OPTIMIZATION OF DNA ENCODING BASED ON COMBINATORIAL CONSTRAINTS. Received October 2007; accepted January 2008
|
|
|
- Jeremy Hunt
- 6 years ago
- Views:
Transcription
1 ICIC Express Letters ICIC International c 2008 ISSN X Volume 2, Number 1, March 2008 pp OPTIMIZATION OF DNA ENCODING BASED ON COMBINATORIAL CONSTRAINTS Jinsong Li, Qiang Zhang, Rui Li and Shihua Zhou Liaoning Key Lab of Intelligent information Processing Dalian University Dalian , P. R. China Corresponding author: zhangq30@yahoo.com Received October 2007; accepted January 2008 Abstract. The design of DNA sequences plays an important role in DNA computing. In this paper, we primarily focus on the combinatorial constraints of DNA sequences in DNA encoding, and then analyze their relationships. We propose the evaluative formulas based on those constraints and transform DNA sequences design into a multi-objective optimize problem. The artificial immune algorithm (AIA)/simulated annealing algorithm (SAA) is presented to solve the problem, and better DNA sequences than the previously known constructions are obtained. Keywords: DNA encoding, Combinatorial constraints, Multi-objective optimize, AIA/SA algorithm 1. Introduction. DNA computing started in 1994 when Adleman solved a hard (NPcomplete) computational problem with an exceptional method [1]. Because of the huge numbers of DNA molecules in a typical test tube, any method of computation based on DNA would seem to have potential for a massive parallelism, capacity, and power. This potential, however, is limited by the constraints imposed by the chemical of DNA. DNA of higher species consists of two complementary chains twisted around each other to form a double helix. Each chain is a linear sequence of nucleotides, or bases- two purines, adenine(a) and guanine (G), and two pyrimidines, thymine (T) and cytosine (C). The purine bases and pyrimidine bases are Watson-Crick (WC) complements of each other, in the sense that Ā = T, T = A, C = G, Ḡ = C (1) Although there are only four kinds of nucleotides in DNA strand, and two kinds of complement ways, base pairs with a wide variety of sequencing constitute the diversity of DNA molecule, and then influences DNA computing. In DNA computation, the core reaction is the specific hybridization between DNA sequences or Watson-Crick complement, which directly influences the reliability of DNA computation with its efficiency and accuracy. However, false hybridization can occur because of the chemical characteristics of DNA molecules [2], which could cause wrong results in DNA computing. For example, as shown in Figure 1, false hybridization happened in 2, 3, 4, and 5. The DNA encoding problem means trying to encode every bit of DNA code words in order to make the DNA strands hybridize with its complement specifically in bio-chemical reaction. But, actually, DNA sequence encoding is a very hard optimization problem. In DNA computing, DNA molecules as data couldn t be generated stochastically, because if the sequences are not carefully chosen, DNA sequence may fold back onto itself, forming a secondary structure which completely inhibits the sequence from participating in the computational process, and how to select the length of DNA sequence according to the scale of problems needs to be considered Hence, in accordance with the specific problem 81
2 82 J. S. LI, Q. ZHANG, R. LI AND S. H. ZHOU Figure 1. Some kinds of hybridization to be solved, what kind of encoding is to be chosen for optimizing the DNA sequence is a fundamental problem in DNA computing. So far, several papers have proposed different algorithms to design DNA sequences [5-15,20,21], for example, Deaton and Garzon introduced a new measure of hybridization likelihoodbasedonhammingdistanceandproposed a theory of error-preventing codes for DNA computing [5]. Marathe et al. proposed dynamic programming algorithms based on Hamming distance and free energy [6]. Frotos et al. proposed template-map method [7]. Hartemink et al. developed the program SCAN [8], Arita et al. developed a sequence design system using GA and a random generate-and-test algorithm [9]. Tanaka et al. developed a support system for sequence design using SA algorithms, and listed up some fitness criteria [10]. Deaton et al. proposed an evolution search method [11]. Soo-Yong Shin et al. developed an evolutionary sequence generation system to minimize the potential errors for DNA computing [12]. In [13,14], the authors used stochastic search algorithms to design codewords that are suitable for DNA computing. Rykov et al. proposed to use cyclic codes over to construct DNA codes [15]. In [20,21], Zhou et al. used the Particle Swarm Optimization to get the DNA sequence that fit fordna computing. In a sense, DNA encoding is essentially a multi-objective optimization problem. The modern intelligent optimization methods (Genetic Algorithm, Artificial Immune Algorithm, Particle Swarm Optimization,Ant Colony Optimization Algorithm et al) present effective ways to solve complex optimization problems. Various optimization algorithms inthednaencodingareprovedtobeverygoodapplication. In Soo-Yong Shin s paper, better sequences were generated by using GA algorithm to solve the multi-objective optimization problem [12]. In [13] Wang used GA/SA to get DNA sequences. However, in the traditional genetic algorithm, probability of crossover and mutation are constant, the individual with better fitness value and the population diversity may be damaged. So in this paper, we use Artificial Immune Algorithm (AIA) [17-18], in combination with simulated annealing algorithm to solve the multi-objective optimization problem, and we get a better result of the DNA sequence than the previously best known results [12,13], the DNA sequence with good structures can limit non-specific hybirdzation, and have similar thermodynamic stability. 2. DNA Encoding and Its Constraints DNA encoding [5]. The DNA encoding problem can be described as follows: the encoding alphabet of DNA sequence is the set of nucleic acid bases Σ = {A, T, C, G},and in the encoding set Z of DNA strands whose length is L, Z = Σ 1 = {< b 1,b 2,,b n >, b i Σ,i =1, 2,,L} and Z =4 l,searchthesubsetw of Z which satisfies for x i,x j W, τ(x i,x j ) >k,wherek is a positive integer, and τ is the expected criterion of evaluating the encoding, namely the encodings should satisfy constraint conditions.
3 ICIC EXPRESS LETTERS, VOL.2, NO.1, DNA encoding constraints. Theoretically, the DNA encoding should satisfy two kinds of constraints, one is combinatorial constraints; the other is thermodynamic constraints Combinatorial constraints. Anyone of computing paradigm can be attributed to the information transmission and processing. Error-correcting codes in information theory effectively solved the problem of electronic computer encoding. Its mathematical basis is to measure distance between the two vertexes in the binary hypercube space by hamming distance. Due to the special nature of DNA Computing, we give their definition [6]: In the following discussion, x i (1 i<m),x j (1 j<m) are supposed as the DNA sequence whose length is n, and m denotes the count of DNA sequence in alphabet = {A, C, G, T }. For convenience, DNA strand is oriented from the 5 0 to 3 0 end. x c whose orientation is the 3 0 to 5 0 end, is supposed as Watson-Crick complement of a strand x. x 0 denotes the reverse of strand x. 1) Hamming Distance: the number of position i at which the ith letter in x i differs from the ith letter in x j. 2) Reverse Hamming Distance: the Hamming Distance between the sequence x i and x r j.ifx j = q 1 q 2 q n,thenx r j = q n q n 1 q 1. 3) Complement Hamming Distance: the Hamming Distance between the sequence x i and x c j. For binary sequences, their complement sequence is to change all 0 to 1 and 1 to 0, but in the DNA computing, the nucleotides in DNA strand must be changed into the nucleotides of their Watson-Crick Complement and meanwhile, the direction of DNA strand change from to ) Reverse Complement Hamming Distance Constraint: the Hamming Distance between x c i and x r j. 5) Continuity [12,22]: If one letter repeats continuously in DNA sequence, and then the DNA strand structure will become unstable, we should try to avoid in DNA encoding. 6) Hairpin structure [12]: the hairpin structure is caused by DNA sequence folding back onto itself due to self-hybridization. The hairpin structure is not expect in most ways of DNA computing, in our paper, one of goal of DNA encoding is try to reduce the number of its occurrences Thermodynamic constraints. 1) Free energy: Free energy is released energy in the transformation that the single stranded DNA molecules spontaneously transform from high-energy state to double stranded molecules of lower-energy state. Free energy change G is an important parameter evaluating the DNA molecular thermodynamic stability. G is usually calculated approximately by Nearest Neighbor Model parameters that defined in [19]: Xn 1 G = θ + w(b i,b i+1 ) (2) i=1 where θ is modificatory value, ω is the specifically minus value of the sequence b i b i+1. 2) Melting Temperature: Melting temperature is also important for efficiency and reliability of the DNA reaction, there are many factors that influence it., such as the component of DNA, the concentration of DNA, the ph of liquor and so on, the follows (3) is to calculated melting temperature in the nearest neighbor model with SantaLucia s unified NN parameters [19] 3. Algorithm. T m = H /( S + R ln C t ) (3)
