Cambridge University Press
|
|
|
- Patience Whitehead
- 6 years ago
- Views:
Transcription
1 Figure 1.1. Model of RNAi pathway in C. elegans. Transmembrane protein SID-1 allows dsrna to enter the cell. In the cytoplasm,dsrna gets processed by DCR-1,existing in a complex with RDE-4,RDE-1 and DRH-1. The resulting sirna is shown bound to RDE-1. Double stranded sirna is unwound by a helicase,allowing complex to bind single strands of sirna. RDE-2 and MUT-7 might help in bringing together target mrna and antisense sirna (white) bound to.,guided by sirna (white),degrades mrna. Antisense sirna also serves as a primer for RdRp,which uses mrna as a template and produces more antisense sirnas (orange). Sense sirnas (blue) do not accumulate in C. elegans.
2 d-sirnas dsrnas sirna amplification RdRP hairpin expression construct genomic sequence pol II + _ RdRP pri-mirna Drosha pol II or III large dsrna mirna precursor tncrna precursor shrna large dsrna mirna precursor tncrna precursor shrna sirna pool mirna tncrna sirna target gene pol II etc. substrates products + sirna-dependent DNA methylation Silencing mechnisms target mrna chemically-synthesized or in vitro-transcribed sirna Figure 2.1. Sequence dependent regulation of gene expression. Expression of particular genomic sequences produces various dsrnas. is responsible for processing the dsrnas into small RNAs. The small RNA is then incorporated into,guiding the protein complex to a specific mrna. enforces specific gene suppression either by cleavage of the mrna or inhibition of translation. The degree of complementarity between the small RNA and the mrna determines the fate of the mrna; perfect or near perfect complementarity generally results in cleavage of the mrna,whereas,a few mismatches results in suppression of translation. In a sequence specific manner,small RNAs also regulate chromatin structure and hence gene expression. Some of the processes depicted here may not be present in all organisms or cell types. From a practical standpoint small RNAs can be used to investigate gene function; several types of small dsrnas can either be introduced into the cell or produced inside the cell. In all cases the small RNA is funneled into the RNAi pathway and triggers specific gene suppression. H.s. Helicase DUF 283 PAZ RNase III RNase III dsrna BD 1922 aa M.m aa D.m aa D.m aa C.e. DCR aa S.p. C188.13c 1374 aa A.t. Carpel Factory 1909 aa Figure 2.2. Alignment of homologs. Most proteins contain six conserved domains: a DExH helicase domain; a domain of unknown function (DUF283); a PAZ domain; two RNase III catalytic domains; and finally a double stranded RNA binding domain. Spacing between the domains is different for the homologs,which may explain the different size classes of small RNAs found in some species.
3 RDE-4 DICER ATP mrna ATP Figure 3.1. Model for mrna degradation in the cytoplasm by RNAi. Introduced dsrnas (red) are recognized by RDE-4/R2D2,a dsrna binding protein. These dsrnas are then processed by into 21-23nt duplexes that can associate with an enzyme complex called. After unwinding of the sirnas, becomes competent to target homologous mrna transcripts for degradation. Figure 3.2. Amplification of dsrna by an RNA-dependent RNA polymerase. In certain organisms, new dsrnas can be generated by RDRPs,primed by sirnas on mrna targets. The new dsrnas can be used subsequently by to create more sirnas,which can lead to additional rounds of amplification.
4 Figure 3.3. Model for gene silencing in the nucleus by RNAi. RNAi can also silence the transcription of targeted genes in certain organisms. In this model a signal can direct a putative nuclear RNAi silencing complex (N),composed of chromatin modifying proteins,to the targeted locus, silencing gene expression at the level of transcription. Figure 5.4. Microtubule Associated Protein 2 (MAP2) suppression in primary cortical neurons by cognate 21nt-siRNAs. A. Double fluorescence staining of neurons transfected with non-specific sirna or with MAP2-siRNA. Upper panels-staining with MAP2 monoclonal antibody (green); lower panels-staining with actin-bound toxin phalloidin (red). B. Distribution of MAP2 expression levels in control and targeted cells,two different sirna (sirna1 and sirna2) show a very similar effect. In each experiment,at least 70 random neurons per experimental condition were analyzed and gene expression was quantified in both control and targeted cells. The figure is reprinted from: Krichevsky,A. M. and Kosik,K. S. RNAi functions in cultured mammalian neurons. Proc Natl Acad SciUSA., 99(18): (2002).
5 Figure 6.2. A model for the sirna-mediated RNAi mechanism depicting major steps in (1) initial duplex recognition by the pre- complex; (2) ATP-dependent activation and sirna unwinding,(3) target recognition,(4) target cleavage and product release. The desired outcome is directed by antisense strand entry into the to produce on-target silencing effects. Undesirable off-target effects are thought to be directed by sense strand identity to unrelated sequences but can be minimized using rational design coupled with comprehensive sequence analyses.
6 Non-transfected HeLa Clone 39 HeLa Clone % of Non-Transfected Real Time PCR Immunofluorescence NT Clone Name Figure Long Term Silencing of GAPDH with CMV Puro Plasmid. HeLa cells were transfected with a CMV puro plasmid expressing GAPDH-specific sirnas. The cells were cloned,and clonal populations were selected in 2.5 µg/ml puromycin. Three weeks after selection,gapdh expression was analyzed by (A) RT-PCR or (B) immunofluorescence. Expression levels of several cell clones are shown. Green: GAPDH. Blue: DAPI stained nuclei.
7 Figure Method for identifying effective shrna sequences. To screen for sirnas that are effective against a gene of interest,the gene to be targeted is cloned into an expression vector as a translational fusion to a fluorescent protein. This construct is then co-transfected with test and control sirna sequences against the gene. If the sirna sequence is effective,then expression of the fusion protein will be reduced,resulting in a loss of fluorescence. Figure RNAi in the neuroepithelium of E10 mouse embryos. E10 mouse embryos were injected,into the lumen of the telencephalic neural tube,with the two reporter plasmids pegfp- N2 (for GFP) and psvpaxd (for βgal),either without (a c and g, Control) or with (d-f and g, sirna) βgal-directed esirnas,followed by directional electroporation and whole embryo culture for 24 hours. (a-f) Horizontal cryosections through the targeted region of the telencenphalon were analysed by double fluorescence for expression of GFP (green; a and d) and βgal immunoreactivity (red; b and e). Co-expression of GFP and βgal in neuroepithelial cells appears yellow in the merge (c and f, arrowheads). Note the lack of βgal expression in neuroepithelial cells in the presence of βgal-directed esirnas. Upper and lower dashed lines indicate the lumenal (apical) surface and basal border of the neuroepithelium,respectively. Asterisks in (b and e) indicate signal due to the cross-reaction of the secondary antibody used to detect βgal with the basal lamina and underlying mesenchymal cells. Scale bar in (f),20 µm. (g) Quantitation of the percentage of GFP-expressing neuroepithelial cells that also express βgal without (Control) or with (sirna) application of βgaldirected esirnas. Data are the mean of three embryos analyzed as in (a-f); bars indicate S.D. (Reprinted figure with permission from PNAS).
