Enhancing motif finding models using multiple sources of genome-wide data
|
|
|
- Sharleen Marilynn Dixon
- 6 years ago
- Views:
Transcription
1 Enhancing motif finding models using multiple sources of genome-wide data Heejung Shim 1, Oliver Bembom 2, Sündüz Keleş 3,4 1 Department of Human Genetics, University of Chicago, 2 Division of Biostatistics, University of California, Berkeley, 3 Department of Statistics, 4 Department of Biostatistics and Medical Informatics, University of Wisconsin. April 21, Overview The SUCcESS package implements the CTCM model (particularly logistic regression model) proposed by Shim and Keleş (2008) for integrating quantitative information into motif finding as well as its extension to use multiple data sources at a time (e.g., ChIP-seq, nucleosome occupancy, or conservation score). We implemented them as an extended module for cosmo, developed by Bembom et al. (2007), implementing an algorithm which supervises detection of motifs using a set of constraints for a position weight matrix. Note that although this package provides all functions implemented in cosmo, we ask that you instead use the latest version of cosmo for the supervised motif detection algorithm. This package is for running SUCcESS option only. This vignette just provides the instructions for how to run SUCcESS: you will need to consult Shim and Keleş (2008) for methodological details and the cosmo vignette (you can find it in the document directory of this package) for detailed options. 1
2 2 Brief Background In addition to ChIP-seq (chromatin immunoprecipitation followed by sequencing), various sources of genomic data have been available to improve detection of transcription factor binding sites. For example, functional regions are prone to be conserved across related species (Kellis et al., 2003), and thus conservation scores obtained by multi-species sequencing can provide information about which regions are more likely to have motifs. As another example, nucleosomes are know to interact with transcription factor bindings and less likely occupy in active regulatory regions (Segal et al., 2006)(Lee et al., 2004). Thus, nucleosome occupancy measured by either ChIP-seq experiments for histone modifications (Lee et al., 2004) or MNase-seq (micrococcal nuclease digestion followed by sequencing) (Shivaswamy et al., 2008)(Albert et al., 2007) can be used as an additional genomic data. Any other data type which provides information on transcription factor binding sites can be integrated into our approach. Our previous paper (Shim and Keleş, 2008) introduced the method for using ChIP-chip only. Here we extend one of the proposed three models to use those multiple data sources at a time. Let T ik = (Tik 1,..., T ik M) denote M genomic data for kth base pair in sequence i. Then, the logistic regression model in Shim and Keleş (2008) can be extended to log ( P r(zik = 1 T ik ) ) 1 P r(z ik = 1 T ik ) = β 0 + β 1 T 1 ik β M T M ik, (1) where Z ik is the indicator variable denoting whether a motif starts at position k in sequence i and β j 0 for j = 1,..., M. When multiple sources of genomic data are provided by users, SUCcESS will use the extended model. 3 Input Data SUCcESS requires two types of input files: for (1) sequence data and (2) additional genomic data. The sequence data should be provided in FASTA format. Here is an example with two sequences. >seq1 tcgtagcccaa >seq2 tcgtagggcccgt 2
3 Two types of input formats can be used for the genomic data depending on the resolution of data. In both formats, each row contains information on each sequence, with one row per sequence. The sequence-level input format lists one number at each row as shown in an example below In the base pair-level input format, each row includes a list of numbers each of which corresponds to each base-pair in the sequence. The list should have the same length as the corresponding sequence. The following example corresponds to the sequence data in FASTA format above For neither sequence-level nor base pair-level genomic data (e.g., probe-level), the value of a measurement unit could be assigned to all bases within the unit or interpolation with simple averaging could be employed to obtain values at the bases between units if necessary. We assume that the genomic data at a particular position of a given sequence have a direct relationship with the probability that the position of the sequence is a motif start site. For the data who do not have a direct relationship with the motif start site probability (e.g., nucleosome occupancy), you will need to transform them. When multiple sources genomic data are available, each genomic data should be provided separately in an appropriate format. 4 run SUCcESS This section will describe how to run SUCcESS using simulated data. package can be loaded using the command The >library(success) and SUCcESS can be run using for example >seqfile = system.file("exfiles/seqdata", package="success") >genomicfile1 = system.file("exfiles/genomicdata1", package="success") >genomicfile2 = system.file("exfiles/genomicdata2", package="success") >ress = success(seqs=seqfile, ChIP=c(genomicFile1, genomicfile2), models = c("tcm"), maxw = 10, minw= 9, starts = 2, nummotifs = 2) 3
4 Sequence data and multiple genomic data can be provided using the arguments seqs and ChIP, respectively. SUCcESS allows users to specify the model types to be considered among the three possible candidates (OOPS ZOOPS and TCM) using the argument models (models = c("zoops") by default). The arguments minw and maxw can be used to provide the minimum and maximum motif widths to be considered (minw = 6 and maxw = 15 by default), respectively. SUCcESS sets the number of starting values to be two times of the number given by the argument starts (starts = 5 by default). SUCcESS can detect multiple motifs whose number is given by nummotifs (1 by default) as MEME (Bailey and Elkan, 1995) does. Please consult the description of cosmo function in the cosmo vignette for the other arguments. The print function shows the estimated position weight matrices > print(ress) Motif 1: A C G T Motif 2: A C G T and the estimated coefficients in the logistic regression model are listed in a more detailed summery of the results (slot Estimated beta). > summary(ress) Input dataset: Sequence Length 1 seq seq
5 3 seq seq seq5 100 Candidate orders for background Markov model: order kldiv e e e Inf 5 4 Inf 6 5 Inf 7 6 Inf Motif 1: Candidate models considered: conset model width wcrit modcrit concrit 1 1 TCM NA NA 2 1 TCM NA NA Selected model: choice crit critval Constraint 1 likcv NA Model TCM lik NA Width 10 bic NumSites 8 lik Markov Order 0 likcv Estimated position weight matrix: A C G T
