3.5b. Regulation of Gene Expression CAMPBELL BIOLOGY IN FOCUS. Urry Cain Wasserman Minorsky Jackson Reece
|
|
|
- Cameron Barnett
- 6 years ago
- Views:
Transcription
1 CAMPBELL BIOLOGY IN FOCUS Urry Cain Wasserman Minorsky Jackson Reece 3.5b Regulation of Gene Expression Lecture Presentations by Kathleen Fitzpatrick and Nicole Tunbridge
2 Overview: Differential Expression of Genes Prokaryotes and eukaryotes alter gene expression in response to their changing environment Multicellular eukaryotes also develop and maintain multiple cell types Gene expression is often regulated at the transcription stage, but control at other stages is important, too
3 Figure 15.1
4 Concept 15.1: Bacteria often respond to environmental change by regulating transcription Natural selection has favored bacteria that produce only the gene products needed by the cell A cell can regulate the production of enzymes by feedback inhibition or by gene regulation Gene expression in bacteria is controlled by a mechanism described as the operon model
5 Figure 15.2 Precursor Feedback inhibition trpe gene Enzyme 1 trpd gene Enzyme 2 Regulation of gene expression trpc gene trpb gene Enzyme 3 trpa gene Tryptoph an (a) Regulation of enzyme activity (b) Regulation of enzyme production
6 Operons: The Basic Concept A group of functionally related genes can be coordinately controlled by a single on-off switch The regulatory switch is a segment of DNA called an operator usually positioned within the promoter An operon is the entire stretch of DNA that includes the operator, the promoter, and the genes that they control
7 The operon can be switched off by a protein repressor The repressor prevents gene transcription by binding to the operator and blocking RNA polymerase The repressor is the product of a separate regulatory gene
8 The repressor can be in an active or inactive form, depending on the presence of other molecules A corepressor is a molecule that cooperates with a repressor protein to switch an operon off For example, E. coli can synthesize the amino acid tryptophan
9 By default the trp operon is on and the genes for tryptophan synthesis are transcribed When tryptophan is present, it binds to the trp repressor protein, which then turns the operon off The repressor is active only in the presence of its corepressor tryptophan; thus the trp operon is turned off (repressed) if tryptophan levels are high
10 Figure 15.3 trp operon Promoter Promoter Genes of operon DNA trpe trpr Regulator y gene mrn 5 A 3 Operator RNA Start polymeras codon e mrna 5 trpd trpc trpb trpa C B A Stop codon E D Prote Inactive Polypeptide subunits that make in repressor up (a) Tryptophan absent, repressor inactive, operon onenzymes for tryptophan synthesis DNA No RNA mad e mrn A Prote in Tryptophan (corepressor) Activ e repre ssor (b) Tryptophan present, repressor active, operon off
11 Figure 15.3a DN A Promot er trp R mr NA Regulat ory gene 3 5 Prot ein Promot er Operat RNA or Start polymer codon mrna ase 5 trp operon Genes of trp operon trp E D Stop codon E D trp C trp B trp A C B A Inactive Polypeptide subunits that repress make up or enzymes for tryptophan (a) Tryptophan absent, repressor inactive, synthesis operon on
12 Figure 15.3b DN A mr NA Prot ein No RN A ma de Acti ve Tryptopha repr n ess (b) Tryptophan(corepress present, repressor active, or operon off or)
13 Repressible and Inducible Operons: Two Types of Negative Gene Regulation A repressible operon is one that is usually on; binding of a repressor to the operator shuts off transcription The trp operon is a repressible operon An inducible operon is one that is usually off; a molecule called an inducer inactivates the repressor and turns on transcription
14 The lac operon is an inducible operon and contains genes that code for enzymes used in the hydrolysis and metabolism of lactose By itself, the lac repressor is active and switches the lac operon off A molecule called an inducer inactivates the repressor to turn the lac operon on
15 For the lac operon, the inducer is allolactose, formed from lactose that enters the cell Enzymes of the lactose pathway are called inducible enzymes Analogously, the enzymes for tryptophan synthesis are said to be repressible enzymes
16 Figure 15.4 Regulator y gene DNA laci mrn A Promoter Operator IacZ No RNA made 3 RNA polymerase 5 Active represso (a) Lactose absent,rrepressor active, operon off Protein DNA lac oper on IacZ IacI IacY IacA RNA polymerase mrn A Protein 3 5 mrna 5 β-galactosidase Inactive Allolacto represso se r (inducer) (b) Lactose present, repressor inactive, operon on Education, Inc Pearson Permease Transacet ylase
17 Figure 15.4a Regulat ory gene DN laci A Promot er Operat or Iac Z No RNA mad e 3 mr RNA 5 NA polymera se Active Prote repres in sor (a) Lactose absent, repressor active, operon off
18 Figure 15.4b DN A lac ope ron Iac Iac I mr NA Z 3 5 Prote in Iac A RNA polymerase mrna 5 Inactiv e repres sor Allolac tose (induc present, repressor (b) Lactose er) operon on inactive, Iac Y βgalactosida se Permea se Transac etylase
19 Inducible enzymes usually function in catabolic pathways; their synthesis is induced by a chemical signal Repressible enzymes usually function in anabolic pathways; their synthesis is repressed by high levels of the end product Regulation of the trp and lac operons involves negative control of genes because operons are switched off by the active form of the repressor
20 Positive Gene Regulation E. coli will preferentially use glucose when it is present in the environment When glucose is scarce, CAP (catabolite activator protein) acts as an activator of transcription CAP is activated by binding with cyclic AMP (camp) Activated CAP attaches to the promoter of the lac operon and increases the affinity of RNA polymerase, thus accelerating transcription
21 When glucose levels increase, CAP detaches from the lac operon, and transcription proceeds at a very low rate, even if lactose is present CAP helps regulate other operons that encode enzymes used in catabolic pathways
22 Figure 15.5 Promoter DNA laci CAP-binding site cam P IacZ Operator RNA polymerase Active binds and transcribes CAP Inactive Inactiv lac e repressor CAP Allolactos e scarce (camp level high): (a) Lactose present, glucose abundant lac mrna synthesized Promoter DNA laci IacZ Operator RNA polym erase less Inactiv likely Inactive e to lac CAP bind repressor (b) Lactose present, glucose present (camp level low): little lac mrna synthesized CAP-binding site
23 Figure 15.5a Prom oter DN lac A I CAP-binding site camp Ia cz Oper Activ e CAP Inactiv e CAP Allolactose (a) Lactose present, glucose abundant scarce (camplac level high): mrna synthesized RNA polymer ator ase binds and transcri Inactive lac bes repressor
24 Figure 15.5b Prom oter DN A CAPbinding site lac I Inac tive CAP (b) Lactose present, glucose present level low): little (camp lac mrna synthesized RN A pol ym era se les s like ly to bin Ia cz Oper ator Inacti ve lac repre ssor
25 Concept 15.2: Eukaryotic gene expression is regulated at many stages All organisms must regulate which genes are expressed at any given time In multicellular organisms regulation of gene expression is essential for cell specialization
26 Differential Gene Expression Almost all the cells in an organism are genetically identical Differences between cell types result from differential gene expression, the expression of different genes by cells with the same genome Abnormalities in gene expression can lead to diseases, including cancer Gene expression is regulated at many stages
27 Figure 15.6 Signa l NUCLEUS Chromatin Chromatin modification: DNA unpacking involving histone acetylation and Gene available DNA demethylation for DNA Gen e RNA transcription Transcriptio n Exo n Primary transcript Intro n RNA processing Tail Cap mrna in nucleus Transport to cytoplasm CYTOPLASM mrna in cytoplasm Degradation of mrna Translation Polypeptide Protein processing, such as cleavage and chemical modification Degradatio n of protein Active protein Transport to cellular destination Cellular function (such as enzymatic activity, structural support)
28 Figure 15.6a Si gn al D N A R N A C ap NUC Chro LEUS matin Chromatin modification : DNA unpacking Gene involving availabl histone G e acetylation en Transcfor and e transcri ription E DNA ption xo demethylati Primar y on In n tr RNA transc ript o processi Ta n ng mrna in il nucleus Transport to cytoplasm CYTOPLAS M
29 Figure 15.6b CYTOPLAS mrna in M cytoplasm Degra dation of mrna Degra datio n of protei n Transl ation Polype ptide Protein processing, such as cleavage Active and chemical protei modification n Transpo rt to cellular Cellular destinati functionon (such as enzymatic activity, structural
30 In all organisms, a common control point for gene expression is at transcription The greater complexity of eukaryotic cell structure and function provides opportunities for regulating gene expression at many additional stages
31 Regulation of Chromatin Structure The structural organization of chromatin packs DNA into a compact form and also helps regulate gene expression in several ways The location of a gene promoter relative to nucleosomes and scaffold or lamina attachment sites can influence gene transcription
32 Genes within highly condensed heterochromatin are usually not expressed Chemical modifications to histone proteins and DNA can influence chromatin structure and gene expression
