Molecular characterisation of an Australian isolate of Bonamia exitiosa
|
|
|
- Sharlene Sherman
- 6 years ago
- Views:
Transcription
1 DISEASES OF AQUATIC ORGANISMS Vol. 71: 81 85, 2006 Published July 11 Dis Aquat Org Molecular characterisation of an Australian isolate of Bonamia exitiosa Serge Corbeil 1, *, Isabelle Arzul 2, Maeva Robert 2, Franck C. J. Berthe 3, Nathalie Besnard-Cochennec 4, Mark St. J. Crane 1 1 AAHL Fish Diseases Laboratory, Australian Animal Health Laboratory, CSIRO Livestock Industries, Private Bag 24, Geelong, Victoria 3220, Australia 2 IFREMER Laboratoire Génétique et Pathologie, Ronce-les-bains BP 133, La Tremblade, France 3 Department of Pathology & Microbiology, Atlantic Veterinary College, University of Prince Edward Island, Canada 4 IFREMER, BP7004, Taravao, Tahiti, French Polynesia ABSTRACT: An Australian (New South Wales) isolate of Bonamia was characterised at the molecular level by sequencing the 18S-ITS-1 region of the small subunit rrna operon obtained from flat oysters Ostrea angasi shown to be infected by histological examination. Sequence data alignment with homologous genes from 2 other isolates of Bonamia (New Zealand and France) revealed high levels of nucleotide identity with both isolates. However, the Australian Bonamia is shown to be more closely related to the New Zealand isolate, suggesting the existence of an oceanic subgroup of Bonamia. KEY WORDS: Bonamia species Ostrea angasi Flat oyster Small subunit rrna operon Resale or republication not permitted without written consent of the publisher INTRODUCTION Translocation of live molluscs is generally recognised as a major underlying cause of the spread of molluscan pathogens and diseases (Berthe et al. 1999). Bonamiosis, an infectious disease caused by haplosporidian parasites of the genus Bonamia, is one such example; in the northern hemisphere, the disease caused by B. ostreae is believed to have been introduced into Europe via European flat oysters Ostrea edulis imported from California, USA (Cigarría & Elston 1997, Grizel 1997). These studies were based on available documentation of trade and transfer of oyster spat between California and Europe. Subsequent molecular characterisation and sequencing of the small subunit (ssu) rrna operon of Bonamia species confirmed the broad northern hemisphere distribution of B. ostreae (Carnegie et al. 2000, Cochennec et al. 2000). Conversely, since its redesignation as B. exitiosa (Hine et al. 2001, Berthe & Hine 2003), the exact distribution and geographical range of this antipodean Bonamia is unclear (Hine 1996, Campalans et al. 2000, Burreson et al. 2004, Kroeck & Montes 2005), but this information is of central importance to better address management of disease caused by this serious pathogen (OIE 2003). Sequence comparison of the 18S gene and Internal Transcribed Spacer 1 (ITS-1) region of the ssu rrna operon has helped clarify the taxonomic position of these intra-haemocytic parasites and has led to the development of diagnostic methods capable of detecting and distinguishing different isolates resources essential for the development of health surveillance, disease monitoring and oyster stock translocation programmes (Carnegie & Cochennec 2004). A parasite belonging to the genus Bonamia was first reported in Australian flat oysters Ostrea angasi from Port Phillip Bay (Victoria), Georges Bay (Tasmania) and Albany (Western Australia) in the early 1990s (Hine & Jones 1994), with no further certitude regarding species affiliation. Specification of Bonamia sp. is important in order to include or exclude B. exitiosa, as well as exclude Mikrocytos roughleyi, which was redescribed recently as a Bonamia bona fide (Cochen- * serge.corbeil@csiro.au Inter-Research
2 82 Dis Aquat Org 71: 81 85, 2006 nec-laureau et al. 2003). A recent epidemiological study of bonamiosis carried out in southern New South Wales (NSW), Australia (Heasman et al. 2004), has allowed us to identify and characterise a Bonamia isolate at the molecular level and to establish its relatedness to other described Bonamia species. MATERIALS AND METHODS Australian isolate of Bonamia sp. Ethanol (70%)- fixed, Bonamia-infected flat oysters Ostrea angasi, collected from the Merimbula estuary in NSW (during 2003), were provided by Dr. Symon Dworjanyn (NSW Fisheries, Port Stephens) (Heasman et al. 2004). A total of 200 oysters, ranging from 80 to 120 mm in diameter, were sampled from the site, and histological sections were examined by light microscopy. Of several samples confirmed positive, 3 were used for sequencing analysis. DNA extraction. Oyster tissues (mantle and gills) were rinsed in phosphate-buffered saline (PBS; ph 7.4), and then DNA was extracted using a QiaAmp DNA minikit (Qiagen) according to the manufacturer s instructions. DNA was eluted and resuspended in a final volume of 100 µl sterile deionised water. Table 1. Primer sequences used for amplifying and sequencing Bonamia rdna 18S-ITS-1 gene sequences. The positions of the primer sequences correspond to the Australian Bonamia sp. 18S-ITS-1 rrna sequence deposited in GenBank (Accession Number DQ312295) Name Sequence 5-3 Primers position 18S For-A TGCATGTCCAAGTATAAACACG S Rev-A AACGCGATCCGACAAAATA S For-B TCGCGGGAGTGCATATTAG S Rev-B GGATGTGGTAGCCGTTTCTC S For-C TGAGAAACGGCTACCACATC S Rev-C GGGGGCAGATATCCAACTAC S For-D CGACTAAGCATTGGGCTACC S Rev-D GTGGGGTCTCGTCCGTTAT S For-E CGAGACCCCACCCATCTAAC S Rev-E CAAAGGGCAGGGACGTAATC S For-F GCTGTTAAAACGCTCGTAGTTG S Rev-F ATCATTACTCCAGCTCAAAACCA S For-G CGGGCCAGAGGTAAAATTC S Rev-G CCCCCTGAGTCCGAAAAC S For-J GTGCATGGCCGTTCTTAGTC S Rev-J TTGCCCCAATCTTCCATCT S For-M GAATGCGGGTTCGATTC S Rev-M TTGGGTAATTTGCGCACC S For-N CTTCGGCGCCGCCAC S Rev-N GCTTTCGCATAAGTTAG ITS For-7 ATTACGTCCCTGCCCTTTGT ITS Rev-7 ACGAGGTTCGCGGTTTTTAT ITS For-8 CGTAACAAGGTTTCCGTAGGT ITS Rev-8 TGCTTTTTGCGTTTGTGTAGT PCR amplification of the rrna 18S gene and ITS-1 region. Taq DNA polymerase (Promega) was used for PCR assays. In all PCR assays, the primers (Table 1) were used at a final concentration of 1 µm. Aliquots (2 µl) of DNA samples were added to 23 µl of reaction mixture for amplification. Thermocycling conditions for the 18S gene PCR were: 95 C for 2 min (1 cycle), followed by 94 C for 30 s, 55 C for 45 s and 72 C for 45 s (40 cycles), followed by 72 C for 5 min (1 cycle). PCR cycling conditions for the ITS region were: 95 C for 2 min (1 cycle), 94 C for 30 s, 50 C for 45 s and 72 C for 45 s (40 cycles), followed by 72 C for 5 min (1 cycle). The resulting amplicons were resolved by electrophoresis in 2% agarose gels in 1 TAE (0.004 M tris-acetate, M EDTA) buffer. Gels were stained with ethidium bromide (0.5 µg ml 1 ). Amplicons were then extracted using a gel extraction kit (Qiagen) before sequencing. Sequence determination and comparison. The 18S rrna gene and the ITS region (GenBank Accession Number DQ312295) of the Australian Bonamia isolate were determined by direct sequencing of the PCR product using an ABI PRISM Ready Reaction Big dye Termination Cycle Sequencing Kit (Perkin-Elmer) and an ABI PRISM Model 377XL DNA sequencer (Sequencing Analysis software Version 2.6) (Perkin-Elmer). All sequencing was performed on 3 different samples. Twelve pairs of primers were used to amplify overlapping fragments covering the 18S-ITS-1 gene sequences (Table 1). Confirmation of the 5 - end of the 18S gene was performed by cloning the PCR product, amplified by the primers Suni (CAA CCTGGT TGA TCC TGC CC, Besnard-Cochennec 2001) and Boas (Cochennec et al. 2000), into the TA cloning vector (Invitrogen). Sequence determination of the cloned fragments was performed using the plasmid primers Topo F (GAC CAT GAT TAC GCC AAG C) and Topo R (CCC AGT CAC GAC GTT G) and the Big Dye V3 sequencing kit (Applied Biosystem). Bonamia sequences were aligned using the CLUSTAL W algorithm (Thompson et al. 1994, Biomanager, Australian National Genomic Information Service). Nucleotide sequences from Bonamia ostreae and B. exitiosa (Accession Number AF and AF337563, respectively) were obtained from the GenBank database. The B. exitiosa ITS-1 sequence was obtained from Besnard-Cochennec (2001), and the Chilean Bonamia sp. ITS-1 sequence (Accession Number AY539840) was obtained from the GenBank database.
