Package qcqpcr. February 8, 2018
|
|
|
- Aubrie Cooper
- 6 years ago
- Views:
Transcription
1 Type Package Title Version 1.5 Date Author Package February 8, 2018 Maintainer Quality control of chromatin immunoprecipitation libraries (ChIP-seq) by quantitative polymerase chain reaction (qpcr). This function calculates Enrichment value with respect to reference for each histone modification (specific to 'Vii7' software < This function is applicable to full panel of histone modifications described by International Human Epigenomic Consortium (IHEC). Imports ggplot2 License MIT + file LICENSE Depends R (>= 2.10) NeedsCompilation no Repository CRAN Date/Publication :44:50 UTC R topics documented: -package Index 5 1
2 2 -package -package Details Quality control of chromatin immunoprecipitation libraries (ChIP-seq) by quantitative polymerase chain reaction (qpcr). This function calculates Enrichment value with respect to reference for each histone modification (specific to Vii7 software < This function is applicable to full panel of histone modifications described by International Human Epigenomic Consortium (IHEC). Quality control of chromatin immunoprecipitation libraries (ChIP-seq) by quantitative polymerase chain reaction (qpcr). This function calculates Enrichment value with respect to reference for each histone modification (specific to Vii7 software < 7-real-time-pcr-system/viia-7-software.html>). This function is applicable to full panel of histone modifications described by International Human Epigenomic Consortium (IHEC). The DESCRIPTION file: Package: Type: Package Title: Version: 1.5 Date: Author: Maintainer: : Quality control of chromatin immunoprecipitation libraries (ChIP-seq) by quantitative polymerase chain Imports: ggplot2 License: MIT + file LICENSE Depends: R (>= 2.10) NeedsCompilation: no Packaged: :24:48 UTC; epigenomics_lab_01 Index of help topics: -package qpcr ChIP-seq Results Histone ChIP-Seq qpcr Analyzer (xx, Title for you graph ) Author(s)
3 3 Examples library(ggplot2) data() (,'Title for your graph') Quality control of chromatin immunoprecipitation libraries (ChIP-seq) by quantitative polymerase chain reaction (qpcr). This function calculates Enrichment value with respect to reference for each histone modification (specific to Vii7 software < This function is applicable to full panel of histone modifications described by International Human Epigenomic Consortium (IHEC). Usage (xx, Name) Arguments xx Name Your Data Set... Title for you graph, (ie: Projects name and date) Author(s) Examples library(ggplot2) data() (,'Title for your graph')
4 4 qpcr ChIP-seq Results A dataset containing the the CT values of ChIPseq libraries. Usage Format A data frame with 82 rows and 11 variables Source Hirst Lab, UBC Vancouver, Canada
5 Index Topic datasets, 4 Topic qpcr, 3 -package, 2, 3 -package, 2, 4 5
Package hhi. April 13, 2018
 Type Package Package hhi April 13, 2018 Title Calculate and Visualize the Herfindahl-Hirschman Index Version 1.1.0 Author Philip D. Waggoner Maintainer Philip D. Waggoner
Type Package Package hhi April 13, 2018 Title Calculate and Visualize the Herfindahl-Hirschman Index Version 1.1.0 Author Philip D. Waggoner Maintainer Philip D. Waggoner
Package ChannelAttribution
 Type Package Package ChannelAttribution February 5, 2018 Title Markov Model for the Online Multi-Channel Attribution Problem Version 1.12 Date 2018-02-05 Author Davide Altomare, David Loris Maintainer
Type Package Package ChannelAttribution February 5, 2018 Title Markov Model for the Online Multi-Channel Attribution Problem Version 1.12 Date 2018-02-05 Author Davide Altomare, David Loris Maintainer
Data and Metadata Models Recommendations Version 1.2 Developed by the IHEC Metadata Standards Workgroup
 Data and Metadata Models Recommendations Version 1.2 Developed by the IHEC Metadata Standards Workgroup 1. Introduction The data produced by IHEC is illustrated in Figure 1. Figure 1. The space of epigenomic
Data and Metadata Models Recommendations Version 1.2 Developed by the IHEC Metadata Standards Workgroup 1. Introduction The data produced by IHEC is illustrated in Figure 1. Figure 1. The space of epigenomic
Package subgroup. February 20, 2015
 Type Package Package subgroup February 20, 2015 Title Methods for exploring treatment effect heterogeneity in subgroup analysis of clinical trials Version 1.1 Date 2014-07-31 Author I. Manjula Schou Maintainer
Type Package Package subgroup February 20, 2015 Title Methods for exploring treatment effect heterogeneity in subgroup analysis of clinical trials Version 1.1 Date 2014-07-31 Author I. Manjula Schou Maintainer
Package MetNorm. February 19, 2015
 Type Package Package MetNorm February 19, 2015 Title Statistical Methods for Normalizing Metabolomics Data Version 0.1 Date 2015-02-08 Author Alysha M De Livera Maintainer Alysha M De Livera
Type Package Package MetNorm February 19, 2015 Title Statistical Methods for Normalizing Metabolomics Data Version 0.1 Date 2015-02-08 Author Alysha M De Livera Maintainer Alysha M De Livera
Package revengc. August 18, Type Package
 Type Package Package revengc August 18, 2017 Title Reverse Engineering Censored, Decoupled Residential Data for Population Density Estimation Version 1.0.0 Author Samantha Duchscherer [aut, cre], UT-Battelle,
Type Package Package revengc August 18, 2017 Title Reverse Engineering Censored, Decoupled Residential Data for Population Density Estimation Version 1.0.0 Author Samantha Duchscherer [aut, cre], UT-Battelle,
Package geno2proteo. December 12, 2017
 Type Package Package geno2proteo December 12, 2017 Title Finding the DNA and Protein Sequences of Any Genomic or Proteomic Loci Version 0.0.1 Date 2017-12-12 Author Maintainer biocviews
Type Package Package geno2proteo December 12, 2017 Title Finding the DNA and Protein Sequences of Any Genomic or Proteomic Loci Version 0.0.1 Date 2017-12-12 Author Maintainer biocviews
The ENCODE Encyclopedia. & Variant Annotation Using RegulomeDB and HaploReg
 The ENCODE Encyclopedia & Variant Annotation Using RegulomeDB and HaploReg Jill E. Moore Weng Lab University of Massachusetts Medical School October 10, 2015 Where s the Encyclopedia? ENCODE: Encyclopedia
