This Technical Note describes these characterization studies in more detail.
|
|
|
- Dwight Wilkerson
- 6 years ago
- Views:
Transcription
1 Technical Note Characterization of SOMAmer Reagents Binding Specificity in the SOMAscan 1.3k Assay Summary Slow Off-Rate Modified Aptamer (SOMAmer) reagents are identified via the SELEX process against epitopes on a single target protein in buffer. However, many proteins share structural and/or functional properties, and thus could be bound by a SOMAmer reagent originally selected against another protein. In order to test the specificity of each SOMAmer reagent used in the SOMAscan 1.3k assay for its initial target protein, we performed a series of characterization steps. These steps included selection of similar proteins via standard in silico methods, obtaining any identified relevant relative proteins (where available), direct experimental SOMAmer binding experiments with those related proteins in buffer, and/or pull-down assays followed by mass spectrometry-based analysis of the protein(s) bound by the SOMAmer reagent from biological matrices. In the current SOMAscan assay 51% of the SOMAmer reagents bind only their original target, and another 9% bind related proteins with >10x weaker affinity than their target protein (24% of the SOMAmer reagents in the menu cannot be tested, as no highly related proteins are available for testing) (See Figure 6). This Technical Note describes these characterization studies in more detail. Introduction Slow Off-Rate Modified Aptamer (SOMAmer) reagents - the protein-specific binding reagents that enable the SOMAscan assay are initially identified via the SELEX process by their ability to strongly bind to a single purified protein. 1 3 Each modified DNA-based SOMAmer selected for the SOMAscan assay has a unique three-dimensional folded structure that binds a specific structural epitope on the surface of its target protein with high affinity. This high-affinity binding between a SOMAmer reagent and its target protein is mediated both by electrostatic contacts contributed by the DNA backbone (similar to unmodified aptamers) as well as hydrophobic contacts enabled by the modified bases in the SOMAmer sequence. 4,5 Because there are many proteins that share structural and functional features, it is possible that the conformational epitope to which a SOMAmer reagent binds is present on other proteins related to the original protein used to select the SOMAmer reagent. Indeed, we have observed that a minority of SOMAmer reagents - are able to bind with some degree of affinity to highly similar proteins, presumably through such a shared structural epitope, although not always with the same high affinity. Because the SOMAscan assay is performed in a complex biological sample containing thousands of different proteins, experimentally determining which reagents may target a shared epitope within a protein species may be extremely valuable in interpreting biomarker discovery data derived from the SOMAscan assay. 1
2 As a first step, we used publicly available databases of known human protein sequences 6 and sequence alignment tools (e. g., BLAST) 7 to identify those relevant relative proteins that share significant homology with proteins used to select the SOMAmer reagents in the current version of the SOMAscan assay (SOMAscan 1.3k). Any proteins with significant homology to the SOMAmer target protein (i.e., proteins with greater than 40% sequence identity with the target protein) were obtained for direct experimental testing. It should be noted that the breadth of related proteins that can be tested is limited to those that are available in the SomaLogic inventory or commercially available as full-length proteins from reliable vendors. In cases where a protein was not available or where a sequence homology search returned no significant results (~20% of the 1.3k menu), we confirmed SOMAmer reagent pull-down of the target protein in buffer but performed no additional testing. If related proteins were available, affinity capture experiments similar to immunoprecipitation were performed. SOMAmer reagents were immobilized on streptavidin coated beads and then incubated with either the target protein or the identified related protein. The SOMAmer-protein complexes were then washed and the protein labeled by conjugation of NHS-AlexaFluor TM 647. The complexes were then eluted and the recovery of bound protein vs. input protein was analyzed by SDS-PAGE and fluorescent imaging. When any binding to proteins other than the SELEX target was observed (at 100 nm in buffer), we performed solution affinity measurements to determine whether the SOMAmer reagent has similar or disparate affinities for the target protein and related protein. If the solution K d was within 10-fold of that for the SELEX target, the reagent was reported to bind the SELEX target and other proteins with similar affinity. If the measured affinity differed by greater than 10-fold, we reported that the reagent binds to the protein(s) other than the SELEX target with at least 10-fold weaker affinity. Although this is a broad statement regarding specific affinity, we have chosen not to report exact K d values because of the high variability observed in both the quality and the reported concentrations of commercially obtained purified proteins. We have also identified a small number of SOMAmer reagents (76 out of 1305) that have been flagged for further performance review. For example, the original SELEX target protein may no longer be commercially available, or pulldown results were ambiguous due to protein sample impurity. Specificity characterization results for these SOMAmer reagents will be communicated as soon as unambiguous target binding data are available and reviewed. Another method that we have used to test SOMAmer reagent specificity is to identify, by liquid chromatography and tandem mass spectrometry (LC-MS/MS), the proteins bound by each reagent after incubation with human plasma. We have begun employing this mode of specificity characterization on SOMAmer reagents selected to bind to proteins with high endogenous concentrations in human plasma. We have noted specific enrichment of the intended endogenous target protein in the SOMAscan menu annotation separately from the specificity results generated using purified proteins, when observed. 2
3 Results Highly specific target binding For 73% of cases in which proteins related to the SELEX target were available for testing, we observed binding of the SOMAmer reagent to the specific SELEX target and not to any of the related proteins. For example, SOMAmer was tested against the intended target, tissue inhibitor of metalloproteinase-1 (TIMP-1), and the related proteins TIMP-2 (60% identical), TIMP-3 (31% identical), and TIMP-4 (40% identical) (Figure 1A). Figure 1. High specificity of a TIMP-1 SOMAmer Reagent. A. Binding to purified TIMP-1, but not to TIMP-2, TIMP-3, or TIMP-4 in a pulldown experiment. Pulldown eluate is shown in the lanes labeled Pulldown and a sample of protein input is shown in lanes labeled Input. B. TIMP-1-specific peptide sequences enriched from human plasma using a TIMP-1 SOMAmer reagent. No binding was observed with TIMP-2 or TIMP-4, and very weak binding was observed with TIMP-3 that could not be measured by solution-affinity methods (at protein concentrations up to 100 nm). When this same TIMP-1 SOMAmer reagent was used in affinity enrichment from human plasma, four unique peptides corresponding to endogenous TIMP-1 were identified by LC-MS/MS in the enriched sample and no peptides corresponding to any other member of the TIMP protein family were identified (Figure 1B). Additionally, no peptides corresponding to TIMP-1 were identified in any other plasma pulldown samples performed using 142 different SOMAmer reagents, including a TIMP-2-specific SOMAmer reagent. Another example of highly specific binding is shown in Figure 2, in which a SOMAmer reagent specific for matrix metalloproteinase-10 (MMP-10) does not bind MMP-12 (61% identical), MMP-13 (57% identical), MMP-3 (80% identical), MMP-1 (61% identical), or MMP-8 (50% identical). 3