3 Corbeil et al.: Australian Bonamia exitiosa isolate consensus GATTAAGCCATGCATGTCCAAGTATAAACACGTTTGTACTGTGAAACTGCAGATGGCTCATTACAACAGTTATAGTTTATTTGACATTGAACTGTTACAC Bonamia_os...T consensus GGATAACCGTAGTAACCTAGGGCTAATACGTGACAAACCCTGC-TCG-CGGGAGTGCATATTAGCTGAAAACCAACTTTGGTTGAATAATAATATTTGTC Bonamia_sp...A... Bonamia_os...C...G consensus GGATCGCGTTGGCCTCGCCAGCGACATGTCATTCAAGTTTCTGACCTATCAGCTTGACGGTAGGGTATTGGCCTACCGTGGCTTTGACGGGTAACGGGGA Bonamia_sp...G... Bonamia_os...T consensus ATGCGGGTTCGATTCCGGAGAGGCAGCCTGAGAAACGGCTACCACATCCACGGGAGGCAGCAGGTGCGCAAATTACCCAATTCTGACTCAGAGAGGTAGT Bonamia_os consensus GACAAGAAATAACGATCGGCGGCCTTC-GGTTGCCTATTCGGAATGAGAACAATGTAAAAGCCTTATCGAATTCCAGCGGAGGGCAAGCCTGGTGCCAGC Bonamia_os...AT...AA.T...T...C...A consensus AGCCGCGGTAATACCAGCTCCGCTAGCGTATACTAAAGTTGTTGCTGTTAAAACGCTCGTAGTTGGATATCTGCCCCC-GCCCGGGCCGGACTCGC CG Bonamia_ex...C..G Bonamia_sp Bonamia_os...G.G...C...T.GTC.G..A consensus C ---GACGCACCTGCGCCTGCGGCCGGCCGCCGGGGCATAATTCAGGAACGCCGGTCTGGCCATTTAATTGGTCGGGCCGCTGGTCCTGATCCTTTAC Bonamia_ex A.TCGC... Bonamia_sp CG... Bonamia_os C.G...C..A...G...AGA-...C consensus TTTGAGAAAATTAAAGTGCTCAAAGCAGGCTCGCGCCTGAATGCATTAGCATGGAATAATAAGACACGACTTCGGCGCCGCCACTCGTGGCGGGTGTTTT Bonamia_os...T T consensus GTTGGTTTTGAGCTGGAGTAATGATTGATAGAAACAATTGGGGGTGCTAGTATCGCCGGGCCAGAGGTAAAATTCTTTAATTCCGGTGAGACTAACTTAT Bonamia_os..C consensus GCGAAAGCATTCACCAAGCGTGTTTTCTTTAATCAAGAACTAAAGTTGGGGGATCGAAGACGATCAGATACCGTCGTAGTCCCAACCATAAACGATGTCG Bonamia_os...A consensus ACTAAGCATTGGGCTACCAAACTTCCTCAGCACTTTATGAGAAATCAAAGTTTTCGGACTCAGGGGGAAGTATGCTCGCAAGAGTGAAACTTAAAGGAAT Bonamia_os...T...TC consensus TGACGGAAGGGCACCACAAGATGTGGAGCCTGCGGCTTAATTTGATTCAACACGGGAAAACTTACCAGGTCCAGACATAGTAAGGATTGACAGATTAAAG Bonamia_os...T...C consensus TTCTTTCTTGATTCTATGCATGGTGGTGCATGGCCGTTCTTAGTTCCTAGGGTGACCCCTCTGGTTAATTCCGATAACGGACGAGACCCCACCCATCTAA Bonamia_os...C consensus CTAGCCGGCGCTAACCCGGCGCCCAGCGCCCGTTAGCGGGGTGCAGCATTGCGCGCCCGGCTTCTTAGAGGGACTATCTGTGTCTCCAGCAGATGGAAGA Bonamia_os...T.G...A consensus TTGGGGCAATAACAGGTCAGGATGCCCTTAGATGCTCTGGGCTGCACGCGCGCTACAATGATGCGTTCAACGAGTTTGACCCGGCTTGACAAGGCCGGGT Bonamia_os...G consensus AATCTTCAACGCGCATCCAAGTGGGGATAGATGATTGCAATTGTTCATCTTGAACAAGGAATATCTAGTAAACGCAAGTCATCAACTTGCATTGATTACG Bonamia_os...C...T consensus TCCCTGCCCTTTG Fig. 1. Bonamia spp. Alignment of 18S rrna partial gene sequences from Australia Bonamia_ex... Bonamia_sp... (Bonamia sp., Bonamia_sp), New Zealand (B. exitiosa, Bonamia_ex) and France (B. Bonamia_os... ostreae, Bonamia_os). Nucleotide mismatches are indicated below the consensus sequence; dashes represent nucleotide deletions with respect to consensus
4 84 Dis Aquat Org 71: 81 85, consensus TT TTATT T TGCA AATAA AACAT ATAAA AACCGCGAACCT T Bon_NSW TAT..GT...A.-...A-...A...TT CGT.C Bon_NZ TAT..GT...A.-...T-...A...TT CGT.C Bon_Chile --C..TA...G.A...AC..C..C...GC.A...TATA...A.TTC.G Bon_France CAC..CATT.T.TTG..CG-...ATTC...CACA.T consensus Bon_NSW Bon_NZ Bon_Chile Bon_France TTTT ATTAAT TTGCTGACTACCGATTTAC -ACA---CAAACGCA A--AAG AT T...A...T...T...A...C..A.-...A...T...T...A...C..A.-...T.AA...A...AC...AAG...T...T.TAT.T.T..A.T...GCAAACT...G...A...G..A...T...GCGC...G..T.T consensus A ATA G TTTGCA-AGACA TGC-CGACATCAATCTTTGCA Bon_NSW.C...T.ACT Bon_NZ.C...T.ACT Bon_Chile.T...C.CGA...ACA... Bon_France GTGC.A.GAA...GC.A.GTTCT...G.TG..AA.C..CCGA.. Fig. 2. Bonamia spp. Alignment of ITS-1 rrna sequences from the Australian Bonamia sp.(bon_nsw), the New Zealand B. exitiosa (Bon_NZ), the Chilean Bonamia sp. (Bon_Chile) and the French B. ostreae (Bon_France). Nucleotide identities are presented on the consensus lines; dashes represent nucleotide deletions RESULTS AND DISCUSSION The aim of this study was to explore the relationship, at a molecular level, of an Australian isolate of Bonamia sp. with previously described species of the genus, B. ostreae and B. exitiosa. In order to evaluate the relatedness of this Australian isolate to isolates from New Zealand and France, sequencing of the 18S gene and ITS-1 region of the rrna operon of Bonamia sp. was performed. The sequences obtained from the 3 NSW Bonamia isolates were identical. A nucleotide sequence comparison between the isolates from Australia, France and New Zealand was conducted. Results indicated that a higher level of nucleotide identity exists between the Bonamia sp. from Australia and B. exitiosa (New Zealand) than between Bonamia sp. and B. ostreae from France (Fig. 1). A 1598 base pair (bp) sequence of the 18S gene of the Australian isolate presents 4 base pair mismatches and 11 base pair deletions when compared with the B. exitiosa sequence. On the other hand, differences between Bonamia sp. and B. ostreae sequences are more numerous, with 41 base pair mismatches, 5 base pair insertions and 10 base pair deletions. Alignment of the ITS-1 rrna sequences from Bonamia sp. and B. exitiosa presents further evidence of the genetic similarity between the Australian and New Zealand Bonamia. The isolates present only 1 base pair mismatch (Fig. 2). In addition, the ITS-1 sequence of a recently characterised Bonamia isolate from Chile (southern hemisphere) shows a higher level of similarity with the oceanic isolates than B. ostreae from France (northern hemisphere) (Fig. 2). It has been shown that the Chilean flat oyster Ostrea chilensis originates from New Zealand (Ó Foighil et al. 1999); it is therefore logical that their respective Bonamia isolates are closely related. Sequence analysis of the Australian isolate of Bonamia sp. demonstrates closer affinity with B. exitiosa compared to B. ostreae. In the light of this information, it is suggested that this Bonamia isolate from Australia be considered as a member of B. exitiosa. It has been hypothesised (Hine & Jones 1994, Hine 1996) that, in this region, the parasite may have evolved in New Zealand and spread to Australia when commercial-size oysters were shipped live from New Zealand and laid in Victorian and Tasmanian waters prior to consumption (Dartnall 1969). With regards to surveillance of bonamiosis, the genetic similarity of the New Zealand and Australian isolates will allow the development of differential molecular diagnostic tests at various levels of specificity. A PCRrestriction fragment length polymorphism (PCR-RFLP) assay has been proposed to differentiate B. ostreae, B. exitiosa and B. roughleyi (Cochennec-Laureau et al. 2003). With ITS-1 sequences, it should be possible to design specific primers that would allow amplification of a given species in a single-step assay. However, more isolates need to be examined at the molecular level to provide further information on the diversity of Bonamia species before clear delineation of species and possible subspecies is justified. This is particularly true since recent reports of Bonamia sp. outside New