The ENCODE Encyclopedia & Variant Annotation Using RegulomeDB and HaploReg Jill E. Moore Weng Lab University of Massachusetts Medical School October 10, 2015 Where s the Encyclopedia? ENCODE: Encyclopedia
Package pairedci. February 20, 2015
 Type Package Package pairedci February 20, 2015 Title Confidence intervals for the ratio of locations and for the ratio of scales of two paired samples Version 0.5-4 Date 2012-11-24 Author Cornelia Froemke,
Type Package Package pairedci February 20, 2015 Title Confidence intervals for the ratio of locations and for the ratio of scales of two paired samples Version 0.5-4 Date 2012-11-24 Author Cornelia Froemke,
Decoding Chromatin States with Epigenome Data Advanced Topics in Computa8onal Genomics
 Decoding Chromatin States with Epigenome Data 02-715 Advanced Topics in Computa8onal Genomics HMMs for Decoding Chromatin States Epigene8c modifica8ons of the genome have been associated with Establishing
Decoding Chromatin States with Epigenome Data 02-715 Advanced Topics in Computa8onal Genomics HMMs for Decoding Chromatin States Epigene8c modifica8ons of the genome have been associated with Establishing
Green Center Computational Core ChIP- Seq Pipeline, Just a Click Away
 Green Center Computational Core ChIP- Seq Pipeline, Just a Click Away Venkat Malladi Computational Biologist Computational Core Cecil H. and Ida Green Center for Reproductive Biology Science Introduc
Green Center Computational Core ChIP- Seq Pipeline, Just a Click Away Venkat Malladi Computational Biologist Computational Core Cecil H. and Ida Green Center for Reproductive Biology Science Introduc
Package stepwise. R topics documented: February 20, 2015
 Package stepwise February 20, 2015 Title Stepwise detection of recombination breakpoints Version 0.3 Author Jinko Graham , Brad McNeney , Francoise Seillier-Moiseiwitsch
Package stepwise February 20, 2015 Title Stepwise detection of recombination breakpoints Version 0.3 Author Jinko Graham , Brad McNeney , Francoise Seillier-Moiseiwitsch
Package ABC.RAP. October 20, 2016
 Title Array Based CpG Region Analysis Pipeline Version 0.9.0 Package ABC.RAP October 20, 2016 Author Abdulmonem Alsaleh [cre, aut], Robert Weeks [aut], Ian Morison [aut], RStudio [ctb] Maintainer Abdulmonem
Title Array Based CpG Region Analysis Pipeline Version 0.9.0 Package ABC.RAP October 20, 2016 Author Abdulmonem Alsaleh [cre, aut], Robert Weeks [aut], Ian Morison [aut], RStudio [ctb] Maintainer Abdulmonem
Introduction to ChIP Seq data analyses. Acknowledgement: slides taken from Dr. H
 Introduction to ChIP Seq data analyses Acknowledgement: slides taken from Dr. H Wu @Emory ChIP seq: Chromatin ImmunoPrecipitation it ti + sequencing Same biological motivation as ChIP chip: measure specific
Introduction to ChIP Seq data analyses Acknowledgement: slides taken from Dr. H Wu @Emory ChIP seq: Chromatin ImmunoPrecipitation it ti + sequencing Same biological motivation as ChIP chip: measure specific
Ensembl Funcgen: A Database and API for Epigenomics and Gene Regulation Data.
 Ensembl Funcgen: A Database and API for Epigenomics and Gene Regulation Data. Nathan Johnson Ensembl Regulation EBI is an Outstation of the European Molecular Biology Laboratory.! Workshop Overview http://www.ebi.ac.uk/~njohnson/courses/23.05.2013-
Ensembl Funcgen: A Database and API for Epigenomics and Gene Regulation Data. Nathan Johnson Ensembl Regulation EBI is an Outstation of the European Molecular Biology Laboratory.! Workshop Overview http://www.ebi.ac.uk/~njohnson/courses/23.05.2013-
Package SLqPCR. R topics documented: May 9, Type Package. Title Functions for analysis of real-time quantitative PCR data at SIRS-Lab GmbH
 Package SLqPCR May 9, 2018 Type Package Title Functions for analysis of real-time quantitative PCR data at SIRS-Lab GmbH Version 1.46.0 Date 2007-19-04 Author Matthias Kohl Maintainer Matthias Kohl
Package SLqPCR May 9, 2018 Type Package Title Functions for analysis of real-time quantitative PCR data at SIRS-Lab GmbH Version 1.46.0 Date 2007-19-04 Author Matthias Kohl Maintainer Matthias Kohl
A peer-reviewed version of this preprint was published in PeerJ on 16 March 2018.
 A peer-reviewed version of this preprint was published in PeerJ on 16 March 2018. View the peer-reviewed version (peerj.com/articles/4473), which is the preferred citable publication unless you specifically
A peer-reviewed version of this preprint was published in PeerJ on 16 March 2018. View the peer-reviewed version (peerj.com/articles/4473), which is the preferred citable publication unless you specifically
Package forams. R topics documented: July 4, Type Package
 Type Package Package forams July 4, 2015 Title Foraminifera and Community Ecology Analyses Version 2.0-5 Date 2015-07-03 Author Maintainer SHE, FORAM Index and ABC Method analyses
Type Package Package forams July 4, 2015 Title Foraminifera and Community Ecology Analyses Version 2.0-5 Date 2015-07-03 Author Maintainer SHE, FORAM Index and ABC Method analyses
ChIP-Seq Data Analysis. J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015
 ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA
ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA
Enhancing motif finding models using multiple sources of genome-wide data
 Enhancing motif finding models using multiple sources of genome-wide data Heejung Shim 1, Oliver Bembom 2, Sündüz Keleş 3,4 1 Department of Human Genetics, University of Chicago, 2 Division of Biostatistics,
Enhancing motif finding models using multiple sources of genome-wide data Heejung Shim 1, Oliver Bembom 2, Sündüz Keleş 3,4 1 Department of Human Genetics, University of Chicago, 2 Division of Biostatistics,
Instruction Manual Version DNA Elution Module. Cat. No. C (mc-magme-002)
 Instruction Manual Version 2-01.14 DNA Elution Module Cat. No. C01010120 (mc-magme-002) diagenode headquarters Diagenode s.a. BELGIUM EUROPE LIEGE SCIENCE PARK Rue Bois Saint-Jean, 3 4102 Seraing - Belgium
Instruction Manual Version 2-01.14 DNA Elution Module Cat. No. C01010120 (mc-magme-002) diagenode headquarters Diagenode s.a. BELGIUM EUROPE LIEGE SCIENCE PARK Rue Bois Saint-Jean, 3 4102 Seraing - Belgium
EpiQuik Quantitative PCR Fast Kit Base Catalog # P-1029
 EpiQuik Quantitative PCR Fast Kit Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE Uses: The EpiQuik Quantitative PCR Fast Kit is designed for quantitative real time analysis of DNA samples