6 Estimated beta: beta Motif occurrences: E-value: seq pos orient motif prob 1 seq AACACTTATC e+00 2 seq TAGATTTCCA e+00 3 seq AAGACTTATC e+00 4 seq AAGACTTACC e-01 5 seq AATACTTACC e-01 6 seq TGTAAGTCTT e-01 7 seq AAAACTCACC e-01 8 seq AAAATGCATC e-19 Motif 2: Candidate models considered: conset model width wcrit modcrit concrit 1 1 TCM NA NA 2 1 TCM NA NA Selected model: choice crit critval Constraint 1 likcv NA Model TCM lik NA Width 10 bic NumSites 8 lik Markov Order 0 likcv Estimated position weight matrix: 6
7 A C G T Estimated beta: beta Motif occurrences: E-value: seq pos orient motif prob 1 seq GGGCCTTACT seq2 2-1 CGTAGGGCCC seq CGTGAAGCGG seq CGTAATCCAC seq CTGCCATAGC seq CAGTATTACG seq ATACCTTACG seq GGTGATGTGG Sequence logo of the estimated motifs can be generated one at a time using the command: > plot(ress, index = 1) The argument index indicates which motif is used to make a plot. The sequence logo of the first motif (index = 1) is shown in Figure 1. The argument type = "prob" makes the plot function produce a plot of the posterior probabilities along each sequence. The plot in Figure 2 is created by the command: > plot(ress, type = "prob", index = 1) 7
8 2 1.5 Information content Position Figure 1: Sequence logo (motif 1) seq1 seq2 seq3 seq4 seq Figure 2: Posterior probabilities (motif 1) 8
9 References Albert, I., Mavrich, T. N., Tomsho, L. P., Qi, J., Zanton, S. J., Schuster, S. C. and Pugh, B. F. (2007) Translational and rotational settings of h2a.z nucleosomes across the saccharomycescerevisiae genome. Nature, 446, Bailey, T. and Elkan, C. (1995) Unsupervised learning of multiple motifs in biopolymers using expectation maximization. Machine Learning, 21, Bembom, O., Keleş, S. and van der Laan, M. J. (2007) Supervised detection of conserved motifs in DNA sequences with cosmo. Statistical Applications in Genetics and Molecular Biology, 6, Article 8. Kellis, M., Patterson, N., Endrizzi, M., Birren, B. and Lander, E. S. (2003) Sequencing and comparison of yeast species to identify genes and regulatory elements. Nature, 423, Lee, C.-K., Shibata, Y., Rao, B., Strahl, B. D. and Lieb, J. D. (2004) Evidence for nucleosome depletion at active regulatory regions genomewide. Nature Genetics, 36, Segal, E., Fondufe-Mittendorf, Y., Chen, L., Thastrom, A., Field, Y., Moore, I. K., Wang, J. Z. and Widom, J. (2006) A genomic code for nucleosome positioning. Nature, 442, Shim, H. and Keleş, S. (2008) Integrating quantitative information from ChIPchip experiments into motif finding. Biostatistics, 9, Shivaswamy, S., Bhinge, A., Zhao, Y., Jones, S., Hirst, M. and Iyer, V. R. (2008) Dynamic remodeling of individual nucleosomes across a eukaryotic genome in response to transcriptional perturbation. PLoS Biology, 6, e65. 9
Epigenetics and DNase-Seq
 Epigenetics and DNase-Seq BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2018 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material, are licensed under CC BY-NC 4.0 by Anthony
Epigenetics and DNase-Seq BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2018 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material, are licensed under CC BY-NC 4.0 by Anthony
Supplementary Information: Conserved nucleosome positioning defines replication origins
 Supplementary Information: Conserved nucleosome positioning defines replication origins Matthew L. Eaton 1, Kyriaki Galani 2, Sukhyun Kang 2, Stephen P. Bell 2, and David M. MacAlpine 1 1 Department of
Supplementary Information: Conserved nucleosome positioning defines replication origins Matthew L. Eaton 1, Kyriaki Galani 2, Sukhyun Kang 2, Stephen P. Bell 2, and David M. MacAlpine 1 1 Department of
Improvement of TRANSFAC Matrices Using Multiple Local Alignment of Transcription Factor Binding Site Sequences
 68 Genome Informatics 16(1): 68 72 (2005) Improvement of TRANSFAC Matrices Using Multiple Local Alignment of Transcription Factor Binding Site Sequences Yutao Fu 1 Zhiping Weng 1,2 bibin@bu.edu zhiping@bu.edu
68 Genome Informatics 16(1): 68 72 (2005) Improvement of TRANSFAC Matrices Using Multiple Local Alignment of Transcription Factor Binding Site Sequences Yutao Fu 1 Zhiping Weng 1,2 bibin@bu.edu zhiping@bu.edu
2/10/17. Contents. Applications of HMMs in Epigenomics
 2/10/17 I529: Machine Learning in Bioinformatics (Spring 2017) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2017 Background:
2/10/17 I529: Machine Learning in Bioinformatics (Spring 2017) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2017 Background:
Bioinformatics of Transcriptional Regulation
 Bioinformatics of Transcriptional Regulation Carl Herrmann IPMB & DKFZ c.herrmann@dkfz.de Wechselwirkung von Maßnahmen und Auswirkungen Einflussmöglichkeiten in einem Dialog From genes to active compounds
Bioinformatics of Transcriptional Regulation Carl Herrmann IPMB & DKFZ c.herrmann@dkfz.de Wechselwirkung von Maßnahmen und Auswirkungen Einflussmöglichkeiten in einem Dialog From genes to active compounds
THE ANALYSIS OF CHROMATIN CONDENSATION STATE AND TRANSCRIPTIONAL ACTIVITY USING DNA MICROARRAYS 1. INTRODUCTION
 JOURNAL OF MEDICAL INFORMATICS & TECHNOLOGIES Vol.6/2003, ISSN 1642-6037 Piotr WIDŁAK *, Krzysztof FUJAREWICZ ** DNA microarrays, chromatin, transcription THE ANALYSIS OF CHROMATIN CONDENSATION STATE AND
JOURNAL OF MEDICAL INFORMATICS & TECHNOLOGIES Vol.6/2003, ISSN 1642-6037 Piotr WIDŁAK *, Krzysztof FUJAREWICZ ** DNA microarrays, chromatin, transcription THE ANALYSIS OF CHROMATIN CONDENSATION STATE AND
Sequence logos for DNA sequence alignments
 Sequence logos for DNA sequence alignments Oliver Bembom Division of Biostatistics, University of California, Berkeley October, 202 Introduction An alignment of DNA or amino acid sequences is commonly
Sequence logos for DNA sequence alignments Oliver Bembom Division of Biostatistics, University of California, Berkeley October, 202 Introduction An alignment of DNA or amino acid sequences is commonly
Characterizing DNA binding sites high throughput approaches Biol4230 Tues, April 24, 2018 Bill Pearson Pinn 6-057
 Characterizing DNA binding sites high throughput approaches Biol4230 Tues, April 24, 2018 Bill Pearson wrp@virginia.edu 4-2818 Pinn 6-057 Reviewing sites: affinity and specificity representation binding
Characterizing DNA binding sites high throughput approaches Biol4230 Tues, April 24, 2018 Bill Pearson wrp@virginia.edu 4-2818 Pinn 6-057 Reviewing sites: affinity and specificity representation binding
Gene expression analysis. Biosciences 741: Genomics Fall, 2013 Week 5. Gene expression analysis