4 Figure 2. Exclusive binding of a MMP-10 specific SOMAmer reagent to MMP-10 but not to MMP-12, MMP-13, MMP-3, MMP-1, or MMP-8. Pulldown eluate is shown in the lanes labeled Pulldown and a sample of protein input is shown in lanes labeled Input. Binding to shared epitopes with disparate affinity In any instance where we observed binding to proteins other than the SELEX target (20% of the 1,305 reagents) in initial pulldown tests, we followed up with measurements of solution affinity. We typically measure the association of radiolabeled SOMAmer reagent with protein and then capture the complex using a protein-affinity chromatography medium. Saturation binding curves are then generated by titrating increasing amounts of protein in the presence of a constant, limiting amount of SOMAmer reagent. The Kd is determined to be the protein concentration at which the half maximal binding is observed. For example, initial pulldown tests indicated that SOMAmer binds not only to the SELEX target pyrophosphatase 1 (PPA1), but also the related protein PPA2, which shares 68% amino acid sequence identity (Figure 3A). Solution affinity measurements determined that the SOMAmer affinity was greater than 10-fold stronger for PPA1 than for PPA2 (Figure 3B). 4
5 Figure 3. PPA1-specific SOMAmer reagent binds to PPA1 with greater than 10-fold stronger affinity than PPA2. A. PPA1 SOMAmer reagent binds to purified PPA1 and PPA2 in a pulldown assay. Pulldown eluate is shown in the lanes labeled Pulldown and a sample of protein input is shown in lanes labeled Input. B. PPA1 SOMAmer reagent binds to PPA1 with greater than 10x stronger affinity (K d = 0.7 nm) than PPA2 (K d = > 100 nm). Protein concentration is shown in M units on the x-axis and the fraction of reagent bound, as measured by radioactivity, is shown on the y-axis. Binding to shared epitopes with similar affinity We observed that 10% of the 1,305 reagents bound to members of a protein family with highly similar affinities. As previously noted, this recognition most often occurs when proteins share extensive sequence identity. Presumably, the binding epitope to which the SOMAmer reagent was selected is highly conserved and biochemically indistinguishable by solution equilibrium binding affinities. Because the reagents in the SOMAscan assay are typically selected only against a single target protein in the absence of protein competitor strategies, it is not surprising that certain reagents bind to epitopes shared amongst a highly related family of proteins. For example, SOMAmer binds to the SELEX target calcium/calmodulin dependent protein kinase II delta (CAMK2D) as well as the related proteins CAMK2A (91% identical) and CAMK2B (87% identical) (Figure 4A). Solution affinity comparisons determined that this reagent has a similar binding affinity, of approximately 2 nm, for all three proteins (Figure 4B). 5
6 Figure 4. SOMAmer reagent selected against CAMK2D binds to CAMK2A and CAMK2B with similar affinities. A. CAMK2D SOMAmer reagent binds to purified CAMK2D, CAMK2A, and CAMK2B in a pulldown assay. Pulldown eluate is shown in the lanes labeled Pulldown and a sample of protein input is shown in lanes labeled Input. B. CAMK2D SOMAmer reagent binds to CAMK2D, CAMK2A, and CAMK2B with a similar affinity (Kd = 2 nm). Protein concentration is shown in M units on the x-axis and the fraction of reagent bound, as measured by radioactivity, is shown on the y-axis. In cases where a SOMAscan signal results from a reagent with documented binding to multiple proteins, the SOMAscan end user may use this information in follow-on studies to further dissect the biological pathways being affected. For example, although the analyte name (which was derived from the name of the original SELEX target protein) is listed as CAMK2D, it is possible that the SOMAscan signal is a result of SOMAmer binding to these three proteins separately or cumulatively, depending on their relative concentrations in the sample. Based on these results, we have begun to perform SELEX experiments with competition strategies to select SOMAmer reagents that can distinguish between such highly related family members for use in future SOMAscan assay versions. One such strategy was utilized to develop reagents that could distinguish the 90% identical proteins GDF-11 and GDF-8. The initial reagent selected against GDF-11 binds to both proteins with comparable affinities (K d ~ 100 pm). With such high amino acid sequence identity, this cross-reactivity was not surprising. To identify specific reagents for GDF-11 and GDF-8, special selection conditions were employed in order to compete for sequence binding between GDF- 11 and GDF-8 SOMAmer reagent libraries. The results include several reagents that bind GDF-11 with very high affinity and with minimal, if any, binding to GDF-8. An example of one of the reagents is shown in Figure 5: Binding to GDF-11 is shown in red and binding to GDF-8 is shown in green. Using similar selection-counter selection strategies, we have also identified GDF-8 specific SOMAmer reagents. 6
7 Figure 5. Radiolabeled affinity curves for binding of a GDF-11 specific SOMAmer reagent to GDF-11 (red) and to GDF-8 (green). Protein concentration is shown in M units on the x-axis and the fraction of reagent bound, as measured by radioactivity, is shown on the y-axis. In summary, we were able to test binding to related proteins for 76% of the 1,305 SOMAmer reagents on the current SOMAscan 1.3K menu. We were unable to detect binding to any related proteins for 51% of the menu (Figure 6). When binding to related proteins was detected, about half of these SOMAmer reagents exhibited binding to at least one related protein with similar affinity while the other half bound to related proteins, but with at least 10-fold weaker affinity. Specific target enrichment from human plasma has been confirmed for 115 of the SOMAmer reagents on the current menu. We were either unable to identify or obtain related proteins to test for shared epitope binding for 24% of the SOMAmer reagent menu. Figure 6. Summary of SOMAscan reagents specificity characterization 7
8 Conclusions In our ongoing commitment to improving the quality of the SOMAscan multiplexed proteomic assay, we have cataloged the specificity of all SOMAmer reagents used in the current SOMAscan 1.3K Assay by performing protein sequence homology searches, followed by experimentally comparing the ability of SOMAmer reagents to bind to their cognate versus related protein targets in buffer. SOMAscan assay end users may use these data to determine which, if any, proteins other than the indicated SELEX target could result in a positive signal in the SOMAscan assay. We have also begun to query SOMAmer reagent specificity in a complex biological sample using tandem mass spectrometry. This has resulted in a richer set of characterization criteria for all reagents used in the SOMAscan 1.3K assay. Many of the analytes detected in the SOMAscan assay are expected to be at relatively low concentrations in human plasma, thus making it challenging to purify adequate amounts of these low abundance proteins for peptide identification by LC-MS/MS. Through ongoing method development in affinity-capture as well as exploration of other biological matrices, such as cell lysate, that may have higher endogenous concentrations of certain proteins relative to plasma, we plan to expand our mass spectrometry-based characterization of SOMAmer reagents to cover a greater portion of the SOMAscan menu. References 1. Tuerk, C. & Gold, L. Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science 249, (1990). 2. Ellington, A. D. & Szostak, J. W. In vitro selection of RNA molecules that bind specific ligands. Nature 346, (1990). 3. Ozer, A., Pagano, J. M. & Lis, J. T. New Technologies Provide Quantum Changes in the Scale, Speed, and Success of SELEX Methods and Aptamer Characterization. Mol. Ther. Nucleic Acids 3, e183 (2014). 4. Gelinas, A. D., Davies, D. R. & Janjic, N. Embracing proteins: structural themes in aptamer-protein complexes. Curr. Opin. Struct. Biol. 36, (2016). 5. Jarvis, T. C. et al. Non-helical DNA Triplex Forms a Unique Aptamer Scaffold for High Affinity Recognition of Nerve Growth Factor. Structure 23, (2015). 6. UniProt Consortium, T. U. UniProt: a hub for protein information. Nucleic Acids Res. 43, D (2015). 7. Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, (1990). All trademarks, service marks, trade names and product names are the property of their respective owners, including SomaLogic and SOMAmer, which are registered trademarks of SomaLogic, Inc. Copyright 2016 SomaLogic, Inc. SSM-067 Effective: 1/11/ Wilderness Place Boulder, CO U.S. Technical Support: Main Number:
TECHNICAL NOTE Phosphorylation Site Specific Aptamers for Cancer Biomarker Peptide Enrichment and Detection in Mass Spectrometry Based Proteomics