4 84 J. S. LI, Q. ZHANG, R. LI AND S. H. ZHOU 3.1. Artificial immune algorithm (AIA). Theimmunesystemisthebasicandremarkable defence system against bacteria, viruses and other disease-causing organisms. It can produce millions of antibodies from hundreds of antibody genes and can protect animals which are infected by foreign molecules [17-18]. The Artificial Immune Algorithm (AIA) was inspired by the immune system. In AIA, the objective function and constraints operate as the antigens, and the solutions to the objective function operate as the antibodies. Similar to Genetic Algorithm (GA), AIA starts by creating antibodies randomly in a feasible space, and finally reaches the optimum vie natural selection, crossover and mutation. Compared with GA, AIA has an affinity calculation function, which can describe the relationship not only between the antigen and the antibody but also between antibodies, which gives AIA the unique characteristic of guaranteeing the survival of variant offspring that can match the antigen better. The higher the affinity, the stronger the binding, and thus the better the immune recognition and response Design of antigen. For comparing with the previous works [12,13], we propose the evaluation function according to Section 2. 1) Hamming Distance Constraint (HDC): A large Hamming distance should be held between any two sequences. f Hamming (i) = min {H(x i,x j )} (4) 1 j<m,j6=i where f Hamming (i) indicates the Hamming evaluation function of theith individual. 2) Reverse Hamming Distance Constraint (RHDC): f Reverse (i) = min {H(x i,x β j )} (5) 1 j<m 3) Reverse Complement Constraint (RCC): f Reverse Complement (i) = min 1 j<m {H(x0 i,x β j )} (6) 4) Continuity Constraint (CC): f Con (i) = nx j=1 (j 1)N (i) j (7) where N (i) j denotes the number of times to whichthesamebaseappearsisj-times continuously in sequence. 5) Hairpin Constraint (HC): f Hairpin (i) = n 2 pinlen X r=5 n pinlen br/2c X c=pinlen+dr/2e Hairpin(x i,c) (8) where r is the minimum length to form hairpin ring,pinlen denotes the minimum sequence length of hybridization to form hairpin, Hairpin(x i,c) is 1, when the reverse-complement distance of two sequence which sequence x i is folded around the cth base is more than pinlen/2,otherwiseis0. 6) GC Content Constraint: f GC (i) = λ GC(i) GC(i) defined (9) where GC(i) defined is the target value of GC content of DNA sequence x i,andgc(i) is the real GC content, λ is the parameter that is used to adjust the weight of the constraint and other constraints. In this paper, we use evaluation function of GC content to evaluate melting temperature, due to the (10). Where Length is the of length DNA sequence. T m RatioGC 500/length (10)
5 ICIC EXPRESS LETTERS, VOL.2, NO.1, ) Free energy Constraint: Besides the concentration of reactant, free energy is influenced by the component of DNA. Under the similar conditions, the more degree of hybridization,the more change of free energy, and hamming distance etc could reflect the degree of hybridization in some extent, so in our paper, the free energy is not considered. We formulate the evaluation function as a maximum problem, and use the weighted sum to deal with the every evaluation function of constraints selected. f j {f Hamming (i),f Reverse (i),f Reverse Complement (i),f Con (i),f Hairpin (i),f GC (i)} (11) 6X fitness(i) = W i f i (12) i= Steps of algorithm. AIA introduces a new operator -immune operator based on the framework of the original genetic algorithm, and including new steps, one is inoculating bacterin which improves the fitness of the population and the other is immune choice which prevents the degenerateness of the population. But those steps can t overcome the disadvantage of genetic algorithm which is difficulttobreakawayfromthelocaloptimal value. The simulated algorithm (SA) can improve the premature convergence default in AIA algorithm. So in this paper, we use simulated algorithm when the antibodies update, and SA operator is designed as follows: ½ 1 fitness(new) fitness(old) P (new old) = exp{λ(fitnew(new) fitneww(old))/t } fitness(new) <fitnew(oid) Steps of Algorithm: Step1: Recognize the invasion of antigens, which corresponds to input data of the scheduling problem. Here multi-objective optimize problem is considered as the antigens, and IGA parameters are inputted. Step2: Generate the antibodies through searching the memory antibodies of the problem, or generating randomly. In our experiment, the population size is 20. Step3: Evaluate the fitness and calculates the affinities between antigen and antibody, antibody and antibody. Step4: Chooses the antigen with larger affinity to input the memory cells and take place of the old antigen. Step5: Generate antibody s expectation, the antigen with lower expectation will be restrained and the antigen with higher expectation will be prompted. Step6: The antibodies that will perform the next generation to optimization are proliferated by selection, crossover and mutation with certain probabilities. Then some more antibodies are generated from memory cells. Step7: If the termination condition is not true, adjust the parameter in SA operator, go to Step 3, otherwise the system is over. ¾ 4. Result and Analysis Simulation results. AIA/SA mentioned above is implemented with Matlab 7.1. The AIA/SA parameters used in our example are: the population size is 20, the generation number is 300, DNA sequence length is 20, probability of crossover is 0.6, the mutation rate is 0.01, initial temperature and cooling schedule about SA are set to be 180 and 0.95, respectively, and the required individual count is 7. Figure 2 illustrates the results of simulation. From Figure 2, we can see that the algorithm has good convergence, and it could get optimal solutions with less generation times comparing with the algorithm in [13]
6 86 J. S. LI, Q. ZHANG, R. LI AND S. H. ZHOU Figure 2. Evolution graph of AIA/SA algorithms 4.2. Comparisons. Table 1 shows the fitness value of DNA sequence based on combinatorial constraints. In our example, better fitness value is obtained by comparing with sequences in the previous works [12,13]. Table 1. The comparison of sequences based on Combinatorial Constraints DNA Sequence(5-3 ) HDC RHDC RCC CC Hairpin GC(%) Fitness DNA sequences in our system AGCTGAGAGCGTGTTATCGA TATCACTCTAGACAGTGCGC CAGTCTCCGTCATCAGATAC CAGCGTGTCGATTATCGTGT GATGCAGACGCACGTATACA GCTATATGCGACGACAGTAC ATAGAGATCCTACGCTGGCT Sequences in Wang s paper [13] ACTATAGACAGCATGCCGCA ACAGACAGTGCTACAGCACG TGATCTGCACATGTCTAGTG TACGCGCACACATGAAAGTG ACGACTGACGTAGCCATGAT ATAGCTGTACGTCGTGTCAG TCTTGCGTATCTCTGCTGAG Sequences in Soo-Yong Shin s paper[12] GAGTTAGATGTCACGTCACG AGGCGAGTAGGGGTATATC TTATGATTCCACTGGCGCTC CCTGTCAACATTGACGCTCA CGCTCCATCCTTGATCGTTT ATCGTACTCATGGTCCCTAC CTTCGCTGCTGATAACCTCA
7 ICIC EXPRESS LETTERS, VOL.2, NO.1, Conclusions. In this paper, we have studied the combinatorial constraints of DNA encoding, and transformed the DNA encoding to multi-objective optimization problem. AIA/SA algorithms are presented to solve the optimization problem, and good sequences are obtained to improve the reliability of DNA computation. Finally, we have proved the efficiency of our algorithms by comparing with the previous works. In future work, we will examine ways to further improve our algorithm, and introduce the thermodynamic constraints in our algorithm. Acknowledgment. This work was supported by National Natural Science Foundation of China (Grant Nos , , ) and Program from New Century Excellent Talents in University of China (NCET ) REFERENCES [1] L. M. Adleman, Molecular computation of solution to combinatorial problems, Science, vol.66, no.11, pp , [2] R. Deaton, R. C. Murphy, M. Garzon, D. R. Franceshetti and S. E. Stevens, Good encodings for DNA-based solutions to combinatorial problems, Proc. of the 2nd DIMACS Workshop on DNA Based Computers, pp , [3] R. Deaton and M. Garzon, Thermodynamic Constraints on DNA-based Computing, in Computing with Bio-Molecus: Theory and Experiments, Springer-Verlag, Singapore, pp , [4] R. Deaton, D. R. Franceschetti, M. Garzon, J. A. Rose, R. C. Murphy and S. E. Stevens, Information transfer through hybridization reaction in DNA based computing, Proc. of the Second Annual Conference, Stanford University, pp , [5] M. Garzon, P. Neathery, R. Deaton, R. J. Murphy, D. R. Franceschetti and S. E Stevens, A new metric for DNA Computing, Proc. of the 2nd Annual Genetic Programming Conference, pp , [6] A. Marathe, A. E. Condon and R. M. Corn, On combinatorial DNA word design, Proc. of the 5th DIMACS Workshop on DNA Based Computers, pp.75-89, [7] A. G. Frutos, et al., Demonstration of a word design strategy for DNA computing on surface, Nucleic Acids Res, vol.l25, pp , 1997 [8]A.J.Hartemink,D.K.Gifford and J. Khodor, Automated constraint-based nucleotide sequence selection for DNA computation, Proc. of the 4th DIMACS Workshop on DNA Based Computers, pp , [9]M.Arita,A.Nishikawa,M.Hagiya,K.Komiya,H.GouzuandK.Sakamoto,Improvingsequence design for DNA computing, Proc. of the Genetic and Evolutionary Computation Conference, pp , [10] F. Tanaka, M. Nakatsugawa, M. Yamamoto, T. Shiba and A. Ohuchi, Developing support system for sequence design in DNA computing, Proc. of the 7th international Workshop on DNA-Based Computers, pp , 2001 [11] R. Deaton, R. C. Murphy, J. A. Rose, M. Garzon, D. R. Franceschetti and S. E. Stevens, A DNA based implementation of an evolutionary search for good encodings for DNA computation, Proc. of the 1997 IEEE International Conference on Evolutionary Computation, pp , [12] S. Y. Shin, D. M. Kim, I. H. Lee and B.-T. Zhang, Evolutionary sequence generation for reliable DNA computing, Congress on Evolutionary Computation, vol.1, pp.79-84, [13] W. Wang, X. D. Zheng, Q. Zhang and J. Xu, The optimization of DNA encodings based on GA/SA algorithms, Progress in Natural Science, vol.17, no.6, pp , [14] D. C. Tulpan, H. H. Hoos and A. E. Condon, Stochastic local search algorithms for DNA word design, in DNA computing: 8th International Workshop on DNA-Based Computers, M. Hagiya and A. Ohuchi (eds.), Springer LNCS, vol.2568, pp , [15] D. C. Tulpan and H. H. Hoos, Hybrid randomized neighbors improve stochastic local search for DNA code design, in Advances in Artificial Intelligence: 16th Conference of the Canadian Society for Computational Studies of Intelligence, Y. Xiang and B. Chaib-draa (eds.), Springer LNCS, vol.2671, pp , [16] V. Rykov, A. J. Macula, D. Torney and P. White, DNA sequences and quaternary cyclic codes, Proc. of the IEEE International Symposium on Information Theory, Washington DC, pp.248, [17] J. E. Hunt and D. E. Cooke, Learning using an artificial immune system, Journal of Network and Computer Applications, vol.19, pp , 1996.
