WORKSHOP. Transcriptional circuitry and the regulatory conformation of the genome. Ofir Hakim Faculty of Life Sciences
|
|
|
- Milo Blankenship
- 6 years ago
- Views:
Transcription
1 WORKSHOP Transcriptional circuitry and the regulatory conformation of the genome Ofir Hakim Faculty of Life Sciences Chromosome conformation capture (3C)
2 Most GR Binding Sites Are Distant From Regulated Genes Transcription Pol II ChIP Genes TF1 GR TF2 AP1 Histone mod H3K4Me2 Regulatory elements DHS chr10:79,970,001-80,260,000
3 Distribution of GR Binding Sites 8,373 GR Binding Sites Position of GR relative to TSS (>5kb upstream of TSS) -5kb +5kb TSS (-2.5kb to +2.5 kb) (of last exon) 1.4% John et al., Nature Genet., 2011
4 The genome is not randomly organized Chromosomes territories Nature Reviews Genetics 6, 429 ES cells Macrophages Cell-type specificity Albumin β-actin Chromosome 5 Hepperger, et al. Chromosoma. 2008
5 Functional compartmentalization Eraf Beta globin RNA polymerase II Early Late Active chromatin DNaseI Expression foci Gene position is correlated with gene activity Replication timing Hutchison and Weintraub, Cell (1985 ) Osborne et al. Nature Genet. (2004) Schermelleh et al. Chromosome Res. (2001)
6 Increase resolution Increase throughput Reduce bias
7 Chromosome Conformation Capture (3C) 1. Crosslinking With Formaldehyde Cells Cross link FA%-0%-2% we commonly use 1% Time- commonly 10 minutes Temperature- commonly 37 C
8 2. lysis? Where is the chromatin? ` Cell lysis 0.3% SDS 1h Supernatant Pellet Cells/nuclei/debris In vivo formaldehyde cross-linking: it is time for black box analysis Gavrilov et al., PLoS One. 2013;8(3):e60403 Gavrilov et al., Brief Funct Genomics, in press
9 3. Endonuclease digestion Cross link Endonuclease digestion Search the literature for validated enzymes capable digesting in 3C conditions (SDS, Triton ) 6 b.p. cutter 4 b.p. butter We will discuss resolution issues
10 4. Ligation Cross link Endonuclease digestion Ligation Quality control From Splinter et al., Methods 2012
11 4. Ligation Cross link Endonuclease digestion Ligation Low DTT in ligation buffer is important for the following DNA precipitation 5mM 2mM Multiple ligases can work in low DTT concentrations suitable for 3C Schwartz et al., Biotechniques in press
12 5. Reverse cross-links Cross link Endonuclease digestion Ligation Reverse cross-links This is the 3C DNA library
13 Chromosome Conformation Capture (3C) Enhancer Principle Coverage Detection Resolution Limitations Examples Contacts between two defined regions Commonly <1Mb Locus-specific PCR High Low throughput and coverage Determine interaction between a known promoter and enhancer Dekker et al., Nature Reviews Genetics 14, (2013)
14 Carbon Copy 3C (5C) Principle Coverage Detection Resolution Limitations Examples All against all Commonly <1Mb Multiplex PCR, HT-sequencing High Limited coverage Determine comprehensively higher-order chromosome structure in a defined region Multiplex PCR Limb-specific contacts Berlivet et al., PLoS Genet. (2013)
15 Circular 3C (4C) Second digestion 3C DNA Ligation PCR Sequencing
16 Circular 3C (4C) Principle Coverage Detection Resolution Limitations Examples All contacts with a point of interest Genome-wide PCR, HT-sequencing High Limited to one view point All genes and genomic elements associated with a known LCR
17 Lipocalin2 Hierarchical (Lcn2) Contact as a bait Landscape for 4C View point Avital Sarusi
18 Resolution Lipocalin2 - proximal (Lcn2) as a range bait for 10kb- 4C 1000kb High contact frequency 6b.p. cutter Bait 4b.p. cutter
19 Resolution - far cis (Mb) and trans Low contact frequency 6b.p. cutter Bait 100k.b. 100k.b. 4b.p. cutter 100k.b. 100k.b.
20 Mknk2 Long-Range Contacts High-Resolution Contacts 1h after induction Bait GR ChIP DHS Rachel Deitch, Dana Raz, Moran Tal Analysis method- van de Werken et al., Nat. Methods 2012
21 ChIA-PET 3C Principle All contacts associated with a given protein Coverage Genome wide * Immunoprecipitation Detection Resolution Limitations Examples paired end HT-sequencing High Rely on one factor, disregarding other contacts Map chromatin interaction network of a known transcription factor Keifer-Kwon et al., 2013
22 Capture-C Sub-sampling of Hi-C Principle Coverage Detection Resolution Limitations Examples Focused all against all Up to genome-wide HT-sequencing High Limited coverage Determine comprehensively higher-order chromosome structure in multiple defined regions Hughes et al., 2014
23 Chromosome Circular 3C (4C) Chromosome 2 View point
24 Correlation with 4C signal What does a C signal mean? DNA FISH R 2 = nuclei d=<0.5 micron 3D distance threshold (microns) Bait
25 Hi-C Principle All against all Coverage Genome-wide * Detection Paired end HT-sequencing Resolution Low * Limitations Examples All intra- and inter- chromosomal associations Dekker et al., Nature Reviews Genetics 14, (2013)
26 Resolution is critical ~ 20 million read pairs ~ 1 billion read pairs Liebermann-Aiden et al., 2009 Dixon et al., 2012
27 Topological associated domain (TAD) Negabase-sized local chromatin interaction domains Dixon et al., 2012
28 Domains within domains Khalor et al., 2011
29 Chromosomes are organized in territories Chromosomes are organized in territories Nature Reviews Genetics 6, 429 Liebermann-Aiden et al., 2009
30 Preferential contacts within and between chromosomes There are two compartments :A and B Are there sub-compartment structures? Liebermann-Aiden et al., 2009
31 Genomic associations: Gene density, Gene activity Compartment A B Compartment A is gene rich Compartment A Genes Spearman s ρ = Expression Spearman s ρ = Accessible chromatin, Spearman s ρ = H3K36 trimethylation, Spearman s ρ = (active) H3K27 trimethylation, Spearman s ρ = (repressive ) A is more closely associated with open, accessible, actively transcribed chromatin. Liebermann-Aiden et al., 2009 Shopland, et al 2006
32 Contacts between TADs are cell-type specific 5 Ifng contacts Global % of genes 0
33 Contacts between TADs are cell-type specific Ifng Chr10 PPARg Chr6 High enrichment for adipogenic genes Lpin1 Chr12 PPARg 34% Lpin1 Ifng Avital Sarusi
Lieberman-Aiden et al. (2009) Comprehensive Mapping of Long-Range Interactions Reveals Folding Principles of the Human Genome. Science 326:
 Lieberman-Aiden et al. (2009) Comprehensive Mapping of Long-Range Interactions Reveals Folding Principles of the Human Genome. Science 326: 289-293. : Understanding the 3D conformation of the genome can