 TECHNICAL NOTE Phosphorylation Site Specific Aptamers for Cancer Biomarker Peptide Enrichment and Detection in Mass Spectrometry Based Proteomics INTRODUCTION The development of renewable capture reagents
TECHNICAL NOTE Phosphorylation Site Specific Aptamers for Cancer Biomarker Peptide Enrichment and Detection in Mass Spectrometry Based Proteomics INTRODUCTION The development of renewable capture reagents
Characterization of Aptamer Binding using SensíQ SPR Platforms
 Characterization of Aptamer Binding using SensíQ SPR Platforms APPLICATION NOTE INTRODUCTION Aptamers have the potential to provide a better solution in diagnostics and other research areas than traditional
Characterization of Aptamer Binding using SensíQ SPR Platforms APPLICATION NOTE INTRODUCTION Aptamers have the potential to provide a better solution in diagnostics and other research areas than traditional
SOMAscan Proteomic Assay Technical White Paper
 SOMAscan Proteomic Assay Technical White Paper Introduction The SOMAscan assay is a highly multiplexed, sensitive, quantitative, and reproducible proteomic tool for discovering previously undetected biomarkers
SOMAscan Proteomic Assay Technical White Paper Introduction The SOMAscan assay is a highly multiplexed, sensitive, quantitative, and reproducible proteomic tool for discovering previously undetected biomarkers
Technical White Paper SomaLogic, Inc. SSM-002, Rev. 3 Effective: 9/21/2015 DCN Page 1
 Technical White Paper 2015 SomaLogic, Inc. SSM-002, Rev. 3 Effective: 9/21/2015 DCN 15-310 Page 1 SOMAscan Proteomic Assay Technical White Paper Introduction The SOMAscan TM assay is a highly multiplexed,
Technical White Paper 2015 SomaLogic, Inc. SSM-002, Rev. 3 Effective: 9/21/2015 DCN 15-310 Page 1 SOMAscan Proteomic Assay Technical White Paper Introduction The SOMAscan TM assay is a highly multiplexed,
Lecture 8: Affinity Chromatography-III
 Lecture 8: Affinity Chromatography-III Key words: Chromatography; Affinity chromatography; Protein Purification During this lecture, we shall be studying few more examples of affinity chromatography. The
Lecture 8: Affinity Chromatography-III Key words: Chromatography; Affinity chromatography; Protein Purification During this lecture, we shall be studying few more examples of affinity chromatography. The
Proteomics. Manickam Sugumaran. Department of Biology University of Massachusetts Boston, MA 02125
 Proteomics Manickam Sugumaran Department of Biology University of Massachusetts Boston, MA 02125 Genomic studies produced more than 75,000 potential gene sequence targets. (The number may be even higher
Proteomics Manickam Sugumaran Department of Biology University of Massachusetts Boston, MA 02125 Genomic studies produced more than 75,000 potential gene sequence targets. (The number may be even higher
Combinatorial RNA libraries
 SELEX and Artificial Ribozymes Some definitions SELEX: Systematic Evolution of Ligands by EXponential Enrichment (alternatively: in vitro selection, in vitro evolution) Aptamer: nucleic acid ligand (from
SELEX and Artificial Ribozymes Some definitions SELEX: Systematic Evolution of Ligands by EXponential Enrichment (alternatively: in vitro selection, in vitro evolution) Aptamer: nucleic acid ligand (from
Kinetics Review. Tonight at 7 PM Phys 204 We will do two problems on the board (additional ones than in the problem sets)
 Quiz 1 Kinetics Review Tonight at 7 PM Phys 204 We will do two problems on the board (additional ones than in the problem sets) I will post the problems with solutions on Toolkit for those that can t make
Quiz 1 Kinetics Review Tonight at 7 PM Phys 204 We will do two problems on the board (additional ones than in the problem sets) I will post the problems with solutions on Toolkit for those that can t make
Application Note. Authors. Abstract
 Automated, High Precision Tryptic Digestion and SISCAPA-MS Quantification of Human Plasma Proteins Using the Agilent Bravo Automated Liquid Handling Platform Application Note Authors Morteza Razavi, N.
Automated, High Precision Tryptic Digestion and SISCAPA-MS Quantification of Human Plasma Proteins Using the Agilent Bravo Automated Liquid Handling Platform Application Note Authors Morteza Razavi, N.
ENCODE RBP Antibody Characterization Guidelines
 ENCODE RBP Antibody Characterization Guidelines Approved on November 18, 2016 Background An integral part of the ENCODE Project is to characterize the antibodies used in the experiments. This document
ENCODE RBP Antibody Characterization Guidelines Approved on November 18, 2016 Background An integral part of the ENCODE Project is to characterize the antibodies used in the experiments. This document
ProteoMiner Protein Enrichment Technology
 Sample Preparation ProteoMiner Protein Enrichment Technology Digging Deeper in the Proteome Detect More Proteins With ProteoMiner Technology Than With Immunodepletion ProteoMiner Technology vs. an Agilent
Sample Preparation ProteoMiner Protein Enrichment Technology Digging Deeper in the Proteome Detect More Proteins With ProteoMiner Technology Than With Immunodepletion ProteoMiner Technology vs. an Agilent
MagSi Beads. Magnetic Silica Beads for Life Science and Biotechnology study
 MagSi Beads Magnetic Silica Beads for Life Science and Biotechnology study MagnaMedics Diagnostics B.V. / Rev. 9.2 / 2012 Wide range of products for numerous applications MagnaMedics separation solutions
MagSi Beads Magnetic Silica Beads for Life Science and Biotechnology study MagnaMedics Diagnostics B.V. / Rev. 9.2 / 2012 Wide range of products for numerous applications MagnaMedics separation solutions
How to run Alpha assay: How to setup an Alpha assay Make your own assay!
 How to run Alpha assay: How to setup an Alpha assay Make your own assay! 1 2009 PerkinElmer AlphaLISA kits - recommendations before starting the assay Samples: Phenol red and hemoglobin: choose AlphaLISA
How to run Alpha assay: How to setup an Alpha assay Make your own assay! 1 2009 PerkinElmer AlphaLISA kits - recommendations before starting the assay Samples: Phenol red and hemoglobin: choose AlphaLISA
MSD Immuno-Dot-Blot Assays. A division of Meso Scale Diagnostics, LLC.
 MSD Immuno-Dot-Blot Assays Example: High Throughput Western Blots Replacements Traditional Western Blots High content Molecular weight and immunoreactivity Labor and protein intensive Inherently low throughput
MSD Immuno-Dot-Blot Assays Example: High Throughput Western Blots Replacements Traditional Western Blots High content Molecular weight and immunoreactivity Labor and protein intensive Inherently low throughput
Application Note AN001
 Testing hybridoma supernatants with the Spots On Dots Antibody Screening Kit Application Note AN1 Table of Contents Overview... 2 Figure 1. Screening of hybridomas raised against peptide antigens... 3
Testing hybridoma supernatants with the Spots On Dots Antibody Screening Kit Application Note AN1 Table of Contents Overview... 2 Figure 1. Screening of hybridomas raised against peptide antigens... 3
Discovery and Humanization of Novel High Affinity Neutralizing Monoclonal Antibodies to Human IL-17A
 Discovery and Humanization of Novel High Affinity Neutralizing Monoclonal Antibodies to Human IL-17A Contacts: Marty Simonetti martysimonetti@gmail.com Kirby Alton kirby.alton@abeomecorp.com Rick Shimkets
Discovery and Humanization of Novel High Affinity Neutralizing Monoclonal Antibodies to Human IL-17A Contacts: Marty Simonetti martysimonetti@gmail.com Kirby Alton kirby.alton@abeomecorp.com Rick Shimkets
Newsletter Issue 7 One-STrEP Analysis of Protein:Protein-Interactions
 www.iba-biotagnology.com Newsletter Issue 7 One-STrEP Analysis of Protein:Protein-Interactions Strep-tag and One-STrEP-tag PPI Analysis with the co-precipitation/ mass spectrometry approach 3 Background
www.iba-biotagnology.com Newsletter Issue 7 One-STrEP Analysis of Protein:Protein-Interactions Strep-tag and One-STrEP-tag PPI Analysis with the co-precipitation/ mass spectrometry approach 3 Background
IMMUNOPRECIPITATION TROUBLESHOOTING TIPS
 IMMUNOPRECIPITATION TROUBLESHOOTING TIPS Creative Diagnostics Abstract Immunoprecipitation (IP) is the technique of precipitating a protein antigen out of solution using an antibody that specifically binds
IMMUNOPRECIPITATION TROUBLESHOOTING TIPS Creative Diagnostics Abstract Immunoprecipitation (IP) is the technique of precipitating a protein antigen out of solution using an antibody that specifically binds
Introduction to Assay Development
 Introduction to Assay Development A poorly designed assay can derail a drug discovery program before it gets off the ground. While traditional bench-top assays are suitable for basic research and target