8 88 J. S. LI, Q. ZHANG, R. LI AND S. H. ZHOU [18] G. Lun and C. Chueh, Multi-objective optimal design of truss structure with immune algorithm, Computers and Structures, vol.82, pp , [19] J. SantaLucia, et al., Improved nearest-neighbor parameters for predicting DNA duplex stability, Biochemistry, vol.35, pp , [20] S. H. Zhou, Q. Zhang, J. Zhao and J. S. Li, Optimization of DNA encodings based on free energy, ICIC Express Letters, vol.1, pp ,2007. [21] S. H. Zhou, Q. Zhang, J. Zhao and J. S. Li, DNA encodings based on multiobject particle swarm, Journal of Computational and Theoretical Nanoscience, vol.4, pp , [22] C. C. Chang, T. Lu, Y. F. Chang and C. T. Lee, Reversible data hiding schemes for deoxyribonucleic acid (DNA) medium, Int. J. Innovative Computing, Information and Control, vol.3, no.5, pp , 2007.
DEOXYRIBONUCLEIC ACID (DNA) computing is a
 IEEE TRANSACTIONS ON EVOLUTIONARY COMPUTATION, VOL. 9, NO. 2, APRIL 2005 143 Multiobjective Evolutionary Optimization of DNA Sequences for Reliable DNA Computing Soo-Yong Shin, In-Hee Lee, Dongmin Kim,
IEEE TRANSACTIONS ON EVOLUTIONARY COMPUTATION, VOL. 9, NO. 2, APRIL 2005 143 Multiobjective Evolutionary Optimization of DNA Sequences for Reliable DNA Computing Soo-Yong Shin, In-Hee Lee, Dongmin Kim,
Towards a General-Purpose Sequence Design System in DNA Computing
 Towards a General-Purpose Sequence Design System in DNA Computing Fumiaki Tanaka, Masashi Nakatsugawa, Masahito Yamamoto, Toshikazu Shiba and Azuma Ohuchi Graduate School of Engineering, Hokkaido University,
Towards a General-Purpose Sequence Design System in DNA Computing Fumiaki Tanaka, Masashi Nakatsugawa, Masahito Yamamoto, Toshikazu Shiba and Azuma Ohuchi Graduate School of Engineering, Hokkaido University,
Sequence Design for DNA Computing
 Sequence Design for DNA Computing 2004. 10. 16 Advanced AI Soo-Yong Shin and Byoung-Tak Zhang Biointelligence Laboratory DNA Hydrogen bonds Hybridization Watson-Crick Complement A single-stranded DNA molecule
Sequence Design for DNA Computing 2004. 10. 16 Advanced AI Soo-Yong Shin and Byoung-Tak Zhang Biointelligence Laboratory DNA Hydrogen bonds Hybridization Watson-Crick Complement A single-stranded DNA molecule
DNASequenceGenerator: A Program for the construction of DNA sequences
 DNASequenceGenerator: A Program for the construction of DNA sequences Udo Feldkamp, Sam Saghafi 2, Wolfgang Banzhaf, Hilmar Rauhe Chair of Systems Analysis, University of Dortmund, Germany {feldkamp, banzhaf,
DNASequenceGenerator: A Program for the construction of DNA sequences Udo Feldkamp, Sam Saghafi 2, Wolfgang Banzhaf, Hilmar Rauhe Chair of Systems Analysis, University of Dortmund, Germany {feldkamp, banzhaf,
TravelingSalesmanProblemBasedonDNAComputing
 TravelingSalesmanProblemBasedonDNAComputing Yan Li Weifang University College of computer and communication Engineering Weifang, P.R.China jk97@zjnu.cn Abstract Molecular programming is applied to traveling
TravelingSalesmanProblemBasedonDNAComputing Yan Li Weifang University College of computer and communication Engineering Weifang, P.R.China jk97@zjnu.cn Abstract Molecular programming is applied to traveling
Evolutionary Algorithms
 Evolutionary Algorithms Evolutionary Algorithms What is Evolutionary Algorithms (EAs)? Evolutionary algorithms are iterative and stochastic search methods that mimic the natural biological evolution and/or
Evolutionary Algorithms Evolutionary Algorithms What is Evolutionary Algorithms (EAs)? Evolutionary algorithms are iterative and stochastic search methods that mimic the natural biological evolution and/or
Routing Optimization of Fourth Party Logistics with Reliability Constraints based on Messy GA
 Journal of Industrial Engineering and Management JIEM, 2014 7(5) : 1097-1111 Online ISSN: 2013-0953 Print ISSN: 2013-8423 http://dx.doi.org/10.3926/jiem.1126 Routing Optimization of Fourth Party Logistics
Journal of Industrial Engineering and Management JIEM, 2014 7(5) : 1097-1111 Online ISSN: 2013-0953 Print ISSN: 2013-8423 http://dx.doi.org/10.3926/jiem.1126 Routing Optimization of Fourth Party Logistics
An Improved Immune Genetic Algorithm for Capacitated Vehicle Routing Problem
 Send Orders for Reprints to reprints@benthamscience.ae 560 The Open Cybernetics & Systemics Journal, 2014, 8, 560-565 Open Access An Improved Immune Genetic Algorithm for Capacitated Vehicle Routing Problem
Send Orders for Reprints to reprints@benthamscience.ae 560 The Open Cybernetics & Systemics Journal, 2014, 8, 560-565 Open Access An Improved Immune Genetic Algorithm for Capacitated Vehicle Routing Problem
A Viral Systems Algorithm for the Traveling Salesman Problem
 Proceedings of the 2012 International Conference on Industrial Engineering and Operations Management Istanbul, Turkey, July 3 6, 2012 A Viral Systems Algorithm for the Traveling Salesman Problem Dedy Suryadi,
Proceedings of the 2012 International Conference on Industrial Engineering and Operations Management Istanbul, Turkey, July 3 6, 2012 A Viral Systems Algorithm for the Traveling Salesman Problem Dedy Suryadi,
translation The building blocks of proteins are? amino acids nitrogen containing bases like A, G, T, C, and U Complementary base pairing links
 The actual process of assembling the proteins on the ribosome is called? translation The building blocks of proteins are? Complementary base pairing links Define and name the Purines amino acids nitrogen
The actual process of assembling the proteins on the ribosome is called? translation The building blocks of proteins are? Complementary base pairing links Define and name the Purines amino acids nitrogen
STRUCTURAL OPTIMIZATION USING ARTIFICIAL IMMUNE SYSTEM (AIS)
 Blucher Mechanical Engineering Proceedings May 2014, vol. 1, num. 1 www.proceedings.blucher.com.br/evento/10wccm STRUCTURAL OPTIMIZATION USING ARTIFICIAL IMMUNE SYSTEM (AIS) Sai Sushank Botu 1, S V Barai
Blucher Mechanical Engineering Proceedings May 2014, vol. 1, num. 1 www.proceedings.blucher.com.br/evento/10wccm STRUCTURAL OPTIMIZATION USING ARTIFICIAL IMMUNE SYSTEM (AIS) Sai Sushank Botu 1, S V Barai
HARDWARE ACCELERATION OF MULTI-DEME GENETIC ALGORITHM FOR DNA CODEWORD SEARCHING
 HARDWARE ACCELERATION OF MULTI-DEME GENETIC ALGORITHM FOR DNA CODEWORD SEARCHING Qinru Qiu 1, Daniel Burns 2, Prakash Mukre 1, Qing Wu 1 1 Department of Electrical and Computer Engineering, Binghamton
HARDWARE ACCELERATION OF MULTI-DEME GENETIC ALGORITHM FOR DNA CODEWORD SEARCHING Qinru Qiu 1, Daniel Burns 2, Prakash Mukre 1, Qing Wu 1 1 Department of Electrical and Computer Engineering, Binghamton
Introduction to DNA Computing
 Introduction to DNA Computing The lecture notes were prepared according to Leonard Adleman s seminal paper Molecular Computation of Solutions to Combinatorial Problems and Keith Devlin s explanatory article
Introduction to DNA Computing The lecture notes were prepared according to Leonard Adleman s seminal paper Molecular Computation of Solutions to Combinatorial Problems and Keith Devlin s explanatory article
Hardware Acceleration of Multi-deme Genetic Algorithm for the Application of DNA Codeword Searching
 Hardware Acceleration of Multi-deme Genetic Algorithm for the Application of DNA Codeword Searching Qinru Qiu*, Daniel Burns**, Prakash Mukre*, Qing Wu* *Department of Electrical and Computer Engineering,
Hardware Acceleration of Multi-deme Genetic Algorithm for the Application of DNA Codeword Searching Qinru Qiu*, Daniel Burns**, Prakash Mukre*, Qing Wu* *Department of Electrical and Computer Engineering,
Microarray Probe Design Using ɛ-multi-objective Evolutionary Algorithms with Thermodynamic Criteria
 Microarray Probe Design Using ɛ-multi-objective Evolutionary Algorithms with Thermodynamic Criteria Soo-Yong Shin, In-Hee Lee, and Byoung-Tak Zhang Biointelligence Laboratory, School of Computer Science
Microarray Probe Design Using ɛ-multi-objective Evolutionary Algorithms with Thermodynamic Criteria Soo-Yong Shin, In-Hee Lee, and Byoung-Tak Zhang Biointelligence Laboratory, School of Computer Science
Chapter 13 - Concept Mapping
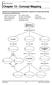 Chapter 13 - Concept Mapping Using the terms and phrases provided below, complete the concept map showing the discovery of DNA structure. amount of base pairs five-carbon sugar purine DNA polymerases Franklin
Chapter 13 - Concept Mapping Using the terms and phrases provided below, complete the concept map showing the discovery of DNA structure. amount of base pairs five-carbon sugar purine DNA polymerases Franklin
Nucleic Acids: DNA and RNA
 Nucleic Acids: DNA and RNA Living organisms are complex systems. Hundreds of thousands of proteins exist inside each one of us to help carry out our daily functions. These proteins are produced locally,
Nucleic Acids: DNA and RNA Living organisms are complex systems. Hundreds of thousands of proteins exist inside each one of us to help carry out our daily functions. These proteins are produced locally,
What Are the Chemical Structures and Functions of Nucleic Acids?