Lieberman-Aiden et al. (2009) Comprehensive Mapping of Long-Range Interactions Reveals Folding Principles of the Human Genome. Science 326: 289-293. : Understanding the 3D conformation of the genome can
Supplementary Figure 2
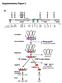 Supplementary Figure 2 a SBS-C1 SBS-C2 SBS-C3 SBS-C4 SBS-C5 SBS-C6 SBS-C7 SBS-C8 SBS-C9 LCR CNS-1 CNS-2 Il5 Rad50 Il13 Il4 Kif3a Sept8 0 50 100 150 200kb Sau3AI 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17
Supplementary Figure 2 a SBS-C1 SBS-C2 SBS-C3 SBS-C4 SBS-C5 SBS-C6 SBS-C7 SBS-C8 SBS-C9 LCR CNS-1 CNS-2 Il5 Rad50 Il13 Il4 Kif3a Sept8 0 50 100 150 200kb Sau3AI 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17
Structure meets function: Chromosome folding in mammals
 Transcription Regulation And Gene Expression in Eukaryotes FS 2016 Graduate Course G2 P Matthias and RG Clerc Pharmazentrum Hörsaal 2 16h15-18h00 Structure meets function: Chromosome folding in mammals
Transcription Regulation And Gene Expression in Eukaryotes FS 2016 Graduate Course G2 P Matthias and RG Clerc Pharmazentrum Hörsaal 2 16h15-18h00 Structure meets function: Chromosome folding in mammals
Bioinformatics of Transcriptional Regulation
 Bioinformatics of Transcriptional Regulation Carl Herrmann IPMB & DKFZ c.herrmann@dkfz.de Wechselwirkung von Maßnahmen und Auswirkungen Einflussmöglichkeiten in einem Dialog From genes to active compounds
Bioinformatics of Transcriptional Regulation Carl Herrmann IPMB & DKFZ c.herrmann@dkfz.de Wechselwirkung von Maßnahmen und Auswirkungen Einflussmöglichkeiten in einem Dialog From genes to active compounds
Measuring Protein-DNA interactions
 Measuring Protein-DNA interactions How is Biological Complexity Achieved? Mediated by Transcription Factors (TFs) 2 Transcription Factors are genetic switches 3 Regulation of Gene Expression by Transcription
Measuring Protein-DNA interactions How is Biological Complexity Achieved? Mediated by Transcription Factors (TFs) 2 Transcription Factors are genetic switches 3 Regulation of Gene Expression by Transcription
Mapping long-range promoter contacts in human cells with high-resolution capture Hi-C
 CORRECTION NOTICE Nat. Genet. 47, 598 606 (2015) Mapping long-range promoter contacts in human cells with high-resolution capture Hi-C Borbala Mifsud, Filipe Tavares-Cadete, Alice N Young, Robert Sugar,
CORRECTION NOTICE Nat. Genet. 47, 598 606 (2015) Mapping long-range promoter contacts in human cells with high-resolution capture Hi-C Borbala Mifsud, Filipe Tavares-Cadete, Alice N Young, Robert Sugar,
SUPPLEMENTAL MATERIALS. Chromatin immunoprecipitation assays. Single-cell suspensions of pituitary
 SUPPLEMENTAL MATERIALS METHODS Chromatin immunoprecipitation assays. Single-cell suspensions of pituitary and liver cells were prepared from the indicated transgenic mice, as described (Ho et al, 2002).
SUPPLEMENTAL MATERIALS METHODS Chromatin immunoprecipitation assays. Single-cell suspensions of pituitary and liver cells were prepared from the indicated transgenic mice, as described (Ho et al, 2002).
The role of erna in chromatin looping
 The role of erna in chromatin looping Introduction Enhancers are cis-acting DNA regulatory elements that activate transcription of target genes 1,2. It is estimated that 400 000 to >1 million alleged enhancers
The role of erna in chromatin looping Introduction Enhancers are cis-acting DNA regulatory elements that activate transcription of target genes 1,2. It is estimated that 400 000 to >1 million alleged enhancers
Introduction to genome biology
 Introduction to genome biology Lisa Stubbs Deep transcritpomes for traditional model species from ENCODE (and modencode) Deep RNA-seq and chromatin analysis on 147 human cell types, as well as tissues,
Introduction to genome biology Lisa Stubbs Deep transcritpomes for traditional model species from ENCODE (and modencode) Deep RNA-seq and chromatin analysis on 147 human cell types, as well as tissues,
Introduction to genome biology
 Introduction to genome biology Lisa Stubbs We ve found most genes; but what about the rest of the genome? Genome size* 12 Mb 95 Mb 170 Mb 1500 Mb 2700 Mb 3200 Mb #coding genes ~7000 ~20000 ~14000 ~26000
Introduction to genome biology Lisa Stubbs We ve found most genes; but what about the rest of the genome? Genome size* 12 Mb 95 Mb 170 Mb 1500 Mb 2700 Mb 3200 Mb #coding genes ~7000 ~20000 ~14000 ~26000
Permutation Clustering of the DNA Sequence Facilitates Understanding of the Nonlinearly Organized Genome
 RESEARCH PROPOSAL Permutation Clustering of the DNA Sequence Facilitates Understanding of the Nonlinearly Organized Genome Qiao JIN School of Medicine, Tsinghua University Advisor: Prof. Xuegong ZHANG
RESEARCH PROPOSAL Permutation Clustering of the DNA Sequence Facilitates Understanding of the Nonlinearly Organized Genome Qiao JIN School of Medicine, Tsinghua University Advisor: Prof. Xuegong ZHANG
Chapter 24: Promoters and Enhancers
 Chapter 24: Promoters and Enhancers A typical gene transcribed by RNA polymerase II has a promoter that usually extends upstream from the site where transcription is initiated the (#1) of transcription
Chapter 24: Promoters and Enhancers A typical gene transcribed by RNA polymerase II has a promoter that usually extends upstream from the site where transcription is initiated the (#1) of transcription
High-throughput genome scaffolding from in vivo DNA interaction frequency
 correction notice Nat. Biotechnol. 31, 1143 1147 (213) High-throughput genome scaffolding from in vivo DNA interaction frequency Noam Kaplan & Job Dekker In the version of this supplementary file originally
correction notice Nat. Biotechnol. 31, 1143 1147 (213) High-throughput genome scaffolding from in vivo DNA interaction frequency Noam Kaplan & Job Dekker In the version of this supplementary file originally
Multiple choice questions (numbers in brackets indicate the number of correct answers)
 1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
Differential Gene Expression
 Biology 4361 Developmental Biology Differential Gene Expression September 28, 2006 Chromatin Structure ~140 bp ~60 bp Transcriptional Regulation: 1. Packing prevents access CH 3 2. Acetylation ( C O )
Biology 4361 Developmental Biology Differential Gene Expression September 28, 2006 Chromatin Structure ~140 bp ~60 bp Transcriptional Regulation: 1. Packing prevents access CH 3 2. Acetylation ( C O )
Applied Bioinformatics - Lecture 16: Transcriptomics