Introduction to Assay Development A poorly designed assay can derail a drug discovery program before it gets off the ground. While traditional bench-top assays are suitable for basic research and target
Module I: Introduction
 Module I: Introduction 20.109 Lecture 1 3 February, 2011 Introduction to: Module Overview Fundamental concepts and techniques in molecular biology Appreciating nucleic acids (RNA in particular) as more
Module I: Introduction 20.109 Lecture 1 3 February, 2011 Introduction to: Module Overview Fundamental concepts and techniques in molecular biology Appreciating nucleic acids (RNA in particular) as more
Figure S6. Detection of anti-gfp antibodies in anti-dna and normal plasma without competition DNA--9
 Supplementary Information Ultrasensitive antibody detection by agglutination-pcr (ADAP) Cheng-ting Tsai 1 *, Peter V. Robinson 1 *, Carole A. Spencer 2 and Carolyn R. Bertozzi 3,4ǂ Department of 1 Chemistry,
Supplementary Information Ultrasensitive antibody detection by agglutination-pcr (ADAP) Cheng-ting Tsai 1 *, Peter V. Robinson 1 *, Carole A. Spencer 2 and Carolyn R. Bertozzi 3,4ǂ Department of 1 Chemistry,
Protein Purification Products. Complete Solutions for All of Your Protein Purification Applications
 Protein Purification Products Complete Solutions for All of Your Protein Purification Applications FLAG-Tagged Protein Products EXPRESS with the pcmv-dykddddk Vector Set Fuse your protein of interest to
Protein Purification Products Complete Solutions for All of Your Protein Purification Applications FLAG-Tagged Protein Products EXPRESS with the pcmv-dykddddk Vector Set Fuse your protein of interest to
Supporting Information
 Supporting Information Wiley-VCH 2006 69451 Weinheim, Germany RNA ligands that distinguish metabolite-induced conformations in the TPP riboswitch Günter Mayer, Marie-Sophie L. Raddatz, Julia D. Grunwald,
Supporting Information Wiley-VCH 2006 69451 Weinheim, Germany RNA ligands that distinguish metabolite-induced conformations in the TPP riboswitch Günter Mayer, Marie-Sophie L. Raddatz, Julia D. Grunwald,
Small-Molecule Drug Target Identification/Deconvolution Technologies
 Small-Molecule Drug Target Identification/Deconvolution Technologies Case-Studies Shantani Target ID Technology Tool Box Target Deconvolution is not Trivial = A single Tool / Technology May Not necessarily
Small-Molecule Drug Target Identification/Deconvolution Technologies Case-Studies Shantani Target ID Technology Tool Box Target Deconvolution is not Trivial = A single Tool / Technology May Not necessarily
Application Manual ProteoSpin Abundant Serum Protein Depletion Kit
 Application Manual ProteoSpin Abundant Serum Protein Depletion Kit For Use with P/N 17300 ProteoSpin Abundant Serum Protein Depletion Kit Basic Features Efficient removal of highly abundant proteins 1
Application Manual ProteoSpin Abundant Serum Protein Depletion Kit For Use with P/N 17300 ProteoSpin Abundant Serum Protein Depletion Kit Basic Features Efficient removal of highly abundant proteins 1
Microarray Industry Products
 Via Nicaragua, 12-14 00040 Pomezia (Roma) Phone: +39 06 91601628 Fax: +39 06 91612477 info@lifelinelab.com www.lifelinelab.com Microarray Industry Products Page 10 NBT / BCPIP Chromogenic phosphatase
Via Nicaragua, 12-14 00040 Pomezia (Roma) Phone: +39 06 91601628 Fax: +39 06 91612477 info@lifelinelab.com www.lifelinelab.com Microarray Industry Products Page 10 NBT / BCPIP Chromogenic phosphatase
IMMUNOPRECIPITATION (IP)
 1 IMMUNOPRECIPITATION (IP) Overview and Technical Tips 2 CONTENTS 3 7 8 9 12 13 17 18 19 20 Introduction Factors Influencing IP General Protocol Modifications Of IP Protocols Troubleshooting Contact Us
1 IMMUNOPRECIPITATION (IP) Overview and Technical Tips 2 CONTENTS 3 7 8 9 12 13 17 18 19 20 Introduction Factors Influencing IP General Protocol Modifications Of IP Protocols Troubleshooting Contact Us
Ab Initio SERVER PROTOTYPE FOR PREDICTION OF PHOSPHORYLATION SITES IN PROTEINS*
 COMPUTATIONAL METHODS IN SCIENCE AND TECHNOLOGY 9(1-2) 93-100 (2003/2004) Ab Initio SERVER PROTOTYPE FOR PREDICTION OF PHOSPHORYLATION SITES IN PROTEINS* DARIUSZ PLEWCZYNSKI AND LESZEK RYCHLEWSKI BiolnfoBank
COMPUTATIONAL METHODS IN SCIENCE AND TECHNOLOGY 9(1-2) 93-100 (2003/2004) Ab Initio SERVER PROTOTYPE FOR PREDICTION OF PHOSPHORYLATION SITES IN PROTEINS* DARIUSZ PLEWCZYNSKI AND LESZEK RYCHLEWSKI BiolnfoBank
NPTEL VIDEO COURSE PROTEOMICS PROF. SANJEEVA SRIVASTAVA
 LECTURE-06 PROTEIN PURIFICATION AND PEPTIDE ISOLATION USING CHROMATOGRAPHY TRANSCRIPT Welcome to the proteomics course. Today, we will talk about protein purification and peptide isolation using chromatography
LECTURE-06 PROTEIN PURIFICATION AND PEPTIDE ISOLATION USING CHROMATOGRAPHY TRANSCRIPT Welcome to the proteomics course. Today, we will talk about protein purification and peptide isolation using chromatography
Purification of alpha-1 antitrypsin using an antibody based affinity chromatography medium
 Purification of alpha-1 antitrypsin using an antibody based affinity chromatography medium Ulrika Meyer a, Hanna Wlad a, Sven Blokland b, Frank J.M. Detmers b and Henrik Ihre a a GE Healthcare Bio-Sciences
Purification of alpha-1 antitrypsin using an antibody based affinity chromatography medium Ulrika Meyer a, Hanna Wlad a, Sven Blokland b, Frank J.M. Detmers b and Henrik Ihre a a GE Healthcare Bio-Sciences
TAG-LITE RECEPTOR LIGAND BINDING ASSAY
 TAG - L I T E TAG-LITE RECEPTOR LIGAND BINDING ASSAY PROTOCOL ASSAY PRINCIPLE The Tag-lite Ligand Binding Assay is a homogeneous alternative to radio ligand binding assays for HTS and compound profiling.
TAG - L I T E TAG-LITE RECEPTOR LIGAND BINDING ASSAY PROTOCOL ASSAY PRINCIPLE The Tag-lite Ligand Binding Assay is a homogeneous alternative to radio ligand binding assays for HTS and compound profiling.
BIOC 463A Expt. 4: Column Chromatographic Methods Column Chromatography
 Column Chromatography Chromatography is the process use to separate molecules based on SOME physical property of the molecule: Mass (i.e. size) Charge Affinity for ligands or substrates Hydrophobic interactions
Column Chromatography Chromatography is the process use to separate molecules based on SOME physical property of the molecule: Mass (i.e. size) Charge Affinity for ligands or substrates Hydrophobic interactions
PureSpeed Tips. Superior Protein Purity and Concentration Protein Purification in a Pipette Tip
 PureSpeed Tips PureSpeed Protein Tips Highest purity and concentration Fast as little as 15 minutes Process many samples at once Superior Protein Purity and Concentration Protein Purification in a Pipette
PureSpeed Tips PureSpeed Protein Tips Highest purity and concentration Fast as little as 15 minutes Process many samples at once Superior Protein Purity and Concentration Protein Purification in a Pipette
Green Fluorescent Protein (GFP) Purification. Hydrophobic Interaction Chromatography
 Green Fluorescent Protein (GFP) Purification Hydrophobic Interaction Chromatography What is the GFP gene? GFP is a green fluorescent protein that is normally found in jellyfish. It has been engineered
Green Fluorescent Protein (GFP) Purification Hydrophobic Interaction Chromatography What is the GFP gene? GFP is a green fluorescent protein that is normally found in jellyfish. It has been engineered
For the quantitative detection of human IL6 in serum, plasma, cell culture supernatants and urine.
 m andw da a For the quantitative detection of human IL6 in serum, plasma, cell culture supernatants and urine. general information Catalogue Number Product Name Species cross-reactivity Range (calibration
m andw da a For the quantitative detection of human IL6 in serum, plasma, cell culture supernatants and urine. general information Catalogue Number Product Name Species cross-reactivity Range (calibration
Thermo Scientific Mass Spectrometric Immunoassay (MSIA) Pipette Tips. Next generation immunoaffinity. Robust quantitative platform
 Thermo Scientific Mass Spectrometric Immunoassay (MSIA) Pipette Tips Next generation immunoaffinity Robust quantitative platform Immunoaffinity sample preparation Thermo Scientific Mass Spectrometric Immunoassay
Thermo Scientific Mass Spectrometric Immunoassay (MSIA) Pipette Tips Next generation immunoaffinity Robust quantitative platform Immunoaffinity sample preparation Thermo Scientific Mass Spectrometric Immunoassay
2. Sample dilution: Tissue lysate and cell lysate sample should be diluted at least 5-fold with 1x Sample Diluent Buffer.