8 A B Figure Chicken embryos are a good model system for developmental studies due to their accessibility. Chicken embryos can be accessed in ovo (A) through a window in the eggshell that can be resealed after manipulations with a coverslip and melted paraffin. As an alternative approach, chicken embryos can be used as ex ovo cultures (B). With both methods embryos can be kept alive throughout embryonic development. Figure The pzjm RNAi vector. The tet operator (Tet Op),dual T7 terminators (red octagons),tetinducible T7 promoters (T7 arrows),ribosomal DNA spacer (rdna),actin poly(a) addition sequence (ACT polya),phleomycin resistance gene (BLE),splice acceptor site (SAS),aldolase poly(a) addition sequence (ALD polya). The plasmid is shown in linearized form,after cleavage in the rdna spacer, and is not drawn to scale.
9 Figure (a) Albino and wild type (yellow) colonies obtained by transformation of the wild type strain with carb sequences. Segregation of albino (b) and wild-type (c) transformants after a cycle of vegetative growth. Colonies showing different phenotypes (arrows) are obtained from spores of the original transformants. Photographs were taken after illumination with blue light for 24 hours. Figure ACMV-[CM]-infected N. benthamiana showing recovery phenotype. N. benthamiana plants imaged at 2-weeks post inoculation [(WPI) (control-a-left; Infected-A-right)] and at 5-WPI (control-b-left; Infected-B-right).
10 Figure ACMV-[CM]-infected GFP silenced GFPtransgenic N. benthamiana (line 16C). Plant photographed using dissecting microscope (A) Normal light and (B) UV filter. Symptom-less recovered leaves appeared red under UV light. Figure Effect of anti-ptgs activity of AC2 gene of EACMCV and ICMV; and AC4 gene of ACMV-[CM] and SLCMV. Leaf of GFP-transgenic N. benthamiana (line 16C) plant agroinfiltrated with pbin-gfp alone (A),or bacterial mixture harboring pbin-gfp along with the following viral gene constructs,p1/hc-pro of TEV (B); AC4 of ACMV-[CM] (C),AC2 of EACMCV (D),AC4 of SLCMV (E) and AC2 of ICMV (F). Leaves were photographed 7 days after infiltration using a dissecting microscope.
Learning Objectives. Define RNA interference. Define basic terminology. Describe molecular mechanism. Define VSP and relevance
 Learning Objectives Define RNA interference Define basic terminology Describe molecular mechanism Define VSP and relevance Describe role of RNAi in antigenic variation A Nobel Way to Regulate Gene Expression
Learning Objectives Define RNA interference Define basic terminology Describe molecular mechanism Define VSP and relevance Describe role of RNAi in antigenic variation A Nobel Way to Regulate Gene Expression
Plants Fight it out Intrinsic defence mechanism The magic world of Gene silencing
 I LOVE YOU Plants Fight it out Intrinsic defence mechanism The magic world of Gene silencing Over expression of Chalcone synthase gene to get Purple Petunias Napoli, Lemieux & Jorgensen,1990 Desired Effect
I LOVE YOU Plants Fight it out Intrinsic defence mechanism The magic world of Gene silencing Over expression of Chalcone synthase gene to get Purple Petunias Napoli, Lemieux & Jorgensen,1990 Desired Effect
Concepts and Methods in Developmental Biology
 Biology 4361 Developmental Biology Concepts and Methods in Developmental Biology June 16, 2009 Conceptual and Methodological Tools Concepts Genomic equivalence Differential gene expression Differentiation/de-differentiation
Biology 4361 Developmental Biology Concepts and Methods in Developmental Biology June 16, 2009 Conceptual and Methodological Tools Concepts Genomic equivalence Differential gene expression Differentiation/de-differentiation
Thermo Scientific Dharmacon SMARTvector 2.0 Lentiviral shrna Particles
 Thermo Scientific Dharmacon SMARTvector 2.0 Lentiviral shrna Particles Long-term gene silencing shrna-specific design algorithm High titer, purified particles Thermo Scientific Dharmacon SMARTvector shrna
Thermo Scientific Dharmacon SMARTvector 2.0 Lentiviral shrna Particles Long-term gene silencing shrna-specific design algorithm High titer, purified particles Thermo Scientific Dharmacon SMARTvector shrna
pdsipher and pdsipher -GFP shrna Vector User s Guide
 pdsipher and pdsipher -GFP shrna Vector User s Guide NOTE: PLEASE READ THE ENTIRE PROTOCOL CAREFULLY BEFORE USE Page 1. Introduction... 1 2. Vector Overview... 1 3. Vector Maps 2 4. Materials Provided...
pdsipher and pdsipher -GFP shrna Vector User s Guide NOTE: PLEASE READ THE ENTIRE PROTOCOL CAREFULLY BEFORE USE Page 1. Introduction... 1 2. Vector Overview... 1 3. Vector Maps 2 4. Materials Provided...
MISSION shrna Library: Next Generation RNA Interference
 Page 1 of 6 Page 1 of 6 Return to Web Version MISSION shrna Library: Next Generation RNA Interference By: Stephanie Uder, Henry George, Betsy Boedeker, LSI Volume 6 Article 2 Introduction The technology
Page 1 of 6 Page 1 of 6 Return to Web Version MISSION shrna Library: Next Generation RNA Interference By: Stephanie Uder, Henry George, Betsy Boedeker, LSI Volume 6 Article 2 Introduction The technology
Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product.
 Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
DNA Transcription. Dr Aliwaini
 DNA Transcription 1 DNA Transcription-Introduction The synthesis of an RNA molecule from DNA is called Transcription. All eukaryotic cells have five major classes of RNA: ribosomal RNA (rrna), messenger
DNA Transcription 1 DNA Transcription-Introduction The synthesis of an RNA molecule from DNA is called Transcription. All eukaryotic cells have five major classes of RNA: ribosomal RNA (rrna), messenger
sirna Overview and Technical Tips
 1 sirna Overview and Technical Tips 2 CONTENTS 3 4 5 7 8 10 11 13 14 18 19 20 21 Introduction Applications How Does It Work? Handy Tips Troubleshooting Conclusions Further References Contact Us 3 INTRODUCTION
1 sirna Overview and Technical Tips 2 CONTENTS 3 4 5 7 8 10 11 13 14 18 19 20 21 Introduction Applications How Does It Work? Handy Tips Troubleshooting Conclusions Further References Contact Us 3 INTRODUCTION
RNA Interference and the World of Small RNAs
 RNA Interference and the World of Small RNAs O, I die, Horatio; The potent poison quite o'er-crows my spirit: I cannot live to hear the news from England; But I do prophesy the election lights On Fortinbras:
RNA Interference and the World of Small RNAs O, I die, Horatio; The potent poison quite o'er-crows my spirit: I cannot live to hear the news from England; But I do prophesy the election lights On Fortinbras:
Gene Expression: Transcription
 Gene Expression: Transcription The majority of genes are expressed as the proteins they encode. The process occurs in two steps: Transcription = DNA RNA Translation = RNA protein Taken together, they make
Gene Expression: Transcription The majority of genes are expressed as the proteins they encode. The process occurs in two steps: Transcription = DNA RNA Translation = RNA protein Taken together, they make
Eukaryotic Transcription
 Eukaryotic Transcription I. Differences between eukaryotic versus prokaryotic transcription. II. (core vs holoenzyme): RNA polymerase II - Promotor elements. - General Pol II transcription factors (GTF).