 Gene expression analysis Biosciences 741: Genomics Fall, 2013 Week 5 Gene expression analysis From EST clusters to spotted cdna microarrays Long vs. short oligonucleotide microarrays vs. RT-PCR Methods
Gene expression analysis Biosciences 741: Genomics Fall, 2013 Week 5 Gene expression analysis From EST clusters to spotted cdna microarrays Long vs. short oligonucleotide microarrays vs. RT-PCR Methods
XPRIME-EM: Eliciting Expert Prior Information for Motif Exploration Using the Expectation- Maximization Algorithm
 Brigham Young University BYU ScholarsArchive All Theses and Dissertations 2012-06-22 XPRIME-EM: Eliciting Expert Prior Information for Motif Exploration Using the Expectation- Maximization Algorithm Wei
Brigham Young University BYU ScholarsArchive All Theses and Dissertations 2012-06-22 XPRIME-EM: Eliciting Expert Prior Information for Motif Exploration Using the Expectation- Maximization Algorithm Wei
Transcriptional Regulatory Code of a Eukaryotic Genome
 Supplementary Information for Transcriptional Regulatory Code of a Eukaryotic Genome Christopher T. Harbison 1,2*, D. Benjamin Gordon 1*, Tong Ihn Lee 1, Nicola J. Rinaldi 1,2, Kenzie Macisaac 3, Timothy
Supplementary Information for Transcriptional Regulatory Code of a Eukaryotic Genome Christopher T. Harbison 1,2*, D. Benjamin Gordon 1*, Tong Ihn Lee 1, Nicola J. Rinaldi 1,2, Kenzie Macisaac 3, Timothy
Mixture modeling for genome-wide localization of transcription factors
 Mixture modeling for genome-wide localization of transcription factors Sündüz Keleş 1,2 and Heejung Shim 1 1 Department of Statistics 2 Department of Biostatistics & Medical Informatics University of Wisconsin,
Mixture modeling for genome-wide localization of transcription factors Sündüz Keleş 1,2 and Heejung Shim 1 1 Department of Statistics 2 Department of Biostatistics & Medical Informatics University of Wisconsin,
ChIP. November 21, 2017
 ChIP November 21, 2017 functional signals: is DNA enough? what is the smallest number of letters used by a written language? DNA is only one part of the functional genome DNA is heavily bound by proteins,
ChIP November 21, 2017 functional signals: is DNA enough? what is the smallest number of letters used by a written language? DNA is only one part of the functional genome DNA is heavily bound by proteins,
The ChIP-Seq project. Giovanna Ambrosini, Philipp Bucher. April 19, 2010 Lausanne. EPFL-SV Bucher Group
 The ChIP-Seq project Giovanna Ambrosini, Philipp Bucher EPFL-SV Bucher Group April 19, 2010 Lausanne Overview Focus on technical aspects Description of applications (C programs) Where to find binaries,
The ChIP-Seq project Giovanna Ambrosini, Philipp Bucher EPFL-SV Bucher Group April 19, 2010 Lausanne Overview Focus on technical aspects Description of applications (C programs) Where to find binaries,
CS273B: Deep learning for Genomics and Biomedicine
 CS273B: Deep learning for Genomics and Biomedicine Lecture 2: Convolutional neural networks and applications to functional genomics 09/28/2016 Anshul Kundaje, James Zou, Serafim Batzoglou Outline Anatomy
CS273B: Deep learning for Genomics and Biomedicine Lecture 2: Convolutional neural networks and applications to functional genomics 09/28/2016 Anshul Kundaje, James Zou, Serafim Batzoglou Outline Anatomy
DOUBLE-STRAND DNA BREAK PREDICTION USING EPIGENOME MARKS AT KILOBASE RESOLUTION
 DOUBLE-STRAND DNA BREAK PREDICTION USING EPIGENOME MARKS AT KILOBASE RESOLUTION Raphaël MOURAD, Assist. Prof. Centre de Biologie Intégrative Université Paul Sabatier, Toulouse III INTRODUCTION Double-strand
DOUBLE-STRAND DNA BREAK PREDICTION USING EPIGENOME MARKS AT KILOBASE RESOLUTION Raphaël MOURAD, Assist. Prof. Centre de Biologie Intégrative Université Paul Sabatier, Toulouse III INTRODUCTION Double-strand
A Brief History. Bootstrapping. Bagging. Boosting (Schapire 1989) Adaboost (Schapire 1995)
 A Brief History Bootstrapping Bagging Boosting (Schapire 1989) Adaboost (Schapire 1995) What s So Good About Adaboost Improves classification accuracy Can be used with many different classifiers Commonly
A Brief History Bootstrapping Bagging Boosting (Schapire 1989) Adaboost (Schapire 1995) What s So Good About Adaboost Improves classification accuracy Can be used with many different classifiers Commonly
NGS Approaches to Epigenomics
 I519 Introduction to Bioinformatics, 2013 NGS Approaches to Epigenomics Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Background: chromatin structure & DNA methylation Epigenomic
I519 Introduction to Bioinformatics, 2013 NGS Approaches to Epigenomics Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Background: chromatin structure & DNA methylation Epigenomic
ChIP-Seq Data Analysis. J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015
 ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA
ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA
ChIP-Seq Tools. J Fass UCD Genome Center Bioinformatics Core Wednesday September 16, 2015
 ChIP-Seq Tools J Fass UCD Genome Center Bioinformatics Core Wednesday September 16, 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA or
ChIP-Seq Tools J Fass UCD Genome Center Bioinformatics Core Wednesday September 16, 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA or
ChIP-Seq Data Analysis. J Fass UCD Genome Center Bioinformatics Core Wednesday December 17, 2014
 ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday December 17, 2014 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind
ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday December 17, 2014 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind
CSC 2427: Algorithms in Molecular Biology Lecture #14
 CSC 2427: Algorithms in Molecular Biology Lecture #14 Lecturer: Michael Brudno Scribe Note: Hyonho Lee Department of Computer Science University of Toronto 03 March 2006 Microarrays Revisited In the last
CSC 2427: Algorithms in Molecular Biology Lecture #14 Lecturer: Michael Brudno Scribe Note: Hyonho Lee Department of Computer Science University of Toronto 03 March 2006 Microarrays Revisited In the last
Chapter 17 Lecture. Concepts of Genetics. Tenth Edition. Regulation of Gene Expression in Eukaryotes
 Chapter 17 Lecture Concepts of Genetics Tenth Edition Regulation of Gene Expression in Eukaryotes Chapter Contents 17.1 Eukaryotic Gene Regulation Can Occur at Any of the Steps Leading from DNA to Protein
Chapter 17 Lecture Concepts of Genetics Tenth Edition Regulation of Gene Expression in Eukaryotes Chapter Contents 17.1 Eukaryotic Gene Regulation Can Occur at Any of the Steps Leading from DNA to Protein
BIOINF/BENG/BIMM/CHEM/CSE 184: Computational Molecular Biology. Lecture 2: Microarray analysis