 Applied Bioinformatics - Lecture 16: Transcriptomics David Hendrix Oregon State University Feb 15th 2016 Transcriptomics High-throughput Sequencing (deep sequencing) High-throughput sequencing (also
Applied Bioinformatics - Lecture 16: Transcriptomics David Hendrix Oregon State University Feb 15th 2016 Transcriptomics High-throughput Sequencing (deep sequencing) High-throughput sequencing (also
Supporting Information
 Supporting Information Gómez-Marín et al. 10.1073/pnas.1505463112 SI Materials and Methods Generation of BAC DK74B2-six2a::GFP-six3a::mCherry-iTol2. BAC clone (number 74B2) from DanioKey zebrafish BAC
Supporting Information Gómez-Marín et al. 10.1073/pnas.1505463112 SI Materials and Methods Generation of BAC DK74B2-six2a::GFP-six3a::mCherry-iTol2. BAC clone (number 74B2) from DanioKey zebrafish BAC
Supplemental Data. Noncoding Transcription by RNA Polymerase Pol IVb/Pol V Mediates Transcriptional Silencing of Overlapping and Adjacent Genes
 Cell, Volume 135 Supplemental Data Noncoding Transcription by RNA Polymerase Pol IVb/Pol V Mediates Transcriptional Silencing of Overlapping and Adjacent Genes Andrzej T. Wierzbicki, Jeremy R. Haag, and
Cell, Volume 135 Supplemental Data Noncoding Transcription by RNA Polymerase Pol IVb/Pol V Mediates Transcriptional Silencing of Overlapping and Adjacent Genes Andrzej T. Wierzbicki, Jeremy R. Haag, and
2/10/17. Contents. Applications of HMMs in Epigenomics
 2/10/17 I529: Machine Learning in Bioinformatics (Spring 2017) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2017 Background:
2/10/17 I529: Machine Learning in Bioinformatics (Spring 2017) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2017 Background:
Gene expression analysis. Biosciences 741: Genomics Fall, 2013 Week 5. Gene expression analysis
 Gene expression analysis Biosciences 741: Genomics Fall, 2013 Week 5 Gene expression analysis From EST clusters to spotted cdna microarrays Long vs. short oligonucleotide microarrays vs. RT-PCR Methods
Gene expression analysis Biosciences 741: Genomics Fall, 2013 Week 5 Gene expression analysis From EST clusters to spotted cdna microarrays Long vs. short oligonucleotide microarrays vs. RT-PCR Methods
Bio5488 Practice Midterm (2018) 1. Next-gen sequencing
 1. Next-gen sequencing 1. You have found a new strain of yeast that makes fantastic wine. You d like to sequence this strain to ascertain the differences from S. cerevisiae. To accurately call a base pair,
1. Next-gen sequencing 1. You have found a new strain of yeast that makes fantastic wine. You d like to sequence this strain to ascertain the differences from S. cerevisiae. To accurately call a base pair,
ChIP-seq analysis 2/28/2018
 ChIP-seq analysis 2/28/2018 Acknowledgements Much of the content of this lecture is from: Furey (2012) ChIP-seq and beyond Park (2009) ChIP-seq advantages + challenges Landt et al. (2012) ChIP-seq guidelines
ChIP-seq analysis 2/28/2018 Acknowledgements Much of the content of this lecture is from: Furey (2012) ChIP-seq and beyond Park (2009) ChIP-seq advantages + challenges Landt et al. (2012) ChIP-seq guidelines
EPIGENTEK. EpiQuik Chromatin Immunoprecipitation Kit. Base Catalog # P-2002 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE
 EpiQuik Chromatin Immunoprecipitation Kit Base Catalog # P-2002 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Chromatin Immunoprecipitation Kit is suitable for combining the specificity of
EpiQuik Chromatin Immunoprecipitation Kit Base Catalog # P-2002 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Chromatin Immunoprecipitation Kit is suitable for combining the specificity of
NGS Approaches to Epigenomics
 I519 Introduction to Bioinformatics, 2013 NGS Approaches to Epigenomics Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Background: chromatin structure & DNA methylation Epigenomic
I519 Introduction to Bioinformatics, 2013 NGS Approaches to Epigenomics Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Background: chromatin structure & DNA methylation Epigenomic
ChampionChIP Quick, High Throughput Chromatin Immunoprecipitation Assay System
 ChampionChIP Quick, High Throughput Chromatin Immunoprecipitation Assay System Liyan Pang, Ph.D. Application Scientist 1 Topics to be Covered Introduction What is ChIP-qPCR? Challenges Facing Biological
ChampionChIP Quick, High Throughput Chromatin Immunoprecipitation Assay System Liyan Pang, Ph.D. Application Scientist 1 Topics to be Covered Introduction What is ChIP-qPCR? Challenges Facing Biological
Multi-omics in biology: integration of omics techniques
 31/07/17 Летняя школа по биоинформатике 2017 Multi-omics in biology: integration of omics techniques Konstantin Okonechnikov Division of Pediatric Neurooncology German Cancer Research Center (DKFZ) 2 Short
31/07/17 Летняя школа по биоинформатике 2017 Multi-omics in biology: integration of omics techniques Konstantin Okonechnikov Division of Pediatric Neurooncology German Cancer Research Center (DKFZ) 2 Short
Genome 373: High- Throughput DNA Sequencing. Doug Fowler
 Genome 373: High- Throughput DNA Sequencing Doug Fowler Tasks give ML unity We learned about three tasks that are commonly encountered in ML Models/Algorithms Give ML Diversity Classification Regression
Genome 373: High- Throughput DNA Sequencing Doug Fowler Tasks give ML unity We learned about three tasks that are commonly encountered in ML Models/Algorithms Give ML Diversity Classification Regression
Chromatin. Structure and modification of chromatin. Chromatin domains
 Chromatin Structure and modification of chromatin Chromatin domains 2 DNA consensus 5 3 3 DNA DNA 4 RNA 5 ss RNA forms secondary structures with ds hairpins ds forms 6 of nucleic acids Form coiling bp/turn
Chromatin Structure and modification of chromatin Chromatin domains 2 DNA consensus 5 3 3 DNA DNA 4 RNA 5 ss RNA forms secondary structures with ds hairpins ds forms 6 of nucleic acids Form coiling bp/turn
THE ANALYSIS OF CHROMATIN CONDENSATION STATE AND TRANSCRIPTIONAL ACTIVITY USING DNA MICROARRAYS 1. INTRODUCTION
 JOURNAL OF MEDICAL INFORMATICS & TECHNOLOGIES Vol.6/2003, ISSN 1642-6037 Piotr WIDŁAK *, Krzysztof FUJAREWICZ ** DNA microarrays, chromatin, transcription THE ANALYSIS OF CHROMATIN CONDENSATION STATE AND
JOURNAL OF MEDICAL INFORMATICS & TECHNOLOGIES Vol.6/2003, ISSN 1642-6037 Piotr WIDŁAK *, Krzysztof FUJAREWICZ ** DNA microarrays, chromatin, transcription THE ANALYSIS OF CHROMATIN CONDENSATION STATE AND
HiChIP Protocol Mumbach et al. (CHANG), p. 1 Chang Lab, Stanford University. HiChIP Protocol