EpiQuik Quantitative PCR Fast Kit Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE Uses: The EpiQuik Quantitative PCR Fast Kit is designed for quantitative real time analysis of DNA samples
Package disco. March 3, 2017
 Package disco March 3, 2017 Title Discordance and Concordance of Transcriptomic Responses Version 0.5 Concordance and discordance of homologous gene regulation allows comparing reaction to stimuli in different
Package disco March 3, 2017 Title Discordance and Concordance of Transcriptomic Responses Version 0.5 Concordance and discordance of homologous gene regulation allows comparing reaction to stimuli in different
Introduction to genome biology
 Introduction to genome biology Lisa Stubbs We ve found most genes; but what about the rest of the genome? Genome size* 12 Mb 95 Mb 170 Mb 1500 Mb 2700 Mb 3200 Mb #coding genes ~7000 ~20000 ~14000 ~26000
Introduction to genome biology Lisa Stubbs We ve found most genes; but what about the rest of the genome? Genome size* 12 Mb 95 Mb 170 Mb 1500 Mb 2700 Mb 3200 Mb #coding genes ~7000 ~20000 ~14000 ~26000
ideal ChIP-seq kit for Transcription Factors
 ideal ChIP-seq kit for Transcription Factors Cat. No. C01010054 (10 rxns) C01010055 (24 rxns) C01010170 (100 rxns) USER GUIDE Version 3 22_03_2017 Please read this manual carefully before starting your
ideal ChIP-seq kit for Transcription Factors Cat. No. C01010054 (10 rxns) C01010055 (24 rxns) C01010170 (100 rxns) USER GUIDE Version 3 22_03_2017 Please read this manual carefully before starting your
Sanger vs Next-Gen Sequencing
 Tools and Algorithms in Bioinformatics GCBA815/MCGB815/BMI815, Fall 2017 Week-8: Next-Gen Sequencing RNA-seq Data Analysis Babu Guda, Ph.D. Professor, Genetics, Cell Biology & Anatomy Director, Bioinformatics
Tools and Algorithms in Bioinformatics GCBA815/MCGB815/BMI815, Fall 2017 Week-8: Next-Gen Sequencing RNA-seq Data Analysis Babu Guda, Ph.D. Professor, Genetics, Cell Biology & Anatomy Director, Bioinformatics
Package ForestTools. September 8, 2017
 Type Package Title Analyzing Remotely Sensed Forest Data Version 0.1.5 Date 2017-09-07 Author Andrew Plowright Package ForestTools September 8, 2017 Maintainer Andrew Plowright
Type Package Title Analyzing Remotely Sensed Forest Data Version 0.1.5 Date 2017-09-07 Author Andrew Plowright Package ForestTools September 8, 2017 Maintainer Andrew Plowright
Chromatin and Transcription
 Chromatin and Transcription Chromatin Structure Chromatin Represses Transcription Nucleosome Positioning Histone Acetylation Chromatin Remodeling Histone Methylation CHIP Analysis Chromatin and Elongation
Chromatin and Transcription Chromatin Structure Chromatin Represses Transcription Nucleosome Positioning Histone Acetylation Chromatin Remodeling Histone Methylation CHIP Analysis Chromatin and Elongation
Auto ideal ChIP-seq kit for Histones
 Auto ideal ChIP-seq kit for Histones Cat. No. C01010057 (24 rxns) Cat. No. C01010171 (100 rxns) Version 2 I 07.15 Contacts DIAGENODE HEADQUARTERS Diagenode s.a. BELGIUM EUROPE LIEGE SCIENCE PARK Rue Bois
Auto ideal ChIP-seq kit for Histones Cat. No. C01010057 (24 rxns) Cat. No. C01010171 (100 rxns) Version 2 I 07.15 Contacts DIAGENODE HEADQUARTERS Diagenode s.a. BELGIUM EUROPE LIEGE SCIENCE PARK Rue Bois
Supplementary Figure 1. HiChIP provides confident 1D factor binding information.
 Supplementary Figure 1 HiChIP provides confident 1D factor binding information. a, Reads supporting contacts called using the Mango pipeline 19 for GM12878 Smc1a HiChIP and GM12878 CTCF Advanced ChIA-PET
Supplementary Figure 1 HiChIP provides confident 1D factor binding information. a, Reads supporting contacts called using the Mango pipeline 19 for GM12878 Smc1a HiChIP and GM12878 CTCF Advanced ChIA-PET
ChIP-seq analysis. adapted from J. van Helden, M. Defrance, C. Herrmann, D. Puthier, N. Servant
 ChIP-seq analysis adapted from J. van Helden, M. Defrance, C. Herrmann, D. Puthier, N. Servant http://biow.sb-roscoff.fr/ecole_bioinfo/training_material/chip-seq/documents/presentation_chipseq.pdf A model
ChIP-seq analysis adapted from J. van Helden, M. Defrance, C. Herrmann, D. Puthier, N. Servant http://biow.sb-roscoff.fr/ecole_bioinfo/training_material/chip-seq/documents/presentation_chipseq.pdf A model
The ChIP-Seq project. Giovanna Ambrosini, Philipp Bucher. April 19, 2010 Lausanne. EPFL-SV Bucher Group
 The ChIP-Seq project Giovanna Ambrosini, Philipp Bucher EPFL-SV Bucher Group April 19, 2010 Lausanne Overview Focus on technical aspects Description of applications (C programs) Where to find binaries,
The ChIP-Seq project Giovanna Ambrosini, Philipp Bucher EPFL-SV Bucher Group April 19, 2010 Lausanne Overview Focus on technical aspects Description of applications (C programs) Where to find binaries,
#9005. SimpleChIP Plus Enzymatic Chromatin IP Kit (Magnetic Beads) n 1 Kit. (30 Immunoprecipitations) Store at RT, 4 C, and 20 C
 Store at RT, 4 C, and 20 C #9005 SimpleChIP Plus Enzymatic Chromatin IP Kit (Magnetic Beads) 3 n 1 Kit (30 Immunoprecipitations) rev. 09/05/17 Orders n 877-616-CELL (2355) orders@cellsignal.com Support
Store at RT, 4 C, and 20 C #9005 SimpleChIP Plus Enzymatic Chromatin IP Kit (Magnetic Beads) 3 n 1 Kit (30 Immunoprecipitations) rev. 09/05/17 Orders n 877-616-CELL (2355) orders@cellsignal.com Support
ab ChIP Kit Magnetic One-Step
 ab156907 ChIP Kit Magnetic One-Step Instructions for Use For selective enrichment of a chromatin fraction containing specific DNA sequences in a high throughput format using chromatin isolated from various
ab156907 ChIP Kit Magnetic One-Step Instructions for Use For selective enrichment of a chromatin fraction containing specific DNA sequences in a high throughput format using chromatin isolated from various
Purification of nanogram-range immunoprecipitated DNA in ChIP-seq application
 Zhong et al. BMC Genomics (2017) 18:985 DOI 10.1186/s12864-017-4371-5 METHODOLOGY Purification of nanogram-range immunoprecipitated DNA in ChIP-seq application Jian Zhong 1, Zhenqing Ye 2, Samuel W. Lenz
Zhong et al. BMC Genomics (2017) 18:985 DOI 10.1186/s12864-017-4371-5 METHODOLOGY Purification of nanogram-range immunoprecipitated DNA in ChIP-seq application Jian Zhong 1, Zhenqing Ye 2, Samuel W. Lenz