Eukaryotic Transcription I. Differences between eukaryotic versus prokaryotic transcription. II. (core vs holoenzyme): RNA polymerase II - Promotor elements. - General Pol II transcription factors (GTF).
Improving CRISPR-Cas9 Gene Knockout with a Validated Guide RNA Algorithm
 Improving CRISPR-Cas9 Gene Knockout with a Validated Guide RNA Algorithm Anja Smith Director R&D Dharmacon, part of GE Healthcare Imagination at work crrna:tracrrna program Cas9 nuclease Active crrna is
Improving CRISPR-Cas9 Gene Knockout with a Validated Guide RNA Algorithm Anja Smith Director R&D Dharmacon, part of GE Healthcare Imagination at work crrna:tracrrna program Cas9 nuclease Active crrna is
SureSilencing sirna Array Technology Overview
 SureSilencing sirna Array Technology Overview Pathway-Focused sirna-based RNA Interference Topics to be Covered Who is SuperArray? Brief Introduction to RNA Interference Challenges Facing RNA Interference
SureSilencing sirna Array Technology Overview Pathway-Focused sirna-based RNA Interference Topics to be Covered Who is SuperArray? Brief Introduction to RNA Interference Challenges Facing RNA Interference
MIT Department of Biology 7.013: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr.
 MIT Department of Biology 7.01: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel iv) Would Xba I be useful for cloning? Why or why not?
MIT Department of Biology 7.01: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel iv) Would Xba I be useful for cloning? Why or why not?
TOOLS sirna and mirna. User guide
 TOOLS sirna and mirna User guide Introduction RNA interference (RNAi) is a powerful tool for suppression gene expression by causing the destruction of specific mrna molecules. Small Interfering RNAs (sirnas)
TOOLS sirna and mirna User guide Introduction RNA interference (RNAi) is a powerful tool for suppression gene expression by causing the destruction of specific mrna molecules. Small Interfering RNAs (sirnas)
Genetics - Problem Drill 19: Dissection of Gene Function: Mutational Analysis of Model Organisms
 Genetics - Problem Drill 19: Dissection of Gene Function: Mutational Analysis of Model Organisms No. 1 of 10 1. The mouse gene knockout is based on. (A) Homologous recombination (B) Site-specific recombination
Genetics - Problem Drill 19: Dissection of Gene Function: Mutational Analysis of Model Organisms No. 1 of 10 1. The mouse gene knockout is based on. (A) Homologous recombination (B) Site-specific recombination
RNA synthesis/transcription I Biochemistry 302. February 6, 2004 Bob Kelm
 RNA synthesis/transcription I Biochemistry 302 February 6, 2004 Bob Kelm Overview of RNA classes Messenger RNA (mrna) Encodes protein Relatively short half-life ( 3 min in E. coli, 30 min in eukaryotic
RNA synthesis/transcription I Biochemistry 302 February 6, 2004 Bob Kelm Overview of RNA classes Messenger RNA (mrna) Encodes protein Relatively short half-life ( 3 min in E. coli, 30 min in eukaryotic
Differential Gene Expression
 Biology 4361 Developmental Biology Differential Gene Expression September 28, 2006 Chromatin Structure ~140 bp ~60 bp Transcriptional Regulation: 1. Packing prevents access CH 3 2. Acetylation ( C O )
Biology 4361 Developmental Biology Differential Gene Expression September 28, 2006 Chromatin Structure ~140 bp ~60 bp Transcriptional Regulation: 1. Packing prevents access CH 3 2. Acetylation ( C O )
Molecular Cell Biology - Problem Drill 11: Recombinant DNA
 Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
sirna Transfection Into Primary Neurons Using Fuse-It-siRNA
 sirna Transfection Into Primary Neurons Using Fuse-It-siRNA This Application Note describes a protocol for sirna transfection into sensitive, primary cortical neurons using Fuse-It-siRNA. This innovative
sirna Transfection Into Primary Neurons Using Fuse-It-siRNA This Application Note describes a protocol for sirna transfection into sensitive, primary cortical neurons using Fuse-It-siRNA. This innovative
Cell type-specific delivery of sirnas with aptamer-sirna chimeras
 Cell type-specific delivery of sirnas with aptamer-sirna chimeras Sullenger, B. A. et al Duke Center for Translational Research, Duke University Nature Biotechnology, 2006, 24, 1005 Julia Vargas November
Cell type-specific delivery of sirnas with aptamer-sirna chimeras Sullenger, B. A. et al Duke Center for Translational Research, Duke University Nature Biotechnology, 2006, 24, 1005 Julia Vargas November
Gene Expression and Heritable Phenotype. CBS520 Eric Nabity
 Gene Expression and Heritable Phenotype CBS520 Eric Nabity DNA is Just the Beginning DNA was determined to be the genetic material, and the structure was identified as a (double stranded) double helix.
Gene Expression and Heritable Phenotype CBS520 Eric Nabity DNA is Just the Beginning DNA was determined to be the genetic material, and the structure was identified as a (double stranded) double helix.
RNA Interference (RNAi) (see also sirna, micrna, shrna, etc.)
 Biochemistry 412 RNA Interference (RNAi) (see also sirna, micrna, shrna, etc.) April 3, 2007 The Discovery of the RNA Interference (RNAi) Phenomenon 1. Gene-specific inhibition of expression by anti-sense
Biochemistry 412 RNA Interference (RNAi) (see also sirna, micrna, shrna, etc.) April 3, 2007 The Discovery of the RNA Interference (RNAi) Phenomenon 1. Gene-specific inhibition of expression by anti-sense
Figure S1. Immunoblotting showing proper expression of tagged R and AVR proteins in N. benthamiana.