 HiChIP Protocol Mumbach et al. (CHANG), p. 1 HiChIP Protocol Citation: Mumbach et. al., HiChIP: Efficient and sensitive analysis of protein-directed genome architecture. Nature Methods (2016). Cell Crosslinking
HiChIP Protocol Mumbach et al. (CHANG), p. 1 HiChIP Protocol Citation: Mumbach et. al., HiChIP: Efficient and sensitive analysis of protein-directed genome architecture. Nature Methods (2016). Cell Crosslinking
H3K36me3 polyclonal antibody
 H3K36me3 polyclonal antibody Cat. No. C15410192 Type: Polyclonal ChIP-grade/ChIP-seq grade Source: Rabbit Lot #: A1845P Size: 50 µg/32 µl Concentration: 1.6 μg/μl Specificity: Human, mouse, Arabidopsis,
H3K36me3 polyclonal antibody Cat. No. C15410192 Type: Polyclonal ChIP-grade/ChIP-seq grade Source: Rabbit Lot #: A1845P Size: 50 µg/32 µl Concentration: 1.6 μg/μl Specificity: Human, mouse, Arabidopsis,
A more efficient, sensitive and robust method of chromatin immunoprecipitation (ChIP)
 A more efficient, sensitive and robust method of chromatin immunoprecipitation (ChIP) ADVANCEMENTS IN EPIGENETICS Introducing ChIP and Chromatrap Chromatrap is a more efficient, sensitive and robust method
A more efficient, sensitive and robust method of chromatin immunoprecipitation (ChIP) ADVANCEMENTS IN EPIGENETICS Introducing ChIP and Chromatrap Chromatrap is a more efficient, sensitive and robust method
EPIGENTEK. EpiQuik Tissue Chromatin Immunoprecipitation Kit. Base Catalog # P-2003 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE
 EpiQuik Tissue Chromatin Immunoprecipitation Kit Base Catalog # P-2003 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Tissue Chromatin Immunoprecipitation Kit is suitable for combining the specificity
EpiQuik Tissue Chromatin Immunoprecipitation Kit Base Catalog # P-2003 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Tissue Chromatin Immunoprecipitation Kit is suitable for combining the specificity
ChIP-seq and RNA-seq. Farhat Habib
 ChIP-seq and RNA-seq Farhat Habib fhabib@iiserpune.ac.in Biological Goals Learn how genomes encode the diverse patterns of gene expression that define each cell type and state. Protein-DNA interactions
ChIP-seq and RNA-seq Farhat Habib fhabib@iiserpune.ac.in Biological Goals Learn how genomes encode the diverse patterns of gene expression that define each cell type and state. Protein-DNA interactions
Year III Pharm.D Dr. V. Chitra
 Year III Pharm.D Dr. V. Chitra 1 Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Only one strand of DNA serves
Year III Pharm.D Dr. V. Chitra 1 Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Only one strand of DNA serves
Le proteine regolative variano nei vari tipi cellulari e in funzione degli stimoli ambientali
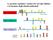 Le proteine regolative variano nei vari tipi cellulari e in funzione degli stimoli ambientali Tipo cellulare 1 Tipo cellulare 2 Tipo cellulare 3 DNA-protein Crosslink Lisi Frammentazione Immunopurificazione
Le proteine regolative variano nei vari tipi cellulari e in funzione degli stimoli ambientali Tipo cellulare 1 Tipo cellulare 2 Tipo cellulare 3 DNA-protein Crosslink Lisi Frammentazione Immunopurificazione
ChIP DNA Clean & Concentrator Catalog Nos. D5201 & D5205
 Page 0 INSTRUCTION MANUAL ChIP DNA Clean & Concentrator Catalog Nos. D5201 & D5205 Highlights Quick (2 minute) recovery of ultra-pure DNA from chromatin immunoprecipitation (ChIP) assays, cell lysates,
Page 0 INSTRUCTION MANUAL ChIP DNA Clean & Concentrator Catalog Nos. D5201 & D5205 Highlights Quick (2 minute) recovery of ultra-pure DNA from chromatin immunoprecipitation (ChIP) assays, cell lysates,
The ENCODE Encyclopedia. & Variant Annotation Using RegulomeDB and HaploReg
 The ENCODE Encyclopedia & Variant Annotation Using RegulomeDB and HaploReg Jill E. Moore Weng Lab University of Massachusetts Medical School October 10, 2015 Where s the Encyclopedia? ENCODE: Encyclopedia
The ENCODE Encyclopedia & Variant Annotation Using RegulomeDB and HaploReg Jill E. Moore Weng Lab University of Massachusetts Medical School October 10, 2015 Where s the Encyclopedia? ENCODE: Encyclopedia
ChIP-Seq Tools. J Fass UCD Genome Center Bioinformatics Core Wednesday September 16, 2015
 ChIP-Seq Tools J Fass UCD Genome Center Bioinformatics Core Wednesday September 16, 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA or
ChIP-Seq Tools J Fass UCD Genome Center Bioinformatics Core Wednesday September 16, 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA or
ChIP-Seq Data Analysis. J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015
 ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA
ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA
APPLICATION NOTE. Abstract. Introduction
 From minuscule amounts to magnificent results: reliable ChIP-seq data from 1, cells with the True MicroChIP and the MicroPlex Library Preparation kits Abstract Diagenode has developed groundbreaking solutions
From minuscule amounts to magnificent results: reliable ChIP-seq data from 1, cells with the True MicroChIP and the MicroPlex Library Preparation kits Abstract Diagenode has developed groundbreaking solutions
ChIP-Seq Data Analysis. J Fass UCD Genome Center Bioinformatics Core Wednesday December 17, 2014
 ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday December 17, 2014 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind
ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday December 17, 2014 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind
Supplementary Figure 1. HiChIP provides confident 1D factor binding information.
 Supplementary Figure 1 HiChIP provides confident 1D factor binding information. a, Reads supporting contacts called using the Mango pipeline 19 for GM12878 Smc1a HiChIP and GM12878 CTCF Advanced ChIA-PET
Supplementary Figure 1 HiChIP provides confident 1D factor binding information. a, Reads supporting contacts called using the Mango pipeline 19 for GM12878 Smc1a HiChIP and GM12878 CTCF Advanced ChIA-PET
REGULATION OF PROTEIN SYNTHESIS. II. Eukaryotes
 REGULATION OF PROTEIN SYNTHESIS II. Eukaryotes Complexities of eukaryotic gene expression! Several steps needed for synthesis of mrna! Separation in space of transcription and translation! Compartmentation
REGULATION OF PROTEIN SYNTHESIS II. Eukaryotes Complexities of eukaryotic gene expression! Several steps needed for synthesis of mrna! Separation in space of transcription and translation! Compartmentation
Supplemental Figure 1.