 Mouse IL-6 (Lysate) ELISA Kit Catalog No: CKM054 Size: 1 x 96 tests I. Introduction The Cell Sciences Mouse IL-6 ELISA (Enzyme-Linked Immunosorbent Assay) kit is an in vitro enzyme-linked immunosorbent
Mouse IL-6 (Lysate) ELISA Kit Catalog No: CKM054 Size: 1 x 96 tests I. Introduction The Cell Sciences Mouse IL-6 ELISA (Enzyme-Linked Immunosorbent Assay) kit is an in vitro enzyme-linked immunosorbent
COMPAS for the Analysis of SELEX Experiments
 COMPAS for the Analysis of SELEX Experiments COMPAS (COMmon PAtternS) is a software tool that was especially developed to harness the technology of next generation sequencing (NGS) to bring light into
COMPAS for the Analysis of SELEX Experiments COMPAS (COMmon PAtternS) is a software tool that was especially developed to harness the technology of next generation sequencing (NGS) to bring light into
Drug Discovery Research Clinical Screening. Comparison of ELISA and AlphaScreen Assay Technologies for Measurement of Protein Expression Levels
 Drug Discovery Research Clinical Screening FLAG Expression Measurement Application Note Comparison of ELISA and AlphaScreen Assay Technologies for Measurement of Protein Expression Levels ALPHASCREEN APPLICATION
Drug Discovery Research Clinical Screening FLAG Expression Measurement Application Note Comparison of ELISA and AlphaScreen Assay Technologies for Measurement of Protein Expression Levels ALPHASCREEN APPLICATION
FRAUNHOFER IME SCREENINGPORT
 FRAUNHOFER IME SCREENINGPORT Detection technologies used in drug discovery Introduction Detection technologies in drug discovery is unlimited only a biased snapshot can be presented Differences can presented
FRAUNHOFER IME SCREENINGPORT Detection technologies used in drug discovery Introduction Detection technologies in drug discovery is unlimited only a biased snapshot can be presented Differences can presented
Rat IGF-1 ELISA Kit (rigf-1-elisa)
 Rat IGF-1 ELISA Kit (rigf-1-elisa) Cat. No. EK0377 96 Tests in 8 x 12 divisible strips Background Insulin-like growth factor 1 (IGF-1), also known as somatomedin C, is a polypeptide protein hormone similar
Rat IGF-1 ELISA Kit (rigf-1-elisa) Cat. No. EK0377 96 Tests in 8 x 12 divisible strips Background Insulin-like growth factor 1 (IGF-1), also known as somatomedin C, is a polypeptide protein hormone similar
SURFACE PLASMON RESONANCE-BASED SYSTEMS
 SURFACE PLASMON RESONANCE-BASED SYSTEMS ADVANCED METHODS IN BIOENGINEERING LABORATORY 3/1/2011 1 Schedule Week 1: Introduction Reagents preparation Ligand immobilization of Protocol 1 Week 2: Kinetics
SURFACE PLASMON RESONANCE-BASED SYSTEMS ADVANCED METHODS IN BIOENGINEERING LABORATORY 3/1/2011 1 Schedule Week 1: Introduction Reagents preparation Ligand immobilization of Protocol 1 Week 2: Kinetics
Enrichment of EGFR/PI3K/AKT/PTEN Proteins for Research using Immunoprecipitation and with Mass Spectrometry-based Analysis
 Enrichment of /PI3K/AKT/ Proteins for Research using Immunoprecipitation and with Mass Spectrometry-based Analysis Bhavin Patel, Scott Meier, Kay Opperman, Paul Haney, Barbara Kaboord, John Rogers Thermo
Enrichment of /PI3K/AKT/ Proteins for Research using Immunoprecipitation and with Mass Spectrometry-based Analysis Bhavin Patel, Scott Meier, Kay Opperman, Paul Haney, Barbara Kaboord, John Rogers Thermo
Proteomics and Protein Microarrays
 Introduction to BioMEMS & Medical Microdevices Proteomics and Protein Microarrays Companion lecture to the textbook: Fundamentals of BioMEMS and Medical Microdevices, by Prof., http://saliterman.umn.edu/
Introduction to BioMEMS & Medical Microdevices Proteomics and Protein Microarrays Companion lecture to the textbook: Fundamentals of BioMEMS and Medical Microdevices, by Prof., http://saliterman.umn.edu/
Tag-lite Tachykinin NK1 labeled Cells, ready-to-use (transformed & labeled), 200 tests* (Part# C1TT1NK1)
 TAG - L I T E Tag-lite Tachykinin NK1 Receptor Ligand Binding Assay protocol Tachykinin NK1 labeled cells for: 200 tests Part#: C1TT1NK1 Lot#: 03-1 March 2016 (See expiration date on package label) Store
TAG - L I T E Tag-lite Tachykinin NK1 Receptor Ligand Binding Assay protocol Tachykinin NK1 labeled cells for: 200 tests Part#: C1TT1NK1 Lot#: 03-1 March 2016 (See expiration date on package label) Store
Gene Regulation Solutions. Microarrays and Next-Generation Sequencing
 Gene Regulation Solutions Microarrays and Next-Generation Sequencing Gene Regulation Solutions The Microarrays Advantage Microarrays Lead the Industry in: Comprehensive Content SurePrint G3 Human Gene
Gene Regulation Solutions Microarrays and Next-Generation Sequencing Gene Regulation Solutions The Microarrays Advantage Microarrays Lead the Industry in: Comprehensive Content SurePrint G3 Human Gene
MBios 478: Mass Spectrometry Applications [Dr. Wyrick] Slide #1. Lecture 25: Mass Spectrometry Applications
![MBios 478: Mass Spectrometry Applications [Dr. Wyrick] Slide #1. Lecture 25: Mass Spectrometry Applications MBios 478: Mass Spectrometry Applications [Dr. Wyrick] Slide #1. Lecture 25: Mass Spectrometry Applications](/thumbs/74/71368870.jpg) MBios 478: Mass Spectrometry Applications [Dr. Wyrick] Slide #1 Lecture 25: Mass Spectrometry Applications Measuring Protein Abundance o ICAT o DIGE Identifying Post-Translational Modifications Protein-protein
MBios 478: Mass Spectrometry Applications [Dr. Wyrick] Slide #1 Lecture 25: Mass Spectrometry Applications Measuring Protein Abundance o ICAT o DIGE Identifying Post-Translational Modifications Protein-protein
Applications of HTRF and Tag-lite Assays for HTP Antibody Screening
 Applications of HTRF and Tag-lite Assays for HTP Antibody Screening Brigitte Devaux, PhD Bristol Myers Squibb, Redwood City CA HTRF Symposium April 25, 2013 1 Introduction Generate human therapeutic antibodies
Applications of HTRF and Tag-lite Assays for HTP Antibody Screening Brigitte Devaux, PhD Bristol Myers Squibb, Redwood City CA HTRF Symposium April 25, 2013 1 Introduction Generate human therapeutic antibodies
ARBRE-P4EU Consensus Protein Quality Guidelines for Biophysical and Biochemical Studies Minimal information to provide
 ARBRE-P4EU Consensus Protein Quality Guidelines for Biophysical and Biochemical Studies Minimal information to provide Protein name and full primary structure, by providing a NCBI (or UniProt) accession
ARBRE-P4EU Consensus Protein Quality Guidelines for Biophysical and Biochemical Studies Minimal information to provide Protein name and full primary structure, by providing a NCBI (or UniProt) accession
The World Leader in SPR Technology. Jimmy Page, PhD, Biacore, Inc.
 The World Leader in SPR Technology Jimmy Page, PhD, Biacore, Inc. Objectives of Biacore Experiments Yes/No Data» Is there binding?» Ligand Fishing Concentration Analysis: How MUCH? Active Concentration
The World Leader in SPR Technology Jimmy Page, PhD, Biacore, Inc. Objectives of Biacore Experiments Yes/No Data» Is there binding?» Ligand Fishing Concentration Analysis: How MUCH? Active Concentration
Supplementary Figure 1. Utilization of publicly available antibodies in different applications.