IMMUNOPRECIPITATION (IP)
 1 IMMUNOPRECIPITATION (IP) Overview and Technical Tips 2 CONTENTS 3 7 8 9 12 13 17 18 19 20 Introduction Factors Influencing IP General Protocol Modifications Of IP Protocols Troubleshooting Contact Us
1 IMMUNOPRECIPITATION (IP) Overview and Technical Tips 2 CONTENTS 3 7 8 9 12 13 17 18 19 20 Introduction Factors Influencing IP General Protocol Modifications Of IP Protocols Troubleshooting Contact Us
Epigenetic Regulation of Cell Type Specific Expression Patterns in the Human Mammary Epithelium
 Epigenetic Regulation of Cell Type Specific Expression Patterns in the Human Mammary Epithelium Reo Maruyama 1,2,3., Sibgat Choudhury 1,2,3., Adam Kowalczyk 4,5, Marina Bessarabova 6, Bryan Beresford-Smith
Epigenetic Regulation of Cell Type Specific Expression Patterns in the Human Mammary Epithelium Reo Maruyama 1,2,3., Sibgat Choudhury 1,2,3., Adam Kowalczyk 4,5, Marina Bessarabova 6, Bryan Beresford-Smith
Introduction to Bioinformatics
 Introduction to Bioinformatics Richard Corbett Canada s Michael Smith Genome Sciences Centre Vancouver, British Columbia June 28, 2017 Our mandate is to advance knowledge about cancer and other diseases
Introduction to Bioinformatics Richard Corbett Canada s Michael Smith Genome Sciences Centre Vancouver, British Columbia June 28, 2017 Our mandate is to advance knowledge about cancer and other diseases
ab Plant Chromatin Extraction Kit
 ab156906 Plant Chromatin Extraction Kit Instructions for Use For isolating chromatin or DNA-protein complex from plants in a simple and rapid format This product is for research use only and is not intended
ab156906 Plant Chromatin Extraction Kit Instructions for Use For isolating chromatin or DNA-protein complex from plants in a simple and rapid format This product is for research use only and is not intended
ideal ChIP-seq kit for Histones
 ideal ChIP-seq kit for Histones Cat. No. C01010050 (10 rxns) Version 5 I 12.16 Ordering information diagenode headquarters Diagenode s.a. BELGIUM EUROPE LIEGE SCIENCE PARK Rue Bois Saint-Jean, 3 4102 Seraing
ideal ChIP-seq kit for Histones Cat. No. C01010050 (10 rxns) Version 5 I 12.16 Ordering information diagenode headquarters Diagenode s.a. BELGIUM EUROPE LIEGE SCIENCE PARK Rue Bois Saint-Jean, 3 4102 Seraing
Bioinformatics of Transcriptional Regulation
 Bioinformatics of Transcriptional Regulation Carl Herrmann IPMB & DKFZ c.herrmann@dkfz.de Wechselwirkung von Maßnahmen und Auswirkungen Einflussmöglichkeiten in einem Dialog From genes to active compounds
Bioinformatics of Transcriptional Regulation Carl Herrmann IPMB & DKFZ c.herrmann@dkfz.de Wechselwirkung von Maßnahmen und Auswirkungen Einflussmöglichkeiten in einem Dialog From genes to active compounds
Epigenetic CHaracterization and Observation (ECHO) Proposers Day
 Epigenetic CHaracterization and Observation (ECHO) Proposers Day Eric Van Gieson, Ph.D. Program Manager Biological Technologies Office (BTO) February 23, 2018 ECHO Program Overview Determining Exposure
Epigenetic CHaracterization and Observation (ECHO) Proposers Day Eric Van Gieson, Ph.D. Program Manager Biological Technologies Office (BTO) February 23, 2018 ECHO Program Overview Determining Exposure
Perm-seq: Mapping Protein-DNA Interactions in Segmental Duplication and Highly Repetitive Regions of Genomes with Prior- Enhanced Read Mapping
 RESEARCH ARTICLE Perm-seq: Mapping Protein-DNA Interactions in Segmental Duplication and Highly Repetitive Regions of Genomes with Prior- Enhanced Read Mapping Xin Zeng 1,BoLi 2, Rene Welch 1, Constanza
RESEARCH ARTICLE Perm-seq: Mapping Protein-DNA Interactions in Segmental Duplication and Highly Repetitive Regions of Genomes with Prior- Enhanced Read Mapping Xin Zeng 1,BoLi 2, Rene Welch 1, Constanza
Package powerpkg. February 20, 2015
 Type Package Package powerpkg February 20, 2015 Title Power analyses for the affected sib pair and the TDT design Version 1.5 Date 2012-10-11 Author Maintainer (1) To estimate the power
Type Package Package powerpkg February 20, 2015 Title Power analyses for the affected sib pair and the TDT design Version 1.5 Date 2012-10-11 Author Maintainer (1) To estimate the power
Package VLF. February 19, 2015
 Type Package Package VLF February 19, 2015 Title Frequency Matrix Approach for Assessing Very Low Frequency Variants in Sequence Records Version 1.0 Date 2013-10-25 Author Maintainer Taryn B. T. Athey
Type Package Package VLF February 19, 2015 Title Frequency Matrix Approach for Assessing Very Low Frequency Variants in Sequence Records Version 1.0 Date 2013-10-25 Author Maintainer Taryn B. T. Athey
Le proteine regolative variano nei vari tipi cellulari e in funzione degli stimoli ambientali
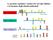 Le proteine regolative variano nei vari tipi cellulari e in funzione degli stimoli ambientali Tipo cellulare 1 Tipo cellulare 2 Tipo cellulare 3 DNA-protein Crosslink Lisi Frammentazione Immunopurificazione
Le proteine regolative variano nei vari tipi cellulari e in funzione degli stimoli ambientali Tipo cellulare 1 Tipo cellulare 2 Tipo cellulare 3 DNA-protein Crosslink Lisi Frammentazione Immunopurificazione
Package DNABarcodes. March 1, 2018
 Type Package Package DNABarcodes March 1, 2018 Title A tool for creating and analysing DNA barcodes used in Next Generation Sequencing multiplexing experiments Version 1.9.0 Date 2014-07-23 Author Tilo
Type Package Package DNABarcodes March 1, 2018 Title A tool for creating and analysing DNA barcodes used in Next Generation Sequencing multiplexing experiments Version 1.9.0 Date 2014-07-23 Author Tilo
Introductory Next Gen Workshop
 Introductory Next Gen Workshop http://www.illumina.ucr.edu/ http://www.genomics.ucr.edu/ Workshop Objectives Workshop aimed at those who are new to Illumina sequencing and will provide: - a basic overview
Introductory Next Gen Workshop http://www.illumina.ucr.edu/ http://www.genomics.ucr.edu/ Workshop Objectives Workshop aimed at those who are new to Illumina sequencing and will provide: - a basic overview
WORKSHOP. Transcriptional circuitry and the regulatory conformation of the genome. Ofir Hakim Faculty of Life Sciences
 WORKSHOP Transcriptional circuitry and the regulatory conformation of the genome Ofir Hakim Faculty of Life Sciences Chromosome conformation capture (3C) Most GR Binding Sites Are Distant From Regulated