 Supplemental Data. Charron et al. Dynamic landscapes of four histone modifications during de-etiolation in Arabidopsis. Plant Cell (2009). 10.1105/tpc.109.066845 Supplemental Figure 1. Immunodetection
Supplemental Data. Charron et al. Dynamic landscapes of four histone modifications during de-etiolation in Arabidopsis. Plant Cell (2009). 10.1105/tpc.109.066845 Supplemental Figure 1. Immunodetection
Non-coding Function & Variation, MPRAs II. Mike White Bio /5/18
 Non-coding Function & Variation, MPRAs II Mike White Bio 5488 3/5/18 MPRA Review Problem 1: Where does your CRE DNA come from? DNA synthesis Genomic fragments Targeted regulome capture Problem 2: How do
Non-coding Function & Variation, MPRAs II Mike White Bio 5488 3/5/18 MPRA Review Problem 1: Where does your CRE DNA come from? DNA synthesis Genomic fragments Targeted regulome capture Problem 2: How do
DNA:CHROMATIN INTERACTIONS
 DNA:CHROMATIN INTERACTIONS Exploring transcription factor binding and the epigenomic landscape Chris Seward Introductions Cell and Developmental Biology PhD Candidate in Dr. Lisa Stubbs Laboratory Currently
DNA:CHROMATIN INTERACTIONS Exploring transcription factor binding and the epigenomic landscape Chris Seward Introductions Cell and Developmental Biology PhD Candidate in Dr. Lisa Stubbs Laboratory Currently
ChIP-seq and RNA-seq
 ChIP-seq and RNA-seq Biological Goals Learn how genomes encode the diverse patterns of gene expression that define each cell type and state. Protein-DNA interactions (ChIPchromatin immunoprecipitation)
ChIP-seq and RNA-seq Biological Goals Learn how genomes encode the diverse patterns of gene expression that define each cell type and state. Protein-DNA interactions (ChIPchromatin immunoprecipitation)
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Endogenous gene tagging to study subcellular localization and chromatin binding. a, b, Schematic of experimental set-up to endogenously tag RNAi factors using the CRISPR Cas9 technology,
Supplementary Figure 1 Endogenous gene tagging to study subcellular localization and chromatin binding. a, b, Schematic of experimental set-up to endogenously tag RNAi factors using the CRISPR Cas9 technology,
M Keramatipour 2. M Keramatipour 1. M Keramatipour 4. M Keramatipour 3. M Keramatipour 5. M Keramatipour
 Molecular Cloning Methods Mohammad Keramatipour MD, PhD keramatipour@tums.ac.ir Outline DNA recombinant technology DNA cloning co Cell based PCR PCR-based Some application of DNA cloning Genomic libraries
Molecular Cloning Methods Mohammad Keramatipour MD, PhD keramatipour@tums.ac.ir Outline DNA recombinant technology DNA cloning co Cell based PCR PCR-based Some application of DNA cloning Genomic libraries
nature methods A paired-end sequencing strategy to map the complex landscape of transcription initiation
 nature methods A paired-end sequencing strategy to map the complex landscape of transcription initiation Ting Ni, David L Corcoran, Elizabeth A Rach, Shen Song, Eric P Spana, Yuan Gao, Uwe Ohler & Jun
nature methods A paired-end sequencing strategy to map the complex landscape of transcription initiation Ting Ni, David L Corcoran, Elizabeth A Rach, Shen Song, Eric P Spana, Yuan Gao, Uwe Ohler & Jun
Supplementary Information
 Supplementary Information Super-resolution imaging of fluorescently labeled, endogenous RNA Polymerase II in living cells with CRISPR/Cas9-mediated gene editing Won-Ki Cho 1, Namrata Jayanth 1, Susan Mullen
Supplementary Information Super-resolution imaging of fluorescently labeled, endogenous RNA Polymerase II in living cells with CRISPR/Cas9-mediated gene editing Won-Ki Cho 1, Namrata Jayanth 1, Susan Mullen
Analysis of 3 dimensional interactions in DNA and chromatin
 Analysis of 3 dimensional interactions in DNA and chromatin Riina Kaukonen Degree project in Biology, Master of Science (2 years), 2010 Examensarbete I biologi 30 hp till masterexamen, 2010 Biology, Department
Analysis of 3 dimensional interactions in DNA and chromatin Riina Kaukonen Degree project in Biology, Master of Science (2 years), 2010 Examensarbete I biologi 30 hp till masterexamen, 2010 Biology, Department
EPIGENTEK. EpiQuik Methylated DNA Immunoprecipitation Kit. Base Catalog # P-2019 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE
 EpiQuik Methylated DNA Immunoprecipitation Kit Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik MeDIP Kit can be used for immunoprecipitating the methylated DNA from a broad range
EpiQuik Methylated DNA Immunoprecipitation Kit Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik MeDIP Kit can be used for immunoprecipitating the methylated DNA from a broad range
Gene expression DNA RNA. Protein DNA. Replication. Initiation Elongation Processing Export. DNA RNA Protein. Transcription. Degradation.
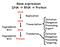 Gene expression DNA RNA Protein DNA DNA Degradation RNA Degradation Protein Replication Transcription Translation Initiation Elongation Processing Export Initiation Elongation Processing Targeting Chapter
Gene expression DNA RNA Protein DNA DNA Degradation RNA Degradation Protein Replication Transcription Translation Initiation Elongation Processing Export Initiation Elongation Processing Targeting Chapter
Control of Eukaryotic Genes. AP Biology
 Control of Eukaryotic Genes The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions? Evolution
Control of Eukaryotic Genes The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions? Evolution
Chapter 17 Lecture. Concepts of Genetics. Tenth Edition. Regulation of Gene Expression in Eukaryotes
 Chapter 17 Lecture Concepts of Genetics Tenth Edition Regulation of Gene Expression in Eukaryotes Chapter Contents 17.1 Eukaryotic Gene Regulation Can Occur at Any of the Steps Leading from DNA to Protein
Chapter 17 Lecture Concepts of Genetics Tenth Edition Regulation of Gene Expression in Eukaryotes Chapter Contents 17.1 Eukaryotic Gene Regulation Can Occur at Any of the Steps Leading from DNA to Protein
RNA was isolated using NucleoSpin RNA II (Macherey-Nagel, Bethlehem, PA) according to the
 Supplementary Methods RT-PCR and real-time PCR analysis RNA was isolated using NucleoSpin RNA II (Macherey-Nagel, Bethlehem, PA) according to the manufacturer s protocol and quantified by measuring the
Supplementary Methods RT-PCR and real-time PCR analysis RNA was isolated using NucleoSpin RNA II (Macherey-Nagel, Bethlehem, PA) according to the manufacturer s protocol and quantified by measuring the
Charles Girardot, Furlong Lab. MACS, CisGenome, SISSRs and other peak calling algorithms: differences and practical use
 Charles Girardot, Furlong Lab MACS, CisGenome, SISSRs and other peak calling algorithms: differences and practical use ChIP-Seq signal properties Only 5 ends of ChIPed fragments are sequenced Shifted read