 BIOINF/BENG/BIMM/CHEM/CSE 184: Computational Molecular Biology Lecture 2: Microarray analysis Genome wide measurement of gene transcription using DNA microarray Bruce Alberts, et al., Molecular Biology
BIOINF/BENG/BIMM/CHEM/CSE 184: Computational Molecular Biology Lecture 2: Microarray analysis Genome wide measurement of gene transcription using DNA microarray Bruce Alberts, et al., Molecular Biology
An introduction to the nucpos package
 An introduction to the nucpos package Hiroaki Kato *, Takeshi Urano Department of Biochemistry, Shimane University School of Medicine October 30, 2018 1 About nucpos nucpos, a derivative of NuPoP, is an
An introduction to the nucpos package Hiroaki Kato *, Takeshi Urano Department of Biochemistry, Shimane University School of Medicine October 30, 2018 1 About nucpos nucpos, a derivative of NuPoP, is an
Distinct Modes of Regulation by Chromatin Encoded through Nucleosome Positioning Signals
 Encoded through Nucleosome Positioning Signals Yair Field 1., Noam Kaplan 1., Yvonne Fondufe-Mittendorf 2., Irene K. Moore 2, Eilon Sharon 1, Yaniv Lubling 1, Jonathan Widom 2 *, Eran Segal 1,3 * 1 Department
Encoded through Nucleosome Positioning Signals Yair Field 1., Noam Kaplan 1., Yvonne Fondufe-Mittendorf 2., Irene K. Moore 2, Eilon Sharon 1, Yaniv Lubling 1, Jonathan Widom 2 *, Eran Segal 1,3 * 1 Department
Protein Architecture: Conserved Functional Domains
 PROTOCOL Protein Motif Analysis compiled by John R. Finnerty Protein Architecture: Conserved Functional Domains Proteins are like machines in that different parts of the protein perform different sub-functions,
PROTOCOL Protein Motif Analysis compiled by John R. Finnerty Protein Architecture: Conserved Functional Domains Proteins are like machines in that different parts of the protein perform different sub-functions,
A Statistical Framework for the Analysis of ChIP- Seq Data
 University of Wisconsin-Madison From the SelectedWorks of Sunduz Keles Fall November 19, 2009 A Statistical Framework for the Analysis of ChIP- Seq Data Pei Fen Kuan, University of North Carolina at Chapel
University of Wisconsin-Madison From the SelectedWorks of Sunduz Keles Fall November 19, 2009 A Statistical Framework for the Analysis of ChIP- Seq Data Pei Fen Kuan, University of North Carolina at Chapel
Genome-wide analysis of nucleosome occupancy surrounding Saccharomyces cerevisiae origins of replication
 Genome-wide analysis of nucleosome occupancy surrounding Saccharomyces cerevisiae origins of replication by Nicolas Matthew Berbenetz A thesis submitted in conformity with the requirements for the degree
Genome-wide analysis of nucleosome occupancy surrounding Saccharomyces cerevisiae origins of replication by Nicolas Matthew Berbenetz A thesis submitted in conformity with the requirements for the degree
Lecture 5: Regulation
 Machine Learning in Computational Biology CSC 2431 Lecture 5: Regulation Instructor: Anna Goldenberg Central Dogma of Biology Transcription DNA RNA protein Process of producing RNA from DNA Constitutive
Machine Learning in Computational Biology CSC 2431 Lecture 5: Regulation Instructor: Anna Goldenberg Central Dogma of Biology Transcription DNA RNA protein Process of producing RNA from DNA Constitutive
Perm-seq: Mapping Protein-DNA Interactions in Segmental Duplication and Highly Repetitive Regions of Genomes with Prior- Enhanced Read Mapping
 RESEARCH ARTICLE Perm-seq: Mapping Protein-DNA Interactions in Segmental Duplication and Highly Repetitive Regions of Genomes with Prior- Enhanced Read Mapping Xin Zeng 1,BoLi 2, Rene Welch 1, Constanza
RESEARCH ARTICLE Perm-seq: Mapping Protein-DNA Interactions in Segmental Duplication and Highly Repetitive Regions of Genomes with Prior- Enhanced Read Mapping Xin Zeng 1,BoLi 2, Rene Welch 1, Constanza
BIOINFORMATICS. NORMAL: accurate nucleosome positioning using a modified Gaussian mixture model
 BIOINFORMATICS Vol. 28 ISMB 2012, pages i242 i249 doi:10.1093/bioinformatics/bts206 : accurate nucleosome positioning using a modified Gaussian mixture model Anton Polishko 1,, Nadia Ponts 2,3, Karine
BIOINFORMATICS Vol. 28 ISMB 2012, pages i242 i249 doi:10.1093/bioinformatics/bts206 : accurate nucleosome positioning using a modified Gaussian mixture model Anton Polishko 1,, Nadia Ponts 2,3, Karine
Machine Learning. HMM applications in computational biology
 10-601 Machine Learning HMM applications in computational biology Central dogma DNA CCTGAGCCAACTATTGATGAA transcription mrna CCUGAGCCAACUAUUGAUGAA translation Protein PEPTIDE 2 Biological data is rapidly
10-601 Machine Learning HMM applications in computational biology Central dogma DNA CCTGAGCCAACTATTGATGAA transcription mrna CCUGAGCCAACUAUUGAUGAA translation Protein PEPTIDE 2 Biological data is rapidly
Sequence Analysis. II: Sequence Patterns and Matrices. George Bell, Ph.D. WIBR Bioinformatics and Research Computing
 Sequence Analysis II: Sequence Patterns and Matrices George Bell, Ph.D. WIBR Bioinformatics and Research Computing Sequence Patterns and Matrices Multiple sequence alignments Sequence patterns Sequence
Sequence Analysis II: Sequence Patterns and Matrices George Bell, Ph.D. WIBR Bioinformatics and Research Computing Sequence Patterns and Matrices Multiple sequence alignments Sequence patterns Sequence
ChIP-seq analysis 2/28/2018
 ChIP-seq analysis 2/28/2018 Acknowledgements Much of the content of this lecture is from: Furey (2012) ChIP-seq and beyond Park (2009) ChIP-seq advantages + challenges Landt et al. (2012) ChIP-seq guidelines
ChIP-seq analysis 2/28/2018 Acknowledgements Much of the content of this lecture is from: Furey (2012) ChIP-seq and beyond Park (2009) ChIP-seq advantages + challenges Landt et al. (2012) ChIP-seq guidelines
Lecture 7: April 7, 2005
 Analysis of Gene Expression Data Spring Semester, 2005 Lecture 7: April 7, 2005 Lecturer: R.Shamir and C.Linhart Scribe: A.Mosseri, E.Hirsh and Z.Bronstein 1 7.1 Promoter Analysis 7.1.1 Introduction to
Analysis of Gene Expression Data Spring Semester, 2005 Lecture 7: April 7, 2005 Lecturer: R.Shamir and C.Linhart Scribe: A.Mosseri, E.Hirsh and Z.Bronstein 1 7.1 Promoter Analysis 7.1.1 Introduction to
Supplementary Information for: Using DNA mechanics to predict in vitro nucleosome positioning
 Supplementary Information for: Using DNA mechanics to predict in vitro nucleosome positioning Alexandre V. Morozov, Karissa Fortney, Daria A. Gaykalova, Vasily M. Studitsky, Jonathan Widom, and Eric D.
Supplementary Information for: Using DNA mechanics to predict in vitro nucleosome positioning Alexandre V. Morozov, Karissa Fortney, Daria A. Gaykalova, Vasily M. Studitsky, Jonathan Widom, and Eric D.