 THE NUCLEIC ACIDS What Are the Chemical Structures and Functions of Nucleic Acids? Nucleic acids are polymers specialized for the storage, transmission, and use of genetic information. DNA = deoxyribonucleic
THE NUCLEIC ACIDS What Are the Chemical Structures and Functions of Nucleic Acids? Nucleic acids are polymers specialized for the storage, transmission, and use of genetic information. DNA = deoxyribonucleic
BIO 101 : The genetic code and the central dogma
 BIO 101 : The genetic code and the central dogma NAME Objectives The purpose of this exploration is to... 1. design experiments to decipher the genetic code; 2. visualize the process of protein synthesis;
BIO 101 : The genetic code and the central dogma NAME Objectives The purpose of this exploration is to... 1. design experiments to decipher the genetic code; 2. visualize the process of protein synthesis;
Name 10 Molecular Biology of the Gene Test Date Study Guide You must know: The structure of DNA. The major steps to replication.
 Name 10 Molecular Biology of the Gene Test Date Study Guide You must know: The structure of DNA. The major steps to replication. The difference between replication, transcription, and translation. How
Name 10 Molecular Biology of the Gene Test Date Study Guide You must know: The structure of DNA. The major steps to replication. The difference between replication, transcription, and translation. How
Chapter 3 Nucleic Acids, Proteins, and Enzymes
 3 Nucleic Acids, Proteins, and Enzymes Chapter 3 Nucleic Acids, Proteins, and Enzymes Key Concepts 3.1 Nucleic Acids Are Informational Macromolecules 3.2 Proteins Are Polymers with Important Structural
3 Nucleic Acids, Proteins, and Enzymes Chapter 3 Nucleic Acids, Proteins, and Enzymes Key Concepts 3.1 Nucleic Acids Are Informational Macromolecules 3.2 Proteins Are Polymers with Important Structural
GENETIC ALGORITHMS. Narra Priyanka. K.Naga Sowjanya. Vasavi College of Engineering. Ibrahimbahg,Hyderabad.
 GENETIC ALGORITHMS Narra Priyanka K.Naga Sowjanya Vasavi College of Engineering. Ibrahimbahg,Hyderabad mynameissowji@yahoo.com priyankanarra@yahoo.com Abstract Genetic algorithms are a part of evolutionary
GENETIC ALGORITHMS Narra Priyanka K.Naga Sowjanya Vasavi College of Engineering. Ibrahimbahg,Hyderabad mynameissowji@yahoo.com priyankanarra@yahoo.com Abstract Genetic algorithms are a part of evolutionary
Immune Programming. Payman Samadi. Supervisor: Dr. Majid Ahmadi. March Department of Electrical & Computer Engineering University of Windsor
 Immune Programming Payman Samadi Supervisor: Dr. Majid Ahmadi March 2006 Department of Electrical & Computer Engineering University of Windsor OUTLINE Introduction Biological Immune System Artificial Immune
Immune Programming Payman Samadi Supervisor: Dr. Majid Ahmadi March 2006 Department of Electrical & Computer Engineering University of Windsor OUTLINE Introduction Biological Immune System Artificial Immune
By the end of today, you will have an answer to: How can 1 strand of DNA serve as a template for replication?
 Name: Period: Date: KIPP NYC College Prep Genetics and Biotech UNIT 9: Introduction to DNA Lecture 4: DNA Modeling and Intro to Replication By the end of today, you will have an answer to: How can 1 strand
Name: Period: Date: KIPP NYC College Prep Genetics and Biotech UNIT 9: Introduction to DNA Lecture 4: DNA Modeling and Intro to Replication By the end of today, you will have an answer to: How can 1 strand
Clustering over DNA Strings
 Proceedings of the 5th WSEAS Int. Conf. on COMPUTATIONAL INTELLIGENCE, MAN-MACHINE SYSTEMS AND CYBERNETICS, Venice, Italy, November 20-22, 2006 184 NURIA GOMEZ BLAS Clustering over DNA Strings EUGENIO
Proceedings of the 5th WSEAS Int. Conf. on COMPUTATIONAL INTELLIGENCE, MAN-MACHINE SYSTEMS AND CYBERNETICS, Venice, Italy, November 20-22, 2006 184 NURIA GOMEZ BLAS Clustering over DNA Strings EUGENIO
DNA. translation. base pairing rules for DNA Replication. thymine. cytosine. amino acids. The building blocks of proteins are?
 2 strands, has the 5-carbon sugar deoxyribose, and has the nitrogen base Thymine. The actual process of assembling the proteins on the ribosome is called? DNA translation Adenine pairs with Thymine, Thymine
2 strands, has the 5-carbon sugar deoxyribose, and has the nitrogen base Thymine. The actual process of assembling the proteins on the ribosome is called? DNA translation Adenine pairs with Thymine, Thymine
DNA vs. RNA B-4.1. Compare DNA and RNA in terms of structure, nucleotides and base pairs.
