Nature Biotechnology: doi: /nbt.3943
|
|
|
- Lucinda Hutchinson
- 6 years ago
- Views:
Transcription
1 Supplementary Figure 1. Distribution of sequence depth across the bacterial artificial chromosomes (BACs). The x-axis denotes the sequencing depth (X) of each BAC and y-axis denotes the number of BACs corresponding to depth.
2 Supplementary Figure 2. Estimation of the pearl millet genome size based on K-mer statistics. The frequency distribution of 17-mers within the raw genomic reads displays 2 major peaks (A and B). Peak A resembles a Gaussian distribution and represents k-mers of ~0-10X coverage which arise by chance due to sequencing errors. Peak B, corresponding to k-mers of ~20-50X coverage, represents the majority of the genome and resembles a Poisson distribution with minor differences due to sequencing errors, heterozygosity and repetitive DNA. The total genome size of pearl millet was estimated by obtaining the multiplication product of 17 bp and the k-mer frequency (value at y-axis) corresponding to the coverage (value at x-axis) at Peak B (i.e. 17X Peak B frequency).
3 Supplementary Figure 3. Percent similarity and align rate for long PacBio reads vs scaffold sequences.
4 Supplementary Figure 4. GC content distributions in selected grass genomes. The mean GC content of pearl millet is higher than barley (Hordeum vulgare), foxtail millet (Setaria italica), rice (Oryza sativa), and sorghum (Sorghum bicolor). GC content is represented on the x-axis and the proportion of the bin number divided by the total windows on the y-axis.
5 Supplementary Figure 5. Scatterplot showing GC content versus sequencing depth. The graph indicates that no GC bias is present in the sequence data generated.
6 Supplementary Figure 6. GC content distribution of whole genome CDS (left) and 384 pearl millet expanded families (right) in grasses. The GC content in whole genome CDS and 384 expanded gene families were found similar.
7 Percentage of genome Sequence divergence rate (%) Supplementary Figure 7. Distribution of divergence rates of different transposable element (TE) types in the pearl millet genome. DNA- DNA elements; LINE- long interspersed nuclear elements; LTR- long terminal repeat transposable elements; SINE- short interspersed nuclear elements. Divergence rates were high among LTR elements.
8 a.
9 b. Supplementary Figure 8. Length distribution of mrna, CDS, exons and introns in sequenced cereal genomes. (a) The x-axis indicates the length, and the y-axis indicates the gene percentage in the corresponding length window. For example, in the figure Distribution of mrna length, the coordinate indicated by the red arrow is (300, 5.995), meaning that 5.995% of all gene models have lengths in the range bp. (b) Boxplots indicate that the average lengths of different gene features in pearl millet are similar to those in the other four cereal species
10 Supplementary Figure 9. An overview of orthologous and paralogous genes in pearl millet. The number of orthologs and paralogs in the pearl millet genome is shown in comparison to those gene classes in the genomes of ten select plant species [Arabidopsis (Arabidopsis thaliana), Brachypodium (Brachypodium distachyon), banana (Musa acuminata), barley, foxtail millet, maize, rice, sorghum, soybean (Glycine max) and bread wheat]. Reciprocal pair-wise comparisons of the 38,579 pearl millet gene models with 385,891 gene models from ten select plant species identified 17,949 orthologous groups (Supplementary Table 12), among which 5,232 contained only a single pearl millet gene, suggestive of simple orthology (Supplementary Table 13).
11 a. b. Supplementary Figure 10. Length distribution for each expanded family. (a) 20 expanded gene families randomly plotted. (b) 384 expanded gene families. Red dots represent pearl millet, and blue represent rice.
12 Supplementary Figure 11. Phylogeny of NBS-LRR genes. Heavily tandemized gene groups can be observed at one telomere of Pg1 and Pg4 followed by Pg7.
13 Supplementary Figure 12. (a) Indels and (b) gene numbers in PMiGAP lines, parental lines of mapping population and B- and R- lines. Large number of indels were observed in the genome compared to CDS regions in PMiGAP lines.
14 Frequency with SV Supplementary Figure 13. Distribution of structural variations in PMiGAP lines. Structural variations such as deletions (DEL), insertions (INS), inversions (INV), intra-chromosomal translocations (ITX) and others were determined using BreakDancer software. Among different structural variations DEL were found in large number across the genome in the PMiGAP lines.
15 Frequency with SV Supplementary Figure 14. Distribution of structural variations across 38 parents of mapping populations. Deletions were found in large numbers across the genome in the parental lines.
16 Frequency with SV Supplementary Figure 15. Distribution of structural variations in B and R lines. The number of structural variations identified in the B- and R- lines is significantly lower than that identified in case of PMiGAP lines and parental lines of mapping populations. This can be attributed to the RAD-seq approach adopted for sequencing these lines.
17 Supplementary Figure 16. Population structure of 345 PMiGAP lines and 31 wild samples. A set of million SNPs were used to determine the number of sub-populations in PMiGAP lines using STRUCTURE analysis. Population structure, when K=2, mainly separate all the
18 cultivated from the wild relatives. At K=3, cultivated are separated but the separation does not correspond to clear geographical pattern, a large fraction of the cultivated share a blue and red component. At K=4 a further division inside the cultivated show ancestry almost admixed for all cultivated. At K=5, wild populations are split in two well defined clusters and a cluster presenting strong similarity to the cultivated. This last cluster is the closest to the cultivated in the PCA, and correspond to the central area of the wild populations, Niger and Mali. The two other well defined cluster correspond to wild from East Africa and Senegal/Mauritania/West Mali. A new cultivated cluster among widely across cultivated appear at K=6 and 7. Structure can be influenced by number of samples as well as imbalance between cultivated and wild populations. Among the East group, two individuals show a closer proximity to the cultivated in the PCA analysis (PE08743 from Soudan and PE08721 from Chad). These two individuals show admixed ancestry 48% cultivated and 24% cultivated respectively. These two sample were sampled at latitude and respectively. At this latitude wild populations are in sympatry with cultivated varieties and hybridization could occur.
19 a. b.
20 c. d. Supplementary Figure 17. Cross entropy analysis of population structure. (a) For all wild and cultivated samples, Cross entropy decreased slowly from K=2 to K=7. But, when we considered wild samples alone, (a) We found a strong signal of structure inside the wild populations separating wild individuals from the East, Center and West of the Sahel. The different genetic group are represented (c). The wild pearl millet accessions sequenced in this study clustered in to three main groups West African (green), Central Africa (violet) and East African (blue). Note that we used a similar scale here for these two graphics. To better assess if there is any geographical structuration inside the cultivated (d), we performed a geographically explicit model using TESS3. We made the alpha parameter varied from 5% to 60% allowing to weight the geographical position of the samples in the structure inference. The cross entropy do not vary much and is flat or slightly increase from K=1 (no structure) to K=3. Altogether, these analysis suggest a marked signal of structure inside the wild populations, but not inside the cultivated.
21 a.
22 b. Supplementary Figure 18. Diversity in PMiGAP lines and wild species accessions of pearl millet. a. Diversity metrics, presented as average pair-wise nucleotide diversity (current [θπ] and historical [θω]), are shown across all seven pseudomolecules. b. FST and -log ratio of the diversity loss across the 7 pseudomolecules.