5 Corbeil et al.: Australian Bonamia exitiosa isolate 85 Zealand and Australia have significantly broadened the recognised geographical distribution of this parasite (Campalans et al. 2000, Burreson et al. 2004, Kroeck & Montes 2005). With regards to those recent isolates from Chile, the USA and Argentina, it is believed that the sequence data provided in this paper will help to clarify taxonomic relationships within the genus Bonamia and, more particularly, the species B. exitiosa. If B. exitiosa appears to be widely distributed within this geographical area, then, tools providing sound strain or subspecies identification will assist decision making concerning translocation of live molluscs. Together with robust epidemiological studies and biological evidence of biodiversity, the implementation of such tools could help avert the detrimental affects of translocation of live molluscs. Acknowledgements. This study was partly funded by the Fisheries Research and Development Corporation (Project Number 2003/622) and the European Community Reference Laboratory for Molluscs Diseases. The authors thank S. Dworjanyn and I. Pirozzi (NSW Fisheries) for the collection of wild Ostrea angasi from estuaries in NSW and Jacqueline Campalans (Escuela de Ciencias del Mar, Universidad Catolica de Valparaiso, Chile) for the Chilean Bonamia isolate. We also thank T. Pye (CSIRO Livestock Industries, Australia) for the DNA sequencing. LITERATURE CITED Berthe FCJ, Hine PM (2003) Bonamia exitiosa Hine et al., 2001 is proposed instead of B. exitiosus as the valid name of Bonamia sp. infecting flat oysters Ostrea chilensis in New Zealand. Dis Aquat Org 57:181 Berthe FCJ, Burreson EM, Hine M (1999) Use of molecular tools for mollusc disease diagnosis. Bull Eur Assoc Fish Pathol 19: Besnard-Cochennec N (2001) Bonamia ostreae, parasite de l huître plate, Ostrea edulis: sa position taxonomique parmis les parasites du groupe microcell, analyses des interactions hote/parasite chez plusieurs populations d huîtres plates. These de doctorat, Université de La Rochelle Burreson EM, Stokes NA, Carnegie RB, Bishop MJ (2004) Bonamia sp. (Haplosporidia) found in nonnative oysters Crassostrea ariakensis in Bogue Sound, North Carolina. J Aquat Anim Health 16:1 9 Campalans M, Rojas P, Gonzalez M (2000) Haemocytic parasitosis in the farmed oyster Tiostrea chilensis. Bull Eur Assoc Fish Pathol 20:31 33 Editorial responsibility: Albert K. Sparks, Seattle, Washington, USA Carnegie RB, Cochennec-Laureau N (2004) Microcell parasites of oysters: recent insights and future trends. Aquat Living Resour 17: Carnegie RB, Barber BJ, Culloty SC, Figueras AJ, Distel DL (2000) Development of a PCR assay for the detection of the oyster pathogen Bonamia ostreae and support for its inclusion in the Haplosporidia. Dis Aquat Org 42: Cigarría J, Elston R (1997) Independent introduction of Bonamia ostreae, a parasite of Ostrea edulis, to Spain. Dis Aquat Org 29: Cochennec N, Le Roux F, Berthe FCJ, Gerard A (2000) Detection of Bonamia ostreae based on small subunit ribosomal probe. J Invertebr Pathol 76:26 32 Cochennec-Laureau N, Reece KS, Berthe FCJ, Hine PM (2003) Mikrocytos roughleyi taxonomic affiliation leads to the genus Bonamia (Haplosporidia). Dis Aquat Org 54: Dartnall AJ (1969) New Zealand seastars in Tasmania. Pap Proc R Soc Tasman 103:53 55 Grizel H (1997) Les maladies des mollusques bivalves: risques et prévention. Rev Sci Tech Off Int Epizooties 16: Heasman M, Diggles BK, Hurwood D, Mather P, Pirozzi I, Dworjanyn S (2004) Paving the way for continued rapid development of the flat oyster (Ostrea angasi) farming industry in New South Wales. Final Report to the Department of Transport & Regional Services Project No. NT002/0195 June 2004 NSW Fisheries Final Report Series No. 66. NSW Fisheries, Nelson Bay Hine PM (1996) Southern hemisphere mollusc diseases and an overview of associated risk assessment problems. Rev Sci Tech Off Int Epizooties 15: Hine PM, Jones JB (1994) Bonamia and other aquatic parasites of importance to New Zealand. NZ J Zool 21:49 56 Hine PM, Cochennec-Laureau N, Berthe FCJ (2001) Bonamia exitiosus n. sp. (Haplosporidia) infecting flat oysters Ostrea chilensis (Philippi) in New Zealand. Dis Aquat Org 47:63 72 Kroeck MA, Montes J (2005) Occurrence of the haemocyte parasite Bonamia sp. in flat oysters Ostrea puelchana farmed in San Antonio Bay (Argentina). Dis Aquat Org 63: Ó Foighil D, Marshall BA, Hilbish TJ, Pino MA (1999) Trans- Pacific range extention by rafting is inferred for the flat oyster Ostrea chilensis. Biol Bull (Woods Hole) 196: OIE (Office International des Epizooties) (2003) Bonamiosis. Manual of diagnostic tests for aquatic animals 2003, 4th edn. OIE, Paris. Also available at: normes/fmanual/a_00037.htm Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22: Submitted: December 17, 2005; Accepted: February 20, 2006 Proofs received from author(s): June 16, 2006
RAPPORT FRA HAVFORSKNINGEN
 ISSN 1893-4536 (online) RAPPORT FRA HAVFORSKNINGEN Nr. 11-2016 Feb 2016 www.imr.no ISSN 1893-4536 (online) Health surveillance of the flat oyster populations in Aust-Agder County, southern Norway in the
ISSN 1893-4536 (online) RAPPORT FRA HAVFORSKNINGEN Nr. 11-2016 Feb 2016 www.imr.no ISSN 1893-4536 (online) Health surveillance of the flat oyster populations in Aust-Agder County, southern Norway in the
Detection of Bonamia ostreae in larvae of flat oysters, Ostrea edulis
 Detection of Bonamia ostreae in larvae of flat oysters, Ostrea edulis Isabelle Arzul, Aimé Langlade, Bruno Chollet, Maeva Robert, Emmanuelle Omnes, Sophie Lerond, Yann Couraleau, Jean-Pierre Joly, Cyrille
Detection of Bonamia ostreae in larvae of flat oysters, Ostrea edulis Isabelle Arzul, Aimé Langlade, Bruno Chollet, Maeva Robert, Emmanuelle Omnes, Sophie Lerond, Yann Couraleau, Jean-Pierre Joly, Cyrille
Occurrence of the haemocyte parasite Bonamia sp. in flat oysters Ostrea puelchana farmed in San Antonio Bay (Argentina)
 DISEASES OF AQUATIC ORGANISMS Vol. 63: 231 235, 2005 Published February 28 Dis Aquat Org Occurrence of the haemocyte parasite Bonamia sp. in flat oysters Ostrea puelchana farmed in San Antonio Bay (Argentina)
DISEASES OF AQUATIC ORGANISMS Vol. 63: 231 235, 2005 Published February 28 Dis Aquat Org Occurrence of the haemocyte parasite Bonamia sp. in flat oysters Ostrea puelchana farmed in San Antonio Bay (Argentina)
Perkinsus olseni and P. chesapeaki, sympatric in clams Ruditapes decussatus from Leucate lagoon, France
 www.ifremer.fr Perkinsus olseni and P. chesapeaki, sympatric in clams Ruditapes decussatus from Leucate lagoon, France lfremer I. Arzul, B. Chollet, J. Michel, M. Robert, L. Miossec, J-P. Joly, C. François
www.ifremer.fr Perkinsus olseni and P. chesapeaki, sympatric in clams Ruditapes decussatus from Leucate lagoon, France lfremer I. Arzul, B. Chollet, J. Michel, M. Robert, L. Miossec, J-P. Joly, C. François
Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH).