 SUPPLEMENTARY FIGURES Figure S1. Immunoblotting showing proper expression of tagged R and AVR proteins in N. benthamiana. YFP:, :CFP, HA: and (A) and HA, CFP and YFPtagged and AVR1-CO39 (B) were expressed
SUPPLEMENTARY FIGURES Figure S1. Immunoblotting showing proper expression of tagged R and AVR proteins in N. benthamiana. YFP:, :CFP, HA: and (A) and HA, CFP and YFPtagged and AVR1-CO39 (B) were expressed
ARN interférence Technologies : sirna, shrna, mirna
 ARN interférence Technologies : sirna, shrna, mirna LabCluster Tour 2014 - Delphine Ayache sigma-aldrich.com Methods for Modulating Gene Expression DNA RNA Protein Transcription Translation Genome Editing:
ARN interférence Technologies : sirna, shrna, mirna LabCluster Tour 2014 - Delphine Ayache sigma-aldrich.com Methods for Modulating Gene Expression DNA RNA Protein Transcription Translation Genome Editing:
Supplementary Figure 1. (a) The qrt-pcr for lnc-2, lnc-6 and lnc-7 RNA level in DU145, 22Rv1, wild type HCT116 and HCT116 Dicer ex5 cells transfected
 Supplementary Figure 1. (a) The qrt-pcr for lnc-2, lnc-6 and lnc-7 RNA level in DU145, 22Rv1, wild type HCT116 and HCT116 Dicer ex5 cells transfected with the sirna against lnc-2, lnc-6, lnc-7, and the
Supplementary Figure 1. (a) The qrt-pcr for lnc-2, lnc-6 and lnc-7 RNA level in DU145, 22Rv1, wild type HCT116 and HCT116 Dicer ex5 cells transfected with the sirna against lnc-2, lnc-6, lnc-7, and the
XactEdit Cas9 Nuclease with NLS User Manual
 XactEdit Cas9 Nuclease with NLS User Manual An RNA-guided recombinant endonuclease for efficient targeted DNA cleavage Catalog Numbers CE1000-50K, CE1000-50, CE1000-250, CE1001-250, CE1001-1000 Table of
XactEdit Cas9 Nuclease with NLS User Manual An RNA-guided recombinant endonuclease for efficient targeted DNA cleavage Catalog Numbers CE1000-50K, CE1000-50, CE1000-250, CE1001-250, CE1001-1000 Table of
TRANSCRIPTION COMPARISON OF DNA & RNA TRANSCRIPTION. Umm AL Qura University. Sugar Ribose Deoxyribose. Bases AUCG ATCG. Strand length Short Long
 Umm AL Qura University TRANSCRIPTION Dr Neda Bogari TRANSCRIPTION COMPARISON OF DNA & RNA RNA DNA Sugar Ribose Deoxyribose Bases AUCG ATCG Strand length Short Long No. strands One Two Helix Single Double
Umm AL Qura University TRANSCRIPTION Dr Neda Bogari TRANSCRIPTION COMPARISON OF DNA & RNA RNA DNA Sugar Ribose Deoxyribose Bases AUCG ATCG Strand length Short Long No. strands One Two Helix Single Double
Supplemental Information. Boundary Formation through a Direct. Threshold-Based Readout. of Mobile Small RNA Gradients
 Developmental Cell, Volume 43 Supplemental Information Boundary Formation through a Direct Threshold-Based Readout of Mobile Small RNA Gradients Damianos S. Skopelitis, Anna H. Benkovics, Aman Y. Husbands,
Developmental Cell, Volume 43 Supplemental Information Boundary Formation through a Direct Threshold-Based Readout of Mobile Small RNA Gradients Damianos S. Skopelitis, Anna H. Benkovics, Aman Y. Husbands,
DNA Transcription. Visualizing Transcription. The Transcription Process
 DNA Transcription By: Suzanne Clancy, Ph.D. 2008 Nature Education Citation: Clancy, S. (2008) DNA transcription. Nature Education 1(1) If DNA is a book, then how is it read? Learn more about the DNA transcription
DNA Transcription By: Suzanne Clancy, Ph.D. 2008 Nature Education Citation: Clancy, S. (2008) DNA transcription. Nature Education 1(1) If DNA is a book, then how is it read? Learn more about the DNA transcription
Chromatographic Separation of the three forms of RNA Polymerase II.
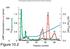 Chromatographic Separation of the three forms of RNA Polymerase II. α-amanitin α-amanitin bound to Pol II Function of the three enzymes. Yeast Pol II. RNA Polymerase Subunit Structures 10-7 Subunit structure.
Chromatographic Separation of the three forms of RNA Polymerase II. α-amanitin α-amanitin bound to Pol II Function of the three enzymes. Yeast Pol II. RNA Polymerase Subunit Structures 10-7 Subunit structure.
Figure S1. Figure S2. Figure S3 HB Anti-FSP27 (COOH-terminal peptide) Ab. Anti-GST-FSP27(45-127) Ab.
 / 36B4 mrna ratio Figure S1 * 2. 1.6 1.2.8 *.4 control TNFα BRL49653 Figure S2 Su bw AT p iw Anti- (COOH-terminal peptide) Ab Blot : Anti-GST-(45-127) Ab β-actin Figure S3 HB2 HW AT BA T Figure S4 A TAG
/ 36B4 mrna ratio Figure S1 * 2. 1.6 1.2.8 *.4 control TNFα BRL49653 Figure S2 Su bw AT p iw Anti- (COOH-terminal peptide) Ab Blot : Anti-GST-(45-127) Ab β-actin Figure S3 HB2 HW AT BA T Figure S4 A TAG
CHAPTER 20 DNA TECHNOLOGY AND GENOMICS. Section A: DNA Cloning
 Section A: DNA Cloning 1. DNA technology makes it possible to clone genes for basic research and commercial applications: an overview 2. Restriction enzymes are used to make recombinant DNA 3. Genes can
Section A: DNA Cloning 1. DNA technology makes it possible to clone genes for basic research and commercial applications: an overview 2. Restriction enzymes are used to make recombinant DNA 3. Genes can
Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling
 Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling Glendining KA 1, Markie D 2, Gardner RJM 4, Franz EA 3, Robertson SP 4, Jasoni CL 1 Supplementary
Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling Glendining KA 1, Markie D 2, Gardner RJM 4, Franz EA 3, Robertson SP 4, Jasoni CL 1 Supplementary
Differential Gene Expression
 Biology 4361 Developmental Biology Differential Gene Expression June 19, 2008 Differential Gene Expression Overview Chromatin structure Gene anatomy RNA processing and protein production Initiating transcription:
Biology 4361 Developmental Biology Differential Gene Expression June 19, 2008 Differential Gene Expression Overview Chromatin structure Gene anatomy RNA processing and protein production Initiating transcription:
CHAPTER 9 DNA Technologies
 CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
RNA Interference (RNAi) (see also mirna, sirna, micrna, shrna, etc.)