WORKSHOP Transcriptional circuitry and the regulatory conformation of the genome Ofir Hakim Faculty of Life Sciences Chromosome conformation capture (3C) Most GR Binding Sites Are Distant From Regulated
Introduction to the UCSC genome browser
 Introduction to the UCSC genome browser Dominik Beck NHMRC Peter Doherty and CINSW ECR Fellow, Senior Lecturer Lowy Cancer Research Centre, UNSW and Centre for Health Technology, UTS SYDNEY NSW AUSTRALIA
Introduction to the UCSC genome browser Dominik Beck NHMRC Peter Doherty and CINSW ECR Fellow, Senior Lecturer Lowy Cancer Research Centre, UNSW and Centre for Health Technology, UTS SYDNEY NSW AUSTRALIA
CM581A2: NEXT GENERATION SEQUENCING PLATFORMS AND LIBRARY GENERATION
 CM581A2: NEXT GENERATION SEQUENCING PLATFORMS AND LIBRARY GENERATION Fall 2015 Instructors: Coordinator: Carol Wilusz, Associate Professor MIP, CMB Instructor: Dan Sloan, Assistant Professor, Biology,
CM581A2: NEXT GENERATION SEQUENCING PLATFORMS AND LIBRARY GENERATION Fall 2015 Instructors: Coordinator: Carol Wilusz, Associate Professor MIP, CMB Instructor: Dan Sloan, Assistant Professor, Biology,
Introduction Bioo Scientific
 Next Generation Sequencing Catalog 2014-2015 Introduction Bioo Scientific Bioo Scientific is a global life science company headquartered in Austin, TX, committed to providing innovative products and superior
Next Generation Sequencing Catalog 2014-2015 Introduction Bioo Scientific Bioo Scientific is a global life science company headquartered in Austin, TX, committed to providing innovative products and superior
ChIP-seq analysis of protein DNA interactions on the Ion Proton System
 APPLICATION NOTE Ion Proton System ChIP-seq analysis of protein DNA interactions on the Ion Proton System Key findings Optimized ChIP-seq protocol for limited cell populations The protocol combines the
APPLICATION NOTE Ion Proton System ChIP-seq analysis of protein DNA interactions on the Ion Proton System Key findings Optimized ChIP-seq protocol for limited cell populations The protocol combines the
IMGM Laboratories GmbH. Sales Manager
 IMGM Laboratories GmbH Dr. Jennifer K. Kuhn Sales Manager About IMGM Laboratories IMGM Laboratories was founded in 2001 IMGM operates as professional provider of advanced genomic services from research
IMGM Laboratories GmbH Dr. Jennifer K. Kuhn Sales Manager About IMGM Laboratories IMGM Laboratories was founded in 2001 IMGM operates as professional provider of advanced genomic services from research
The SVM2CRM data User s Guide
 The SVM2CRM data User s Guide Guidantonio Malagoli Tagliazucchi, Silvio Bicciato Center for genome research University of Modena and Reggio Emilia, Modena October 31, 2017 October 31, 2017 Contents 1 Overview
The SVM2CRM data User s Guide Guidantonio Malagoli Tagliazucchi, Silvio Bicciato Center for genome research University of Modena and Reggio Emilia, Modena October 31, 2017 October 31, 2017 Contents 1 Overview
Simplify your pathogen testing. Amplify your confidence.
 Simplify your pathogen testing. Amplify your confidence. 2 One protocol. Fewer steps. Better results. Be confident in the knowledge that your pathogen testing process is accurate and reliable with the
Simplify your pathogen testing. Amplify your confidence. 2 One protocol. Fewer steps. Better results. Be confident in the knowledge that your pathogen testing process is accurate and reliable with the
Gene Expression on the Fluidigm BioMark HD
 Gene Expression on the Fluidigm BioMark HD Overview Introduction to Fluidigm James Miller Advantages of the technology Running a Fluidigm gene expression project Paul Lacaze Assay design, chemistry, experimental
Gene Expression on the Fluidigm BioMark HD Overview Introduction to Fluidigm James Miller Advantages of the technology Running a Fluidigm gene expression project Paul Lacaze Assay design, chemistry, experimental
Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017
 Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017 Agenda What is Functional Genomics? RNA Transcription/Gene Expression Measuring Gene
Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017 Agenda What is Functional Genomics? RNA Transcription/Gene Expression Measuring Gene
Package GerminaR. October 25, 2017
 Type Package Package GerminaR October 25, 2017 Title Germination Indexes for Seed Germination Variables for Ecophysiological Studies Version 1.2 Date 2017-10-25 BugReports https://github.com/flavjack/germinar/issues
Type Package Package GerminaR October 25, 2017 Title Germination Indexes for Seed Germination Variables for Ecophysiological Studies Version 1.2 Date 2017-10-25 BugReports https://github.com/flavjack/germinar/issues
Przemyslaw Stempor, Julie Ahringer
 SOFTWARE TOOL ARTICLE SeqPlots - Interactive software for exploratory data analyses, pattern discovery and visualization in genomics [version 1; referees: 2 approved, 1 approved with reservations] Przemyslaw
SOFTWARE TOOL ARTICLE SeqPlots - Interactive software for exploratory data analyses, pattern discovery and visualization in genomics [version 1; referees: 2 approved, 1 approved with reservations] Przemyslaw
Welcome to the NGS webinar series
 Welcome to the NGS webinar series Webinar 1 NGS: Introduction to technology, and applications NGS Technology Webinar 2 Targeted NGS for Cancer Research NGS in cancer Webinar 3 NGS: Data analysis for genetic
Welcome to the NGS webinar series Webinar 1 NGS: Introduction to technology, and applications NGS Technology Webinar 2 Targeted NGS for Cancer Research NGS in cancer Webinar 3 NGS: Data analysis for genetic
The Next Generation of Transcription Factor Binding Site Prediction
 The Next Generation of Transcription Factor Binding Site Prediction Anthony Mathelier*, Wyeth W. Wasserman* Centre for Molecular Medicine and Therapeutics at the Child and Family Research Institute, Department
The Next Generation of Transcription Factor Binding Site Prediction Anthony Mathelier*, Wyeth W. Wasserman* Centre for Molecular Medicine and Therapeutics at the Child and Family Research Institute, Department
MIqPCR: Minimum Information about a Quantitative Polymerase Chain Reaction experiment Version 1.0, 4th April, 2008.
