://www.ncbi.nlm.nih.gov/ Ensembl database: http://useast.ensembl.org/index.html
Week 1 BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers Web resources: NCBI database: http://www.ncbi.nlm.nih.gov/ Ensembl database: http://useast.ensembl.org/index.html
Applied Bioinformatics
 Applied Bioinformatics In silico and In clinico characterization of genetic variations Assistant Professor Department of Biomedical Informatics Center for Human Genetics Research ATCAAAATTATGGAAGAA ATCAAAATCATGGAAGAA
Applied Bioinformatics In silico and In clinico characterization of genetic variations Assistant Professor Department of Biomedical Informatics Center for Human Genetics Research ATCAAAATTATGGAAGAA ATCAAAATCATGGAAGAA
Lecture for Wednesday. Dr. Prince BIOL 1408
 Lecture for Wednesday Dr. Prince BIOL 1408 THE FLOW OF GENETIC INFORMATION FROM DNA TO RNA TO PROTEIN Copyright 2009 Pearson Education, Inc. Genes are expressed as proteins A gene is a segment of DNA that
Lecture for Wednesday Dr. Prince BIOL 1408 THE FLOW OF GENETIC INFORMATION FROM DNA TO RNA TO PROTEIN Copyright 2009 Pearson Education, Inc. Genes are expressed as proteins A gene is a segment of DNA that
Lecture 11: Gene Prediction
 Lecture 11: Gene Prediction Study Chapter 6.11-6.14 1 Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and end positions of genes in a genome Where are
Lecture 11: Gene Prediction Study Chapter 6.11-6.14 1 Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and end positions of genes in a genome Where are
Materials Protein synthesis kit. This kit consists of 24 amino acids, 24 transfer RNAs, four messenger RNAs and one ribosome (see below).
 Protein Synthesis Instructions The purpose of today s lab is to: Understand how a cell manufactures proteins from amino acids, using information stored in the genetic code. Assemble models of four very
Protein Synthesis Instructions The purpose of today s lab is to: Understand how a cell manufactures proteins from amino acids, using information stored in the genetic code. Assemble models of four very
The Nature of Genes. The Nature of Genes. Genes and How They Work. Chapter 15/16
 Genes and How They Work Chapter 15/16 The Nature of Genes Beadle and Tatum proposed the one gene one enzyme hypothesis. Today we know this as the one gene one polypeptide hypothesis. 2 The Nature of Genes
Genes and How They Work Chapter 15/16 The Nature of Genes Beadle and Tatum proposed the one gene one enzyme hypothesis. Today we know this as the one gene one polypeptide hypothesis. 2 The Nature of Genes
GENETICS - CLUTCH CH.15 GENOMES AND GENOMICS.
 !! www.clutchprep.com CONCEPT: OVERVIEW OF GENOMICS Genomics is the study of genomes in their entirety Bioinformatics is the analysis of the information content of genomes - Genes, regulatory sequences,
!! www.clutchprep.com CONCEPT: OVERVIEW OF GENOMICS Genomics is the study of genomes in their entirety Bioinformatics is the analysis of the information content of genomes - Genes, regulatory sequences,
Ensembl Tools. EBI is an Outstation of the European Molecular Biology Laboratory.
 Ensembl Tools EBI is an Outstation of the European Molecular Biology Laboratory. Questions? We ve muted all the mics Ask questions in the Chat box in the webinar interface I will check the Chat box periodically
Ensembl Tools EBI is an Outstation of the European Molecular Biology Laboratory. Questions? We ve muted all the mics Ask questions in the Chat box in the webinar interface I will check the Chat box periodically
Analyzed Fungi Neurospora crassa mutants. Mutants were UNABLE to grow without Arginine (an amino acid) Other biochemical experiments indicated:
 From Gene to Protein Beadle and Tatum Analyzed Fungi Neurospora crassa mutants Mutants were UNABLE to grow without Arginine (an amino acid) Other biochemical experiments indicated: Precursor Ornithine
From Gene to Protein Beadle and Tatum Analyzed Fungi Neurospora crassa mutants Mutants were UNABLE to grow without Arginine (an amino acid) Other biochemical experiments indicated: Precursor Ornithine
High-throughput BRCA1 mutation scanning using HR-MCA MCA. Evaluation project :
 Evaluation project : ESHG 17 june 2007 High-throughput BRCA1 mutation scanning using HR-MCA MCA Department of Clinical Genetics Leiden Institute of Biology and Medical Genetics, Prague, Czech Republic
Evaluation project : ESHG 17 june 2007 High-throughput BRCA1 mutation scanning using HR-MCA MCA Department of Clinical Genetics Leiden Institute of Biology and Medical Genetics, Prague, Czech Republic
7.03 Problem Set 3 Due before 5 PM on Wednesday, October 18 Hand in answers in recitation section or in the box outside of
 7.03 Problem Set 3 Due before 5 PM on Wednesday, October 18 Hand in answers in recitation section or in the box outside of 68-120 1. The following DNA sequence fragment comes from the middle of a bacterial
7.03 Problem Set 3 Due before 5 PM on Wednesday, October 18 Hand in answers in recitation section or in the box outside of 68-120 1. The following DNA sequence fragment comes from the middle of a bacterial
1. DNA, RNA structure. 2. DNA replication. 3. Transcription, translation
 1. DNA, RNA structure 2. DNA replication 3. Transcription, translation DNA and RNA are polymers of nucleotides DNA is a nucleic acid, made of long chains of nucleotides Nucleotide Phosphate group Nitrogenous
1. DNA, RNA structure 2. DNA replication 3. Transcription, translation DNA and RNA are polymers of nucleotides DNA is a nucleic acid, made of long chains of nucleotides Nucleotide Phosphate group Nitrogenous
CS273B: Deep Learning in Genomics and Biomedicine. Recitation 1 30/9/2016
 CS273B: Deep Learning in Genomics and Biomedicine. Recitation 1 30/9/2016 Topics Genetic variation Population structure Linkage disequilibrium Natural disease variants Genome Wide Association Studies Gene
CS273B: Deep Learning in Genomics and Biomedicine. Recitation 1 30/9/2016 Topics Genetic variation Population structure Linkage disequilibrium Natural disease variants Genome Wide Association Studies Gene
Browsing Genes and Genomes with Ensembl
 Browsing Genes and Genomes with Ensembl Victoria Newman Ensembl Outreach Officer EMBL-EBI Objectives What is Ensembl? What type of data can you get in Ensembl? How to navigate the Ensembl browser website.
Browsing Genes and Genomes with Ensembl Victoria Newman Ensembl Outreach Officer EMBL-EBI Objectives What is Ensembl? What type of data can you get in Ensembl? How to navigate the Ensembl browser website.
Authors: Vivek Sharma and Ram Kunwar
 Molecular markers types and applications A genetic marker is a gene or known DNA sequence on a chromosome that can be used to identify individuals or species. Why we need Molecular Markers There will be
Molecular markers types and applications A genetic marker is a gene or known DNA sequence on a chromosome that can be used to identify individuals or species. Why we need Molecular Markers There will be
RNA and PROTEIN SYNTHESIS. Chapter 13
 RNA and PROTEIN SYNTHESIS Chapter 13 DNA Double stranded Thymine Sugar is RNA Single stranded Uracil Sugar is Ribose Deoxyribose Types of RNA 1. Messenger RNA (mrna) Carries copies of instructions from
RNA and PROTEIN SYNTHESIS Chapter 13 DNA Double stranded Thymine Sugar is RNA Single stranded Uracil Sugar is Ribose Deoxyribose Types of RNA 1. Messenger RNA (mrna) Carries copies of instructions from
132 Grundlagen der Bioinformatik, SoSe 14, D. Huson, June 22, This exposition is based on the following source, which is recommended reading:
 132 Grundlagen der Bioinformatik, SoSe 14, D. Huson, June 22, 214 1 Gene Prediction Using HMMs This exposition is based on the following source, which is recommended reading: 1. Chris Burge and Samuel
132 Grundlagen der Bioinformatik, SoSe 14, D. Huson, June 22, 214 1 Gene Prediction Using HMMs This exposition is based on the following source, which is recommended reading: 1. Chris Burge and Samuel
Biological Sciences 50 Practice Exam 2
 NAME: Fall 2005 TF: Biological Sciences 50 Practice Exam 2 A. Be sure to write your name on the top of each of page of the examination. B. Write each answer only on the same page as the pertinent question.