 Biochemistry 412 RNA Interference (RNAi) (see also mirna, sirna, micrna, shrna, etc.) April 8, 2008 The Discovery of the RNA Interference (RNAi) Phenomenon 1. Gene-specific inhibition of expression by
Biochemistry 412 RNA Interference (RNAi) (see also mirna, sirna, micrna, shrna, etc.) April 8, 2008 The Discovery of the RNA Interference (RNAi) Phenomenon 1. Gene-specific inhibition of expression by
Transcriptional Regulation in Eukaryotes
 Transcriptional Regulation in Eukaryotes Concepts, Strategies, and Techniques Michael Carey Stephen T. Smale COLD SPRING HARBOR LABORATORY PRESS NEW YORK 2000 Cold Spring Harbor Laboratory Press, 0-87969-537-4
Transcriptional Regulation in Eukaryotes Concepts, Strategies, and Techniques Michael Carey Stephen T. Smale COLD SPRING HARBOR LABORATORY PRESS NEW YORK 2000 Cold Spring Harbor Laboratory Press, 0-87969-537-4
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10163 Supplementary Table 1 Efficiency of vector construction. Process wells recovered efficiency (%) Recombineering* 480 461 96 Intermediate plasmids 461 381 83 Recombineering efficiency
doi:10.1038/nature10163 Supplementary Table 1 Efficiency of vector construction. Process wells recovered efficiency (%) Recombineering* 480 461 96 Intermediate plasmids 461 381 83 Recombineering efficiency
WORKING WITH THE FIGURES. 1. In Figure 8-3, why are the arrows for genes 1 and 2 pointing in opposite directions?
 8 RNA: Transcription and Processing WORKING WITH THE FIGURES 1. In Figure 8-3, why are the arrows for genes 1 and 2 pointing in opposite directions? The arrows for genes 1 and 2 indicate the direction
8 RNA: Transcription and Processing WORKING WITH THE FIGURES 1. In Figure 8-3, why are the arrows for genes 1 and 2 pointing in opposite directions? The arrows for genes 1 and 2 indicate the direction
PROTEOMICS AND FUNCTIONAL GENOMICS PRESENTATION KIMBERLY DONG
 PROTEOMICS AND FUNCTIONAL GENOMICS PRESENTATION KIMBERLY DONG A Lentiviral RNAi Library for Human and Mouse Genes Applied to an Arrayed Viral High-Content Screen Jason Moffat,1,2,4,10 Dorre A. Grueneberg,1,10
PROTEOMICS AND FUNCTIONAL GENOMICS PRESENTATION KIMBERLY DONG A Lentiviral RNAi Library for Human and Mouse Genes Applied to an Arrayed Viral High-Content Screen Jason Moffat,1,2,4,10 Dorre A. Grueneberg,1,10
Custom RNAi Services. GeneCust Europe. GeneCust Europe
 GeneCust Europe Laboratoire de Biotechnologie du Luxembourg S.A. 2 route de Remich L-5690 Ellange Luxembourg Tél. : +352 27620411 Fax : +352 27620412 Email : info@genecust.com Web : www.genecust.com Custom
GeneCust Europe Laboratoire de Biotechnologie du Luxembourg S.A. 2 route de Remich L-5690 Ellange Luxembourg Tél. : +352 27620411 Fax : +352 27620412 Email : info@genecust.com Web : www.genecust.com Custom
Gene Expression Technology
 Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
2054, Chap. 14, page 1
 2054, Chap. 14, page 1 I. Recombinant DNA technology (Chapter 14) A. recombinant DNA technology = collection of methods used to perform genetic engineering 1. genetic engineering = deliberate modification
2054, Chap. 14, page 1 I. Recombinant DNA technology (Chapter 14) A. recombinant DNA technology = collection of methods used to perform genetic engineering 1. genetic engineering = deliberate modification
Optimization of RNAi Targets on the Human Transcriptome Ahmet Arslan Kurdoglu Computational Biosciences Program Arizona State University
 Optimization of RNAi Targets on the Human Transcriptome Ahmet Arslan Kurdoglu Computational Biosciences Program Arizona State University my background Undergraduate Degree computer systems engineer (ASU
Optimization of RNAi Targets on the Human Transcriptome Ahmet Arslan Kurdoglu Computational Biosciences Program Arizona State University my background Undergraduate Degree computer systems engineer (ASU
Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit
 Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
NAME TA SEC Problem Set 4 FRIDAY October 15, Answers to this problem set must be inserted into the box outside
 MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel NAME TA SEC 7.012 Problem Set 4 FRIDAY October 15,
MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel NAME TA SEC 7.012 Problem Set 4 FRIDAY October 15,
AP Biology Gene Expression/Biotechnology REVIEW
 AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
Chapter 18: Regulation of Gene Expression. 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer
 Chapter 18: Regulation of Gene Expression 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer Gene Regulation Gene regulation refers to all aspects of controlling
Chapter 18: Regulation of Gene Expression 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer Gene Regulation Gene regulation refers to all aspects of controlling
The 5 end problem. 3 5 Now what? RNA primers. DNA template. elongate. excise primers, elongate, ligate
 1 2 3 DNA Replication Viruses must replicate their genomes to make new progeny This always requires expression of at least one virus protein, sometimes many (hence always delayed after infection) DNA is
1 2 3 DNA Replication Viruses must replicate their genomes to make new progeny This always requires expression of at least one virus protein, sometimes many (hence always delayed after infection) DNA is
Supplementary Information Design of small molecule-responsive micrornas based on structural requirements for Drosha processing
 Supplementary Information Design of small molecule-responsive micrornas based on structural requirements for Drosha processing Chase L. Beisel, Yvonne Y. Chen, Stephanie J. Culler, Kevin G. Hoff, & Christina
Supplementary Information Design of small molecule-responsive micrornas based on structural requirements for Drosha processing Chase L. Beisel, Yvonne Y. Chen, Stephanie J. Culler, Kevin G. Hoff, & Christina
Multiple choice questions (numbers in brackets indicate the number of correct answers)
 1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX
 Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX INTRODUCTION The CRISPR/Cas genome editing system consists of a single guide RNA
Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX INTRODUCTION The CRISPR/Cas genome editing system consists of a single guide RNA
Comparison of Commercial Transfection Reagents: Cell line optimized transfection kits for in vitro cancer research.