 MIqPCR: Minimum Information about a Quantitative Polymerase Chain Reaction experiment Version 1.0, 4th April, 2008. RDML-Consortium This module identifies the minimum information required to report the
MIqPCR: Minimum Information about a Quantitative Polymerase Chain Reaction experiment Version 1.0, 4th April, 2008. RDML-Consortium This module identifies the minimum information required to report the
ChIPnorm: A Statistical Method for Normalizing and Identifying Differential Regions in Histone Modification ChIP-seq Libraries
 ChIPnorm: A Statistical Method for Normalizing and Identifying Differential Regions in Histone Modification ChIP-seq Libraries Nishanth Ulhas Nair., Avinash Das Sahu 2., Philipp Bucher 3,4 *, Bernard M.
ChIPnorm: A Statistical Method for Normalizing and Identifying Differential Regions in Histone Modification ChIP-seq Libraries Nishanth Ulhas Nair., Avinash Das Sahu 2., Philipp Bucher 3,4 *, Bernard M.
Package bc3net. November 28, 2016
 Version 1.0.4 Date 2016-11-26 Package bc3net November 28, 2016 Title Gene Regulatory Network Inference with Bc3net Author Ricardo de Matos Simoes [aut, cre], Frank Emmert-Streib [aut] Maintainer Ricardo
Version 1.0.4 Date 2016-11-26 Package bc3net November 28, 2016 Title Gene Regulatory Network Inference with Bc3net Author Ricardo de Matos Simoes [aut, cre], Frank Emmert-Streib [aut] Maintainer Ricardo
Mentor: Dr. Bino John
 MAT Shiksha Mantri Mentor: Dr. Bino John Model-based Analysis y of Tiling-arrays g y for ChIP-chip X. Shirley Liu et al., PNAS (2006) vol. 103 no. 33 12457 12462 Tiling Arrays Subtype of microarray chips
MAT Shiksha Mantri Mentor: Dr. Bino John Model-based Analysis y of Tiling-arrays g y for ChIP-chip X. Shirley Liu et al., PNAS (2006) vol. 103 no. 33 12457 12462 Tiling Arrays Subtype of microarray chips
Functional Genomics Research Stream. Research Meeting: June 19, 2012 SYBR Green qpcr, Research Update
 Functional Genomics Research Stream Research Meeting: June 19, 2012 SYBR Green qpcr, Research Update Updates Alternate Lab Meeting Fridays 11:30-1:00 WEL 4.224 Welcome to attend either one Lab Log thanks
Functional Genomics Research Stream Research Meeting: June 19, 2012 SYBR Green qpcr, Research Update Updates Alternate Lab Meeting Fridays 11:30-1:00 WEL 4.224 Welcome to attend either one Lab Log thanks
ChIP-chip protocol adapted for the mod-encode project
 ChIP-chip protocol adapted for the mod-encode project Version 1.2 : August 2007 Nicolas Nègre, Xiaochun Ni, Sergey Lavrov, Giacomo Cavalli and Kevin P. White University of Chicago, Department of Human
ChIP-chip protocol adapted for the mod-encode project Version 1.2 : August 2007 Nicolas Nègre, Xiaochun Ni, Sergey Lavrov, Giacomo Cavalli and Kevin P. White University of Chicago, Department of Human
PRODUCT DESCRIPTIONS AND METRICS
 PRODUCT DESCRIPTIONS AND METRICS Adobe PDM - Adobe Analytics (2015v1) The Products and Services described in this PDM are either On-demand Services or Managed Services (as outlined below) and are governed
PRODUCT DESCRIPTIONS AND METRICS Adobe PDM - Adobe Analytics (2015v1) The Products and Services described in this PDM are either On-demand Services or Managed Services (as outlined below) and are governed
Package goseq. R topics documented: December 23, 2017
 Package goseq December 23, 2017 Version 1.30.0 Date 2017/09/04 Title Gene Ontology analyser for RNA-seq and other length biased data Author Matthew Young Maintainer Nadia Davidson ,
Package goseq December 23, 2017 Version 1.30.0 Date 2017/09/04 Title Gene Ontology analyser for RNA-seq and other length biased data Author Matthew Young Maintainer Nadia Davidson ,
Chromatrap spin column ChIP kit for qpcr
 V 6.4 Chromatrap spin column ChIP kit for qpcr A solid phase chromatin immunoprecipitation assay (ChIP) Protocol v6.4 Catalogue no 500071, 500117, 500115, 500116, 500166, 500168, 500167, 500169 ADVANCEMENTS
V 6.4 Chromatrap spin column ChIP kit for qpcr A solid phase chromatin immunoprecipitation assay (ChIP) Protocol v6.4 Catalogue no 500071, 500117, 500115, 500116, 500166, 500168, 500167, 500169 ADVANCEMENTS
ipure kit v2 Magnetic DNA Purification kit for epigenetic applications Cat. No. C (x24) C (x100) Version 2 I 07.15
 ipure kit v2 Magnetic DNA Purification kit for epigenetic applications Cat. No. C03010014 (x24) C03010015 (x100) Version 2 I 07.15 Ordering information diagenode headquarters Diagenode s.a. BELGIUM EUROPE
ipure kit v2 Magnetic DNA Purification kit for epigenetic applications Cat. No. C03010014 (x24) C03010015 (x100) Version 2 I 07.15 Ordering information diagenode headquarters Diagenode s.a. BELGIUM EUROPE
Myers Lab ChIP-seq Protocol v Modified January 10, 2014
 Myers Lab ChIP-seq Protocol V011014 1 Contact information: Dr. Florencia Pauli Behn HudsonAlpha Institute for Biotechnology 601 Genome Way Huntsville, AL 35806 Telephone: 256-327-5229 Email: fpauli@hudsonalpha.org
Myers Lab ChIP-seq Protocol V011014 1 Contact information: Dr. Florencia Pauli Behn HudsonAlpha Institute for Biotechnology 601 Genome Way Huntsville, AL 35806 Telephone: 256-327-5229 Email: fpauli@hudsonalpha.org
Universal Real-Time PCR Reagents.