23 Supplementary Figure 19. Genome wide linkage disequilibrium decay in the PMiGAP lines, parents of mapping populations, and B- and R- lines. Linkage disequilibrium decays more rapidly across the PMiGAP lines compared to the B- and R- lines and parental lines.
24 a.
25 b.
26 c.
27 d. Supplementary Figure 20. Association mapping for the grain number per panicle. The figure represent the -log(p-value) of the association in function of the position of the SNP on the pseudomolecule for the control as well as early and late stress field trials. We represented here the result for the grains per panicle (GNP). (a) results for Pg1 in 2011, (b) results for Pg1 in 2012, (c) results for Pg5 in 2011, (d) results for Pg5 in 2012.
28 Supplementary Figure 21. Hierarchical clustering of predicted hybrid performance. The lower bottom of clusters shown are the best hybrid combinations for pearl millet breeding
29 Supplementary Figure 22. Length and N50 distribution of the BAC clones. 29
30 Supplementary Figure 23. QQ plot of expected and observed P-value. The expected and observed p-value for the number of grains per panicle (GNP) and fresh stover yield (FSWTHA) are reported for the two years (2011, 2012) and the three field trials (control, early stress, late stress). P-value are reported on a -log scale. A value of 4 corresponds to a P-value of Expected 30
31 and observed p-value fit the x=y relationship. The marker trait association with are significant when P-value above
3I03 - Eukaryotic Genetics Repetitive DNA
 Repetitive DNA Satellite DNA Minisatellite DNA Microsatellite DNA Transposable elements LINES, SINES and other retrosequences High copy number genes (e.g. ribosomal genes, histone genes) Multifamily member
Repetitive DNA Satellite DNA Minisatellite DNA Microsatellite DNA Transposable elements LINES, SINES and other retrosequences High copy number genes (e.g. ribosomal genes, histone genes) Multifamily member
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Number and length distributions of the inferred fosmids.
 Supplementary Figure 1 Number and length distributions of the inferred fosmids. Fosmid were inferred by mapping each pool s sequence reads to hg19. We retained only those reads that mapped to within a
Supplementary Figure 1 Number and length distributions of the inferred fosmids. Fosmid were inferred by mapping each pool s sequence reads to hg19. We retained only those reads that mapped to within a
Genomics and Transcriptomics of Spirodela polyrhiza
 Genomics and Transcriptomics of Spirodela polyrhiza Doug Bryant Bioinformatics Core Facility & Todd Mockler Group, Donald Danforth Plant Science Center Desired Outcomes High-quality genomic reference sequence
Genomics and Transcriptomics of Spirodela polyrhiza Doug Bryant Bioinformatics Core Facility & Todd Mockler Group, Donald Danforth Plant Science Center Desired Outcomes High-quality genomic reference sequence
Structural variation. Marta Puig Institut de Biotecnologia i Biomedicina Universitat Autònoma de Barcelona
 Structural variation Marta Puig Institut de Biotecnologia i Biomedicina Universitat Autònoma de Barcelona Genetic variation How much genetic variation is there between individuals? What type of variants
Structural variation Marta Puig Institut de Biotecnologia i Biomedicina Universitat Autònoma de Barcelona Genetic variation How much genetic variation is there between individuals? What type of variants
Phenotypic response conferred by the Lr22a leaf rust resistance gene against ten Swiss P. triticina isolates.
 Supplementary Figure 1 Phenotypic response conferred by the leaf rust resistance gene against ten Swiss P. triticina isolates. The third leaf of Thatcher (left) and RL6044 (right) is shown ten days after
Supplementary Figure 1 Phenotypic response conferred by the leaf rust resistance gene against ten Swiss P. triticina isolates. The third leaf of Thatcher (left) and RL6044 (right) is shown ten days after
Analysis of large deletions in human-chimp genomic alignments. Erika Kvikstad BioInformatics I December 14, 2004
 Analysis of large deletions in human-chimp genomic alignments Erika Kvikstad BioInformatics I December 14, 2004 Outline Mutations, mutations, mutations Project overview Strategy: finding, classifying indels
Analysis of large deletions in human-chimp genomic alignments Erika Kvikstad BioInformatics I December 14, 2004 Outline Mutations, mutations, mutations Project overview Strategy: finding, classifying indels
Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions. Supplementary Material
 Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions Joshua N. Burton 1, Andrew Adey 1, Rupali P. Patwardhan 1, Ruolan Qiu 1, Jacob O. Kitzman 1, Jay Shendure 1 1 Department
Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions Joshua N. Burton 1, Andrew Adey 1, Rupali P. Patwardhan 1, Ruolan Qiu 1, Jacob O. Kitzman 1, Jay Shendure 1 1 Department
Runs of Homozygosity Analysis Tutorial
 Runs of Homozygosity Analysis Tutorial Release 8.7.0 Golden Helix, Inc. March 22, 2017 Contents 1. Overview of the Project 2 2. Identify Runs of Homozygosity 6 Illustrative Example...............................................
Runs of Homozygosity Analysis Tutorial Release 8.7.0 Golden Helix, Inc. March 22, 2017 Contents 1. Overview of the Project 2 2. Identify Runs of Homozygosity 6 Illustrative Example...............................................