 DNA vs. RNA B-4.1 Compare DNA and RNA in terms of structure, nucleotides and base pairs. Key Concepts l Nucleic Acids: l deoxyribonucleic acid (DNA) l ribonucleic acid (RNA) l Nucleotides: l nitrogen base,
DNA vs. RNA B-4.1 Compare DNA and RNA in terms of structure, nucleotides and base pairs. Key Concepts l Nucleic Acids: l deoxyribonucleic acid (DNA) l ribonucleic acid (RNA) l Nucleotides: l nitrogen base,
Changing Mutation Operator of Genetic Algorithms for optimizing Multiple Sequence Alignment
 International Journal of Information and Computation Technology. ISSN 0974-2239 Volume 3, Number 11 (2013), pp. 1155-1160 International Research Publications House http://www. irphouse.com /ijict.htm Changing
International Journal of Information and Computation Technology. ISSN 0974-2239 Volume 3, Number 11 (2013), pp. 1155-1160 International Research Publications House http://www. irphouse.com /ijict.htm Changing
Deterministic Crowding, Recombination And Self-Similarity
 Deterministic Crowding, Recombination And Self-Similarity Bo Yuan School of Information Technology and Electrical Engineering The University of Queensland Brisbane, Queensland 4072 Australia E-mail: s4002283@student.uq.edu.au
Deterministic Crowding, Recombination And Self-Similarity Bo Yuan School of Information Technology and Electrical Engineering The University of Queensland Brisbane, Queensland 4072 Australia E-mail: s4002283@student.uq.edu.au
SOLVING TRAVELLING SALESMAN PROBLEM IN A SIMULATION OF GENETIC ALGORITHMS WITH DNA. Angel Goñi Moreno
 International Journal "Information Theories & Applications" Vol.15 / 2008 357 SOLVING TRAVELLING SALESMAN PROBLEM IN A SIMULATION OF GENETIC ALGORITHMS WITH DNA Angel Goñi Moreno Abstract: In this paper
International Journal "Information Theories & Applications" Vol.15 / 2008 357 SOLVING TRAVELLING SALESMAN PROBLEM IN A SIMULATION OF GENETIC ALGORITHMS WITH DNA Angel Goñi Moreno Abstract: In this paper
chapter 12 DNA and RNA Biology Mr. Hines
 chapter 12 DNA and RNA Biology Mr. Hines Transformation What is transformation? Process in which one strain of bacteria is changed by a gene or genes from another strain of bacteria. 12.1 DNA Remember
chapter 12 DNA and RNA Biology Mr. Hines Transformation What is transformation? Process in which one strain of bacteria is changed by a gene or genes from another strain of bacteria. 12.1 DNA Remember
Evolutionary Algorithms and Simulated Annealing in the Topological Configuration of the Spanning Tree
 Evolutionary Algorithms and Simulated Annealing in the Topological Configuration of the Spanning Tree A. SADEGHEIH Department of Industrial Engineering University of Yazd, P.O.Box: 89195-741 IRAN, YAZD
Evolutionary Algorithms and Simulated Annealing in the Topological Configuration of the Spanning Tree A. SADEGHEIH Department of Industrial Engineering University of Yazd, P.O.Box: 89195-741 IRAN, YAZD
A Genetic Algorithm on Inventory Routing Problem
 A Genetic Algorithm on Inventory Routing Problem Artvin Çoruh University e-mail: nevin.aydin@gmail.com Volume 3 No 3 (2014) ISSN 2158-8708 (online) DOI 10.5195/emaj.2014.31 http://emaj.pitt.edu Abstract
A Genetic Algorithm on Inventory Routing Problem Artvin Çoruh University e-mail: nevin.aydin@gmail.com Volume 3 No 3 (2014) ISSN 2158-8708 (online) DOI 10.5195/emaj.2014.31 http://emaj.pitt.edu Abstract
DNA STRUCTURE & REPLICATION
 DNA STRUCTURE & REPLICATION A MODEL OF DNA In 1953, two scientists named Watson & Crick built a model of DNA that demonstrates its exact structure and function. They called this model a double helix, which
DNA STRUCTURE & REPLICATION A MODEL OF DNA In 1953, two scientists named Watson & Crick built a model of DNA that demonstrates its exact structure and function. They called this model a double helix, which
A New DNA Implementation of Finite State Machines *
 International Journal of Computer Science & Applications Vol. III, No. I, pp. 51-60 2006 Technomathematics Research Foundation A New DNA Implementation of Finite State Machines * A. Nowzari-Dalini,1,2,
International Journal of Computer Science & Applications Vol. III, No. I, pp. 51-60 2006 Technomathematics Research Foundation A New DNA Implementation of Finite State Machines * A. Nowzari-Dalini,1,2,
MINIMIZE THE MAKESPAN FOR JOB SHOP SCHEDULING PROBLEM USING ARTIFICIAL IMMUNE SYSTEM APPROACH
 MINIMIZE THE MAKESPAN FOR JOB SHOP SCHEDULING PROBLEM USING ARTIFICIAL IMMUNE SYSTEM APPROACH AHMAD SHAHRIZAL MUHAMAD, 1 SAFAAI DERIS, 2 ZALMIYAH ZAKARIA 1 Professor, Faculty of Computing, Universiti Teknologi
MINIMIZE THE MAKESPAN FOR JOB SHOP SCHEDULING PROBLEM USING ARTIFICIAL IMMUNE SYSTEM APPROACH AHMAD SHAHRIZAL MUHAMAD, 1 SAFAAI DERIS, 2 ZALMIYAH ZAKARIA 1 Professor, Faculty of Computing, Universiti Teknologi
Intelligent DNA Chips: Logical Operation of Gene Expression Profiles on DNA Computers
 Genome Informatics 11: 33 42 (2000) 33 Intelligent DNA Chips: Logical Operation of Gene Expression Profiles on DNA Computers Yasubumi Sakakibara 1 Akira Suyama 2 yasu@j.dendai.ac.jp suyama@dna.c.u-tokyo.ac.jp
Genome Informatics 11: 33 42 (2000) 33 Intelligent DNA Chips: Logical Operation of Gene Expression Profiles on DNA Computers Yasubumi Sakakibara 1 Akira Suyama 2 yasu@j.dendai.ac.jp suyama@dna.c.u-tokyo.ac.jp
COMBINED-OBJECTIVE OPTIMIZATION IN IDENTICAL PARALLEL MACHINE SCHEDULING PROBLEM USING PSO
 COMBINED-OBJECTIVE OPTIMIZATION IN IDENTICAL PARALLEL MACHINE SCHEDULING PROBLEM USING PSO Bathrinath S. 1, Saravanasankar S. 1 and Ponnambalam SG. 2 1 Department of Mechanical Engineering, Kalasalingam
COMBINED-OBJECTIVE OPTIMIZATION IN IDENTICAL PARALLEL MACHINE SCHEDULING PROBLEM USING PSO Bathrinath S. 1, Saravanasankar S. 1 and Ponnambalam SG. 2 1 Department of Mechanical Engineering, Kalasalingam
Storage Allocation and Yard Trucks Scheduling in Container Terminals Using a Genetic Algorithm Approach
 Storage Allocation and Yard Trucks Scheduling in Container Terminals Using a Genetic Algorithm Approach Z.X. Wang, Felix T.S. Chan, and S.H. Chung Abstract Storage allocation and yard trucks scheduling
Storage Allocation and Yard Trucks Scheduling in Container Terminals Using a Genetic Algorithm Approach Z.X. Wang, Felix T.S. Chan, and S.H. Chung Abstract Storage allocation and yard trucks scheduling
Bio-inspired Models of Computation. An Introduction
 Bio-inspired Models of Computation An Introduction Introduction (1) Natural Computing is the study of models of computation inspired by the functioning of biological systems Natural Computing is not Bioinformatics
Bio-inspired Models of Computation An Introduction Introduction (1) Natural Computing is the study of models of computation inspired by the functioning of biological systems Natural Computing is not Bioinformatics
AN ALGORITHM FOR REMOTE SENSING IMAGE CLASSIFICATION BASED ON ARTIFICIAL IMMUNE B CELL NETWORK
 AN ALGORITHM FOR REMOTE SENSING IMAGE CLASSIFICATION BASED ON ARTIFICIAL IMMUNE B CELL NETWORK Shizhen Xu a, *, Yundong Wu b, c a Insitute of Surveying and Mapping, Information Engineering University 66
AN ALGORITHM FOR REMOTE SENSING IMAGE CLASSIFICATION BASED ON ARTIFICIAL IMMUNE B CELL NETWORK Shizhen Xu a, *, Yundong Wu b, c a Insitute of Surveying and Mapping, Information Engineering University 66
TIMETABLING EXPERIMENTS USING GENETIC ALGORITHMS. Liviu Lalescu, Costin Badica
 TIMETABLING EXPERIMENTS USING GENETIC ALGORITHMS Liviu Lalescu, Costin Badica University of Craiova, Faculty of Control, Computers and Electronics Software Engineering Department, str.tehnicii, 5, Craiova,
TIMETABLING EXPERIMENTS USING GENETIC ALGORITHMS Liviu Lalescu, Costin Badica University of Craiova, Faculty of Control, Computers and Electronics Software Engineering Department, str.tehnicii, 5, Craiova,
Multi-Layer Data Encryption using Residue Number System in DNA Sequence
 International Journal of Computer Applications (975 8887) Multi-Layer Data Encryption using Residue Number System in DNA Sequence M. I. Youssef Faculty Of Engineering, Department Of Electrical Engineering
International Journal of Computer Applications (975 8887) Multi-Layer Data Encryption using Residue Number System in DNA Sequence M. I. Youssef Faculty Of Engineering, Department Of Electrical Engineering
An Efficient and Effective Immune Based Classifier
 Journal of Computer Science 7 (2): 148-153, 2011 ISSN 1549-3636 2011 Science Publications An Efficient and Effective Immune Based Classifier Shahram Golzari, Shyamala Doraisamy, Md Nasir Sulaiman and Nur
Journal of Computer Science 7 (2): 148-153, 2011 ISSN 1549-3636 2011 Science Publications An Efficient and Effective Immune Based Classifier Shahram Golzari, Shyamala Doraisamy, Md Nasir Sulaiman and Nur
CEng 713 Evolutionary Computation, Lecture Notes
 CEng 713 Evolutionary Computation, Lecture Notes Introduction to Evolutionary Computation Evolutionary Computation Elements of Evolution: Reproduction Random variation Competition Selection of contending
CEng 713 Evolutionary Computation, Lecture Notes Introduction to Evolutionary Computation Evolutionary Computation Elements of Evolution: Reproduction Random variation Competition Selection of contending
Click here to read the case study about protein synthesis.