 Bisulfite Treatment of DNA Dilute DNA sample to 2µg DNA in 50µl ddh 2 O. Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH). Incubate in a 37ºC water bath for 30 minutes. To 55µl samples
Bisulfite Treatment of DNA Dilute DNA sample to 2µg DNA in 50µl ddh 2 O. Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH). Incubate in a 37ºC water bath for 30 minutes. To 55µl samples
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
ISSN (online)
 Nr. 18 2017 R A P P O R T F R A H A V F O R S K N I N G E N ISSN 1893-4536 (online) www.imr.no The surveillance and control programme for bonamiosis and marteiliosis in European flat oysters, Ostrea edulis,
Nr. 18 2017 R A P P O R T F R A H A V F O R S K N I N G E N ISSN 1893-4536 (online) www.imr.no The surveillance and control programme for bonamiosis and marteiliosis in European flat oysters, Ostrea edulis,
Mycobacterium tuberculosis End-Point PCR Kit Product# EP42100
 3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: 866-667-4362 (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com Mycobacterium tuberculosis End-Point PCR Kit Product# EP42100
3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: 866-667-4362 (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com Mycobacterium tuberculosis End-Point PCR Kit Product# EP42100
Identification of a Cucumber mosaic virus Subgroup II Strain Associated with Virus-like Symptoms on Hosta in Ohio
 2013 Plant Management Network. Accepted for publication 18 December 2012. Published. Identification of a Cucumber mosaic virus Subgroup II Strain Associated with Virus-like Symptoms on Hosta in Ohio John
2013 Plant Management Network. Accepted for publication 18 December 2012. Published. Identification of a Cucumber mosaic virus Subgroup II Strain Associated with Virus-like Symptoms on Hosta in Ohio John
Material and methods. Samples
 290 Material and methods Samples Six strains, derived from single females collected in the wild, from different D. gouveai population were analyzed: J79L67 (Ibotirama/BA); J18E2 (Pireno polis/go); H27S1
290 Material and methods Samples Six strains, derived from single females collected in the wild, from different D. gouveai population were analyzed: J79L67 (Ibotirama/BA); J18E2 (Pireno polis/go); H27S1
SuperScript IV Reverse Transcriptase as a better alternative to AMV-based enzymes
 WHITE PAPER SuperScript IV Reverse Transcriptase SuperScript IV Reverse Transcriptase as a better alternative to AMV-based enzymes Abstract Reverse transcriptases (RTs) from avian myeloblastosis virus
WHITE PAPER SuperScript IV Reverse Transcriptase SuperScript IV Reverse Transcriptase as a better alternative to AMV-based enzymes Abstract Reverse transcriptases (RTs) from avian myeloblastosis virus
Mortality and oyster herpesvirus infections in Tomales Bay,California, USA
 Mortality and oyster herpesvirus infections in Tomales Bay,California, USA Colleen A. Burge 1, Robyn M. Estes-Strenge 1, Daniel P. Cheney 2, Frederick J. Griffin 3, Kimberly S. Reece 4, Tristan Renault
Mortality and oyster herpesvirus infections in Tomales Bay,California, USA Colleen A. Burge 1, Robyn M. Estes-Strenge 1, Daniel P. Cheney 2, Frederick J. Griffin 3, Kimberly S. Reece 4, Tristan Renault
SUPPLEMENTARY MATERIAL AND METHODS
 SUPPLEMENTARY MATERIAL AND METHODS Amplification of HEV ORF1, ORF2 and ORF3 genome regions Total RNA was extracted from 200 µl EDTA plasma using Cobas AmpliPrep total nucleic acid isolation kit (Roche,
SUPPLEMENTARY MATERIAL AND METHODS Amplification of HEV ORF1, ORF2 and ORF3 genome regions Total RNA was extracted from 200 µl EDTA plasma using Cobas AmpliPrep total nucleic acid isolation kit (Roche,
Diagnosis and Quantification of Strawberry Vein Banding Virus Using Molecular Approaches
 Diagnosis and Quantification of Strawberry Vein Banding Virus Using Molecular Approaches Ali Mahmoudpour Department of Plant Pathology, University of California, Davis, CA, 95616, USA Current Address:
Diagnosis and Quantification of Strawberry Vein Banding Virus Using Molecular Approaches Ali Mahmoudpour Department of Plant Pathology, University of California, Davis, CA, 95616, USA Current Address:
HiPer RT-PCR Teaching Kit
 HiPer RT-PCR Teaching Kit Product Code: HTBM024 Number of experiments that can be performed: 5 Duration of Experiment: Protocol: 4 hours Agarose Gel Electrophoresis: 45 minutes Storage Instructions: The
HiPer RT-PCR Teaching Kit Product Code: HTBM024 Number of experiments that can be performed: 5 Duration of Experiment: Protocol: 4 hours Agarose Gel Electrophoresis: 45 minutes Storage Instructions: The
Intended Use Norgen s Giardia intestinalis End-Point RT-PCR Kit is designed for the detection of Giardia
 3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: 866-667-4362 (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com Giardia intestinalis End-Point RT-PCR Kit Product# EP43800 Product
3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: 866-667-4362 (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com Giardia intestinalis End-Point RT-PCR Kit Product# EP43800 Product
OIE Reference Laboratory Reports Activities
 OIE Reference Laboratory Reports Activities Activities in 2014 This report has been submitted : 2015-01-19 12:51:35 Name of disease (or topic) for which you are a designated OIE Reference Laboratory: Infection
OIE Reference Laboratory Reports Activities Activities in 2014 This report has been submitted : 2015-01-19 12:51:35 Name of disease (or topic) for which you are a designated OIE Reference Laboratory: Infection
The Expression of Recombinant Sheep Prion Protein (RecShPrPC) and its Detection Using Western Blot and Immuno-PCR
 The Expression of Recombinant Sheep Prion Protein (RecShPrPC) and its Detection Using Western Blot and Immuno-PCR S. Thomas, C. S. Fernando, J. Roach, U. DeSilva and C. A. Mireles DeWitt The objective
The Expression of Recombinant Sheep Prion Protein (RecShPrPC) and its Detection Using Western Blot and Immuno-PCR S. Thomas, C. S. Fernando, J. Roach, U. DeSilva and C. A. Mireles DeWitt The objective
qpcr Kit, DNA-free Product components 100 rxn 250 rxn Product description
 qpcr Kit, DNA-free For the PCR detection and identification of bacterial and fungal DNA using custom primers Product code A8514 Product components 100 rxn 250 rxn A 2.5x mastermix (3 mm MgCl 2 final concentration)
qpcr Kit, DNA-free For the PCR detection and identification of bacterial and fungal DNA using custom primers Product code A8514 Product components 100 rxn 250 rxn A 2.5x mastermix (3 mm MgCl 2 final concentration)
Cold Fusion Cloning Kit. Cat. #s MC100A-1, MC101A-1. User Manual
 Fusion Cloning technology Cold Fusion Cloning Kit Store the master mixture and positive controls at -20 C Store the competent cells at -80 C. (ver. 120909) A limited-use label license covers this product.
Fusion Cloning technology Cold Fusion Cloning Kit Store the master mixture and positive controls at -20 C Store the competent cells at -80 C. (ver. 120909) A limited-use label license covers this product.
FUTURE STRATEGIES FOR A NORWEGIAN PRODUCTION OF SEED OYSTER, Ostrea edulis FOR EXPORT
 FUTURE STRATEGIES FOR A NORWEGIAN PRODUCTION OF SEED OYSTER, Ostrea edulis FOR EXPORT PART REPORT, as an input on the closing of the project Norske setteskjell av flatøsters (Ostrea edulis) til et internasjonalt
FUTURE STRATEGIES FOR A NORWEGIAN PRODUCTION OF SEED OYSTER, Ostrea edulis FOR EXPORT PART REPORT, as an input on the closing of the project Norske setteskjell av flatøsters (Ostrea edulis) til et internasjonalt
Molecular Cell Biology - Problem Drill 11: Recombinant DNA
 Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
HiPer Random Amplification of Polymorphic DNA (RAPD) Teaching Kit
 HiPer Random Amplification of Polymorphic DNA (RAPD) Teaching Kit Product Code: HTBM031 Number of experiments that can be performed: 5 Duration of Experiment: Protocol: 3.5 hours Agarose Gel Electrophoresis:
HiPer Random Amplification of Polymorphic DNA (RAPD) Teaching Kit Product Code: HTBM031 Number of experiments that can be performed: 5 Duration of Experiment: Protocol: 3.5 hours Agarose Gel Electrophoresis:
LaNe RAGE: a new tool for genomic DNA flanking sequence determination
 Electronic Journal of Biotechnology ISSN: 0717-3458 Vol.8 No.2, Issue of August 15, 2005 2005 by Pontificia Universidad Católica de Valparaíso -- Chile Received July 2, 2004 / Accepted March 29, 2005 SHORT
Electronic Journal of Biotechnology ISSN: 0717-3458 Vol.8 No.2, Issue of August 15, 2005 2005 by Pontificia Universidad Católica de Valparaíso -- Chile Received July 2, 2004 / Accepted March 29, 2005 SHORT
MightyAmp DNA Polymerase Ver.3
 Cat. # R076A For Research Use MightyAmp DNA Polymerase Ver.3 Product Manual Table of Contents I. Description... 3 II. Components... 3 III. Storage... 3 IV. General PCR Reaction Mix... 3 V. Primer Design...
Cat. # R076A For Research Use MightyAmp DNA Polymerase Ver.3 Product Manual Table of Contents I. Description... 3 II. Components... 3 III. Storage... 3 IV. General PCR Reaction Mix... 3 V. Primer Design...
FMF NIRCA PROTOCOL STEP 1.
 FMF NIRCA PROTOCOL STEP 1. After you have isolated patient s DNA and DNA from a healthy donor (wild type), you perform a nested PCR. The primers used to amplify exon 2 and exon 10 of the mefv gene are
FMF NIRCA PROTOCOL STEP 1. After you have isolated patient s DNA and DNA from a healthy donor (wild type), you perform a nested PCR. The primers used to amplify exon 2 and exon 10 of the mefv gene are
3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: (905) Fax: (905)
 3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: 866-667-4362 (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com E.coli O157:H7 End-Point PCR Kit Product# EP41300 Product Insert
3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: 866-667-4362 (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com E.coli O157:H7 End-Point PCR Kit Product# EP41300 Product Insert
Table of Contents. Description Kit Components Reagents not supplied in the kit Equipment required Storage...