 Universal Real-Time PCR Reagents Universal Real-Time PCR Reagents On any instrument. Under any conditions. At any time. In the Tube (FPO) Sso7d (FPO) Processivity (FPO) It s time to rethink what you know
Universal Real-Time PCR Reagents Universal Real-Time PCR Reagents On any instrument. Under any conditions. At any time. In the Tube (FPO) Sso7d (FPO) Processivity (FPO) It s time to rethink what you know
Multi-omics in biology: integration of omics techniques
 31/07/17 Летняя школа по биоинформатике 2017 Multi-omics in biology: integration of omics techniques Konstantin Okonechnikov Division of Pediatric Neurooncology German Cancer Research Center (DKFZ) 2 Short
31/07/17 Летняя школа по биоинформатике 2017 Multi-omics in biology: integration of omics techniques Konstantin Okonechnikov Division of Pediatric Neurooncology German Cancer Research Center (DKFZ) 2 Short
Figure S1: NUN preparation yields nascent, unadenylated RNA with a different profile from Total RNA.
 Summary of Supplemental Information Figure S1: NUN preparation yields nascent, unadenylated RNA with a different profile from Total RNA. Figure S2: rrna removal procedure is effective for clearing out
Summary of Supplemental Information Figure S1: NUN preparation yields nascent, unadenylated RNA with a different profile from Total RNA. Figure S2: rrna removal procedure is effective for clearing out
Package ASGSCA. February 24, 2018
 Type Package Package ASGSCA February 24, 2018 Title Association Studies for multiple SNPs and multiple traits using Generalized Structured Equation Models Version 1.12.0 Date 2014-07-30 Author Hela Romdhani,
Type Package Package ASGSCA February 24, 2018 Title Association Studies for multiple SNPs and multiple traits using Generalized Structured Equation Models Version 1.12.0 Date 2014-07-30 Author Hela Romdhani,
Package dfpk. October 24, 2017
 Type Package Package dfpk October 24, 2017 Title Bayesian Dose-Finding Designs using Pharmacokinetics (PK) for Phase I Clinical Trials Version 3.4.0 Date 2017-10-24 Author Artemis Toumazi ,
Type Package Package dfpk October 24, 2017 Title Bayesian Dose-Finding Designs using Pharmacokinetics (PK) for Phase I Clinical Trials Version 3.4.0 Date 2017-10-24 Author Artemis Toumazi ,
Whole Transcriptome Analysis of Illumina RNA- Seq Data. Ryan Peters Field Application Specialist
 Whole Transcriptome Analysis of Illumina RNA- Seq Data Ryan Peters Field Application Specialist Partek GS in your NGS Pipeline Your Start-to-Finish Solution for Analysis of Next Generation Sequencing Data
Whole Transcriptome Analysis of Illumina RNA- Seq Data Ryan Peters Field Application Specialist Partek GS in your NGS Pipeline Your Start-to-Finish Solution for Analysis of Next Generation Sequencing Data
A PROTEIN INTERACTION NETWORK OF THE MALARIA PARASITE PLASMODIUM FALCIPARUM
 A PROTEIN INTERACTION NETWORK OF THE MALARIA PARASITE PLASMODIUM FALCIPARUM DOUGLAS J. LACOUNT, MARISSA VIGNALI, RAKESH CHETTIER, AMIT PHANSALKAR, RUSSELL BELL, JAY R. HESSELBERTH, LORI W. SCHOENFELD,
A PROTEIN INTERACTION NETWORK OF THE MALARIA PARASITE PLASMODIUM FALCIPARUM DOUGLAS J. LACOUNT, MARISSA VIGNALI, RAKESH CHETTIER, AMIT PHANSALKAR, RUSSELL BELL, JAY R. HESSELBERTH, LORI W. SCHOENFELD,
Introduction to ChIP- Seq data analyses
 Introduction to ChIP- Seq data analyses Outline Introduction to ChIP- seq experiment. Biological motivation. Experimental procedure. Method and software for ChIP- seq peak calling. Protein binding ChIP-
Introduction to ChIP- Seq data analyses Outline Introduction to ChIP- seq experiment. Biological motivation. Experimental procedure. Method and software for ChIP- seq peak calling. Protein binding ChIP-
Genome-Wide Localization of Protein-DNA Binding and Histone Modification by a Bayesian Change-Point Method with ChIP-seq Data
 Genome-Wide Localization of Protein-DNA Binding and Histone Modification by a Bayesian Change-Point Method with ChIP-seq Data Haipeng Xing 1. *, Yifan Mo 1,2., Will Liao 1,2., Michael Q. Zhang 2,3 1 Applied
Genome-Wide Localization of Protein-DNA Binding and Histone Modification by a Bayesian Change-Point Method with ChIP-seq Data Haipeng Xing 1. *, Yifan Mo 1,2., Will Liao 1,2., Michael Q. Zhang 2,3 1 Applied
EpiQ Chromatin Analysis Kit Primer Design and qpcr Optimization Guide
 EpiQ Chromatin Analysis Kit Primer Design and qpcr Optimization Guide Table of Contents I. INTRODUCTION II. PRIMER DESIGN A. Finding the target gene using the UCSC Genome Bioinformatics Site B. Identifying
EpiQ Chromatin Analysis Kit Primer Design and qpcr Optimization Guide Table of Contents I. INTRODUCTION II. PRIMER DESIGN A. Finding the target gene using the UCSC Genome Bioinformatics Site B. Identifying
Leveraging biological replicates to improve analysis in ChIP-seq experiments
 , http://dx.doi.org/10.5936/csbj.201401002 CSBJ Leveraging biological replicates to improve analysis in ChIP-seq experiments Yajie Yang a,b, Justin Fear a,b, Jianhong Hu c, Irina Haecker d, Lei Zhou a,
, http://dx.doi.org/10.5936/csbj.201401002 CSBJ Leveraging biological replicates to improve analysis in ChIP-seq experiments Yajie Yang a,b, Justin Fear a,b, Jianhong Hu c, Irina Haecker d, Lei Zhou a,
Nature Genetics: doi: /ng Supplementary Figure 1. Analysis of DHS STARR-seq in the P5424 cell line.
 Supplementary Figure 1 Analysis of DHS STARR-seq in the P5424 cell line. (a) Luciferase enhancer assays of proximal DHSs defined as active or inactive enhancers by STARR-seq in P5424 cells. For each candidate,
Supplementary Figure 1 Analysis of DHS STARR-seq in the P5424 cell line. (a) Luciferase enhancer assays of proximal DHSs defined as active or inactive enhancers by STARR-seq in P5424 cells. For each candidate,
A Polycomb Group Protein Is Retained at Specific Sites on Chromatin in Mitosis
 A Polycomb Group Protein Is Retained at Specific Sites on Chromatin in Mitosis The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters.
A Polycomb Group Protein Is Retained at Specific Sites on Chromatin in Mitosis The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters.
Gene Regulation Solutions. Microarrays and Next-Generation Sequencing
 Gene Regulation Solutions Microarrays and Next-Generation Sequencing Gene Regulation Solutions The Microarrays Advantage Microarrays Lead the Industry in: Comprehensive Content SurePrint G3 Human Gene
Gene Regulation Solutions Microarrays and Next-Generation Sequencing Gene Regulation Solutions The Microarrays Advantage Microarrays Lead the Industry in: Comprehensive Content SurePrint G3 Human Gene
Package EcoTroph. February 19, 2015
 Type Package Title EcoTroph R package Version 1.6 Date 2013-08-31 Package EcoTroph February 19, 2015 Author J. Guitton, M. Colleter, P. Gatti, and D. Gascuel Maintainer Jerome Guitton
Type Package Title EcoTroph R package Version 1.6 Date 2013-08-31 Package EcoTroph February 19, 2015 Author J. Guitton, M. Colleter, P. Gatti, and D. Gascuel Maintainer Jerome Guitton
Package xbreed. April 3, 2017
 Package xbreed April 3, 2017 Title Genomic Simulation of Purebred and Crossbred Populations Version 1.0.1 Description Simulation of purebred and crossbred genomic data as well as pedigree and phenotypes
Package xbreed April 3, 2017 Title Genomic Simulation of Purebred and Crossbred Populations Version 1.0.1 Description Simulation of purebred and crossbred genomic data as well as pedigree and phenotypes
Package sitreee. July 14, 2017
 Version 0.0-1 Date 2017-07-14 Title Sitree Extensions Author Clara Anton Fernandez [aut, cre] Package sitreee July 14, 2017 Maintainer Clara Anton Fernandez Depends R (>= 3.1.0), sitree
Version 0.0-1 Date 2017-07-14 Title Sitree Extensions Author Clara Anton Fernandez [aut, cre] Package sitreee July 14, 2017 Maintainer Clara Anton Fernandez Depends R (>= 3.1.0), sitree
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/2/3/e5473/dc Supplementary Materials for FOXA defines cancer cell specificity Gaihua Zhang, Yongbing Zhao, Yi Liu, Li-Pin Kao, Xiao Wang, Benjamin Skerry, Zhaoyu
advances.sciencemag.org/cgi/content/full/2/3/e5473/dc Supplementary Materials for FOXA defines cancer cell specificity Gaihua Zhang, Yongbing Zhao, Yi Liu, Li-Pin Kao, Xiao Wang, Benjamin Skerry, Zhaoyu
Supporting Information. SI Material and Methods
 Supporting Information SI Material and Methods RNA extraction and qrt-pcr RNA extractions were done using the classical phenol-chloroform method. Total RNA samples were treated with Turbo DNA-free (Ambion)
Supporting Information SI Material and Methods RNA extraction and qrt-pcr RNA extractions were done using the classical phenol-chloroform method. Total RNA samples were treated with Turbo DNA-free (Ambion)
Whole genome Bisulfite Sequencing for Methylation Analysis Preparing Samples for the Illumina Sequencing Platform
 Whole genome Bisulfite Sequencing for Methylation Analysis Preparing Samples for the Illumina Sequencing Platform Introduction, 2 Sample Prep Workflow, 3 Best Practices, 4 DNA Input Recommendations, 6
Whole genome Bisulfite Sequencing for Methylation Analysis Preparing Samples for the Illumina Sequencing Platform Introduction, 2 Sample Prep Workflow, 3 Best Practices, 4 DNA Input Recommendations, 6
SOLiD Total RNA-Seq Kit SOLiD RNA Barcoding Kit
 SOLiD Total RNA-Seq Kit SOLiD RNA Barcoding Kit Agenda SOLiD Total RNAseq Kit Overview Kit Configurations Barcoding Kit Introduction New Small RNA and WT Workflow Small RNA Workflow Step-by-step Workflow
SOLiD Total RNA-Seq Kit SOLiD RNA Barcoding Kit Agenda SOLiD Total RNAseq Kit Overview Kit Configurations Barcoding Kit Introduction New Small RNA and WT Workflow Small RNA Workflow Step-by-step Workflow
Considerations for Illumina library preparation. Henriette O Geen June 20, 2014 UCD Genome Center
 Considerations for Illumina library preparation Henriette O Geen June 20, 2014 UCD Genome Center Diversity of applications De novo genome Sequencing ranscriptome Expression Splice Isoform bundance Genotyping
Considerations for Illumina library preparation Henriette O Geen June 20, 2014 UCD Genome Center Diversity of applications De novo genome Sequencing ranscriptome Expression Splice Isoform bundance Genotyping
Controlling DNA. Ethical guidelines for the use of DNA technology. Module Type: Discussion, literature review, and debate
 Ethical guidelines for the use of DNA technology Author: Tara Cornelisse, Ph.D. Candidate, Environmental Studies, University of California Santa Cruz. Field-tested with: 11 th -12 th grade students in
Ethical guidelines for the use of DNA technology Author: Tara Cornelisse, Ph.D. Candidate, Environmental Studies, University of California Santa Cruz. Field-tested with: 11 th -12 th grade students in
Package microrna. R topics documented: January 21, Version Author R. Gentleman, S. Falcon
 Version 1.36.0 Author R. Gentleman, S. Falcon Package microrna January 21, 2018 Title Data and functions for dealing with micrornas Different data resources for micrornas and some functions for manipulating
Version 1.36.0 Author R. Gentleman, S. Falcon Package microrna January 21, 2018 Title Data and functions for dealing with micrornas Different data resources for micrornas and some functions for manipulating
Biotools: an R function to predict spatial gene diversity via an individual-based approach
 Biotools: an R function to predict spatial gene diversity via an individual-based approach A.R. da Silva 1, G. Malafaia 2 and I.P.P. Menezes 3 1 Laboratório de Estatística Aplicada, Instituto Federal Goiano,
Biotools: an R function to predict spatial gene diversity via an individual-based approach A.R. da Silva 1, G. Malafaia 2 and I.P.P. Menezes 3 1 Laboratório de Estatística Aplicada, Instituto Federal Goiano,
A pipeline for ChIP-seq data analysis (Prot 56)
 A pipeline for ChIP-seq data analysis (Prot 56) Ruhi Ali 1, Florence M.G. Cavalli 1, Juan M. Vaquerizas 1 and Nicholas M. Luscombe 2,3,4. 1. European Bioinformatics Institute. Wellcome Trust Genome Campus,
A pipeline for ChIP-seq data analysis (Prot 56) Ruhi Ali 1, Florence M.G. Cavalli 1, Juan M. Vaquerizas 1 and Nicholas M. Luscombe 2,3,4. 1. European Bioinformatics Institute. Wellcome Trust Genome Campus,
Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate
 Supplementary Figure Legends Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate BC041951 in gastric cancer. (A) The flow chart for selected candidate lncrnas in 660 up-regulated
Supplementary Figure Legends Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate BC041951 in gastric cancer. (A) The flow chart for selected candidate lncrnas in 660 up-regulated