33 Histone Modifications and DNA Methylation In histone acetylation, acetyl groups are attached to positively charged lysines in histone tails This generally loosens chromatin structure, promoting the initiation of transcription The addition of methyl groups (methylation) can condense chromatin and lead to reduced transcription
34 Figure 15.7 Nucleosome Unacetylated histones Histone tails Acetylated histones
35 DNA methylation is the addition of methyl groups to certain bases in DNA, usually cytosine Individual genes are usually more heavily methylated in cells where they are not expressed Once methylated, genes usually remain so through successive cell divisions After replication, enzymes methylate the correct daughter strand so that the methylation pattern is inherited
36 Epigenetic Inheritance Though chromatin modifications do not alter DNA sequence, they may be passed to future generations of cells The inheritance of traits transmitted by mechanisms not directly involving the nucleotide sequence is called epigenetic inheritance Epigenetic modifications can be reversed, unlike mutations in DNA sequence
37 Regulation of Transcription Initiation Chromatin-modifying enzymes provide initial control of gene expression by making a region of DNA either more or less able to bind the transcription machinery
38 Organization of a Typical Eukaryotic Gene Associated with most eukaryotic genes are multiple control elements, segments of noncoding DNA that serve as binding sites for transcription factors that help regulate transcription Control elements and the transcription factors they bind are critical for the precise regulation of gene expression in different cell types
39 Animation: mrna Degradation Right click slide / Select play
40 Figure 15.8 Enhancer (distal control elements) Proximal control elements Transcripti on start site Exon DNA Upstream Intro n Prom oter Primary RNA transcript (pre-mrna) Exon 5 Intro n Exon Tran scrip tion Exon Intro n G P P 5 Ca p Exon Coding segment P 5 UT R Transcript ion terminatio n region Dow nstrea Poly-A m signal Cleaved 3 end of primary transcript RNA processing Intron RNA mrna Poly-A signal sequence Intro Exon n Sta rt cod on Sto p cod on AAA AA A 3 UT R Pol y-a tail 3
41 Figure 15.8a DN A Enhance r (distal control element Upstre s) am Proxi Transcr mal iption contr start Ex ol site on eleme nts Pro mo ter Intr on Ex on Poly-A signal seque Intrnce Ex on on Transc ription termin ation region Downstre am
42 Figure 15.8b-1 Proxi mal contro l eleme nts DN A Transcr iption start Ex site on Pro mo ter Intr on Ex on Poly-A signal sequen Intrce Ex on on
43 Figure 15.8b-2 Proxi mal contro l eleme nts DN A Primary RNA transcri pt (premrna) Transcr iption start Ex site on Intr on Pro mo ter 5 Ex on Intr on Ex on Poly-A signal sequen Intrce Ex on Tra nsc ript Ex ion Intr on on on PolyA signa Ex l on Cleaved 3 end of primary transcri pt
44 Figure 15.8b-3 Proxi mal contro l eleme nts DN A Primary RNA transcri pt (premrna) mr NA Transcr iption start Ex site on Intr on Pro mo ter 5 Ex on Intr on on Tra nsc ript Ex ion Intr on on RNA process ing Intron RNA G P P P 5 Ca p Ex on Poly-A signal sequen Intrce Ex 5 U T Coding segmen t St art co on PolyA signa Ex l on Cleaved 3 end of primary transcri pt AAA AAA St op co 3 U T Po ly- 3
45 The Roles of Transcription Factors To initiate transcription, eukaryotic RNA polymerase requires the assistance of proteins called transcription factors General transcription factors are essential for the transcription of all protein-coding genes In eukaryotes, high levels of transcription of particular genes depend on interaction between control elements and specific transcription factors
46 Enhancers and Specific Transcription Factors Proximal control elements are located close to the promoter Distal control elements, groupings of which are called enhancers, may be far away from a gene or even located in an intron
47 An activator is a protein that binds to an enhancer and stimulates transcription of a gene Activators have two domains, one that binds DNA and a second that activates transcription Bound activators facilitate a sequence of proteinprotein interactions that result in transcription of a given gene
48 Figure 15.9 Activation domain DNA DNA-binding domain
49 Bound activators are brought into contact with a group of mediator proteins through DNA bending The mediator proteins in turn interact with proteins at the promoter These protein-protein interactions help to assemble and position the initiation complex on the promoter
50 Animation: Transcription Initiation Right click slide / Select play
51 Figure 15.UN01 Chromatin modification Transcription RNA processing mrna degradation Translation Protein processing and degradation
52 Figure Activators DN A Enhancer Distal control element Promoter TATA box Gene
53 Figure Promoter Activators DN A Enhancer Distal control element DNAbending protein Gene TATA box General transcription factors Group of mediator proteins
54 Figure Promoter Activators DN A Enhancer Distal control element DNAbending protein Gene TATA box General transcription factors Group of mediator proteins RNA polymerase II RNA polymerase II Transcription initiation complex RNA synthesis
55 Some transcription factors function as repressors, inhibiting expression of a particular gene by a variety of methods Some activators and repressors act indirectly by influencing chromatin structure to promote or silence transcription
56 Combinatorial Control of Gene Activation A particular combination of control elements can activate transcription only when the appropriate activator proteins are present
57 Figure Enhancer Control element s Enhancer (a) LIVER CELL NUCLEUS Available activators Albumin gene expresse d Crystallin gene not expresse Promoter Albumin gene Promoter Crystallin gene (b) LENS CELL NUCLEUS Available activators Albumin gene not expresse d Crystallin gene expresse d
58 Figure 15.11a (a) LIVER CELL NUCLEUS Availa ble activat ors Albumi n gene expres sed Crystal lin gene
59 Figure 15.11b (b) LENS CELL NUCLEUS Availa ble activat ors Albumi n gene not expres sed Crystal lin gene
60 Coordinately Controlled Genes in Eukaryotes Unlike the genes of a prokaryotic operon, each of the co-expressed eukaryotic genes has a promoter and control elements These genes can be scattered over different chromosomes, but each has the same combination of control elements Copies of the activators recognize specific control elements and promote simultaneous transcription of the genes
61 Mechanisms of Post-Transcriptional Regulation Transcription alone does not account for gene expression Regulatory mechanisms can operate at various stages after transcription Such mechanisms allow a cell to fine-tune gene expression rapidly in response to environmental changes
62 RNA Processing In alternative RNA splicing, different mrna molecules are produced from the same primary transcript, depending on which RNA segments are treated as exons and which as introns
63 Figure 15.UN02 Chromatin modification Transcription RNA processing mrna degradation Translation Protein processing and degradation
64 Figure Exons DNA Troponin T gene Primary RNA transcript RNA splicing mrna or
65 mrna Degradation The life span of mrna molecules in the cytoplasm is important in determining the pattern of protein synthesis in a cell Eukaryotic mrna generally survives longer than prokaryotic mrna Nucleotide sequences that influence the life span of mrna in eukaryotes reside in the untranslated region (UTR) at the 3 end of the molecule
66 Initiation of Translation The initiation of translation of selected mrnas can be blocked by regulatory proteins that bind to sequences or structures of the mrna Alternatively, translation of all mrnas in a cell may be regulated simultaneously For example, translation initiation factors are simultaneously activated in an egg following fertilization
67 Protein Processing and Degradation After translation, various types of protein processing, including cleavage and chemical modification, are subject to control The length of time each protein functions in a cell is regulated by means of selective degradation To mark a particular protein for destruction, the cell commonly attaches molecules of ubiquitin to the protein, which triggers its destruction
68 Concept 15.3: Noncoding RNAs play multiple roles in controlling gene expression Only a small fraction of DNA encodes proteins, and a very small fraction of the non-protein-coding DNA consists of genes for RNA such as rrna and trna A significant amount of the genome may be transcribed into noncoding RNAs (ncrnas) Noncoding RNAs regulate gene expression at several points
69 Effects on mrnas by MicroRNAs and Small Interfering RNAs MicroRNAs (mirnas) are small single-stranded RNA molecules that can bind to complementary mrna sequences These can degrade the mrna or block its translation
70 Figure 15.UN03 Chromatin modification Transcription RNA processing mrna degradation Translation Protein processing and degradation
71 Figure mirna mirnaprotein complex 1 The mirna binds to a target mrna. mrna degraded Translation blocked 2 If bases are completely complementary, mrna is degraded. If match is less than complete, translation is blocked Pearson Education, Inc.