Simultaneous profiling of transcriptome and DNA methylome from a single cell
 Additional file 1: Supplementary materials Simultaneous profiling of transcriptome and DNA methylome from a single cell Youjin Hu 1, 2, Kevin Huang 1, 3, Qin An 1, Guizhen Du 1, Ganlu Hu 2, Jinfeng Xue
Additional file 1: Supplementary materials Simultaneous profiling of transcriptome and DNA methylome from a single cell Youjin Hu 1, 2, Kevin Huang 1, 3, Qin An 1, Guizhen Du 1, Ganlu Hu 2, Jinfeng Xue
CS273B: Deep Learning in Genomics and Biomedicine. Recitation 1 30/9/2016
 CS273B: Deep Learning in Genomics and Biomedicine. Recitation 1 30/9/2016 Topics Genetic variation Population structure Linkage disequilibrium Natural disease variants Genome Wide Association Studies Gene
CS273B: Deep Learning in Genomics and Biomedicine. Recitation 1 30/9/2016 Topics Genetic variation Population structure Linkage disequilibrium Natural disease variants Genome Wide Association Studies Gene
TIGR THE INSTITUTE FOR GENOMIC RESEARCH
 Introduction to Genome Annotation: Overview of What You Will Learn This Week C. Robin Buell May 21, 2007 Types of Annotation Structural Annotation: Defining genes, boundaries, sequence motifs e.g. ORF,
Introduction to Genome Annotation: Overview of What You Will Learn This Week C. Robin Buell May 21, 2007 Types of Annotation Structural Annotation: Defining genes, boundaries, sequence motifs e.g. ORF,
Genome Sequencing-- Strategies
 Genome Sequencing-- Strategies Bio 4342 Spring 04 What is a genome? A genome can be defined as the entire DNA content of each nucleated cell in an organism Each organism has one or more chromosomes that
Genome Sequencing-- Strategies Bio 4342 Spring 04 What is a genome? A genome can be defined as the entire DNA content of each nucleated cell in an organism Each organism has one or more chromosomes that
Genomic resources and gene/qtl discovery in cereals
 Genomic resources and gene/qtl discovery in cereals Roberto Tuberosa Dept. of Agroenvironmental Sciences & Technology University of Bologna, Italy The ABDC Congress 1-4 March 2010 Gudalajara, Mexico Outline
Genomic resources and gene/qtl discovery in cereals Roberto Tuberosa Dept. of Agroenvironmental Sciences & Technology University of Bologna, Italy The ABDC Congress 1-4 March 2010 Gudalajara, Mexico Outline
Genome Assembly Using de Bruijn Graphs. Biostatistics 666
 Genome Assembly Using de Bruijn Graphs Biostatistics 666 Previously: Reference Based Analyses Individual short reads are aligned to reference Genotypes generated by examining reads overlapping each position
Genome Assembly Using de Bruijn Graphs Biostatistics 666 Previously: Reference Based Analyses Individual short reads are aligned to reference Genotypes generated by examining reads overlapping each position
LECTURE 12: INSIGHTS FROM GENOME SEQUENCING
 LECTURE 12: INSIGHTS FROM GENOME SEQUENCING Read Chapter 10 (p366-375) DOE s Genomics and its implications (link at course website) STS (p359), SNP, SSR (integrates the following) Linkage, Physical, Sequence
LECTURE 12: INSIGHTS FROM GENOME SEQUENCING Read Chapter 10 (p366-375) DOE s Genomics and its implications (link at course website) STS (p359), SNP, SSR (integrates the following) Linkage, Physical, Sequence
Genomes summary. Bacterial genome sizes
 Genomes summary 1. >930 bacterial genomes sequenced. 2. Circular. Genes densely packed. 3. 2-10 Mbases, 470-7,000 genes 4. Genomes of >200 eukaryotes (45 higher ) sequenced. 5. Linear chromosomes 6. On
Genomes summary 1. >930 bacterial genomes sequenced. 2. Circular. Genes densely packed. 3. 2-10 Mbases, 470-7,000 genes 4. Genomes of >200 eukaryotes (45 higher ) sequenced. 5. Linear chromosomes 6. On
 Genome Annotation Genome annotation What is the function of each part of the genome? Where are the genes? What is the mrna sequence (transcription, splicing) What is the protein sequence? What does
Genome Annotation Genome annotation What is the function of each part of the genome? Where are the genes? What is the mrna sequence (transcription, splicing) What is the protein sequence? What does
Personal Genomics Platform White Paper Last Updated November 15, Executive Summary
 Executive Summary Helix is a personal genomics platform company with a simple but powerful mission: to empower every person to improve their life through DNA. Our platform includes saliva sample collection,
Executive Summary Helix is a personal genomics platform company with a simple but powerful mission: to empower every person to improve their life through DNA. Our platform includes saliva sample collection,
Sequence Assembly and Alignment. Jim Noonan Department of Genetics
 Sequence Assembly and Alignment Jim Noonan Department of Genetics james.noonan@yale.edu www.yale.edu/noonanlab The assembly problem >>10 9 sequencing reads 36 bp - 1 kb 3 Gb Outline Basic concepts in genome
Sequence Assembly and Alignment Jim Noonan Department of Genetics james.noonan@yale.edu www.yale.edu/noonanlab The assembly problem >>10 9 sequencing reads 36 bp - 1 kb 3 Gb Outline Basic concepts in genome
Variation detection based on second generation sequencing data. Xin LIU Department of Science and Technology, BGI
 Variation detection based on second generation sequencing data Xin LIU Department of Science and Technology, BGI liuxin@genomics.org.cn 2013.11.21 Outline Summary of sequencing techniques Data quality
Variation detection based on second generation sequencing data Xin LIU Department of Science and Technology, BGI liuxin@genomics.org.cn 2013.11.21 Outline Summary of sequencing techniques Data quality
Association mapping of Sclerotinia stalk rot resistance in domesticated sunflower plant introductions
 Association mapping of Sclerotinia stalk rot resistance in domesticated sunflower plant introductions Zahirul Talukder 1, Brent Hulke 2, Lili Qi 2, and Thomas Gulya 2 1 Department of Plant Sciences, NDSU
Association mapping of Sclerotinia stalk rot resistance in domesticated sunflower plant introductions Zahirul Talukder 1, Brent Hulke 2, Lili Qi 2, and Thomas Gulya 2 1 Department of Plant Sciences, NDSU
3. human genomics clone genes associated with genetic disorders. 4. many projects generate ordered clones that cover genome
 Lectures 30 and 31 Genome analysis I. Genome analysis A. two general areas 1. structural 2. functional B. genome projects a status report 1. 1 st sequenced: several viral genomes 2. mitochondria and chloroplasts
Lectures 30 and 31 Genome analysis I. Genome analysis A. two general areas 1. structural 2. functional B. genome projects a status report 1. 1 st sequenced: several viral genomes 2. mitochondria and chloroplasts
De novo Genome Assembly
 De novo Genome Assembly A/Prof Torsten Seemann Winter School in Mathematical & Computational Biology - Brisbane, AU - 3 July 2017 Introduction The human genome has 47 pieces MT (or XY) The shortest piece
De novo Genome Assembly A/Prof Torsten Seemann Winter School in Mathematical & Computational Biology - Brisbane, AU - 3 July 2017 Introduction The human genome has 47 pieces MT (or XY) The shortest piece
Supplementary Note: Detecting population structure in rare variant data
 Supplementary Note: Detecting population structure in rare variant data Inferring ancestry from genetic data is a common problem in both population and medical genetic studies, and many methods exist to
Supplementary Note: Detecting population structure in rare variant data Inferring ancestry from genetic data is a common problem in both population and medical genetic studies, and many methods exist to
Toward a better understanding of plant genomes structure: combining NGS and optical mapping technology to improve the sunflower assembly
 Toward a better understanding of plant genomes structure: combining NGS and optical mapping technology to improve the sunflower assembly Céline CHANTRY-DARMON 1 CNRGV The French Plant Genomic Center Created
Toward a better understanding of plant genomes structure: combining NGS and optical mapping technology to improve the sunflower assembly Céline CHANTRY-DARMON 1 CNRGV The French Plant Genomic Center Created
A tutorial introduction into the MIPS PlantsDB barley&wheat databases. Manuel Spannagl&Kai Bader transplant user training Poznan June 2013
 A tutorial introduction into the MIPS PlantsDB barley&wheat databases Manuel Spannagl&Kai Bader transplant user training Poznan June 2013 MIPS PlantsDB tutorial - some exercises Please go to: http://mips.helmholtz-muenchen.de/plant/genomes.jsp
A tutorial introduction into the MIPS PlantsDB barley&wheat databases Manuel Spannagl&Kai Bader transplant user training Poznan June 2013 MIPS PlantsDB tutorial - some exercises Please go to: http://mips.helmholtz-muenchen.de/plant/genomes.jsp