NAME: Fall 2005 TF: Biological Sciences 50 Practice Exam 2 A. Be sure to write your name on the top of each of page of the examination. B. Write each answer only on the same page as the pertinent question.
Lecture 19A. DNA computing
 Lecture 19A. DNA computing What exactly is DNA (deoxyribonucleic acid)? DNA is the material that contains codes for the many physical characteristics of every living creature. Your cells use different
Lecture 19A. DNA computing What exactly is DNA (deoxyribonucleic acid)? DNA is the material that contains codes for the many physical characteristics of every living creature. Your cells use different
Grundlagen der Bioinformatik, SoSe 11, D. Huson, July 4, This exposition is based on the following source, which is recommended reading:
 Grundlagen der Bioinformatik, SoSe 11, D. Huson, July 4, 211 155 12 Gene Prediction Using HMMs This exposition is based on the following source, which is recommended reading: 1. Chris Burge and Samuel
Grundlagen der Bioinformatik, SoSe 11, D. Huson, July 4, 211 155 12 Gene Prediction Using HMMs This exposition is based on the following source, which is recommended reading: 1. Chris Burge and Samuel
Sample to Insight. Dr. Bhagyashree S. Birla NGS Field Application Scientist
 Dr. Bhagyashree S. Birla NGS Field Application Scientist bhagyashree.birla@qiagen.com NGS spans a broad range of applications DNA Applications Human ID Liquid biopsy Biomarker discovery Inherited and somatic
Dr. Bhagyashree S. Birla NGS Field Application Scientist bhagyashree.birla@qiagen.com NGS spans a broad range of applications DNA Applications Human ID Liquid biopsy Biomarker discovery Inherited and somatic
Furthermore, 8 reference probes are included in this probemix, detecting several different autosomal chromosomal locations.
 SALSA MLPA mix P328-A2 EYS Lot A2-0217. As compared to version A1, two reference s have been replaced and one length has been adjusted. Retinitis Pigmentosa-25 (RP25) is characterised by progressive peripheral
SALSA MLPA mix P328-A2 EYS Lot A2-0217. As compared to version A1, two reference s have been replaced and one length has been adjusted. Retinitis Pigmentosa-25 (RP25) is characterised by progressive peripheral
CITATION FILE CONTENT / FORMAT
 CITATION 1) For any resultant publications using single samples please cite: Matthew A. Field, Vicky Cho, T. Daniel Andrews, and Chris C. Goodnow (2015). "Reliably detecting clinically important variants
CITATION 1) For any resultant publications using single samples please cite: Matthew A. Field, Vicky Cho, T. Daniel Andrews, and Chris C. Goodnow (2015). "Reliably detecting clinically important variants
Use of Minigenes in a Diagnostic Laboratory. Allan Richards. Cambridge
 Use of Minigenes in a Diagnostic Laboratory Allan Richards Cambridge Minigene Analysis Minigenes are used to assess the effect of sequence variants on the processing of pre-mrna into the final message
Use of Minigenes in a Diagnostic Laboratory Allan Richards Cambridge Minigene Analysis Minigenes are used to assess the effect of sequence variants on the processing of pre-mrna into the final message
American Board of Medical Genetics and Genomics
 American Board of Medical Genetics and Genomics Logbook Guidelines for Certification in Laboratory Genetics and Genomics for the 2019 Examination as of 11/10/2016 Purpose: The purpose of the logbook is
American Board of Medical Genetics and Genomics Logbook Guidelines for Certification in Laboratory Genetics and Genomics for the 2019 Examination as of 11/10/2016 Purpose: The purpose of the logbook is
CH 17 :From Gene to Protein
 CH 17 :From Gene to Protein Defining a gene gene gene Defining a gene is problematic because one gene can code for several protein products, some genes code only for RNA, two genes can overlap, and there
CH 17 :From Gene to Protein Defining a gene gene gene Defining a gene is problematic because one gene can code for several protein products, some genes code only for RNA, two genes can overlap, and there
Axiom mydesign Custom Array design guide for human genotyping applications