72 Another class of small RNAs are called small interfering RNAs (sirnas) sirnas and mirnas are similar but form from different RNA precursors The phenomenon of inhibition of gene expression by sirnas is called RNA interference (RNAi)
73 Chromatin Remodeling and Effects on Transcription by ncrnas In some yeasts RNA produced from centromeric DNA is copied into double-stranded RNA and then processed into sirnas The sirnas, together with a complex of proteins, act as a homing device to target transcripts being made from centromeric sequences Proteins in the complex then recruit enzymes that modify the chromatin to form the highly condensed heterochromatin found at the centromere
74 A class of small ncrnas called piwi-associated RNAs (pirnas) also induce formation of heterochromatin They block expression of transposons, parasitic DNA elements in the genome The role of ncrnas adds to the complexity of the processes involved in regulation of gene expression
75 Concept 15.4: Researchers can monitor expression of specific genes Cells of a given multicellular organism differ from each other because they express different genes from an identical genome The most straightforward way to discover which genes are expressed by cells of interest is to identify the mrnas being made
76 Studying the Expression of Single Genes We can detect mrna in a cell using nucleic acid hybridization, the base pairing of a strand of nucleic acid to its complementary sequence The complementary molecule in this case is a short single-stranded DNA or RNA called a nucleic acid probe Each probe is labeled with a fluorescent tag to allow visualization
77 The technique allows us to see the mrna in place (in situ) in the intact organism and is thus called in situ hybridization
78 Figure μm
79 Another widely used method for comparing the amounts of specific mrnas in several different samples is reverse transcriptase polymerase chain reaction (RT-PCR) RT-PCR turns sample sets of mrnas into doublestranded DNAs with the corresponding sequences
80 RT-PCR relies on the activity of reverse transcriptase, which can synthesize a DNA copy of an mrna, called a complementary DNA (cdna) Once the cdna is produced, PCR is used to make many copies of the sequence of interest, using primers specific to that sequence
81 Figure Test tube containing reverse transcriptase and mrna DNA in nucleus mrnas in cytoplasm
82 Figure DNA in nucleus 1 Test tube containing reverse transcriptase and mrna 2 Reverse transcriptase makes the first DNA strand. mrnas in cytoplasm mrna 5 Reverse transcriptase 3 Poly-A tail A A A A A A 3 T T T T T 5 DNA Primer strand
83 Figure DNA in nucleus 1 Test tube containing reverse transcriptase and mrna 2 Reverse transcriptase makes the first DNA strand. mrnas in cytoplasm mrna 5 3 Poly-A tail A A A A A A 3 T T T T T 5 DNA Primer strand 3 mrma is degraded. 5 3 Reverse transcriptase A A A A A A 3 T T T T T 5
84 Figure DNA in nucleus 1 Test tube containing reverse transcriptase and mrna 2 Reverse transcriptase makes the first DNA strand. mrnas in cytoplasm mrna 5 Reverse transcriptase A A A A A A 3 T T T T T 5 3 DNA Primer strand 3 mrma is degraded DNA polymerase synthesizes the second strand. 5 3 DNA polym erase Poly-A tail A A A A A A 3 T T T T T 5 3 5
85 Figure DNA in nucleus 1 Test tube containing reverse transcriptase and mrna 2 Reverse transcriptase makes the first DNA strand. mrnas in cytoplasm mrna 5 Reverse transcriptase A A A A A A 3 T T T T T 5 3 DNA Primer strand 3 mrma is degraded DNA polymerase synthesizes the second strand. 5 cdna carries complete coding sequence 2014 Pearsonwithout Education, Inc. introns. Poly-A tail 5 3 A A A A A A 3 T T T T T DNA polym erase 5 3 cdna 3 5
86 Figure Techniq ue 1 cdna synthesis 2 PCR amplification 3 Gel electrophoresis Results mrna s cdna s Prime rs βglobin gene Embryonic 1 stages
87 Studying the Expression of Groups of Genes A major goal of biologists is to learn how genes act together to produce and maintain a functioning organism Large groups of genes are studied by a systems approach Such approaches allow networks of expression across a genome to be identified
88 Genome-wide expression studies can be carried out using DNA microarray assays A microarray also called a DNA chip contains tiny amounts of many single-stranded DNA fragments affixed to the slide in a grid mrnas from cells of interest are isolated and made into cdnas labeled with fluorescent molecules
89 cdnas from two different samples are labeled with different fluorescent tags and tested on the same microarray The experiment can identify subsets of genes that are being expressed differently in one sample compared to another
90 Figure Genes in red wells expressed in first tissue. Genes in green wells expressed in second tissue. Genes in yellow wells expressed in both tissues. DNA microarray Genes in black wells not expressed in either tissue.
91 An alternative to microarray analysis is simply to sequence cdna samples from different tissues or stages to discover which genes are expressed This is called RNA sequencing This method is becoming more widespread as the cost of sequencing decreases
92 Studies of genes that are expressed together in some tissues but not others may contribute to a better understanding of diseases and suggest new diagnostic tests or therapies
93 Figure 15.UN04 Enhancer with possible control elements Reporter Promoter gene level5of 0 Relative reporter mrna (% of control)
94 Figure 15.UN05 Genes expressed Promot er Genes Operato r Inactive repressor: no corepressor present Genes not expressed Active repressor: corepressor bound Corepressor
95 Figure 15.UN06 Chromatin modification Genes in highly compacted chromatin are generally not transcribed. Histone acetylation seems to loosen chromatin structure, enhancing transcription. DNA methylation generally reduces transcription. Chromatin modification Transcription RNA processing Transcription Regulation of transcription initiation: DNA control elements in enhancers bind specific transcription factors. Bending of the DNA enables activators to contact proteins at the promoter, initiating transcription. The genes in a coordinately controlled group all share a combination of control elements. RNA processing Alternative RNA splicing: Primary RNA transcript mrna mrna degradation Translation Translation Protein processing and degradation mrna degradatio Each mrna has a n characteristic life span. or Initiation of translation can be controlled via regulation of initiation factors. Protein processing and degradation Protein processing and degradation are subject to regulation.
96 Figure 15.UN06a Chromatin modificati on Trans criptio n RNA processin g mrn A degra dation Transl ation Protei n proce ssing and degra
97 Figure 15.UN06b Chromatin modification Genes in highly compacted chromatin are not generally Histone acetylation transcribed. seems to loosen chromatin structure, enhancing transcription. DNA methylation generally reduces transcription. mrna degrad Each mrna has ation a characteristic life span. RNA processing Alternative RNA splicing: Primary RNA transcript mrn A o r Transla tion Initiation of translation can be controlled via regulation of initiation factors. Protein processing and degradation Protein processing and degradation are subject to regulation.