BENG 183 Trey Ideker. Genome Assembly and Physical Mapping
 BENG 183 Trey Ideker Genome Assembly and Physical Mapping Reasons for sequencing Complete genome sequencing!!! Resequencing (Confirmatory) E.g., short regions containing single nucleotide polymorphisms
BENG 183 Trey Ideker Genome Assembly and Physical Mapping Reasons for sequencing Complete genome sequencing!!! Resequencing (Confirmatory) E.g., short regions containing single nucleotide polymorphisms
Organization and evolution of transposable elements along the bread wheat chromosome 3B
 Daron et al. Genome Biology 2014, 15:546 RESEARCH Open Access Organization and evolution of transposable elements along the bread wheat chromosome 3B Josquin Daron 1,2, Natasha Glover 1,2, Lise Pingault
Daron et al. Genome Biology 2014, 15:546 RESEARCH Open Access Organization and evolution of transposable elements along the bread wheat chromosome 3B Josquin Daron 1,2, Natasha Glover 1,2, Lise Pingault
Variant Detection in Next Generation Sequencing Data. John Osborne Sept 14, 2012
 + Variant Detection in Next Generation Sequencing Data John Osborne Sept 14, 2012 + Overview My Bias Talk slanted towards analyzing whole genomes using Illumina paired end reads with open source tools
+ Variant Detection in Next Generation Sequencing Data John Osborne Sept 14, 2012 + Overview My Bias Talk slanted towards analyzing whole genomes using Illumina paired end reads with open source tools
Themes. Homo erectus. Jin and Su, Nature Reviews Genetics (2000)
 HC70A & SAS70A Winter 2009 Genetic Engineering in Medicine, Agriculture, and Law Tracking Human Ancestry Professor John Novembre Themes Global patterns of human genetic diversity Tracing our ancient ancestry
HC70A & SAS70A Winter 2009 Genetic Engineering in Medicine, Agriculture, and Law Tracking Human Ancestry Professor John Novembre Themes Global patterns of human genetic diversity Tracing our ancient ancestry
Sequence assembly. Jose Blanca COMAV institute bioinf.comav.upv.es
 Sequence assembly Jose Blanca COMAV institute bioinf.comav.upv.es Sequencing project Unknown sequence { experimental evidence result read 1 read 4 read 2 read 5 read 3 read 6 read 7 Computational requirements
Sequence assembly Jose Blanca COMAV institute bioinf.comav.upv.es Sequencing project Unknown sequence { experimental evidence result read 1 read 4 read 2 read 5 read 3 read 6 read 7 Computational requirements
Conifer Translational Genomics Network Coordinated Agricultural Project
 Conifer Translational Genomics Network Coordinated Agricultural Project Genomics in Tree Breeding and Forest Ecosystem Management ----- Module 2 Genes, Genomes, and Mendel Nicholas Wheeler & David Harry
Conifer Translational Genomics Network Coordinated Agricultural Project Genomics in Tree Breeding and Forest Ecosystem Management ----- Module 2 Genes, Genomes, and Mendel Nicholas Wheeler & David Harry
Human SNP haplotypes. Statistics 246, Spring 2002 Week 15, Lecture 1
 Human SNP haplotypes Statistics 246, Spring 2002 Week 15, Lecture 1 Human single nucleotide polymorphisms The majority of human sequence variation is due to substitutions that have occurred once in the
Human SNP haplotypes Statistics 246, Spring 2002 Week 15, Lecture 1 Human single nucleotide polymorphisms The majority of human sequence variation is due to substitutions that have occurred once in the
Einführung in die Genetik
 Einführung in die Genetik Prof. Dr. Kay Schneitz (EBio Pflanzen) http://plantdev.bio.wzw.tum.de schneitz@wzw.tum.de Prof. Dr. Claus Schwechheimer (PlaSysBiol) http://wzw.tum.de/sysbiol claus.schwechheimer@wzw.tum.de
Einführung in die Genetik Prof. Dr. Kay Schneitz (EBio Pflanzen) http://plantdev.bio.wzw.tum.de schneitz@wzw.tum.de Prof. Dr. Claus Schwechheimer (PlaSysBiol) http://wzw.tum.de/sysbiol claus.schwechheimer@wzw.tum.de
Next-Generation Sequencing. Technologies
 Next-Generation Next-Generation Sequencing Technologies Sequencing Technologies Nicholas E. Navin, Ph.D. MD Anderson Cancer Center Dept. Genetics Dept. Bioinformatics Introduction to Bioinformatics GS011062
Next-Generation Next-Generation Sequencing Technologies Sequencing Technologies Nicholas E. Navin, Ph.D. MD Anderson Cancer Center Dept. Genetics Dept. Bioinformatics Introduction to Bioinformatics GS011062
Genome annotation. Erwin Datema (2011) Sandra Smit (2012, 2013)
 Genome annotation Erwin Datema (2011) Sandra Smit (2012, 2013) Genome annotation AGACAAAGATCCGCTAAATTAAATCTGGACTTCACATATTGAAGTGATATCACACGTTTCTCTAAT AATCTCCTCACAATATTATGTTTGGGATGAACTTGTCGTGATTTGCCATTGTAGCAATCACTTGAA
Genome annotation Erwin Datema (2011) Sandra Smit (2012, 2013) Genome annotation AGACAAAGATCCGCTAAATTAAATCTGGACTTCACATATTGAAGTGATATCACACGTTTCTCTAAT AATCTCCTCACAATATTATGTTTGGGATGAACTTGTCGTGATTTGCCATTGTAGCAATCACTTGAA
RNA-Sequencing analysis
 RNA-Sequencing analysis Markus Kreuz 25. 04. 2012 Institut für Medizinische Informatik, Statistik und Epidemiologie Content: Biological background Overview transcriptomics RNA-Seq RNA-Seq technology Challenges
RNA-Sequencing analysis Markus Kreuz 25. 04. 2012 Institut für Medizinische Informatik, Statistik und Epidemiologie Content: Biological background Overview transcriptomics RNA-Seq RNA-Seq technology Challenges
% Viability. isw2 ino isw2 ino isw2 ino isw2 ino mM HU 4-NQO CPT
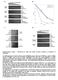 a Drug concentration b 1.3% MMS nhp1 nhp1 8 nhp1 mag1.5% MMS.3% MMS nhp1 nhp1 ino8 9 ino8 9 % Viability 4.5% MMS ino8 9 ino8 9 2.5.1.15 % MMS c d nhp1 nhp1 nhp1 nhp1 nhp1 nhp1 Control (YPD) γ IR (1 gy)
a Drug concentration b 1.3% MMS nhp1 nhp1 8 nhp1 mag1.5% MMS.3% MMS nhp1 nhp1 ino8 9 ino8 9 % Viability 4.5% MMS ino8 9 ino8 9 2.5.1.15 % MMS c d nhp1 nhp1 nhp1 nhp1 nhp1 nhp1 Control (YPD) γ IR (1 gy)
7 Gene Isolation and Analysis of Multiple
 Genetic Techniques for Biological Research Corinne A. Michels Copyright q 2002 John Wiley & Sons, Ltd ISBNs: 0-471-89921-6 (Hardback); 0-470-84662-3 (Electronic) 7 Gene Isolation and Analysis of Multiple
Genetic Techniques for Biological Research Corinne A. Michels Copyright q 2002 John Wiley & Sons, Ltd ISBNs: 0-471-89921-6 (Hardback); 0-470-84662-3 (Electronic) 7 Gene Isolation and Analysis of Multiple
Detecting ancient admixture using DNA sequence data
 Detecting ancient admixture using DNA sequence data October 10, 2008 Jeff Wall Institute for Human Genetics UCSF Background Origin of genus Homo 2 2.5 Mya Out of Africa (part I)?? 1.6 1.8 Mya Further spread
Detecting ancient admixture using DNA sequence data October 10, 2008 Jeff Wall Institute for Human Genetics UCSF Background Origin of genus Homo 2 2.5 Mya Out of Africa (part I)?? 1.6 1.8 Mya Further spread
Linking Genetic Variation to Important Phenotypes
 Linking Genetic Variation to Important Phenotypes BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2018 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material, are licensed under
Linking Genetic Variation to Important Phenotypes BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2018 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material, are licensed under
Advanced Plant Technology Program Vocabulary
 Advanced Plant Technology Program Vocabulary A Below you ll find a list of words (and their simplified definitions) that our researchers use on a daily basis. Abiotic stress (noun): Stress brought on by
Advanced Plant Technology Program Vocabulary A Below you ll find a list of words (and their simplified definitions) that our researchers use on a daily basis. Abiotic stress (noun): Stress brought on by
UC Davis UC Davis Previously Published Works
 UC Davis UC Davis Previously Published Works Title Annotation-based genome-wide SNP discovery in the large and complex Aegilops tauschii genome using next-generation sequencing without a reference genome
UC Davis UC Davis Previously Published Works Title Annotation-based genome-wide SNP discovery in the large and complex Aegilops tauschii genome using next-generation sequencing without a reference genome
4.1. Genetics as a Tool in Anthropology
 4.1. Genetics as a Tool in Anthropology Each biological system and every human being is defined by its genetic material. The genetic material is stored in the cells of the body, mainly in the nucleus of
4.1. Genetics as a Tool in Anthropology Each biological system and every human being is defined by its genetic material. The genetic material is stored in the cells of the body, mainly in the nucleus of
Comparative genomics with maize and other grasses: from genes to genomes!