 Click here to read the case study about protein synthesis. Big Question: How do cells use the genetic information stored in DNA to make millions of different proteins the body needs? Key Concept: Genetics
Click here to read the case study about protein synthesis. Big Question: How do cells use the genetic information stored in DNA to make millions of different proteins the body needs? Key Concept: Genetics
How do we know what the structure and function of DNA is? - Double helix, base pairs, sugar, and phosphate - Stores genetic information
 DNA: CH 13 How do we know what the structure and function of DNA is? - Double helix, base pairs, sugar, and phosphate - Stores genetic information Discovering DNA s Function 1928: Frederick Griffith studied
DNA: CH 13 How do we know what the structure and function of DNA is? - Double helix, base pairs, sugar, and phosphate - Stores genetic information Discovering DNA s Function 1928: Frederick Griffith studied
DNA and RNA. Gene Composition. Gene Composition Introduction to DNA
 DNA and RNA 12.1 Introduction to DNA Gene Composition We know now that genes dictate characteristics of organisms. But what is it about the genes that produce this control? Not until the late 1920s did
DNA and RNA 12.1 Introduction to DNA Gene Composition We know now that genes dictate characteristics of organisms. But what is it about the genes that produce this control? Not until the late 1920s did
Nucleic Acids, Proteins, and Enzymes
 3 Nucleic Acids, Proteins, and Enzymes Chapter 3 Nucleic Acids, Proteins, and Enzymes Key Concepts 3.1 Nucleic Acids Are Informational Macromolecules 3.2 Proteins Are Polymers with Important Structural
3 Nucleic Acids, Proteins, and Enzymes Chapter 3 Nucleic Acids, Proteins, and Enzymes Key Concepts 3.1 Nucleic Acids Are Informational Macromolecules 3.2 Proteins Are Polymers with Important Structural
Chapter 9: DNA: The Molecule of Heredity
 Chapter 9: DNA: The Molecule of Heredity What is DNA? Answer: Molecule that carries the blueprint of life General Features: DNA is packages in chromosomes (DNA + Proteins) Gene = Functional segment of
Chapter 9: DNA: The Molecule of Heredity What is DNA? Answer: Molecule that carries the blueprint of life General Features: DNA is packages in chromosomes (DNA + Proteins) Gene = Functional segment of
Review of ORGANIC CHEMISTRY
 Nucleic Acids: DNA Review of ORGANIC CHEMISTRY Definition: Contains CARBON (C) and Hydrogen (H) Large polymers can be made of smaller individual monomers. Ex: For carbohydrates, polysaccharides are large
Nucleic Acids: DNA Review of ORGANIC CHEMISTRY Definition: Contains CARBON (C) and Hydrogen (H) Large polymers can be made of smaller individual monomers. Ex: For carbohydrates, polysaccharides are large
DNA - DEOXYRIBONUCLEIC ACID
 DNA - DEOXYRIBONUCLEIC ACID blueprint of life (has the instructions for making an organism) established by James Watson and Francis Crick codes for your genes shape of a double helix made of repeating
DNA - DEOXYRIBONUCLEIC ACID blueprint of life (has the instructions for making an organism) established by James Watson and Francis Crick codes for your genes shape of a double helix made of repeating
Intelligent Techniques Lesson 4 (Examples about Genetic Algorithm)
 Intelligent Techniques Lesson 4 (Examples about Genetic Algorithm) Numerical Example A simple example will help us to understand how a GA works. Let us find the maximum value of the function (15x - x 2
Intelligent Techniques Lesson 4 (Examples about Genetic Algorithm) Numerical Example A simple example will help us to understand how a GA works. Let us find the maximum value of the function (15x - x 2
sensors ISSN
 Sensors 2010, 10, 330-341; doi:10.3390/s100100330 OPEN ACCESS sensors ISSN 1424-8220 www.mdpi.com/journal/sensors Article An Evolution Based Biosensor Receptor DNA Sequence Generation Algorithm Eungyeong
Sensors 2010, 10, 330-341; doi:10.3390/s100100330 OPEN ACCESS sensors ISSN 1424-8220 www.mdpi.com/journal/sensors Article An Evolution Based Biosensor Receptor DNA Sequence Generation Algorithm Eungyeong
Name Class Date. Information and Heredity, Cellular Basis of Life Q: What is the structure of DNA, and how does it function in genetic inheritance?
 12 DNA Big idea Information and Heredity, Cellular Basis of Life Q: What is the structure of DNA, and how does it function in genetic inheritance? WHAT I KNOW WHAT I LEARNED 12.1 How did scientists determine
12 DNA Big idea Information and Heredity, Cellular Basis of Life Q: What is the structure of DNA, and how does it function in genetic inheritance? WHAT I KNOW WHAT I LEARNED 12.1 How did scientists determine
On DNA Computers Controlling Gene Expression Levels
 Proceedings of the 44th IEEE Conference on Decision and Control, and the European Control Conference 2005 Seville, Spain, December 2-5, 2005 MoC.5 On DNA Computers Controlling Gene Expression Levels Olgica
Proceedings of the 44th IEEE Conference on Decision and Control, and the European Control Conference 2005 Seville, Spain, December 2-5, 2005 MoC.5 On DNA Computers Controlling Gene Expression Levels Olgica
The Metaphor. Individuals living in that environment Individual s degree of adaptation to its surrounding environment
 Genetic Algorithms Sesi 14 Optimization Techniques Mathematical Programming Network Analysis Branch & Bound Simulated Annealing Tabu Search Classes of Search Techniques Calculus Base Techniqes Fibonacci
Genetic Algorithms Sesi 14 Optimization Techniques Mathematical Programming Network Analysis Branch & Bound Simulated Annealing Tabu Search Classes of Search Techniques Calculus Base Techniqes Fibonacci
Intro. ANN & Fuzzy Systems. Lecture 36 GENETIC ALGORITHM (1)
 Lecture 36 GENETIC ALGORITHM (1) Outline What is a Genetic Algorithm? An Example Components of a Genetic Algorithm Representation of gene Selection Criteria Reproduction Rules Cross-over Mutation Potential
Lecture 36 GENETIC ALGORITHM (1) Outline What is a Genetic Algorithm? An Example Components of a Genetic Algorithm Representation of gene Selection Criteria Reproduction Rules Cross-over Mutation Potential
Molecular Genetics I DNA
 Molecular Genetics I DNA Deoxyribonucleic acid is the molecule that encodes the characteristics of living things. It is the molecule that is passed from a mother cell to daughter cells, and the molecule
Molecular Genetics I DNA Deoxyribonucleic acid is the molecule that encodes the characteristics of living things. It is the molecule that is passed from a mother cell to daughter cells, and the molecule
Genetic Algorithms in Matrix Representation and Its Application in Synthetic Data
 Genetic Algorithms in Matrix Representation and Its Application in Synthetic Data Yingrui Chen *, Mark Elliot ** and Joe Sakshaug *** * ** University of Manchester, yingrui.chen@manchester.ac.uk University
Genetic Algorithms in Matrix Representation and Its Application in Synthetic Data Yingrui Chen *, Mark Elliot ** and Joe Sakshaug *** * ** University of Manchester, yingrui.chen@manchester.ac.uk University
Journal of Chemical and Pharmaceutical Research, 2014, 6(5): Research Article
 Available online www.jocpr.com Journal of Chemical and Pharmaceutical Research, 2014, 6(5): 693-697 Research Article ISSN : 0975-7384 CODEN(USA) : JCPRC5 Study on multi-resource constraints vehicle scheduling
Available online www.jocpr.com Journal of Chemical and Pharmaceutical Research, 2014, 6(5): 693-697 Research Article ISSN : 0975-7384 CODEN(USA) : JCPRC5 Study on multi-resource constraints vehicle scheduling
Chapter 6. Genes and DNA. Table of Contents. Section 1 What Does DNA Look Like? Section 2 How DNA Works
 Genes and DNA Table of Contents Section 1 What Does DNA Look Like? Section 1 What Does DNA Look Like? Objectives List three important events that led to understanding the structure of DNA. Describe the
Genes and DNA Table of Contents Section 1 What Does DNA Look Like? Section 1 What Does DNA Look Like? Objectives List three important events that led to understanding the structure of DNA. Describe the
PowerPoint Notes on Chapter 9 - DNA: The Genetic Material
 PowerPoint Notes on Chapter 9 - DNA: The Genetic Material Section 1 Identifying the Genetic Material Objectives Relate Griffith s conclusions to the observations he made during the transformation experiments.
PowerPoint Notes on Chapter 9 - DNA: The Genetic Material Section 1 Identifying the Genetic Material Objectives Relate Griffith s conclusions to the observations he made during the transformation experiments.
AGENDA for 10/11/13 AGENDA: HOMEWORK: Due end of the period OBJECTIVES:
 AGENDA for 10/11/13 AGENDA: 1. Finish 1.2.3 DNA Analysis Analyzing DNA Samples Using Current Forensic Methods OBJECTIVES: 1. Demonstrate the steps of gel electrophoresis 2. Analyze restriction fragment
AGENDA for 10/11/13 AGENDA: 1. Finish 1.2.3 DNA Analysis Analyzing DNA Samples Using Current Forensic Methods OBJECTIVES: 1. Demonstrate the steps of gel electrophoresis 2. Analyze restriction fragment
Structural Bioinformatics (C3210) DNA and RNA Structure
 Structural Bioinformatics (C3210) DNA and RNA Structure Importance of DNA/RNA 3D Structure Nucleic acids are essential materials found in all living organisms. Their main function is to maintain and transmit
Structural Bioinformatics (C3210) DNA and RNA Structure Importance of DNA/RNA 3D Structure Nucleic acids are essential materials found in all living organisms. Their main function is to maintain and transmit
Chapter 8 DNA STRUCTURE AND CHROMOSOMAL ORGANIZATION
 Chapter 8 DNA STRUCTURE AND CHROMOSOMAL ORGANIZATION Chapter Summary Even though DNA has been known as a biochemical compound for over 100 years, it was not implicated as the carrier of hereditary information
Chapter 8 DNA STRUCTURE AND CHROMOSOMAL ORGANIZATION Chapter Summary Even though DNA has been known as a biochemical compound for over 100 years, it was not implicated as the carrier of hereditary information
Nucleic acids. What important polymer is located in the nucleus? is the instructions for making a cell's.