Bayesian Variable Selection and Data Integration for Biological Regulatory Networks
 Bayesian Variable Selection and Data Integration for Biological Regulatory Networks Shane T. Jensen Department of Statistics The Wharton School, University of Pennsylvania stjensen@wharton.upenn.edu Gary
Bayesian Variable Selection and Data Integration for Biological Regulatory Networks Shane T. Jensen Department of Statistics The Wharton School, University of Pennsylvania stjensen@wharton.upenn.edu Gary
L8: Downstream analysis of ChIP-seq and ATAC-seq data
 L8: Downstream analysis of ChIP-seq and ATAC-seq data Shamith Samarajiwa CRUK Bioinformatics Autumn School September 2017 Summary Downstream analysis for extracting meaningful biology : Normalization and
L8: Downstream analysis of ChIP-seq and ATAC-seq data Shamith Samarajiwa CRUK Bioinformatics Autumn School September 2017 Summary Downstream analysis for extracting meaningful biology : Normalization and
Recombinant DNA: Basics and Advanced Applications. Code: ECTS Credits: 9. Degree Type Year Semester
 2018/2019 Recombinant DNA: Basics and Advanced Applications Code: 42895 ECTS Credits: 9 Degree Type Year Semester 4313794 Biochemistry, Molecular Biology and Biomedicine OT 0 A Contact Use of languages
2018/2019 Recombinant DNA: Basics and Advanced Applications Code: 42895 ECTS Credits: 9 Degree Type Year Semester 4313794 Biochemistry, Molecular Biology and Biomedicine OT 0 A Contact Use of languages
Measuring Protein-DNA interactions
 Measuring Protein-DNA interactions How is Biological Complexity Achieved? Mediated by Transcription Factors (TFs) 2 Transcription Factors are genetic switches 3 Regulation of Gene Expression by Transcription
Measuring Protein-DNA interactions How is Biological Complexity Achieved? Mediated by Transcription Factors (TFs) 2 Transcription Factors are genetic switches 3 Regulation of Gene Expression by Transcription
ChIP-seq data analysis with Chipster. Eija Korpelainen CSC IT Center for Science, Finland
 ChIP-seq data analysis with Chipster Eija Korpelainen CSC IT Center for Science, Finland chipster@csc.fi What will I learn? Short introduction to ChIP-seq Analyzing ChIP-seq data Central concepts Analysis
ChIP-seq data analysis with Chipster Eija Korpelainen CSC IT Center for Science, Finland chipster@csc.fi What will I learn? Short introduction to ChIP-seq Analyzing ChIP-seq data Central concepts Analysis
Huijuan Feng, Shining Ma,Chao Ye & Zhixing Feng
 Huijuan Feng, Shining Ma,Chao Ye & Zhixing Feng Background-Author introduction Research interest: Methods for gene mapping of complex traits Inference of population structure from genetic data Genome variation
Huijuan Feng, Shining Ma,Chao Ye & Zhixing Feng Background-Author introduction Research interest: Methods for gene mapping of complex traits Inference of population structure from genetic data Genome variation
Examination Assignments
 Bioinformatics Institute of India H-109, Ground Floor, Sector-63, Noida-201307, UP. INDIA Tel.: 0120-4320801 / 02, M. 09818473366, 09810535368 Email: info@bii.in, Website: www.bii.in INDUSTRY PROGRAM IN
Bioinformatics Institute of India H-109, Ground Floor, Sector-63, Noida-201307, UP. INDIA Tel.: 0120-4320801 / 02, M. 09818473366, 09810535368 Email: info@bii.in, Website: www.bii.in INDUSTRY PROGRAM IN
File S1. Program overview and features
 File S1 Program overview and features Query list filtering. Further filtering may be applied through user selected query lists (Figure. 2B, Table S3) that restrict the results and/or report specifically
File S1 Program overview and features Query list filtering. Further filtering may be applied through user selected query lists (Figure. 2B, Table S3) that restrict the results and/or report specifically
Computational Analysis of Ultra-high-throughput sequencing data: ChIP-Seq
 Computational Analysis of Ultra-high-throughput sequencing data: ChIP-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne Data flow in ChIP-Seq data analysis Level 1:
Computational Analysis of Ultra-high-throughput sequencing data: ChIP-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne Data flow in ChIP-Seq data analysis Level 1:
Transcription factor binding site prediction in vivo using DNA sequence and shape features
 Transcription factor binding site prediction in vivo using DNA sequence and shape features Anthony Mathelier, Lin Yang, Tsu-Pei Chiu, Remo Rohs, and Wyeth Wasserman anthony.mathelier@gmail.com @AMathelier
Transcription factor binding site prediction in vivo using DNA sequence and shape features Anthony Mathelier, Lin Yang, Tsu-Pei Chiu, Remo Rohs, and Wyeth Wasserman anthony.mathelier@gmail.com @AMathelier
Nucleosomes: what, why and where? Rob Brewster
 Nucleosomes: what, why and where? Rob Brewster What is a nucleosome? Outline how is DNA packaged/organized in Eukaryotes? Why do nucleosomes form? DNA is seff, how do ~100 bp loops form so readily? Where
Nucleosomes: what, why and where? Rob Brewster What is a nucleosome? Outline how is DNA packaged/organized in Eukaryotes? Why do nucleosomes form? DNA is seff, how do ~100 bp loops form so readily? Where
Chromatin immunoprecipitation: five steps to great results
 Chromatin immunoprecipitation: five steps to great results Introduction The discovery and use of antibodies in life science research has been critical to many advancements across applications, including
Chromatin immunoprecipitation: five steps to great results Introduction The discovery and use of antibodies in life science research has been critical to many advancements across applications, including
Decoding Chromatin States with Epigenome Data Advanced Topics in Computa8onal Genomics
 Decoding Chromatin States with Epigenome Data 02-715 Advanced Topics in Computa8onal Genomics HMMs for Decoding Chromatin States Epigene8c modifica8ons of the genome have been associated with Establishing
Decoding Chromatin States with Epigenome Data 02-715 Advanced Topics in Computa8onal Genomics HMMs for Decoding Chromatin States Epigene8c modifica8ons of the genome have been associated with Establishing
QTL mapping in mice. Karl W Broman. Department of Biostatistics Johns Hopkins University Baltimore, Maryland, USA.
 QTL mapping in mice Karl W Broman Department of Biostatistics Johns Hopkins University Baltimore, Maryland, USA www.biostat.jhsph.edu/ kbroman Outline Experiments, data, and goals Models ANOVA at marker
QTL mapping in mice Karl W Broman Department of Biostatistics Johns Hopkins University Baltimore, Maryland, USA www.biostat.jhsph.edu/ kbroman Outline Experiments, data, and goals Models ANOVA at marker
Applications of HMMs in Epigenomics
 I529: Machine Learning in Bioinformatics (Spring 2013) Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Contents Background:
I529: Machine Learning in Bioinformatics (Spring 2013) Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Contents Background:
The Next Generation of Transcription Factor Binding Site Prediction
 The Next Generation of Transcription Factor Binding Site Prediction Anthony Mathelier*, Wyeth W. Wasserman* Centre for Molecular Medicine and Therapeutics at the Child and Family Research Institute, Department
The Next Generation of Transcription Factor Binding Site Prediction Anthony Mathelier*, Wyeth W. Wasserman* Centre for Molecular Medicine and Therapeutics at the Child and Family Research Institute, Department