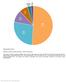 Supplementary Figure 1 Utilization of publicly available antibodies in different applications. The fraction of publicly available antibodies toward human protein targets is shown with data for the following
Supplementary Figure 1 Utilization of publicly available antibodies in different applications. The fraction of publicly available antibodies toward human protein targets is shown with data for the following
PXBioVisioN. Protease Inhibition. Analysis of Peptides as Surrogates for Protease Activity to Monitor Protease Inhibition in-vivo
 PXBioVisioN Protease Inhibition Analysis of Peptides as Surrogates for Protease Activity to Monitor Protease Inhibition in-vivo Peptides are Products of Protease Activity Peptidomics is defined as the
PXBioVisioN Protease Inhibition Analysis of Peptides as Surrogates for Protease Activity to Monitor Protease Inhibition in-vivo Peptides are Products of Protease Activity Peptidomics is defined as the
Immunoassays and Mass Spectrometry: Transitioning Technology
 TECHNICAL BRIEF Immunoassays and Mass Spectrometry: Transitioning Technology This technical brief presents data obtained when measuring eicosanoids by immunoassay and mass spectrometry. It briefly considers
TECHNICAL BRIEF Immunoassays and Mass Spectrometry: Transitioning Technology This technical brief presents data obtained when measuring eicosanoids by immunoassay and mass spectrometry. It briefly considers
MSD GOLD Streptavidin and Avidin Plates
 MSD GOLD Streptavidin and Avidin Plates SECTOR Plates QUICKPLEX Plates 96-well Streptavidin L15SA L55SA 96-well Small Spot Streptavidin L45SA N/A 96-well High Bind Avidin L15AB L55AB FOR RESEARCH USE ONLY.
MSD GOLD Streptavidin and Avidin Plates SECTOR Plates QUICKPLEX Plates 96-well Streptavidin L15SA L55SA 96-well Small Spot Streptavidin L45SA N/A 96-well High Bind Avidin L15AB L55AB FOR RESEARCH USE ONLY.
Rapid GST Inclusion Body Solubilization and Renaturation Kit
 Product Manual Rapid GST Inclusion Body Solubilization and Renaturation Kit Catalog Number AKR-110 FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Bacteria are widely used for His
Product Manual Rapid GST Inclusion Body Solubilization and Renaturation Kit Catalog Number AKR-110 FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Bacteria are widely used for His
colorimetric sandwich ELISA kit datasheet
 colorimetric sandwich ELISA kit datasheet For the quantitative detection of mouse Interleukin 6 (IL6) concentrations in serum, plasma and cell culture supernatants. general information Catalogue Number
colorimetric sandwich ELISA kit datasheet For the quantitative detection of mouse Interleukin 6 (IL6) concentrations in serum, plasma and cell culture supernatants. general information Catalogue Number
Biomarker Discovery using Surface Plasmon Resonance Imaging
 F e a t u r e A r t i c l e Feature Article Biomarker Discovery using Surface Plasmon Resonance Imaging Elodie LY-MORIN, Sophie BELLON, Géraldine MÉLIZZI, Chiraz FRYDMAN Surface Plasmon Resonance (SPR)
F e a t u r e A r t i c l e Feature Article Biomarker Discovery using Surface Plasmon Resonance Imaging Elodie LY-MORIN, Sophie BELLON, Géraldine MÉLIZZI, Chiraz FRYDMAN Surface Plasmon Resonance (SPR)
Tyrosine Kinase Assay Kit, Red*
 Rh Tyrosine Kinase Assay Kit, Red* 1.0 INTRODUCTION Part # P2882, P2883 Lit. # L0531 Rev. 11/02 Page 1 of 6 The phosphorylation of proteins by protein tyrosine kinases (PTKs) is critical to the normal
Rh Tyrosine Kinase Assay Kit, Red* 1.0 INTRODUCTION Part # P2882, P2883 Lit. # L0531 Rev. 11/02 Page 1 of 6 The phosphorylation of proteins by protein tyrosine kinases (PTKs) is critical to the normal
Peptide libraries: applications, design options and considerations. Laura Geuss, PhD May 5, 2015, 2:00-3:00 pm EST
 Peptide libraries: applications, design options and considerations Laura Geuss, PhD May 5, 2015, 2:00-3:00 pm EST Overview 1 2 3 4 5 Introduction Peptide library basics Peptide library design considerations
Peptide libraries: applications, design options and considerations Laura Geuss, PhD May 5, 2015, 2:00-3:00 pm EST Overview 1 2 3 4 5 Introduction Peptide library basics Peptide library design considerations
Improving the Limit of Detection of Lateral Flow Assays using 3DNA Technology
 Improving the Limit of Detection of Lateral Flow Assays using 3DNA Technology Introduction Lateral Flow (LF) and similar assays represent a unique growing class of Point of Care (POC) tests designed to
Improving the Limit of Detection of Lateral Flow Assays using 3DNA Technology Introduction Lateral Flow (LF) and similar assays represent a unique growing class of Point of Care (POC) tests designed to
ChIP-Seq Data Analysis. J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015
 ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA
ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA
Affinity Chromatography. Teaching Kit Manual. GeNei TM. Cat No. New Cat No. KT Revision No.:
 Affinity Chromatography Teaching Kit Manual Cat No. New Cat No. KT41 106192 Revision No.: 00010905 CONTENTS Page No. Objective 3 Principle 3 Kit Description 4 Materials Provided 6 Procedure 7 Result 12
Affinity Chromatography Teaching Kit Manual Cat No. New Cat No. KT41 106192 Revision No.: 00010905 CONTENTS Page No. Objective 3 Principle 3 Kit Description 4 Materials Provided 6 Procedure 7 Result 12
Efficient Multi-Well Protein Purification Strategies
 Application Note PN 33576 Efficient Multi-Well Protein Purification Strategies Introduction Many tools and techniques are available today for protein purification. Development of a purification process
Application Note PN 33576 Efficient Multi-Well Protein Purification Strategies Introduction Many tools and techniques are available today for protein purification. Development of a purification process
Strep-Tactin XT Spin Column
 Strep-Tactin XT Spin Column Purification Protocol Last date of revision Last June date 2017 of revision June 2017 Version PR90-0001 Version PR90-0001 For research use only Important licensing information
Strep-Tactin XT Spin Column Purification Protocol Last date of revision Last June date 2017 of revision June 2017 Version PR90-0001 Version PR90-0001 For research use only Important licensing information
SUMOylated protein capture kit
 SUMOylated protein capture kit Cat no. A010-100 Capture and detect SUMOylated proteins High capacity, high specificity SUMO binding matrix Fast, convenient protein isolation using purification system provided
SUMOylated protein capture kit Cat no. A010-100 Capture and detect SUMOylated proteins High capacity, high specificity SUMO binding matrix Fast, convenient protein isolation using purification system provided
FirePlex mirna Assay. Multiplex microrna profiling from low sample inputs
 FirePlex mirna Assay Multiplex microrna profiling from low sample inputs Abstract We introduce a new assay for multiplex microrna (mirna) discovery and verification that enables simultaneous profiling
FirePlex mirna Assay Multiplex microrna profiling from low sample inputs Abstract We introduce a new assay for multiplex microrna (mirna) discovery and verification that enables simultaneous profiling
Masayoshi Honda, Jeehae Park, Robert A. Pugh, Taekjip Ha, and Maria Spies
 Molecular Cell, Volume 35 Supplemental Data Single-Molecule Analysis Reveals Differential Effect of ssdna-binding Proteins on DNA Translocation by XPD Helicase Masayoshi Honda, Jeehae Park, Robert A. Pugh,
Molecular Cell, Volume 35 Supplemental Data Single-Molecule Analysis Reveals Differential Effect of ssdna-binding Proteins on DNA Translocation by XPD Helicase Masayoshi Honda, Jeehae Park, Robert A. Pugh,
Expression Array System
 Integrated Science for Gene Expression Applied Biosystems Expression Array System Expression Array System SEE MORE GENES The most complete, most sensitive system for whole genome expression analysis. The
Integrated Science for Gene Expression Applied Biosystems Expression Array System Expression Array System SEE MORE GENES The most complete, most sensitive system for whole genome expression analysis. The
NAG-1 (GDF-15, MIC-1) Prostate Cancer ELISA kit
 NAG-1 (GDF-15, MIC-1) Prostate Cancer ELISA kit Catalog Number: NG1 Store at -0 C. FOR RESEARCH USE ONLY v. 1081 Introduction This sandwich ELISA kit is for determination of NAG-1 (GDF-15, MIC-1) levels
NAG-1 (GDF-15, MIC-1) Prostate Cancer ELISA kit Catalog Number: NG1 Store at -0 C. FOR RESEARCH USE ONLY v. 1081 Introduction This sandwich ELISA kit is for determination of NAG-1 (GDF-15, MIC-1) levels
Technical tips Session 5
 Technical tips Session 5 Chromatine Immunoprecipitation (ChIP): This is a powerful in vivo method to quantitate interaction of proteins associated with specific regions of the genome. It involves the immunoprecipitation
Technical tips Session 5 Chromatine Immunoprecipitation (ChIP): This is a powerful in vivo method to quantitate interaction of proteins associated with specific regions of the genome. It involves the immunoprecipitation
CBI Toolbox Tour 2015
 CBI Toolbox Tour 2015 Thermophoresis (NanoTemper) NT.115 & NT.LabelFree Images: NanoTemper Circular Dichroism Jasco J-1500 Spectrometer Six Position Turreted Peltier Temperature Control System Automated
CBI Toolbox Tour 2015 Thermophoresis (NanoTemper) NT.115 & NT.LabelFree Images: NanoTemper Circular Dichroism Jasco J-1500 Spectrometer Six Position Turreted Peltier Temperature Control System Automated
Protein Sequence Analysis. BME 110: CompBio Tools Todd Lowe April 19, 2007 (Slide Presentation: Carol Rohl)
 Protein Sequence Analysis BME 110: CompBio Tools Todd Lowe April 19, 2007 (Slide Presentation: Carol Rohl) Linear Sequence Analysis What can you learn from a (single) protein sequence? Calculate it s physical
Protein Sequence Analysis BME 110: CompBio Tools Todd Lowe April 19, 2007 (Slide Presentation: Carol Rohl) Linear Sequence Analysis What can you learn from a (single) protein sequence? Calculate it s physical
Protein Microarrays?
 Protein Microarrays? Protein Chemistry/Proteomics Unit and the Neuroscience Program,Biomedicum Helsinki E-Mail: marc.baumann@helsinki.fi (http://research.med.helsinki.fi/corefacilities/proteinchem) What
Protein Microarrays? Protein Chemistry/Proteomics Unit and the Neuroscience Program,Biomedicum Helsinki E-Mail: marc.baumann@helsinki.fi (http://research.med.helsinki.fi/corefacilities/proteinchem) What
EPIGENTEK. EpiQuik Histone Demethylase LSD1 Activity/Inhibition Assay Kit. Base Catalog # P-3076 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE
 EpiQuik Histone Demethylase LSD1 Activity/Inhibition Assay Kit Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Histone Demethylase LSD1 Activity/Inhibition Assay Kit is very suitable
EpiQuik Histone Demethylase LSD1 Activity/Inhibition Assay Kit Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Histone Demethylase LSD1 Activity/Inhibition Assay Kit is very suitable
GSI Equine IL6 ELISA Kit- Cell Lysate DataSheet
 IL-6 is an interleukin that acts as both a pro-inflammatory and anti-inflammatory cytokine and is produced by T cells, macrophages, fibroblasts, osteoblasts, endothelial and other cells (1,2,3). IL-6 induces
IL-6 is an interleukin that acts as both a pro-inflammatory and anti-inflammatory cytokine and is produced by T cells, macrophages, fibroblasts, osteoblasts, endothelial and other cells (1,2,3). IL-6 induces
Effect of Hydrogen Ion Concentration and Buffer Composition on Ligand Binding Characteristics and Polymerization of Cow s Milk Folate Binding Protein
 Bioscience Reports, Vol. 21, No. 6, December 2001 ( 2002) Effect of Hydrogen Ion Concentration and Buffer Composition on Ligand Binding Characteristics and Polymerization of Cow s Milk Folate Binding Protein
Bioscience Reports, Vol. 21, No. 6, December 2001 ( 2002) Effect of Hydrogen Ion Concentration and Buffer Composition on Ligand Binding Characteristics and Polymerization of Cow s Milk Folate Binding Protein
Aims: -Purification of a specific protein. -Study of protein-protein interactions
 Aims: -Purification of a specific protein -Study of protein-protein interactions This is a reliable method for purifying total IgG from crude protein mixtures such as serum. Protein A (linked to resin
Aims: -Purification of a specific protein -Study of protein-protein interactions This is a reliable method for purifying total IgG from crude protein mixtures such as serum. Protein A (linked to resin
RayBio Human bfgf ELISA Kit
 RayBio Human bfgf ELISA Kit User Manual (for Cell Lysate and Tissue Lysate) (Revised Mar 1, 2012) RayBio Human bfgf ELISA Kit Protocol (Cat#: ELH-bFGF-001C) RayBiotech, Inc. We Provide You With Excellent
RayBio Human bfgf ELISA Kit User Manual (for Cell Lysate and Tissue Lysate) (Revised Mar 1, 2012) RayBio Human bfgf ELISA Kit Protocol (Cat#: ELH-bFGF-001C) RayBiotech, Inc. We Provide You With Excellent
What is Nano-Bio? Non-Covalent Interactions
 - - What is Nano-Bio? Physicist: Biotech: Biologists: -study of molecular interactions -application of nano-tools to study biological systems. -application of nano-tools to detect, treat, and prevent disease
- - What is Nano-Bio? Physicist: Biotech: Biologists: -study of molecular interactions -application of nano-tools to study biological systems. -application of nano-tools to detect, treat, and prevent disease
Product. Ni-NTA His Bind Resin. Ni-NTA His Bind Superflow. His Bind Resin. His Bind Magnetic Agarose Beads. His Bind Column. His Bind Quick Resin
 Novagen offers a large variety of affinity supports and kits for the purification of recombinant proteins containing popular peptide fusion tags, including His Tag, GST Tag, S Tag and T7 Tag sequences.
Novagen offers a large variety of affinity supports and kits for the purification of recombinant proteins containing popular peptide fusion tags, including His Tag, GST Tag, S Tag and T7 Tag sequences.
Gene Expression Technology
 Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Elecrtonic Supplementary Information. Application of quantum dot barcodes prepared using biological self-assembly to multiplexed immunoassays
 Elecrtonic Supplementary Information Application of quantum dot barcodes prepared using biological self-assembly to multiplexed immunoassays Sakandar Rauf, Andrew Glidle and Jonathan M Cooper Department
Elecrtonic Supplementary Information Application of quantum dot barcodes prepared using biological self-assembly to multiplexed immunoassays Sakandar Rauf, Andrew Glidle and Jonathan M Cooper Department
Systematic Evolution of Ligands by Exponential Enrichment SELEX
 Systematic Evolution of Ligands by Exponential Enrichment SELEX SELEX Systematic evolution of ligands by exponential amplification * Isolation of new RNs with novel properties * New therapeutics, New diagnostics
Systematic Evolution of Ligands by Exponential Enrichment SELEX SELEX Systematic evolution of ligands by exponential amplification * Isolation of new RNs with novel properties * New therapeutics, New diagnostics
Affinity Chromatography Media. Cellufine Sulfate. Technical Data Sheet
 Affinity Chromatography Media Cellufine Sulfate Technical Data Sheet Cellufine Sulfate For concentration, purification and depyrogenation of virus, viral/microbial antigens and heparin-binding proteins
Affinity Chromatography Media Cellufine Sulfate Technical Data Sheet Cellufine Sulfate For concentration, purification and depyrogenation of virus, viral/microbial antigens and heparin-binding proteins
RayBio Human HGF ELISA Kit
 RayBio Human HGF ELISA Kit User Manual (for Cell Lysate and Tissue Lysate) (Revised Mar 1, 2012) RayBio Human HGF ELISA Kit Protocol (Cat#: ELH-HGF-001C) RayBiotech, Inc. We Provide You With Excellent
RayBio Human HGF ELISA Kit User Manual (for Cell Lysate and Tissue Lysate) (Revised Mar 1, 2012) RayBio Human HGF ELISA Kit Protocol (Cat#: ELH-HGF-001C) RayBiotech, Inc. We Provide You With Excellent
Hossain_Supplemental Figure 1
 Hossain_Supplemental Figure 1 GFP-PACT GFP-PACT Motif I GFP-PACT Motif II A. MG132 (1µM) GFP Tubulin GFP-PACT Pericentrin GFP-PACT GFP-PACT Pericentrin Fig. S1. Expression and localization of Orc1 PACT
Hossain_Supplemental Figure 1 GFP-PACT GFP-PACT Motif I GFP-PACT Motif II A. MG132 (1µM) GFP Tubulin GFP-PACT Pericentrin GFP-PACT GFP-PACT Pericentrin Fig. S1. Expression and localization of Orc1 PACT
CSE : Computational Issues in Molecular Biology. Lecture 19. Spring 2004
 CSE 397-497: Computational Issues in Molecular Biology Lecture 19 Spring 2004-1- Protein structure Primary structure of protein is determined by number and order of amino acids within polypeptide chain.
CSE 397-497: Computational Issues in Molecular Biology Lecture 19 Spring 2004-1- Protein structure Primary structure of protein is determined by number and order of amino acids within polypeptide chain.
MANUALL TUBEs: Tandem Ubiquitin Binding Entities. Biotinylated K63-TUBE 1. Catalog Numbers: UM304
 - 1 - MANUALL Biotinylated K63-TUBE 1 Catalog Numbers: UM304 all products are for research use only not intended for human or animal diagnostic or therapeutic uses LifeSensors, Inc., 271 Great Valley Parkway,
- 1 - MANUALL Biotinylated K63-TUBE 1 Catalog Numbers: UM304 all products are for research use only not intended for human or animal diagnostic or therapeutic uses LifeSensors, Inc., 271 Great Valley Parkway,
MSD MULTI-SPOT Assay System
 MSD MULTI-SPOT Assay System Human MMP 2-Plex Ultra-Sensitive Kit 1-Plate Kit 5-Plate Kit 25-Plate Kit K15033C-1 K15033C-2 K15033C-4 17335-v5-2012Mar 1 MSD Biomarker Assays Human MMP 2-Plex Ultra-Sensitive
MSD MULTI-SPOT Assay System Human MMP 2-Plex Ultra-Sensitive Kit 1-Plate Kit 5-Plate Kit 25-Plate Kit K15033C-1 K15033C-2 K15033C-4 17335-v5-2012Mar 1 MSD Biomarker Assays Human MMP 2-Plex Ultra-Sensitive
[4] SCHEMA-Guided Protein Recombination
![[4] SCHEMA-Guided Protein Recombination [4] SCHEMA-Guided Protein Recombination](/thumbs/78/77868499.jpg) [4] SCHEMA-guided protein recombination 35 [4] SCHEMA-Guided Protein Recombination By Jonathan J. Silberg, Jeffrey B. Endelman, and Frances H. Arnold Introduction SCHEMA is a scoring function that predicts
[4] SCHEMA-guided protein recombination 35 [4] SCHEMA-Guided Protein Recombination By Jonathan J. Silberg, Jeffrey B. Endelman, and Frances H. Arnold Introduction SCHEMA is a scoring function that predicts
Immunoassay Methods. Assay Guidance Manual Assay Guidance Manual Assay Guidance Manual. Abstract
 Karen L. Cox, BS Eli Lilly & Company, Indianapolis, IN cox_karen_l@lilly.com Viswanath Devanarayan, PhD AbbVie viswanath.devanarayan@abbvie.com Aidas Kriauciunas Eli Lilly & Company, Indianapolis, IN Joseph
Karen L. Cox, BS Eli Lilly & Company, Indianapolis, IN cox_karen_l@lilly.com Viswanath Devanarayan, PhD AbbVie viswanath.devanarayan@abbvie.com Aidas Kriauciunas Eli Lilly & Company, Indianapolis, IN Joseph
UV Fluorescence Polarization as a Means to Investigate Protein Conformational and Mass Change
 A p p l i c a t i o n N o t e UV Fluorescence Polarization as a Means to Investigate Protein Conformational and Mass Change Using Intrinsic Tryptophan Fluorescence in Conjunction with UV-capable Polarizers
A p p l i c a t i o n N o t e UV Fluorescence Polarization as a Means to Investigate Protein Conformational and Mass Change Using Intrinsic Tryptophan Fluorescence in Conjunction with UV-capable Polarizers
Purification of (recombinant) proteins. Pekka Lappalainen, Institute of Biotechnology, University of Helsinki
 Purification of (recombinant) proteins Pekka Lappalainen, Institute of Biotechnology, University of Helsinki Physical properties of proteins that can be applied for purification -size -charge (isoelectric
Purification of (recombinant) proteins Pekka Lappalainen, Institute of Biotechnology, University of Helsinki Physical properties of proteins that can be applied for purification -size -charge (isoelectric
RayBio Human NF-κB p65 Transcription Factor Activity Assay Kit
 RayBio Human NF-κB p65 Transcription Factor Activity Assay Kit Catalog #: TFEH-p65 User Manual Mar 13, 2017 3607 Parkway Lane, Suite 200 Norcross, GA 30092 Tel: 1-888-494-8555 (Toll Free) or 770-729-2992,
RayBio Human NF-κB p65 Transcription Factor Activity Assay Kit Catalog #: TFEH-p65 User Manual Mar 13, 2017 3607 Parkway Lane, Suite 200 Norcross, GA 30092 Tel: 1-888-494-8555 (Toll Free) or 770-729-2992,
Enhancers mutations that make the original mutant phenotype more extreme. Suppressors mutations that make the original mutant phenotype less extreme
 Interactomics and Proteomics 1. Interactomics The field of interactomics is concerned with interactions between genes or proteins. They can be genetic interactions, in which two genes are involved in the
Interactomics and Proteomics 1. Interactomics The field of interactomics is concerned with interactions between genes or proteins. They can be genetic interactions, in which two genes are involved in the
RayBio Mouse IL-6 ELISA Kit
 RayBio Mouse IL-6 ELISA Kit User Manual (for Cell Lysate and Tissue Lysate) (Revised Mar 1, 2012) RayBio Mouse IL-6 ELISA Kit Protocol (Cat#: ELM-IL6-001C) RayBiotech, Inc. We Provide You With Excellent
RayBio Mouse IL-6 ELISA Kit User Manual (for Cell Lysate and Tissue Lysate) (Revised Mar 1, 2012) RayBio Mouse IL-6 ELISA Kit Protocol (Cat#: ELM-IL6-001C) RayBiotech, Inc. We Provide You With Excellent
Angewandte. In Vitro Selection of Structure-Switching Signaling Aptamers** Razvan Nutiu and Yingfu Li* Chemie
 Fluorescent Probes In Vitro Selection of Structure-Switching Signaling Aptamers** Razvan Nutiu and Yingfu Li* Signaling aptamers are aptamer probes that couple target binding to fluorescent-signal generation.
Fluorescent Probes In Vitro Selection of Structure-Switching Signaling Aptamers** Razvan Nutiu and Yingfu Li* Signaling aptamers are aptamer probes that couple target binding to fluorescent-signal generation.
Chromatographic Separation of the three forms of RNA Polymerase II.
 Chromatographic Separation of the three forms of RNA Polymerase II. α-amanitin α-amanitin bound to Pol II Function of the three enzymes. Yeast Pol II. RNA Polymerase Subunit Structures 10-7 Subunit structure.
Chromatographic Separation of the three forms of RNA Polymerase II. α-amanitin α-amanitin bound to Pol II Function of the three enzymes. Yeast Pol II. RNA Polymerase Subunit Structures 10-7 Subunit structure.
Tony Mire-Sluis Vice President, Corporate, Product and Device Quality Amgen Inc
 The Regulatory Implications of the ever increasing power of Mass Spectrometry and its role in the Analysis of Biotechnology Products Where do we draw the line? Tony Mire-Sluis Vice President, Corporate,
The Regulatory Implications of the ever increasing power of Mass Spectrometry and its role in the Analysis of Biotechnology Products Where do we draw the line? Tony Mire-Sluis Vice President, Corporate,