98 Figure 15.UN06c Transcription Regulation of transcription initiation: DNA control elements in enhancers Bending of bind specific the DNAtranscription factors. enables activators to contact proteins at the promoter, initiating transcription. The genes in a coordinately controlled group all share a combination of control
99 Figure 15.UN07 Enhancer Promoter Gene 1 Gene 2 Gene 3 Gene 4 Gene 5
Regulation of Gene Expression
 CAMPBELL BIOLOGY IN FOCUS URRY CAIN WASSERMAN MINORSKY REECE 15 Regulation of Gene Expression Lecture Presentations by Kathleen Fitzpatrick and Nicole Tunbridge, Simon Fraser University SECOND EDITION
CAMPBELL BIOLOGY IN FOCUS URRY CAIN WASSERMAN MINORSKY REECE 15 Regulation of Gene Expression Lecture Presentations by Kathleen Fitzpatrick and Nicole Tunbridge, Simon Fraser University SECOND EDITION
Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression Edited by Shawn Lester PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley
Chapter 18 Regulation of Gene Expression Edited by Shawn Lester PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley
BIOLOGY. Chapter 16 GenesExpression
 BIOLOGY Chapter 16 GenesExpression CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson 18 Gene Expression 2014 Pearson Education, Inc. Figure 16.1 Differential Gene Expression results
BIOLOGY Chapter 16 GenesExpression CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson 18 Gene Expression 2014 Pearson Education, Inc. Figure 16.1 Differential Gene Expression results
Chapter 18: Regulation of Gene Expression. 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer
 Chapter 18: Regulation of Gene Expression 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer Gene Regulation Gene regulation refers to all aspects of controlling
Chapter 18: Regulation of Gene Expression 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer Gene Regulation Gene regulation refers to all aspects of controlling
Regulation of Gene Expression
 Slide 1 Chapter 18 Regulation of Gene Expression PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions
Slide 1 Chapter 18 Regulation of Gene Expression PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions
Control of Eukaryotic Gene Expression (Learning Objectives)
 Control of Eukaryotic Gene Expression (Learning Objectives) 1. Compare and contrast chromatin and chromosome: composition, proteins involved and level of packing. Explain the structure and function of
Control of Eukaryotic Gene Expression (Learning Objectives) 1. Compare and contrast chromatin and chromosome: composition, proteins involved and level of packing. Explain the structure and function of
Chapter 18: Regulation of Gene Expression. Gene Regulation. Transcription Factors 3/21/2017
 Chapter 18: Regulation of Gene Expression 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer Gene Regulation Gene regulation refers to all aspects of controlling
Chapter 18: Regulation of Gene Expression 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer Gene Regulation Gene regulation refers to all aspects of controlling
Section C: The Control of Gene Expression
 Section C: The Control of Gene Expression 1. Each cell of a multicellular eukaryote expresses only a small fraction of its genes 2. The control of gene expression can occur at any step in the pathway from
Section C: The Control of Gene Expression 1. Each cell of a multicellular eukaryote expresses only a small fraction of its genes 2. The control of gene expression can occur at any step in the pathway from
32 Gene regulation in Eukaryotes Lecture Outline 11/28/05. Gene Regulation in Prokaryotes and Eukarykotes
 3 Gene regulation in Eukaryotes Lecture Outline /8/05 Gene regulation in eukaryotes Chromatin remodeling More kinds of control elements Promoters, Enhancers, and Silencers Combinatorial control Cell-specific
3 Gene regulation in Eukaryotes Lecture Outline /8/05 Gene regulation in eukaryotes Chromatin remodeling More kinds of control elements Promoters, Enhancers, and Silencers Combinatorial control Cell-specific
CELL BIOLOGY - CLUTCH CH. 7 - GENE EXPRESSION.
 !! www.clutchprep.com CONCEPT: CONTROL OF GENE EXPRESSION BASICS Gene expression is the process through which cells selectively to express some genes and not others Every cell in an organism is a clone
!! www.clutchprep.com CONCEPT: CONTROL OF GENE EXPRESSION BASICS Gene expression is the process through which cells selectively to express some genes and not others Every cell in an organism is a clone
CHAPTER 13 LECTURE SLIDES
 CHAPTER 13 LECTURE SLIDES Prepared by Brenda Leady University of Toledo To run the animations you must be in Slideshow View. Use the buttons on the animation to play, pause, and turn audio/text on or off.
CHAPTER 13 LECTURE SLIDES Prepared by Brenda Leady University of Toledo To run the animations you must be in Slideshow View. Use the buttons on the animation to play, pause, and turn audio/text on or off.
GENE REGULATION slide shows by Kim Foglia modified Slides with blue edges are Kim s
 GENE REGULATION slide shows by Kim Foglia modified Slides with blue edges are Kim s 2007-2008 Bacterial metabolism Bacteria need to respond quickly to changes in their environment STOP GO if they have
GENE REGULATION slide shows by Kim Foglia modified Slides with blue edges are Kim s 2007-2008 Bacterial metabolism Bacteria need to respond quickly to changes in their environment STOP GO if they have
Control of Gene Expression
 Control of Gene Expression 1 How Gene Regulation Works 2 Control of Gene Expression Controlling gene expression is often accomplished by controlling transcription initiation Regulatory proteins bind to
Control of Gene Expression 1 How Gene Regulation Works 2 Control of Gene Expression Controlling gene expression is often accomplished by controlling transcription initiation Regulatory proteins bind to
Control of Eukaryotic Genes. AP Biology
 Control of Eukaryotic Genes The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions? Evolution
Control of Eukaryotic Genes The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions? Evolution
Chapter 18. Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression 2007-2008 Control of Prokaryotic (Bacterial) Genes 2007- Bacterial metabolism Bacteria need to respond quickly to changes in their environment STOP GO if they have
Chapter 18 Regulation of Gene Expression 2007-2008 Control of Prokaryotic (Bacterial) Genes 2007- Bacterial metabolism Bacteria need to respond quickly to changes in their environment STOP GO if they have
Gene Expression. Lesson 6
 Gene Expression Lesson 6 Regulation of gene expression Gene regulation turning on or off specific genes depending on the requirements of an organism Housekeeping genes are always switched on (vital life
Gene Expression Lesson 6 Regulation of gene expression Gene regulation turning on or off specific genes depending on the requirements of an organism Housekeeping genes are always switched on (vital life
Differences between prokaryotes & eukaryotes. Gene function
 GENE REGULATION Differences between prokaryotes & eukaryotes Gene function Description of Prokaryotic Chromosome and E.coli Review Differences between Prokaryotic & Eukaryotic Chromosomes Four differences
GENE REGULATION Differences between prokaryotes & eukaryotes Gene function Description of Prokaryotic Chromosome and E.coli Review Differences between Prokaryotic & Eukaryotic Chromosomes Four differences
Chapter 11: Regulation of Gene Expression
 Chapter Review 1. It has long been known that there is probably a genetic link for alcoholism. Researchers studying rats have begun to elucidate this link. Briefly describe the genetic mechanism found
Chapter Review 1. It has long been known that there is probably a genetic link for alcoholism. Researchers studying rats have begun to elucidate this link. Briefly describe the genetic mechanism found
Control of Eukaryotic Genes
 Control of Eukaryotic Genes 2007-2008 The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions?
Control of Eukaryotic Genes 2007-2008 The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions?
AP Biology Gene Expression/Biotechnology REVIEW
 AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
Unit IX Problem 3 Genetics: Basic Concepts in Molecular Biology
 Unit IX Problem 3 Genetics: Basic Concepts in Molecular Biology - The central dogma (principle) of molecular biology: Information from DNA are transcribed to mrna which will be further translated to synthesize
Unit IX Problem 3 Genetics: Basic Concepts in Molecular Biology - The central dogma (principle) of molecular biology: Information from DNA are transcribed to mrna which will be further translated to synthesize
Control of Eukaryotic Genes. AP Biology
 Control of Eukaryotic Genes The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions? Evolution
Control of Eukaryotic Genes The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions? Evolution
Gene Expression and Regulation - 1
 Gene Expression and Regulation - 1 We have been discussing the molecular structure of DNA and its function in DNA replication and in transcription. Earlier we discussed how genes interact in transmission
Gene Expression and Regulation - 1 We have been discussing the molecular structure of DNA and its function in DNA replication and in transcription. Earlier we discussed how genes interact in transmission
Control of Eukaryotic Genes. AP Biology
 Control of Eukaryotic Genes The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions? Evolution
Control of Eukaryotic Genes The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions? Evolution
Unit 7. Genetic Regulation, Development, and Biotechnology. AP Biology
 Unit 7 Genetic Regulation, Development, and Biotechnology The BIG Questions How are genes turned on & off in eukaryotes and prokaryotes? How do cells with the same genes differentiate to perform completely
Unit 7 Genetic Regulation, Development, and Biotechnology The BIG Questions How are genes turned on & off in eukaryotes and prokaryotes? How do cells with the same genes differentiate to perform completely
EUKARYOTIC GENE CONTROL
 EUKARYOTIC GENE CONTROL THE BIG QUESTIONS How are genes turned on and off? How do cells with the same DNA/ genes differentiate to perform completely different and specialized functions? GENE EXPRESSION
EUKARYOTIC GENE CONTROL THE BIG QUESTIONS How are genes turned on and off? How do cells with the same DNA/ genes differentiate to perform completely different and specialized functions? GENE EXPRESSION
Control of Eukaryotic Genes. AP Biology
 Control of Eukaryotic Genes The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions? Evolution
Control of Eukaryotic Genes The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions? Evolution
GENE REGULATION IN PROKARYOTES
 GENE REGULATION IN PROKARYOTES Prepared by Brenda Leady, University of Toledo Copyright (c) The McGraw-Hill Companies, Inc. Permission required for reproduction or display. 1 Gene regulation refers to
GENE REGULATION IN PROKARYOTES Prepared by Brenda Leady, University of Toledo Copyright (c) The McGraw-Hill Companies, Inc. Permission required for reproduction or display. 1 Gene regulation refers to
Prokaryotic Transcription
 Prokaryotic Transcription Transcription Basics DNA is the genetic material Nucleic acid Capable of self-replication and synthesis of RNA RNA is the middle man Nucleic acid Structure and base sequence are
Prokaryotic Transcription Transcription Basics DNA is the genetic material Nucleic acid Capable of self-replication and synthesis of RNA RNA is the middle man Nucleic acid Structure and base sequence are
Chapter 2. An Introduction to Genes and Genomes
 PowerPoint Lectures for Introduction to Biotechnology, Second Edition William J.Thieman and Michael A.Palladino Chapter 2 An Introduction to Genes and Genomes Lectures by Lara Dowland Chapter Contents
PowerPoint Lectures for Introduction to Biotechnology, Second Edition William J.Thieman and Michael A.Palladino Chapter 2 An Introduction to Genes and Genomes Lectures by Lara Dowland Chapter Contents
Chapter 13 - Regulation of Gene Expression
 Chapter 13 - Regulation of Gene Expression 1. Describe the typical components of an operon in an E. coli (prokaryotic) cell. (p. 238-239) a. regulator gene - b. promoter - c. operator - d. structural gene
Chapter 13 - Regulation of Gene Expression 1. Describe the typical components of an operon in an E. coli (prokaryotic) cell. (p. 238-239) a. regulator gene - b. promoter - c. operator - d. structural gene
There are four major types of introns. Group I introns, found in some rrna genes, are self-splicing: they can catalyze their own removal.
 1 2 Continuous genes - Intron: Many eukaryotic genes contain coding regions called exons and noncoding regions called intervening sequences or introns. The average human gene contains from eight to nine
1 2 Continuous genes - Intron: Many eukaryotic genes contain coding regions called exons and noncoding regions called intervening sequences or introns. The average human gene contains from eight to nine
AP Biology. The BIG Questions. Chapter 19. Prokaryote vs. eukaryote genome. Prokaryote vs. eukaryote genome. Why turn genes on & off?
 The BIG Questions Chapter 19. Control of Eukaryotic Genome How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions?
The BIG Questions Chapter 19. Control of Eukaryotic Genome How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions?
Division Ave. High School AP Biology
 Control of Eukaryotic Genes 2007-2008 The BIG Questions n How are genes turned on & off in eukaryotes? n How do cells with the same genes differentiate to perform completely different, specialized functions?
Control of Eukaryotic Genes 2007-2008 The BIG Questions n How are genes turned on & off in eukaryotes? n How do cells with the same genes differentiate to perform completely different, specialized functions?
Themes: RNA and RNA Processing. Messenger RNA (mrna) What is a gene? RNA is very versatile! RNA-RNA interactions are very important!
 Themes: RNA is very versatile! RNA and RNA Processing Chapter 14 RNA-RNA interactions are very important! Prokaryotes and Eukaryotes have many important differences. Messenger RNA (mrna) Carries genetic
Themes: RNA is very versatile! RNA and RNA Processing Chapter 14 RNA-RNA interactions are very important! Prokaryotes and Eukaryotes have many important differences. Messenger RNA (mrna) Carries genetic
Chapter 19. Control of Eukaryotic Genome. AP Biology
 Chapter 19. Control of Eukaryotic Genome The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions?
Chapter 19. Control of Eukaryotic Genome The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions?
DNA makes RNA makes Proteins. The Central Dogma
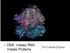 DNA makes RNA makes Proteins The Central Dogma TRANSCRIPTION DNA RNA transcript RNA polymerase RNA PROCESSING Exon RNA transcript (pre-mrna) Intron Aminoacyl-tRNA synthetase NUCLEUS CYTOPLASM FORMATION
DNA makes RNA makes Proteins The Central Dogma TRANSCRIPTION DNA RNA transcript RNA polymerase RNA PROCESSING Exon RNA transcript (pre-mrna) Intron Aminoacyl-tRNA synthetase NUCLEUS CYTOPLASM FORMATION
Unit 6: Molecular Genetics & DNA Technology Guided Reading Questions (100 pts total)
 Name: AP Biology Biology, Campbell and Reece, 7th Edition Adapted from chapter reading guides originally created by Lynn Miriello Chapter 16 The Molecular Basis of Inheritance Unit 6: Molecular Genetics
Name: AP Biology Biology, Campbell and Reece, 7th Edition Adapted from chapter reading guides originally created by Lynn Miriello Chapter 16 The Molecular Basis of Inheritance Unit 6: Molecular Genetics
Chapter 18: Regulation of Gene Expression
 Chapter 18: Regulation of Gene Expression Regulation of Metabolism Shuts off transcription Types of Feedback Negative feedback = body s response is to reduce the stimulus Ex: regulation of body temp, blood
Chapter 18: Regulation of Gene Expression Regulation of Metabolism Shuts off transcription Types of Feedback Negative feedback = body s response is to reduce the stimulus Ex: regulation of body temp, blood
Transcription and Post Transcript Modification
 Transcription and Post Transcript Modification You Should Be Able To 1. Describe transcription. 2. Compare and contrast eukaryotic + prokaryotic transcription. 3. Explain mrna processing in eukaryotes.
Transcription and Post Transcript Modification You Should Be Able To 1. Describe transcription. 2. Compare and contrast eukaryotic + prokaryotic transcription. 3. Explain mrna processing in eukaryotes.
B. Incorrect! Centromeric DNA is largely heterochromatin, which is inactive DNA.
 MCAT Biology - Problem Drill 06: Molecular Biology of Eukaryotes Question No. 1 of 10 1. Which type of DNA would have the highest level of expression? Question #01 (A) Heterochromatin. (B) Centromeric
MCAT Biology - Problem Drill 06: Molecular Biology of Eukaryotes Question No. 1 of 10 1. Which type of DNA would have the highest level of expression? Question #01 (A) Heterochromatin. (B) Centromeric
Gene Expression: Transcription, Translation, RNAs and the Genetic Code
 Lecture 28-29 Gene Expression: Transcription, Translation, RNAs and the Genetic Code Central dogma of molecular biology During transcription, the information in a DNA sequence (a gene) is copied into a
Lecture 28-29 Gene Expression: Transcription, Translation, RNAs and the Genetic Code Central dogma of molecular biology During transcription, the information in a DNA sequence (a gene) is copied into a
Transcription in Eukaryotes
 Transcription in Eukaryotes Biology I Hayder A Giha Transcription Transcription is a DNA-directed synthesis of RNA, which is the first step in gene expression. Gene expression, is transformation of the
Transcription in Eukaryotes Biology I Hayder A Giha Transcription Transcription is a DNA-directed synthesis of RNA, which is the first step in gene expression. Gene expression, is transformation of the
Find this material useful? You can help our team to keep this site up and bring you even more content consider donating via the link on our site.