 Table of Contents Description... 2 Kit Components... 2 Reagents not supplied in the kit... 2 Equipment required... 2 Storage... 2 References... 2 Principle... 3 Protocol... 4 Identification of HPV types...
Table of Contents Description... 2 Kit Components... 2 Reagents not supplied in the kit... 2 Equipment required... 2 Storage... 2 References... 2 Principle... 3 Protocol... 4 Identification of HPV types...
Pasteurella multocida
 BACTOTYPE PCR Amplification Kit Pasteurella multocida Labor Diagnostik Leipzig Manual Technology The product group BACTOTYPE PCR Amplification Kit comprises optimised systems for the identification of
BACTOTYPE PCR Amplification Kit Pasteurella multocida Labor Diagnostik Leipzig Manual Technology The product group BACTOTYPE PCR Amplification Kit comprises optimised systems for the identification of
Guide-it sgrna In Vitro Transcription and Screening Systems User Manual
 Clontech Laboratories, Inc. Guide-it sgrna In Vitro Transcription and Screening Systems User Manual Cat. Nos. 631438, 631439 & 631440 (042114) Clontech Laboratories, Inc. A Takara Bio Company 1290 Terra
Clontech Laboratories, Inc. Guide-it sgrna In Vitro Transcription and Screening Systems User Manual Cat. Nos. 631438, 631439 & 631440 (042114) Clontech Laboratories, Inc. A Takara Bio Company 1290 Terra
Supplemental Data Supplemental Figure 1.
 Supplemental Data Supplemental Figure 1. Silique arrangement in the wild-type, jhs, and complemented lines. Wild-type (WT) (A), the jhs1 mutant (B,C), and the jhs1 mutant complemented with JHS1 (Com) (D)
Supplemental Data Supplemental Figure 1. Silique arrangement in the wild-type, jhs, and complemented lines. Wild-type (WT) (A), the jhs1 mutant (B,C), and the jhs1 mutant complemented with JHS1 (Com) (D)
BIOLOGY Dr.Locke Lecture# 27 An Introduction to Polymerase Chain Reaction (PCR)
 BIOLOGY 207 - Dr.Locke Lecture# 27 An Introduction to Polymerase Chain Reaction (PCR) Required readings and problems: Reading: Open Genetics, Chapter 8.1 Problems: Chapter 8 Optional Griffiths (2008) 9
BIOLOGY 207 - Dr.Locke Lecture# 27 An Introduction to Polymerase Chain Reaction (PCR) Required readings and problems: Reading: Open Genetics, Chapter 8.1 Problems: Chapter 8 Optional Griffiths (2008) 9
Sexing Bovine Preimplantation Embryos by PCR
 Sexing Bovine Preimplantation Embryos by PCR Katherine E.M. Hendricks 1, Leydson F. Martins 2, Justin M. Fear 1 and Peter J. Hansen 1 1 Dept. of Animal Sciences, University of Florida and Department of
Sexing Bovine Preimplantation Embryos by PCR Katherine E.M. Hendricks 1, Leydson F. Martins 2, Justin M. Fear 1 and Peter J. Hansen 1 1 Dept. of Animal Sciences, University of Florida and Department of
i) Wild animals: Those animals that do not live under human supervision or control and do not have their phenotype selected by humans.
 The OIE Validation Recommendations provide detailed information and examples in support of the OIE Validation Standard that is published as Chapter 1.1.6 of the Terrestrial Manual, or Chapter 1.1.2 of
The OIE Validation Recommendations provide detailed information and examples in support of the OIE Validation Standard that is published as Chapter 1.1.6 of the Terrestrial Manual, or Chapter 1.1.2 of
High Pure PCR Template Preparation Kit for preparation of 100 nucleic acid samples Cat. No
 for preparation of 100 nucleic acid samples Cat. No. 1 796 88 Principle Cells are lysed during a short incubation with Proteinase K in the presence of a chaotropic salt (guanidine HCl), which immediately
for preparation of 100 nucleic acid samples Cat. No. 1 796 88 Principle Cells are lysed during a short incubation with Proteinase K in the presence of a chaotropic salt (guanidine HCl), which immediately
Microbial Diversity and Assessment (III) Spring, 2007 Guangyi Wang, Ph.D. POST103B
 Microbial Diversity and Assessment (III) Spring, 2007 Guangyi Wang, Ph.D. POST103B guangyi@hawaii.edu http://www.soest.hawaii.edu/marinefungi/ocn403webpage.htm Overview of Last Lecture Taxonomy (three
Microbial Diversity and Assessment (III) Spring, 2007 Guangyi Wang, Ph.D. POST103B guangyi@hawaii.edu http://www.soest.hawaii.edu/marinefungi/ocn403webpage.htm Overview of Last Lecture Taxonomy (three
XactEdit Cas9 Nuclease with NLS User Manual
 XactEdit Cas9 Nuclease with NLS User Manual An RNA-guided recombinant endonuclease for efficient targeted DNA cleavage Catalog Numbers CE1000-50K, CE1000-50, CE1000-250, CE1001-250, CE1001-1000 Table of
XactEdit Cas9 Nuclease with NLS User Manual An RNA-guided recombinant endonuclease for efficient targeted DNA cleavage Catalog Numbers CE1000-50K, CE1000-50, CE1000-250, CE1001-250, CE1001-1000 Table of
Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC
 Supplementary Appendixes Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC ACG TAG CTC CGG CTG GA-3 for vimentin, /5AmMC6/TCC CTC GCG CGT GGC TTC CGC
Supplementary Appendixes Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC ACG TAG CTC CGG CTG GA-3 for vimentin, /5AmMC6/TCC CTC GCG CGT GGC TTC CGC
Engineering D66N mutant using quick change site directed mutagenesis. Harkewal Singh 09/01/2010
 Engineering D66N mutant using quick change site directed mutagenesis Harkewal Singh 09/01/2010 1 1- What is quick change site directed mutagenesis? 2- An overview of the kit contents. 3- A brief information
Engineering D66N mutant using quick change site directed mutagenesis Harkewal Singh 09/01/2010 1 1- What is quick change site directed mutagenesis? 2- An overview of the kit contents. 3- A brief information
Using Genetics for Species Identification
 Using Genetics for Species Identification John Hyde NOAA Southwest Fisheries Science Center La Jolla, California USA December 6, 2013 2 Important Point to Consider Not all specimens need to be genetically
Using Genetics for Species Identification John Hyde NOAA Southwest Fisheries Science Center La Jolla, California USA December 6, 2013 2 Important Point to Consider Not all specimens need to be genetically
STUDY OF VNTR HUMAN POLYMORPHISMS BY PCR
 STUDY OF VNTR HUMAN POLYMORPHISMS BY PCR Ref. PCR1 1. OBJECTIVE OF THE EXPERIMENT The objective of this experiment is to introduce students to the principles and practice of Polymerase Chain Reaction (PCR)
STUDY OF VNTR HUMAN POLYMORPHISMS BY PCR Ref. PCR1 1. OBJECTIVE OF THE EXPERIMENT The objective of this experiment is to introduce students to the principles and practice of Polymerase Chain Reaction (PCR)
Bacterial 16S rdna PCR Kit Fast (800)
 Cat. # RR182A For Research Use Bacterial 16S rdna PCR Kit Fast (800) Product Manual Table of Contents I. Description... 3 II. Components... 3 III. Materials Required but not Provided... 4 IV. Storage...
Cat. # RR182A For Research Use Bacterial 16S rdna PCR Kit Fast (800) Product Manual Table of Contents I. Description... 3 II. Components... 3 III. Materials Required but not Provided... 4 IV. Storage...
Roche Molecular Biochemicals Technical Note No. LC 10/2000
 Roche Molecular Biochemicals Technical Note No. LC 10/2000 LightCycler Overview of LightCycler Quantification Methods 1. General Introduction Introduction Content Definitions This Technical Note will introduce
Roche Molecular Biochemicals Technical Note No. LC 10/2000 LightCycler Overview of LightCycler Quantification Methods 1. General Introduction Introduction Content Definitions This Technical Note will introduce
Supporting Information. Copyright Wiley-VCH Verlag GmbH & Co. KGaA, Weinheim, 2006
 Supporting Information Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Supporting Information for Expanding the Genetic
Supporting Information Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Supporting Information for Expanding the Genetic
Supplementary Information
 Supplementary Information A general solution for opening double-stranded DNA for isothermal amplification Gangyi Chen, Juan Dong, Yi Yuan, Na Li, Xin Huang, Xin Cui* and Zhuo Tang* Supplementary Materials
Supplementary Information A general solution for opening double-stranded DNA for isothermal amplification Gangyi Chen, Juan Dong, Yi Yuan, Na Li, Xin Huang, Xin Cui* and Zhuo Tang* Supplementary Materials
Multiple choice questions (numbers in brackets indicate the number of correct answers)
 1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
Cat # Box1 Box2. DH5a Competent E. coli cells CCK-20 (20 rxns) 40 µl 40 µl 50 µl x 20 tubes. Choo-Choo Cloning TM Enzyme Mix
 Molecular Cloning Laboratories User Manual Version 3.3 Product name: Choo-Choo Cloning Kits Cat #: CCK-10, CCK-20, CCK-096, CCK-384 Description: Choo-Choo Cloning is a highly efficient directional PCR
Molecular Cloning Laboratories User Manual Version 3.3 Product name: Choo-Choo Cloning Kits Cat #: CCK-10, CCK-20, CCK-096, CCK-384 Description: Choo-Choo Cloning is a highly efficient directional PCR
 PRODUCT INFORMATION Long PCR Enzyme Mix #K0182 500 u Lot Exp. 00.0000 Store at -20 C. CERTIFICATE OF ANALYSIS Long PCR Enzyme Mix is functionally tested in PCR amplification of 47.4 kb fragment from lambda
PRODUCT INFORMATION Long PCR Enzyme Mix #K0182 500 u Lot Exp. 00.0000 Store at -20 C. CERTIFICATE OF ANALYSIS Long PCR Enzyme Mix is functionally tested in PCR amplification of 47.4 kb fragment from lambda
Manipulating DNA. Nucleic acids are chemically different from other macromolecules such as proteins and carbohydrates.