 Original Paper Open Access Comparative genomics with maize and other grasses: from genes to genomes! James C. Schnable 1 and Eric Lyons 2 * 1 Department of Plant and Microbial Biology, University of California-Berkeley,
Original Paper Open Access Comparative genomics with maize and other grasses: from genes to genomes! James C. Schnable 1 and Eric Lyons 2 * 1 Department of Plant and Microbial Biology, University of California-Berkeley,
Why can GBS be complicated? Tools for filtering, error correction and imputation.
 Why can GBS be complicated? Tools for filtering, error correction and imputation. Edward Buckler USDA-ARS Cornell University http://www.maizegenetics.net Many Organisms Are Diverse Humans are at the lower
Why can GBS be complicated? Tools for filtering, error correction and imputation. Edward Buckler USDA-ARS Cornell University http://www.maizegenetics.net Many Organisms Are Diverse Humans are at the lower
Section 10.3 Outline 10.3 How Is the Base Sequence of a Messenger RNA Molecule Translated into Protein?
 Section 10.3 Outline 10.3 How Is the Base Sequence of a Messenger RNA Molecule Translated into Protein? Messenger RNA Carries Information for Protein Synthesis from the DNA to Ribosomes Ribosomes Consist
Section 10.3 Outline 10.3 How Is the Base Sequence of a Messenger RNA Molecule Translated into Protein? Messenger RNA Carries Information for Protein Synthesis from the DNA to Ribosomes Ribosomes Consist
Biotechnology Explorer
 Biotechnology Explorer C. elegans Behavior Kit Bioinformatics Supplement explorer.bio-rad.com Catalog #166-5120EDU This kit contains temperature-sensitive reagents. Open immediately and see individual
Biotechnology Explorer C. elegans Behavior Kit Bioinformatics Supplement explorer.bio-rad.com Catalog #166-5120EDU This kit contains temperature-sensitive reagents. Open immediately and see individual
SNP calling and VCF format
 SNP calling and VCF format Laurent Falquet, Oct 12 SNP? What is this? A type of genetic variation, among others: Family of Single Nucleotide Aberrations Single Nucleotide Polymorphisms (SNPs) Single Nucleotide
SNP calling and VCF format Laurent Falquet, Oct 12 SNP? What is this? A type of genetic variation, among others: Family of Single Nucleotide Aberrations Single Nucleotide Polymorphisms (SNPs) Single Nucleotide
Supplementary Table 1: Oligo designs. A list of ATAC-seq oligos used for PCR.
 Ad1_noMX: Ad2.1_TAAGGCGA Ad2.2_CGTACTAG Ad2.3_AGGCAGAA Ad2.4_TCCTGAGC Ad2.5_GGACTCCT Ad2.6_TAGGCATG Ad2.7_CTCTCTAC Ad2.8_CAGAGAGG Ad2.9_GCTACGCT Ad2.10_CGAGGCTG Ad2.11_AAGAGGCA Ad2.12_GTAGAGGA Ad2.13_GTCGTGAT
Ad1_noMX: Ad2.1_TAAGGCGA Ad2.2_CGTACTAG Ad2.3_AGGCAGAA Ad2.4_TCCTGAGC Ad2.5_GGACTCCT Ad2.6_TAGGCATG Ad2.7_CTCTCTAC Ad2.8_CAGAGAGG Ad2.9_GCTACGCT Ad2.10_CGAGGCTG Ad2.11_AAGAGGCA Ad2.12_GTAGAGGA Ad2.13_GTCGTGAT
Unit 1: DNA and the Genome Sub-topic 6: Mutation
 Unit 1: DNA and the Genome Sub-topic 6: Mutation Page 1 of 24 On completion of this topic I will be able to state that: mutations are random changes in the genome, causing no protein or an altered protein
Unit 1: DNA and the Genome Sub-topic 6: Mutation Page 1 of 24 On completion of this topic I will be able to state that: mutations are random changes in the genome, causing no protein or an altered protein
Supplementary Figures Montero et al._supplementary Figure 1
 Montero et al_suppl. Info 1 Supplementary Figures Montero et al._supplementary Figure 1 Montero et al_suppl. Info 2 Supplementary Figure 1. Transcripts arising from the structurally conserved subtelomeres
Montero et al_suppl. Info 1 Supplementary Figures Montero et al._supplementary Figure 1 Montero et al_suppl. Info 2 Supplementary Figure 1. Transcripts arising from the structurally conserved subtelomeres
Complementary Technologies for Precision Genetic Analysis
 Complementary NGS, CGH and Workflow Featured Publication Zhu, J. et al. Duplication of C7orf58, WNT16 and FAM3C in an obese female with a t(7;22)(q32.1;q11.2) chromosomal translocation and clinical features
Complementary NGS, CGH and Workflow Featured Publication Zhu, J. et al. Duplication of C7orf58, WNT16 and FAM3C in an obese female with a t(7;22)(q32.1;q11.2) chromosomal translocation and clinical features
Axiom mydesign Custom Array design guide for human genotyping applications
 TECHNICAL NOTE Axiom mydesign Custom Genotyping Arrays Axiom mydesign Custom Array design guide for human genotyping applications Overview In the past, custom genotyping arrays were expensive, required
TECHNICAL NOTE Axiom mydesign Custom Genotyping Arrays Axiom mydesign Custom Array design guide for human genotyping applications Overview In the past, custom genotyping arrays were expensive, required
Quality Measures for CytoChip Microarrays
 Quality Measures for CytoChip Microarrays How to evaluate CytoChip Oligo data quality in BlueFuse Multi software. Data quality is one of the most important aspects of any microarray experiment. This technical
Quality Measures for CytoChip Microarrays How to evaluate CytoChip Oligo data quality in BlueFuse Multi software. Data quality is one of the most important aspects of any microarray experiment. This technical
TRANSPOSON INSERTION SITE VERIFICATION
 TRANSPOSON INSERTION SITE VERIFICATION Transposon and T-DNA insertion in Arabidopsis genes can be identified using the Arabidopsis thaliana Insertion Database (ATIdb) (http://atidb.org/cgi-perl/gbrowse/atibrowse).