 Find this material useful? You can help our team to keep this site up and bring you even more content consider donating via the link on our site. Still having trouble understanding the material? Check
Find this material useful? You can help our team to keep this site up and bring you even more content consider donating via the link on our site. Still having trouble understanding the material? Check
Control of Metabolic Processes
 Control of Metabolic Processes Harriet Wilson, Lecture Notes Bio. Sci. 4 - Microbiology Sierra College As described earlier, the metabolic processes occurring within living organisms (glycolysis, respiration,
Control of Metabolic Processes Harriet Wilson, Lecture Notes Bio. Sci. 4 - Microbiology Sierra College As described earlier, the metabolic processes occurring within living organisms (glycolysis, respiration,
Biotechnology Unit 3: DNA to Proteins. From DNA to RNA
 From DNA to RNA Biotechnology Unit 3: DNA to Proteins I. After the discovery of the structure of DNA, the major question remaining was how does the stored in the 4 letter code of DNA direct the and of
From DNA to RNA Biotechnology Unit 3: DNA to Proteins I. After the discovery of the structure of DNA, the major question remaining was how does the stored in the 4 letter code of DNA direct the and of
M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION
 M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION Chapter 7 Microbial Genetics Lecture prepared by Mindy Miller-Kittrell, University of Tennessee, Knoxville The Structure and Replication
M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION Chapter 7 Microbial Genetics Lecture prepared by Mindy Miller-Kittrell, University of Tennessee, Knoxville The Structure and Replication
Unit 6 Exam Review Game January 29, 2019 JEOPARDY! Unit 6 Exam Review Game January 29 th, 2019
 JOPARY! Unit 6 xam Review ame January 29 th, 2019 ow to play JOPARY! The class will be split into two teams, the R team and the RN team. When a question is displayed, the R team will use the A keys on
JOPARY! Unit 6 xam Review ame January 29 th, 2019 ow to play JOPARY! The class will be split into two teams, the R team and the RN team. When a question is displayed, the R team will use the A keys on
Computational Biology I LSM5191 (2003/4)
 Computational Biology I LSM5191 (2003/4) Aylwin Ng, D.Phil Lecture Notes: Transcriptome: Molecular Biology of Gene Expression I Flow of information: DNA to polypeptide DNA Start Exon1 Intron Exon2 Termination
Computational Biology I LSM5191 (2003/4) Aylwin Ng, D.Phil Lecture Notes: Transcriptome: Molecular Biology of Gene Expression I Flow of information: DNA to polypeptide DNA Start Exon1 Intron Exon2 Termination
Unit 8: Genomics Guided Reading Questions (150 pts total)
 Name: AP Biology Biology, Campbell and Reece, 7th Edition Adapted from chapter reading guides originally created by Lynn Miriello Chapter 18 The Genetics of Viruses and Bacteria Unit 8: Genomics Guided
Name: AP Biology Biology, Campbell and Reece, 7th Edition Adapted from chapter reading guides originally created by Lynn Miriello Chapter 18 The Genetics of Viruses and Bacteria Unit 8: Genomics Guided
Unit II Problem 3 Genetics: Summary of Basic Concepts in Molecular Biology
 Unit II Problem 3 Genetics: Summary of Basic Concepts in Molecular Biology - The central dogma (principle) of molecular biology: Information from DNA are transcribed to mrna which will be further translated
Unit II Problem 3 Genetics: Summary of Basic Concepts in Molecular Biology - The central dogma (principle) of molecular biology: Information from DNA are transcribed to mrna which will be further translated
30 Gene expression: Transcription
 30 Gene expression: Transcription Gene structure. o Exons coding region of DNA. o Introns non-coding region of DNA. o Introns are interspersed between exons of a single gene. o Promoter region helps enzymes
30 Gene expression: Transcription Gene structure. o Exons coding region of DNA. o Introns non-coding region of DNA. o Introns are interspersed between exons of a single gene. o Promoter region helps enzymes
Transcription is the first stage of gene expression
 Transcription is the first stage of gene expression RNA synthesis is catalyzed by RNA polymerase, which pries the DNA strands apart and hooks together the RNA nucleotides The RNA is complementary to the
Transcription is the first stage of gene expression RNA synthesis is catalyzed by RNA polymerase, which pries the DNA strands apart and hooks together the RNA nucleotides The RNA is complementary to the
GENE REGULATION. Gene regulation occurs at the level of transcription or production of mrna
 GENE REGULATION Virtually every cell in your body contains a complete set of genes But they are not all turned on in every tissue Each cell in your body expresses only a small subset of genes at any time
GENE REGULATION Virtually every cell in your body contains a complete set of genes But they are not all turned on in every tissue Each cell in your body expresses only a small subset of genes at any time
Gene Expression: Transcription
 Gene Expression: Transcription The majority of genes are expressed as the proteins they encode. The process occurs in two steps: Transcription = DNA RNA Translation = RNA protein Taken together, they make
Gene Expression: Transcription The majority of genes are expressed as the proteins they encode. The process occurs in two steps: Transcription = DNA RNA Translation = RNA protein Taken together, they make
Big Idea 3C Basic Review
 Big Idea 3C Basic Review 1. A gene is a. A sequence of DNA that codes for a protein. b. A sequence of amino acids that codes for a protein. c. A sequence of codons that code for nucleic acids. d. The end
Big Idea 3C Basic Review 1. A gene is a. A sequence of DNA that codes for a protein. b. A sequence of amino acids that codes for a protein. c. A sequence of codons that code for nucleic acids. d. The end
A. Incorrect! This feature does help with it suitability as genetic material.
 College Biology - Problem Drill 08: Gene Structures and Functions No. 1 of 10 1. Which of the statements below is NOT true in explaining why DNA is a suitable genetic material? #01 (A) Its double helix
College Biology - Problem Drill 08: Gene Structures and Functions No. 1 of 10 1. Which of the statements below is NOT true in explaining why DNA is a suitable genetic material? #01 (A) Its double helix
Chapter 17. From Gene to Protein. Slide 1. Slide 2. Slide 3. Gene Expression. Which of the following is the best example of gene expression? Why?
 Slide 1 Chapter 17 From Gene to Protein PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
Slide 1 Chapter 17 From Gene to Protein PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
Genetics Biology 331 Exam 3B Spring 2015
 Genetics Biology 331 Exam 3B Spring 2015 MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1) DNA methylation may be a significant mode of genetic regulation
Genetics Biology 331 Exam 3B Spring 2015 MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1) DNA methylation may be a significant mode of genetic regulation
Matakuliah Genetika (BIO612206) Jurusan Biologi FMIPA Universitas Lampung. Priyambodo, M.Sc. staff.unila.ac.id/priyambodo
 Matakuliah Genetika (BIO612206) Jurusan Biologi FMIPA Universitas Lampung Priyambodo, M.Sc. staff.unila.ac.id/priyambodo Prokariotik Eukariotik staff.unila.ac.id/priyambodo Regulasi ekspresi gen pada
Matakuliah Genetika (BIO612206) Jurusan Biologi FMIPA Universitas Lampung Priyambodo, M.Sc. staff.unila.ac.id/priyambodo Prokariotik Eukariotik staff.unila.ac.id/priyambodo Regulasi ekspresi gen pada
Gene Regulation Biology
 Gene Regulation Biology Potential and Limitations of Cell Re-programming in Cancer Research Eric Blanc KCL April 13, 2010 Eric Blanc (KCL) Gene Regulation Biology April 13, 2010 1 / 21 Outline 1 The Central
Gene Regulation Biology Potential and Limitations of Cell Re-programming in Cancer Research Eric Blanc KCL April 13, 2010 Eric Blanc (KCL) Gene Regulation Biology April 13, 2010 1 / 21 Outline 1 The Central
Fig Ch 17: From Gene to Protein
 Fig. 17-1 Ch 17: From Gene to Protein Basic Principles of Transcription and Translation RNA is the intermediate between genes and the proteins for which they code Transcription is the synthesis of RNA
Fig. 17-1 Ch 17: From Gene to Protein Basic Principles of Transcription and Translation RNA is the intermediate between genes and the proteins for which they code Transcription is the synthesis of RNA
From DNA to Protein: Genotype to Phenotype
 12 From DNA to Protein: Genotype to Phenotype 12.1 What Is the Evidence that Genes Code for Proteins? The gene-enzyme relationship is one-gene, one-polypeptide relationship. Example: In hemoglobin, each
12 From DNA to Protein: Genotype to Phenotype 12.1 What Is the Evidence that Genes Code for Proteins? The gene-enzyme relationship is one-gene, one-polypeptide relationship. Example: In hemoglobin, each
How to Use This Presentation
 How to Use This Presentation To View the presentation as a slideshow with effects select View on the menu bar and click on Slide Show. To advance through the presentation, click the right-arrow key or
How to Use This Presentation To View the presentation as a slideshow with effects select View on the menu bar and click on Slide Show. To advance through the presentation, click the right-arrow key or
7.1 The lac Operon 7-1
 7.1 The lac Operon The lac operon was the first operon discovered It contains 3 genes coding for E. coli proteins that permit the bacteria to use the sugar lactose Galactoside permease (lacy) which transports
7.1 The lac Operon The lac operon was the first operon discovered It contains 3 genes coding for E. coli proteins that permit the bacteria to use the sugar lactose Galactoside permease (lacy) which transports
I. Prokaryotic Gene Regulation. Figure 1: Operon. Operon:
 I. Prokaryotic Gene Regulation Figure 1: Operon Operon: a) Regulatory Elements consist of an Operator that serves as the on-off switch for the genes of the operon. Also contains a promoter for the Structural
I. Prokaryotic Gene Regulation Figure 1: Operon Operon: a) Regulatory Elements consist of an Operator that serves as the on-off switch for the genes of the operon. Also contains a promoter for the Structural
Nucleic acids deoxyribonucleic acid (DNA) ribonucleic acid (RNA) nucleotide
 Nucleic Acids Nucleic acids are molecules that store information for cellular growth and reproduction There are two types of nucleic acids: - deoxyribonucleic acid (DNA) and ribonucleic acid (RNA) These
Nucleic Acids Nucleic acids are molecules that store information for cellular growth and reproduction There are two types of nucleic acids: - deoxyribonucleic acid (DNA) and ribonucleic acid (RNA) These
Gene Circuits -2. Shu-Ping Lin, Ph.D. Institute of Biomedical Engineering
 Gene Circuits -2 Shu-Ping Lin, Ph.D. Institute of Biomedical Engineering E-mail: splin@dragon.nchu.edu.tw Website: http://web.nchu.edu.tw/pweb/users/splin/ Genes Flow of information in gene expression
Gene Circuits -2 Shu-Ping Lin, Ph.D. Institute of Biomedical Engineering E-mail: splin@dragon.nchu.edu.tw Website: http://web.nchu.edu.tw/pweb/users/splin/ Genes Flow of information in gene expression
Gene Regulation in Eukaryotes
 Gene Regulation in Eukaryotes The latest estimates are that a human cell, a eukaryotic cell, contains 20,000 25,000 genes. Some of these are expressed in all cells all the time. These so-called housekeeping
Gene Regulation in Eukaryotes The latest estimates are that a human cell, a eukaryotic cell, contains 20,000 25,000 genes. Some of these are expressed in all cells all the time. These so-called housekeeping
Transcription. The sugar molecule found in RNA is ribose, rather than the deoxyribose found in DNA.