 Lesson Overview 14.3 Studying the Human Genome Nucleic acids are chemically different from other macromolecules such as proteins and carbohydrates. Nucleic acids are chemically different from other macromolecules
Lesson Overview 14.3 Studying the Human Genome Nucleic acids are chemically different from other macromolecules such as proteins and carbohydrates. Nucleic acids are chemically different from other macromolecules
MMLV Reverse Transcriptase 1st-Strand cdna Synthesis Kit
 MMLV Reverse Transcriptase 1st-Strand cdna Synthesis Kit Cat. No. MM070150 Available exclusively thru Lucigen. lucigen.com/epibio www.lucigen.com MA265E MMLV Reverse Transcriptase 1st-Strand cdna Synthesis
MMLV Reverse Transcriptase 1st-Strand cdna Synthesis Kit Cat. No. MM070150 Available exclusively thru Lucigen. lucigen.com/epibio www.lucigen.com MA265E MMLV Reverse Transcriptase 1st-Strand cdna Synthesis
Polymerase Chain Reaction (PCR)
 Laboratory for Environmental Pathogens Research Department of Environmental Sciences University of Toledo Polymerase Chain Reaction (PCR) Background information The polymerase chain reaction (PCR) is an
Laboratory for Environmental Pathogens Research Department of Environmental Sciences University of Toledo Polymerase Chain Reaction (PCR) Background information The polymerase chain reaction (PCR) is an
Supplementary Appendix
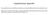 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Fredricks DN, Fiedler TL, Marrazzo JM. Molecular identification
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Fredricks DN, Fiedler TL, Marrazzo JM. Molecular identification
Supplemental Data. The Survival Advantage of Olfaction. in a Competitive Environment. Supplemental Experimental Procedures
 Supplemental Data The Survival Advantage of Olfaction in a Competitive Environment Kenta Asahina, Viktoryia Pavlenkovich, and Leslie B. Vosshall Supplemental Experimental Procedures Experimental animals
Supplemental Data The Survival Advantage of Olfaction in a Competitive Environment Kenta Asahina, Viktoryia Pavlenkovich, and Leslie B. Vosshall Supplemental Experimental Procedures Experimental animals
CHAPTER 3 DEVELOPMENT OF DENV GROUP SPECIFIC REAL TIME RT-PCR
 CHAPTER 3 DEVELOPMENT OF DENV GROUP SPECIFIC REAL TIME RT-PCR 28 Dengue is diagnosed by either detecting virus or antibody to the virus in blood. Isolation of virus in cell culture or in infant mouse brain
CHAPTER 3 DEVELOPMENT OF DENV GROUP SPECIFIC REAL TIME RT-PCR 28 Dengue is diagnosed by either detecting virus or antibody to the virus in blood. Isolation of virus in cell culture or in infant mouse brain
Comparing the Agilent 2100 Bioanalyzer Performance to Traditional DNA Analysis Techniques
 Comparing the Agilent 2100 Bioanalyzer Performance to Traditional DNA Analysis Techniques Application Note Author Deborah Vitale Agilent Technologies, Inc. Palo Alto, CA, USA Abstract This Application
Comparing the Agilent 2100 Bioanalyzer Performance to Traditional DNA Analysis Techniques Application Note Author Deborah Vitale Agilent Technologies, Inc. Palo Alto, CA, USA Abstract This Application
Pinpoint Slide DNA Isolation System Catalog No. D3001
 INSTRUCTIONS Pinpoint Slide DNA Isolation System Catalog No. D3001 Highlights Easily isolates genomic DNA in any targeted microscopic tissue area on a slide. The simple procedure combines Pinpoint tissue
INSTRUCTIONS Pinpoint Slide DNA Isolation System Catalog No. D3001 Highlights Easily isolates genomic DNA in any targeted microscopic tissue area on a slide. The simple procedure combines Pinpoint tissue
Computational Biology I LSM5191
 Computational Biology I LSM5191 Lecture 5 Notes: Genetic manipulation & Molecular Biology techniques Broad Overview of: Enzymatic tools in Molecular Biology Gel electrophoresis Restriction mapping DNA
Computational Biology I LSM5191 Lecture 5 Notes: Genetic manipulation & Molecular Biology techniques Broad Overview of: Enzymatic tools in Molecular Biology Gel electrophoresis Restriction mapping DNA
Guide-it Indel Identification Kit User Manual
 Clontech Laboratories, Inc. Guide-it Indel Identification Kit User Manual Cat. No. 631444 (120114) Clontech Laboratories, Inc. A Takara Bio Company 1290 Terra Bella Avenue, Mountain View, CA 94043, USA
Clontech Laboratories, Inc. Guide-it Indel Identification Kit User Manual Cat. No. 631444 (120114) Clontech Laboratories, Inc. A Takara Bio Company 1290 Terra Bella Avenue, Mountain View, CA 94043, USA
Supporting Online Information
 Supporting Online Information Isolation of Human Genomic DNA Sequences with Expanded Nucleobase Selectivity Preeti Rathi, Sara Maurer, Grzegorz Kubik and Daniel Summerer* Department of Chemistry and Chemical
Supporting Online Information Isolation of Human Genomic DNA Sequences with Expanded Nucleobase Selectivity Preeti Rathi, Sara Maurer, Grzegorz Kubik and Daniel Summerer* Department of Chemistry and Chemical
Genomics and Gene Recognition Genes and Blue Genes
 Genomics and Gene Recognition Genes and Blue Genes November 1, 2004 Prokaryotic Gene Structure prokaryotes are simplest free-living organisms studying prokaryotes can give us a sense what is the minimum
Genomics and Gene Recognition Genes and Blue Genes November 1, 2004 Prokaryotic Gene Structure prokaryotes are simplest free-living organisms studying prokaryotes can give us a sense what is the minimum
Product# EP Product Description
 3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: 866-667-4362 (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com Chlamydia End-Point PCR Kit Product# EP31400 Product Insert Intended
3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: 866-667-4362 (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com Chlamydia End-Point PCR Kit Product# EP31400 Product Insert Intended
PCR Laboratory Exercise
 PCR Laboratory Exercise Advance Protocol (updated 1/2018) Introduction Detection of TPA-25 Alu by PCR A Human DNA Fingerprinting Lab Protocol 1994 Cold Spring Harbor Laboratory DNA Learning Center In this
PCR Laboratory Exercise Advance Protocol (updated 1/2018) Introduction Detection of TPA-25 Alu by PCR A Human DNA Fingerprinting Lab Protocol 1994 Cold Spring Harbor Laboratory DNA Learning Center In this
Fungal rdna (D1/D2) PCR Kit Fast
 Cat. # RR184A For Research Use Fungal rdna (D1/D2) PCR Kit Fast Product Manual Table of Contents I. Description... 3 II. Components... 3 III. Materials Required but not Provided... 4 IV. Storage... 4 V.
Cat. # RR184A For Research Use Fungal rdna (D1/D2) PCR Kit Fast Product Manual Table of Contents I. Description... 3 II. Components... 3 III. Materials Required but not Provided... 4 IV. Storage... 4 V.