TRANSPOSON INSERTION SITE VERIFICATION Transposon and T-DNA insertion in Arabidopsis genes can be identified using the Arabidopsis thaliana Insertion Database (ATIdb) (http://atidb.org/cgi-perl/gbrowse/atibrowse).
CHAPTER 21 LECTURE SLIDES
 CHAPTER 21 LECTURE SLIDES Prepared by Brenda Leady University of Toledo To run the animations you must be in Slideshow View. Use the buttons on the animation to play, pause, and turn audio/text on or off.
CHAPTER 21 LECTURE SLIDES Prepared by Brenda Leady University of Toledo To run the animations you must be in Slideshow View. Use the buttons on the animation to play, pause, and turn audio/text on or off.
F. graminearum F. verticillioides
 F. oxysporum F. verticillioides Assembly (Mb) 0.0 7.0 13.9 20.9 27.9 34.8 41.8 Assembly (Mb) 0.0 10.8 21.7 32.5 43.3 54.2 65.0 0.0 6.2 12.3 18.5 24.6 30.8 37 Assembly (Mb) 0.0 7.0 13.9 20.9 27.9 34.8 41.8
F. oxysporum F. verticillioides Assembly (Mb) 0.0 7.0 13.9 20.9 27.9 34.8 41.8 Assembly (Mb) 0.0 10.8 21.7 32.5 43.3 54.2 65.0 0.0 6.2 12.3 18.5 24.6 30.8 37 Assembly (Mb) 0.0 7.0 13.9 20.9 27.9 34.8 41.8
Linkage Disequilibrium. Adele Crane & Angela Taravella
 Linkage Disequilibrium Adele Crane & Angela Taravella Overview Introduction to linkage disequilibrium (LD) Measuring LD Genetic & demographic factors shaping LD Model predictions and expected LD decay
Linkage Disequilibrium Adele Crane & Angela Taravella Overview Introduction to linkage disequilibrium (LD) Measuring LD Genetic & demographic factors shaping LD Model predictions and expected LD decay
CSE182-L16. LW statistics/assembly
 CSE182-L16 LW statistics/assembly Silly Quiz Who are these people, and what is the occasion? Genome Sequencing and Assembly Sequencing A break at T is shown here. Measuring the lengths using electrophoresis
CSE182-L16 LW statistics/assembly Silly Quiz Who are these people, and what is the occasion? Genome Sequencing and Assembly Sequencing A break at T is shown here. Measuring the lengths using electrophoresis
Supplementary Figure 1. The tree of Chinese jujube and its growing environment. The jujube has a very long lifecycle, even more than 1000 productive
 Supplementary Figure 1. The tree of Chinese jujube and its growing environment. The jujube has a very long lifecycle, even more than 1000 productive years. It is well adapted to drought and salinity. Supplementary
Supplementary Figure 1. The tree of Chinese jujube and its growing environment. The jujube has a very long lifecycle, even more than 1000 productive years. It is well adapted to drought and salinity. Supplementary
Genome-wide association studies (GWAS) Part 1
 Genome-wide association studies (GWAS) Part 1 Matti Pirinen FIMM, University of Helsinki 03.12.2013, Kumpula Campus FIMM - Institiute for Molecular Medicine Finland www.fimm.fi Published Genome-Wide Associations
Genome-wide association studies (GWAS) Part 1 Matti Pirinen FIMM, University of Helsinki 03.12.2013, Kumpula Campus FIMM - Institiute for Molecular Medicine Finland www.fimm.fi Published Genome-Wide Associations
De Novo Assembly of High-throughput Short Read Sequences
 De Novo Assembly of High-throughput Short Read Sequences Chuming Chen Center for Bioinformatics and Computational Biology (CBCB) University of Delaware NECC Third Skate Genome Annotation Workshop May 23,
De Novo Assembly of High-throughput Short Read Sequences Chuming Chen Center for Bioinformatics and Computational Biology (CBCB) University of Delaware NECC Third Skate Genome Annotation Workshop May 23,
Genome Sequencing and Structural Variation
 Genome Sequencing and Structural Variation Institut für Medizinische Genetik und Humangenetik Charité Universitätsmedizin Berlin Genomics: Lecture #10 Today Structural Variation Deletions Duplications
Genome Sequencing and Structural Variation Institut für Medizinische Genetik und Humangenetik Charité Universitätsmedizin Berlin Genomics: Lecture #10 Today Structural Variation Deletions Duplications
Integrating and Leveraging Cereal and Grass Genome Resources. David Marshall Information and Computational Sciences Group
 Integrating and Leveraging Cereal and Grass Genome Resources David Marshall Information and Computational Sciences Group David.Marshall@hutton.ac.uk What I m going to cover Brief overview of the new barley
Integrating and Leveraging Cereal and Grass Genome Resources David Marshall Information and Computational Sciences Group David.Marshall@hutton.ac.uk What I m going to cover Brief overview of the new barley
Association Mapping in Plants PLSC 731 Plant Molecular Genetics Phil McClean April, 2010
 Association Mapping in Plants PLSC 731 Plant Molecular Genetics Phil McClean April, 2010 Traditional QTL approach Uses standard bi-parental mapping populations o F2 or RI These have a limited number of
Association Mapping in Plants PLSC 731 Plant Molecular Genetics Phil McClean April, 2010 Traditional QTL approach Uses standard bi-parental mapping populations o F2 or RI These have a limited number of
Contrasting Evolutionary Patterns of the Rp1 Resistance Gene Family in Different Species of Poaceae. Research article. Abstract.