 Comparison of Commercial Transfection Reagents: Cell line optimized transfection kits for in vitro cancer research. by Altogen Labs, 11200 Manchaca Road, Suite 203 Austin TX 78748 USA Tel. (512) 433-6177
Comparison of Commercial Transfection Reagents: Cell line optimized transfection kits for in vitro cancer research. by Altogen Labs, 11200 Manchaca Road, Suite 203 Austin TX 78748 USA Tel. (512) 433-6177
RNA Structure and the Versatility of RNA. Mitesh Shrestha
 RNA Structure and the Versatility of RNA Mitesh Shrestha Ribonucleic Acid (RNA) Nitrogenous Bases (Adenine, Uracil, Guanine, Cytosine) Ribose Sugar Ribonucleic Acid (RNA) Phosphate Group RNA world Hypothesis
RNA Structure and the Versatility of RNA Mitesh Shrestha Ribonucleic Acid (RNA) Nitrogenous Bases (Adenine, Uracil, Guanine, Cytosine) Ribose Sugar Ribonucleic Acid (RNA) Phosphate Group RNA world Hypothesis
Chapter 11: Regulation of Gene Expression
 Chapter Review 1. It has long been known that there is probably a genetic link for alcoholism. Researchers studying rats have begun to elucidate this link. Briefly describe the genetic mechanism found
Chapter Review 1. It has long been known that there is probably a genetic link for alcoholism. Researchers studying rats have begun to elucidate this link. Briefly describe the genetic mechanism found
REGULATION OF PROTEIN SYNTHESIS. II. Eukaryotes
 REGULATION OF PROTEIN SYNTHESIS II. Eukaryotes Complexities of eukaryotic gene expression! Several steps needed for synthesis of mrna! Separation in space of transcription and translation! Compartmentation
REGULATION OF PROTEIN SYNTHESIS II. Eukaryotes Complexities of eukaryotic gene expression! Several steps needed for synthesis of mrna! Separation in space of transcription and translation! Compartmentation
Quantitative Real Time PCR USING SYBR GREEN
 Quantitative Real Time PCR USING SYBR GREEN SYBR Green SYBR Green is a cyanine dye that binds to double stranded DNA. When it is bound to D.S. DNA it has a much greater fluorescence than when bound to
Quantitative Real Time PCR USING SYBR GREEN SYBR Green SYBR Green is a cyanine dye that binds to double stranded DNA. When it is bound to D.S. DNA it has a much greater fluorescence than when bound to
Review of Protein (one or more polypeptide) A polypeptide is a long chain of..
 Gene expression Review of Protein (one or more polypeptide) A polypeptide is a long chain of.. In a protein, the sequence of amino acid determines its which determines the protein s A protein with an enzymatic
Gene expression Review of Protein (one or more polypeptide) A polypeptide is a long chain of.. In a protein, the sequence of amino acid determines its which determines the protein s A protein with an enzymatic
Figure S1. MUT-16 localization in L4 hermaphrodite, adult hermaphrodite, and adult male germlines. MUT-16 DAPI. Phillips et al. S-1. male.
 Supplementary Material for Phillips et al. Figure S1. MUT-16 localization in L4 hermaphrodite, adult hermaphrodite, and adult male germlines. Figure S2. Mutator foci and P granules localize independently
Supplementary Material for Phillips et al. Figure S1. MUT-16 localization in L4 hermaphrodite, adult hermaphrodite, and adult male germlines. Figure S2. Mutator foci and P granules localize independently
SIBC504: TRANSCRIPTION & RNA PROCESSING Assistant Professor Dr. Chatchawan Srisawat
 SIBC504: TRANSCRIPTION & RNA PROCESSING Assistant Professor Dr. Chatchawan Srisawat TRANSCRIPTION: AN OVERVIEW Transcription: the synthesis of a single-stranded RNA from a doublestranded DNA template.
SIBC504: TRANSCRIPTION & RNA PROCESSING Assistant Professor Dr. Chatchawan Srisawat TRANSCRIPTION: AN OVERVIEW Transcription: the synthesis of a single-stranded RNA from a doublestranded DNA template.
TRANSCRIPTION AND PROCESSING OF RNA
 TRANSCRIPTION AND PROCESSING OF RNA 1. The steps of gene expression. 2. General characterization of transcription: steps, components of transcription apparatus. 3. Transcription of eukaryotic structural
TRANSCRIPTION AND PROCESSING OF RNA 1. The steps of gene expression. 2. General characterization of transcription: steps, components of transcription apparatus. 3. Transcription of eukaryotic structural
A 5 P degradation hot spot influences molecular farming of anticancerogenic nuclease TBN1 in tobacco cells
 Plant Cell, Tissue and Organ Culture (PCTOC) A 5 P degradation hot spot influences molecular farming of anticancerogenic nuclease TBN1 in tobacco cells Anna Týcová a,b, Rajen J. J. Piernikarczyk c, Michael
Plant Cell, Tissue and Organ Culture (PCTOC) A 5 P degradation hot spot influences molecular farming of anticancerogenic nuclease TBN1 in tobacco cells Anna Týcová a,b, Rajen J. J. Piernikarczyk c, Michael
sirna transfection optimization with the Agilent 2100 bioanalyzer Application Note A new method for effective gene silencing
 sirna transfection optimization with the Agilent 2100 bioanalyzer A new method for effective gene silencing Application Note Abstract Marc Valer Carsten Buhlmann Ioanna Andreou Martin Weber When working
sirna transfection optimization with the Agilent 2100 bioanalyzer A new method for effective gene silencing Application Note Abstract Marc Valer Carsten Buhlmann Ioanna Andreou Martin Weber When working
over time using live cell microscopy. The time post infection is indicated in the lower left corner.
 Title of file for HTML: Supplementary Information Description: Supplementary Figures and Supplementary Table Title of file for HTML: Supplementary Movie 1 Description: Fusion of NBs. BSR cells were infected
Title of file for HTML: Supplementary Information Description: Supplementary Figures and Supplementary Table Title of file for HTML: Supplementary Movie 1 Description: Fusion of NBs. BSR cells were infected
Supplementary Figure 1. Isolation of GFPHigh cells.
 Supplementary Figure 1. Isolation of GFP High cells. (A) Schematic diagram of cell isolation based on Wnt signaling activity. Colorectal cancer (CRC) cell lines were stably transduced with lentivirus encoding
Supplementary Figure 1. Isolation of GFP High cells. (A) Schematic diagram of cell isolation based on Wnt signaling activity. Colorectal cancer (CRC) cell lines were stably transduced with lentivirus encoding
Make the protein through the genetic dogma process.
 Make the protein through the genetic dogma process. Coding Strand 5 AGCAATCATGGATTGGGTACATTTGTAACTGT 3 Template Strand mrna Protein Complete the table. DNA strand DNA s strand G mrna A C U G T A T Amino
Make the protein through the genetic dogma process. Coding Strand 5 AGCAATCATGGATTGGGTACATTTGTAACTGT 3 Template Strand mrna Protein Complete the table. DNA strand DNA s strand G mrna A C U G T A T Amino
Lecture 25 (11/15/17)
 Lecture 25 (11/15/17) Reading: Ch9; 328-332 Ch25; 990-995, 1005-1012 Problems: Ch9 (study-guide: applying); 1,2 Ch9 (study-guide: facts); 7,8 Ch25 (text); 1-3,5-7,9,10,13-15 Ch25 (study-guide: applying);
Lecture 25 (11/15/17) Reading: Ch9; 328-332 Ch25; 990-995, 1005-1012 Problems: Ch9 (study-guide: applying); 1,2 Ch9 (study-guide: facts); 7,8 Ch25 (text); 1-3,5-7,9,10,13-15 Ch25 (study-guide: applying);
Contents... vii. List of Figures... xii. List of Tables... xiv. Abbreviatons... xv. Summary... xvii. 1. Introduction In vitro evolution...
 vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
BIOLOGY. Chapter 16 GenesExpression
 BIOLOGY Chapter 16 GenesExpression CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson 18 Gene Expression 2014 Pearson Education, Inc. Figure 16.1 Differential Gene Expression results
BIOLOGY Chapter 16 GenesExpression CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson 18 Gene Expression 2014 Pearson Education, Inc. Figure 16.1 Differential Gene Expression results
High-throughput functional genomics using CRISPR Cas9
 Nature Reviews Genetics AOP, published online 9 April 2015; doi:10.1038/nrg3899 REVIEWS High-throughput functional genomics using CRISPR Cas9 Ophir Shalem, Neville E. Sanjana and Feng Zhang Abstract Forward
Nature Reviews Genetics AOP, published online 9 April 2015; doi:10.1038/nrg3899 REVIEWS High-throughput functional genomics using CRISPR Cas9 Ophir Shalem, Neville E. Sanjana and Feng Zhang Abstract Forward
DICER-LIKE2 Plays a Primary Role in Transitive Silencing of Transgenes in Arabidopsis
 DICER-LIKE2 Plays a Primary Role in Transitive Silencing of Transgenes in Arabidopsis Sizolwenkosi Mlotshwa 1, Gail J. Pruss 1, Angela Peragine 2, Matthew W. Endres 1, Junjie Li 3, Xuemei Chen 3, R. Scott
DICER-LIKE2 Plays a Primary Role in Transitive Silencing of Transgenes in Arabidopsis Sizolwenkosi Mlotshwa 1, Gail J. Pruss 1, Angela Peragine 2, Matthew W. Endres 1, Junjie Li 3, Xuemei Chen 3, R. Scott
Division Ave. High School AP Biology
 Control of Eukaryotic Genes 2007-2008 The BIG Questions n How are genes turned on & off in eukaryotes? n How do cells with the same genes differentiate to perform completely different, specialized functions?
Control of Eukaryotic Genes 2007-2008 The BIG Questions n How are genes turned on & off in eukaryotes? n How do cells with the same genes differentiate to perform completely different, specialized functions?
Alternative Cleavage and Polyadenylation of RNA
 Developmental Cell 18 Supplemental Information The Spen Family Protein FPA Controls Alternative Cleavage and Polyadenylation of RNA Csaba Hornyik, Lionel C. Terzi, and Gordon G. Simpson Figure S1, related
Developmental Cell 18 Supplemental Information The Spen Family Protein FPA Controls Alternative Cleavage and Polyadenylation of RNA Csaba Hornyik, Lionel C. Terzi, and Gordon G. Simpson Figure S1, related
Transcription in Eukaryotes
 Transcription in Eukaryotes Biology I Hayder A Giha Transcription Transcription is a DNA-directed synthesis of RNA, which is the first step in gene expression. Gene expression, is transformation of the
Transcription in Eukaryotes Biology I Hayder A Giha Transcription Transcription is a DNA-directed synthesis of RNA, which is the first step in gene expression. Gene expression, is transformation of the
Methods of Biomaterials Testing Lesson 3-5. Biochemical Methods - Molecular Biology -
 Methods of Biomaterials Testing Lesson 3-5 Biochemical Methods - Molecular Biology - Chromosomes in the Cell Nucleus DNA in the Chromosome Deoxyribonucleic Acid (DNA) DNA has double-helix structure The
Methods of Biomaterials Testing Lesson 3-5 Biochemical Methods - Molecular Biology - Chromosomes in the Cell Nucleus DNA in the Chromosome Deoxyribonucleic Acid (DNA) DNA has double-helix structure The
IP-Free Electra DAUGHTER Vectors Mammalian with CMV promoter
 IP-Free Electra DAUGHTER Vectors Mammalian with CMV promoter Electra cloning DNA2.0 has developed a simple one-tube universal cloning process that can be performed in a 5 minute bench-top reaction with
IP-Free Electra DAUGHTER Vectors Mammalian with CMV promoter Electra cloning DNA2.0 has developed a simple one-tube universal cloning process that can be performed in a 5 minute bench-top reaction with
Recombinant DNA Technology. The Role of Recombinant DNA Technology in Biotechnology. yeast. Biotechnology. Recombinant DNA technology.
 PowerPoint Lecture Presentations prepared by Mindy Miller-Kittrell, North Carolina State University C H A P T E R 8 Recombinant DNA Technology The Role of Recombinant DNA Technology in Biotechnology Biotechnology?
PowerPoint Lecture Presentations prepared by Mindy Miller-Kittrell, North Carolina State University C H A P T E R 8 Recombinant DNA Technology The Role of Recombinant DNA Technology in Biotechnology Biotechnology?
Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate
 Supplementary Figure Legends Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate BC041951 in gastric cancer. (A) The flow chart for selected candidate lncrnas in 660 up-regulated
Supplementary Figure Legends Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate BC041951 in gastric cancer. (A) The flow chart for selected candidate lncrnas in 660 up-regulated
RNA interferance as new approaches for pest management
 RNA interferance as new approaches for pest management Nabil Killiny Assistant professor Plant Pathology Citrus Research and Education Center Outlines Introduction Mechanism Applications Conclusion Introduction:
RNA interferance as new approaches for pest management Nabil Killiny Assistant professor Plant Pathology Citrus Research and Education Center Outlines Introduction Mechanism Applications Conclusion Introduction:
Transcription Eukaryotic Cells
 Transcription Eukaryotic Cells Packet #20 1 Introduction Transcription is the process in which genetic information, stored in a strand of DNA (gene), is copied into a strand of RNA. Protein-encoding genes
Transcription Eukaryotic Cells Packet #20 1 Introduction Transcription is the process in which genetic information, stored in a strand of DNA (gene), is copied into a strand of RNA. Protein-encoding genes
The phenomenon of RNAi was first discovered in
 RNA interference Gregory J. Hannon Cold Spring Harbour Laboratory, 1 Bungtown Road, Cold Spring Harbour, New York 11724, USA (e-mail: hannon@cshl.org) A conserved biological response to double-stranded
RNA interference Gregory J. Hannon Cold Spring Harbour Laboratory, 1 Bungtown Road, Cold Spring Harbour, New York 11724, USA (e-mail: hannon@cshl.org) A conserved biological response to double-stranded
Chapter 31. Transcription and RNA processing
 Chapter 31 Transcription and RNA processing RNA polymerase (RNAP) E. coli promoters Components of E. coli RNA Polymerase Holoenzyme (α 2 ββ'ωσ) Structure of prokaryotic RNAP The closed and open state of
Chapter 31 Transcription and RNA processing RNA polymerase (RNAP) E. coli promoters Components of E. coli RNA Polymerase Holoenzyme (α 2 ββ'ωσ) Structure of prokaryotic RNAP The closed and open state of
Description: Nuclear morphology and dynamics in nontargeting sirna transfected cells. HeLa Kyoto
 Title of file for HTML: Supplementary Information Description: Supplementary Figures and Supplementary Tables Title of file for HTML: Supplementary Movie 1 Description: Nuclear morphology and dynamics
Title of file for HTML: Supplementary Information Description: Supplementary Figures and Supplementary Tables Title of file for HTML: Supplementary Movie 1 Description: Nuclear morphology and dynamics
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table.
 Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
DNA makes RNA makes Proteins. The Central Dogma
 DNA makes RNA makes Proteins The Central Dogma TRANSCRIPTION DNA RNA transcript RNA polymerase RNA PROCESSING Exon RNA transcript (pre-mrna) Intron Aminoacyl-tRNA synthetase NUCLEUS CYTOPLASM FORMATION
DNA makes RNA makes Proteins The Central Dogma TRANSCRIPTION DNA RNA transcript RNA polymerase RNA PROCESSING Exon RNA transcript (pre-mrna) Intron Aminoacyl-tRNA synthetase NUCLEUS CYTOPLASM FORMATION
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Dynamic Phosphorylation of HP1 Regulates Mitotic Progression in Human Cells Supplementary Figures Supplementary Figure 1. NDR1 interacts with HP1. (a) Immunoprecipitation using
SUPPLEMENTARY INFORMATION Dynamic Phosphorylation of HP1 Regulates Mitotic Progression in Human Cells Supplementary Figures Supplementary Figure 1. NDR1 interacts with HP1. (a) Immunoprecipitation using
2. Outline the levels of DNA packing in the eukaryotic nucleus below next to the diagram provided.
 AP Biology Reading Packet 6- Molecular Genetics Part 2 Name Chapter 19: Eukaryotic Genomes 1. Define the following terms: a. Euchromatin b. Heterochromatin c. Nucleosome 2. Outline the levels of DNA packing
AP Biology Reading Packet 6- Molecular Genetics Part 2 Name Chapter 19: Eukaryotic Genomes 1. Define the following terms: a. Euchromatin b. Heterochromatin c. Nucleosome 2. Outline the levels of DNA packing
Functional characterisation
 Me Me Ac Ac Functional characterisation How can we know if measured changes in DNA methylation and function (phenotype) and linked, and in what way? DNMT Dianne Ford Professor of Molecular Nutritional
Me Me Ac Ac Functional characterisation How can we know if measured changes in DNA methylation and function (phenotype) and linked, and in what way? DNMT Dianne Ford Professor of Molecular Nutritional
Computational Biology I LSM5191
 Computational Biology I LSM5191 Lecture 5 Notes: Genetic manipulation & Molecular Biology techniques Broad Overview of: Enzymatic tools in Molecular Biology Gel electrophoresis Restriction mapping DNA
Computational Biology I LSM5191 Lecture 5 Notes: Genetic manipulation & Molecular Biology techniques Broad Overview of: Enzymatic tools in Molecular Biology Gel electrophoresis Restriction mapping DNA
Prokaryotic Transcription
 Prokaryotic Transcription Transcription Basics DNA is the genetic material Nucleic acid Capable of self-replication and synthesis of RNA RNA is the middle man Nucleic acid Structure and base sequence are
Prokaryotic Transcription Transcription Basics DNA is the genetic material Nucleic acid Capable of self-replication and synthesis of RNA RNA is the middle man Nucleic acid Structure and base sequence are
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Legends for Supplementary Tables. Supplementary Table 1. An excel file containing primary screen data. Worksheet 1, Normalized quantification data from a duplicated screen: valid
SUPPLEMENTARY INFORMATION Legends for Supplementary Tables. Supplementary Table 1. An excel file containing primary screen data. Worksheet 1, Normalized quantification data from a duplicated screen: valid
RNA-mediated Double-Stranded Break repair in mammalian cells
 RNA-mediated Double-Stranded Break repair in mammalian cells A thesis Presented to The Academic Faculty By Matthew B Taylor In Fulfillment of the requirements for the Honors Thesis Option for a Bachelor
RNA-mediated Double-Stranded Break repair in mammalian cells A thesis Presented to The Academic Faculty By Matthew B Taylor In Fulfillment of the requirements for the Honors Thesis Option for a Bachelor
Supplemental Data. Regulating Gene Expression. through RNA Nuclear Retention
 Supplemental Data Regulating Gene Expression through RNA Nuclear Retention Kannanganattu V. Prasanth, Supriya G. Prasanth, Zhenyu Xuan, Stephen Hearn, Susan M. Freier, C. Frank Bennett, Michael Q. Zhang,
Supplemental Data Regulating Gene Expression through RNA Nuclear Retention Kannanganattu V. Prasanth, Supriya G. Prasanth, Zhenyu Xuan, Stephen Hearn, Susan M. Freier, C. Frank Bennett, Michael Q. Zhang,
Differential Gene Expression
 Biology 4361 - Developmental Biology Differential Gene Expression June 18, 2009 Differential Gene Expression Overview Chromatin structure Gene anatomy RNA processing and protein production Initiating transcription:
Biology 4361 - Developmental Biology Differential Gene Expression June 18, 2009 Differential Gene Expression Overview Chromatin structure Gene anatomy RNA processing and protein production Initiating transcription:
qpcr Quantitative PCR or Real-time PCR Gives a measurement of PCR product at end of each cycle real time
 qpcr qpcr Quantitative PCR or Real-time PCR Gives a measurement of PCR product at end of each cycle real time Differs from endpoint PCR gel on last cycle Used to determines relative amount of template
qpcr qpcr Quantitative PCR or Real-time PCR Gives a measurement of PCR product at end of each cycle real time Differs from endpoint PCR gel on last cycle Used to determines relative amount of template
Supplemental Information. Natural RNA Polymerase Aptamers. Regulate Transcription in E. coli
 Molecular Cell, Volume 67 Supplemental Information Natural RNA Polymerase Aptamers Regulate Transcription in E. coli Nadezda Sedlyarova, Philipp Rescheneder, Andrés Magán, Niko Popitsch, Natascha Rziha,
Molecular Cell, Volume 67 Supplemental Information Natural RNA Polymerase Aptamers Regulate Transcription in E. coli Nadezda Sedlyarova, Philipp Rescheneder, Andrés Magán, Niko Popitsch, Natascha Rziha,
Transport of Potato Lipoxygenase into the Vacuole Larsen, Mia Kruse Guldstrand; Welinder, Karen Gjesing; Jørgensen, Malene
 Aalborg Universitet Transport of Potato Lipoxygenase into the Vacuole Larsen, Mia Kruse Guldstrand; Welinder, Karen Gjesing; Jørgensen, Malene Publication date: 2009 Document Version Publisher's PDF, also
Aalborg Universitet Transport of Potato Lipoxygenase into the Vacuole Larsen, Mia Kruse Guldstrand; Welinder, Karen Gjesing; Jørgensen, Malene Publication date: 2009 Document Version Publisher's PDF, also