 Transcription RNA (ribonucleic acid) is a key intermediary between a DNA sequence and a polypeptide. RNA is an informational polynucleotide similar to DNA, but it differs from DNA in three ways: RNA generally
Transcription RNA (ribonucleic acid) is a key intermediary between a DNA sequence and a polypeptide. RNA is an informational polynucleotide similar to DNA, but it differs from DNA in three ways: RNA generally
DNA REPLICATION. DNA structure. Semiconservative replication. DNA structure. Origin of replication. Replication bubbles and forks.
 DNA REPLICATION 5 4 Phosphate 3 DNA structure Nitrogenous base 1 Deoxyribose 2 Nucleotide DNA strand = DNA polynucleotide 2004 Biology Olympiad Preparation Program 2 2004 Biology Olympiad Preparation Program
DNA REPLICATION 5 4 Phosphate 3 DNA structure Nitrogenous base 1 Deoxyribose 2 Nucleotide DNA strand = DNA polynucleotide 2004 Biology Olympiad Preparation Program 2 2004 Biology Olympiad Preparation Program
(c) 2014 Dr. Alice Heicklen & Dr. Deborah Mowshowitz, Columbia University, New York, NY. Last update 02/26/ :57 PM
 C2006/F2402 '14 OUTLINE OF LECTURE #11 (c) 2014 Dr. Alice Heicklen & Dr. Deborah Mowshowitz, Columbia University, New York, NY. Last update 02/26/2014 12:57 PM Handouts: 10C -- Typical Eukaryotic Gene,
C2006/F2402 '14 OUTLINE OF LECTURE #11 (c) 2014 Dr. Alice Heicklen & Dr. Deborah Mowshowitz, Columbia University, New York, NY. Last update 02/26/2014 12:57 PM Handouts: 10C -- Typical Eukaryotic Gene,
CH 17 :From Gene to Protein
 CH 17 :From Gene to Protein Defining a gene gene gene Defining a gene is problematic because one gene can code for several protein products, some genes code only for RNA, two genes can overlap, and there
CH 17 :From Gene to Protein Defining a gene gene gene Defining a gene is problematic because one gene can code for several protein products, some genes code only for RNA, two genes can overlap, and there
Chapter 19 Genetic Regulation of the Eukaryotic Genome. A. Bergeron AP Biology PCHS
 Chapter 19 Genetic Regulation of the Eukaryotic Genome A. Bergeron AP Biology PCHS 2 Do Now - Eukaryotic Transcription Regulation The diagram below shows five genes (with their enhancers) from the genome
Chapter 19 Genetic Regulation of the Eukaryotic Genome A. Bergeron AP Biology PCHS 2 Do Now - Eukaryotic Transcription Regulation The diagram below shows five genes (with their enhancers) from the genome
NUCLEIC ACID METABOLISM. Omidiwura, B.R.O
 NUCLEIC ACID METABOLISM Omidiwura, B.R.O Nucleic Acids Nucleic acids are molecules that store information for cellular growth and reproduction There are two types of nucleic acids: - deoxyribonucleic acid
NUCLEIC ACID METABOLISM Omidiwura, B.R.O Nucleic Acids Nucleic acids are molecules that store information for cellular growth and reproduction There are two types of nucleic acids: - deoxyribonucleic acid
Regulation of bacterial gene expression
 Regulation of bacterial gene expression Gene Expression Gene Expression: RNA and protein synthesis DNA ----------> RNA ----------> Protein transcription translation! DNA replication only occurs in cells
Regulation of bacterial gene expression Gene Expression Gene Expression: RNA and protein synthesis DNA ----------> RNA ----------> Protein transcription translation! DNA replication only occurs in cells
Developmental Biology BY1101 P. Murphy
 Developmental Biology BY1101 P. Murphy Lecture 7 Cellular differentiation and the regulation of gene expression. In this lecture we looked at two main questions: How is gene expression regulated? (revision
Developmental Biology BY1101 P. Murphy Lecture 7 Cellular differentiation and the regulation of gene expression. In this lecture we looked at two main questions: How is gene expression regulated? (revision
Enzyme that uses RNA as a template to synthesize a complementary DNA
 Biology 105: Introduction to Genetics PRACTICE FINAL EXAM 2006 Part I: Definitions Homology: Comparison of two or more protein or DNA sequence to ascertain similarities in sequences. If two genes have
Biology 105: Introduction to Genetics PRACTICE FINAL EXAM 2006 Part I: Definitions Homology: Comparison of two or more protein or DNA sequence to ascertain similarities in sequences. If two genes have
Biology. Biology. Slide 1 of 39. End Show. Copyright Pearson Prentice Hall
 Biology Biology 1 of 39 12-3 RNA and Protein Synthesis 2 of 39 Essential Question What is transcription and translation and how do they take place? 3 of 39 12 3 RNA and Protein Synthesis Genes are coded
Biology Biology 1 of 39 12-3 RNA and Protein Synthesis 2 of 39 Essential Question What is transcription and translation and how do they take place? 3 of 39 12 3 RNA and Protein Synthesis Genes are coded
Biology. Biology. Slide 1 of 39. End Show. Copyright Pearson Prentice Hall
 Biology Biology 1 of 39 12-3 RNA and Protein Synthesis 2 of 39 12 3 RNA and Protein Synthesis Genes are coded DNA instructions that control the production of proteins. Genetic messages can be decoded by
Biology Biology 1 of 39 12-3 RNA and Protein Synthesis 2 of 39 12 3 RNA and Protein Synthesis Genes are coded DNA instructions that control the production of proteins. Genetic messages can be decoded by
Lecture 9 Controlling gene expression
 Lecture 9 Controlling gene expression BIOLOGY Campbell, Reece and Mitchell Chapter 18 334- (352-356) Every cell in your body contains the same number of genes approximately 35, 000 DNA is wound around
Lecture 9 Controlling gene expression BIOLOGY Campbell, Reece and Mitchell Chapter 18 334- (352-356) Every cell in your body contains the same number of genes approximately 35, 000 DNA is wound around
Chapter 14 Regulation of Transcription
 Chapter 14 Regulation of Transcription Cis-acting sequences Distance-independent cis-acting elements Dissecting regulatory elements Transcription factors Overview transcriptional regulation Transcription
Chapter 14 Regulation of Transcription Cis-acting sequences Distance-independent cis-acting elements Dissecting regulatory elements Transcription factors Overview transcriptional regulation Transcription
CHAPTERS , 17: Eukaryotic Genetics
 CHAPTERS 14.1 14.6, 17: Eukaryotic Genetics 1. Review the levels of DNA packing within the eukaryote nucleus. Label each level. (A similar diagram is on pg 188 of your textbook.) 2. How do the coding regions
CHAPTERS 14.1 14.6, 17: Eukaryotic Genetics 1. Review the levels of DNA packing within the eukaryote nucleus. Label each level. (A similar diagram is on pg 188 of your textbook.) 2. How do the coding regions
Chapter 12 Packet DNA 1. What did Griffith conclude from his experiment? 2. Describe the process of transformation.
 Chapter 12 Packet DNA and RNA Name Period California State Standards covered by this chapter: Cell Biology 1. The fundamental life processes of plants and animals depend on a variety of chemical reactions
Chapter 12 Packet DNA and RNA Name Period California State Standards covered by this chapter: Cell Biology 1. The fundamental life processes of plants and animals depend on a variety of chemical reactions
Chapter 17 Lecture. Concepts of Genetics. Tenth Edition. Regulation of Gene Expression in Eukaryotes
 Chapter 17 Lecture Concepts of Genetics Tenth Edition Regulation of Gene Expression in Eukaryotes Chapter Contents 17.1 Eukaryotic Gene Regulation Can Occur at Any of the Steps Leading from DNA to Protein
Chapter 17 Lecture Concepts of Genetics Tenth Edition Regulation of Gene Expression in Eukaryotes Chapter Contents 17.1 Eukaryotic Gene Regulation Can Occur at Any of the Steps Leading from DNA to Protein
Wednesday, November 22, 17. Exons and Introns
 Exons and Introns Introns and Exons Exons: coded regions of DNA that get transcribed and translated into proteins make up 5% of the genome Introns and Exons Introns: non-coded regions of DNA Must be removed
Exons and Introns Introns and Exons Exons: coded regions of DNA that get transcribed and translated into proteins make up 5% of the genome Introns and Exons Introns: non-coded regions of DNA Must be removed
GENES AND CHROMOSOMES V. Lecture 7. Biology Department Concordia University. Dr. S. Azam BIOL 266/
 1 GENES AND CHROMOSOMES V Lecture 7 BIOL 266/4 2014-15 Dr. S. Azam Biology Department Concordia University 2 CELL NUCLEUS AND THE CONTROL OF GENE EXPRESSION An Overview of Gene Regulation in Eukaryotes
1 GENES AND CHROMOSOMES V Lecture 7 BIOL 266/4 2014-15 Dr. S. Azam Biology Department Concordia University 2 CELL NUCLEUS AND THE CONTROL OF GENE EXPRESSION An Overview of Gene Regulation in Eukaryotes
BIOLOGY 111. CHAPTER 6: DNA: The Molecule of Life
 BIOLOGY 111 CHAPTER 6: DNA: The Molecule of Life Chromosomes and Inheritance Learning Outcomes 6.1 Describe the structure of the DNA molecule and how this structure allows for the storage of information,
BIOLOGY 111 CHAPTER 6: DNA: The Molecule of Life Chromosomes and Inheritance Learning Outcomes 6.1 Describe the structure of the DNA molecule and how this structure allows for the storage of information,
DNA and Gene Expression (chaps 12-15)
 Name: Class: _ Date: _ DNA and Gene Expression (chaps 12-15) Multiple Choice Identify the choice that best completes the statement or answers the question. 1 Who first confirmed that the replication of
Name: Class: _ Date: _ DNA and Gene Expression (chaps 12-15) Multiple Choice Identify the choice that best completes the statement or answers the question. 1 Who first confirmed that the replication of
Videos. Bozeman Transcription and Translation: Drawing transcription and translation:
 Videos Bozeman Transcription and Translation: https://youtu.be/h3b9arupxzg Drawing transcription and translation: https://youtu.be/6yqplgnjr4q Objectives 29a) I can contrast RNA and DNA. 29b) I can explain
Videos Bozeman Transcription and Translation: https://youtu.be/h3b9arupxzg Drawing transcription and translation: https://youtu.be/6yqplgnjr4q Objectives 29a) I can contrast RNA and DNA. 29b) I can explain
I. Gene Expression Figure 1: Central Dogma of Molecular Biology
 I. Gene Expression Figure 1: Central Dogma of Molecular Biology Central Dogma: Gene Expression: RNA Structure RNA nucleotides contain the pentose sugar Ribose instead of deoxyribose. Contain the bases
I. Gene Expression Figure 1: Central Dogma of Molecular Biology Central Dogma: Gene Expression: RNA Structure RNA nucleotides contain the pentose sugar Ribose instead of deoxyribose. Contain the bases
TRANSCRIPTION AND PROCESSING OF RNA
 TRANSCRIPTION AND PROCESSING OF RNA 1. The steps of gene expression. 2. General characterization of transcription: steps, components of transcription apparatus. 3. Transcription of eukaryotic structural
TRANSCRIPTION AND PROCESSING OF RNA 1. The steps of gene expression. 2. General characterization of transcription: steps, components of transcription apparatus. 3. Transcription of eukaryotic structural
Lesson 8. DNA: The Molecule of Heredity. Gene Expression and Regulation. Introduction to Life Processes - SCI 102 1
 Lesson 8 DNA: The Molecule of Heredity Gene Expression and Regulation Introduction to Life Processes - SCI 102 1 Genes and DNA Hereditary information is found in discrete units called genes Genes are segments
Lesson 8 DNA: The Molecule of Heredity Gene Expression and Regulation Introduction to Life Processes - SCI 102 1 Genes and DNA Hereditary information is found in discrete units called genes Genes are segments
17.5 Eukaryotic Transcription Initiation Is Regulated by Transcription Factors That Bind to Cis-Acting Sites
 17.5 Eukaryotic Transcription Initiation Is Regulated by Transcription Factors That Bind to Cis-Acting Sites 1 Section 17.5 Transcription regulatory proteins, transcription factors, target cis-acting sites
17.5 Eukaryotic Transcription Initiation Is Regulated by Transcription Factors That Bind to Cis-Acting Sites 1 Section 17.5 Transcription regulatory proteins, transcription factors, target cis-acting sites
Unit 6: Molecular Genetics & DNA Technology Guided Reading Questions (100 pts total)
 AP Biology Biology, Campbell and Reece, 10th Edition Adapted from chapter reading guides originally created by Lynn Miriello Name: Chapter 16 The Molecular Basis of Inheritance Concept 16.1 DNA is the
AP Biology Biology, Campbell and Reece, 10th Edition Adapted from chapter reading guides originally created by Lynn Miriello Name: Chapter 16 The Molecular Basis of Inheritance Concept 16.1 DNA is the
Molecular Genetics Student Objectives
 Molecular Genetics Student Objectives Exam 1: Enduring understanding 3.A: Heritable information provides for continuity of life. Essential knowledge 3.A.1: DNA, and in some cases RNA, is the primary source
Molecular Genetics Student Objectives Exam 1: Enduring understanding 3.A: Heritable information provides for continuity of life. Essential knowledge 3.A.1: DNA, and in some cases RNA, is the primary source
Q. No. 1. How can RNA be distinguished from DNA?
 Frequently asked questions (FAQS): Q. No. 1. How can RNA be distinguished from DNA? Ans. RNA and DNA are both nucleic acids, but differ in three main ways. First, unlike DNA which is generally double-stranded,
Frequently asked questions (FAQS): Q. No. 1. How can RNA be distinguished from DNA? Ans. RNA and DNA are both nucleic acids, but differ in three main ways. First, unlike DNA which is generally double-stranded,
Chapter 3. DNA, RNA, and Protein Synthesis
 Chapter 3. DNA, RNA, and Protein Synthesis 4. Transcription Gene Expression Regulatory region (promoter) 5 flanking region Upstream region Coding region 3 flanking region Downstream region Transcription
Chapter 3. DNA, RNA, and Protein Synthesis 4. Transcription Gene Expression Regulatory region (promoter) 5 flanking region Upstream region Coding region 3 flanking region Downstream region Transcription
Name. Student ID. Midterm 2, Biology 2020, Kropf 2004
 Midterm 2, Biology 2020, Kropf 2004 1 1. RNA vs DNA (5 pts) The table below compares DNA and RNA. Fill in the open boxes, being complete and specific Compare: DNA RNA Pyrimidines C,T C,U Purines 3-D structure
Midterm 2, Biology 2020, Kropf 2004 1 1. RNA vs DNA (5 pts) The table below compares DNA and RNA. Fill in the open boxes, being complete and specific Compare: DNA RNA Pyrimidines C,T C,U Purines 3-D structure
Chapter 11. How Genes Are Controlled. Lectures by Edward J. Zalisko
 Chapter 11 How Genes Are Controlled PowerPoint Lectures for Campbell Essential Biology, Fifth Edition, and Campbell Essential Biology with Physiology, Fourth Edition Eric J. Simon, Jean L. Dickey, and
Chapter 11 How Genes Are Controlled PowerPoint Lectures for Campbell Essential Biology, Fifth Edition, and Campbell Essential Biology with Physiology, Fourth Edition Eric J. Simon, Jean L. Dickey, and