 TECHNICAL NOTE Axiom mydesign Custom Genotyping Arrays Axiom mydesign Custom Array design guide for human genotyping applications Overview In the past, custom genotyping arrays were expensive, required
TECHNICAL NOTE Axiom mydesign Custom Genotyping Arrays Axiom mydesign Custom Array design guide for human genotyping applications Overview In the past, custom genotyping arrays were expensive, required
ENGR 213 Bioengineering Fundamentals April 25, A very coarse introduction to bioinformatics
 A very coarse introduction to bioinformatics In this exercise, you will get a quick primer on how DNA is used to manufacture proteins. You will learn a little bit about how the building blocks of these
A very coarse introduction to bioinformatics In this exercise, you will get a quick primer on how DNA is used to manufacture proteins. You will learn a little bit about how the building blocks of these
Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR
 Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR 1 The problem We wish to clone a yet unknown gene from a known
Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR 1 The problem We wish to clone a yet unknown gene from a known
SUPPLEMENTAL MATERIAL GENOTYPING WITH MULTIPLEXING TARGETED RESEQUENCING
 SUPPLEMENTAL MATERIAL GENOTYPING WITH MULTIPLEXING TARGETED RESEQUENCING All of the patients and control subjects were sequenced and genotyped in the same way. Shotgun libraries of approximately 250 bp
SUPPLEMENTAL MATERIAL GENOTYPING WITH MULTIPLEXING TARGETED RESEQUENCING All of the patients and control subjects were sequenced and genotyped in the same way. Shotgun libraries of approximately 250 bp
Computational Genomics. Ron Shamir & Roded Sharan Fall
 Computational Genomics Ron Shamir & Roded Sharan Fall 2012-13 Bioinformatics The information science of biology: organize, store, analyze and visualize biological data Responds to the explosion of biological
Computational Genomics Ron Shamir & Roded Sharan Fall 2012-13 Bioinformatics The information science of biology: organize, store, analyze and visualize biological data Responds to the explosion of biological
From DNA to Protein: Genotype to Phenotype
 12 From DNA to Protein: Genotype to Phenotype 12.1 What Is the Evidence that Genes Code for Proteins? The gene-enzyme relationship is one-gene, one-polypeptide relationship. Example: In hemoglobin, each
12 From DNA to Protein: Genotype to Phenotype 12.1 What Is the Evidence that Genes Code for Proteins? The gene-enzyme relationship is one-gene, one-polypeptide relationship. Example: In hemoglobin, each
SALSA MLPA probemix P074-A3 Androgen Receptor (AR) Lot A Compared to previous lot A2-0712, three reference probes have been replaced.
 SALSA MLPA probemix P074-A3 Androgen Receptor (AR) Lot A3-0814. Compared to previous lot A2-0712, three reference probes have been replaced. The androgen insensitivity syndrome (AIS), formerly known as
SALSA MLPA probemix P074-A3 Androgen Receptor (AR) Lot A3-0814. Compared to previous lot A2-0712, three reference probes have been replaced. The androgen insensitivity syndrome (AIS), formerly known as
BIOLOGY - CLUTCH CH.17 - GENE EXPRESSION.
 !! www.clutchprep.com CONCEPT: GENES Beadle and Tatum develop the one gene one enzyme hypothesis through their work with Neurospora (bread mold). This idea was later revised as the one gene one polypeptide
!! www.clutchprep.com CONCEPT: GENES Beadle and Tatum develop the one gene one enzyme hypothesis through their work with Neurospora (bread mold). This idea was later revised as the one gene one polypeptide
NUCLEOTIDE RESOLUTION STRUCTURAL VARIATION DETECTION USING NEXT- GENERATION WHOLE GENOME RESEQUENCING
 NUCLEOTIDE RESOLUTION STRUCTURAL VARIATION DETECTION USING NEXT- GENERATION WHOLE GENOME RESEQUENCING Ken Chen, Ph.D. kchen@genome.wustl.edu The Genome Center, Washington University in St. Louis The path
NUCLEOTIDE RESOLUTION STRUCTURAL VARIATION DETECTION USING NEXT- GENERATION WHOLE GENOME RESEQUENCING Ken Chen, Ph.D. kchen@genome.wustl.edu The Genome Center, Washington University in St. Louis The path