NRAS Mutation Analysis Reagents (Codons 12 and 13)
 NRAS Mutation Analysis Reagents (Codons 12 and 13) User Manual V1.1 Cat No. GP18 32 reactions 1 CONTENTS Introduction 4 Overview of Mutector TM Assay 5 Materials Provided 6 Materials Required 7 Equipment
NRAS Mutation Analysis Reagents (Codons 12 and 13) User Manual V1.1 Cat No. GP18 32 reactions 1 CONTENTS Introduction 4 Overview of Mutector TM Assay 5 Materials Provided 6 Materials Required 7 Equipment
International Journal of Pharma and Bio Sciences DIAGNOSIS OF PARASITIC DISEASES IN BIVALVE MOLLUSCS ABSTRACT
 Research Article Biotechnology International Journal of Pharma and Bio Sciences ISSN 0975-6299 DIAGNOSIS OF PARASITIC DISEASES IN BIVALVE MOLLUSCS R.CAROLINE JEBA* ANDS.ANITHA Department of Bio-technology,
Research Article Biotechnology International Journal of Pharma and Bio Sciences ISSN 0975-6299 DIAGNOSIS OF PARASITIC DISEASES IN BIVALVE MOLLUSCS R.CAROLINE JEBA* ANDS.ANITHA Department of Bio-technology,
PROTOCOL FOR MICROSATELLITE GENOTYPING BY UNLINKED MARKERS
 PROTOCOL FOR MICROSATELLITE GENOTYPING BY UNLINKED MARKERS Introduction Molecular genotyping of Plasmodium falciparum using polymorphic markers is commonly used to distinguish among different clones of
PROTOCOL FOR MICROSATELLITE GENOTYPING BY UNLINKED MARKERS Introduction Molecular genotyping of Plasmodium falciparum using polymorphic markers is commonly used to distinguish among different clones of
Annual Meeting of the National Reference Laboratories for Mollusc Diseases
 REPORT OF THE Annual Meeting of the National Reference Laboratories for Mollusc Diseases Nantes, 18-19 March 2008 Organised by the Community Reference Laboratory for Mollusc Diseases IFREMER, La Tremblade,
REPORT OF THE Annual Meeting of the National Reference Laboratories for Mollusc Diseases Nantes, 18-19 March 2008 Organised by the Community Reference Laboratory for Mollusc Diseases IFREMER, La Tremblade,
Archimer
 Please note that this is an author-produced PDF of an article accepted for publication following peer review. The definitive publisher-authenticated version is available on the publisher Web site Veterinary
Please note that this is an author-produced PDF of an article accepted for publication following peer review. The definitive publisher-authenticated version is available on the publisher Web site Veterinary
Amplicon Sequencing Template Preparation
 Amplicon Sequencing Template Preparation The DNA sample preparation procedure for Amplicon Sequencing consists of a simple PCR amplification reaction, but uses special Fusion Primers (Figure 1-1). The
Amplicon Sequencing Template Preparation The DNA sample preparation procedure for Amplicon Sequencing consists of a simple PCR amplification reaction, but uses special Fusion Primers (Figure 1-1). The
16S rdna Sequencing Diagnosis of Spirochetemia in Lyme and related Borrelioses
 16S rdna Sequencing Diagnosis of Spirochetemia in Lyme and related Borrelioses Sin Hang Lee, MD, F.R.C.P.(C)* Pathologist, Milford Hospital and Director, Milford Medical Laboratory, Inc. Milford, CT Email:
16S rdna Sequencing Diagnosis of Spirochetemia in Lyme and related Borrelioses Sin Hang Lee, MD, F.R.C.P.(C)* Pathologist, Milford Hospital and Director, Milford Medical Laboratory, Inc. Milford, CT Email:
inventory of Narragansett Bay
 Genetic barcoding of Green Algae Ulva sp. for algal inventory of Narragansett Bay Benjamin Gibson Abstract: Through the use of DNA isolation, PCR amplification, plasmid cloning and plasmid purification,
Genetic barcoding of Green Algae Ulva sp. for algal inventory of Narragansett Bay Benjamin Gibson Abstract: Through the use of DNA isolation, PCR amplification, plasmid cloning and plasmid purification,
Supplemental Materials. DNA preparation. Dehalogenimonas lykanthroporepellens strain BL-DC-9 T (=ATCC
 Supplemental Materials DNA preparation. Dehalogenimonas lykanthroporepellens strain BL-DC-9 T (=ATCC BAA-1523 = JCM 15061) was grown in defined basal medium amended with 0.5 mm 1,1,2- trichloroethane (1,1,2-TCA)
Supplemental Materials DNA preparation. Dehalogenimonas lykanthroporepellens strain BL-DC-9 T (=ATCC BAA-1523 = JCM 15061) was grown in defined basal medium amended with 0.5 mm 1,1,2- trichloroethane (1,1,2-TCA)
Molecular Techniques Third-year Biology
 PLANNING Genetics Lab practices Molecular Techniques. Genetics Lab practices protocol. 2015-16 PCR-DIRECTED MUTAGENESIS, MOLECULAR CLONING AND RESTRICTION ANALYSIS Sessions 1 & 2 (2x3 hours): PCR-directed
PLANNING Genetics Lab practices Molecular Techniques. Genetics Lab practices protocol. 2015-16 PCR-DIRECTED MUTAGENESIS, MOLECULAR CLONING AND RESTRICTION ANALYSIS Sessions 1 & 2 (2x3 hours): PCR-directed
strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular
![strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular](/thumbs/72/66361212.jpg) Additional file 2 Identification of AOX1 in P. pastoris GS115 with a Mut s phenotype Results and Discussion The HBsAg producing strain was originally identified as a Mut s (methanol utilization slow) strain
Additional file 2 Identification of AOX1 in P. pastoris GS115 with a Mut s phenotype Results and Discussion The HBsAg producing strain was originally identified as a Mut s (methanol utilization slow) strain
OIE Reference Laboratory Reports Activities
 OIE Reference Laboratory Reports Activities Activities in 2016 This report has been submitted : 2017-05-18 07:24:55 Name of disease (or topic) for which you are a designated OIE Reference Laboratory: Infection
OIE Reference Laboratory Reports Activities Activities in 2016 This report has been submitted : 2017-05-18 07:24:55 Name of disease (or topic) for which you are a designated OIE Reference Laboratory: Infection
GeneCopoeia TM. All-in-One qpcr Mix For universal quantitative real-time PCR. User Manual
 GeneCopoeia TM Expressway to Discovery All-in-One qpcr Mix For universal quantitative real-time PCR Cat. No. AOPR-0200 (200 qpcr reactions) Cat. No. AOPR-0600 (600 qpcr reactions) Cat. No. AOPR-1000 (1000
GeneCopoeia TM Expressway to Discovery All-in-One qpcr Mix For universal quantitative real-time PCR Cat. No. AOPR-0200 (200 qpcr reactions) Cat. No. AOPR-0600 (600 qpcr reactions) Cat. No. AOPR-1000 (1000
Contents. 1 Basic Molecular Microbiology of Bacteria... 1 Exp. 1.1 Isolation of Genomic DNA Introduction Principle...
 Contents 1 Basic Molecular Microbiology of Bacteria... 1 Exp. 1.1 Isolation of Genomic DNA... 1 Introduction... 1 Principle... 1 Reagents Required and Their Role... 2 Procedure... 3 Observation... 4 Result
Contents 1 Basic Molecular Microbiology of Bacteria... 1 Exp. 1.1 Isolation of Genomic DNA... 1 Introduction... 1 Principle... 1 Reagents Required and Their Role... 2 Procedure... 3 Observation... 4 Result
Fatchiyah
 Fatchiyah Email: fatchiya@yahoo.co.id RNAs: mrna trna rrna RNAi DNAs: Protein: genome DNA cdna mikro-makro mono-poly single-multi Analysis: Identification human and animal disease Finger printing Sexing
Fatchiyah Email: fatchiya@yahoo.co.id RNAs: mrna trna rrna RNAi DNAs: Protein: genome DNA cdna mikro-makro mono-poly single-multi Analysis: Identification human and animal disease Finger printing Sexing
Amplified Analysis of DNA by the Autonomous Assembly of Polymers Consisting of DNAzyme Wires
 Supporting information for the paper: Amplified Analysis of DNA by the Autonomous Assembly of Polymers Consisting of DNAzyme Wires Fuan Wang, Johann Elbaz, Ron Orbach, Nimrod Magen and Itamar Willner*
Supporting information for the paper: Amplified Analysis of DNA by the Autonomous Assembly of Polymers Consisting of DNAzyme Wires Fuan Wang, Johann Elbaz, Ron Orbach, Nimrod Magen and Itamar Willner*
Lecture Four. Molecular Approaches I: Nucleic Acids
 Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
HELINI White spot Syndrome virus [WSSV] Real-time PCR Kit
![HELINI White spot Syndrome virus [WSSV] Real-time PCR Kit HELINI White spot Syndrome virus [WSSV] Real-time PCR Kit](/thumbs/72/67570787.jpg) HELINI White spot Syndrome virus [WSSV] Real-time PCR Kit Instruction manual Cat. No: 6001-25/50/100 tests Compatible with: Agilent, Bio-Rad, Roche, Applied Bio systems [ABI], Rotor-gene, Cepheid, Bioer,
HELINI White spot Syndrome virus [WSSV] Real-time PCR Kit Instruction manual Cat. No: 6001-25/50/100 tests Compatible with: Agilent, Bio-Rad, Roche, Applied Bio systems [ABI], Rotor-gene, Cepheid, Bioer,
Chapter 20: Biotechnology
 Name Period The AP Biology exam has reached into this chapter for essay questions on a regular basis over the past 15 years. Student responses show that biotechnology is a difficult topic. This chapter
Name Period The AP Biology exam has reached into this chapter for essay questions on a regular basis over the past 15 years. Student responses show that biotechnology is a difficult topic. This chapter
TIANgel Mini DNA Purification Kit
 TIANgel Mini DNA Purification Kit For DNA purification from agarose and polyacrylamide gels www.tiangen.com/en DP130419 TIANgel Mini DNA Purification Kit Kit Contents (Spin column) Cat. no. DP208 Contents
TIANgel Mini DNA Purification Kit For DNA purification from agarose and polyacrylamide gels www.tiangen.com/en DP130419 TIANgel Mini DNA Purification Kit Kit Contents (Spin column) Cat. no. DP208 Contents
Search for and Analysis of Single Nucleotide Polymorphisms (SNPs) in Rice (Oryza sativa, Oryza rufipogon) and Establishment of SNP Markers
 DNA Research 9, 163 171 (2002) Search for and Analysis of Single Nucleotide Polymorphisms (SNPs) in Rice (Oryza sativa, Oryza rufipogon) and Establishment of SNP Markers Shinobu Nasu, Junko Suzuki, Rieko
DNA Research 9, 163 171 (2002) Search for and Analysis of Single Nucleotide Polymorphisms (SNPs) in Rice (Oryza sativa, Oryza rufipogon) and Establishment of SNP Markers Shinobu Nasu, Junko Suzuki, Rieko
TaKaRa PCR Amplification Kit
 Cat. # R011 For Research Use TaKaRa PCR Amplification Kit Product Manual Table of Contents I. Description... 3 II. Components... 3 III. Storage... 4 IV. Materials Required but not Provided... 4 V. Principle...
Cat. # R011 For Research Use TaKaRa PCR Amplification Kit Product Manual Table of Contents I. Description... 3 II. Components... 3 III. Storage... 4 IV. Materials Required but not Provided... 4 V. Principle...
Blue mussels and Marteilia refringens:
 Blue mussels and Marteilia refringens: is it possible to process mussels from non-approved zones in zones with disease-free status with the proper preventive measures? Lone Madsen 1 & Stig Mellergaard
Blue mussels and Marteilia refringens: is it possible to process mussels from non-approved zones in zones with disease-free status with the proper preventive measures? Lone Madsen 1 & Stig Mellergaard
High-throughput cloning and expression in recalcitrant bacteria
 High-throughput cloning and expression in recalcitrant bacteria Eric R Geertsma & Bert Poolman Supplementary text and figures: Supplementary Figure 1 Frequency of SfiI sites yielding identical 3 extensions
High-throughput cloning and expression in recalcitrant bacteria Eric R Geertsma & Bert Poolman Supplementary text and figures: Supplementary Figure 1 Frequency of SfiI sites yielding identical 3 extensions
Molecular analysis of Fomitiporia mediterranea isolates from esca-affected grapevines in southern Italy
 Phytopathol. Mediterr. (2004) 43, 268 272 Molecular analysis of Fomitiporia mediterranea isolates from esca-affected grapevines in southern Italy CLAUDIO CICCARONE 1, ANTONIO GRANITI 2, ANGELA SCHIAFFINO
Phytopathol. Mediterr. (2004) 43, 268 272 Molecular analysis of Fomitiporia mediterranea isolates from esca-affected grapevines in southern Italy CLAUDIO CICCARONE 1, ANTONIO GRANITI 2, ANGELA SCHIAFFINO
NRAS Codon 61 Mutation Analysis Reagents
 NRAS Codon 61 Mutation Analysis Reagents User Manual V1.0 Cat No. GP19 32 reactions 1 CONTENTS Introduction 4 Overview of Mutector TM Assay 5 Materials Provided 6 Materials Required 7 Equipment Required
NRAS Codon 61 Mutation Analysis Reagents User Manual V1.0 Cat No. GP19 32 reactions 1 CONTENTS Introduction 4 Overview of Mutector TM Assay 5 Materials Provided 6 Materials Required 7 Equipment Required
Identification of the subtypes of Verocytotoxin encoding genes (vtx) of Escherichia coli by conventional PCR
 Identification of the subtypes of Verocytotoxin encoding genes (vtx) of Escherichia coli by conventional PCR 1. Aim and field of application Verocytotoxins (VT), synonymous Shiga toxins, are a toxin family
Identification of the subtypes of Verocytotoxin encoding genes (vtx) of Escherichia coli by conventional PCR 1. Aim and field of application Verocytotoxins (VT), synonymous Shiga toxins, are a toxin family
Axygen AxyPrep Magnetic Bead Purification Kits. A Corning Brand
 Axygen AxyPrep Magnetic Bead Purification Kits A Corning Brand D Sample Prep Solutions for Genomics Obtaining Pure Nucleic Acids from Your Sample is Precious The purification of high quality DNA is the
Axygen AxyPrep Magnetic Bead Purification Kits A Corning Brand D Sample Prep Solutions for Genomics Obtaining Pure Nucleic Acids from Your Sample is Precious The purification of high quality DNA is the
Product Name : Simple mirna Detection Kit
 Product Name : Simple mirna Detection Kit Code No. : DS700 This product is for research use only Kit Contents This kit provides sufficient reagents to perform 20 reactions for detecting microrna. Components
Product Name : Simple mirna Detection Kit Code No. : DS700 This product is for research use only Kit Contents This kit provides sufficient reagents to perform 20 reactions for detecting microrna. Components
For in vitro Veterinary Diagnostics only. DNA Extraction and PCR Detection Kit for Pasteurella multocida.
 For in vitro Veterinary Diagnostics only. DNA Extraction and PCR Detection Kit for Pasteurella multocida www.kylt.eu DIRECTION FOR USE Art. No. 31058 / 31059 Kylt Pasteurella multocida DNA Extraction and
For in vitro Veterinary Diagnostics only. DNA Extraction and PCR Detection Kit for Pasteurella multocida www.kylt.eu DIRECTION FOR USE Art. No. 31058 / 31059 Kylt Pasteurella multocida DNA Extraction and
2.5. Equipment and materials supplied by user PCR based template preparation Influence of temperature on in vitro EGFP synthesis 11
 Manual 15 Reactions LEXSY in vitro Translation Cell-free protein expression kit based on Leishmania tarentolae for PCR-based template generation Cat. No. EGE-2010-15 FOR RESEARCH USE ONLY. NOT INTENDED
Manual 15 Reactions LEXSY in vitro Translation Cell-free protein expression kit based on Leishmania tarentolae for PCR-based template generation Cat. No. EGE-2010-15 FOR RESEARCH USE ONLY. NOT INTENDED
1. COMPONENTS. PyroStart Fast PCR Master Mix (2X) (#K0211 for 250 reactions of 20µl) 2. STORAGE 3. DESCRIPTION
 1 2 1 PyroStart Fast PCR Master Mix (2X) (#K0211 for 250 reactions of 20µl) TABLE OF CONTENTS 1. COMPONENTS... 2 2. STORAGE... 2 3. DESCRIPTION... 2 4. PROTOCOL FOR FAST PCR... 3 4.1. General Considerations...
1 2 1 PyroStart Fast PCR Master Mix (2X) (#K0211 for 250 reactions of 20µl) TABLE OF CONTENTS 1. COMPONENTS... 2 2. STORAGE... 2 3. DESCRIPTION... 2 4. PROTOCOL FOR FAST PCR... 3 4.1. General Considerations...
Biotechnology. Chapter 20. Biology Eighth Edition Neil Campbell and Jane Reece. PowerPoint Lecture Presentations for
 Chapter 20 Biotechnology PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from Joan Sharp Copyright
Chapter 20 Biotechnology PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from Joan Sharp Copyright
OIE guidance on zoning and the development of compartments: practical experience and recommendations to the OIE
 OIE guidance on zoning and the development of compartments: practical experience and recommendations to the OIE Dr. Joanne Constantine Dr. Andrea Osborn Riding the Wave, Vietnam 2015 Objective Critical
OIE guidance on zoning and the development of compartments: practical experience and recommendations to the OIE Dr. Joanne Constantine Dr. Andrea Osborn Riding the Wave, Vietnam 2015 Objective Critical
DNA Visualizer Extraction Kit
 DNA Visualizer Extraction Kit Catalog Number D0006 50 reactions Version: 03 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Intended Use... 3 Background... 3 General Information...
DNA Visualizer Extraction Kit Catalog Number D0006 50 reactions Version: 03 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Intended Use... 3 Background... 3 General Information...
The Isolation and Sequence Analysis of the Cryptochrome (Cry1) Gene From a Petunia hybrida Plant. By Jenalyn Quevedo
 The Isolation and Sequence Analysis of the Cryptochrome (Cry1) Gene From a Petunia hybrida Plant By Jenalyn Quevedo Biology 115L June 6, 2005 1 Abstract The blue-light photoreceptor cryptochrome (cry1)
The Isolation and Sequence Analysis of the Cryptochrome (Cry1) Gene From a Petunia hybrida Plant By Jenalyn Quevedo Biology 115L June 6, 2005 1 Abstract The blue-light photoreceptor cryptochrome (cry1)
MgCl 2 (25 mm) 1.6 ml 1.6 ml 1.6 ml 1.6 ml
 Technical Data Sheet KAPA2G Fast PCR Kit Kit components KK 5008 Product codes KK 5010 KK 5009 KK 5011 KAPA2G Fast DNA polymerase (5 U/μl) 100 U 100 U 250 U 250 U 1. Production Description The KAPA2G Fast
Technical Data Sheet KAPA2G Fast PCR Kit Kit components KK 5008 Product codes KK 5010 KK 5009 KK 5011 KAPA2G Fast DNA polymerase (5 U/μl) 100 U 100 U 250 U 250 U 1. Production Description The KAPA2G Fast