 Nucleic acids DNA - The Double Helix Recall that the nucleus is a small spherical, dense body in a cell. It is often called the "control center" because it controls all the activities of the cell including
Nucleic acids DNA - The Double Helix Recall that the nucleus is a small spherical, dense body in a cell. It is often called the "control center" because it controls all the activities of the cell including
VISHVESHWARAIAH TECHNOLOGICAL UNIVERSITY S.D.M COLLEGE OF ENGINEERING AND TECHNOLOGY. A seminar report on GENETIC ALGORITHMS.
 VISHVESHWARAIAH TECHNOLOGICAL UNIVERSITY S.D.M COLLEGE OF ENGINEERING AND TECHNOLOGY A seminar report on GENETIC ALGORITHMS Submitted by Pranesh S S 2SD06CS061 8 th semester DEPARTMENT OF COMPUTER SCIENCE
VISHVESHWARAIAH TECHNOLOGICAL UNIVERSITY S.D.M COLLEGE OF ENGINEERING AND TECHNOLOGY A seminar report on GENETIC ALGORITHMS Submitted by Pranesh S S 2SD06CS061 8 th semester DEPARTMENT OF COMPUTER SCIENCE
Genetic Algorithms and Sensitivity Analysis in Production Planning Optimization
 Genetic Algorithms and Sensitivity Analysis in Production Planning Optimization CECÍLIA REIS 1,2, LEONARDO PAIVA 2, JORGE MOUTINHO 2, VIRIATO M. MARQUES 1,3 1 GECAD Knowledge Engineering and Decision Support
Genetic Algorithms and Sensitivity Analysis in Production Planning Optimization CECÍLIA REIS 1,2, LEONARDO PAIVA 2, JORGE MOUTINHO 2, VIRIATO M. MARQUES 1,3 1 GECAD Knowledge Engineering and Decision Support
The discovery that DNA is the genetic code involved many experiments.
 Section 1: The discovery that DNA is the genetic code involved many experiments. K What I Know W What I Want to Find Out L What I Learned Vocabulary Review nucleic acid New double helix nucleosome Discovery
Section 1: The discovery that DNA is the genetic code involved many experiments. K What I Know W What I Want to Find Out L What I Learned Vocabulary Review nucleic acid New double helix nucleosome Discovery
What is Evolutionary Computation? Genetic Algorithms. Components of Evolutionary Computing. The Argument. When changes occur...
 What is Evolutionary Computation? Genetic Algorithms Russell & Norvig, Cha. 4.3 An abstraction from the theory of biological evolution that is used to create optimization procedures or methodologies, usually
What is Evolutionary Computation? Genetic Algorithms Russell & Norvig, Cha. 4.3 An abstraction from the theory of biological evolution that is used to create optimization procedures or methodologies, usually
601 CTGTCCACACAATCTGCCCTTTCGAAAGATCCCAACGAAAAGAGAGACCACATGGTCCTT GACAGGTGTGTTAGACGGGAAAGCTTTCTAGGGTTGCTTTTCTCTCTGGTGTACCAGGAA >>>>>>>>>>>>>>>>>>
 BIO450 Primer Design Tutorial The most critical step in your PCR experiment will be designing your oligonucleotide primers. Poor primers could result in little or even no PCR product. Alternatively, they
BIO450 Primer Design Tutorial The most critical step in your PCR experiment will be designing your oligonucleotide primers. Poor primers could result in little or even no PCR product. Alternatively, they
Applying Computational Intelligence in Software Testing
 www.stmjournals.com Applying Computational Intelligence in Software Testing Saumya Dixit*, Pradeep Tomar School of Information and Communication Technology, Gautam Buddha University, Greater Noida, India
www.stmjournals.com Applying Computational Intelligence in Software Testing Saumya Dixit*, Pradeep Tomar School of Information and Communication Technology, Gautam Buddha University, Greater Noida, India
Telemedicine Cryptography using DNA Sequence
 Telemedicine Cryptography using DNA Sequence Ruby Chandrakar 1, Suman Kumar Swarnkar 2 1,2 Bharti College of Engineering Technology,Durg Abstract: Today innovative methods of Data hiding are the most evolving
Telemedicine Cryptography using DNA Sequence Ruby Chandrakar 1, Suman Kumar Swarnkar 2 1,2 Bharti College of Engineering Technology,Durg Abstract: Today innovative methods of Data hiding are the most evolving
A nucleotide consists of: an inorganic phosphate group (attached to carbon 5 of the sugar) a 5C sugar (pentose) a Nitrogenous (N containing) base
 Nucleic Acids! Nucleic acids are found in all living cells and viruses and the two main types are DNA and RNA. They are macromolecules made of chains of nucleotides bonded together. They carry genetic
Nucleic Acids! Nucleic acids are found in all living cells and viruses and the two main types are DNA and RNA. They are macromolecules made of chains of nucleotides bonded together. They carry genetic
AGENDA for 10/10/13 AGENDA: HOMEWORK: Due end of the period OBJECTIVES: Due Fri, 10-11
 AGENDA for 10/10/13 AGENDA: 1. 1.2.3 DNA Analysis Analyzing DNA Samples Using Current Forensic Methods OBJECTIVES: 1. Demonstrate the steps of gel electrophoresis 2. Analyze restriction fragment length
AGENDA for 10/10/13 AGENDA: 1. 1.2.3 DNA Analysis Analyzing DNA Samples Using Current Forensic Methods OBJECTIVES: 1. Demonstrate the steps of gel electrophoresis 2. Analyze restriction fragment length
Structure and Replication
 Structure and Replication 6.A: Students will identify components of DNA, and describe how information for specifying traits of an organism is carried in the DNA 6.B: Students will recognize that components
Structure and Replication 6.A: Students will identify components of DNA, and describe how information for specifying traits of an organism is carried in the DNA 6.B: Students will recognize that components
Some types of Mutagenesis
 Mutagenesis What Is a Mutation? Genetic information is encoded by the sequence of the nucleotide bases in DNA of the gene. The four nucleotides are: adenine (A), thymine (T), guanine (G), and cytosine
Mutagenesis What Is a Mutation? Genetic information is encoded by the sequence of the nucleotide bases in DNA of the gene. The four nucleotides are: adenine (A), thymine (T), guanine (G), and cytosine
DNA RNA PROTEIN SYNTHESIS -NOTES-
 DNA RNA PROTEIN SYNTHESIS -NOTES- THE COMPONENTS AND STRUCTURE OF DNA DNA is made up of units called nucleotides. Nucleotides are made up of three basic components:, called deoxyribose in DNA In DNA, there
DNA RNA PROTEIN SYNTHESIS -NOTES- THE COMPONENTS AND STRUCTURE OF DNA DNA is made up of units called nucleotides. Nucleotides are made up of three basic components:, called deoxyribose in DNA In DNA, there
Ch 10 Molecular Biology of the Gene
 Ch 10 Molecular Biology of the Gene For Next Week Lab -Hand in questions from 4 and 5 by TUES in my mailbox (Biology Office) -Do questions for Lab 6 for next week -Lab practical next week Lecture Read
Ch 10 Molecular Biology of the Gene For Next Week Lab -Hand in questions from 4 and 5 by TUES in my mailbox (Biology Office) -Do questions for Lab 6 for next week -Lab practical next week Lecture Read
THE COMPONENTS & STRUCTURE OF DNA
 THE COMPONENTS & STRUCTURE OF DNA - How do genes work? - What are they made of, and how do they determine the characteristics of organisms? - Are genes single molecules, or are they longer structures made
THE COMPONENTS & STRUCTURE OF DNA - How do genes work? - What are they made of, and how do they determine the characteristics of organisms? - Are genes single molecules, or are they longer structures made
Journal of Global Research in Computer Science PREMATURE CONVERGENCE AND GENETIC ALGORITHM UNDER OPERATING SYSTEM PROCESS SCHEDULING PROBLEM
 Volume, No. 5, December 00 Journal of Global Research in Computer Science RESEARCH PAPER Available Online at www.jgrcs.info PREMATURE CONVERGENCE AND GENETIC ALGORITHM UNDER OPERATING SYSTEM PROCESS SCHEDULING
Volume, No. 5, December 00 Journal of Global Research in Computer Science RESEARCH PAPER Available Online at www.jgrcs.info PREMATURE CONVERGENCE AND GENETIC ALGORITHM UNDER OPERATING SYSTEM PROCESS SCHEDULING
Nucleic acids and protein synthesis
 THE FUNCTIONS OF DNA Nucleic acids and protein synthesis The full name of DNA is deoxyribonucleic acid. Every nucleotide has the same sugar molecule and phosphate group, but each nucleotide contains one
THE FUNCTIONS OF DNA Nucleic acids and protein synthesis The full name of DNA is deoxyribonucleic acid. Every nucleotide has the same sugar molecule and phosphate group, but each nucleotide contains one
Chapter 10. DNA: The Molecule of Heredity. Lectures by Gregory Ahearn. University of North Florida. Copyright 2009 Pearson Education, Inc.
 Chapter 10 DNA: The Molecule of Heredity Lectures by Gregory Ahearn University of North Florida Copyright 2009 Pearson Education, Inc. 10.1 What Is The Structure Of DNA? Deoxyribonucleic acid (DNA) is
Chapter 10 DNA: The Molecule of Heredity Lectures by Gregory Ahearn University of North Florida Copyright 2009 Pearson Education, Inc. 10.1 What Is The Structure Of DNA? Deoxyribonucleic acid (DNA) is
BIOB111 - Tutorial activity for Session 13
 BIOB111 - Tutorial activity for Session 13 General topics for week 7 Session 13: Types of nucleic acids, DNA replication Useful links: 1. Visit this website and use its menu to locate information and practice
BIOB111 - Tutorial activity for Session 13 General topics for week 7 Session 13: Types of nucleic acids, DNA replication Useful links: 1. Visit this website and use its menu to locate information and practice
AN IMPROVED ALGORITHM FOR MULTIPLE SEQUENCE ALIGNMENT OF PROTEIN SEQUENCES USING GENETIC ALGORITHM
 AN IMPROVED ALGORITHM FOR MULTIPLE SEQUENCE ALIGNMENT OF PROTEIN SEQUENCES USING GENETIC ALGORITHM Manish Kumar Department of Computer Science and Engineering, Indian School of Mines, Dhanbad-826004, Jharkhand,
AN IMPROVED ALGORITHM FOR MULTIPLE SEQUENCE ALIGNMENT OF PROTEIN SEQUENCES USING GENETIC ALGORITHM Manish Kumar Department of Computer Science and Engineering, Indian School of Mines, Dhanbad-826004, Jharkhand,
CHAPTER 4, Part 1: LECTURE TOPICS: DNA and RNA - MOLECULES OF HEREDITY
 Chapter 4 Notes: Part 1 Biochemistry 461 Fall 2010 CHAPTER 4, Part 1: LECTURE TOPICS: DNA and RNA - MOLECULES OF HEREDITY 1) DNA/RNA structures, nomenclature, shorthand conventions 2) DNA and RNA as genetic
Chapter 4 Notes: Part 1 Biochemistry 461 Fall 2010 CHAPTER 4, Part 1: LECTURE TOPICS: DNA and RNA - MOLECULES OF HEREDITY 1) DNA/RNA structures, nomenclature, shorthand conventions 2) DNA and RNA as genetic
DNA Structure and Function. Chapter 13
 DNA Structure and Function Chapter 13 Impacts, Issues Here Kitty, Kitty, Kitty, Kitty, Kitty Clones made from adult cells have problems; the cell s DNA must be reprogrammed to function like the DNA of
DNA Structure and Function Chapter 13 Impacts, Issues Here Kitty, Kitty, Kitty, Kitty, Kitty Clones made from adult cells have problems; the cell s DNA must be reprogrammed to function like the DNA of
Genetic Algorithm with Upgrading Operator
 Genetic Algorithm with Upgrading Operator NIDAPAN SUREERATTANAN Computer Science and Information Management, School of Advanced Technologies, Asian Institute of Technology, P.O. Box 4, Klong Luang, Pathumthani
Genetic Algorithm with Upgrading Operator NIDAPAN SUREERATTANAN Computer Science and Information Management, School of Advanced Technologies, Asian Institute of Technology, P.O. Box 4, Klong Luang, Pathumthani
Frederick Griffith. Dead Smooth Bacteria. Live Smooth Bacteria. Live Rough Bacteria. Live R+ dead S Bacteria
 Frederick Griffith Live Smooth Bacteria Live Rough Bacteria Dead Smooth Bacteria Live R+ dead S Bacteria Live Smooth Bacteria Frederick Griffith Live Rough Bacteria Dead Smooth Bacteria Live R+ dead S
Frederick Griffith Live Smooth Bacteria Live Rough Bacteria Dead Smooth Bacteria Live R+ dead S Bacteria Live Smooth Bacteria Frederick Griffith Live Rough Bacteria Dead Smooth Bacteria Live R+ dead S
Feature Selection of Gene Expression Data for Cancer Classification: A Review
 Available online at www.sciencedirect.com ScienceDirect Procedia Computer Science 50 (2015 ) 52 57 2nd International Symposium on Big Data and Cloud Computing (ISBCC 15) Feature Selection of Gene Expression
Available online at www.sciencedirect.com ScienceDirect Procedia Computer Science 50 (2015 ) 52 57 2nd International Symposium on Big Data and Cloud Computing (ISBCC 15) Feature Selection of Gene Expression
DNA, Fantastic! View it at Glenn Wolkenfeld 2012
 DNA, Fantastic! View it at www.sciencemusicvideos.com Glenn Wolkenfeld 2012 Welcome, I m so happy you came by For a lesson bout the essence of b-i-o-l-o-g-y DNA s the topic; it s so fantastic, We re talking
DNA, Fantastic! View it at www.sciencemusicvideos.com Glenn Wolkenfeld 2012 Welcome, I m so happy you came by For a lesson bout the essence of b-i-o-l-o-g-y DNA s the topic; it s so fantastic, We re talking
STUDY GUIDE SECTION 10-1 Discovery of DNA
 STUDY GUIDE SECTION 10-1 Discovery of DNA Name Period Date Multiple Choice-Write the correct letter in the blank. 1. The virulent strain of the bacterium S. pneumoniae causes disease because it a. has
STUDY GUIDE SECTION 10-1 Discovery of DNA Name Period Date Multiple Choice-Write the correct letter in the blank. 1. The virulent strain of the bacterium S. pneumoniae causes disease because it a. has
A HYBRID MODERN AND CLASSICAL ALGORITHM FOR INDONESIAN ELECTRICITY DEMAND FORECASTING
 A HYBRID MODERN AND CLASSICAL ALGORITHM FOR INDONESIAN ELECTRICITY DEMAND FORECASTING Wahab Musa Department of Electrical Engineering, Universitas Negeri Gorontalo, Kota Gorontalo, Indonesia E-Mail: wmusa@ung.ac.id
A HYBRID MODERN AND CLASSICAL ALGORITHM FOR INDONESIAN ELECTRICITY DEMAND FORECASTING Wahab Musa Department of Electrical Engineering, Universitas Negeri Gorontalo, Kota Gorontalo, Indonesia E-Mail: wmusa@ung.ac.id
Adv Biology: DNA and RNA Study Guide
 Adv Biology: DNA and RNA Study Guide Chapter 12 Vocabulary -Notes What experiments led up to the discovery of DNA being the hereditary material? o The discovery that DNA is the genetic code involved many
Adv Biology: DNA and RNA Study Guide Chapter 12 Vocabulary -Notes What experiments led up to the discovery of DNA being the hereditary material? o The discovery that DNA is the genetic code involved many
DNA: The Molecule of Heredity
 DNA: The Molecule of Heredity STRUCTURE AND FUNCTION - a nucleic acid o C, H, O, N, P o Made of nucleotides = smaller subunits o Components of nucleotides: Deoxyribose (simple sugar) Phosphate group Nitrogen
DNA: The Molecule of Heredity STRUCTURE AND FUNCTION - a nucleic acid o C, H, O, N, P o Made of nucleotides = smaller subunits o Components of nucleotides: Deoxyribose (simple sugar) Phosphate group Nitrogen
THE CELLULAR AND MOLECULAR BASIS OF INHERITANCE
 Umm AL Qura University THE CELLULAR AND MOLECULAR BASIS OF INHERITANCE Dr. Neda Bogari www.bogari.net EMERY'S ELEMENTS OF MEDICAL GENETICS Peter Turnpenny and Sian Ellard 13 th edition 2008 COURSE SYLLABUS
Umm AL Qura University THE CELLULAR AND MOLECULAR BASIS OF INHERITANCE Dr. Neda Bogari www.bogari.net EMERY'S ELEMENTS OF MEDICAL GENETICS Peter Turnpenny and Sian Ellard 13 th edition 2008 COURSE SYLLABUS
DNA Structure and Replication, and Virus Structure and Replication Test Review
 DNA Structure and Replication, and Virus Structure and Replication Test Review What does DNA stand for? Deoxyribonucleic Acid DNA is what type of macromolecule? DNA is a nucleic acid The building blocks
DNA Structure and Replication, and Virus Structure and Replication Test Review What does DNA stand for? Deoxyribonucleic Acid DNA is what type of macromolecule? DNA is a nucleic acid The building blocks
DNA Chapter 12. DNA and RNA B.1.4, B.1.9, B.1.21, B.1.26, B DNA and RNA B.1.4, B.1.9, B.1.21, B.1.26, B Griffith s Experiment
 DNA Chapter 12 DNA and RNA B.1.4, B.1.9, B.1.21, B.1.26, B.1.27 To truly understand genetics, biologists after Mendel had to discover the chemical nature of the gene. In 1928, Frederick Griffith was trying
DNA Chapter 12 DNA and RNA B.1.4, B.1.9, B.1.21, B.1.26, B.1.27 To truly understand genetics, biologists after Mendel had to discover the chemical nature of the gene. In 1928, Frederick Griffith was trying
Molecular Biology I. The Chemical Nature of DNA. Dr. Obaidur rahman
 Molecular Biology I The Chemical Nature of DNA Dr. Obaidur rahman Characteristics of Genetic Material The coding instructions of all living organisms are written in the same genetic language that of nucleic
Molecular Biology I The Chemical Nature of DNA Dr. Obaidur rahman Characteristics of Genetic Material The coding instructions of all living organisms are written in the same genetic language that of nucleic