Charles Girardot, Furlong Lab MACS, CisGenome, SISSRs and other peak calling algorithms: differences and practical use ChIP-Seq signal properties Only 5 ends of ChIPed fragments are sequenced Shifted read
Figure S1. Replication initiation sites at efficient ORIs do not coincide with nucleosomedepleted
 Figure S1. Replication initiation sites at efficient ORIs do not coincide with nucleosomedepleted regions in the mouse genome. Typical examples of SNS profiles (Cayrou et al., 11) and nucleosome occupancies
Figure S1. Replication initiation sites at efficient ORIs do not coincide with nucleosomedepleted regions in the mouse genome. Typical examples of SNS profiles (Cayrou et al., 11) and nucleosome occupancies
Supplemental Information. Autoregulatory Feedback Controls. Sequential Action of cis-regulatory Modules. at the brinker Locus
 Developmental Cell, Volume 26 Supplemental Information Autoregulatory Feedback Controls Sequential Action of cis-regulatory Modules at the brinker Locus Leslie Dunipace, Abbie Saunders, Hilary L. Ashe,
Developmental Cell, Volume 26 Supplemental Information Autoregulatory Feedback Controls Sequential Action of cis-regulatory Modules at the brinker Locus Leslie Dunipace, Abbie Saunders, Hilary L. Ashe,
Reading Lecture 8: Lecture 9: Lecture 8. DNA Libraries. Definition Types Construction
 Lecture 8 Reading Lecture 8: 96-110 Lecture 9: 111-120 DNA Libraries Definition Types Construction 142 DNA Libraries A DNA library is a collection of clones of genomic fragments or cdnas from a certain
Lecture 8 Reading Lecture 8: 96-110 Lecture 9: 111-120 DNA Libraries Definition Types Construction 142 DNA Libraries A DNA library is a collection of clones of genomic fragments or cdnas from a certain
The Genetic Code and Transcription. Chapter 12 Honors Genetics Ms. Susan Chabot
 The Genetic Code and Transcription Chapter 12 Honors Genetics Ms. Susan Chabot TRANSCRIPTION Copy SAME language DNA to RNA Nucleic Acid to Nucleic Acid TRANSLATION Copy DIFFERENT language RNA to Amino
The Genetic Code and Transcription Chapter 12 Honors Genetics Ms. Susan Chabot TRANSCRIPTION Copy SAME language DNA to RNA Nucleic Acid to Nucleic Acid TRANSLATION Copy DIFFERENT language RNA to Amino
Intracellular receptors specify complex patterns of gene expression that are cell and gene
 SUPPLEMENTAL RESULTS AND DISCUSSION Some HPr-1AR ARE-containing Genes Are Unresponsive to Androgen Intracellular receptors specify complex patterns of gene expression that are cell and gene specific. For
SUPPLEMENTAL RESULTS AND DISCUSSION Some HPr-1AR ARE-containing Genes Are Unresponsive to Androgen Intracellular receptors specify complex patterns of gene expression that are cell and gene specific. For
SNPs - GWAS - eqtls. Sebastian Schmeier
 SNPs - GWAS - eqtls s.schmeier@gmail.com http://sschmeier.github.io/bioinf-workshop/ 17.08.2015 Overview Single nucleotide polymorphism (refresh) SNPs effect on genes (refresh) Genome-wide association
SNPs - GWAS - eqtls s.schmeier@gmail.com http://sschmeier.github.io/bioinf-workshop/ 17.08.2015 Overview Single nucleotide polymorphism (refresh) SNPs effect on genes (refresh) Genome-wide association
Nature Genetics: doi: /ng.3556 INTEGRATED SUPPLEMENTARY FIGURE TEMPLATE. Supplementary Figure 1
 INTEGRATED SUPPLEMENTARY FIGURE TEMPLATE Supplementary Figure 1 REF6 expression in transgenic lines. (a,b) Expression of REF6 in REF6-HA ref6 and REF6ΔZnF-HA ref6 plants detected by RT qpcr (a) and immunoblot
INTEGRATED SUPPLEMENTARY FIGURE TEMPLATE Supplementary Figure 1 REF6 expression in transgenic lines. (a,b) Expression of REF6 in REF6-HA ref6 and REF6ΔZnF-HA ref6 plants detected by RT qpcr (a) and immunoblot
Nuclear Organization and Gene Expression Dr. David L. Spector
 NUCLEAR ORGANIZATION AND GENE EXPRESSION David L. Spector, Ph.D. Cold Spring Harbor Laboratory One Bungtown Road Cold Spring Harbor, New York 11724 Visit our website at www.cshl.edu/spectorlab 1 Orphanides
NUCLEAR ORGANIZATION AND GENE EXPRESSION David L. Spector, Ph.D. Cold Spring Harbor Laboratory One Bungtown Road Cold Spring Harbor, New York 11724 Visit our website at www.cshl.edu/spectorlab 1 Orphanides
Edexcel (B) Biology A-level
 Edexcel (B) Biology A-level Topic 7: Modern Genetics Notes Using Gene Sequencing Genome = all of an organism s DNA, including mitochondrial/chloroplast DNA. Polymerase chain reaction (PCR) is used to amplify
Edexcel (B) Biology A-level Topic 7: Modern Genetics Notes Using Gene Sequencing Genome = all of an organism s DNA, including mitochondrial/chloroplast DNA. Polymerase chain reaction (PCR) is used to amplify
Next- genera*on Sequencing. Lecture 13
 Next- genera*on Sequencing Lecture 13 ChIP- seq Applica*ons iden%fy sequence varia%ons DNA- seq Iden%fy Pathogens RNA- seq Kahvejian et al, 2008 Protein-DNA interaction DNA is the informa*on carrier of
Next- genera*on Sequencing Lecture 13 ChIP- seq Applica*ons iden%fy sequence varia%ons DNA- seq Iden%fy Pathogens RNA- seq Kahvejian et al, 2008 Protein-DNA interaction DNA is the informa*on carrier of
Complete Sample to Analysis Solutions for DNA Methylation Discovery using Next Generation Sequencing
 Complete Sample to Analysis Solutions for DNA Methylation Discovery using Next Generation Sequencing SureSelect Human/Mouse Methyl-Seq Kyeong Jeong PhD February 5, 2013 CAG EMEAI DGG/GSD/GFO Agilent Restricted
Complete Sample to Analysis Solutions for DNA Methylation Discovery using Next Generation Sequencing SureSelect Human/Mouse Methyl-Seq Kyeong Jeong PhD February 5, 2013 CAG EMEAI DGG/GSD/GFO Agilent Restricted
Wednesday, November 22, 17. Exons and Introns
 Exons and Introns Introns and Exons Exons: coded regions of DNA that get transcribed and translated into proteins make up 5% of the genome Introns and Exons Introns: non-coded regions of DNA Must be removed
Exons and Introns Introns and Exons Exons: coded regions of DNA that get transcribed and translated into proteins make up 5% of the genome Introns and Exons Introns: non-coded regions of DNA Must be removed
Chromatin and Transcription
 Chromatin and Transcription Chromatin Structure Chromatin Represses Transcription Nucleosome Positioning Histone Acetylation Chromatin Remodeling Histone Methylation CHIP Analysis Chromatin and Elongation
Chromatin and Transcription Chromatin Structure Chromatin Represses Transcription Nucleosome Positioning Histone Acetylation Chromatin Remodeling Histone Methylation CHIP Analysis Chromatin and Elongation
Supplementary Information
 Supplementary Information Table of contents: Pages: Supplementary Methods 2-3 Supplementary Figure 1 4 Supplementary Figure 2 5 Supplementary Figure 3 6 Supplementary Figure 4 7 Supplementary Figure 5