2/19/13. Contents. Applications of HMMs in Epigenomics
 2/19/13 I529: Machine Learning in Bioinformatics (Spring 2013) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Background:
2/19/13 I529: Machine Learning in Bioinformatics (Spring 2013) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Background:
Dynamic Remodeling of Individual Nucleosomes Across a Eukaryotic Genome in Response to Transcriptional Perturbation
 Dynamic Remodeling of Individual Nucleosomes Across a Eukaryotic Genome in Response to Transcriptional Perturbation Sushma Shivaswamy 1[, Akshay Bhinge 1[, Yongjun Zhao 2, Steven Jones 2, Martin Hirst
Dynamic Remodeling of Individual Nucleosomes Across a Eukaryotic Genome in Response to Transcriptional Perturbation Sushma Shivaswamy 1[, Akshay Bhinge 1[, Yongjun Zhao 2, Steven Jones 2, Martin Hirst
Permutation Clustering of the DNA Sequence Facilitates Understanding of the Nonlinearly Organized Genome
 RESEARCH PROPOSAL Permutation Clustering of the DNA Sequence Facilitates Understanding of the Nonlinearly Organized Genome Qiao JIN School of Medicine, Tsinghua University Advisor: Prof. Xuegong ZHANG
RESEARCH PROPOSAL Permutation Clustering of the DNA Sequence Facilitates Understanding of the Nonlinearly Organized Genome Qiao JIN School of Medicine, Tsinghua University Advisor: Prof. Xuegong ZHANG
Mixture modeling for genome-wide localization of transcription factors
 Mixture modeling for genome-wide localization of transcription factors Sündüz Keleş Department of Statistics and Department of Biostatistics & Medical Informatics 1300 University Avenue, 1245B Medical
Mixture modeling for genome-wide localization of transcription factors Sündüz Keleş Department of Statistics and Department of Biostatistics & Medical Informatics 1300 University Avenue, 1245B Medical
Learning a Weighted Sequence Model of the Nucleosome Core and Linker Yields More Accurate Predictions in Saccharomyces cerevisiae and Homo sapiens
 Learning a Weighted Sequence Model of the Nucleosome Core and Linker Yields More Accurate Predictions in Saccharomyces cerevisiae and Homo sapiens Sheila M. Reynolds 1, Jeff A. Bilmes 1,2, William Stafford
Learning a Weighted Sequence Model of the Nucleosome Core and Linker Yields More Accurate Predictions in Saccharomyces cerevisiae and Homo sapiens Sheila M. Reynolds 1, Jeff A. Bilmes 1,2, William Stafford
Recombinant DNA: Basics and Advanced Applications
 Recombinant DNA: Basics and Advanced Applications 2015/2016 Code: 42895 ECTS Credits: 9 Degree Type Year Semester 4313794 Biochemistry, Molecular Biology and Biomedicine OT 0 A Contact Name: Antonio Casamayor
Recombinant DNA: Basics and Advanced Applications 2015/2016 Code: 42895 ECTS Credits: 9 Degree Type Year Semester 4313794 Biochemistry, Molecular Biology and Biomedicine OT 0 A Contact Name: Antonio Casamayor
ChIP-seq/Functional Genomics/Epigenomics. CBSU/3CPG/CVG Next-Gen Sequencing Workshop. Josh Waterfall. March 31, 2010
 ChIP-seq/Functional Genomics/Epigenomics CBSU/3CPG/CVG Next-Gen Sequencing Workshop Josh Waterfall March 31, 2010 Outline Introduction to ChIP-seq Control data sets Peak/enriched region identification
ChIP-seq/Functional Genomics/Epigenomics CBSU/3CPG/CVG Next-Gen Sequencing Workshop Josh Waterfall March 31, 2010 Outline Introduction to ChIP-seq Control data sets Peak/enriched region identification
Nucleosome Positioning and Organization Advanced Topics in Computa8onal Genomics
 Nucleosome Positioning and Organization 02-715 Advanced Topics in Computa8onal Genomics Nucleosome Core Nucleosome Core and Linker 147 bp DNA wrapping around nucleosome core Varying lengths of linkers
Nucleosome Positioning and Organization 02-715 Advanced Topics in Computa8onal Genomics Nucleosome Core Nucleosome Core and Linker 147 bp DNA wrapping around nucleosome core Varying lengths of linkers
Exploring Similarities of Conserved Domains/Motifs
 Exploring Similarities of Conserved Domains/Motifs Sotiria Palioura Abstract Traditionally, proteins are represented as amino acid sequences. There are, though, other (potentially more exciting) representations;
Exploring Similarities of Conserved Domains/Motifs Sotiria Palioura Abstract Traditionally, proteins are represented as amino acid sequences. There are, though, other (potentially more exciting) representations;
QTL mapping in mice. Karl W Broman. Department of Biostatistics Johns Hopkins University Baltimore, Maryland, USA.
 QTL mapping in mice Karl W Broman Department of Biostatistics Johns Hopkins University Baltimore, Maryland, USA www.biostat.jhsph.edu/ kbroman Outline Experiments, data, and goals Models ANOVA at marker
QTL mapping in mice Karl W Broman Department of Biostatistics Johns Hopkins University Baltimore, Maryland, USA www.biostat.jhsph.edu/ kbroman Outline Experiments, data, and goals Models ANOVA at marker
ECS 234: Genomic Data Integration ECS 234
 : Genomic Data Integration Heterogeneous Data Integration DNA Sequence Microarray Proteomics >gi 12004594 gb AF217406.1 Saccharomyces cerevisiae uridine nucleosidase (URH1) gene, complete cds ATGGAATCTGCTGATTTTTTTACCTCACGAAACTTATTAAAACAGATAATTTCCCTCATCTGCAAGGTTG
: Genomic Data Integration Heterogeneous Data Integration DNA Sequence Microarray Proteomics >gi 12004594 gb AF217406.1 Saccharomyces cerevisiae uridine nucleosidase (URH1) gene, complete cds ATGGAATCTGCTGATTTTTTTACCTCACGAAACTTATTAAAACAGATAATTTCCCTCATCTGCAAGGTTG
Nucleosome Positioning in Saccharomyces cerevisiae
 MICROBIOLOGY AND MOLECULAR BIOLOGY REVIEWS, June 2011, p. 301 320 Vol. 75, No. 2 1092-2172/11/$12.00 doi:10.1128/mmbr.00046-10 Copyright 2011, American Society for Microbiology. All Rights Reserved. Nucleosome
MICROBIOLOGY AND MOLECULAR BIOLOGY REVIEWS, June 2011, p. 301 320 Vol. 75, No. 2 1092-2172/11/$12.00 doi:10.1128/mmbr.00046-10 Copyright 2011, American Society for Microbiology. All Rights Reserved. Nucleosome
Standardized collection of MNase-seq experiments enables unbiased dataset comparisons
 Rizzo et al. BMC Molecular Biology 212, 13:15 METHODOLOGY ARTICLE Open Access Standardized collection of MNase-seq experiments enables unbiased dataset comparisons Jason M Rizzo 1,2, Jonathan E Bard 2
Rizzo et al. BMC Molecular Biology 212, 13:15 METHODOLOGY ARTICLE Open Access Standardized collection of MNase-seq experiments enables unbiased dataset comparisons Jason M Rizzo 1,2, Jonathan E Bard 2
Array Informatics. Mark Gerstein
 1 Lectures.GersteinLab.org (c) Array Informatics Mark Gerstein CEGS Informatics Developing Tools and Technical Analyses Related to Genome Technologies Main Genome Technologies Tiling Arrays Next Generation
1 Lectures.GersteinLab.org (c) Array Informatics Mark Gerstein CEGS Informatics Developing Tools and Technical Analyses Related to Genome Technologies Main Genome Technologies Tiling Arrays Next Generation
vs. Nucleosomal Pattern Informal presentation of the problem A. Bolshoy 1
 vs. Nucleosomal Pattern Informal presentation of the problem A. Bolshoy 1 What is this about? The following few slides should help me to explain my reflections on extraction of nucleosome positioning pattern;
vs. Nucleosomal Pattern Informal presentation of the problem A. Bolshoy 1 What is this about? The following few slides should help me to explain my reflections on extraction of nucleosome positioning pattern;
Package qcqpcr. February 8, 2018