 Contrasting Evolutionary Patterns of the Rp1 Resistance Gene Family in Different Species of Poaceae Sha Luo,,1 Junhua Peng,,2 Kunpeng Li, 1 Min Wang, 1 and Hanhui Kuang*,1 1 Key Laboratory of Horticulture
Contrasting Evolutionary Patterns of the Rp1 Resistance Gene Family in Different Species of Poaceae Sha Luo,,1 Junhua Peng,,2 Kunpeng Li, 1 Min Wang, 1 and Hanhui Kuang*,1 1 Key Laboratory of Horticulture
Mutations, Genetic Testing and Engineering
 Mutations, Genetic Testing and Engineering Objectives Describe how techniques such as DNA fingerprinting, genetic modifications, and chromosomal analysis are used to study the genomes of organisms (TEKS
Mutations, Genetic Testing and Engineering Objectives Describe how techniques such as DNA fingerprinting, genetic modifications, and chromosomal analysis are used to study the genomes of organisms (TEKS
Introduction to RNA-Seq. David Wood Winter School in Mathematics and Computational Biology July 1, 2013
 Introduction to RNA-Seq David Wood Winter School in Mathematics and Computational Biology July 1, 2013 Abundance RNA is... Diverse Dynamic Central DNA rrna Epigenetics trna RNA mrna Time Protein Abundance
Introduction to RNA-Seq David Wood Winter School in Mathematics and Computational Biology July 1, 2013 Abundance RNA is... Diverse Dynamic Central DNA rrna Epigenetics trna RNA mrna Time Protein Abundance
Single Nucleotide Variant Analysis. H3ABioNet May 14, 2014
 Single Nucleotide Variant Analysis H3ABioNet May 14, 2014 Outline What are SNPs and SNVs? How do we identify them? How do we call them? SAMTools GATK VCF File Format Let s call variants! Single Nucleotide
Single Nucleotide Variant Analysis H3ABioNet May 14, 2014 Outline What are SNPs and SNVs? How do we identify them? How do we call them? SAMTools GATK VCF File Format Let s call variants! Single Nucleotide
The Polymerase Chain Reaction. Chapter 6: Background
 The Polymerase Chain Reaction Chapter 6: Background Invention of PCR Kary Mullis Mile marker 46.58 in April of 1983 Pulled off the road and outlined a way to conduct DNA replication in a tube Worked for
The Polymerase Chain Reaction Chapter 6: Background Invention of PCR Kary Mullis Mile marker 46.58 in April of 1983 Pulled off the road and outlined a way to conduct DNA replication in a tube Worked for
Measurement of Molecular Genetic Variation. Forces Creating Genetic Variation. Mutation: Nucleotide Substitutions
 Measurement of Molecular Genetic Variation Genetic Variation Is The Necessary Prerequisite For All Evolution And For Studying All The Major Problem Areas In Molecular Evolution. How We Score And Measure
Measurement of Molecular Genetic Variation Genetic Variation Is The Necessary Prerequisite For All Evolution And For Studying All The Major Problem Areas In Molecular Evolution. How We Score And Measure
Expressed sequence tag-pcr markers for identification of alien barley chromosome 2H in wheat
 Expressed sequence tag-pcr markers for identification of alien barley chromosome 2H in wheat M.J. Wang 1 *, H.D. Zou 1 *, Z.S. Lin 2, Y. Wu 1, X. Chen 2 and Y.P. Yuan 1 1 College of Plant Science, Jilin
Expressed sequence tag-pcr markers for identification of alien barley chromosome 2H in wheat M.J. Wang 1 *, H.D. Zou 1 *, Z.S. Lin 2, Y. Wu 1, X. Chen 2 and Y.P. Yuan 1 1 College of Plant Science, Jilin
de novo sequencing of the sunflower genome
 de novo sequencing of the sunflower genome Stéphane Muños LIPM INRA Toulouse stephane.munos@toulouse.inra.fr @stephane_munos @SUNRISE_France 39 Sunflower, an important cropfor Europe Million tons of seed
de novo sequencing of the sunflower genome Stéphane Muños LIPM INRA Toulouse stephane.munos@toulouse.inra.fr @stephane_munos @SUNRISE_France 39 Sunflower, an important cropfor Europe Million tons of seed
DNA Collection. Data Quality Control. Whole Genome Amplification. Whole Genome Amplification. Measure DNA concentrations. Pros
 DNA Collection Data Quality Control Suzanne M. Leal Baylor College of Medicine sleal@bcm.edu Copyrighted S.M. Leal 2016 Blood samples For unlimited supply of DNA Transformed cell lines Buccal Swabs Small
DNA Collection Data Quality Control Suzanne M. Leal Baylor College of Medicine sleal@bcm.edu Copyrighted S.M. Leal 2016 Blood samples For unlimited supply of DNA Transformed cell lines Buccal Swabs Small
SNPs - GWAS - eqtls. Sebastian Schmeier
 SNPs - GWAS - eqtls s.schmeier@gmail.com http://sschmeier.github.io/bioinf-workshop/ 17.08.2015 Overview Single nucleotide polymorphism (refresh) SNPs effect on genes (refresh) Genome-wide association
SNPs - GWAS - eqtls s.schmeier@gmail.com http://sschmeier.github.io/bioinf-workshop/ 17.08.2015 Overview Single nucleotide polymorphism (refresh) SNPs effect on genes (refresh) Genome-wide association
Introducing new DNA into the genome requires cloning the donor sequence, delivery of the cloned DNA into the cell, and integration into the genome.
 Key Terms Chapter 32: Genetic Engineering Cloning describes propagation of a DNA sequence by incorporating it into a hybrid construct that can be replicated in a host cell. A cloning vector is a plasmid
Key Terms Chapter 32: Genetic Engineering Cloning describes propagation of a DNA sequence by incorporating it into a hybrid construct that can be replicated in a host cell. A cloning vector is a plasmid
Chapter 15 Gene Technologies and Human Applications
 Chapter Outline Chapter 15 Gene Technologies and Human Applications Section 1: The Human Genome KEY IDEAS > Why is the Human Genome Project so important? > How do genomics and gene technologies affect
Chapter Outline Chapter 15 Gene Technologies and Human Applications Section 1: The Human Genome KEY IDEAS > Why is the Human Genome Project so important? > How do genomics and gene technologies affect
Development of Genomic Tools for RKN Resistance Breeding in Cotton
 Development of Genomic Tools for RKN Resistance Breeding in Cotton Dr. Hongbin Zhang Department of Soil & Crop Sciences and Institute for Plant Genomics & Biotechnology Texas A&M University, College Station,
Development of Genomic Tools for RKN Resistance Breeding in Cotton Dr. Hongbin Zhang Department of Soil & Crop Sciences and Institute for Plant Genomics & Biotechnology Texas A&M University, College Station,
Mutations during meiosis and germ line division lead to genetic variation between individuals
 Mutations during meiosis and germ line division lead to genetic variation between individuals Types of mutations: point mutations indels (insertion/deletion) copy number variation structural rearrangements
Mutations during meiosis and germ line division lead to genetic variation between individuals Types of mutations: point mutations indels (insertion/deletion) copy number variation structural rearrangements
mrna for protein translation
 Biology 1B Evolution Lecture 5 (March 5, 2010), Genetic Drift and Migration Mutation What is mutation? Changes in the coding sequence Changes in gene regulation, or how the genes are expressed as amino
Biology 1B Evolution Lecture 5 (March 5, 2010), Genetic Drift and Migration Mutation What is mutation? Changes in the coding sequence Changes in gene regulation, or how the genes are expressed as amino
Genomics Based Approaches to Genetic Improvement in Sugarcane. Robert Henry
 Genomics Based Approaches to Genetic Improvement in Sugarcane Robert Henry Centre for Plant Conservation Genetics Life Enriched by Plant Biodiversity Food Biomass options Sorghum Sugarcane Grasses Shrubs
Genomics Based Approaches to Genetic Improvement in Sugarcane Robert Henry Centre for Plant Conservation Genetics Life Enriched by Plant Biodiversity Food Biomass options Sorghum Sugarcane Grasses Shrubs
Transcriptome sequencing of two wild barley (Hordeum spontaneum L.) ecotypes differentially adapted to drought stress reveals ecotypespecific
 Bedada et al. BMC Genomics 2014, 15:995 RESEARCH ARTICLE Open Access Transcriptome sequencing of two wild barley (Hordeum spontaneum L.) ecotypes differentially adapted to drought stress reveals ecotypespecific
Bedada et al. BMC Genomics 2014, 15:995 RESEARCH ARTICLE Open Access Transcriptome sequencing of two wild barley (Hordeum spontaneum L.) ecotypes differentially adapted to drought stress reveals ecotypespecific
Understanding Genes & Mutations. John A Phillips III May 16, 2005
 Understanding Genes & Mutations John A Phillips III May 16, 2005 Learning Objectives Understand gene structure Become familiar with genetic & mutation databases Be able to find information on genetic variation
Understanding Genes & Mutations John A Phillips III May 16, 2005 Learning Objectives Understand gene structure Become familiar with genetic & mutation databases Be able to find information on genetic variation
SUPPLEMENTAL MATERIALS
 SUPPLEMENL MERILS Eh-seq: RISPR epitope tagging hip-seq of DN-binding proteins Daniel Savic, E. hristopher Partridge, Kimberly M. Newberry, Sophia. Smith, Sarah K. Meadows, rian S. Roberts, Mark Mackiewicz,
SUPPLEMENL MERILS Eh-seq: RISPR epitope tagging hip-seq of DN-binding proteins Daniel Savic, E. hristopher Partridge, Kimberly M. Newberry, Sophia. Smith, Sarah K. Meadows, rian S. Roberts, Mark Mackiewicz,
Re-sequencing and hybrid assembly strategy of two nematode resistant Beta vulgaris translocation lines. Sarah Christina Jäger
 Faculty of Agricultural and Nutritional Sciences Re-sequencing and hybrid assembly strategy of two nematode resistant Beta vulgaris translocation lines Plant Breeding Institute Sarah Christina Jäger Plant
Faculty of Agricultural and Nutritional Sciences Re-sequencing and hybrid assembly strategy of two nematode resistant Beta vulgaris translocation lines Plant Breeding Institute Sarah Christina Jäger Plant
Next Generation Sequencing Technologies. Rob Mitra 1/30/17
 Next Generation Sequencing Technologies Rob Mitra 1/30/17 Outline Overview of next-generation sequencing How does it work? What technologies are being used? How would one use it in practice? Math basic
Next Generation Sequencing Technologies Rob Mitra 1/30/17 Outline Overview of next-generation sequencing How does it work? What technologies are being used? How would one use it in practice? Math basic
Overview of Human Genetics
 Overview of Human Genetics 1 Structure and function of nucleic acids. 2 Structure and composition of the human genome. 3 Mendelian genetics. Lander et al. (Nature, 2001) MAT 394 (ASU) Human Genetics Spring
Overview of Human Genetics 1 Structure and function of nucleic acids. 2 Structure and composition of the human genome. 3 Mendelian genetics. Lander et al. (Nature, 2001) MAT 394 (ASU) Human Genetics Spring
4/26/2015. Cut DNA either: Cut DNA either:
 Ch.20 Enzymes that cut DNA at specific sequences (restriction sites) resulting in segments of DNA (restriction fragments) Typically 4-8 bp in length & often palindromic Isolated from bacteria (Hundreds
Ch.20 Enzymes that cut DNA at specific sequences (restriction sites) resulting in segments of DNA (restriction fragments) Typically 4-8 bp in length & often palindromic Isolated from bacteria (Hundreds
Recent technology allow production of microarrays composed of 70-mers (essentially a hybrid of the two techniques)
 Microarrays and Transcript Profiling Gene expression patterns are traditionally studied using Northern blots (DNA-RNA hybridization assays). This approach involves separation of total or polya + RNA on
Microarrays and Transcript Profiling Gene expression patterns are traditionally studied using Northern blots (DNA-RNA hybridization assays). This approach involves separation of total or polya + RNA on
Introduction to Plant Genome and Gene Structure. Dr. Jonaliza L. Siangliw Rice Gene Discovery Unit, BIOTEC
 Introduction to Plant Genome and Gene Structure Dr. Jonaliza L. Siangliw Rice Gene Discovery Unit, BIOTEC Defining genome Coined by Hans Winkler (1920) as genom(e) by joining gene and chromosome Lederberg
Introduction to Plant Genome and Gene Structure Dr. Jonaliza L. Siangliw Rice Gene Discovery Unit, BIOTEC Defining genome Coined by Hans Winkler (1920) as genom(e) by joining gene and chromosome Lederberg
ChIP-Seq Data Analysis. J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015
 ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA
ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA
Answer: Sequence overlap is required to align the sequenced segments relative to each other.
 14 Genomes and Genomics WORKING WITH THE FIGURES 1. Based on Figure 14-2, why must the DNA fragments sequenced overlap in order to obtain a genome sequence? Answer: Sequence overlap is required to align
14 Genomes and Genomics WORKING WITH THE FIGURES 1. Based on Figure 14-2, why must the DNA fragments sequenced overlap in order to obtain a genome sequence? Answer: Sequence overlap is required to align
Champions of the Poor of the Semi-Arid Tropics
 ICRISAT and Partners Champions of the Poor of the Semi-Arid Tropics William D. Dar Director General, ICRISAT This presentation The context Our research heartland Major outputs and impacts Moving into the
ICRISAT and Partners Champions of the Poor of the Semi-Arid Tropics William D. Dar Director General, ICRISAT This presentation The context Our research heartland Major outputs and impacts Moving into the
A clone-free, single molecule map of the domestic cow (Bos taurus) genome
 Zhou et al. BMC Genomics (2) 16:644 DOI 1.1186/s12864--1823-7 RESEARCH ARTICLE Open Access A clone-free, single molecule map of the domestic cow (Bos taurus) genome Shiguo Zhou 1, Steve Goldstein 1, Michael
Zhou et al. BMC Genomics (2) 16:644 DOI 1.1186/s12864--1823-7 RESEARCH ARTICLE Open Access A clone-free, single molecule map of the domestic cow (Bos taurus) genome Shiguo Zhou 1, Steve Goldstein 1, Michael
Percent survival. Supplementary fig. S3 A.
 Supplementary fig. S3 A. B. 100 Percent survival 80 60 40 20 Ml 0 0 100 C. Fig. S3 Comparison of leukaemia incidence rate in the triple targeted chimaeric mice and germline-transmission translocator mice
Supplementary fig. S3 A. B. 100 Percent survival 80 60 40 20 Ml 0 0 100 C. Fig. S3 Comparison of leukaemia incidence rate in the triple targeted chimaeric mice and germline-transmission translocator mice
Petar Pajic 1 *, Yen Lung Lin 1 *, Duo Xu 1, Omer Gokcumen 1 Department of Biological Sciences, University at Buffalo, Buffalo, NY.
 The psoriasis associated deletion of late cornified envelope genes LCE3B and LCE3C has been maintained under balancing selection since Human Denisovan divergence Petar Pajic 1 *, Yen Lung Lin 1 *, Duo
The psoriasis associated deletion of late cornified envelope genes LCE3B and LCE3C has been maintained under balancing selection since Human Denisovan divergence Petar Pajic 1 *, Yen Lung Lin 1 *, Duo
Genome Annotation. What Does Annotation Describe??? Genome duplications Genes Mobile genetic elements Small repeats Genetic diversity
 Genome Annotation Genome Sequencing Costliest aspect of sequencing the genome o But Devoid of content Genome must be annotated o Annotation definition Analyzing the raw sequence of a genome and describing
Genome Annotation Genome Sequencing Costliest aspect of sequencing the genome o But Devoid of content Genome must be annotated o Annotation definition Analyzing the raw sequence of a genome and describing