Supplementary Information Table of contents: Pages: Supplementary Methods 2-3 Supplementary Figure 1 4 Supplementary Figure 2 5 Supplementary Figure 3 6 Supplementary Figure 4 7 Supplementary Figure 5
MODULE TSS1: TRANSCRIPTION START SITES INTRODUCTION (BASIC)
 MODULE TSS1: TRANSCRIPTION START SITES INTRODUCTION (BASIC) Lesson Plan: Title JAMIE SIDERS, MEG LAAKSO & WILSON LEUNG Identifying transcription start sites for Peaked promoters using chromatin landscape,
MODULE TSS1: TRANSCRIPTION START SITES INTRODUCTION (BASIC) Lesson Plan: Title JAMIE SIDERS, MEG LAAKSO & WILSON LEUNG Identifying transcription start sites for Peaked promoters using chromatin landscape,
6/256 1/256 0/256 1/256 2/256 7/256 10/256. At3g06290 (SAC3B)
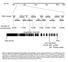 Chr.III M 5M 1M 15M 2M 23M BAC clones F22F7 F1A16 F24F17 F24P17 T8E24 F17A9 F21O3 F17A17 F18C1 F2O1 F28L1 F5E6 F3E22 T1B9 MLP3 Number of recombinants 6/256 1/256 /256 1/256 2/256 7/256 1/256 At3g629 (SAC3B)
Chr.III M 5M 1M 15M 2M 23M BAC clones F22F7 F1A16 F24F17 F24P17 T8E24 F17A9 F21O3 F17A17 F18C1 F2O1 F28L1 F5E6 F3E22 T1B9 MLP3 Number of recombinants 6/256 1/256 /256 1/256 2/256 7/256 1/256 At3g629 (SAC3B)
Identifying multi-locus chromatin contacts in human cells using tethered multiple 3C
 Ay et al. BMC Genomics (2015) 16:121 DOI 10.1186/s12864-015-1236-7 RESEARCH ARTICLE Open Access Identifying multi-locus chromatin contacts in human cells using tethered multiple 3C Ferhat Ay 1, Thanh H
Ay et al. BMC Genomics (2015) 16:121 DOI 10.1186/s12864-015-1236-7 RESEARCH ARTICLE Open Access Identifying multi-locus chromatin contacts in human cells using tethered multiple 3C Ferhat Ay 1, Thanh H
Differential Gene Expression
 Biology 4361 Developmental Biology Differential Gene Expression June 19, 2008 Differential Gene Expression Overview Chromatin structure Gene anatomy RNA processing and protein production Initiating transcription:
Biology 4361 Developmental Biology Differential Gene Expression June 19, 2008 Differential Gene Expression Overview Chromatin structure Gene anatomy RNA processing and protein production Initiating transcription:
The ENCODE Encyclopedia. Zhiping Weng University of Massachusetts Medical School ENCODE 2016: Research Applications and Users Meeting June 8, 2016
 The ENCODE Encyclopedia Zhiping Weng University of Massachusetts Medical School ENCODE 2016: Research Applications and Users Meeting June 8, 2016 The Encyclopedia Of DNA Elements Consortium Goals: - Catalog
The ENCODE Encyclopedia Zhiping Weng University of Massachusetts Medical School ENCODE 2016: Research Applications and Users Meeting June 8, 2016 The Encyclopedia Of DNA Elements Consortium Goals: - Catalog
GENETICS - CLUTCH CH.10 TRANSCRIPTION.
 !! www.clutchprep.com CONCEPT: OVERVIEW OF TRANSCRIPTION Transcription is the process of using DNA as a template to RNA RNA polymerase is the enzyme that transcribes DNA - There are many different types
!! www.clutchprep.com CONCEPT: OVERVIEW OF TRANSCRIPTION Transcription is the process of using DNA as a template to RNA RNA polymerase is the enzyme that transcribes DNA - There are many different types
Functional Genomics II
 Functional Genomics II Jay Hesselberth PhD University of Colorado SOM Biochemistry & Molecular Genetics jay.hesselberth@gmail.com Outline Tues, May. Definition. Yeast genome 3. Gene-centric resources and
Functional Genomics II Jay Hesselberth PhD University of Colorado SOM Biochemistry & Molecular Genetics jay.hesselberth@gmail.com Outline Tues, May. Definition. Yeast genome 3. Gene-centric resources and
7.03, 2005, Lecture 20 EUKARYOTIC GENES AND GENOMES I
 7.03, 2005, Lecture 20 EUKARYOTIC GENES AND GENOMES I For the last several lectures we have been looking at how one can manipulate prokaryotic genomes and how prokaryotic genes are regulated. In the next
7.03, 2005, Lecture 20 EUKARYOTIC GENES AND GENOMES I For the last several lectures we have been looking at how one can manipulate prokaryotic genomes and how prokaryotic genes are regulated. In the next
BS 50 Genetics and Genomics Week of Oct 24
 BS 50 Genetics and Genomics Week of Oct 24 Additional Practice Problems for Section Question 1: The following table contains a list of statements that apply to replication, transcription, both, or neither.
BS 50 Genetics and Genomics Week of Oct 24 Additional Practice Problems for Section Question 1: The following table contains a list of statements that apply to replication, transcription, both, or neither.
A Repressor Complex Governs the Integration of
 Developmental Cell 15 Supplemental Data A Repressor Complex Governs the Integration of Flowering Signals in Arabidopsis Dan Li, Chang Liu, Lisha Shen, Yang Wu, Hongyan Chen, Masumi Robertson, Chris A.
Developmental Cell 15 Supplemental Data A Repressor Complex Governs the Integration of Flowering Signals in Arabidopsis Dan Li, Chang Liu, Lisha Shen, Yang Wu, Hongyan Chen, Masumi Robertson, Chris A.
Epigenetics and DNase-Seq
 Epigenetics and DNase-Seq BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2018 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material, are licensed under CC BY-NC 4.0 by Anthony
Epigenetics and DNase-Seq BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2018 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material, are licensed under CC BY-NC 4.0 by Anthony
Long-range gene regulation
 Long-range gene regulation Short Course in Medical Genetics Melbourne, June 2011 Question Why should long-range regulation of gene expression be of interest to Clinical Scientists and Pathologists interested
Long-range gene regulation Short Course in Medical Genetics Melbourne, June 2011 Question Why should long-range regulation of gene expression be of interest to Clinical Scientists and Pathologists interested
Quantitative and non-quantitative RT-PCR. cdna was generated from 500ng RNA (iscript;
 Supplemental Methods Quantitative and non-quantitative RT-PCR. cdna was generated from 500ng RNA (iscript; Bio-Rad, Hercules, CA, USA) and standard RT-PCR experiments were carried out using the 2X GoTaq
Supplemental Methods Quantitative and non-quantitative RT-PCR. cdna was generated from 500ng RNA (iscript; Bio-Rad, Hercules, CA, USA) and standard RT-PCR experiments were carried out using the 2X GoTaq
Why DNA Isn t Your Destiny
 Technical Journal Club April 7 2015 Why DNA Isn t Your Destiny Despoina Goniotaki Aguzzi Group GENOME: a genome is an organism s complete set of DNA, including all of its genes. Each genome contains all
Technical Journal Club April 7 2015 Why DNA Isn t Your Destiny Despoina Goniotaki Aguzzi Group GENOME: a genome is an organism s complete set of DNA, including all of its genes. Each genome contains all
Nature Genetics: doi: /ng Supplementary Figure 1. ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets.