 Type Package Title Version 1.5 Date 2018-02-02 Author Package February 8, 2018 Maintainer Quality control of chromatin immunoprecipitation libraries (ChIP-seq) by quantitative polymerase
Type Package Title Version 1.5 Date 2018-02-02 Author Package February 8, 2018 Maintainer Quality control of chromatin immunoprecipitation libraries (ChIP-seq) by quantitative polymerase
DIAMANTINA INSTITUTE for Cancer, Immunology and Metabolic Medicine
 DIAMANTINA INSTITUTE for Cancer, Immunology and Metabolic Medicine Defining MYB Transcriptional Network by Genome-wide Chromatin Occupancy Profiling (ChIP-Seq) 2010 E.Glazov, L. Zhao Transcription Factors:
DIAMANTINA INSTITUTE for Cancer, Immunology and Metabolic Medicine Defining MYB Transcriptional Network by Genome-wide Chromatin Occupancy Profiling (ChIP-Seq) 2010 E.Glazov, L. Zhao Transcription Factors:
A genomic code for nucleosome positioning
 doi:10.1038/nature04979 A genomic code for nucleosome positioning Eran Segal 1, Yvonne Fondufe-Mittendorf 2, Lingyi Chen 2, AnnChristine Thåström 2, Yair Field 1, Irene K. Moore 2, Ji-Ping Z. Wang 3 &
doi:10.1038/nature04979 A genomic code for nucleosome positioning Eran Segal 1, Yvonne Fondufe-Mittendorf 2, Lingyi Chen 2, AnnChristine Thåström 2, Yair Field 1, Irene K. Moore 2, Ji-Ping Z. Wang 3 &
Recombinant DNA: Basics and Advanced Applications
 Recombinant DNA: Basics and Advanced Applications 2014/2015 Code: 42895 ECTS Credits: 9 Degree Type Year Semester 4313794 Bioquímica, Biologia Molecular i Biomedicina OT 0 A Contact Name: Antonio Casamayor
Recombinant DNA: Basics and Advanced Applications 2014/2015 Code: 42895 ECTS Credits: 9 Degree Type Year Semester 4313794 Bioquímica, Biologia Molecular i Biomedicina OT 0 A Contact Name: Antonio Casamayor
Bioinformatics. Microarrays: designing chips, clustering methods. Fran Lewitter, Ph.D. Head, Biocomputing Whitehead Institute
 Bioinformatics Microarrays: designing chips, clustering methods Fran Lewitter, Ph.D. Head, Biocomputing Whitehead Institute Course Syllabus Jan 7 Jan 14 Jan 21 Jan 28 Feb 4 Feb 11 Feb 18 Feb 25 Sequence
Bioinformatics Microarrays: designing chips, clustering methods Fran Lewitter, Ph.D. Head, Biocomputing Whitehead Institute Course Syllabus Jan 7 Jan 14 Jan 21 Jan 28 Feb 4 Feb 11 Feb 18 Feb 25 Sequence
Methods and tools for exploring functional genomics data
 Methods and tools for exploring functional genomics data William Stafford Noble Department of Genome Sciences Department of Computer Science and Engineering University of Washington Outline Searching for
Methods and tools for exploring functional genomics data William Stafford Noble Department of Genome Sciences Department of Computer Science and Engineering University of Washington Outline Searching for
Figure S1. nuclear extracts. HeLa cell nuclear extract. Input IgG IP:ORC2 ORC2 ORC2. MCM4 origin. ORC2 occupancy
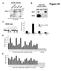 A nuclear extracts B HeLa cell nuclear extract Figure S1 ORC2 (in kda) 21 132 7 ORC2 Input IgG IP:ORC2 32 ORC C D PRKDC ORC2 occupancy Directed against ORC2 C-terminus (sc-272) MCM origin 2 2 1-1 -1kb
A nuclear extracts B HeLa cell nuclear extract Figure S1 ORC2 (in kda) 21 132 7 ORC2 Input IgG IP:ORC2 32 ORC C D PRKDC ORC2 occupancy Directed against ORC2 C-terminus (sc-272) MCM origin 2 2 1-1 -1kb
AC Algorithms for Mining Biological Sequences (COMP 680)
 AC-04-18 Algorithms for Mining Biological Sequences (COMP 680) Instructor: Mathieu Blanchette School of Computer Science and McGill Centre for Bioinformatics, 332 Duff Building McGill University, Montreal,
AC-04-18 Algorithms for Mining Biological Sequences (COMP 680) Instructor: Mathieu Blanchette School of Computer Science and McGill Centre for Bioinformatics, 332 Duff Building McGill University, Montreal,
Computational Genomics
 Computational Genomics http://www.cs.cmu.edu/~02710 Ziv Bar-Joseph zivbj@cs.cmu.edu GHC 8006 Chakra Chennubhotla chakracs@pitt.edu Suite 3064, BST3 Topics Introduction (1 Week) Sequence analysis(4 weeks)
Computational Genomics http://www.cs.cmu.edu/~02710 Ziv Bar-Joseph zivbj@cs.cmu.edu GHC 8006 Chakra Chennubhotla chakracs@pitt.edu Suite 3064, BST3 Topics Introduction (1 Week) Sequence analysis(4 weeks)
A holistic perspective of gene expression during the E. coli growth cycle. DNA structure and nucleosome placement
 A holistic perspective of gene expression during the E. coli growth cycle. Andrew Travers DNA structure and nucleosome placement Haifa, 13/05/12 1 A/T rich base-steps are more deformable than G/C-rich
A holistic perspective of gene expression during the E. coli growth cycle. Andrew Travers DNA structure and nucleosome placement Haifa, 13/05/12 1 A/T rich base-steps are more deformable than G/C-rich
Measuring gene expression
 Measuring gene expression Grundlagen der Bioinformatik SS2018 https://www.youtube.com/watch?v=v8gh404a3gg Agenda Organization Gene expression Background Technologies FISH Nanostring Microarrays RNA-seq
Measuring gene expression Grundlagen der Bioinformatik SS2018 https://www.youtube.com/watch?v=v8gh404a3gg Agenda Organization Gene expression Background Technologies FISH Nanostring Microarrays RNA-seq
University of California, Berkeley
 University of California, Berkeley U.C. Berkeley Division of Biostatistics Working Paper Series Year 2004 Paper 145 Regulatory Motif Finding by Logic Regression Sunduz Keles Mark J. van der Laan Chris
University of California, Berkeley U.C. Berkeley Division of Biostatistics Working Paper Series Year 2004 Paper 145 Regulatory Motif Finding by Logic Regression Sunduz Keles Mark J. van der Laan Chris
Understanding transcriptional regulation by integrative analysis of transcription factor binding data
 Understanding transcriptional regulation by integrative analysis of transcription factor binding data Cheng et al. 2012 Shu Yang Feb. 21, 2013 1 / 26 Introduction 2 / 26 DNA-binding Proteins sequence-specific
Understanding transcriptional regulation by integrative analysis of transcription factor binding data Cheng et al. 2012 Shu Yang Feb. 21, 2013 1 / 26 Introduction 2 / 26 DNA-binding Proteins sequence-specific
Discovering gene regulatory control using ChIP-chip and ChIP-seq. Part 1. An introduction to gene regulatory control, concepts and methodologies
 Discovering gene regulatory control using ChIP-chip and ChIP-seq Part 1 An introduction to gene regulatory control, concepts and methodologies Ian Simpson ian.simpson@.ed.ac.uk http://bit.ly/bio2links
Discovering gene regulatory control using ChIP-chip and ChIP-seq Part 1 An introduction to gene regulatory control, concepts and methodologies Ian Simpson ian.simpson@.ed.ac.uk http://bit.ly/bio2links
Motif Discovery from Large Number of Sequences: a Case Study with Disease Resistance Genes in Arabidopsis thaliana
 Motif Discovery from Large Number of Sequences: a Case Study with Disease Resistance Genes in Arabidopsis thaliana Irfan Gunduz, Sihui Zhao, Mehmet Dalkilic and Sun Kim Indiana University, School of Informatics
Motif Discovery from Large Number of Sequences: a Case Study with Disease Resistance Genes in Arabidopsis thaliana Irfan Gunduz, Sihui Zhao, Mehmet Dalkilic and Sun Kim Indiana University, School of Informatics
Figure 7.1: PWM evolution: The sequence affinity of TFBSs has evolved from single sequences, to PWMs, to larger and larger databases of PWMs.
 Chapter 7 Discussion This thesis presents dry and wet lab techniques to elucidate the involvement of transcription factors (TFs) in the regulation of the cell cycle and myogenesis. However, the techniques
Chapter 7 Discussion This thesis presents dry and wet lab techniques to elucidate the involvement of transcription factors (TFs) in the regulation of the cell cycle and myogenesis. However, the techniques
Genome-Scale Predictions of the Transcription Factor Binding Sites of Cys 2 His 2 Zinc Finger Proteins in Yeast June 17 th, 2005
 Genome-Scale Predictions of the Transcription Factor Binding Sites of Cys 2 His 2 Zinc Finger Proteins in Yeast June 17 th, 2005 John Brothers II 1,3 and Panayiotis V. Benos 1,2 1 Bioengineering and Bioinformatics
Genome-Scale Predictions of the Transcription Factor Binding Sites of Cys 2 His 2 Zinc Finger Proteins in Yeast June 17 th, 2005 John Brothers II 1,3 and Panayiotis V. Benos 1,2 1 Bioengineering and Bioinformatics
CollecTF Documentation
 CollecTF Documentation Release 1.0.0 Sefa Kilic August 15, 2016 Contents 1 Curation submission guide 3 1.1 Data.................................................... 3 1.2 Before you start.............................................
CollecTF Documentation Release 1.0.0 Sefa Kilic August 15, 2016 Contents 1 Curation submission guide 3 1.1 Data.................................................... 3 1.2 Before you start.............................................
Biology 644: Bioinformatics
 Processes Activation Repression Initiation Elongation.... Processes Splicing Editing Degradation Translation.... Transcription Translation DNA Regulators DNA-Binding Transcription Factors Chromatin Remodelers....
Processes Activation Repression Initiation Elongation.... Processes Splicing Editing Degradation Translation.... Transcription Translation DNA Regulators DNA-Binding Transcription Factors Chromatin Remodelers....
Galaxy Platform For NGS Data Analyses
 Galaxy Platform For NGS Data Analyses Weihong Yan wyan@chem.ucla.edu Collaboratory Web Site http://qcb.ucla.edu/collaboratory http://collaboratory.lifesci.ucla.edu Workshop Outline ü Day 1 UCLA galaxy
Galaxy Platform For NGS Data Analyses Weihong Yan wyan@chem.ucla.edu Collaboratory Web Site http://qcb.ucla.edu/collaboratory http://collaboratory.lifesci.ucla.edu Workshop Outline ü Day 1 UCLA galaxy
Predicting Transcription Factors Binding Sites With Deep Learning
 Predicting Transcription Factors Binding Sites With Deep Learning Yurii Toma (yurii.toma@ut.ee) Kaur Alasoo (kaur.alasoo@gmail.com) Dmytro Fishman (dmytrofishman@gmail.com) What is the central dogma of
Predicting Transcription Factors Binding Sites With Deep Learning Yurii Toma (yurii.toma@ut.ee) Kaur Alasoo (kaur.alasoo@gmail.com) Dmytro Fishman (dmytrofishman@gmail.com) What is the central dogma of
A New Algorithm for Protein-Protein Interaction Prediction
 A New Algorithm for Protein-Protein Interaction Prediction Yiwei Li The University of Western Ontario Supervisor: Dr. Lucian Ilie Overview 1 Background 2 Previous approaches Experimental approaches Computational
A New Algorithm for Protein-Protein Interaction Prediction Yiwei Li The University of Western Ontario Supervisor: Dr. Lucian Ilie Overview 1 Background 2 Previous approaches Experimental approaches Computational
Chapter 19 Genetic Regulation of the Eukaryotic Genome. A. Bergeron AP Biology PCHS
 Chapter 19 Genetic Regulation of the Eukaryotic Genome A. Bergeron AP Biology PCHS 2 Do Now - Eukaryotic Transcription Regulation The diagram below shows five genes (with their enhancers) from the genome
Chapter 19 Genetic Regulation of the Eukaryotic Genome A. Bergeron AP Biology PCHS 2 Do Now - Eukaryotic Transcription Regulation The diagram below shows five genes (with their enhancers) from the genome
Next-generation sequencing technologies
 Next-generation sequencing technologies Illumina: Summary https://www.youtube.com/watch?v=fcd6b5hraz8 Illumina platforms: Benchtop sequencers https://www.illumina.com/systems/sequencing-platforms.html
Next-generation sequencing technologies Illumina: Summary https://www.youtube.com/watch?v=fcd6b5hraz8 Illumina platforms: Benchtop sequencers https://www.illumina.com/systems/sequencing-platforms.html
A Novel Approaches For Motif Discovery Using Data Mining Algorithm
 A Novel Approaches For Motif Discovery Using Data Mining Algorithm M Mohamed Divan Masood 1, D Manjula 2 1 Research Scholar, Computer Science and Engineering, Anna University, Chennai. 2Head of the Department,
A Novel Approaches For Motif Discovery Using Data Mining Algorithm M Mohamed Divan Masood 1, D Manjula 2 1 Research Scholar, Computer Science and Engineering, Anna University, Chennai. 2Head of the Department,
A Supervised Learning Approach to the Prediction of Hi-C Data
 A Supervised Learning Approach to the Prediction of Hi-C Data Tyler Derr YUE Lab The Department of Biochemistry & Molecular Biology Institute for Personalized Medicine Pennsylvania State University College
A Supervised Learning Approach to the Prediction of Hi-C Data Tyler Derr YUE Lab The Department of Biochemistry & Molecular Biology Institute for Personalized Medicine Pennsylvania State University College
Bio5488 Practice Midterm (2018) 1. Next-gen sequencing
 1. Next-gen sequencing 1. You have found a new strain of yeast that makes fantastic wine. You d like to sequence this strain to ascertain the differences from S. cerevisiae. To accurately call a base pair,
1. Next-gen sequencing 1. You have found a new strain of yeast that makes fantastic wine. You d like to sequence this strain to ascertain the differences from S. cerevisiae. To accurately call a base pair,
ChIP-seq and RNA-seq. Farhat Habib
 ChIP-seq and RNA-seq Farhat Habib fhabib@iiserpune.ac.in Biological Goals Learn how genomes encode the diverse patterns of gene expression that define each cell type and state. Protein-DNA interactions
ChIP-seq and RNA-seq Farhat Habib fhabib@iiserpune.ac.in Biological Goals Learn how genomes encode the diverse patterns of gene expression that define each cell type and state. Protein-DNA interactions