 Supplementary Figure 1 ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets. Gene structures are shown underneath each panel. Supplementary Figure 2 pref6::ref6-gfp complements
Supplementary Figure 1 ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets. Gene structures are shown underneath each panel. Supplementary Figure 2 pref6::ref6-gfp complements
Gene Expression Microarrays. For microarrays, purity of the RNA was further assessed by
 Supplemental Methods Gene Expression Microarrays. For microarrays, purity of the RNA was further assessed by an Agilent 2100 Bioanalyzer. 500 ng of RNA was reverse transcribed into crna and biotin-utp
Supplemental Methods Gene Expression Microarrays. For microarrays, purity of the RNA was further assessed by an Agilent 2100 Bioanalyzer. 500 ng of RNA was reverse transcribed into crna and biotin-utp
DNA METHYLATION NH 2 H 3. Solutions using bisulfite conversion and immunocapture Ideal for NGS, Sanger sequencing, Pyrosequencing, and qpcr
 NH 2 DNA METHYLATION H 3 C Solutions using bisulfite conversion and immunocapture Ideal for NGS, Sanger sequencing, Pyrosequencing, and qpcr NH 2 N N H PAGE 3 Understanding DNA Methylation DNA methylation
NH 2 DNA METHYLATION H 3 C Solutions using bisulfite conversion and immunocapture Ideal for NGS, Sanger sequencing, Pyrosequencing, and qpcr NH 2 N N H PAGE 3 Understanding DNA Methylation DNA methylation
Enzyme that uses RNA as a template to synthesize a complementary DNA
 Biology 105: Introduction to Genetics PRACTICE FINAL EXAM 2006 Part I: Definitions Homology: Comparison of two or more protein or DNA sequence to ascertain similarities in sequences. If two genes have
Biology 105: Introduction to Genetics PRACTICE FINAL EXAM 2006 Part I: Definitions Homology: Comparison of two or more protein or DNA sequence to ascertain similarities in sequences. If two genes have
Gene Regulation 10/19/05
 10/19/05 Gene Regulation (formerly Gene Prediction - 2) Gene Prediction & Regulation Mon - Overview & Gene structure review: Eukaryotes vs prokaryotes Wed - Regulatory regions: Promoters & enhancers -
10/19/05 Gene Regulation (formerly Gene Prediction - 2) Gene Prediction & Regulation Mon - Overview & Gene structure review: Eukaryotes vs prokaryotes Wed - Regulatory regions: Promoters & enhancers -
Exploring the three-dimensional organization of genomes: interpreting chromatin interaction data
 Exploring the three-dimensional organization of genomes: interpreting chromatin interaction data Job Dekker 1, Marc A. Marti-Renom 2,3 and Leonid A. Mirny 4 Abstract How DNA is organized in three dimensions
Exploring the three-dimensional organization of genomes: interpreting chromatin interaction data Job Dekker 1, Marc A. Marti-Renom 2,3 and Leonid A. Mirny 4 Abstract How DNA is organized in three dimensions
Integrated NGS Sample Preparation Solutions for Limiting Amounts of RNA and DNA. March 2, Steven R. Kain, Ph.D. ABRF 2013
 Integrated NGS Sample Preparation Solutions for Limiting Amounts of RNA and DNA March 2, 2013 Steven R. Kain, Ph.D. ABRF 2013 NuGEN s Core Technologies Selective Sequence Priming Nucleic Acid Amplification
Integrated NGS Sample Preparation Solutions for Limiting Amounts of RNA and DNA March 2, 2013 Steven R. Kain, Ph.D. ABRF 2013 NuGEN s Core Technologies Selective Sequence Priming Nucleic Acid Amplification
Bootcamp: Molecular Biology Techniques and Interpretation
 Bootcamp: Molecular Biology Techniques and Interpretation Bi8 Winter 2016 Today s outline Detecting and quantifying nucleic acids and proteins: Basic nucleic acid properties Hybridization PCR and Designing
Bootcamp: Molecular Biology Techniques and Interpretation Bi8 Winter 2016 Today s outline Detecting and quantifying nucleic acids and proteins: Basic nucleic acid properties Hybridization PCR and Designing
#9003. SimpleChIP Enzymatic Chromatin IP Kit (Magnetic Beads) n 1 Kit. (30 Immunoprecipitations) and 20 C. Store at 4 C, RT
 Store at 4 C, RT and 20 C #9003 SimpleChIP Enzymatic Chromatin IP Kit (Magnetic Beads) 3 n 1 Kit (30 Immunoprecipitations) For Research Use Only. Not For Use In Diagnostic Procedures. rev. 06/15/18 Orders
Store at 4 C, RT and 20 C #9003 SimpleChIP Enzymatic Chromatin IP Kit (Magnetic Beads) 3 n 1 Kit (30 Immunoprecipitations) For Research Use Only. Not For Use In Diagnostic Procedures. rev. 06/15/18 Orders
Supplemental Data. Zhou et al. (2016). Plant Cell /tpc
 Supplemental Figure 1. Confirmation of mutant mapping results. (A) Complementation assay with stably transformed genomic fragments (ComN-N) (2 kb upstream of TSS and 1.5 kb downstream of TES) and CaMV
Supplemental Figure 1. Confirmation of mutant mapping results. (A) Complementation assay with stably transformed genomic fragments (ComN-N) (2 kb upstream of TSS and 1.5 kb downstream of TES) and CaMV
Applications of HMMs in Epigenomics
 I529: Machine Learning in Bioinformatics (Spring 2013) Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Contents Background:
I529: Machine Learning in Bioinformatics (Spring 2013) Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Contents Background:
GENE REGULATION. Gene regulation occurs at the level of transcription or production of mrna
 GENE REGULATION Virtually every cell in your body contains a complete set of genes But they are not all turned on in every tissue Each cell in your body expresses only a small subset of genes at any time
GENE REGULATION Virtually every cell in your body contains a complete set of genes But they are not all turned on in every tissue Each cell in your body expresses only a small subset of genes at any time
2/19/13. Contents. Applications of HMMs in Epigenomics
 2/19/13 I529: Machine Learning in Bioinformatics (Spring 2013) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Background:
2/19/13 I529: Machine Learning in Bioinformatics (Spring 2013) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Background:
